Abstract
Despite the great benefits of plastics in different aspects of life and due to the increase in plastic production and use, plastic wastes are becoming a major environmental concern. It is well known that inappropriate use and disposal lead to the accumulation of plastic litter in different aquatic environments. Microbial biofilm is able to develop on the surface of plastics (plastisphere) in aquatic environments over time. The aim of this study was to describe the bacterial communities associated with plastics in freshwater. Thus, in our first test, a total of six self-designed plastic colonizers were submerged under the surface of the water in Vácszentlászló lake, located in central Hungary, for a period of 3 months. Two plastic colonizers were cultivated monthly. Associated microbial communities were then analyzed as follows: (a) bacterial communities were studied by amplicon sequencing and (b) culturable bacteria were isolated from plastic surfaces and identified by 16S rRNA gene sequencing. Coinciding with these analyses of plastic colonizing communities, surface water samples from the lake were also taken, and in a second test, other materials (eg. wood, glass) associated bacterial communities were also investigated with the same methods. Amplicon sequencing showed notable differences between the plastic and other materials colonizing, and lake waterborne microbial community composition. Using the LB agar, no novel species were found; however, several known pathogenic species were identified. The self-designed plastic colonizer was successfully used during the winter over a 3-month period, suggesting that it could be an appropriate method of choice to study microplastic-associated microbes for longer periods and in variable environmental conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Due to its durability, widespread use, and low cost, plastic production has increased enormously in the last decades (Li et al., 2018), reaching 359 million tons in 2018 (Plasticseurope (2019) ). Despite its notable benefits in our daily life, the unsustainable use and improper waste management have resulted in a global accumulation of plastic debris in the environment. Therefore, its negative impact on the environmental elements is becoming a global concern (Wagner et al., 2014; Li et al., 2018).
Plastics are likely carried to marine environments through rivers, streams, and wastewater treatment plants. Plastics might also find their way to freshwater lakes and accumulate and persist there for decades (Harrison et al., 2018). Lakes which are located near populated areas may contain higher concentrations of microplastics (< 5 mm in size) (Li et al., 2018). Studies have emphasized that the majority of plastic debris collected from freshwater lakes was originated from the households, such as food wrappers, bags, and beverage bottles (Blettler et al., 2017). In freshwater systems these materials may cause loss of biological diversity and their derivates are threatening human health via consumption of fish and drinking water (Blettler et al., 2018; Peng et al., 2017).
Plastics are synthetic polymers mainly derived from the petrol industry. The majority of these artificial materials is resistant to biodegradation, thus in general they mostly undergo mechanical fractionation into smaller fragments (Horton et al., 2017). The plastic fragments of less than 5 mm in diameter have been given the name microplastics (Thompson et al., 2004). Microplastics have been divided into two categories: (a) primary microplastics (microbeads which are synthetic polymers with size 0.1 to 5 mm and used in many products, e.g., personal care cosmetics) and (b) secondary microplastics that results from the degradation of larger plastic particles. Accumulation of microplastics in the oceans has been occurring for decades worldwide even in Antarctica (Oberbeckmann et al., 2014). The estimated number of microplastics floating in the ocean in 2014 was between 15 and 51 trillion particles (Eckert et al., 2018; Peng et al., 2015). One of the most heavily impacted seas by plastic pollution is the Mediterranean Sea, it is estimated that there are currently 1–3 thousand tons of plastic floating on its surface with the majority of plastic fragments that it contains being microplastics (about 250 billion fragments) (Jambeck et al., 2015).
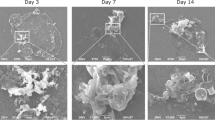
Microplastic threats at a high trophic level have been described in many studies (Nelms et al., 2019). Examining the digestive systems of higher marine mammals has shown the presence of microplastics in the stomach and intestine (Nelms et al., 2019). Ingestion of plastics by seabirds has been reported worldwide; variable seabird species were affected such as petrels, seagulls, and albatross (Franeker & Law, 2015; Cartraud et al., 2019). For seabirds, it may cause internal wounds and obstruction of the stomach; in addition, it may affect reproduction and might increase the death rate (Ryan, 1988; Lavers et al., 2014). Microplastics of different kinds of polymers were detected in 36.5% of examined fish from the English Channel; the abundance of plastic was from one to fifteen fragments per fish (Lusher et al., 2013). Additionally, microplastics can potentially influence the lower trophic level since they can provide a novel surface for microbial biofilm development. It was suggested that the hydrophobicity of microplastic surfaces and the long half-life of plastic potentially enhance biofilm formation (Zettler et al., 2013). Furthermore, the organic polymers, the additives, and the adsorbed contaminants of plastics might offer a carbon source that is essential for microbial metabolism (Zettler et al., 2013; McCormick et al., 2016). Scanning electron microscope images of these biofilms have shown a highly diverse community of bacterial microbiota as well as eukaryotic communities (Bryant et al., 2016; Zettler et al., 2013). This microbial community of heterotrophs, autotrophs, predators, and symbionts has been given the term plastisphere (Zettler et al., 2013). In the aquatic environments, microbes in a biofilm are essential to process the organic matter and to supply energy into the food web. Because of the important role of these biofilms in aquatic environments, it is necessary to understand the potential effect of microplastics on their formation and composition.
Studies have shown that microplastic surfaces are quickly (within hours) colonized by microorganisms (Harrison et al., 2018). However, several factors may affect the development of microplastics associated biofilms such as water temperature, salinity, pressure, light, and oxygen availability (Harrison et al., 2018) in addition to nutrient concentrations (Oberbeckmann et al., 2018). Therefore, biofilm community development on plastic near the water surface differs from that developing on plastics in deep water where the light and oxygen are limited and have higher pressure (Harrison et al., 2018).
Initial studies have revealed that microbial communities associated with plastic litter are also different from those in the surrounding seawater. For example, by DNA sequencing analysis, filamentous cyanobacteria Phormidium and Rivularia were found on plastics but not in the surrounding water in which the unicellular Prochlorococcus was dominated. Moreover, higher community richness was observed in the surrounding seawater, while higher evenness was observed on the plastic surface compared to seawater, suggesting a potential selection of specific microbial taxa by plastic surfaces (Zettler et al., 2013). These differences were also observed with seasonal variations, for instance, in summer the diversity of microbial communities associated with plastic differs from winter (Oberbeckmann et al., 2014).
Concern has been raised by the potential occurrence and transfer of harmful bacterial species on plastic surfaces. Aeromonas salmonicida which is a fish pathogen was isolated from the microplastic surface in the Adriatic sea (Viršek et al., 2017). Toxic eukaryotic species were isolated from plastic surfaces as “hitchhiker organisms” (Debroas et al., 2017) and Vibrio species, which may be responsible for human or animal diseases, were isolated from microplastic surfaces in the North and Baltic seas (Kirstein et al., 2016).
Particular attention should be focused on freshwater environments, as there currently are few published studies of microplastic-associated biofilms and their microbial communities in freshwater systems compared to marine water (Li et al., 2018; McCormick et al., 2016). Different methods have been used to study microbial colonization on microplastic surfaces, most of which were based on incubating microplastic particles in experimental tanks filled with freshwater (Miao et al., 2019; Wu et al., 2019; Wang et al., 2020). In other studies, microplastic particles were collected from sampling sites either by filtering a certain volume of water or by picking microplastic particles from shallow water (Jiang et al., 2018; McCormick et al., 2016); few studies have incubated plastic or microplastic particles in situ to study microbial colonization in the natural environments such as incubating polyethylene (PET) bottles under the water surface for a period of time (Oberbeckmann et al., 2014) or fixing PET bags to Perspex boards and suspended under the water surface for a defined time and then analyzing the associated microbial communities (Lobelle & Cunliffe, 2011).
