Abstract
Quorum quenching (QQ), a mechanism which inhibits, interferes or inactivates quorum sensing, has been investigated for control of biofilms instigated by quorum sensing process. Application of quorum quenchers (QQs) provides the possibility to investigate how different phenotypes of Pseudomonas aeruginosa (non-mucoid, mucoid, and heavily mucoid strains) modulate their gene expression to form biofilms, their quorum sensing (QS) mediated biofilm to be formed, and their virulence expressed. The mRNA expression of the AHL-mediated QS circuit and AHL-mediated virulence factors in P. aeruginosa was investigated in presence of QQs. qPCR analysis showed that farnesol and tyrosol actively reduce the expression of the synthase protein, LasI and RhlI, and prevent production of 3OC12-HSL and C4-HSL, respectively. Also, the use of farnesol and tyrosol significantly moderated gene expression for exo-proteins toxA, aprA, LasB, as well as rhlAB, which are responsible for rhamnolipid production. Our findings were promising, identifying several suppressive regulatory effects of furanone and Candida albicans QS signal molecules, tyrosol, and farnesol on the AHL-mediated P. aeruginosa QS network and related virulence factors.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In humans, Pseudomonas aeruginosa strains can cause infections and diseases such as wound infections, periodontitis, keratitis and chronic pneumonia during cystic fibrosis (CF). These usually occur in individuals with compromised immune systems (Gellatly and Hancock 2013). The diversity of genetic variation amongst P. aeruginosa strains is highlighted by the presence of non-mucoid, mucoid and heavily mucoid phenotypes (Workentine et al. 2013).
An important factor contributing toward the pathogenesis of P. aeruginosa is its remarkable ability to switch between planktonic and sessile modes of growth in the onset of chronic infections. The sessile cells, compared to planktonic cells, do not show an enhanced resistance towards antibiotics. However, they show an increased threshold in their minimum inhibitory concentration (MIC) against the antibiotic of interest when compared as they are enveloped within an extracellular matrix (Lebeaux et al., 2014). The basic assumption regarding the protective role played by the biofilm is that of reduced diffusion of antibiotics into the inner layers of the biofilms and their effective access to the sessile bacteria (Mah et al. 2003; Singh et al. 2021).
P. aeruginosa produces numerous virulence factors that aid in its colonisation during an infection. Virulence factors such as mucoid exopolysaccharides, rhamnolipids, haemolysins, proteases, lipopolysaccharides, pili, and lipases are common tools used by P. aeruginosa to invade, adhere and colonise the host. The arsenal of virulence factors makes P. aeruginosa a potent pathogen (Schaber et al. 2004; Schroeder et al. 2017).
Many processes involving virulence are modulated by the hierarchical quorum sensing (QS) circuits documented in P. aeruginosa. These traits are involved in the pathogenicity of the bacteria (Lee and Zhang 2014). It has been proposed that targeting the QS system of P. aeruginosa by interrupting bacterial communication instead of killing the bacteria by antibiotics would have an anti-pathogenic effect and may aid in the fight against biofilm forming pathogens and antibiotic resistance (Barlow and Nathwani 2005; Rémy et al. 2020).
Conventional P. aeruginosa antibiotics such as ceftazidime, ciprofloxacin, and azithromycin display QQ activity (Skindersoe et al. 2008; Swatton et al. 2016). However, even though the mechanism is not fully understood, it has been speculated that their QQ activity involves inhibiting bacterial protein synthesis which prevents the expression of the inducer protein from synthesising the QS signal molecules (Reuter et al., 2016). Azithromycin is also known to inhibit alginate production in mucoid strains of P. aeruginosa (Imperi, Leoni and Visca, 2014). In a recent study, the newly synthesised P. aeruginosa quorum-sensing autoinducer analogues (AIA-1, -2) were demonstrated to boost the bactericidal efficacy of azithromycin by altering the cell surface hydrophobicity of P. aeruginosa, reducing antibiotic tolerance (Abe et al. 2021). The use of the said antibiotics has been known to induce resistance amongst bacteria, hence they cannot be used as potential QQs.
