Abstract
The variability of Helicobacter pylori morphology and the heterogeneity of virulence factors expressed by these bacteria play a key role as a driving force for adaptation to the hostile stomach environment. The aim of the study was to determine the relationship between the presence of certain genes encoding virulence factors and H. pylori morphology. One reference and 13 clinical H. pylori strains with a known virulence profile (vacA, cagA, babA2, dupA, and iceA) were used in this study. Bacteria were cultured for 1 h and 24 h in stressogenic culture conditions, i.e., serum-free BHI broths at suboptimal conditions (room temperature and atmosphere, without shaking). H. pylori cell morphology was observed by light and scanning electron microscopy. The vacA polymorphism and the cagA and babA2 presence were positively correlated with the reduction in cell size. Exposure to short-time stressogenic conditions caused more intense transformation to coccoid forms in highly pathogenic H. pylori type I strains (35.83% and 47.5% for type I s1m2 and I s1m1, respectively) than in intermediate-pathogenic type III (8.17%) and low pathogenic type II (9.92%) strains. The inverse relationship was observed for the number of rods, which were more common in type III (46.83%) and II (48.42%) strains than in type I s1m2 (19.25%) or I s1m1 (6.58%) strains. Our results suggest that there is a close relationship between the presence of virulence genes of H. pylori strains and their adaptive morphological features.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Helicobacter pylori is a microaerophilic, spiral, Gram-negative rod capable of effectively colonizing the gastric mucosa (Marshall and Warren 1984). Bacteria spread directly from person to person and are responsible for the development of a wide range of gastrointestinal diseases, including chronic (active) gastritis, gastric and/or duodenal ulcers, gastric adenocarcinomas, and mucosa-associated lymphoid tissue lymphomas (Malfertheiner et al. 2017). The presence of gastrointestinal pathologies and their severity is influenced by the type of bacterial strains and individual predispositions of the host, which determine the switching of H. pylori between commensalism and pathogenicity (Larussa et al. 2015). The long-lasting equilibrium between the host and H. pylori, covering at least 60,000 years, has enabled these microbes to develop evolutionary adaptations (Atherton and Blaser 2009; Talebi Bezmin Abadi 2017). The ability to avoid immune clearance and rapid response to changing environmental conditions are key determinants of the persistent host colonization by H. pylori (Larussa et al. 2015; Talebi Bezmin Abadi 2017).
One way of adaptation to environmental changes is the capability of H. pylori to undergo a morphological transformation. This microorganism occurs in two morphological forms, i.e., spiral—live, culturable, and coccoid—viable but nonculturable. The morphological transition from spiral to spherical form is observed under suboptimal environmental conditions, such as aerobiosis, temperature or pH changes, prolonged culture, or exposure to antibiotics or proton pump inhibitors (Loke et al. 2016). In proteomic studies, Loke et al. (2016) found that coccoid forms have high levels of proteins participating in DNA replication and carcinogenesis initiation and should therefore be regarded as highly virulent. The ability of H. pylori to produce elongated/ filamentous forms was also demonstrated, both in vitro (in amiA mutants and also after exposure to aztreonam, elevated NaCl concentrations or during oxygenated CO2-depleted culture) and in vivo (Takeuchi et al. 2006; Chaput et al. 2006; Park et al. 2011; Shao et al. 2012; Singh et al. 2015). It is postulated that increased cell filamentation may be a property of commensal bacteria, while the reduction of elongation is a hallmark of pathogenicity (Rossetti et al. 2013).
