Abstract
Cancer of unknown primary site (CUP) is defined as a heterogeneous group of tumors that appear as metastases, and of which standard diagnostic work-up fails to identify the origin. It is considered a separate entity with a specific biology, and nowadays molecular characteristics and the determination of actionable mutations may be important in a significant group of patients. In this guide, we summarize the diagnostic, therapeutic, and possible new developments in molecular medicine that may help us in the management of this unique disease entity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer of unknown origin (CUP) represents a heterogeneous group of tumors with a histologically confirmed diagnosis of cancer, which originates as metastatic disease, and of which, after a systematic search for the primary tumor, including at least clinical history, physical examination, blood tests and biochemistry, along with computed thoraco-abdominal tomography (CT), the origin of the primary tumor is not found.
Currently CUP represents between 3–5% of diagnosed tumors, being approximately the tenth most frequent tumor in incidence.
Given the poor prognosis of cancer of unknown origin, it is necessary to optimize diagnosis and knowledge of the potential molecular pathways involved, in order to establish therapeutic strategies, and clinical trials are currently underway to explore these pathways.
Metodology
This guideline has been developed with the consensus of ten oncologists highly experienced in the diagnosis and treatment of CUP, and members of the Spanish Research Group on Cancer of Unknown Origin (GECOD) and the Spanish Society of Medical Oncology (SEOM).
In order to assign the levels of evidence and the grade of recommendation for the different states of this treatment guideline, The Infectious Diseases Society of America-US Public Health Service Grading System for ranking Recommendations in Clinical Guidelines have been followed [1].
Epidemiology. biological background and proportional distribution according to occult primary site
CUP accounted for 3–5% of all diagnosed cancer in the historical series but in recent publications its incidence seems to decline. The CUP incidence is influenced by the non-existence of international consensus on the definition, classifications and registration of CUP. There is a lack of a diagnostic process record for these patients [2].
There is no gender difference, and the average age of presentation is 60. Among individuals older than 60 years of age, the incidence-rate for CUP in the digestive tract has increased markedly [2].
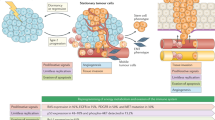
Little is understood about the disease pathogenesis and biology. Two predominant theories exist:
-
Parallel progression model: CUP tumors are metastatic tumors that have arisen from an undetectable or regressed primary lesion. The dissemination of primary tumor cells is an early event where subsequent clonal evolution of the metastasis is distinctly different from that of the primary tumor.
-
No primary tumor exists: CUP is a single metastatic entity. Metastasis occurs without parallel progression, with the tumor microenvironment selectively favoring the outgrowth of tumor cells at the metastatic site, while it avoids the growth of these genetically identical cells at the primary site [3].
The process that generates CUP is driven by multiple interdependent alterations in cell behavior, including chromosomal alterations, self-sufficiency in growth signals, resistance to growth-inhibitory signals, reprogramming of energy metabolism, evasion of apoptosis, limitless replicative potential, sustained angiogenesis, tissue invasion and metastasis and evasion from immune destruction [4].
Prognosis. Subsets of patients. Prognostic groups
The prognosis of patients with CUP is generally poor, since by definition they are aggressive cancers that are metastatic at their onset. However, a correct diagnosis can identify a subgroup of patients around 20% with a more favorable prognosis, in whom a greater benefit can be expected when treated with a suitable treatment.
This group of patients (Table 1), called favorable prognosis, should be treated similarly to known equivalent primary tumors with metastatic spread, and may achieve long-term control of metastatic spread in 30–60% of cases [5].
Unfortunately, most patients with CUP do not belong to these specific subgroups. Eighty percent of all patients diagnosed with CUP have a poor response to treatment and a poor prognosis, with a median overall survival of six months [6].
The prognosis of CUP is classified primarily by performance status and serum lactate dehydrogenase (LDH) level. Those with a good performance status (0–1) and a normal LDH value have a median overall survival of 12 months. Those with one or both prognostic factors have a median overall survival of only 4 months [7].
Other factors predictive of a poor outcome are: male sex, high comorbidity, age over 64 years, smoking history of more than 10 pack-years, weight loss, lymphopenia, low serum albumin concentrations and high alkaline phosphatase concentrations [8].
Diagnostic workup
The diagnostic process in patients with CUP seeks to identify subgroups that can benefit from a specific therapeutic procedure, avoiding prolonged, expensive diagnostic processes of scant therapeutic benefit for the patient [9]. The diagnosis workup is summarized in the Table 2.
Anamnesis and physical examination
The primary clinical work-up includes complete medical history with attention to toxic habits, medical and surgical history, previous diseases, or a family history of neoplasms. Physical examination must include head and neck and rectal examination, testes in males, and pelvic/gynecologic and breast examination in women [10].