Our study aims to help in filling the knowledge gap concerning microplastic biofilms and their associated microbial communities in freshwater systems. The structure of the microbial community and the interactions between microbes and microplastic surface are valuable information needed to support the efforts with regards to biodegradation; in this study, we aim to investigate the community composition of microbial biofilms on polypropylene (PP) microplastic surfaces in a freshwater lake in Hungary. To achieve this, we used self-developed plastic colonizers which are simple and easy to be applied in situ and under different environmental conditions. The aim was to try out this method in situ, which can serve as an easily accessible tool to analyze the plastic-associated bacterial community. We aimed also to compare the structure of the PP microplastic-associated microbial community with the surrounding lake water community. Additionally, we aimed to investigate whether the microbial community is microplastic specific or not comparing to other materials such as glass, wood, stainless steel, and biodegradable plastic.
2 Materials and Methods
2.1 Study Area and Sampling Location
The study was conducted in Vácszentlászló lake, a freshwater lake located near to the village of Vácszentlászló, Pest county in Hungary (47°33′37.0″N 19°33′09.4″E) approximately 20 km from Szent Istvan Campus of Hungarian University of Agricultural and Life Sciences where samples were analyzed. The lake is a shallow reservoir (mean depth: 2 m; area: 47 ha), the water fluctuation level is normally up to 1 m, the water in the lake is eutrophic (for water quality of the sampling lake, see Supplementary Table. 1).
2.2 Plastic Colonizers Design
Self-designed plastic colonizers were prepared as follows: commercially available stainless-steel ball-shaped tea filters (Easy Filter 6.5 × 6cm. No 1082, Mingwei, China) were wrapped in aluminum foil and pre-sterilized in an autoclave at 121 °C for 15 min. Filters were then filled with 3 g (for each) of polypropylene plastic straws cut into less than 5-mm small pieces (see Fig. 1). Six of these plastic colonizers were hung on a wooden pier next to each other with a strong fishing line and submerged under the water surface (around 50-cm depth). Their positions were fixed with lead weights to keep the colonizers under the water’s surface (see Fig. 2). The first study was conducted over a period of three months from December 2018 until the end of February 2019. By the end of the first month, two plastic colonizers were collected. They were transferred to the laboratory directly within 1 hour in a pre-sterilized container filled with ambient lake water. This process was repeated at the end of the second and third months.
2.3 Recovery of Bacterial Biofilm from Plastic Colonizers
In the laboratory, two plastic colonizers were opened (treated as one sample), and plastic particles were transferred to a pre-sterilized stainless-steel mesh using a sterilized spatula, then washed down with sterilized normal saline to remove the stuck debris but keep the plastic associated biofilm. Then, the water-washed plastics were transferred to an Erlenmeyer flask containing sterilized 90 ml distilled water, 30g of glass beads, and 13.5 μl of TWEEN 80. It was incubated at room temperature on a shaker with 170 rpm speed for 1 h to recover the microbial biofilm attached to plastic surfaces.
2.4 DNA Isolation of Plastic-Associated and Lake Water Bacterial Communities
After inoculating aliquots of the microplastic suspension onto LB agar (see below), the remaining suspension (~ 90 ml) was centrifuged at 4 °C and 4000 rpm speed for 25 min, supernatant was discarded, and the pellet was stored at – 80 °C for subsequent DNA isolation. For isolation of community DNA from lake water, 150 ml were centrifuged under the same conditions to have a sufficient pellet. Community DNA was then isolated by using the DNeasy UltraClean Microbial Kit (QIAGEN, Venlo, The Netherlands) according to the manufacturer’s instructions.
2.5 Bacterial Community Assessment
To assess the composition of bacterial communities in plastic colonizers (first month; VMP1, second month; VMP2, and third month; VMP 3) and in lake water (first month; VLW1, second month; VLW 2 and month; VLW 3) Illumina 16S rDNA amplicon sequencing was used (SeqOmics Biotechnology lab. Morahalom, Hungary). For paired-end 16S rDNA amplicon sequencing, the variable V3 and V4 regions of the 16S rRNA gene were amplified by using 16S amplicon PCR forward (5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTA CGGGNGGCWGCAG-3′) and reverse (5′-GTCT CGTGGGCT CGGAGATGTGTATAAGAGAC AGGACTACHVGGGTATCTAATCC-3′) primers, with Illumina adapter overhanging nucleotide sequences (Klindworth et al., 2013). PCR reaction mixture volume was 25 μl with 12.5 ng of DNA, 0.2 μM of each Illumina 16S primers, and 12.5 μl of 2X KAPA HiFi HotStart Ready Mix (KAPABiosystems, London, UK). The temperature profile was as follows: initial denaturation (3 min at 25 °C), 25 denaturation cycles for 0.5 min at 95 °C, the annealing temperature was 25 °C for 0.5 min, and 0.5-min elongation at 72 °C. ProFlex PCR System (Applied Biosystems by Life Technologies, USA) was used for all amplification steps. Analysis of amplicon was performed under UV after electrophoresis in 1% (w/v) agarose gel stained with EtBr. Paired-end fragment reads were generated on an Illumina MiSeq sequencer using MiSeq Reagent Kit v3 (600 cycles). Primary data analysis (base calling) was carried out with Bcl2fastq software (v2.17.1.14, Illumina). Sequences were processed using mothur v1.41.1 (Schloss et al., 2009) as recommended by the MiSeq SOP page (http://www.mothur.org/wiki/MiSeq_SOP) (Kozich et al., 2013). Sequence lengths were screened by setting minimum length to 400 base pairs, then were assorted based on the alignment using SILVA 132 SSURef NR99 database (Quast et al., 2013). Chimera detection was performed with mothur’s uchime command (Edgar et al., 2011), and “split.abund” command was also used to remove singleton reads according to Kunin et al. (2010). Taxonomic assignments were made against SILVA release 132 applying a minimum bootstrap confidence score of 80%. Operational taxonomic units (OTUs) were assigned at a 97% similarity threshold level for prokaryotic species delineation (Tindall et al., 2010).
2.6 Isolation of Culturable Bacteria on LB Agar from Plastic
From a 1-hour shaking suspension of plastic particles, the initial 1 ml was serially diluted up to 10–6 using 9 ml of normal saline. One milliliter of each dilution was plated on LB agar and incubated at 28 °C for 3 days. From every sample, a maximum of eight colonies with different morphologies were purified on LB agar for further identification. LB agar was used to isolate fast-growing cultivable species from the plastic surface to get information about their pathogenicity or find novel ones.
2.7 16S rRNA Gene Sequencing for Identification of Bacterial Isolates
Genomic DNA was isolated by DNeasy UltraClean Microbial Kit according to the manufacturer instructions. Subsequently, the 16S rRNA gene was amplified using primers 8F (5′-AGAGTTTGATCCTGCCTCAG-3′) and 1492R (5′-GGTTACCTTGTTACGACTT-3′) (Turner et al., 1999). Amplification was performed by using an Eppendorf Mastercycler (Eppendorf, Hamburg, Germany). PCR products were cleaned using NucleoSpin® Gel and PCR Clean-up (Machery-Nagel GmbH & Co., Düren, Germany). The complete 16S rRNA gene sequence was determined by using a BigDye Terminator v3.1 Cycle Sequencing kit (Applied Biosystems, Foster City, USA) according to the manufacturer’s instructions. Sequencing products were separated on a model 3130 Genetic Analyzer (Applied Biosystems, Foster City, USA).