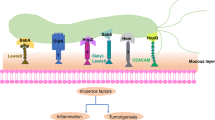
The heterogeneity of H. pylori strains and the variability of virulence factors expressed by these bacteria play a key role in adaptation to the stomach environment of the host (Odenbreit et al. 2009). The production of CagA (cytotoxin-associated gene A) is a feature of more virulent H. pylori strains, because this oncoprotein is capable of modulating host cell physiology (initiation of increased motility and elongated shape, the so-called hummingbird phenotype), induction of carcinogenesis, and IL-8-dependent initiation of inflammation. The vacuolating cytotoxin A (VacA) is a 140-kDa polypeptide responsible for the host tissue damage associated with the formation of anion-selective pores in cell membranes, mitochondria disruption, and eukaryotic cell death (Roesler et al. 2014; Amieva and Peek 2016). The VacA toxicity is determined mainly by two variable regions, called signal (s1 and s2) and middle (m1 and m2). The vacAs1 strains have a higher cytotoxic activity and are responsible for a greater risk of developing gastrointestinal diseases (especially vacAs1m1) than non-vacuolating vacAs2 strains (Roesler et al. 2014). Based on the expression of these two virulence factors, H. pylori strains were classified into three groups, i.e., with the highest virulence potential, cagA+/vacAs1 (type I) and the lowest virulence, cagA−/vacAs2 (type II), and strains with intermediate virulence with genotype, cagA−/vacAs1 or cagA+/vacAs2 (type III) (Xiang et al. 1995). Blood group antigen-binding adhesin A (BabA) is a protein associated with adhesion to the gastric mucosa, by binding to various components of the host (Ansari and Yamaoka 2017). Duodenal ulcer promoting gene A (DupA) is one of many important virulence factors often referred to as important in the promotion of gastrointestinal diseases. The production of this protein is coupled positively with the secretion of proinflammatory cytokine IL-8 (Queiroz et al. 2011). Another important virulence determinant involved in the pathogenicity of H. pylori is induced by contact with epithelium gene A (IceA). Production of this protein is stimulated by contact with the gastric epithelium, which contributes to the increase in IL-8 secretion (Shiota et al. 2012).
The aim of the study was to determine the relationship between the presence of certain genes encoding virulence factors and morphological changes of H. pylori strains.
Materials and methods
Bacterial strains and growth conditions
Virulence genes of all 13 clinical H. pylori strains used in this study (Table 2) have been identified and published previously by Biernat et al. (2010, 2014). In these reports, we have shown that H. pylori with specific virulence factors (cagA, vacA, babA2, dupA, and iceA in Table 2) were associated with the induction of gastric mucosa inflammation and tissue destruction. Detection of vacA, iceA1, and babA2 was carried out using multiplex PCR, whereas the cagA and iceA2 genes were detected by single PCR. H. pylori strains used in the study were formerly isolated from antral biopsy specimens of patients with gastritis. In addition, the reference strain J99 (ATCC 700824) was used as a control strain. The genome sequence of H. pylori J99 is available at GenBank with the accession number AE001439.
Culture methods of H. pylori strains were performed according to Macegoniuk et al. (2017) with minor modifications. The frozen bacterial suspensions of clinical and reference H. pylori strains were stored at − 70 °C in tryptic soy broth (Oxoid) with 15% glycerol. After reviving from frozen stocks, bacterial strains were pre-cultured onto Columbia agar (Difco) enriched with 7% hemolysed horse blood and H. pylori-selective supplement (Oxoid). Examined strains were incubated under microaerophilic conditions (Genbox microaer kits, BioMerieux) at 37 °C for 3 days and then sub-cultured on the same fresh media and incubated for 3 more days under the same conditions. In all of the following experiments, each H. pylori strain was passaged for only two or three times to reduce the risk of phase-variable changing of virulence genes (Odenbreit et al. 2009).
Preparation of cultures for light microscopy
H. pylori strain J99 was used to find the best environment that yielded the lowest average cell length and highest percentage of coccoid forms. Bacteria from solid cultures were inoculated into 2 ml of fresh brain heart infusion (BHI, Sigma) broths to obtain OD600 of 0.3 (early log phase) or 1 (stationary phase). Each bacterial culture was then incubated for 1 h and 24 h under specified conditions: BHI with 7% fetal bovine serum (Gibco) at optimal conditions (37 °C, microaerophilic, 100 rpm), BHI without the serum at optimal conditions, BHI with the serum at suboptimal conditions (room temperature and atmosphere, without shaking), and BHI without the serum at suboptimal conditions.
Cell length, shape determination, and image acquisition
Light microscopy analysis was performed similarly as described elsewhere (Rossetti et al. 2013) with small modifications. For each time point and strain, 50 μL of bacterial suspension was dropped onto a coverslip, stained by Gram’s method and observed under an Olympus BX50 microscope (Olympus Optical) using a × 100 oil immersion objective with numerical aperture of 1.3. The measurement of cell length was performed manually using the CellTool software. The average length of bacterial cells was counted using two independent experiments. From each experiment, three slides were taken and 50 bacterial cells per slide were counted (n = 300 cells/strain for each time point).
