Abstract
Introduction
KRAS was one of the earliest human oncogenes to be described and is one of the most commonly mutated genes in different human cancers, including colorectal cancer. Despite KRAS mutants being known driver mutations, KRAS has proved difficult to target therapeutically, necessitating a comprehensive understanding of the molecular mechanisms underlying KRAS-driven cellular transformation.
Objectives
To investigate the metabolic signatures associated with single copy mutant KRAS in isogenic human colorectal cancer cells and to determine what metabolic pathways are affected.
Methods
Using NMR-based metabonomics, we compared wildtype (WT)-KRAS and mutant KRAS effects on cancer cell metabolism using metabolic profiling of the parental KRAS G13D/+ HCT116 cell line and its isogenic, derivative cell lines KRAS +/– and KRAS G13D/–.
Results
Mutation in the KRAS oncogene leads to a general metabolic remodelling to sustain growth and counter stress, including alterations in the metabolism of amino acids and enhanced glutathione biosynthesis. Additionally, we show that KRASG13D/+ and KRASG13D/− cells have a distinct metabolic profile characterized by dysregulation of TCA cycle, up-regulation of glycolysis and glutathione metabolism pathway as well as increased glutamine uptake and acetate utilization.
Conclusions
Our study showed the effect of a single point mutation in one KRAS allele and KRAS allele loss in an isogenic genetic background, hence avoiding confounding genetic factors. Metabolic differences among different KRAS mutations might play a role in their different responses to anticancer treatments and hence could be exploited as novel metabolic vulnerabilities to develop more effective therapies against oncogenic KRAS.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Kirsten Rat Sarcoma Viral Oncogene Homolog (Kras or kras2) was one of the earliest human oncogenes to be described and is one of the most commonly mutated genes in different human cancers, including colorectal cancer (Dinu et al., 2014; Tsuchida et al., 2016). Activation of Ras is associated with a wide range of biological responses, such as cell growth, proliferation, survival, differentiation and tissue morphogenesis, depending on the cell type and stimuli (Vartanian et al., 2013). KRAS mutations are associated with poor prognosis and resistance to targeted therapies, demonstrating the clinical importance of this gene (Arrington et al., 2012).
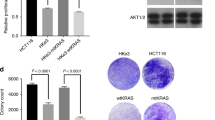
In addition to the well-documented pro-tumorigenic role of mutant Ras alleles, there is some evidence suggesting that the wild type copy of Ras can display tumour-suppressor effects. Zhang et al. showed that following exposure to carcinogenic chemicals, mice harbouring only one copy of wild type Kras2 produced larger lung tumors than mice harbouring both wild type alleles (Zhongqiu Zhang et al., 2001). To investigate the effect of a single copy mutant KRAS, Vartanian et al. analysed the signalling and biological properties of a panel of isogenic cancer cell lines (DLD1, HCT116, and Hec1A) (Vartanian et al., 2013). They demonstrated that a single copy of mutant KRAS results in only modest activation of downstream signalling pathways involving extracellular signal-regulated kinase (ERK) and AKT. One copy of mutant KRAS had a small effect on cell proliferation and cell migration, but a greater effect on cell transformation, as evaluated by growth in soft agar. Interestingly, elimination of the wild type KRAS allele led to increased growth in soft agar, indicating tumor-suppressive characteristics of the wild type KRAS allele under these conditions (Vartanian et al., 2013).
Like many oncogenes, mutant KRAS impacts cellular metabolism and these effects might underlie some of the biological effects of wild type and mutant KRAS outlined above.
Metabonomics (Everett et al., 2019; Lindon et al., 2000) is a novel field in systems biology that produces disease-relevant molecular information downstream of the genome and proteome and has shown early promise in cancer research. We recently demonstrated the power of metabonomics to define the specific profiles of human SW48 colorectal cancer cells carrying different KRAS mutations in codons 12, 13, 61 and 146 compared with their wild-type counterpart (Varshavi et al., 2020).
In this study, we applied NMR-based metabonomics to obtain an understanding of metabolic dysregulation driven by common KRAS mutations and also studied the effects of single copy mutant KRAS on the cellular metabolic profile. Specifically, we compared the metabolic profiles of parental HCT116 KRAS G13D/+ (a cell line isolated from a patient with colorectal cancer in which glycine (G) 13 is mutated to aspartate (D)) and its isogenic derivative cell lines: KRAS +/– and KRAS G13D/–. In the cell line KRAS +/–, the mutated KRAS allele is deleted and the WT allele remains, whereas in KRAS G13D/– the WT KRAS allele is deleted and the G13D mutant allele remains. These isogenic cell lines share the same genetic background, differing only by the mutational status of the KRAS gene and therefore allowed study of the metabolic functions of normal or mutant KRAS proteins and in particular, allowed us to determine the effects of the loss of an allele of the cell line metabolic profile, something that had not been possible in our previous studies (Varshavi et al., 2020).
2 Experimental
2.1 Cell culture and harvesting method
HCT116 colorectal carcinoma isogenic cell lines were obtained from Horizon Discovery Ltd (Cambridge, UK) and maintained at 37 °C with 5% CO2. Cell lines were generated using the Recombinant Adeno-Associated Virus (rAAV) based gene editing platform and were all validated by genotyping—gDNA and cDNA. The study was conducted in vitro with the cells transiently growing as spheroids in low adherence flasks, cultured and harvested as recently described using five biological replicates (Varshavi et al., 2020).
