Abstract
Background
The activities of MYC, the androgen receptor, and its associated pioneer factors demonstrate substantial reprogramming between early and advanced prostate cancer. Although previous studies have shown a shift in cellular metabolic requirements associated with prostate cancer progression, the epigenetic regulation of these processes is incompletely described. Here, we have integrated chromatin immunoprecipitation sequencing (ChIP-seq) and whole-transcriptome sequencing to identify novel regulators of metabolism in advanced prostate tumors characterized by elevated MYC activity.
Results
Using ChIP-seq against MYC, HOXB13, and AR in LNCaP cells, we observed redistribution of co-bound sites suggestive of differential KMT2A activity as a function of MYC expression. In a cohort of 177 laser-capture microdissected foci of prostate tumors, KMT2A expression was positively correlated with MYC activity, AR activity, and HOXB13 expression, but decreased with tumor grade severity. However, KMT2A expression was negatively correlated with these factors in 25 LuCaP patient-derived xenograft models of advanced prostate cancer and 99 laser-capture microdissected foci of metastatic castration-resistant prostate cancer. Stratified by KMT2A expression, ChIP-seq against AR and HOXB13 in 15 LuCaP patient-derived xenografts showed an inverse association with sites involving genes implicated in lipid metabolism, including the arachidonic acid metabolic enzyme PLA2G4F. LuCaP patient-derived xenograft models grown as organoids recapitulated the inverse association between KMT2A expression and fluorine-18 labeled arachidonic acid uptake in vitro.
Conclusions
Our study demonstrates that the epigenetic activity of transcription factor oncogenes exhibits a shift during prostate cancer progression with distinctive phenotypic effects on metabolism. These epigenetically driven changes in lipid metabolism may serve as novel targets for the development of novel imaging agents and therapeutics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Background
Prostate cancers (PCa) are distinctively sensitive to the transcriptional activity of the androgen receptor (AR) during tumorigenesis and in response to hormone-based therapies for advanced disease [1]. In early PCa, the AR’s role in mediating luminal cell terminal differentiation undergoes a distinctive switch as AR remains active in progressively less-differentiated disease [2]. Concurrent with early transformation of these cells, the MYC proto-oncogene reactivates and opposes the antiproliferative functions of AR to drive continued cell growth [3, 4]. Although elevated MYC protein levels are detectable very early in both pre-neoplastic and neoplastic prostate luminal cells, unlike many other tumor types, its mRNA expression remains largely uncoupled from genomic alterations affecting the MYC locus with aneuploidies involving most of chromosome 8q [5,6,7]. By contrast, in advanced and metastatic castration-resistant prostate cancer (mCRPC), MYC frequently undergoes high-level or focal amplification which significantly co-occurs with focal gains of the AR gene body and/or its enhancer [8, 9].
Although increased transcriptional output is canonically associated with increased expression of transcription factors, both MYC and AR show evidence of rewiring or reprogramming upon progression of primary PCa to advanced disease, especially in the context of hormonal therapies used for treating recurrent PCa [10]. Hormone- and context-dependent activities of these master regulators further involve pioneer factors that mediate AR-driven transcriptional programming; GATA2, FOXA1, and HOXB13 drive lineage-specific gene expression that is critical for tumor development [11]. Although somatic mutations to FOXA1 and inherited mutations to HOXB13 are potential drivers, studies of prostate tumors suggest mostly that differential activity of these factors is associated with disease progression [11]. Recently, the mixed-lineage leukemia (MLL) protein complex, more commonly implicated as a driver of MLL fusion-positive leukemias, was also shown to interact with AR signaling in advanced PCa, demonstrating the increasing complexity of characterizing transcriptional drivers of PCa [12]. KMT2A encodes MLL1, a SET domain encoding histone lysine methyltransferase and although it rarely exhibits somatic genomic alterations in PCa, changes to its expression and recruitment to gene promoters and enhancers profoundly impact target gene expression [13].