H. pylori cell forms were classified into five groups: coccoid forms (0.5–1 μm), short rods (1–2 μm), rods (2–4 μm), elongated rods (4–5 μm), and filamentous forms (≥ 5 μm) (Fig. 1). Classification of H. pylori cell length was determined by the observations of others (Nayak and Rose 2007; Hirsch et al. 2012; Chiou et al. 2013). Spherical forms larger than 1 μm, and rods shorter than 1 μm, accounted for less than 1% of the total pool of bacteria and did not affect the final results.
Scanning electron microscopy
In order to precisely determine the morphological structure of H. pylori cells and to confirm observations made using light microscopy, scanning electron microscopy (SEM) was used. The morphology of two H. pylori strains with the greatest variation in the virulence profile, namely 6171/T.II (vacAs2m2, cagA−, dupA−, iceA−, babA2−) and J99 (vacAs1m1, cagA+, dupA+, iceA2+, babA2+), was analyzed by SEM. These strains from solid cultures were inoculated into 2 ml of fresh BHI (Sigma) broths to obtain OD600 of 0.3 (early log phase) and were cultured for 1 h in serum-free BHI broths at suboptimal conditions (room temperature and atmosphere, without shaking). After this time point, microbes in liquid media were prefixed in 2.5% glutaraldehyde for 24 h and pelleted by centrifugation (600g, 5 min) in microcentrifuge tubes. Pellets were washed a total of three times in 0.1 M cacodylate buffer (pH 7.4) after each step cetrifugating bacteria again (600g, 5 min). Microbes were treated with a series of ethanol concentrations (50, 70, 90, 99.99%) and pelleted by centrifugation (600g, 5 min). After dehydration in a graded series of ethanol, the specimens were coated with 10 nm of gold. The samples were examined with a scanning electron microscope (Auriga 60, Zeiss) operating at 2 kV.
Statistical analysis
From two independent experiments, six slides per strain were taken and results were expressed as the mean ± standard deviation of the mean (SDM). All data were analyzed by STATISTICA v. 10.0. The distribution normality of continuous variables was calculated using the Shapiro-Wilk test. As data from bacterial cell sizes were not normally distributed, the correlation between variables was analyzed by the nonparametric Spearman’s rho. The differences in statistical significance between bacterial cell sizes at 1-h and 24-h incubation periods were analyzed by the Wilcoxon test. Cell morphology type frequencies were estimated by the Pearson χ2 analysis. The reduction of bacterial cell size after 1-day incubation period for H. pylori strains with specific virulence factors was analyzed by the Mann-Whitney U test and the Kruskal-Wallis test. The differences were found to be statistically significant at a p ≤ 0.05.
Results
Optimization of conditions inducing H. pylori morphological changes
After 1-h and 24-h incubation periods, there were no significant differences in the cell length distribution of H. pylori J99 in broths with different initial densities of bacteria, but at a density of OD600 = 1 bacterial cells were intensively autoaggregated (data not shown). Therefore, further studies were carried out using a density of OD600 = 0.3 to accurately determine the size and shape of a single bacterial cell.
It was observed that the reduction of H. pylori J99 in size and the changes in morphology were most prominent in serum-free BHI media at suboptimal culture conditions (Table 1); these counted for 1.02 ± 0.02 μm (56.3% coccoid forms and 42.3% short rods) and 0.9 ± 0.04 μm (71% coccoid forms and 29% short rods) at 1-h and 24-h incubation, respectively. Average bacterial cell length was highest in BHI broths with the serum at optimal culture conditions and equaled 2.26 ± 0.02 μm (70.3% rods and 26.7% short rods) after 1 h and 2.16 ± 0.04 μm (60% rods and 34.7% short rods) after 1 day of incubation periods. Other culture conditions contributed to the formation of intermediate-sized cells, i.e., 1.81 ± 0.04 and 1.45 ± 0.03 μm (suboptimal culture conditions, BHI with the serum) and 1.47 ± 0.03 and 1.06 ± 0.02 μm (optimal culture conditions, serum-free BHI) at 1-h and 24-h incubation, respectively.
For this reason, bacterial inoculations (OD600 = 0.3) were incubated in serum-free BHI broths at suboptimal culture conditions for 1 h and 24 h in the studies of the remaining H. pylori strains.
Analysis of cell length variability
Light microscopy observation of tested H. pylori strains at 0 h showed no significant differences in average cell length (p > 0.05), which ranged from 2.49 ± 0.03 to 2.1 ± 0.03 μm (Table 2). After 1-h and 24-h incubation, the average cell length of tested H. pylori strains changed profoundly and ranged between 2.44 ± 0.05 and 1.02 ± 0.02 μm and between 1.5 ± 0.05 and 0.9 ± 0.04 μm, respectively. Despite the gradual gradation in average bacterial cell length, coupled with the increasing number of virulence genes, these differences were not statistically significant (p > 0.05) (data not shown).