2.8 Statistical Analysis
Testing the difference between microbial communities on PP microplastic surfaces (group1) compared to surrounding water (group2) is of high importance to measure the significance level of difference. A t-test is used to compare between the means of two data sets, assuming that the data sets are normally distributed, in large samples (> 30 or 40), the sampling distribution tends to be normal, regardless of the shape of the data (Ghasemi & Zahediasl, 2012), and the two groups of data are also assumed to be independent of one another (Supplementary Table 2).
Thus, a paired samples t-test using SPSS software was used to answer the research question and determine whether there are statistically significant differences between the two data sets:
-
H0: There is no (statistically) significant difference between microbial communities on PP microplastic surfaces compared to the surrounding water.
-
H1: There is a (statistically) significant difference between microbial communities on PP microplastic surfaces compared to the surrounding water.
2.9 Bacterial Communities Associated with Different Materials (Second Test)
In order to assess if the bacterial community structures are plastic specific, some months after the first test, four colonizers were filled with 3 g of polypropylene plastic, biodegradable PLA (polylactic acid straw) plastic, wood, and glass, in addition to one empty (stainless steel). All were installed as mentioned above, for 2-month period (between September and November, in 2019). After that colonizers plus water sample were collected, genomic DNA was isolated, and amplicon sequencing was conducted as above.
3 Results
3.1 Amplicon Sequencing Results: PP Microplastic vs Lake Water
The Illumina 16S rRNA gene amplicon sequencing provided 29718, 28837, 28453, 30742, 29366, and 29386 reads for VMP1, VMP2, VMP3, VLW1, VLW2, and VLW3 respectively.
The rarefaction curves of the samples indicated that the data contained enough sequence depth to ascertain the full bacterial diversity. High sequencing coverage was reached in all samples (see Supplementary Figure 1).
Diverse bacterial communities were found on PP microplastic surfaces and in lake water as well. The amount of different OTUs observed on PP microplastic surfaces after the first, second, and third months were 293, 394, and 345 respectively. Over the same time period, the amount of different OTUs observed in lake water was 134, 352, and 348 (see Supplementary Figures 2, 3, 4, 5, and 6). In the first month (Dec 2018), Cyanobacteria and Proteobacteria dominated the PP microplastic community (VMP1) with the relative abundance of 69.3% and 16.6% respectively. In the second month (January 2019, sample ID: VMP2), Proteobacteria became the most abundant (34.0%), followed by Cyanobacteria (31.3%) and Bacteroidetes (20.6%). In the third month (February 2019, sample ID: VMP3), Bacteroidetes relative abundance increased to 54.2% and Proteobacteria increased to 32.9%, whereas Cyanobacteria notably decreased to 1.2%.
In the lake water, Cyanobacteria were highly dominated in the microbial community of the first month (December 2018, sample ID: VLW1) with a relative abundance of (90.6%). In the second month (January 2019, sample ID: VLW2), Cyanobacteria were also the most dominant but the relative abundance has decreased to 70.0%; Proteobacteria (9.5%) became the second most dominant phylum. After the third month (February 2019, sample ID: VLW3), Cyanobacteria relative abundance decreased again to 53.0%, Proteobacteria relative abundance increased to 19.2%, and Bacteroidetes became the third most dominant phylum with the relative abundance of 16.0%. Supplementary Figure 7 compares the most abundant microbial phyla between PP microplastic surface samples and lake water samples.
At the order level Nostocales which belongs to phylum Cyanobacteria was the most dominant in PP microplastic samples in the first and second months, but notably decreased to 1.21% in the third month in which Flavobacteriales was the dominant order followed by Betaproteobacteriales. All lake water samples were dominated by Nostocales but with a notable decrease in relative abundance with time (see Fig. 3). On the family and genus levels, similar trends can be seen with regards to members of the Phormidiaceae family and genus Planktothrix which also belongs to Cyanobacteria as well as the Flavobacteriaceae family and genus Flavobacterium (Supplementary Figure 8).
Dominant taxonomic orders associated with microplastics and surrounding water based on relative abundance more than 1%; VMP1: microplastic from plastic colonizer, first month; VMP2: microplastic second month; VMP3: microplastic third month; VLW1: surrounding water first month; VLW2: surrounding water second month; VLW3: surrounding water third month
In order to visualize differences between bacterial community’s structures, a heatmap analysis was performed at the genus level (Fig. 4). For some genera, the tendencies of abundance changing were likely the same during the 3-month period in both PP microplastic and surrounding lake water communities, e.g., Albidiferax and Algoriphagus. But in most cases, distinct differences in dominant genera between PP microplastic surface samples compared to the surrounding lake water were observed. For example, the Planktothrix genus, which belongs to Cyanobacteria, is frequently present in all lake water samples across all months, whereas in the PP microplastic samples it decreased in abundance from the first month to the third month. With regard to Flavobacterium, it was less frequent in the first month in both PP microplastic and lake water associated samples, but became more abundant in the second and third months on PP microplastics compared to lake water samples (for the community structures associated with PP microplastic and water at different taxonomic levels, see Supplementary Table 3).
Heatmap of microbial community structure on the genus level with the relative abundance of more than 1%; VMP1: microplastic first month; VMP2: microplastic second month; VMP3: microplastic third month; VLW1: surrounding water first month; VLW2: surrounding water second month; VLW3: surrounding water third month. The color intensity in each panel shows the percentage in a sample, color key is at the right side
Cluster analysis of bacterial communities based on Bray-Curtis similarity demonstrated that the dissimilarities between PP microplastic surfaces (VMP samples) and the surrounding water (VLW samples) increased with time. The bacterial community of the first month (VMP1) on PP microplastic surfaces belongs to a close cluster with the samples of the surrounding water (> 60%). From the second month, the PP microplastic-associated bacterial community became more different compared to surrounding water (around 50% similarity). The highest differences between the microplastic surface and the surrounding water were observed in the third month where the similarity decreased to less than 30% (Fig. 5).
Clustering analysis of bacterial communities in the six samples (three microplastic and water) based on OTU abundance-based Bray-Curtis similarity coefficients; VMP1: microplastic-associated community from plastic colonizers, first month; VMP2: microplastic second month; VMP3: microplastic third month; VLW1: surrounding lake water first month; VLW2: surrounding water second month; VLW3: surrounding water third month
Based on the statistical analysis above, it was found then that the difference in microbial community compositions of PP microplastics compared with surrounding is highly significant when analyzing over the whole study period (P-value = 0.000; F = 49.247; df = 3970), so the H0 hypothesis is rejected.
Again, our hypothesis was tested among the months, below are the results for testing the differences between the PP microplastic-associated bacterial communities (group 1) and surrounding water (group 2) on monthly basis (December, January, February). The results show a highly significance among December (P-value = 0.000), January (P-value = 0.010), and February (P-value = 0.023) (see Table 1).
3.2 Amplicon Sequencing Results of Second Test: PP Microplastic vs Different Materials
At the phylum level, Proteobacteria, Bacteroidetes, Firmicutes, and Verrucomicrobia were the most dominant (> 5%) in plastic associated community respectively; dominant phyla in the degradable plastic community were Proteobacteria, Bacteroidetes, Verrucomicrobia, Cyanobacteria, Firmicutes, and Actinobacteria respectively, whereas in the glass-associated community the following phyla have dominated: Proteobacteria, Bacteroidetes, Verrucomicrobia, Actinobacteria, and Cyanobacteria respectively. The dominant phyla associated with the empty colonizer (metal) were Proteobacteria, Bacteroidetes, Verrucomicrobia, Cyanobacteria, and Actinobacteria respectively.