Laboratory
These consist of complete blood count, liver and kidney function tests, electrolytes (including calcium), and LDH, since they represent important prognostic factors.
-
Serum tumor markers: serum tumor markers are often elevated in a non-specific manner in patients with CUP, consequently their measurement offers no diagnostic or prognostic assistance. Exceptions are determination of serum PSA in male patients with bone metastasis, so as to exclude occult metastatic prostate cancer, germ cell tumor markers (αFP, βHCG) in patients with midline disease, serum αFP in liver-dominant disease, so as to exclude hepatocellular cancer, and CA125 in women with peritoneal involvement.
Radiological examinations and complementary tests to identify the primary in cases of CUP include
Computed tomography (CT) thoraco-abdominal-pelvic is customary since, in addition to attempting to detect the primary, it serves as an extension study and can locate lesions that can be biopsied [11].
Mammography should be performed in cases of adenocarcinoma in women.
Positron emission tomography (PET) CT remains to be fully evaluated in large-scale prospective studies. Currently, as it is more effective in detecting additional metastases rather than the hidden primary, it should be used when radical therapy is contemplated for localized CUP: cervical head/neck nodes, axillary adenopathy and single metastatic lesions [12, 13].
Examinations to be excluded in the absence of symptoms that indicate otherwise
-
Laryngoscopy: useful in cases of cervical lymph node involvement.
-
Bronchoscopy: in cases of radiological findings such as hiliar or mediastinal lymph node involvement, and pulmonary symptoms.
-
Gastroscopy: if abdominal symptoms or positive fecal occult blood test.
-
Colonoscopy: if abdominal symptoms or positive fecal occult blood test, or biopsy with immunohistochemistry CK20 + /CK7 − /CDX2 + .
-
Testicular ultrasound: if retroperitoneal or mediastinal mass.
-
Gynecologic ultrasound: if pelvic or peritoneal metastases, CK7 + on the biopsy tissue.
-
Breast MRI: if adenocarcinoma with negative mammography and metastasis to axillary lymph nodes.
Histological diagnosis
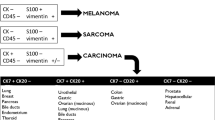
Pathological assessment of malignant tissue samples is crucial for CUP diagnosis. Core biopsy is preferred to fine-needle biopsy or cytology. Further procedures such as surgical biopsies may be considered when the initial sample is inadequate to confirm the diagnosis. Firstly, the pathological evaluation must rule out some special tumors with a specific therapeutic approach (lymphoma, germ-cell tumor, melanoma, or sarcoma) [14]. Thereafter the IHC staining may classify CUP into these five morphological subgroups [15]:
-
Well/moderately-differentiated adenocarcinoma (60%).
-
Poorly-differentiated adenocarcinoma/undifferentiated carcinoma (20–30%).
-
Squamous-cell carcinoma (5%).
-
Poorly-differentiated neoplasms (5%).
-
Neuroendocrine tumors (1–3%).
Immunohistochemistry
IHC plays an essential role in the evaluation of samples of metastatic tumors. It has a relatively low cost compared with other techniques, but it also has limitations. Nowadays, it would be important to reserve tumor material for molecular studies.
Keratins, expressed in epithelial cells, have historically been useful in confirming epithelial origin in poorly differentiated malignancies, although other tumor types may also express keratins. IHC should be applied for determining most likely cell lineage, and in order to exclude high chemo-sensitive and potentially curable tumors, and/or to rule out hormone-sensitive malignancies amenable to specific therapy. Staining for chromogranin A and synaptophysin is needed to profile neuroendocrine differentiation.
Among keratin family members, CK7 and CK20 have most widely been used to predict primary site. Although these expression patterns may be useful to prioritize one site of origin over another and to direct further workup, cases that do not fit these profiles are encountered frequently. In this sense, the CK7 positive and CK20 negative immunophenotype is the most common in CUP; however, this profile is not particularly useful for suggesting a specific anatomical site of origin [16].
The most common CK7 and CK20 profiles and other positive markers are shown in Fig. 1.
We propose a step-by-step algorithm to arrive at a CUP diagnosis (Table 3).
Molecular diagnosis
There are basically two types of strategies when considering the use of a molecular study platform in patients diagnosed with CUP: diagnostic molecular platforms aimed at identifying the primary tumor and Sequencing platforms for tumor mutation profile characterization.
Diagnostic molecular platforms aimed at identifying the primary tumor
These platforms base their results on the performance of a similarity score with the genetic or epigenetic characteristics of already known primary tumors. The procedure is based on the fact that, once the molecular profile of the tumor has been obtained, it is compared with the results of databases of cases of already known locations and histological types. The similarity of the molecular profile of the tumor evaluated with these patterns is assessed and a diagnosis is given that offers one (or several) locations, estimating the probabilities of each (similarity score).