2.2 Metabolite extraction of HCT116 cell lines
Intracellular metabolites were extracted as previously described with some modifications (Dettmer et al., 2011). 800 μl of chilled MeOH/H2O (− 20 °C, 80:20(v/v)) was added to the frozen cell pellet, homogenized for 1 min using Pellet pestles (Sigma-Aldrich, UK). The sample was centrifuged at 4 °C and 7000×g for 5 min and the supernatant was collected into a new Eppendorf tube. The pellet was re-extracted three times with 400 μl cold methanol/water (− 20 °C, 80:20 (v/v)) to produce 2 ml of extract. The supernatants were combined, dried under a gentle flow of N2 gas and stored at -80° C until dissolution for NMR analysis.
2.3 Cell extract sample preparation and NMR acquisition
Samples were prepared, acquired, processed and analysed as described previously (Benjamini, 2010; Varshavi et al., 2018, 2020; Veselkov et al., 2014). Two-dimensional NMR experiments were carried out for selected samples to aid/confirm the identities of the metabolites (Dona et al., 2016). Details for HCT116 KRAS G13D/− are in Supplementary Table 1. Metabolites were identified at MSI level 2 (Sumner et al., 2007) and as previously described and using published methods (Dona et al., 2016; Everett, 2015; Varshavi et al., 2020) (Table 1, Supplementary Table 2).
2.4 Multivariate analysis of HCT116 cell line extracts
PCA was conducted on all samples to establish any obvious outliers in the data and to determine any between group differences. In order to maximize the separation between classes, NMR data were also subject to a supervised dimension-reduction technique, maximum margin criterion (MMC, Supplementary Fig. 1) (Veselkov et al., 2014). Leave-one-out cross-validation with the quadratic as a classifier was applied to check the validity of the MMC model (Supplementary Fig. 2). To determine metabolites responsible for the class separation, one-way ANOVA was applied with a false discovery rate (FDR) of 0.1 i.e. 10%, in order to account for multiple hypothesis testing.
3 Results
3.1 Metabonomics analysis of HCT116 cell line extracts expressing WT or G13D mutant KRAS
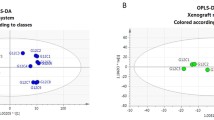
A representative 1H NMR spectrum from an HCT116 KRAS G13D/− extract is shown in Fig. 1 (a, b). PCA of the three genotypes showed high degrees of reproducibility among the biological replicates for each group. Moreover, PCA demonstrated a clear discrimination between all three genotypes: KRAS G13D/+, KRAS ± and KRAS G13D/− as shown in Fig. 2. Supervised maximum margin criterion (MMC) analysis gave correct classification rates of 100% for the models constructed between all classes.
a The aliphatic region of the 600 MHz 1H NMR spectrum from an HCT116 KRAS G13D/− extract. Numbers indicate signals corresponding to individual metabolites. 1. leucine; 2. valine; 3. lactate and threonine; 4. alanine; 5. acetate; 6. glutamate; 7. succinate; 8. glutamine; 9. glutathione; 10. aspartate; 11. asparagine; 12. creatine; 13. O-phosphocholine; 14. taurine; 15. myo-inositol; 16. glycine. b The aromatic region of the 600 MHz 1H NMR spectrum from an HCT116 KRAS G13D/− extract. Numbers indicate signals corresponding to individual metabolites. 18. uridine; 19. NAD; 20. tyrosine; 21. phenylalanine; 22. inosine; 23. ATP; 24. AMP
a PCA score plot of the 600 MHz 1H NMR spectra of extracts of HCT116 cells, b superimposed NMR spectra of cells, with the same colour coding as in the PCA plot; c ‘heat map display of the 600 MHz 1H NMR spectra from 0.8 to 10.0 ppm of KRAS G13D/− (top 5 strips), KRAS G13D/+middle 5 strips) and KRAS ± (bottom 5 strips). Red and blue elements in the spectra indicate NMR signals that are more intense, or less intense, respectively, than the median signal intensity for all the samples
Figure 3 and Supplementary Figs. 3 and 4 show representative ANOVAs built between genotypes (KRAS G13D/+ vs KRAS ±), (KRAS G13D/− vs KRAS ±) and (KRAS G13D/+ vs KRAS G13D/−) respectively. Table 1 and Supplementary Table 3 provide a list of statistically significant metabolites and their relative fold-changes between genotypes. To further understand the biological significance of the metabolite changes in the KRAS mutant clones, we used enrichment analysis (EA) tools in MetaboAnalyst to link metabolites to metabolic pathways (Fig. 4). In both KRAS G13D/− and KRAS G13D/+ cells, ammonia recycling, glutathione metabolism, glycine and serine metabolism pathways and glutamate metabolism were over-represented.
At bottom a ‘heat map display of the 600 MHz 1H NMR spectra of KRAS G13D/+ (top 5 strips) vs the corresponding spectra of KRAS± (bottom 5 strips). Red and blue elements in the spectra indicate NMR signals that are more intense, or less intense, respectively, than the median signal intensity for all the samples. At top, the corresponding ANOVA plot, showing positive peaks for those metabolite signals that are more intense in KRAS G13D/+, and negative peaks for those metabolite signals that are less intense. The signals are colour coded by the p value adjusted for FDR of 0.1
Metabolic pathway analyses related to the metabolites that were discriminating between a KRAS G13D/+ and KRAS±, b KRAS G13D/− and KRAS± and c KRAS G13D/+ and KRAS G13D/−. The horizontal bars summarize the main metabolite sets identified in this analysis; the bars are coloured based on their p values and the length is based on the fold enrichment
KRAS G13D/+ and KRAS G13D/− cells showed elevated levels of valine, isoleucine, leucine, threonine, alanine, proline, glutamate, glutathione, aspartate, asparagine, creatine, taurine and serine relative to HCT116 cell lines harbouring only one WT KRAS allele (KRAS ±).