The statistical analysis of the reduction in bacterial cell size with a specific virulence factor was found to be significant for vacA polymorphism (p = 0.0039) and the presence of cagA (p = 0.0113) and babA2 (p = 0.0179). This significance was not observed for dupA (p = 0.8465) and iceA (p = 0.9485) genes.
The reduction rate of cell length during incubation from 0 to 1 h was positively correlated with the presence of virulence genes (Table 2). The reduction of bacterial cell size counted for 38.68–51.54% (p < 0.005), 17.7–32.72% (p < 0.005), and 2.01–10.22% (p < 0.05, except from 6171/T.II and 7361/T.II strains) for vacAs1m1, vacAs1m2, and vacAs2m2 strains, respectively. The opposite situation was observed during the incubation of bacteria from 1 to 24 h. The reduction rate in size was inversely correlated with the presence of virulence genes. In vacAs2m2 and vacAs1m2 strains, the ranges were 24.75–40.83% (p < 0.005) and 10.27–20.97% (p < 0.05, except for 7042/T.I strain), respectively. In the case of the strain J99 (vacAs1m1), a distinct reduction in cell size was observed (p < 0.005), but among other vacAs1m1 strains, the decrease in cell length ranged from 2.48 to 4.2% and was insignificant (p > 0.05).
Analysis of cell shape variability
Table 3 shows the morphological alternations of all examined H. pylori strains after 1-h incubation under stressogenic culture conditions. Strain 6171/T.II with low virulence (vacAs2m2, cagA−, dupA−, iceA−, babA2−) at 1-h incubation was predominantly in the form of rods (61.3%), but also produced elongated/ filamentous cells (5.7%), short rods (29.3%), and coccoid forms (3.7%) (Fig. 2). A completely different phenotype was demonstrated for a highly pathogenic strain, namely H. pylori J99 (vacAs1m1, cagA+, dupA+, iceA2+, babA2+). Under the same conditions, it mainly formed coccoids (56.3%) and short rods (42.3%), while rod forms counted for only 1.3%, and elongated/ filamentous forms were not observed (Fig. 2).
Differences in cell morphology of H. pylori strains observed under light and scanning electron microscope. Bacteria cultured for 1 h in serum-free BHI broths at suboptimal conditions (room temperature and atmosphere, without shaking). The H. pylori 6171/T.II strain seen mainly as rods (type II, low virulence) under light microscopy (a) and SEM (b), and H. pylori J99 seen mainly as coccoid forms (type I, high virulence) under light microscopy (c) and SEM (d). Coccoid forms of H. pylori are marked by arrows, while the small spherical structures within the squares indicate outer membrane vesicles. Scale bar in light microscopy, 5 μm
For analyzing the differences in the morphological properties, tested strains were classified into four groups, namely type II, type III, type I s1m2 (cagA+, vacAs1m2), and type I s1m1 (cagA+, vacAs1m1). After 1-h incubation, the presence of several times higher numbers of spherical forms in H. pylori type I strains (35.83% and 47.5% for type I s1m2 and I s1m1, respectively) than in type III (8.17%) (p < 0.000) and II (9.92%) (p < 0.000) strains was demonstrated (Fig. 3). The inverse relationship was observed for the number of rods, which were more common in type III (46.83%) and II (48.42%) strains than in type I s1m2 (19.25%) (p < 0.000) or I s1m1 (6.58%) (p < 0.000) strains (Fig. 3). Despite no statistically significant differences in the morphological properties of H. pylori type I s1m1 and I s1m2 strains (p = 0.141), the existence of a feature distinguishing both types was noted. In the case of H. pylori type I s1m1 strains, filamentous/elongated forms were never observed, while in H. pylori type I s1m2 strains, these forms were present and counted for 3% of all examined cells (Table 3).