As Acyl-Homoserine Lactones (AHLs) represent the primary QS signal molecules in P. aeruginosa, utilising small molecules that mimic AHLs to inhibit QS is a promising strategy. Furanones were first identified as QQs that inhibit by mimicking AHL molecules by attaching to the LasR receptor of P. aeruginosa. This interfered with the binding of the AHL molecule, thereby preventing QS mediated gene regulation in P. aeruginosa (Manner and Fallarero., 2018). Since the discovery of the role of furanone in inhibition of the QS system, numerous synthetically produced, structurally diverse furanone derivatives have been synthesised (Irie et al. 2017). Similarly, an AHL analogue, meta-bromo-thiolactone, competitively inhibits QS in P. aeruginosa and prevents biofilm formation and pyocyanin production and protects lung cells against the antagonistic activity of P. aeruginosa (O’Loughlin et al., 2013).
While there are numerous environmental and physiological cues for dispersal of biofilm, chemicals produced by microorganism themselves can be utilised in a strategy to induce biofilm dispersal and inhibit QS in P. aeruginosa. Candida albicans produces two QS molecules, namely farnesol and tyrosol (Kaplan 2010). Farnesol was shown to inhibit the morphological shift from yeast to hyphae at high cell densities while tyrosol was shown to accelerate the transition from yeast to hyphae in C. albicans (Decanis et al. 2011). As P. aeruginosa and C. albicans are known to coexist in numerous nosocomial opportunistic infections (Méar et al. 2013; Doing et al. 2020), the influence of farnesol and tyrosol as potential QQs may be investigated against P. aeruginosa biofilm formation and subsequent virulence production between the non-mucoid and mucoid phenotypes along with other QQs and biofilm dispersal agents.
This study investigated the effect of quorum quenching (QQ) on biofilm formation and virulence factor secretion of three strains of P. aeruginosa: P. aeruginosa NCTC 10,662 (non-mucoid), P. aeruginosa PAO1 (mucoid) and P. aeruginosa RBHi (a heavily mucoid CF isolate).
Materials and methods
Preparation of bacterial inoculum, biofilm formation and dispersal
P. aeruginosa NCTC 10,662 (non-mucoid), P. aeruginosa PAO1 (semi mucoid) and P. aeruginosa RBHi (heavily mucoid) were selected as model phenotypes for this study.
P. aeruginosa PAO1 and P. aeruginosa NCTC 10,662 were obtained from the University of Westminster, London culture collection. P. aeruginosa CF isolate was kindly donated by the culture collection facility at Royal Brompton Hospital, London, UK (referred to as RBHi in this study).
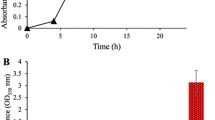
P. aeruginosa strains were subcultured into Lysogeny broth (LB broth) from the slants and incubated aerobically overnight (16–18 h) at 37oC at 180 rpm. The absorbance (OD600) of the overnight growth of P. aeruginosa sp. was readjusted in sterile LB broth to obtain an equivalence absorption according to 0.5 McFarland standards (~ 1.5 × 108 cells) and used for further biofilm growth. A flow-cell (Transmission Flow-cell, FC 281, BioSurface Technologies Corporation (BST), Montana, USA) provided a closed system where bacterial attachment occurred and biofilm structure and hence extracellular polymeric substances (EPS) were produced.
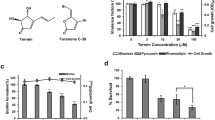
Quorum quenchers used in this study included (Z-)-4-Bromo-5-bromomethylene)-2(5 H)-furanone; E, E – farnesol; and 2,4- hydroxyphenyl)-ethanol (Tyrosol). Equation 1 was used to calculate the MIC50s, concentrations that inhibited 50% of the isolates, for farnesol, tyrosol, and furanone (Quave et al. 2008). A mean of 3 absorbance readings were taken at OD595 nm for each value (n = 3).
ODs24 = Optical density of culture at 595 nm of experimental sample, 24 h post-inoculation.
ODs0 = Optical Density of culture at 595 nm of experimental sample at time point 0.