There are several types of platforms with this approach: those that point to the genomic profile, which determines either the DNA gene expression or microRNA profile; and those that point to the epigenomic profile by characterizing the DNA methylation pattern of a certain number of genes associated with known tumors [17,18,19,20,21].The rate of concordance with respect to the diagnosis of occult primary tumor is around 82–97%.
Despite this, the impact on the clinical benefit of targeted therapy based on molecular studies remains controversial and the level of evidence and grade of recommendation is low, because they are based on short series or retrospective phase II studies [20, 22]. We have two prospective randomized studies completed in the last few years and neither demonstrate a significant benefit, except for the identification of clearly treatable primary tumors [23, 24]. (Level of evidence III, grade of recommendation B).
Sequencing platforms for tumor mutation profile characterization
Sequencing with gene panels using Next Generation Sequencing (NGS) techniques makes it possible to identify mutations that, in a more or less significant percentage of cases, could be associated with a benefit from the use of drugs targeted at these same mutations.
The potential benefit of this approach based on the ultraselection of oncospecific treatments based on molecular profiling can identify actionable genomic alterations in 87% of patients, and in between 30–40% of these patients with clear treatment directed against the molecular target with ESCAT or OncoKB level of 2 or less [25,26,27,28,29,30,31,32,33,34,35,36] (Table 4). The main objection stems from whether the response potential of a drug to a given mutation is conditioned by the tumor type in which the mutation is found. That means the histological type context and tumor location may condition the response to the drug, and not just the mutational profile [31, 37,38,39].
Platforms that perform complete genomic profiling by massive genomic sequencing in the coding region of a predetermined number of cancer-related genes allow the detection of genomic alterations such as mutations, DNA insertions/deletions, copy number variations, and gene recombinations. They also allow the evaluation of repair genes (study of microsatellite instability) and the determination of the tumor mutational burden (TMB), which is of paramount importance for the indication of target treatments and/or immunotherapeutics [26, 27, 32].
There is currently consensus on the therapeutic agnostic indication for the use of NTRK fusion gene inhibitors or, in those tumors with a TMB with more than 10 mutations per megabase, for the use of immunotherapy with pembrolizumab [35, 40, 41]. (Level of evidence III, grade of recommendation B).
Finally, it should be noted that a major international umbrella trial is currently underway with the intention of demonstrating that ultraselection of treatments based on genomic profiling and target therapy may represent a new paradigm in the management of these patients even without remote knowledge of the most likely primary tumor [42].
Recommendations for the use of molecular platforms
There is consensus among experts that the use of molecular platforms (MP) should be an ordinary part of the diagnostic process of CUP [25, 29, 30, 32, 33].
Patients who are candidates for MP should show an absence of clear clinical criteria for a known primary tumor, as well as an absence of clinical-immunohistochemical correlation. Initially, an anatomopathological morphological evaluation and immunohistochemical battery should be performed as precisely as possible, ensuring that after IHC, there is sufficient tissue to perform an MP analysis. If there are no clear results with initial IHC and a second and/or third IHC step, an MP should be used, not so much to identify the most likely primary tumor, but to characterize genomic alterations that may be associated with possible personalized target therapy treatments, especially in fit patients with a 0–1 ECOG and not very high LDH [8]. (Level of evidence III, grade of recommendation B).
Oncospecific systemic treatment according to histopathologic and/or clinical criteria
There is a subgroup of patients, between 10–15% of all CUP, whose form of presentation and/or specific anatomopathological criteria are part of a favorable prognostic group, since they have a specific treatment equivalent to that of the known primary tumors which they resemble. Table 5 shows the different groups of CUP with specific treatment [6]. (Level of evidence IV, grade of recommendation B).
However, most of these patients do not belong to any specific subtype or location, so the treatment of choice is empirical chemotherapy, with combinations containing a platinum agent plus another cytotoxic agent (taxanes, gemcitabine, irinotecan) [45], although this treatment provides poor results with low response rates (RR) of 15–20% and about nine months overall survival. [40, 46,47,48,49,50,51,52,53] Table 6. (Level of evidence II, grade of recommendation A).
It is important to point out that the IHC diagnosis is only indicative of the location of a primary tumor and should be considered by itself insufficient to guide treatment in that direction. Hence the concept of "suggestive primary cancer profile". This is where it makes sense to complement IHC information with NGS to ultra-detect mutations or other actionable genomic alterations. In these cases, a genomic platform could help us to identify a genomic profile and/or an actionable genomic alteration, or biomarkers of response to immunotherapy [27, 54]. (level of evidence III, grade of recommendation B).
Surgical and radiotherapy treatment
When CUP are identified as localized disease, after complete staging (including PET) [55], local treatment can result in long disease-free intervals and improve the prognosis of these patients. The most common sites include liver, bone, lung, skin, adrenal gland, and lymph nodes.