Compared to KRAS ± cells, KRAS G13D/− cells also exhibited higher levels of lactate, glutamine, succinate, choline, betaine, inosine and adenosine 5′-monophosphate (AMP) and lower levels of acetate, creatine phosphate and adenosine 5′-triphosphate ATP while KRAS G13D/+ cells displayed increased levels of glycine and ATP and decreased levels of myo-inositol.
KRAS G13D/− cells showed higher levels of lactate, succinate, choline, betaine, glutamine, glutathione, creatine and inosine, and lower levels of serine, threonine, glycine, acetate, creatine phosphate, phosphocholine, myo-inositol and ATP relative to KRAS G13D/+ cells.
4 Discussion
Altered metabolism has been recently recognized as one of the hallmarks of cancer cells, allowing metabonomics to be widely used as a powerful tool to identify diagnostic cancer biomarkers and new therapeutic targets (Spratlin et al., 2009; Varshavi et al., 2020; Vermeersch & Styczynski, 2013). Oncogenic KRAS is one of the genes that is frequently mutated in colorectal cancer, and mutant KRAS is associated with poor prognosis and resistance to therapeutics (Mayers et al., 2016). About 80% of KRAS mutations are heterozygous (Hartman et al., 2012). However, in many types of cancer the remaining wild-type KRAS gene can be lost during tumour progression. Therefore, wild-type KRAS might serve as a tumour-suppressor gene in the presence of mutant KRAS (Luo et al., 2014; To et al., 2013; Zhang et al., 2001; Zhongqiu Hartman et al., 2012). Mutant KRAS is a known driver mutation for cancer development and progression, making KRAS mutant proteins good candidates for therapeutic development. Yet, until recently, KRAS mutants have proved difficult to target with small molecules, fuelling interest in targets upstream and downstream of KRAS for drug discovery. Understanding the metabolic consequences of WT KRAS allele loss might provide additional information in identifying key vulnerabilities in tumours with mutant KRAS.
In the current study, we applied an exploratory, untargeted, NMR-based metabonomics approach to obtain an understanding of metabolic dysregulation driven by a common human KRAS oncogenic mutation, G13D. The use of isogenic cell lines in this study also afforded us window into the effects of single copy mutant KRAS on metabolic alteration.
To this end, we compared the metabolic profiles of the parental KRAS G13D/+ (parental line isolated from a colorectal cancer patient in which glycine (G) 13 is mutated to aspartate (D)) and its isogenic derivative cell lines KRAS +/– and KRAS G13D/– in human colon cancer cells (HCT116). In the cell line KRAS +/–, the mutated KRAS allele is deleted and the WT one remains, whereas in KRAS G13D/– the WT KRAS allele is deleted and the G13D mutated one remains.
PCA analysis (Fig. 2) showed a clear discrimination not only between mutant-containing (KRAS G13D/+, KRAS G13D/–) and wild-type-containing (KRAS +/–) KRAS, but also between KRAS G13D/+ and KRAS G13D/–. Interestingly, parental cell lines (KRAS G13D/+) clustered more closely to mutant KRAS-deleted lines (KRAS +/–) than to WT KRAS-deleted lines (KRAS G13D/–), indicating that in each isogenic cell line the WT KRAS allele has a stronger effect on the overall metabolic profile than the mutant allele. The main alterations associated with mutant KRAS involved changes in multiple metabolic pathways such as glycolysis, TCA, and most significantly amino acid pathways (Fig. 5).
An overview of metabolic pathways altered in HCT116 cell line expressing the WT or the G13D mutant KRAS. Yellow boxes represent deregulated metabolites. The direction of the arrows shows the regulation direction of metabolites; the red arrows represent metabolites in KRAS G13D/− compared to KRAS ±; the green arrows represent metabolites in KRAS G13D/+ compared to KRAS ±, the blue arrows represent metabolites in KRAS G13D/− compared to KRAS G13D/+.  indicates that the metabolite is only present or only absent in KRAS G13D/− compared to KRAS G13D/+ respectively.
indicates that the metabolite is only present or only absent in KRAS G13D/− compared to KRAS G13D/+ respectively.  indicates that the metabolite is only present and absent in KRAS G13D/− compared to KRAS ± respectively: 3PG (3-phosphoglycerate), AMP (Adenosine monophosphate), ADP (Adenosine diphosphate), ATP (Adenosine triphosphate), α- KG (α-ketoglutarate), DMG (Dimethylglycine), GSH (Glutathione), GTP (Guanosine-5'-triphosphate), HIF-1α (Hypoxia-inducible factor 1-alpha), NADP (Nicotinamide adenine dinucleotide phosphate), NADPH (Nicotinamide adenine dinucleotide phosphate), PCho (Phosphocholine), PHD (Prolyl-hydroxylase), ROS (Reactive Oxygen Species), UDP-GlcNAc (Uridine diphosphate N-acetylglucosamine)
indicates that the metabolite is only present and absent in KRAS G13D/− compared to KRAS ± respectively: 3PG (3-phosphoglycerate), AMP (Adenosine monophosphate), ADP (Adenosine diphosphate), ATP (Adenosine triphosphate), α- KG (α-ketoglutarate), DMG (Dimethylglycine), GSH (Glutathione), GTP (Guanosine-5'-triphosphate), HIF-1α (Hypoxia-inducible factor 1-alpha), NADP (Nicotinamide adenine dinucleotide phosphate), NADPH (Nicotinamide adenine dinucleotide phosphate), PCho (Phosphocholine), PHD (Prolyl-hydroxylase), ROS (Reactive Oxygen Species), UDP-GlcNAc (Uridine diphosphate N-acetylglucosamine)
The amino acids most commonly consumed at a high rate by cancer cells are glutamine and serine (Tsun & Possemato, 2015). The serine levels in HCT116 KRAS G13D/+ or KRAS G13D/– cells were significantly higher compared with isogenic HCT116 cells containing only a wild-type KRAS allele (KRAS +/–).