The impact of virulence type on the morphology distribution of H. pylori strains after 1-h incubation period in stressogenic culture conditions. Cell morphology measured from light microscopy images reports as percent of the total bacterial population examined. Data combined from two independent bacterial cultures incubated in serum-free BHI broths at suboptimal culture conditions for 1 h period (n = 300 cells/strain). Results are given as mean ± SDM (n = 6)
Discussion
H. pylori in the natural habitat is mainly present as spiral forms, whereas culture in artificial in vitro conditions or exposure to suboptimal environmental factors is accompanied by morphological transformation into spherical or filamentous forms (Azevedo et al. 2007; Cellini et al. 2008; Park et al. 2011; Shao et al. 2012; Faghri et al. 2014). This pleomorphic nature is not specific to H. pylori only, since the ability to undergo morphological transformation has been observed for many Gram-negative rods, i.e. Campylobacter jejuni, Enterobacter aerogenes, Escherichia coli, Klebsiella pneumoniae, Legionella pneumophila, Proteus mirabilis, Salmonella enterica, and Vibrio cholerae (Krogfelt et al. 1993; Justice et al. 2006, 2014; Chaiyanan et al. 2007; Krebs and Taylor 2011; Horvath Jr et al. 2012; Ramamurthy et al. 2014; Ghaffar et al. 2015).
The current study has found that adaptational changes in cell morphology were closely related to the virulence profile of H. pylori strains. Rossetti et al. (2013) postulated that increased cell size (including the intensification of the filamentation process) is potentially a feature of commensal microbes, while reduction of elongation intensity can be traced as a pathogenic hallmark. This hypothesis is in agreement with the results obtained in our study. We noted that lower virulence strains had longer cells than highly virulent strains. Moreover, the reduction in cell size of the tested H. pylori strains during short-term exposure to suboptimal conditions was positively correlated with the presence of certain virulence genes, e.g., vacA, cagA, and babA.
It seems that strains with higher adaptability (and higher virulence) are capable of achieving reduced cell size in a shorter time during exposure to stressogenic conditions. The results in this study, showing that H. pylori strains with a higher virulence potential are capable of morphological adaptations in a short period to adverse environmental conditions, are consistent with reports by Vitoriano et al. (2011). The authors undertook co-culture experiments of peptic ulcer disease (PUD) and nonulcer dyspepsia (NUD) H. pylori strains with eukaryotic cell lines. It was demonstrated that after 1 day of co-culture, NUD strains possessed their original spiral shape, whereas PUD strains obtained spherical form as early as after 12-h co-incubation time. It has been suggested that this phenomenon is caused by the more severe destruction of eukaryotic cells and necrosis-dependent changes in environmental conditions and thus faster transformation of H. pylori PUD strains into coccoid forms. There are also scientific reports suggesting that minimization of bacterial cell size is a protective feature that contributes to the reduction of the cellular surface exposed to the immune system clearance (complement deposition or opsonophagocytosis) (Dalia and Weiser 2011; Veyrier et al. 2015). Their suggestions are within the scope of our observations showing that under suboptimal conditions (including nutrient depletion or exposure to the oxygenic and thermic stress), the process of reducing cell size and creating short rods or coccoid forms may be a potentially adaptive feature.
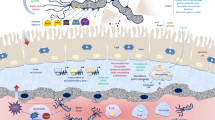
The direct molecular mechanism underlying the relationship between the presence of virulence genes and H. pylori morphology is not known. The presence of H. pylori virulence genes, however, does not have to exert a direct influence on the bacterial phenotype. Modulation of the morphology or physiology of microbial cells could also have an indirect impact. For example, there is a strong relationship between the presence of dupA and H. pylori growth at low pH (Lu et al. 2005; Talebi Bezmin Abadi et al. 2012). Bacterial adaptation in different pH conditions may contribute to the activation or inhibition of different molecular mechanisms. There are many proteins that become active or inactive depending on various pH environments. In a similar fashion, the phenotype of H. pylori may be influenced by other environmental factors. The infection caused by H. pylori strains, especially those producing many virulence factors, contributes to the formation of the inflammatory altered gastric environment, which is characterized by occurrence of numerous immune cells (neutrophils, macrophages, and T cells), proinflammatory cytokines (IL-1β, IL-6, IL-8, and TNF-α), chemokines, and matrix metalloproteinases (Ma et al. 2016). Survival in the inflamed environment requires the presence of multiple genes that protect against the immune clearance, which are most likely co-expressed with the expression of virulence genes, such as cagA, vacA, or babA. Some of these genes, protecting against immune attacks and the presence of unfavorable environmental conditions, could encode information about factors responsible for rapid morphological changes in H. pylori.