ODc24 = Optical density of culture at 595 nm of control sample, 24 h post inoculation.
ODc0 = Optical density of culture at 595 nm of control sample at time point 0.
MIC50 were 125 µg/mL for farnesol and tyrosol, and 5 µg/mL for furanone.
-
1)
Farnesol 125 µg/mL (MIC50).
(1-(1.01–0.51/ 1.59 − 0.56)) x 100 = 51.5%
-
2)
Tyrosol 125 µg/mL (MIC50).
(1-(0.94 − 0.48/ 1.59 − 0.56)) x 100 = 55.4%
-
3)
Furanone 5 µg/mL (MIC50).
(1-(0.89 − 0.45/ 1.59 − 0.56)) x 100 = 57.3%
qPCR analysis of QS network and virulence factors inP. aeruginosa.
RNA extraction from bacterial cells was performed using the Trizol RNA isolation protocol described by Chomczynski and Sacchi in 1987. cDNA preparation was then done by using the QuantiTect Reverse Transcription kit (Qiagen ltd, Manchester, UK).
Samples were prepared using diluted forward and reverse primer (Tables 1 and 2) mix (1/10 dilution). They were then run in duplicate at 95 °C initial denaturation for 2 min. Subsequently, the amplification program involved 40 cycles of denaturation at 95 °C for 15 s, primer annealing at 55 °C for 15 s and extension at 72 °C for 30 s. A final extension was performed at 72 °C for 2 min followed by cooling at 4 °C. A dissociation step at 55 °C was used to generate a melting curve with a 1 °C increase every 5 s till 95 °C to obtain verification of amplified product. Reference genes were as reported in Table 3. SYBR Green qPCR was performed using Rotor Gene Q (Qiagen ltd, Manchester, UK).
Statistical analysis
All experiments were performed in triplicates. All data for assays performed in this study were statistically analysed using GraphPad Prism to determine p values and establish correlation between data sets. All graphs were plotted using GraphPad Prism. p < 0.05 was considered significant.
Results
Quantitative PCR and expression levels of QS genes and QS mediated virulence factors ofP. aeruginosa.
Following the treatment with MIC50 furanone, farnesol and tyrosol, there was a significant reduction (p = 0.0001) in the relative abundance of RNA for all the LasI protein (Fig. 1a) from the three strains of P. aeruginosa. Comparison between farnesol and tyrosol showed that farnesol had a greater effect on LasI compared to tyrosol. The effect of farnesol and tyrosol on LasI was more pronounced in NCTC 10,662 compared to PAO1 and RBHi. This was not the case with LasR. With the exception of RBHi, treatment with farnesol up-regulated the expression of LasR in NCTC 10,662 and PAO1. LasR was also up-regulated with tyrosol in PAO1 (Fig. 1b).
mRNA expression of AHL mediated QS circuit in P. aeruginosa sessile cells grown with QQ. The fold change of mRNA for LasI (a), LasR (b), RhlI (c) and RhlR (d) was determined for P. aeruginosa sessile cells extracted from biofilm treated with furanone, farnesol and tyrosol after ~ 16 h growth. Results are expressed as the mean fold change (control standardised to 1.0) with error bars representing SEM (n = 9)
Reduction in mRNA expression for RhlI protein was seen with all the strains when treated with furanone, farnesol, and tyrosol (Fig. 1c). Similar to LasI, NCTC 10,662 showed the highest reduction in RNA content for RhlI, while all three strains showed a down-regulation of RhlI when treated with farnesol and tyrosol (Fig. 1c). In the case of RBHi, tyrosol up-regulated the expression of RhlR (Fig. 1d), however, down-regulation of RhlR was marginal when treated with farnesol.