Serine is a non-essential amino acid that contributes to a range of metabolic processes, which are vital for the growth and survival of cancer cells, including protein, lipid, nucleotide and glutathione biosynthesis, as well as one-carbon metabolism (Amelio et al., 2014; Yang & Vousden, 2016). Serine can be either taken up from the medium by the transformed cell or be produced from the glycolytic intermediate 3-phosphoglycerate (3PG). The first step in the serine synthesis pathway (SSP) involves the conversion of 3-phosphoglycerate to 3-phosphohydroxypyruvate (3PHP) by the NAD+-dependent enzyme 3-phosphoglycerate dehydrogenase (PHGDH). 3-Hydroxypyruvate can be then transaminated to phosphoserine by the enzyme phosphoserine aminotransferase (PSAT1), using glutamate as amino donor (Amelio et al., 2014; Yang & Vousden, 2016). Serine is subsequently produced from phosphoserine by phosphoserine phosphatase (PSPH). PHGDH, the initial enzyme in the SSP, is reportedly upregulated in breast cancers and melanomas (Labuschagne et al., 2014; Possemato et al., 2011). However, many cancer cells are largely dependent on uptake of exogenous serine, as de novo serine synthesis alone cannot support their proliferation and survival (Jain et al., 2012; Maddocks et al., 2013). The enhanced concentrations of serine in KRAS G13D/– and KRAS G13D/+ cells relative to KRAS +/– cells could, therefore, be due to an increase in serine uptake from the medium and/or biosynthesis. Interestingly, lower levels of serine were observed in KRAS G13D/– versus KRAS G13D/+ cells. This could be associated with either increased consumption or decreased uptake/biosynthesis. The latter is more likely, as serine is involved in adenine synthesis, and its limited availability can result in a decreased concentration of ATP in the cell. This is consistent with our observation, showing lower levels of ATP in KRAS G13D/– cells versus KRAS G13D/+ and KRAS +/– cells. This hypothesis could be tested further by comparison of the serine concentration in the culture media across the cell lines.
Concentrations of glutamine in KRAS G13D/+ and KRAS +/– cells were comparable but increased in KRAS G13D/– cells relative to KRAS +/– and KRAS G13D/+ cells. The increased levels of glutamine in KRAS G13D/– cells could indicate that the WT KRAS-deleted isogenic cell lines are more dependent on glutamine and hence they take it up at a high rate.
Glutamine is a “conditionally essential” amino acid that is synthesized sufficiently under normal circumstances but becomes essential during proliferation. KRAS-mutant cells can be particularly dependent on glutamine as a carbon source to fuel the TCA cycle and as a nitrogen source for nucleotide, hexosamine and nonessential amino acid (NEAA) biosynthesis (Toda et al., 2016). Glutamine is transported into the cell through one of many transporters, such as the neutral amino acid transporter (ASCT2) (Scopelliti et al., 2018). Once imported into the cells, glutamine is converted to glutamate by glutaminase (GLS) in the mitochondria (Chendong Yang et al., 2014). In this study, higher intracellular concentrations of glutamate were observed in KRAS mutant cells (KRAS G13D/+, KRAS G13D/–) relative to wild-type-containing cells (KRAS +/–), indicating increased glutaminolysis.
Glutamate can subsequently be converted to α-ketoglutarate by glutamate dehydrogenase 1 (GLUD1) where it can fuel the TCA cycle or be converted to nonessential amino acids such as aspartate and alanine by transaminases, including glutamate–oxaloacetate transaminase (GOT) and glutamate–pyruvate transaminase (GPT), respectively. Glutamate can also contribute to the synthesis of proline and glutathione (Liu et al., 2012).
In the current study, KRAS G13D/+ and KRAS G13D/– cells showed increased levels of metabolites associated with the glutamate synthase cycle including alanine, aspartate, asparagine, proline and glutathione when compared to KRAS +/– cells.
Elevated levels of aspartate and asparagine in mutant KRAS cells relative to KRAS +/– is consistent with other studies reporting that oncogenic KRAS shifts glutamine metabolism toward aspartate synthesis (Ahn & Metallo, 2015; Lyssiotis et al., 2013; Son et al., 2013b). Aspartate is synthesised by transferring the α-amino group on glutamate to the α-keto group of oxaloacetate. This metabolic alteration is thought to have an important role in regeneration of NADPH and NAD to maintaining redox balance in addition to glycolysis (Son et al., 2013a). Additionally, aspartate and glutamine are the precursors for asparagine. Asparagine is involved in the regulation of the cellular adaptation to depletion of glutamine and other nonessential amino acids, as well as suppression of apoptosis induced by glutamine depletion (Zhang et al., 2014).
Glutathione (GSH) levels were also increased in mutant KRAS cells compared to their WT, isogenic counterpart (KRAS +/–). A higher level of glutathione was also evident in KRAS G13D/– relative to KRAS G13D/+ cells. In agreement with our findings, a recent study (Kerr et al., 2016) found that KRAS mutation and copy number gain is associated with up-regulation of glutathione metabolism. Glutathione is a tripeptide (consisting of glutamate, cysteine and glycine) that is involved in cellular protection against reactive oxygen species (ROS) and reactive nitrogen (RNS) species (Aquilano et al., 2014). GSH deregulation has been reported in many human cancers (Calvert et al., 1998; Estrela et al., 2006; Obrador et al., 2002), possibly because GSH is protective against oxidative damage (Ortega et al., 2010), stress-induced apoptosis (Franco et al., 2009) and multidrug and radiation resistance (Arrick & Nathan, 1984; Estrela et al., 1995; Meister, 1991; Mitchell & Russo, 1987).