An attempt should be made at treatment with surgery of the solitary lesion. If resection is not feasible, definitive local radiation therapy should be proposed. Also, consider radiation therapy for patients with risk factors for residual disease, for example, multiple involved nodes or extracapsular spread, to maximize the chance of local control.
The role of neoadjuvant or adjuvant chemotherapy is undefined. Empirical adjuvant chemotherapy is reasonable in this setting, particularly in patients with poorly differentiated carcinoma or if indicated in a predicted tumor. (Level of evidence IV, grade of recommendation B).
Treatment based on suggesting primary cancer profile
The majority of patients with CUP (80–85%) are not included in the onco-specific treatment subgroups and their prognosis and treatment options are poor. Here arises the concept of “suggesting primary cancer profile” using the molecular cancer classifier (see Diagnostic molecular platforms aimed at identifying the primary tumor) to guide site-specific treatment [40].
Two retrospective non-randomized trials evaluating tumor type-specific therapies support this approach especially in patients with more responsive tumour types [20, 56]. However, two prospective studies (a Japanese phase II prospective and the European GEFCAPI-04 phase III trial) did not confirm the benefit in terms of progression-free survival or overall survival [23, 24].
So far, if primary site identification by these methods improves, outcomes remain to be seen, as these studies have numerous flaws such as non-optimal tumor type-specific therapy, cohorts enriched with resistant and unresponsive cancer types, or they are underpowered to detect survival benefit.
Otherwise, these molecular platforms can identify patients with tumors considered either sensitive or resistant to treatment. The benefits of this approach are most evident in patients predicted to have treatment-responsive tumor types [57]. Emerging non-randomized data favour the identification of certain subgroups of patients in whom a more specific treatment, similar to that for primary tumors which their conditions resemble, may be beneficial, such as renal-cell CUP, lung CUP and colorectal CUP. (Level of evidence III, grade of recommendation B).
Personalized treatment according to actionable genomic alterations
Due to its poor prognosis and response to conventional platinum-based chemotherapy treatments, the discovery of potential therapeutic molecular targets is one of the challenges of CUP.
Retrospective studies have observed that most patients with CUP harbored ≥ 1 oncogenic driver mutation determined by NGS technique [27, 28] and in 65% of patients with CUP actionable mutations by liquid biopsy-based cell-free circulating-tumor DNA [29], suggesting the need for further investigation into the value of molecular profiling.
Actionable genomic alterations matched to targeted therapies (Table 7) have been studied mostly from case reports retrospectively in patients with CUP [26, 58,59,60,61,62,63,64,65,66,67] and also biomarkers of response to immunotherapy such as TMB-high and MSI-high, associated to an important clinical benefit in some cases.
We have to take into consideration that some targeted therapies have already been approved by FDA for refractory and unresectable solid tumors such as larotrectinib and entrectinib (NTRK inhibitors) in NTRK fusion tumors with remarkable response [64] in CUP patients and pembrolizumab for MSI-high and TMB-high solid tumors [67]. (Level of evidence III, grade of recommendation B).
Apart from that there is a lack of prospective data from a large trial comparing platinum based chemotherapy to molecular alteration-based targeted therapy on this group, so nowadays the prognostic and predictive value of molecular profiling is pending and prospective studies examining biomarker-directed therapy on these cases (CUPISCO trial NCT03498521) [34] are ongoing.
Conclusions
CUP represents 3–5% of all tumor diagnoses and is characterized by an aggressive clinical course, an unpredictable metastatic pattern, early dissemination, intrinsic resistance to treatment and poor prognosis. It is a biologically complex entity, with intrinsic genetic alterations, which condition a behavior different from that of the primary tumors which it most resembles.
Given the aggressiveness of most CUPs, it is important to make a diagnosis as early as possible, performing the diagnostic process in a standardized manner, both clinically and pathologically, reserving part of the tumor sample, if possible, to determine actionable mutations, especially in patients with ECOG 0–1.
The limitations facing the diagnosis and treatment of CUP remain a major challenge. Improvements in diagnostic techniques, including the latest generation of IHC markers and genomic molecular platforms, have helped to refine the detection of the possible primary in many cases.
New molecular platforms based on massive gene sequencing offer us a new horizon through the detection of actionable genomic alterations for which personalized targeted treatments are available. The combination of treatment according to the most likely primary tumor and appropriate treatment for actionable mutations will undoubtedly offer an enormous opportunity of benefit for many patients diagnosed with CUP in the future.
References
Dykewicz CA. Centers for Disease Control and Prevention (U.S.), infectious diseases society of America, American society of blood and marrow transplantation. Summary of the guidelines for preventing opportunistic infections among hematopoietic stem cell transplant recipients. Clin Infect Dis Off Publ Infect Dis Soc Am. 2001;33(2):139–44.