Figure 1.
qPCR analysis of expression of genes related to virulence factors of P. aeruginosa showed that treatment with furanone, farnesol, and tyrosol down-regulated the expression of toxA, aprA, rhlAB and LasB when compared to the control as shown in Fig. 2. Farnesol seemed highly effective in down-regulating rhIAB when compared to tyrosol for all the strains of P. aeruginosa (Fig. 2c). LasB showed the highest down-regulation amongst all the genes analysed with furanone treatment (Fig. 2d).
mRNA expression of AHL mediated virulence factors of P. aeruginosa sessile cells grown with QQs. The fold change of mRNA for toxA (a), aprA (b), rhlAB (c) and LasB (d) was determined for P. aeruginosa sessile cells extracted from biofilm treated with furanone, farnesol and tyrosol after ~ 16 h growth. Results are expressed as the mean fold change (control standardised to 1.0) with error bars representing SEM (n = 9)
Data pertaining to cell counts from the control and experimental cultures that underwent treatment with furanone, farnesol and tyrosol is provided as supplementary information to corroborate the regulation of gene expression in P aeruginosa sp.
Discussion
In recent years, QS has been the primary focus of research involving treatment of biofilm mediated chronic infections. It has been well established that QS plays a vital role in development and formation of biofilms which is a recalcitrant mode of growth and aids in the onset of bacterial resistance towards conventional antibiotics (Pletzer et al. 2018). This has led to an inexorable rise in formation of superbugs-related infections that are extremely hard to treat due to the dearth of effective therapies (Fernández et al. 2011). Therefore, recent research has been focused on elucidating novel therapeutic strategies in order to combat biofilm-related infections by attenuating the ability of cell-to-cell communication by targeting the QS signalling system present in bacteria (Römling and Balsalobre., 2012 and Bi et al. 2021). This in turn would aid in arresting biofilm formation and biofilm related chronic infections. Novel therapies that target the QS system in pathogenic bacteria could provide the foundation for the development of next generation anti-virulence therapies.
Three primary strategies can be adopted to combat biofilm formation by inhibiting QS. The most frequently studied strategies are degradation and modification of QS signals to prevent bacterial communication along with competitive inhibition of the receptor protein of the QS circuit as well as impeding the function of synthase protein responsible for producing the QS molecules (LaSarre and Federle., 2013, Bhatt et al. 2021). In the natural environment, blocking communication of ecological niche adversary is essential for survival. This is a promising avenue to explore in order to get a better understanding of the process of social interaction of individual bacterial species or of groups aiming to dictate the niche (Bhagirath et al. 2016). As vital QS is for bacterial coordination and survival, it is also essential for bacteria to interfere with QS of other microbes in order to gain an advantage for survival (Li and Tian., 2012). This naturally existing process of communication interference to gain an advantage over competitors can be exploited to develop novel therapies targeting the QS system of pathogenic bacteria.
Gene expression studies, involving the individual effect of farnesol and tyrosol, were conducted along with furanone. All the three strains showed a significant down-regulation in both the AHL mediated synthases (LasI and RhlR) when treated with furanone (Fig. 1). However, LasR receptor was up-regulated in NCTC 10,662 using farnesol and in PAO1 using farnesol and tyrosol. Similar up-regulation of RhlR receptor was observed in RBHi when treated with tyrosol. This shows that farnesol and tyrosol actively reduce the expression of the synthase protein (LasI and RhlI) and prevent production of N-3-oxo-dodecanoyl homoserine lactone (3OC12-HSL) and C4-HSL respectively. lasR gene is linked to lasI and is considered to be the cognate receptor for 3OC12-HS. Signal synthase, RhlI, generates C4-HSL, and the C4-HSL receptor is called RhlR (Muh et al. 2006). Significant down-regulation in gene expression was observed with the use of farnesol and tyrosol for exo-proteins toxA, aprA and LasB as well as rhlAB which are responsible for the production of rhamnolipid.
Since the introduction of antibiotics as novel treatment against infections in the 1940s, multi drug resistance traits have become synonymous with biofilm forming opportunistic pathogens that dominate the nosocomial setting (Bryers 2008). This necessitates the identification and investigation of novel antimicrobial therapies and their mode of action. Exploiting interspecies and interkingdom interactions/ competition as well as naturally occurring products and their use in synergy could provide a potential solution to combat/ eradicate biofilm related chronic infections and drug resistance.