KRAS G13D/+ and KRAS G13D/– cells also exhibited higher concentrations of proline relative to KRAS +/– cells. Proline is an α-amino acid that is derived from glutamate via the intermediate pyrroline-5-carboxylate (P5C). P5C is subsequently converted to proline by the NAD (P) H-dependent enzyme pyrroline-5-carboxylate reductase (PYCR), which is present in three isoforms: PYCR1, PYCR2 and PYCRL. Up-regulation of mitochondrial PYCR1 and PYCR2 is observed in a number of cancer types, such as prostate cancer and lymphoma (Ahn & Metallo, 2015; De Ingeniis et al., 2012; Ernst et al., 2002). In addition to biosynthesis, proline catabolism is reportedly down-regulated in a number of tumour types. The first step of proline catabolism is catalysed by proline oxidase, which is proposed as a novel mitochondrial tumour suppressor in human cancers (Liu et al., 2012). In their paper, Liu and colleagues propose proline metabolism as a crucial link in the reprogramming of glutamine and glucose metabolism during tumorigenesis. Therefore, the enzymes involved in proline biosynthesis could provide novel targets for cancer therapy.
In our study, we have observed metabolic variation not only between KRAS mutant (KRAS G13D/+, KRAS G13D/–) cells and wild-type (KRAS +/–) cells, but also between KRAS G13D/+ and KRAS G13D/– cells. Interestingly, KRAS G13D/– showed higher levels of succinate relative to both KRAS +/– and KRAS G13D/+. Accumulation of succinate is associated with the inhibition of prolyl hydroxylases. One of the consequences of this inhibition is the stabilization of the transcription factor hypoxia-inducible factor 1-alpha (HIF-1α) and the generation of a pseudo-hypoxic response reducing oxidative phosphorylation and promoting glycolysis (Selak et al. 2005; Tavares et al., 2015). Stabilisation of HIF1α results in the increased expression of lactate dehydrogenase A (LDHA), which increases the conversion of pyruvate to lactate, and the expression of pyruvate dehydrogenase kinase 1 (PDK1), which inhibits pyruvate dehydrogenase (PDH) and decreases mitochondrial oxidation of pyruvate to acetyl coenzyme A (acetyl-CoA) (Kim et al., 2006; Schulze & Harris, 2012; Zhao et al., 2014). One hypothesis is that under metabolic stress (hypoxia or nutrient deprivation), acetyl-CoA production can be compensated by acetyl-CoA synthase 2 (ACCS2), which converts acetate to acetyl-CoA using ATP to drive the net reaction (Kamphorst et al., 2014; Schug et al., 2015). Our results showing increased levels of succinate, lactate and AMP and decreased concentrations of acetate in KRAS G13D/– cells relative to KRAS +/– and KRAS G13D/+ cells support these observations and indicate that the KRAS G13D mutation could impact HIF1α function in the absence of the WT KRAS allele.
In addition, as HIF-1α can also actively repress mitochondrial aerobic metabolism through PDK1 (Papandreou et al., 2006), interruption of the TCA cycle could explain lower levels of ATP in KRAS G13D/− cells compared with KRAS G13D/+ and KRAS ± cells. Interestingly, we also observed increased levels of creatine and depletion of phosphocreatine in KRAS G13D/− cells relative to KRAS G13D/+ and KRAS ± cells. This could indicate that in low ATP states, phosphocreatine can be used as a rapidly mobilisable reserve of high-energy phosphates to sustain the energetic requirements of cancer cells.
5 Conclusion
Overall, our data provide further insights into nutrient use and the regulation of metabolic pathways driven by oncogenic KRAS. In this study, we have used a robust isogenic system to study the role of a single point mutation in one KRAS allele in an isogenic genetic background, hence avoiding confounding genetic factors.
We have demonstrated that the KRASG13D oncogene is associated with general metabolic reprogramming to support growth and counter stress, including alterations in the metabolism of amino acids and enhanced glutathione biosynthesis. Additionally, we show that KRASG13D/+ and KRASG13D/− cells have distinct metabolic profiles characterized by dysregulation of and changes in the TCA cycle, the glycolysis and glutathione metabolism pathways, as well as in glutamine, acetate and phosphocreatine. The metabolic adaptation driven by the loss of the KRAS WT allele that we have initially described in this paper could contribute to differential responses to anticancer treatments and/or reveal pathways that might be exploited for therapeutic intervention.
Data availability
Original NMR data will be deposited in MetaboLights, EBI, Cambridge UK following publication and can be accessed via the code MTBLS3783 at the following url: https://www.ebi.ac.uk/metabolights/.
References
Ahn, C. S., & Metallo, C. M. (2015). Mitochondria as biosynthetic factories for cancer proliferation. Cancer & Metabolism, 3(1), 1.
Amelio, I., Cutruzzolá, F., Antonov, A., Agostini, M., & Melino, G. (2014). Serine and glycine metabolism in cancer(). Trends in Biochemical Sciences, 39(4), 191–198.
Aquilano, K., Baldelli, S., & Ciriolo, M. R. (2014). Glutathione: New roles in redox signaling for an old antioxidant. Frontiers in Pharmacology, 5, 196.
Arrick, B. A., & Nathan, C. F. (1984). Glutathione metabolism as a determinant of therapeutic efficacy: A review. Cancer Research, 44(10), 4224.
Arrington, A. K., Heinrich, E. L., Lee, W., Duldulao, M., Patel, S., Sanchez, J., Garcia-Aguilar, J., & Kim, J. (2012). Prognostic and predictive roles of KRAS mutation in colorectal cancer. International Journal of Molecular Sciences, 13(10), 12153–12168.