Rassy E, Pavlidis N. The currently declining incidence of cancer of unknown primary. Cancer Epidemiol. 2019;61:139–41.
Conway A-M, Mitchell C, Kilgour E, Brady G, Dive C, Cook N. Molecular characterisation and liquid biomarkers in Carcinoma of Unknown Primary (CUP): taking the ‘U’ out of ‘CUP.’ Br J Cancer. 2019;120(2):141–53.
Rassy E, Assi T, Pavlidis N. Exploring the biological hallmarks of cancer of unknown primary: where do we stand today? Br J Cancer. 2020;122(8):1124–32.
Pavlidis N, Pentheroudakis G, Plataniotis G. Cervical lymph node metastases of squamous cell carcinoma from an unknown primary site: a favourable prognosis subset of patients with CUP. Clin Transl Oncol. 2009;11(6):340–8.
Hainsworth JD, Fizazi K. Treatment for patients with unknown primary cancer and favorable prognostic factors. Semin Oncol. 2009;36(1):44–51.
Penel N, Negrier S, Ray-Coquard I, Ferte C, Devos P, Hollebecque A, et al. Development and validation of a bedside score to predict early death in cancer of unknown primary patients. PLoS ONE. 2009;4(8):e6483.
Amela EY, Lauridant-Philippin G, Cousin S, Ryckewaert T, Adenis A, Penel N. Management of “unfavourable” carcinoma of unknown primary site: synthesis of recent literature. Crit Rev Oncol Hematol. 2012;84(2):213–23.
Losa F, Soler G, Casado A, Estival A, Fernández I, Giménez S, et al. SEOM clinical guideline on unknown primary cancer (2017). Clin Transl Oncol Off Publ Fed Span Oncol Soc Natl Cancer Inst Mex. 2018;20(1):89–96.
Natoli C, Ramazzotti V, Nappi O, Giacomini P, Palmeri S, Salvatore M, et al. Unknown primary tumors. Biochim Biophys Acta. 2011;1816(1):13–24.
Greco FA, Oien K, Erlander M, Osborne R, Varadhachary G, Bridgewater J, et al. Cancer of unknown primary: progress in the search for improved and rapid diagnosis leading toward superior patient outcomes. Ann Oncol Off J Eur Soc Med Oncol. 2012;23(2):298–304.
Zhu L, Wang N. 18F-fluorodeoxyglucose positron emission tomography-computed tomography as a diagnostic tool in patients with cervical nodal metastases of unknown primary site: a meta-analysis. Surg Oncol. 2013;22(3):190–4.
Møller AKH, Loft A, Berthelsen AK, Pedersen KD, Graff J, Christensen CB, et al. A prospective comparison of 18F-FDG PET/CT and CT as diagnostic tools to identify the primary tumor site in patients with extracervical carcinoma of unknown primary site. Oncologist. 2012;17(9):1146–54.
Oien KA. Pathologic evaluation of unknown primary cancer. Semin Oncol. 2009;36(1):8–37.
Lin F, Liu H. Immunohistochemistry in undifferentiated neoplasm/tumor of uncertain origin. Arch Pathol Lab Med. 2014;138(12):1583–610.
Conner JR, Hornick JL. Metastatic carcinoma of unknown primary: diagnostic approach using immunohistochemistry. Adv Anat Pathol. 2015;22(3):149–67.
Economopoulou P, Mountzios G, Pavlidis N, Pentheroudakis G. Cancer of Unknown Primary origin in the genomic era: elucidating the dark box of cancer. Cancer Treat Rev. 2015;41(7):598–604.
Kerr SE, Schnabel CA, Sullivan PS, Zhang Y, Singh V, Carey B, et al. Multisite validation study to determine performance characteristics of a 92-gene molecular cancer classifier. Clin Cancer Res Off J Am Assoc Cancer Res. 2012;18(14):3952–60.
Erlander MG, Ma X-J, Kesty NC, Bao L, Salunga R, Schnabel CA. Performance and clinical evaluation of the 92-gene real-time PCR assay for tumor classification. J Mol Diagn JMD. 2011;13(5):493–503.
Moran S, Martínez-Cardús A, Sayols S, Musulén E, Balañá C, Estival-Gonzalez A, et al. Epigenetic profiling to classify cancer of unknown primary: a multicentre, retrospective analysis. Lancet Oncol. 2016;17(10):1386–95.
Hainsworth JD, Greco FA. Gene expression profiling in patients with carcinoma of unknown primary site: from translational research to standard of care. Virchows Arch Int J Pathol. 2014;464(4):393–402.