For the natural materials like wood and water, the most dominant bacterial phyla were Proteobacteria, Firmicutes, Bacteroidetes, and Verrucomicrobia, whereas the following phyla were dominant in the water-associated community: Cyanobacteria, Firmicutes, and Proteobacteria.
On the order level, orders Bacteroidales, Verrucomicrobiales, Clostridiales, Rhodobacterales, Xanthomonadales, and Desulfovibrionales were the most abundant (> 5%) in plastic associated communities; in degradable (PLA) plastic communities, orders Verrucomicrobiales, Bacteroidales, Nostocales, Rhodobacterales, Clostridiales, Xanthomonadales, Desulfobacterales were the most dominant. Furthermore, orders Rhodobacterales, Verrucomicrobiales, Xanthomonadales, Nostocales, Bacteroidales, Desulfobacterales, and Clostridiales were mostly dominated in glass-associated community.
The wood-associated bacterial community was dominated by orders Clostridiales, Bacteroidales, Pseudomonadales, Verrucomicrobiales, Desulfovibrionales, and Erysipelotrichales, whereas water-associated bacterial communities were dominated by the following taxonomic orders Nostocales, Bacillales, Planctomycetales, and Rhizobiales (for the whole taxonomic order structures, see Fig. 6).
3.3 Bacterial Isolates Using LB Agar
Variable bacterial species were isolated during the first test from PP microplastic surfaces on LB agar. A full list of isolated bacterial species that were identified based on 16S rRNA gene sequencing (> 99% similarity) are mentioned in Table 2 taking into consideration that only eight strains were isolated from each plastic sample; some species were found more than once.
4 Discussion
4.1 Microbial Communities of PP Microplastic Compared to Surrounding Water Community
In line with previous studies (Zettler et al., 2013; Miao et al., 2019; Oberbeckmann et al., 2014), microbial community structures of PP microplastic were different from that in surrounding water; significantly different abundances of phyla Cyanobacteria, Proteobacteria, and Bacteroidetes were observed in PP microplastic communities comparing to the water communities; these differences became more obvious over time. The community structures were also changing over the period of 3 months; the dominance of microbial phyla was changing in terms of relative abundance over the 3-month period. For example, Cyanobacteria was the most dominant phylum in the lake water samples over the studied time, but with decreasing relative abundance over the first, second, and third month respectively (91%, 70%, 53%). On PP microplastic surfaces, the relative abundance of Cyanobacteria decreased from 69 to 31%, and 1% over the 3 months, but Bacteroidetes became more and more dominant (2%, 21%, 54% respectively). This result is consistent with Jiang et al. (2018) where they also found that Cyanobacteria and Proteobacteria are suggested to be early microplastic colonizers in freshwater, followed by Bacteroidetes as secondary microplastic colonizers (Jiang et al., 2018), which is also in line with Lee et al. (2008) regarding the early and secondary colonizers in marine water (Lee et al., 2008). Furthermore, the occurrence of Cyanobacteria, which is an oxygen producer, is probably due to its importance for polymer oxidation.
Planktothrix species dominated the investigated samples. The high abundance of bloom-forming Planktothrix agardhii and P. rubescens are regularly observed in European waters (Nõges & Ott, 2003; Willame et al., 2005; Churro et al., 2017). While they cause strong red-colored water-bloom, these species can produce microcystins (MCs), which inhibit eukaryotic protein phosphatases. Bioactive peptides are also produced by this species, which presumably enhance the colonization potential and possible dominance in habitats (Kurmayer et al., 2016). Cyanotoxins as microcystins can cause human and animal poisoning. MCs might be responsible for tumor promotion (Bogialli et al., 2013). This finding supports the hypothesis that microplastics may serve as a vector for pathogenic bacteria (Zettler et al., 2013).
During the 3 months, the Bacteroidetes became more and more dominant on PP microplastic surface samples. The most abundant Flavobacteriaceae family was also observed in the second and third months of lake water but was absent in the first month. Flavobacteriaceae composed around one-third of the identified plastic-associated community (Oberbeckmann et al., 2016). Flavobacteria were described as a major colonizer of diatom detritus (Abell & Bowman, 2005). Therefore, the increased relative abundance of Flavobacteriaceae on PP microplastic surfaces is probably because of the noted algae colonizing of the surface of the plastic particles inside the colonizers; however, we cannot prove this hypothesis since we have no data about eukaryotic members of the community. On PP microplastic surfaces, Flavobacterium lacus was the dominant species in the third month followed by and Flavobacterium chungnamense; however, other members of Flavobacterium were also present. Members of this genus are chemo-organotroph aerobes; can be found in various aquatic habitats such as freshwater, wastewater, and seawater; and can adapt to very cold environments like Antarctic lakes. Flavobacterium was isolated from the surface of low-density polyethylene microplastic and considered a serious fish pathogen (Li et al., 2014; Gong et al., 2019). Flavobacterium is known to degrade complex polymers such as Pentachlorophenol PCP (Saber & Crawford, 1985), diesel (Chaudhary et al., 2019), and nitriles (Egelkamp et al., 2017). Nylon oligomer has been degraded by Flavobacterium sp. KI 725 when provided as the sole source of carbon and nitrogen (Negoro, 2000); therefore, the increased abundance of Flavobacteria on microplastic surfaces with time might be due to a potential role in biodegradation.
Lacihabitans were also present on microplastic surfaces; this genus by the time of writing contains only one valid species Lacihabitans soyangensis; it belongs to the Cytophagaceae family, which is widely distributed in the environment like freshwater, marine water, and soil. Members of this family are known to have the capability to degrade several organic compounds such as starch, chitin, and cellulose (Joung et al., 2014). Bacterial cellulose (BC) is usually abundant in biofilm due to its role in intra- and inter-domain interactions; BC is known to be produced by Proteobacteria which was abundant on microplastic surfaces in this study; therefore, this might be the reason for the occurrence of Lacihabitans spp. on microplastics.
Algoriphagus marisflavi was first isolated and identified in estuarian water in the Yellow Sea (Korea); it can grow at low temperatures 4 °C and at 0–2.0 % (w/v) NaCl (Park et al., 2017). Members of the genus Algoriphagus were isolated from different habitats: seawater, freshwater, marine sediments, and algae (Nedashkovskaya et al., 2004; Liu et al., 2009). We suggest that its presence in the plastisphere in our study might also be because of the notable abundance of algae on the microplastic surfaces. Burkholderiaceae was identified on PP microplastic surfaces over the 3-month period as the second most dominant with the relative abundance of 7.9%, 18.8%, and 15.3% respectively. Members of Burkholderiaceae were also identified in bacterial communities associated with plastic surfaces in different environments such as wastewater treatment systems (Pal et al., 2012) and drinking water facilities (Kalmbach et al., 2000). Therefore, the occurrence of these family members might be connected to the plastic surface rather than the availability of nutrients.
Members of genus Gemmobacter have been isolated from diverse environments; freshwater spring, snow samples, birds, planktonic seaweeds, and marine environments (Chen et al., 2013; Liu et al., 2014; Yoo et al., 2019); Gemmobacter belongs to the family Rhodobacteraceae which was frequently identified on plastics (Zettler et al., 2013; Bryant et al., 2016) and was described as a primary biofilm colonizer in seawater (Elifantz et al., 2013). Additionally, Rhodobacteraceae was suggested as one of the candidate bacterial families for plastic degradation (Line & Eva, 2019).
The PVC group member uncultured Planctomycetales bacteria’s (closest relative Tundrisphaera lichenicola 89.7% 16S rRNA similarity) abundance showed an increasing trend within the plastic colonizer community. Planctomycetes usually prefer a surface-attached lifestyle. DeLong et al. (1993) found a greater abundance on the surface of marine aggregates than in the bacterioplankton (DeLong et al., 1993). Analyzing four lakes in Germany, Allgaier et al. (2006) found a complete lack of planctomycetes among the free-living microorganisms, while the group appeared abundant among the surface colonizers (Allgaier et al., 2006).