Benjamini, Y. (2010). Discovering the false discovery rate. Journal of the Royal Statistical Society: Series B (statistical Methodology), 72(4), 405–416.
Calvert, P., Yao, K.-S., Hamilton, T. C., & O’Dwyer, P. J. (1998). Clinical studies of reversal of drug resistance based on glutathione. Chemico-Biological Interactions, 111, 213–224.
De Ingeniis, J., Ratnikov, B., Richardson, A. D., Scott, D. A., Aza-Blanc, P., De, S. K., Kazanov, M., Pellecchia, M., Ronai, Z. E., Osterman, A. L., & Smith, J. W. (2012). Functional specialization in proline biosynthesis of melanoma. PLoS ONE, 7(9), e45190.
Dettmer, K., Nürnberger, N., Kaspar, H., Gruber, M. A., Almstetter, M. F., & Oefner, P. J. (2011). Metabolite extraction from adherently growing mammalian cells for metabolomics studies: Optimization of harvesting and extraction protocols. Analytical and Bioanalytical Chemistry, 399(3), 1127–1139.
Dinu, D., Dobre, M., Panaitescu, E., Bîrlă, R., Iosif, C., Hoara, P., Caragui, A., Boeriu, M., Constantinoiu, S., & Ardeleanu, C. (2014). Prognostic significance of KRAS gene mutations in colorectal cancer—preliminary study. Journal of Medicine and Life, 7(4), 581–587.
Dona, A. C., Kyriakides, M., Scott, F., Shephard, E. A., Varshavi, D., Veselkov, K., & Everett, J. R. (2016). A guide to the identification of metabolites in NMR-based metabonomics/metabolomics experiments. Computational and Structural Biotechnology Journal, 14, 135–153.
Ernst, T., Hergenhahn, M., Kenzelmann, M., Cohen, C. D., Bonrouhi, M., Weninger, A., Klaren, R., Grone, E. F., Wiesel, M., Gudemann, C., Kuster, J., Schott, W., Staehler, G., Kretzler, M., Hollstein, M., & Grone, H. J. (2002). Decrease and gain of gene expression are equally discriminatory markers for prostate carcinoma: A gene expression analysis on total and microdissected prostate tissue. American Journal of Pathology, 160(6), 2169–2180.
Estrela, J. M., Obrador, E., Navarro, J., De La Vega, M. C. L., & Pellicer, J. A. (1995). Elimination of Ehrlich tumours by ATP-induced growth inhibition, glutathione depletion and X-rays. Nature Medicine, 1(1), 84–88.
Estrela, J. M., Ortega, A., & Obrador, E. (2006). Glutathione in cancer biology and therapy. Critical Reviews in Clinical Laboratory Sciences, 43(2), 143–181.
Everett, J. R. (2015). A new paradigm for known metabolite identification in metabonomics/metabolomics: metabolite identification efficiency. Computational and Structural Biotechnology Journal, 13, 131–144.
Everett, J. R., Holmes, E., Veselkov, K. A., Lindon, J. C., & Nicholson, J. K. (2019). A unified conceptual framework for metabolic phenotyping in diagnosis and prognosis. Trends in Pharmacological Sciences, 40(10), 763–773.
Franco, R., Sánchez-Olea, R., Reyes-Reyes, E. M., & Panayiotidis, M. I. (2009). Environmental toxicity, oxidative stress and apoptosis: Ménage à Trois. Mutation Research/genetic Toxicology and Environmental Mutagenesis, 674(1–2), 3–22.
Hartman, D. J., Davison, J. M., Foxwell, T. J., Nikiforova, M. N., & Chiosea, S. I. (2012). Mutant allele-specific imbalance modulates prognostic impact of KRAS mutations in colorectal adenocarcinoma and is associated with worse overall survival. International Journal of Cancer, 131(8), 1810–1817.
Jain, M., Nilsson, R., Sharma, S., Madhusudhan, N., Kitami, T., Souza, A. L., Kafri, R., Kirschner, M. W., Clish, C. B., & Mootha, V. K. (2012). Metabolite profiling identifies a key role for glycine in rapid cancer cell proliferation. Science, 336(6084), 1040–1044.
Kamphorst, J. J., Chung, M. K., Fan, J., & Rabinowitz, J. D. (2014). Quantitative analysis of acetyl-CoA production in hypoxic cancer cells reveals substantial contribution from acetate. Cancer & Metabolism, 2(1), 23.
Kerr, E., Gaude, E., Turrell, F., Frezza, C., & Martins, C. P. (2016). Mutant Kras copy number defines metabolic reprogramming and therapeutic susceptibilities. Nature, 531(7592), 110–113.
Kim, J.-W., Tchernyshyov, I., Semenza, G. L., & Dang, C. V. (2006). HIF-1-mediated expression of pyruvate dehydrogenase kinase: A metabolic switch required for cellular adaptation to hypoxia. Cell Metabolism, 3(3), 177–185.
Labuschagne, C. F., van den Broek, N. J. F., Mackay, G. M., Vousden, K. H., & Maddocks, O. D. K. (2014). Serine, but not glycine, supports one-carbon metabolism and proliferation of cancer cells. Cell Reports, 7(4), 1248–1258.
Lindon, J., Nicholson, J., Holmes, E., & Everett, J. (2000). Metabonomics: Metabolic processes studied by NMR spectroscopy of biofluids. Concepts in Magnetic Resonance, 12(5), 289–320.
Liu, W., Le, A., Hancock, C., Lane, A. N., Dang, C. V., Fan, T. W. M., & Phang, J. M. (2012). Reprogramming of proline and glutamine metabolism contributes to the proliferative and metabolic responses regulated by oncogenic transcription factor c-MYC. Proceedings of the National Academy of Sciences, 109(23), 8983–8988.