A primary physiological function of a prostate luminal cell is to secrete prostatic fluid, a major component of which is citrate [14, 15]. This distinctive metabolic requirement for generating citrate precursors is maintained during PCa tumorigenesis with elevated flux through the tricarboxylic acid cycle despite tumorigenic hypoxic conditions [15]. In addition, MYC and AR cooperate to facilitate de novo synthesis of polyunsaturated long-chain fatty acids to accommodate increased demand for phospholipids on account of increased cell division events [16,17,18]. The 20-carbon arachidonic acid (ArA) is routinely incorporated into phospholipids; ArA is normally acquired dietarily but can be released from phospholipids by deacetylating phospholipases A (PLA) for generating eicosanoids and prostaglandins [19,20,21]. PLA2G4F encodes a Group IV cytosolic phospholipase A2 (cPLA2) with high specificity for ArA [21]. Although cPLA2 proteins are ubiquitously expressed, the relationship between cellular ArA requirements and the effects of AR and MYC on PCa progression are not known.
Here, we report that in prostate cancer cells, differential binding of AR and HOXB13 on account of varying MYC levels centers on genes regulated by KMT2A/MLL1, which in turn is inversely proportional to the expression of PLA2G4F in cohorts of advanced, but not localized, PCa. Patient-derived prostate cancer xenografts grown as organoids have increased uptake of ArA in models with increased MYC activity and PLA2G4F expression. Our data demonstrate a phenotypic ramification of epigenetic reprogramming that accompanies disease progression, with potential implications for the design of novel therapeutics and imaging agents.
2 Results
2.1 AR and HOXB13 co-localize with MYC at genes regulated by KMT2A
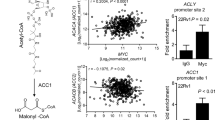
Although MYC activity is a well-known driver of prostate tumorigenesis, its relationship with individual genes and molecular processes remains actively investigated, with current efforts focusing on identifying effector genes or co-factors necessary for its positive association with androgen receptor (AR) activity [7, 10]. Using an institutional cohort of 177 laser capture microdissected (LCM) foci of primary prostate cancer (PCa), we examined the range of global gene expression that tracked with MYC transcript abundance. While the top 1% of correlating genes were strongly enriched for ribosomal biogenesis genes, AR and multiple AR target genes were also represented (Fig. 1A). Given the known role of MYC in mediating the AR-HOXB13 signaling axis, we analyzed a series of chromatin immunoprecipitation sequencing (ChIP-seq) experiments using antibodies against AR, HOXB13, and MYC, where the level of MYC was modulated by MYC-targeting hairpins (shMYC) [22, 23]. The overlap between MYC and AR or MYC and HOXB13 binding identified targets that may contribute to its function in primary PCa (outlined in Fig. 1B), which in turn pinpointed potential master regulators that regulate tumorigenesis.
Identification of transcription factors associated with MYC redistribution at AR- and HOXB13-occupied sites. A Ranked order depiction of genes whose expression correlate with MYC in an institutional cohort of 177 laser-capture microdissected prostate cancer tumor foci. B Schematic of the strategy used to identify upstream transcription regulators based on co-occupied peaks in LNCaP ChIP-seq data. anti-MYC [22], anti-AR, and anti-HOXB13 ChIP-seq data [23] used for analyses. C, D Bar plots showing the total peak number of MYC co-occupied sites, characterized as “core” or “redistributed” peaks, for anti-AR (C) and anti-HOXB13 (D) ChIP-seq. E Venn diagraph showing the overlap between transcription regulators identified by MYC/AR and MYC/HOXB13 co-occupied genes and nominated Ingenuity Upstream Regulator Analysis
Comparison of MYC and AR ChIP-seq data showed ~ 2600 and ~ 3200 MYC binding sites that colocalized with AR binding sites in LNCaP/NTC and LNCaP/shMYC cells, respectively; of these, 67% of peaks were shared (2154 of 3213 sites; Fig. 1C). For HOXB13 ChIP-seq, ~ 850 and ~ 1050 MYC binding sites were co-occupied in LNCaP/NTC and LNCaP/shMYC cells, respectively, with ~ 63% overlap (663 of 1049 sites, Fig. 1D). Co-bound sites were categorized as being core or redistributed and annotated to the nearest gene body or transcription start site. Here, we defined core peaks as co-occupied binding sites enriched in both MYC high- and low-expressing cells, and redistributed peaks as co-occupied binding sites enriched in MYC low-expressing cells. The corresponding lists of genes were then used as input for Ingenuity Pathway Analysis (IPA) to identify potential upstream transcriptional regulators (Supplementary Table 1).