The distribution of another PVC group member uncultured Chthoniobacterales bacteria (closest relative Terrimicrobium sacchariphilum 91.42% 16S rRNA similarity) was not so specific since it was present in all samples.
Members of the Mycobacterium genus were also identified on PP microplastic surfaces in our study. This genus contains many pathogenic species associated with pulmonary diseases; additionally, Mycobacterium has been reported as a potential degrader of polyethylene (Sudhakar et al., 2008; Da Costa et al., 2015).
Based on the former publications about the identified species, it can be verified that potential fish and human pathogenic strains can be isolated from microplastic surfaces. These results are in line with studies that mention the potential occurrence of pathogens on microplastic surfaces (Keswani et al., 2016; Zettler et al., 2013; Wu et al., 2019).
Aeromonas bestiarum species was identified as a fish pathogen in carp and trout (Vet et al., 2010). Aeromonas sobria strains have also been described as a pathogen of silver carp as well as a rare human pathogen (Dar et al., 2016). Plastic colonizing Shewanella putrefaciens is a potential opportunistic human pathogen; strains of this species were reported as a causative agent of osteitis, erysipelas, abscess, and rare cases bacteremia and soft tissue infections (Hochedez et al., 2013). Strains of Bacillus simplex were reported as a suspected cause of human brain abscess (Pesce et al., 2016).
The majority of methods used to study the microbial colonization of microplastics in water mainly focused on the collection of microplastic particles through a mesh or net or the collection of plastic litter (Zettler et al., 2013; Frère et al., 2018; McCormick et al., 2016; Viršek et al., 2017; Jiang et al., 2018). Other methods were used in which plastic particles were incubated in situ but mostly for a short incubation periods (Oberbeckmann et al., 2018; Dudek et al., 2020; Oberbeckmann et al., 2014); other studies incubate the plastic particles in a controlled environment in vitro (Dussud et al., 2018; Kirstein et al., 2019). In contrast, the method of plastic colonizers used in this study can be used in situ in different environments like freshwater lakes and rivers. Periodic changes in microbial communities and the possible alteration of microplastic surfaces might also be monitored since it is expected to stand for changeable environmental conditions; thus, it can be used to monitor the changes over the seasons. It might also be used in marine environments to study microbial communities that associate with microplastics. Future studies based on this same easy-to-use colonizing method can be compared with each other. Furthermore, based on our results, the limitations of the plastic colonizers have to be tested in saline water and for long-term periods (minimum one year) to verify the stability of the complex structure.
4.2 Microbial Communities of PP Microplastic Compared to Different Materials
The amplicon sequencing results of the different materials have shown that bacterial communities associated with different materials (plastic, degradable plastic, glass, metal, and wood) are clearly different from the surrounding water on both phylum and order level; in comparison, the differences in bacterial communities associated with polypropylene plastic, degradable plastic, and glass were smaller; mainly, these differences were in the relative abundance of different taxa. As shown in Fig. 6, clustering shows that community structures of polypropylene plastic and degradable poly lactic acid plastic were the closest to each other, followed by glass, wood, water, and finally metal surface.
5 Conclusions
Our results have verified that the structure of PP microplastic-associated bacterial communities is different from that surrounding natural freshwater, especially in terms of relative abundance, and these differences increase over time. When comparing plastic with other materials, similarities in the structure of bacterial communities were observed especially with degradable plastic and glass, fewer similarities were observed with wood and metal-associated communities. The self-developed plastic colonizer method used in this study might be appropriate and reproducible under different conditions and can be used with different plastic materials which are recommended for further research. It is also important for future studies to follow the colonizing order of plastic-associated bacteria in a long-term monitoring project. Our results support the idea that human pathogenic bacteria as well as fish pathogens can be transferred as “hitchhikers” on microplastic surfaces.
Data Availability
All data generated or analyzed during this study are included in this article (and its supplementary information files). No additional data is associated with this study.
References
Abell, G. C. J., & Bowman, J. P. (2005). Colonization and community dynamics of class flavobacteria on diatom detritus in experimental mesocosms based on Southern Ocean Seawater Q. FEMS Microbiology Ecology, 53, 379–391. https://doi.org/10.1016/j.femsec.2005.01.008
Allgaier, M., Allgaier, M., Grossart, H.-p., & Grossart, H.-p. (2006). Diversity and seasonal dynamics of Actinobacteria populations in four lakes in Northeastern Germany. Microbiology, 72(5), 3489–3497. https://doi.org/10.1128/AEM.72.5.3489
Blettler, M. C. M., Ulla, M. A., Rabuffetti, A. P., & Garello, N. (2017). Plastic pollution in freshwater ecosystems: Macro-, meso-, and microplastic debris in a floodplain lake. Environmental Monitoring and Assessment, 189(11). https://doi.org/10.1007/s10661-017-6305-8
Blettler, M. C. M., Abrial, E., Khan, F. R., Sivri, N., & Espinola, L. A. (2018). Freshwater plastic pollution : Recognizing research biases and identifying knowledge gaps. Water Research, 143, 416–424. https://doi.org/10.1016/j.watres.2018.06.015
Bogialli, S., Di Gregorio, F. N., Lucentini, L., Ferretti, E., Ottaviani, M., Ungaro, N., Abis, P. P., & De Grazia, M. C. (2013). Management of a toxic cyanobacterium bloom (Planktothrix rubescens) affecting an Italian drinking water basin: A case study. Environmental Science and Technology, 47(1), 574–583. https://doi.org/10.1021/es302260p
Bryant, J. A., Clemente, T. M., Viviani, D. A., Fong, A. A., Thomas, K. A., Kemp, P., Karl, D. M., White, A. E., & DeLong, E. F. (2016). Diversity and activity of communities inhabiting plastic debris in the North Pacific Gyre. MSystems, 1(3), e00024–e00016. https://doi.org/10.1128/mSystems.00024-16
Cartraud, A. E., Le Corre, M., Turquet, J., & Tourmetz, J. (2019). Plastic ingestion in seabirds of the Western Indian Ocean. Marine Pollution Bulletin, 140(February), 308–314. https://doi.org/10.1016/j.marpolbul.2019.01.065
Chaudhary, D. K., Kim, D. U., Kim, D., & Kim, J. (2019). Flavobacterium petrolei sp. nov., a novel psychrophilic, diesel-degrading bacterium isolated from oil-contaminated Arctic soil. Scientific Reports, 9(1), 1–9. https://doi.org/10.1038/s41598-019-40667-7
Chen, W.M., Cho, N., Huang, W., Young, C., and Sheu, S.Y.. (2013). “Description of Gemmobacter fontiphilus sp. Nov., isolated from a freshwater spring, reclassification of Catellibacterium nectariphilum as Gemmobacter nectariphilus comb. nov., Catellibacterium changlense as Gemmobacter changlensis comb. nov., Catellibacte.” International Journal of Systematic and Evolutionary Microbiology 63 (Pt_2): 470–78. doi: 10.1099/ijs.0.042051-0.