Limitations
Our study has some limitations. Firstly, the analysis was conducted with selected virulence genes (cagA, vacA, babA2, dupA, iceA). The selection of these virulence determinants was dictated by literature data indicating the significance of the abovementioned genes in the development of gastrointestinal disorders (Zambon et al. 2003; Torres et al. 2009; Shiota et al. 2010, 2012). However, it cannot be ruled out that the presence of other genes, not investigated in this study, also influences H. pylori morphology. On the other hand, this method seems to be surprisingly sensitive because it detected differences in cell morphology in relatively low amounts of tested virulence determinants. Secondly, the analysis was performed with s1/2 and m1/2 vacA alleles only. The relevance of the vacA intermediate (i) region in the cytotoxic activity of VacA has been demonstrated (Rhead et al. 2007). In the current study, H. pylori strains were categorized into three groups, namely types I, II, and III. This classification does not include the i-region, but only the s- and m-regions (Xiang et al. 1995). Hence, the impact of this region has not been investigated. The link between the morphology of H. pylori cells and the presence of virulence genes needs to be explored in different populations.
Conclusion
Based on microscopic observations, we demonstrated that the morphology of H. pylori strains might be influenced by the virulence profile. Less virulent H. pylori strains had a lower potential for the formation of coccoid forms than highly pathogenic strains. Furthermore, strains with lower pathogenicity were characterized by the presence of filamentous/ elongated forms, not observed in highly virulent strains.
References
Amieva M, Peek RM (2016) Pathobiology of Helicobacter pylori-induced gastric cancer. Gastroenterology 150:64–78
Ansari S, Yamaoka Y (2017) Helicobacter pylori BabA in adaptation for gastric colonization. World J Gastroenterol 23:4158–4169
Atherton JC, Blaser MJ (2009) Coadaptation of Helicobacter pylori and humans: ancient history, modern implications. J Clin Invest 119:2475–2487
Azevedo NF, Almeida C, Cerqueira L, Dias S, Keevil CW, Vieira MJ (2007) Coccoid form of Helicobacter pylori as a morphological manifestation of cell adaptation to the environment. Appl Environ Microbiol 73:3423–3427
Biernat M, Iwańczak B, Grabińska J, Junka A, Łaczmański Ł, Gościniak G (2010) The prevalence of dupA gene of H. pylori strains in Polish children. Helicobacter 15:366
Biernat M, Gościniak G, Iwańczak B (2014) Prevalence of Helicobacter pylori cagA, vacA, iceA, babA2 genotypes in Polish children and adolescents with gastroduodenal disease. Postepy Hig Med Dosw 68:1015–1021
Cellini L, Grande R, Di Campli E, Traini T, Di Giulio M, Lannutti SN, Lattanzio R (2008) Dynamic colonization of Helicobacter pylori in human gastric mucosa. Scand J Gastroenterol 43:178–185
Chaiyanan S, Chaiyanan S, Grim C, Maugel T, Huq A, Colwell RR (2007) Ultrastructure of coccoid viable but non-culturable Vibrio cholerae. Environ Microbiol 9:393–402
Chaput C, Ecobichon C, Cayet N, Girardin SE, Werts C, Guadagnini S, Prévost MC, Mengin-Lecreulx D, Labigne A, Boneca IG (2006) Role of AmiA in the morphological transition of Helicobacter pylori and in immune escape. PLoS Pathog 2:e97
Chiou PY, Luo CH, Chang KC, Lin NT (2013) Maintenance of the cell morphology by MinC in Helicobacter pylori. PLoS One 8:e71208
Dalia AB, Weiser JN (2011) Minimization of bacterial size allows for complement evasion and is overcome by the agglutinating effect of antibody. Cell Host Microbe 10:486–496
Faghri J, Poursina F, Moghim S, Zarkesh Esfahani H, Nasr Esfahani B, Fazeli H, Mirzaei N, Jamshidian A, Ghasemian Safaei H (2014) Morphological and bactericidal effects of different antibiotics on Helicobacter pylori. Jundishapur J Microbiol 7:e8704
Ghaffar NM, Connerton PL, Connerton IF (2015) Filamentation of Campylobacter in broth cultures. Front Microbiol 6:657