Hainsworth JD, Rubin MS, Spigel DR, Boccia RV, Raby S, Quinn R, et al. Molecular gene expression profiling to predict the tissue of origin and direct site-specific therapy in patients with carcinoma of unknown primary site: a prospective trial of the Sarah Cannon research institute. J Clin Oncol Off J Am Soc Clin Oncol. 2013;31(2):217–23.
Hayashi H, Kurata T, Takiguchi Y, Arai M, Takeda K, Akiyoshi K, et al. Randomized phase II trial comparing site-specific treatment based on gene expression profiling with carboplatin and paclitaxel for patients with cancer of unknown primary site. J Clin Oncol Off J Am Soc Clin Oncol. 2019;37(7):570–9.
Fizazi K, Maillard A, Penel N, Baciarello G, Allouache D, Daugaard G, et al. A phase III trial of empiric chemotherapy with cisplatin and gemcitabine or systemic treatment tailored by molecular gene expression analysis in patients with carcinomas of an unknown primary (CUP) site (GEFCAPI 04). Ann Oncol. 2019;30:v851.
Tothill R, Li J, Mileshkin L, Doig K, Siganakis T, Cowin P, et al. Massively-parallel sequencing assists the diagnosis and guided treatment of cancers of unknown primary. J Pathol. 2013;1:231.
Gatalica Z, Millis SZ, Vranic S, Bender R, Basu GD, Voss A, et al. Comprehensive tumor profiling identifies numerous biomarkers of drug response in cancers of unknown primary site: analysis of 1806 cases. Oncotarget. 2014;5(23):12440–7.
Ross JS, Wang K, Gay L, Otto GA, White E, Iwanik K, et al. Comprehensive genomic profiling of carcinoma of unknown primary site: new routes to targeted therapies. JAMA Oncol. 2015;1(1):40–9.
Varghese AM, Arora A, Capanu M, Camacho N, Won HH, Zehir A, et al. Clinical and molecular characterization of patients with cancer of unknown primary in the modern era. Ann Oncol Off J Eur Soc Med Oncol. 2017;28(12):3015–21.
Kato S, Krishnamurthy N, Banks KC, De P, Williams K, Williams C, et al. Utility of genomic analysis in circulating tumor DNA from patients with carcinoma of unknown primary. Cancer Res. 2017;77(16):4238–46.
Clynick B, Dessauvagie B, Strerrett G, Harvey N, Subrramanian S, Herron J, et al. Detection of therapeutic targets in carcinomas of unknown primary. ESMO 2017 # 1705-PD. Ann Oncol. 2017;28(85):v595-604.
Oien KA, Dennis JL. Diagnostic work-up of carcinoma of unknown primary: from immunohistochemistry to molecular profiling. Ann Oncol Off J Eur Soc Med Oncol. 2012;23(Suppl 10):x271-277.
Ross JS, Sokol ES, Moch H, Mileshkin L, Baciarello G, Losa F, et al. Comprehensive genomic profiling of carcinoma of unknown primary origin: retrospective molecular classification considering the CUPISCO study design. Oncologist. 2021;26(3):e394-402.
Löffler H, Pfarr N, Kriegsmann M, Endris V, Hielscher T, Lohneis P, et al. Molecular driver alterations and their clinical relevance in cancer of unknown primary site. Oncotarget. 2016;7(28):44322–9.
Krämer A, Losa F, Gay LM, Sokol E, Page DR, Frampton GM, et al. Genomic profiling of carcinomas of unknown primary (CUP) to support clinical decisions. J Clin Oncol. 2018;36(15_suppl):e24162–e24162.
Mosele F, Remon J, Mateo J, Westphalen CB, Barlesi F, Lolkema MP, et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: a report from the ESMO Precision Medicine Working Group. Ann Oncol. 2020;31(11):1491–505.
Mateo J, Chakravarty D, Dienstmann R, Jezdic S, Gonzalez-Perez A, Lopez-Bigas N, et al. A framework to rank genomic alterations as targets for cancer precision medicine: the ESMO scale for clinical actionability of molecular targets (ESCAT). Ann Oncol. 2018;29(9):1895–902.
Fizazi K, Greco FA, Pavlidis N, Daugaard G, Oien K, Pentheroudakis G. Cancers of unknown primary site: ESMO clinical practice guidelines for diagnosis, treatment and follow-up†. Ann Oncol. 2015;1(26):v133–8.
Introduction | Metastatic malignant disease of unknown primary origin in adults: diagnosis and management | Guidance | NICE [Internet]. NICE; [cited 2021 Jul 5]. Available from: https://www.nice.org.uk/guidance/cg104/chapter/introduction
NCCN Clinical Practice Guidelines in Oncology. Occult Primary (Cancer of Unknown Primary [CUP]). Version 2.2021. NCCN.org.
Rassy E, Bakouny Z, Choueiri TK, Van Allen EM, Fizazi K, Greco FA, et al. The role of site-specific therapy for cancers of unknown of primary: A meta-analysis. Eur J Cancer. 2020;1(127):118–22.