This upstream regulator analysis identified nine shared transcription regulators among MYC/AR bound sites and four shared transcription regulators among MYC/HOXB13 bound sites (Fig. 1E). Of these, three were also shared by HOXB13 and AR: BCOR (BCL6 corepressor), BMI1 (B lymphoma Mo-MLV insertion region 1), and KMT2A (H3K4 lysine methyltransferase 2A) (Fig. 1E). BCOR and BMI1 encode members of the polycomb repressor complex 1 and are involved in cell differentiation; however, only BMI1 is implicated in PCa [24, 25]. Recently, a positive relationship between MYC and KMT2A has been documented and given that changes in KMT2A expression are associated with advanced disease, we selected KMT2A for further analysis [13, 26, 27].
2.2 KMT2A is positively associated with MYC activity in primary, but not advanced, PCa
Our ChIP-seq analyses of genetically-modified prostate cancer cell lines suggested that KMT2A activity may positively mediate the tumorigenic function of MYC. To address this potential interaction further, we analyzed whole-transcriptome sequencing (WTS) data from LCM primary prostate tumor foci (N = 177). Case-by-case, KMT2A expression was positively correlated with single-sample gene set enrichment analysis (ssGSEA) scores for MYC activity (Fig. 2A; r = 0.48, 95% CI 0.36 to 0.59, Pr < 0.0001). We next asked whether AR activity or HOXB13 expression was altered in the context of KMT2A. We observed a statistically significant and positive correlation between KMT2A and the ssGSEA scores for AR activity (Fig. 2B; r = 0.50, 95% CI 0.38 to 0.60, Pr < 0.0001) and the expression of HOXB13 (Fig. 2C; r = 0.32, 95% CI 0.18 to 0.45, Pr < 0.0001). However, when we subdivided these tumor foci by histological aggressivity (i.e., Gleason pattern, Gp), these positive associations generally decreased in effect size (Fig. 2A–C) from Gp3 (N = 48) to Gp4 (N = 101) to Gp5 (N = 28).
Decreasing association of KMT2A expression with prostate cancer drivers during primary disease progression. A–C Pearson correlation of the log2 CPM expression level for KMT2A with a 54-gene ssGSEA MYC activity score (A), a 266-gene ssGSEA AR activity score (B), or the log2 CPM expression level for HOXB13 (C) in a cohort of 177 laser capture microdissected foci of human prostate tumors. The error bars represent the 95% confidence bands for linear regression. For each correlation, the foci are subdivided into Gp3 (left), Gp4 (middle, including intraductal tumors), and Gp5 (right)
To validate this finding, we examined a separate cohort of LCM pairs of adjacent Gp3 and Gp4 tumor foci (from Gleason score 7 PCa) previously analyzed by Affymetrix microarray [28]. Across all foci (N = 26), and separately, KMT2A expression was positively correlated with MYC activity, AR activity, and HOXB13 expression (Supplementary Fig. 1). Interestingly however, analysis of WTS data from the TCGA-PRAD cohort (N = 484) did not recapitulate these associations (Supplementary Fig. 2), likely due in part to the bulk and admixed nature of TCGA tumor cases.
Nonetheless, these data suggest that at least in a subset of primary PCa, increased expression of KMT2A may contribute to tumor development, and that its expression in relationship to MYC activity modulates cancer progression. To assess this relationship in advanced disease, we examined gene expression profiles from WTS of two additional cohorts: a panel of metastatic prostate cancer patient derived xenografts (LuCaP PDX series, N = 25, [23, 29,30,31,32]) and the West Coast Dream Team-Prostate Cancer Foundation (WCDT-PCF) cohort of metastatic castration-resistant prostate cancer (mCRPC, N = 99, [8]). As depicted in Fig. 2A–C, the association between MYC activity, AR activity, HOXB13 expression and KMT2A expression decreased with dedifferentiation associated with higher Gleason patterns. Continuing this trend, we observed no strong association in LuCaP tumors (Fig. 3A), and this association was inversely related in WCDT-PCF tumors (Fig. 3B; r = − 0.35, 95% CI − 0.51 to − 0.16, Pr = 0.0004). Consistent with this inverse tendency, KMT2A expression was also negatively correlated with AR activity and HOXB13 expression in both the LuCaP PDX (Fig. 3C, D) and WCDT-PCF (Fig. 3E, F) cohorts. As the KMT2A expression and MYC activity relationship become inverted upon progression to metastatic disease, the tight association with AR and HOXB13 lineage-specific drivers is presumably lost, suggesting a context-dependent role for KMT2A in PCa with increased MYC activity; increased KMT2A expression (and likely its activity) may drive development of primary disease but not advanced PCa.