Hirsch C, Tegtmeyer N, Rohde M, Rowland M, Oyarzabal OA, Backert S (2012) Live Helicobacter pylori in the root canal of endodontic-infected deciduous teeth. J Gastroenterol 47:936–940
Horvath DJ Jr, Dabdoub SM, Li B, Vanderbrink BA, Justice SS (2012) New paradigms of urinary tract infections: implications for patient management. Indian J Urol 28:154–158
Justice SS, Hunstad DA, Seed PC, Hultgren SJ (2006) Filamentation by Escherichia coli subverts innate defenses during urinary tract infection. Proc Natl Acad Sci 103:19884–19889
Justice SS, Harrison A, Becknell B, Mason KM (2014) Bacterial differentiation, development, and disease: mechanisms for survival. FEMS Microbiol Lett 360:1–8
Krebs SJ, Taylor RK (2011) Nutrient-dependent, rapid transition of Vibrio cholerae to coccoid morphology and expression of the toxin co-regulated pilus in this form. Microbiology 157:2942–2953
Krogfelt KA, Poulsen LK, Molin S (1993) Identification of coccoid Escherichia coli BJ4 cells in the large intestine of streptomycin-treated mice. Infect Immun 61:5029–5034
Larussa T, Leone I, Suraci E, Imeneo M, Luzza F (2015) Helicobacter pylori and T helper cells: mechanisms of immune escape and tolerance. J Immunol Res 2015:1–10
Loke MF, Ng CG, Vilashni Y, Lim J, Ho B (2016) Understanding the dimorphic lifestyles of human gastric pathogen Helicobacter pylori using the SWATH-based proteomics approach. Sci Rep 6:26784
Lu H, Hsu PI, Graham DY, Yamaoka Y (2005) Duodenal ulcer promoting gene of Helicobacter pylori. Gastroenterology 128:833–848
Ma HY, Liu XZ, Liang CM (2016) Inflammatory microenvironment contributes to epithelial-mesenchymal transition in gastric cancer. World J Gastroenterol 22:6619–6628
Macegoniuk K, Grela E, Biernat M, Psurski M, Gościniak G, Dziełak A, Mucha A, Wietrzyk J, Berlicki Ł, Grabowiecka A (2017) Aminophosphinates against Helicobacter pylori ureolysis - biochemical and whole-cell inhibition characteristics. PLoS One 12:e0182437
Malfertheiner P, Megraud F, O’Morain CA, Gisbert JP, Kuipers EJ, Axon AT, Bazzoli F, Gasbarrini A, Atherton J, Graham DY, Hunt R, Moayyedi P, Rokkas T, Rugge M, Selgrad M, Suerbaum S, Sugano K, El-Omar EM, European Helicobacter and Microbiota Study Group and Consensus panel (2017) Management of Helicobacter pylori infection - the Maastricht V/Florence consensus report. Gut 66:6–30
Marshall BJ, Warren JR (1984) Unidentified curved bacilli in the stomach of patients with gastritis and peptic ulceration. Lancet 1:1311–1315
Nayak AK, Rose JB (2007) Detection of Helicobacter pylori in sewage and water using a new quantitative PCR method with SYBR® green. J Appl Microbiol 103:1931–1941
Odenbreit S, Swoboda K, Barwig I, Ruhl S, Borén T, Koletzko S, Haas R (2009) Outer membrane protein expression profile in Helicobacter pylori clinical isolates. Infect Immun 77:3782–3790
Park SA, Ko A, Lee NG (2011) Stimulation of growth of the human gastric pathogen Helicobacter pylori by atmospheric level of oxygen under high carbon dioxide tension. BMC Microbiol 11:96
Queiroz DM, Rocha GA, Rocha AM, Moura SB, Saraiva IE, Gomes LI, Soares TF, Melo FF, Cabral MM, Oliveira CA (2011) DupA polymorphisms and risk of Helicobacter pylori-associated diseases. Int J Med Microbiol 301:225–228
Ramamurthy T, Ghosh A, Pazhani GP, Shinoda S (2014) Current perspectives on viable but non-culturable (VBNC) pathogenic bacteria. Front Public Health 2:103
Rhead JL, Letley DP, Mohammadi M, Hussein N, Mohagheghi MA, Eshagh Hosseini M, Atherton JC (2007) A new Helicobacter pylori vacuolating cytotoxin determinant, the intermediate region, is associated with gastric cancer. Gastroenterology 133:926–936
Roesler BM, Rabelo-Gonçalves EMA, Zeitune JMR (2014) Virulence factors of Helicobacter pylori: a review. Clin Med Insights Gastroenterol 7:9–17
Rossetti V, Ammann TW, Thurnheer T, Bagheri HC, Belibasakis GN (2013) Phenotypic diversity of multicellular filamentation in oral streptococci. PLoS One 8:e76221