Colomer R, Mondejar R, Romero-Laorden N, Alfranca A, Sanchez-Madrid F, Quintela-Fandino M. When should we order a next generation sequencing test in a patient with cancer? EClin Med. 2020;25:100487.
Pauli C, Bochtler T, Mileshkin L, Baciarello G, Losa F, Ross JS, et al. A challenging task: identifying patients with cancer of unknown primary (CUP) according to ESMO guidelines: the CUPISCO trial experience. Oncologist. 2021;26(5):e769–79.
Hainsworth JD, Greco FA. Adenocarcinoma of unknown primary site. www.uptodate.com ©2021 UpToDate, Inc.
Olivier T, Fernandez E, Labidi-Galy I, Dietrich P-Y, Rodriguez-Bravo V, Baciarello G, et al. Redefining cancer of unknown primary: is precision medicine really shifting the paradigm? Cancer Treat Rev. 2021;1(97):102204.
Greco FA, Pavlidis N. Treatment for patients with unknown primary carcinoma and unfavorable prognostic factors. Semin Oncol. 2009;36(1):65–74.
Culine S, Lortholary A, Voigt J-J, Bugat R, Théodore C, Priou F, et al. Cisplatin in combination with either gemcitabine or irinotecan in carcinomas of unknown primary site: results of a randomized phase II study–trial for the French Study Group on Carcinomas of Unknown Primary (GEFCAPI 01). J Clin Oncol Off J Am Soc Clin Oncol. 2003;21(18):3479–82.
Greco FA, Erland JB, Morrissey LH, Burris HA, Hermann RC, Steis R, et al. Carcinoma of unknown primary site: phase II trials with docetaxel plus cisplatin or carboplatin. Ann Oncol Off J Eur Soc Med Oncol. 2000;11(2):211–5.
Huebner G, Link H, Kohne CH, Stahl M, Kretzschmar A, Steinbach S, et al. Paclitaxel and carboplatin vs gemcitabine and vinorelbine in patients with adeno- or undifferentiated carcinoma of unknown primary: a randomised prospective phase II trial. Br J Cancer. 2009;100(1):44–9.
Dowell JE, Garrett AM, Shyr Y, Johnson DH, Hande KR. A randomized Phase II trial in patients with carcinoma of an unknown primary site. Cancer. 2001;91(3):592–7.
Briasoulis E, Fountzilas G, Bamias A, Dimopoulos MA, Xiros N, Aravantinos G, et al. Multicenter phase-II trial of irinotecan plus oxaliplatin [IROX regimen] in patients with poor-prognosis cancer of unknown primary: a hellenic cooperative oncology group study. Cancer Chemother Pharmacol. 2008;62(2):277–84.
Schuette K, Folprecht G, Kretzschmar A, Link H, Koehne C-H, Gruenwald V, et al. Phase II trial of capecitabine and oxaliplatin in patients with adeno- and undifferentiated carcinoma of unknown primary. Onkologie. 2009;32(4):162–6.
Møller AKH, Pedersen KD, Abildgaard J, Petersen BL, Daugaard G. Capecitabine and oxaliplatin as second-line treatment in patients with carcinoma of unknown primary site. Acta Oncol Stockh Swed. 2010;49(4):431–5.
Hainsworth JD, Spigel DR, Burris HA, Shipley D, Farley C, Macias-Perez IM, et al. Oxaliplatin and capecitabine in the treatment of patients with recurrent or refractory carcinoma of unknown primary site: a phase 2 trial of the Sarah Cannon Oncology Research Consortium. Cancer. 2010;116(10):2448–54.
Gatalica Z, Xiu J, Swensen J, Vranic S. Comprehensive analysis of cancers of unknown primary for the biomarkers of response to immune checkpoint blockade therapy. Eur J Cancer Oxf Engl. 2018;1990(94):179–86.
Rades D, Kühnel G, Wildfang I, Börner AR, Schmoll HJ, Knapp W. Localised disease in cancer of unknown primary (CUP): the value of positron emission tomography (PET) for individual therapeutic management. Ann Oncol Off J Eur Soc Med Oncol. 2001;12(11):1605–9.
Hainsworth JD, Rubin MS, Spigel DR, Boccia RV, Raby S, Quinn R, et al. Molecular gene expression profiling to predict the tissue of origin and direct site-specific therapy in patients with carcinoma of unknown primary site: a prospective trial of the sarah cannon research institute. J Clin Oncol. 2012. https://doi.org/10.1200/jco.2012.43.3755.
Rassy E, Pavlidis N. Progress in refining the clinical management of cancer of unknown primary in the molecular era. Nat Rev Clin Oncol. 2020;17(9):541–54.