Conclusions
This study demonstrated the effect of quorum quenchers derived from C. albicans and the subsequent changes in gene expression of biofilm formation and virulence factors in P. aeruginosa.
Our findings have shown promise in identifying several suppressive regulatory effects of C. albicans QS signal molecules on AHL mediated P. aeruginosa QS network and related virulence factors. It is well known that conventional antibiotics cause resistance in bacteria. This study provides a novel option for the control of P. aeruginosa biofilm related infections and diseases, as well as a foundation for future research on the mechanisms of biofilm inhibition in different P. aeruginosa phenotypes from the perspective of QQ. The QQs used in this study provide the opportunity for application of these molecules as an alternative to traditional antibiotics. Future challenges for the research include studying the effects of the QQs in vivo. This brings the knowledge obtained from this research closer to real-life application.
Further research will extend the understanding of mechanistic action of the QQ.
Quantification of P. aeruginosa virulence factors such as rhamnolipid, pyoverdine, pyocyanin, and elastolytic activity, as well as the use of the crystal violet method to measure biofilm formation upon treatment with quorum quenchers, may help to improve the findings.
References
Abe M, Murakami K, Hiroshima Y, Amoh T, Sebe M, Kataoka K, Fujii H (2021) Autoinducer Analogs Can Provide Bactericidal Activity to Macrolides in Pseudomonas aeruginosa through Antibiotic Tolerance Reduction. Antibiotics 11:10. https://. doi
Aghamollaei H, Moghaddam M, Kooshki H, Heiat M, Mirnejad R, Barzi N (2015) Detection of Pseudomonas aeruginosa by a triplex polymerase chain reaction assay based on lasI/R and gyrB genes. J Infect Public Health 8:314–322. https://doi.org/10.1016/j.jiph.2015.03.003
Barlow G, Nathwani D (2005) Is antibiotic resistance a problem? A practical guide for hospital clinicians. Postgrad Med J 81:680–692. https://doi.org/10.1136/pgmj.2005.035113
Bhagirath A, Li Y, Somayajula D, Dadashi M, Badr S, Duan K (2016) Cystic fibrosis lung environment and Pseudomonas aeruginosa infection. BMC Pulm Med 16 https://doi.org/10.1186/s12890-016-0339-5
Bhatt P, Zhou X, Huang Y, Zhang W, Chen S (2021) Characterization of the role of esterases in the biodegradation of organophosphate, carbamate, and pyrethroid pesticides. J Hazard Mater 411:125026. doi: https://doi.org/10.1016/j.jhazmat.2020.125026
Bi Y, Xia G, Shi C, Wan J, Liu L, Chen Y, Wu Y, Zhang W, Zhou M, He H, Liu R (2021) Therapeutic strategies against bacterial biofilms. Fundam res 1:193–212. https://doi.org/10.1016/j.fmre.2021.02.003
Bryers J (2008) Medical biofilms. Biotechnol Bioeng 100:1–18. https://doi.org/10.1002/bit.21838
Decanis N, Tazi N, Correia A, Vilanova M, Rouabhia M (2011) Farnesol, a Fungal Quorum-Sensing Molecule Triggers Candida Albicans Morphological Changes by Downregulating the Expression of Different Secreted Aspartyl Proteinase Genes. Open Microbiol J 5:119–126. https://doi.org/10.2174/1874285801105010119
Doing G, Koeppen K, Occipinti P, Harty CE, Hogan DA (2020) Conditional antagonism in co-cultures of Pseudomonas aeruginosa and Candida albicans: An intersection of ethanol and phosphate signalling distilled from dual-seq transcriptomics. PLoS Genet 16:e1008783. https://doi.org/10.1371/journal.pgen.1008783