Shao C, Zhou Y, Sun Y, Wang H, Qu W, Yu H, Chen C, Jia J (2012) Analysis of aztreonam-inducing proteome changes in nondividing filamentous Helicobacter pylori. Curr Microbiol 65:108–115
Shiota S, Matsunari O, Watada M, Hanada K, Yamaoka Y (2010) Systematic review and meta-analysis: the relationship between the Helicobacter pylori dupA gene and clinical outcomes. Gut Pathog 2:13
Shiota S, Watada M, Matsunari O, Iwatani S, Suzuki R, Yamaoka Y (2012) Helicobacter pylori iceA, clinical outcomes, and correlation with cagA: a meta-analysis. PLoS One 7:e30354
Singh M, Prasad K, Yachha S (2015) Elongated Helicobacter pylori in gastric mucosa of children associated with gastric disease. J Pediatr Infect Dis 3:35–39
Takeuchi H, Nakazawa T, Okamoto T, Shirai M, Kimoto M, Nishioka M, Con SA, Morimoto N, Sugiura T (2006) Cell elongation and cell death of Helicobacter pylori is modulated by the disruption of cdrA (cell division-related gene A). Microbiol Immunol 50:487–497
Talebi Bezmin Abadi A (2017) Strategies used by Helicobacter pylori to establish persistent infection. World J Gastroenterol 23:2870–2882
Talebi Bezmin Abadi A, Taghvaei T, Wolfram L, Kusters JG (2012) Infection with Helicobacter pylori strains lacking dupA is associated with an increased risk of gastric ulcer and gastric cancer development. J Med Microbiol 61:23–30
Torres LE, Melián K, Moreno A, Alonso J, Sabatier CA, Hernández M, Bermúdez L, Rodríguez BL (2009) Prevalence of vacA, cagA and babA2 genes in Cuban Helicobacter pylori isolates. World J Gastroenterol 15:204–210
Veyrier FJ, Biais N, Morales P, Belkacem N, Guilhen C, Ranjeva S, Sismeiro O, Péhau-Arnaudet G, Rocha EP, Werts C, Taha MK, Boneca IG (2015) Common cell shape evolution of two nasopharyngeal pathogens. PLoS Genet 11:e1005338
Vitoriano I, Saraiva-Pava KD, Rocha-Gonçalves A, Santos A, Lopes AI, Oleastro M, Roxo-Rosa M (2011) Ulcerogenic Helicobacter pylori strains isolated from children: a contribution to get insight into the virulence of the bacteria. PLoS One 6:e26265
Xiang Z, Censini S, Bayeli PF, Telford JL, Figura N, Rappuoli R, Covacci A (1995) Analysis of expression of CagA and VacA virulence factors in 43 strains of Helicobacter pylori reveals that clinical isolates can be divided into two major types and that CagA is not necessary for expression of the vacuolating cytotoxin. Infect Immun 63:94–98
Zambon CF, Navaglia F, Basso D, Rugge M, Plebani M (2003) Helicobacter pylori babA2, cagA, and s1 vacA genes work synergistically in causing intestinal metaplasia. J Clin Pathol 56:287–291
Acknowledgements
The H. pylori reference strain J99 (ATCC 700824) was obtained by the courtesy of Dr. A. Zawilak-Pawlik (Hirszfeld Institute of Immunology and Experimental Therapy, Polish Academy of Sciences, Poland). We would like to thank Dr. Łukasz Łaczmański (Bacteriophage Laboratory, Institute of Immunology and Experimental Therapy, Polish Academy of Sciences, Poland) for carrying out the statistical analysis. We also thank Dr. Jakub Grzesiak (Electron Microscopy Laboratory, Wroclaw Research Centre EIT+, Poland) for providing technical assistance in electron microscopy.
Authorship
PK designed the study, performed research, analyzed data, contributed new methods, and wrote the paper. MB and GG analyzed data and wrote the paper.
Funding
The study was supported by Wroclaw Medical University grant No: STM.A130.17.034 and partly from grant No: ST.A130.16.031. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
This study was presented during the IV Scientific Conference “Vectors and pathogens in the past and in the future”, 24 November 2017, Wroclaw, Poland.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Krzyżek, P., Biernat, M.M. & Gościniak, G. Intensive formation of coccoid forms as a feature strongly associated with highly pathogenic Helicobacter pylori strains. Folia Microbiol 64, 273–281 (2019). https://doi.org/10.1007/s12223-018-0665-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-018-0665-5