Inverse association of KMT2A expression with prostate cancer drivers in metastatic disease. A–F Pearson correlation of the log2 CPM expression level for KMT2A with the 54-gene ssGSEA MYC activity score (A, B), a 266-gene ssGSEA AR activity score (C, E), or the log2 CPM expression level for HOXB13 (D, F) in the LuCaP series of patient-derived xenografts (N = 25) (A, C, D) or the West Coast Dream Team-Prostate Cancer Foundation metastatic CRPC cohort (N = 99) (B, E, F). The error bars represent the 95% confidence bands for linear regression
2.3 Fluorine-18 labeled arachidonic acid uptake is increased in CRPC with low KMT2A expression and high MYC activity
Because KMT2A was differentially expressed relative to MYC activity in primary PCa versus mCRPC, we next examined potential functional implications of this relationship using the LuCaP PDX cohorts as a model of advanced disease in a series of ChIP-seq experiments. As depicted in Fig. 4A, we stratified this cohort based on mean KMT2A expression (log2 CPM) across the entire LuCaP series, with ChIP-seq data available for 15 of these LuCaP models; nine were classified KMT2A-high and six were classified KMT2A-low. From ChIP-seq experiments with antibodies against AR and HOXB13, peaks were merged within each subgroup to create a union set of sites for each factor based on KMT2A status. For AR and HOXB13, we identified 180 and 203 differentially enriched peaks (cutoff of P < 0.1), respectively (Supplementary Table 2); the majority of which enriched in KMT2A-low samples (Fig. 4B). This enrichment is consistent with increased MYC-associated transcriptional activity of AR and HOXB13 in tumors with lower KMT2A expression (see Fig. 3B).
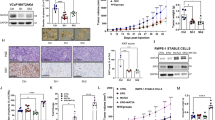
Inverse relationship between PLA2G4F and KMT2A expression in metastatic prostate cancer. A Expression of KMT2A across the LuCaP PDX models in our study displayed as a heatmap, indicating which models are classified as KMT2A high or low. B Heatmap of anti-AR and anti-HOXB13 ChIP-seq [29] peak signals (aggregated across samples as indicated) around TSS regions (± 3 kb) in LuCaP PDX's with differential KMT2A expression. Each row is a peak ranked by KMT2A low to high. The location of PLA2G4F on heatmap is marked by a black bar. C Functional enrichment from Ingenuity Upstream Pathway Analysis of anti-AR and anti-HOXB13 ChIP-seq differential peaks overlapping genes for pathways with P < 0.1. Red indicates that PLA2G4F is present in the pathway gene set. D Integrative genome viewer tracks of the PLA2G4F gene locus from anti-AR and anti-HOXB13 ChIP-seq. LNCaP DNase-hypersensitivity regions shown in blue. E–H Spearman correlation of the log2 CPM expression level of PLA2G4F with log2 CPM expression level of KMT2A (E, G) or the 54-gene ssGSEA MYC activity score (F, H) in the LuCaP (E, F) or the West Coast Dream Team-Prostate Cancer Foundation metastatic CRPC cohort (G, H). Samples with log2 CPM PLA2G4F < 0 were excluded. The error bars represent the 95% confidence bands for linear regression
We then sought to identify potential phenotypic effects in the KMT2A-low setting. We generated a list of differentially enriched genes for each transcription factor (TF) and histone mark and used these lists as input for IPA. IPA identified eight shared pathways, and examination of genes giving rise to pathway enrichment revealed strong representation of genes involved in fatty acid synthesis (Fig. 4C). The most common shared gene appearing in four of the eight pathways from both ChIP series, was PLA2G4F (phospholipase A2 group IVF), a key component of the arachidonic acid (ArA) metabolic pathway [33]. However, this enrichment was inversely proportional to KMT2A levels, with AR and HOXB13 being recruited to the PLA2G4F locus in the six KMT2A-low LuCaPs and mostly absent from the PLA2G4F locus in the nine KMT2A-high LuCaP models (Fig. 4D). Consistent with AR and HOXB13 binding, PLA2G4F mRNA expression was negatively correlated with KMT2A expression across all LuCaP PDXs (Fig. 4E) and maintained a negative correlation with MYC activity as well (Fig. 4F). This trend was also maintained in the WCDT-PCF mCRPC cohort (Fig. 4G, H). Interestingly, both KMT2A expression and MYC activity were positively correlated with PLA2G4F expression in primary PCa, with decreasing association stepwise from Gp3 to Gp4 to G5 (Supp Fig. 3). Taken together, these results further indicate a disparate role for KMT2A in MYC-driven advanced vs. localized PCa.