Luo, F., Poulogiannis, G., Ye, H., Hamoudi, R., Dong, G., Zhang, W., Ibrahim, A. E. K., & Arends, M. J. (2014). Wild-type K-ras has a tumour suppressor effect on carcinogen-induced murine colorectal adenoma formation. International Journal of Experimental Pathology, 95(1), 8–15.
Lyssiotis, C. A., Son, J., Cantley, L. C., & Kimmelman, A. C. (2013). Pancreatic cancers rely on a novel glutamine metabolism pathway to maintain redox balance. Cell Cycle, 12(13), 1987–1988.
Maddocks, O. D. K., Berkers, C. R., Mason, S. M., Zheng, L., Blyth, K., Gottlieb, E., & Vousden, K. H. (2013). Serine starvation induces stress and p53-dependent metabolic remodelling in cancer cells. Nature, 493(7433), 542–546.
Mayers, J. R., Torrence, M. E., Danai, L. V., Papagiannakopoulos, T., Davidson, S. M., Bauer, M. R., Lau, A. N., Ji, B. W., Dixit, P. D., Hosios, A. M., Muir, A., Chin, C. R., Freinkman, E., Jacks, T., Wolpin, B. M., Vitkup, D., & Vander Heiden, M. G. (2016). Tissue of origin dictates branched-chain amino acid metabolism in mutant Kras-driven cancers. Science, 353(6304), 1161–1165.
Meister, A. (1991). Glutathione deficiency produced by inhibition of its synthesis, and its reversal; applications in research and therapy. Pharmacology & Therapeutics, 51(2), 155–194.
Mitchell, J. B., & Russo, A. (1987). The role of glutathione in radiation and drug induced cytotoxicity. The British Journal of Cancer. Supplement, 8, 96–104.
Obrador, E., Carretero, J., Ortega, A., Medina, I., Rodilla, V., Pellicer, J. A., & Estrela, J. M. (2002). γ-Glutamyl transpeptidase overexpression increases metastatic growth of B16 melanoma cells in the mouse liver. Hepatology, 35(1), 74–81.
Ortega, Á. L., Mena, S., & Estrela, J. M. (2010). Oxidative and nitrosative stress in the metastatic microenvironment. Cancers, 2(2), 274–304.
Papandreou, I., Cairns, R. A., Fontana, L., Lim, A. L., & Denko, N. C. (2006). HIF-1 mediates adaptation to hypoxia by actively downregulating mitochondrial oxygen consumption. Cell Metabolism, 3, 187–197.
Possemato, R., Marks, K. M., Shaul, Y. D., Pacold, M. E., Kim, D., Birsoy, K., Sethumadhavan, S., Woo, H.-K., Jang, H. G., Jha, A. K., Chen, W. W., Barrett, F. G., Stransky, N., Tsun, Z.-Y., Cowley, G. S., Barretina, J., Kalaany, N. Y., Hsu, P. P., Ottina, K., … Sabatini, D. M. (2011). Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature, 476(7360), 346–350.
Schug, Z. T., Peck, B., Jones, D. T., Zhang, Q., Grosskurth, S., Alam, I. S., Goodwin, L. M., Smethurst, E., Mason, S., Blyth, K., McGarry, L., James, D., Shanks, E., Kalna, G., Saunders, R. E., Jiang, M., Howell, M., Lassailly, F., Thin, M. Z., … Gottlieb, E. (2015). Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell, 27(1), 57–71.
Schulze, A., & Harris, A. L. (2012). How cancer metabolism is tuned for proliferation and vulnerable to disruption. Nature, 491(7424), 364–373.
Scopelliti, A. J., Font, J., Vandenberg, R. J., Boudker, O., & Ryan, R. M. (2018). Structural characterisation reveals insights into substrate recognition by the glutamine transporter ASCT2/SLC1A5. Nature Communications, 9, 38.
Selak, M. A., Armour, S. M., MacKenzie, E. D., Boulahbel, H., Watson, D. G., Mansfield, K. D., Pan, Y., Simon, M. C., Thompson, C. B., & Gottlieb, E. (2005). Succinate links TCA cycle dysfunction to oncogenesis by inhibiting HIF-α prolyl hydroxylase. Cancer Cell, 7(1), 77–85.
Son, J., Lyssiotis, C. A., Ying, H., Wang, X., Hua, S., & Ligorio, M. (2013a). Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature, 496, 101–105.
Son, J., Lyssiotis, C. A., Ying, H., Wang, X., Hua, S., Ligorio, M., Perera, R. M., Ferrone, C. R., Mullarky, E., Shyh-Chang, N., Kang, Y. A., Fleming, J. B., Bardeesy, N., Asara, J. M., Haigis, M. C., DePinho, R. A., Cantley, L. C., & Kimmelman, A. C. (2013b). Glutamine supports pancreatic cancer growth through a Kras-regulated metabolic pathway. Nature, 496(7443), 101–105.
Spratlin, J. L., Serkova, N. J., & Eckhardt, S. G. (2009). Clinical applications of metabolomics in oncology: A review. Clinical Cancer Research, 15(2), 431–440.
Sumner, L. W., Amberg, A., Barrett, D., Beale, M. H., Beger, R., Daykin, C. A., Fan, T. W. M., Fiehn, O., Goodacre, R., Griffin, J. L., Hankemeier, T., Hardy, N., Harnly, J., Higashi, R., Kopka, J., Lane, A. N., Lindon, J. C., Marriott, P., Nicholls, A. W., … Viant, M. R. (2007). Proposed minimum reporting standards for chemical analysis. Metabolomics, 3(3), 211–221.