Churro, C., Azevedo, J., Vasconcelos, V., & Silva, A. (2017). Detection of a Planktothrix agardhii bloom in Portuguese marine coastalwaters. Toxins, 9(12), 1–13. https://doi.org/10.3390/toxins9120391
Dar, G. H., Dar, S. A., Kamili, A. N., Chishti, M. Z., & Ahmad, F. (2016). Microbial pathogenesis detection and characterization of potentially pathogenic Aeromonas sobria isolated from Fi Sh Hypophthalmichthys molitrix (Cypriniformes: Cyprinidae ). Microbial Pathogenesis, 91, 136–140. https://doi.org/10.1016/j.micpath.2015.10.017
DeLong, E., Franks, D., & Alldredge, A. (1993). Phylogenetic diversity of aggregate-attached vs. free-living marine bacterial assemblages. Limnology and Oceanography, 38(5), 924–934. https://doi.org/10.2307/2838082
Debroas, D., Mone, A., & Ter Halle, A. (2017). Plastics in the North Atlantic garbage patch: A boat-microbe for hitchhikers and plastic degraders. Science of the Total Environment, 599–600, 1222–1232. https://doi.org/10.1016/j.scitotenv.2017.05.059
Dudek, K. L., Cruz, B. N., Polidoro, B., & Neuer, S. (2020). Microbial colonization of microplastics in the Caribbean Sea. Limnology and Oceanography Letters, 5(1), 5–17. https://doi.org/10.1002/lol2.10141
Dussud, C., Hudec, C., George, M., Fabre, P., Higgs, P., Bruzaud, S., Delort, A. M., et al. (2018). Colonization of non-biodegradable and biodegradable plastics by marine microorganisms. Frontiers in Microbiology, 9, 1–13. https://doi.org/10.3389/fmicb.2018.01571
Eckert, E. M., Di Cesare, A., Kettner, M. T., Arias-Andres, M., Fontaneto, D., Grossart, H. P., & Corno, G. (2018). Microplastics increase impact of treated wastewater on freshwater microbial community. Environmental Pollution, 234, 495–502. https://doi.org/10.1016/j.envpol.2017.11.070
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C., & Knight, R. (2011). UCHIME improves sensitivity and speed of Chimera detection. Bioinformatics, 27(16), 2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Egelkamp, R., Schneider, D., Hertel, R., & Daniel, R. (2017). Nitrile-degrading bacteria isolated from compost. Frontiers in Environmental Science, 5. https://doi.org/10.3389/fenvs.2017.00056
Elifantz, H., Horn, G., Ayon, M., Cohen, Y., & Minz, D. (2013). Rhodobacteraceae are the key members of the microbial community of the initial biofilm formed in Eastern Mediterranean Coastal Seawater. FEMS Microbiology Ecology, 85(2), 348–357. https://doi.org/10.1111/1574-6941.12122
Franeker, Jan A. Van, and Kara Lavender Law. (2015). Seabirds, gyres and global trends in plastic pollution. Environmental Pollution, 203, 89–96. https://doi.org/10.1016/j.envpol.2015.02.034
Frère, L., Maignien, L., Chalopin, M., Huvet, A., Rinnert, E., Morrison, H., Kerninon, S., et al. (2018). Microplastic bacterial communities in the Bay of Brest: Influence of polymer type and size. Environmental Pollution, 242, 614–625. https://doi.org/10.1016/j.envpol.2018.07.023
Da Costa, F., Ana, R., Fedrizzi, T., Lopes, M. L., Pecorari, M., Oliveira, W. L., Costa, D., Giacobazzi, E., Da Costa, J. R., & Bahia, et al. (2015). Characterization of 17 strains belonging to the Mycobacterium simiae complex and description of Mycobacterium paraense sp. nov. International Journal of Systematic and Evolutionary Microbiology, 65(2), 656–662. https://doi.org/10.1099/ijs.0.068395-0
Ghasemi, A., & Zahediasl, S. (2012). Normality tests for statistical analysis: A guide for non-statisticians. International Journal of Endocrinology and Metabolism, 10(2), 486–489. https://doi.org/10.5812/ijem.3505
Gong, M., Yang, G., Zhuang, L., & Zeng, E. Y. (2019). Microbial biofilm formation and community structure on low-density polyethylene microparticles in lake water microcosms. Environmental Pollution, 252, 94–102. https://doi.org/10.1016/j.envpol.2019.05.090
Harrison, J. P., Hoellein, T. J., Sapp, M., Tagg, A. S., Ju-Nam, Y., & Ojeda, J. J. (2018). Microplastic-associated biofilms: A comparison of freshwater and marine environments. In M. Wagner & S. Lambert (Eds.), Freshwater Microplastics : Emerging Environmental Contaminants? (pp. 181–201). Springer International Publishing. https://doi.org/10.1007/978-3-319-61615-5_9
Hochedez, P., Vignier, N., Barreau, M., Olive, C., Baubion, E., and The, R.. (2013). “Human infection with Shewanella putrefaciens and S. algae: Report of 16 cases in Martinique and review of the literature” 89 (1): 151–56. doi: https://doi.org/10.4269/ajtmh.13-0055.
Horton, A. A., Walton, A., Spurgeon, D. J., Lahive, E., & Svendsen, C. (2017). Microplastics in freshwater and terrestrial environments: Evaluating the current understanding to identify the knowledge gaps and future research priorities. Science of the Total Environment, 586(February), 127–141. https://doi.org/10.1016/j.scitotenv.2017.01.190
Jambeck, J. R., Geyer, R., Wilcox, C., Siegler, T. R., Perryman, M., Andrady, A., Narayan, R., & Law, K. L. (2015). Plastic waste inputs from land into the ocean. Science. Vol., 347. https://doi.org/10.1126/science.1260352
Jiang, P., Zhao, S., Zhu, L., & Li, D. (2018). Microplastic-associated bacterial assemblages in the intertidal zone of the Yangtze Estuary. Science of the Total Environment, 624, 48–54. https://doi.org/10.1016/j.scitotenv.2017.12.105
Joung, Y., Kim, H., Kang, H., Lee, B. I., Ahn, T. S., & Joh, K. (2014). Lacihabitans soyangensis gen. nov., sp. nov., a new member of the family Cytophagaceae, isolated from a freshwater reservoir. International Journal of Systematic and Evolutionary Microbiology, 64, 3188–3194. https://doi.org/10.1099/ijs.0.058511-0
Kalmbach, S., Szewzyk, U., Manz, W., & Bendinger, B. (2000). In situ probing reveals Aquabacterium commune as a widespread and highly abundant bacterial species in drinking water biofilms. Water Research, 34(2), 575–581. https://doi.org/10.1016/S0043-1354(99)00179-7
Keswani, A., Oliver, D. M., Gutierrez, T., & Quilliam, R. S. (2016). Microbial hitchhikers on marine plastic debris: Human exposure risks at bathing waters and beach environments. Marine Environmental Research, 118, 10–19. https://doi.org/10.1016/j.marenvres.2016.04.006
Kirstein, I. V., Kirmizi, S., Wichels, A., Garin-Fernandez, A., Erler, R., Löder, M., & Gerdts, G. (2016). Dangerous hitchhikers? Evidence for potentially pathogenic Vibrio spp. on microplastic particles. Marine Environmental Research, 120, 1–8. https://doi.org/10.1016/j.marenvres.2016.07.004
Kirstein, I. V., Wichels, A., Gullans, E., Krohne, G., & Gerdts, G. (2019). The plastisphere – uncovering tightly attached plastic ‘specific’ microorganisms. PLoS ONE, 14(4), 1–17. https://doi.org/10.1371/journal.pone.0215859
Klindworth, A., Quast, C., Pruesse, E., Schweer, T., Horn, M., & Glo, F. O. (2013). Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Research, 41(1), 1–11. https://doi.org/10.1093/nar/gks808
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K., & Schloss, P. D. (2013). Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the Miseq Illumina sequencing platform. Applied and Environmental Microbiology, 79(17), 5112–5120. https://doi.org/10.1128/AEM.01043-13
Kunin, V., Engelbrektson, A., Ochman, H., & Hugenholtz, P. (2010). Wrinkles in the rare biosphere: Pyrosequencing errors can lead to artificial inflation of diversity estimates. Environmental Microbiology, 12(1), 118–123. https://doi.org/10.1111/j.1462-2920.2009.02051.x
Kurmayer, R., Deng, L., & Entfellner, E. (2016). Role of toxic and bioactive secondary metabolites in colonization and bloom formation by filamentous Cyanobacteria planktothrix. Harmful Algae, 54, 69–86. https://doi.org/10.1016/j.hal.2016.01.004
Lavers, J. L., Bond, A. L., & Hutton, I. (2014). Plastic ingestion by flesh-footed shearwaters (Puffinus carneipes): Implications for fledgling body condition and the accumulation of plastic-derived chemicals. Environmental Pollution, 187, 124–129. https://doi.org/10.1016/j.envpol.2013.12.020
Lee, J. W., Nam, J. H., Kim, Y. H., Lee, K. H., & Lee, D. H. (2008). Bacterial communities in the initial stage of marine biofilm formation on artificial surfaces. Journal of Microbiology, 46(2), 174–182. https://doi.org/10.1007/s12275-008-0032-3
Li, A., Liu, H., Sun, B., Zhou, Y., and Xin, Y.. (2014). “Flavobacterium lacus sp. nov., isolated from a high-altitude lake, and emended description of Flavobacterium filum.” International Journal of Systematic and Evolutionary Microbiology 64 (PART 3): 933–39. doi: 10.1099/ijs.0.056689-0.