Fernández L, Breidenstein E, Hancock R (2011) Creeping baselines and adaptive resistance to antibiotics. Drug Resist Updat 14:1–21. https://doi.org/10.1016/j.drup.2011.01.001
Gellatly S, Hancock R (2013) Pseudomonas aeruginosa: new insights into pathogenesis and host defenses. Pathog Dis 67:159–173. https://doi.org/10.1111/2049-632x.12033
Irie Y, Roberts A, Kragh K, Gordon V, Hutchison J, Allen R, Melaugh G, Bjarnsholt T, West S, Diggle S (2017) The Pseudomonas aeruginosa PSL Polysaccharide Is a Social but Noncheatable Trait in Biofilms. mBio 8. https://doi.org/10.1128/mbio.00374-17
Jack A, Khan S, Powell L, Pritchard M, Beck K, Sadh H, Sutton L, Cavaliere A, Florance H, Rye P, Thomas D, Hill K (2018) Alginate Oligosaccharide-Induced Modification of the lasI-lasR and rhlI-rhlR Quorum-Sensing Systems in Pseudomonas aeruginosa. Antimicrob Agents Chemother 62. https://doi.org/10.1128/aac.02318-17
Kaplan J (2010) Biofilm Dispersal: Mechanisms, Clinical Implications, and Potential Therapeutic Uses. J Dent Res 89:205–218. https://doi.org/10.1177/0022034509359403
LaSarre B, Federle M (2013) Exploiting Quorum Sensing to Confuse Bacterial Pathogens. MMBR 77:73–111. https://doi.org/10.1128/mmbr.00046-12
Lebeaux D, Ghigo J, Beloin C (2014) Biofilm-Related Infections: Bridging the Gap between Clinical Management and Fundamental Aspects of Recalcitrance toward Antibiotics. MMBR 78:510–543. https://doi.org/10.1128/mmbr.00013-14
Lee J, Zhang L (2014) The hierarchy quorum sensing network in Pseudomonas aeruginosa. Protein Cell 6:26–41. https://doi.org/10.1007/s13238-014-0100-x
Li Y, Tian X (2012) Quorum Sensing and Bacterial Social Interactions in Biofilms. Sensors 12:2519–2538. https://doi.org/10.3390/s120302519
Mah T, Pitts B, Pellock B, Walker G, Stewart P, O’Toole G (2003) A genetic basis for Pseudomonas aeruginosa biofilm antibiotic resistance. Nature 426:306–310. https://doi.org/10.1038/nature02122
Manner S, Fallarero A (2018) Screening of Natural Product Derivatives Identifies Two Structurally Related Flavonoids as Potent Quorum Sensing Inhibitors against Gram-Negative Bacteria. Int J Mol Sci 19:1346. https://doi.org/10.3390/ijms19051346
Méar J, Kipnis E, Faure E, Dessein R, Schurtz G, Faure K, Guery B (2013) Candida albicans and Pseudomonas aeruginosa interactions: More than an opportunistic criminal association? Med Mal Infect 43:146–151. https://doi.org/10.1016/j.medmal.2013.02.005
Muh U, Hare BJ, Duerkop BA, Schuster M, Hanzelka BL, Heim R, Olson ER, Greenberg EP (2006) A structurally unrelated mimic of a Pseudomonas aeruginosa acyl-homoserine lactone quorum-sensing signal. PNAS 103:16948–16952. https://doi.org/10.1073/pnas.0608348103
O’Loughlin C, Miller L, Siryaporn A, Drescher K, Semmelhack M, Bassler B (2013) A quorum-sensing inhibitor blocks Pseudomonas aeruginosa virulence and biofilm formation. PNAS 110:17981–17986. https://doi.org/10.1073/pnas.1316981110
Pletzer D, Mansour S, Hancock R (2018) Synergy between conventional antibiotics and anti-biofilm peptides in a murine, sub-cutaneous abscess model caused by recalcitrant ESKAPE pathogens. PLoS Pathog 14:e1007084. https://doi.org/10.1371/journal.ppat.1007084
Pourakbari B, Yaslianifard S, Yaslianifard S, Mahmoudi S, Keshavarz-Valian S, Mamishi S (2016) Evaluation of efflux pumps gene expression in resistant Pseudomonas aeruginosa isolates in an Iranian referral hospital. Iran J Microbiol 8:249–256 PMID: 28210464; PMCID: PMC5296939