Tavares, L. C., Jarak, I., Nogueira, F. N., Oliveira, P. J., & Carvalho, R. A. (2015). Metabolic evaluations of cancer metabolism by NMR-based stable isotope tracer methodologies. European Journal of Clinical Investigation, 45, 37–43.
To, M. D., Del Rosario, R., Westcott, P. M. K., Banta, K. L., & Balmain, A. (2013). Interactions between wildtype and mutant ras genes in lung and skin carcinogenesis. Oncogene, 32(34), 4028–4033.
Toda, K., Kawada, K., Iwamoto, M., Inamoto, S., Sasazuki, T., Shirasawa, S., Hasegawa, S., & Sakai, Y. (2016). Metabolic alterations caused by KRAS mutations in colorectal cancer contribute to cell adaptation to glutamine depletion by upregulation of asparagine synthetase()(). Neoplasia, 18(11), 654–665.
Tsuchida, N., Murugan, A. K., & Grieco, M. (2016). Kirsten Ras oncogene: significance of its discovery in human cancer research. Oncotarget, 7(29), 46717–46733.
Tsun, Z.-Y., & Possemato, R. (2015). Amino acid management in cancer. Seminars in Cell & Developmental Biology, 43, 22–32.
Varshavi, D., McCarthy, N., Veselkov, K., Keun, H. C., & Everett, J. R. (2020). Metabolic characterization of colorectal cancer cells harbouring different KRAS mutations in codon 12, 13, 61 and 146 using human SW48 isogenic cell lines. Metabolomics, 16(4), 51.
Varshavi, D., Scott, F. H., Varshavi, D., Veeravalli, S., Phillips, I. R., Veselkov, K., Strittmatter, N., Takats, Z., Shephard, E. A., & Everett, J. R. (2018). Metabolic biomarkers of ageing in C57BL/6J wild-type and Flavin-containing monooxygenase 5 (FMO5)-knockout mice. Frontiers in Molecular Biosciences, 5, 28.
Vartanian, S., Bentley, C., Brauer, M. J., Li, L., Shirasawa, S., Sasazuki, T., Kim, J.-S., Haverty, P., Stawiski, E., Modrusan, Z., Waldman, T., & Stokoe, D. (2013). Identification of mutant K-Ras-dependent phenotypes using a panel of isogenic cell lines. The Journal of Biological Chemistry, 288(4), 2403–2413.
Vermeersch, K. A., & Styczynski, M. P. (2013). Applications of metabolomics in cancer research. J Carcinog, 12, 9.
Veselkov, K. A., Mirnezami, R., Strittmatter, N., Goldin, R. D., Kinross, J., Speller, A. V. M., Abramov, T., Jones, E. A., Darzi, A., Holmes, E., Nicholson, J. K., & Takats, Z. (2014). Chemo-informatic strategy for imaging mass spectrometry-based hyperspectral profiling of lipid signatures in colorectal cancer. Proceedings of the National Academy of Sciences, 111(3), 1216–1221.
Yang, C., Ko, B., Hensley, C. T., Jiang, L., Wasti, A. T., Kim, J., Sudderth, J., Calvaruso, M. A., Lumata, L., Mitsche, M., Rutter, J., Merritt, M. E., & DeBerardinis, R. J. (2014). Glutamine oxidation maintains the TCA cycle and cell survival during impaired mitochondrial pyruvate transport. Molecular Cell, 56(3), 414–424.
Yang, M., & Vousden, K. H. (2016). Serine and one-carbon metabolism in cancer. Nature Reviews Cancer, 16(10), 650–662.
Zhang, J., Fan, J., Venneti, S., Cross, J. R., Takagi, T., & Bhinder, B. (2014). Asparagine plays a critical role in regulating cellular adaptation to glutamine depletion. Molecular Cell, 56, 205–218.
Zhang, Z., Wang, Y., Vikis, H. G., Johnson, L., Liu, G., Li, J., Anderson, M. W., Sills, R. C., Hong, H. L., Devereux, T. R., Jacks, T., Guan, K.-L., & You, M. (2001). Wildtype Kras2 can inhibit lung carcinogenesis in mice. Nature Genetics, 29(1), 25–33.
Zhao, D., Li, F.-L., Cheng, Z.-L., & Lei, Q.-Y. (2014). Impact of acetylation on tumor metabolism. Molecular & Cellular Oncology, 1(3), e963452.
Acknowledgements
We thank Professors Jeremy Nicholson and Elaine Holmes for access to NMR facilities at Imperial College London and we thank Dr. Beatriz Jimenez and her team there for technical assistance.
Funding
We thank the University of Greenwich and Horizon Discovery for funding.
Author information
Authors and Affiliations
Contributions
Conceptualizing, refining and funding research ideas: JRE and NMcC. Literature search: Dorna V. Creating research design: Dorna V, NMcC and JRE. Collection of data: Dorna V, Dorsa V, JRE. Data analysis: Dorna V, Dorsa V, JRE. Statistical software: KV and colleagues. Drafting manuscript: Dorna V, Dorsa V. Editing and approving manuscript: Dorna V, Dorsa V, NMcC, KV, HK, JRE.
Corresponding author
Ethics declarations
Conflict of interest
Nicola McCarthy is an employee of Horizon Discovery. All other authors declare that they have no conflict of interest.
Research involved in animal or human rights
No animals or human subjects were involved in this study, which complied with all University ethical and safety standards.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Varshavi, D., Varshavi, D., McCarthy, N. et al. Metabonomics study of the effects of single copy mutant KRAS in the presence or absence of WT allele using human HCT116 isogenic cell lines. Metabolomics 17, 104 (2021). https://doi.org/10.1007/s11306-021-01852-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-021-01852-w