Li, J., Liu, H., & Paul Chen, J. (2018). Microplastics in freshwater systems : A review on occurrence , environmental effects , and methods for microplastics detection. Water Research, 137, 362–374. https://doi.org/10.1016/j.watres.2017.12.056
Liu, J.J., Zhang, X.Q., Chi, F.T., Pan, J., Sun, C., and Wu, M.. (2014). “Gemmobacter megaterium sp. nov., isolated from coastal planktonic seaweeds.” International Journal of Systematic and Evolutionary Microbiology 64 (PART 1): 66–71. doi: 10.1099/ijs.0.050955-0.
Liu, Y., Li, H., Jiang, J. T., Liu, Y. H., Song, X. F., Xu, C. J., & Liu, Z. P. (2009). Algoriphagus aquatilis sp. nov., isolated from a freshwater lake. International Journal of Systematic and Evolutionary Microbiology, 59(7), 1759–1763. https://doi.org/10.1099/ijs.0.005215-0
Lobelle, D., & Cunliffe, M. (2011). Early microbial biofilm formation on marine plastic debris. Marine Pollution Bulletin, 62(1), 197–200. https://doi.org/10.1016/j.marpolbul.2010.10.013
Lusher, A. L., McHugh, M., & Thompson, R. C. (2013). Occurrence of microplastics in the gastrointestinal tract of pelagic and demersal fish from the English channel. Marine Pollution Bulletin, 67(1–2), 94–99. https://doi.org/10.1016/j.marpolbul.2012.11.028
McCormick, A. R., Hoellein, T. J., London, M. G., Hittie, J., Scott, J. W., & Kelly, J. J. (2016). Microplastic in surface waters of urban rivers: Concentration, sources, and associated bacterial assemblages. Ecosphere, 7(11). https://doi.org/10.1002/ecs2.1556
Miao, L., Wang, P., Hou, J., Yu, Y., Liu, Z., Liu, S., & Li, T. (2019). Science of the total environment distinct community structure and microbial functions of biofilms colonizing microplastics. Science of the Total Environment, 650, 2395–2402. https://doi.org/10.1016/j.scitotenv.2018.09.378
Nedashkovskaya, O. I., Vancanneyt, M., Van Trappen, S., Vandemeulebroecke, K., Lysenko, A. M., Rohde, M., Falsen, E., Frolova, G. M., Mikhailov, V. V., & Swings, J. (2004). Description of Algoriphagus aquimarinus sp. nov., Algoriphagus chordae sp. nov. and Algoriphagus winogradskyi sp. nov., from sea water and algae, transfer of Hongiella halophila Yi and Chun 2004 to the genus Algoriphagus as Algoriphagus halophilus comb. N. International Journal of Systematic and Evolutionary Microbiology, 54(5), 1757–1764. https://doi.org/10.1099/ijs.0.02915-0
Negoro, S. (2000). Biodegradation of nylon oligomers. Applied Microbiology and Biotechnology, 54(4), 461–466. https://doi.org/10.1007/s002530000434
Nelms, S. E., Barnett, J., Brownlow, A., Davison, N. J., Deaville, R., & Galloway, T. S. (2019). Microplastics in marine mammals stranded around the British Coast: Ubiquitous but transitory? Scientific Reports, 1–8. https://doi.org/10.1038/s41598-018-37428-3
Nõges, P., & Ott, I. (2003). Occurrence, coexistence and competition of Limnothrix redekei and Planktothrix agardhii: Analysis of Danish-Estonian Lake Database. Algological Studies/Archiv Für Hydrobiologie, Supplement Volumes, 109(April 2014), 429–441. https://doi.org/10.1127/1864-1318/2003/0109-0429
Oberbeckmann, S., Kreikemeyer, B., Labrenz, M., & Harrison, J. P. (2018). Environmental factors support the formation of specific bacterial assemblages on microplastics. Frontiers in Microbiology, 8(January), 1–12. https://doi.org/10.3389/fmicb.2017.02709
Oberbeckmann, S., Loeder, M. G. J., Gerdts, G., & Osborn, A. M. (2014). Spatial and seasonal variation in diversity and structure of microbial biofilms on marine plastics in Northern European waters. FEMS Microbiol Ecol, 90, 478–492. https://doi.org/10.1111/1574-6941.12409
Oberbeckmann, S., Mark Osborn, A., & Duhaime, M. B. (2016). Microbes on a bottle: Substrate, season and geography influence community composition of microbes colonizing marine plastic debris. PLoS ONE, 11(8), 1–24. https://doi.org/10.1371/journal.pone.0159289
Pal, L., Kraigher, B., Brajer-humar, B., Levstek, M., & Mandic-mulec, I. (2012). Bioresource technology total bacterial and ammonia-oxidizer community structure in moving bed biofilm reactors treating municipal wastewater and inorganic synthetic wastewater. Bioresource Technology, 110, 135–143. https://doi.org/10.1016/j.biortech.2012.01.130
Park, S., Park, J. M., & Yoon, J. H. (2017). Algoriphagus Marisflavi sp. nov., isolated from water of an estuary environment. International Journal of Systematic and Evolutionary Microbiology, 67(10), 4168–4174. https://doi.org/10.1099/ijsem.0.002273
Peng, C. C., Binti, F., & Yusuff, M. (2015). Feeding behaviour of tilapia ( Oreochromis sp.) fingerlings to microbeads. Phil. J. of Nat. Sci, 24(March), 21–26.
Peng, G., Zhu, B., Yang, D., Lei, S., Shi, H., & Li, D. (2017). Microplastics in sediments of the Changjiang Estuary , China *. Environmental Pollution, 225, 283–290.
Pesce, A., Toccaceli, G., & Andrea, G. D. (2016). Uncommon strain for an intracranial infection : Bacillus simplex as suspected cause of brain abscess. Journal of Neuroinfectious Diseases, 7(1), 6–8. https://doi.org/10.4172/2314-7326.1000209
Plasticseurope (2019). 2019. “Plastics - the Facts 2019.” https://www.plasticseurope.org/en/resources/market-data.