Quave CL, Plano LR, Pantuso T, Bennett BC (2008) Effects of extracts from Italian medicinal plants on planktonic growth, biofilm formation and adherence of methicillin-resistant Staphylococcus aureus. J Ethnopharmacol 118(3):418–428 https:\\doi:. https://doi.org/10.1016/j.jep.2008.05.005
Rémy B, Plener L, Decloquement P, Armstrong N, Elias M, Daudé D, Chabrière É (2020) Lactonase Specificity Is Key to Quorum Quenching in Pseudomonas aeruginosa. Front microbiol 11:762. https://doi.org/10.3389/fmicb.2020.00762
Reuter K, Steinbach A, Helms V (2016) Interfering with Bacterial Quorum Sensing. Perspect. Medicinal Chem 8. PMC.S13209. https://doi.org/10.4137/pmc.s13209
Römling U, Balsalobre C (2012) Biofilm infections, their resilience to therapy and innovative treatment strategies. J Intern Med 272:541–561. https://doi.org/10.1111/joim.12004
Sabharwal N, Dhall S, Chhibber S, Harjai K (2014) Molecular detection of virulence genes as markers in Pseudomonas aeruginosa isolated from urinary tract infections. Int J Mol Epidemiol Genet 5:125–134 PMID: 25379131
Schaber J, Carty N, McDonald N, Graham E, Cheluvappa R, Griswold J, Hamood A (2004) Analysis of quorum sensing-deficient clinical isolates of Pseudomonas aeruginosa. J Med Microbiol 53:841–853. https://doi.org/10.1099/jmm.0.45617-0
Schroeder M, Brooks B, Brooks A (2017) The Complex Relationship between Virulence and Antibiotic Resistance. Genes 8:39. https://doi.org/10.3390/genes8010039
Singh S, Datta S, Narayanan KB, Rajnish KN (2021) Bacterial exo-polysaccharides in biofilms: role in antimicrobial resistance and treatments. J Genet Eng Biotechnol 19:140. https://doi.org/10.1186/s43141-021-00242-y
Skindersoe M, Alhede M, Phipps R, Yang L, Jensen P, Rasmussen T, Bjarnsholt T, Tolker-Nielsen T, Hoiby N, Givskov M (2008) Effects of Antibiotics on Quorum Sensing in Pseudomonas aeruginosa. Antimicrob Agents Chemother 52:3648–3663. https://doi.org/10.1128/aac.01230-07
Swatton J, Davenport P, Maunders E, Griffin J, Lilley K, Welch M (2016) Impact of Azithromycin on the Quorum Sensing-Controlled Proteome of Pseudomonas aeruginosa. PLoS ONE 11:e0147698. https://doi.org/10.1371/journal.pone.0147698
Workentine M, Sibley C, Glezerson B, Purighalla S, Norgaard-Gron J, Parkins M, Rabin H, Surette M (2013) Phenotypic Heterogeneity of Pseudomonas aeruginosa Populations in a Cystic Fibrosis Patient. PLoS ONE 8:e60225. https://doi.org/10.1371/journal.pone.0060225
Zhu H, Bandara R, Conibear T, Thuruthyil S, Rice S, Kjelleberg S, Givskov M, Willcox M (2004) Pseudomonas aeruginosa with LasI Quorum-Sensing Deficiency during Corneal Infection. IOVS 45:1897. https://doi.org/10.1167/iovs.03-0980
Funding
This research was supported by the Cavendish Scholarship, University of Westminster, London, UK.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Rachith Kalgudi designed, conducted, analysed and wrote the experiments, discussion and manuscript. Tajalli Keshavarz and Godfrey Kyazze oversaw the research. Roya Tamimi contributed to the writing of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors have no relevant financial or non-financial interests to disclose.
Ethical approval
This article does not report any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kalgudi, R., Tamimi, R., Kyazze, G. et al. Effect of quorum quenchers on virulence factors production and quorum sensing signalling pathway of non-mucoid, mucoid, and heavily mucoid Pseudomonas aeruginosa. World J Microbiol Biotechnol 38, 163 (2022). https://doi.org/10.1007/s11274-022-03339-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03339-9