Tan DS-W, Montoya J, Ng Q-S, Chan K-S, Lynette O, Sakktee Krisna S, et al. Molecular profiling for druggable genetic abnormalities in carcinoma of unknown primary. J Clin Oncol Off J Am Soc Clin Oncol. 2013;31(14):237–9.
Yamada T, Ohtsubo K, Ishikawa D, Nanjo S, Takeuchi S, Mouri H, et al. Cancer of unknown primary site with epidermal growth factor receptor mutation for which gefitinib proved effective. Gan To Kagaku Ryoho. 2012;39(8):1291–4.
Taniwaki M, Yamasaki M, Kawata K, Kawamoto K, Funaishi K, Matsumoto Y, et al. ROS1-rearranged putative lung adenocarcinoma presenting as carcinoma of unknown primary site: a case report. Oncotarget. 2018;9(81):35278–82.
Hainsworth JD, Anthony GF. Lung adenocarcinoma with anaplastic lymphoma kinase (ALK) rearrangement presenting as carcinoma of unknown primary site: recognition and treatment implications. Drugs-Real World Outcomes. 2016;3(1):115–20.
Kato S, Okamura R, Mareboina M, Lee S, Goodman A, Patel SP, et al. Revisiting epidermal growth factor receptor (EGFR) amplification as a target for anti-EGFR therapy: analysis of cell-free circulating tumor DNA in patients with advanced malignancies. JCO Precis Oncol. 2019;3:1.
Hainsworth JD, Meric-Bernstam F, Swanton C, Hurwitz H, Spigel DR, Sweeney C, et al. Targeted therapy for advanced solid tumors on the basis of molecular profiles: results from MyPathway, an open-label, phase IIa multiple basket study. J Clin Oncol Off J Am Soc Clin Oncol. 2018;36(6):536–42.
Okamura R, Boichard A, Kato S, Sicklick JK, Bazhenova L, Kurzrock R (2018) Analysis of NTRK alterations in pan-cancer adult and pediatric malignancies: implications for NTRK-targeted therapeutics. JCO Precis Oncol. 2018;2018:PO.18.00183. https://doi.org/10.1200/PO.18.00183
Gatalica Z, Xiu J, Swensen J, Vranic S. Molecular characterization of cancers with NTRK gene fusions. Mod Pathol Off J U S Can Acad Pathol Inc. 2019;32(1):147–53.
Cocco E, Scaltriti M, Drilon A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat Rev Clin Oncol. 2018;15(12):731–47.
Brahmer JR, Tykodi SS, Chow LQM, Hwu W-J, Topalian SL, Hwu P, et al. Safety and activity of anti-PD-L1 antibody in patients with advanced cancer. N Engl J Med. 2012;366(26):2455–65.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Ferran Losa: Advisory or Consultancy: Roche, Amgen; Speaker Bureau: Roche, Amgen,Merck, Sanofi, Servier and Travel/Accommodation or Expenses: Roche, Amgen,Merck, Sanofi. Isaura Fernandez. Consultant or Advisory: Roche, Astra Zeneca; Speaker Bureau: Pfizer, AstraZeneca, Roche,MSD, Clovis,Glaxo. Olatz Etxaniz. Consultant or Advisory: Pfizer, Merck,Ipsen,Roche; Speaker Bureau: Pfizer,Ipsen,BMS; Expenses: Clovis, Pfizer,Janssen,BMS. Alejandra Giménez. Consultant or Advisory: Ipsen; Speaker Bureau: Merck, Ipsen, Sanofi; Expenses: Merck, Amgem, Roche. Paula Gomila. No disclosures. Lara Iglesias. Consultant or Advisory: Merck, MSD, BMS, Lilly, Roche, Bayer, Sanofi. Speaker Bureau: Merck, MSD, BMS, Roche, Bayer, Sanofi, Eisai, Kyowa-kirin. Federico Longo. Advisory/Consultant: Roche, Ferre. Esteban Nogales. Consultant or Advisory: Roche, Daiichi Sankyo; Speaking: Pfizer, AstraZeneca, Roche, Kyowa Kirin. Antonio Sánchez: No disclosures. Gemma Soler. Advisory or Consultancy: Amgem,Roche,Kiowa; Speaker Bureau: Amgem,Merck,Roche, Sanofi, Servier and Travel/Accommodation or Expenses: Amgem, Merck,Roche, Sanofi.
Ethical approval
The present study was conducted in accordance with the ethical standards established in the 1964 declaration of Helsinki and its later amendments.
Informed consent
For this type of study informed consent is not required.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Losa, F., Fernández, I., Etxaniz, O. et al. SEOM—GECOD clinical guideline for unknown primary cancer (2021). Clin Transl Oncol 24, 681–692 (2022). https://doi.org/10.1007/s12094-022-02806-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12094-022-02806-x