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., & Glöckner, F. O. (2013). The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Research, 41(D1), 590–596. https://doi.org/10.1093/nar/gks1219
Line, R. and Eva, S.. 2019. “Bacterial candidates for colonization and degradation of marine plastic debris.” Environmental Science & Technology, 11636–43.
Ryan, P. G. (1988). Effects of ingested plastic on seabird feeding: Evidence from chickens. Marine Pollution Bulletin, 19(3), 125–128. https://doi.org/10.1016/0025-326X(88)90708-4
Saber, D. L., & Crawford, R. L. (1985). Isolation and characterization of Flavobacterium strains that degrade pentachlorophenol. Applied and Environmental Microbiology, 50(6), 1512–1518. https://doi.org/10.1128/aem.50.6.1512-1518.1985
Schloss, P. D., Westcott, S. L., Ryabin, T., Hall, J. R., Hartmann, M., Hollister, E. B., & Ryan A. Lesniewski, et al. (2009). Introducing Mothur: Open-source, platform-independent, community-supported software for describing and comparing microbial communities. Applied and Environmental Microbiology, 75(23), 7537–7541. https://doi.org/10.1128/AEM.01541-09
Sudhakar, M., Mukesh, D., Sriyutha Murthy, P., & Venkatesan, R. (2008). Marine microbe-mediated biodegradation of low- and high-density polyethylenes. International Biodeterioration and Biodegradation, 61(3), 203–213. https://doi.org/10.1016/j.ibiod.2007.07.011
Thompson, R. C., Olson, Y., Mitchell, R. P., Davis, A., Rowland, S. J., John, A. W. G., McGonigle, D., & Russell, A. E. (2004). Lost at sea: Where is all the plastic? Science, 304(5672), 838. https://doi.org/10.1126/science.1094559
Tindall, B. J., Rosselló-Móra, R., Busse, H. J., Ludwig, W., & Kämpfer, P. (2010). Notes on the characterization of prokaryote strains for taxonomic purposes. International Journal of Systematic and Evolutionary Microbiology, 60(1), 249–266. https://doi.org/10.1099/ijs.0.016949-0
Turner, S., Pryer, K. M., Miao, V. P. W., & Palmer, J. D. (1999). Investigating deep phylogenetic relationships among Cyanobacteria and plastids by small subunit RRNA sequence analysis. Journal of Eukaryotic Microbiology, 46(4), 327–338. https://doi.org/10.1111/j.1550-7408.1999.tb04612.x
Vet, B., Pulawy, I., Kozi, A., & Diseases, F. (2010). Serotyping of Aeromonas species isolated from Polish fish farms in relation to species and virulence phenotype of the bacteria. Bull Vet Inst Pulawy, 81(8), 315–320.
Viršek, M. K., Lovšin, M. N., Koren, Š., Kržan, A., & Peterlin, M. (2017). Microplastics as a vector for the transport of the bacterial fish pathogen species Aeromonas salmonicida. Marine Pollution Bulletin. https://doi.org/10.1016/j.marpolbul.2017.08.024
Wagner, M., Scherer, C., Alvarez-muñoz, D., Brennholt, N., Bourrain, X., Buchinger, S., Fries, E., et al. (2014). Microplastics in freshwater ecosystems: What we know and what we need to know. Environmental Sciences Europe, 1–9. https://doi.org/10.1186/s12302-014-0012-7
Wang, L., Tong, J., Li, Y., Zhu, J., Zhang, W., Niu, L., & Zhang, H. (2020). Bacterial and fungal assemblages and functions associated with biofilms differ between diverse types of plastic debris in a freshwater system. Environmental Research, 110371. https://doi.org/10.1016/j.envres.2020.110371
Willame, R., Jurczak, T., Iffly, J. F., Kull, T., Meriluoto, J., & Hoffmann, L. (2005). Distribution of hepatotoxic cyanobacterial blooms in Belgium and Luxembourg. Hydrobiologia, 551(1), 99–117. https://doi.org/10.1007/s10750-005-4453-2
Wu, X., Pan, J., Li, M., Li, Y., Bartlam, M., & Wang, Y. (2019). Selective enrichment of bacterial pathogens by microplastic biofilm. Water Research, 165, 114979. https://doi.org/10.1016/j.watres.2019.114979
Yoo, Y., Lee, D. W., Lee, H., Kwon, B. O., Khim, J. S., Yim, U. H., Park, H., et al. (2019). Gemmobacter lutimaris sp. nov., a marine bacterium isolated from a tidal flat. International Journal of Systematic and Evolutionary Microbiology, 69(6), 1676–1681. https://doi.org/10.1099/ijsem.0.003375
Zettler, E. R., Mincer, T. J., & Amaral-zettler, L. A. (2013). Life in the ‘ plastisphere ’: microbial communities on plastic marine debris. Environ. Sci. Technol, 47, 7137–7146. https://doi.org/10.1021/es401288x
Funding
Open access funding provided by Hungarian University of Agriculture and Life Sciences. Project no. 2020-1.1.2-PIACI-KFI-2021-00239 has been implemented with the support provided by the Ministry of Innovation and Technology of Hungary from the National Research, Development and Innovation Fund, financed under the PIACI KFI funding scheme. This research was also supported by the Ministry of Innovation and Technology within the framework of the Thematic Excellence Programme 2020, National Challenges Subprogramme (TKP2020-NKA-16) and by the UNKP-18-4-SZIE-18 new national Excellence Program of the Ministry of Human Capacities (István Szabó).
Author information
Authors and Affiliations
Contributions
Conceptualization: István Szabó, Sándor Szoboszlay, Balázs Kriszt
Data curation: István Szabó, Al-Omari Jafar, Gábor Soma Szerdahelyi, Milan Farkas, Sándor Szoboszlay, Al-Omari Yazid, Péter Márton Szabó, Rózsa Sebők
Formal analysis: István Szabó, Al-Omari Jafar, Gábor Soma Szerdahelyi
Funding acquisition: Sándor Szoboszlay, Balázs Kriszt, István Szabó
Investigation: István Szabó, Al-Omari Jafar, Gábor Soma Szerdahelyi
Methodology: Istvan Szabo, Al-Omari Jafar, Gábor Soma Szerdahelyi, Milan Farkas, Al-Omari Yazid, Péter Márton Szabó, Rózsa Sebők, Jeffrey Griffitts
Resources: Sándor Szoboszlay, István Szabó, Balázs Kriszt
Writing: original draft: Al-Omari Jafar,
Writing: review & editing: István Szabó, Jeffrey Griffitts
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
ESM 1
(DOCX 17 kb)
ESM 2
(DOCX 13 kb)
ESM 3
(DOCX 20 kb)
ESM 4
Supplementary Figure 1. Rarefaction curves of the 97% OTUs for all microplastics and water samples. (JPG 563 kb)
ESM 5
Supplementary Figures 2, 3, 4, 5, and 6: shared species, sequences, and richness between microplastic and water samples (JPG 707 kb)
ESM 6
(JPG 530 kb)
ESM 7
(JPG 534 kb)
ESM 8
(JPG 535 kb)
ESM 9
(JPG 693 kb)
ESM 10
Supplementary Figure 7. Dominant bacterial phyla and orders associated with microplastics and surrounding water. (JPG 539 kb)
ESM 11
Supplementary Figure 8. Dominant bacterial families associated with microplastics and surrounding water. (JPG 579 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Szabó, I., Al-Omari, J., Szerdahelyi, G.S. et al. In Situ Investigation of Plastic-Associated Bacterial Communities in a Freshwater Lake of Hungary. Water Air Soil Pollut 232, 493 (2021). https://doi.org/10.1007/s11270-021-05445-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11270-021-05445-0