Due to the involvement of the protein product of PLA2G4F, cytosolic phospholipase A2 zeta, in ArA metabolism, we hypothesized that MYC activity-high, KMTA2-low, and PLA2G4F-high tumors have an increased cellular demand for ArA, due in part to its presence in phospholipids. We assessed uptake of fluorine-18 labeled ArA ([18F]ArA) over a two-hour time course in LuCaP PDX’s grown as organoids. As presented in Fig. 5A, the rate of [18F]ArA uptake varied across LuCaPs, ranging from 6 to 88% per million cells in KMT2A-high LuCaP models and 27–154% per million cells in the KMT2A-low LuCaP models. Overall, KMT2A-low models demonstrated greater median [18F]ArA uptake at two hours (54%) than the KMT2A-high models (22%) (P = 0.026, Mann–Whitney U test; Fig. 5B). This uptake tracked in part with the expression of PLA2G4F, with strong positive correlations observed between both the uptake of [18F]ArA at two hours (Fig. 5C) and the rate of uptake over two hours (Fig. 5D) versus PLA2G4F expression across the LuCaP cohort. These findings indicate that differential co-regulation of KMT2A expression, and its effects on PLA2G4F expression, by the chromatin binding activities of MYC, AR, and HOXB13, is modulated during PCa progression and is associated with a shift in cellular metabolic requirements.
Arachidonic acid (ArA) metabolism is associated with KMT2A expression. A Time course of [18F]ArA uptake in LuCaP PDX organoids (N = 15) classified as either KMT2A-high (red) or KMT2A-low (blue). Individual organoids are given by light shaded lines, with each line representing the median uptake of 2–5 replicates. Dark shaded lines represent the median uptake for KMT2A-high or KMT2A-low PDX organoids. Results are expressed as median uptake per 1 million cells. B. Percent [18F]ArA uptake in LuCaPs based on KMT2A expression at 120 min. Line at median. Each organoid (representing 2–5 replicate experiments) plotted as open circles (N = 9 high, N = 6 low). P = 0.026 by Mann–Whitney U test. C, D Spearman correlation of the log2 CPM expression level of PLA2G4F with the percent [18F]ArA uptake in LuCaPs at 120 min (C) or the rate of uptake over 2 h, measured by the slope of linear regression (D). Cases are colored by KMT2A expression status. The error bars represent the 95% confidence bands for linear regression of each scatter plot
3 Discussion
Although compounds targeting MYC continue to be developed, many efforts to dissect and target the tumorigenic properties of MYC in PCa have shifted to its effector genes or co-factors necessary for its activity [34]. Thus, in this study, we started by assessing sites differentially bound by MYC, AR, and HOXB13 in the setting of high or low MYC expression using LNCaP PCa cells to identify novel transcriptional regulators that likely contributed to changes in MYC activity over the course of human PCa progression. From that analysis we identified KMT2A/MLL1, whose expression in association with MYC activity shifted from directly to inversely proportional as PCa progressed from localized to metastatic. Integrating ChIP-seq and WTS data from LuCaP PDX models revealed that AR and HOXB13 direct the regulation of lipid metabolism as a function of KMT2A levels.
The consequence of this finding, that MYC indirectly, but positively, regulates lipid metabolism in part through PLA2G4F, agrees with numerous reports that dysregulated MYC function drives glycolysis and lipogenesis, such that inhibition of MYC activity reduced accumulation of intracellular lipid droplets [17, 35]. The implications of increased lipid metabolism are two-fold: increased synthesis and turnover of membrane lipids associated with cell division, and fatty acid oxidation as an alternative energy source [36]. With respect to PCa specifically, studies have linked changes in ArA metabolism to PCa development and progression in vitro, with these studies suggesting that ArA is preferentially metabolized by PCa cells [37, 38]. However, to the best of our knowledge, the current study is the first to demonstrate an epigenetic link between ArA metabolism, uptake, and MYC activity in human prostate tumors.
The central premise of our finding relies on the observation that MYC is rewired from opposing PLA2G4F expression in early cancer to being positively associated with it in progressively advanced disease. The stepwise histologic progression from lower to higher Gleason grades is reflective of dedifferentiation, which in turn is a direct consequence of altered AR activity [39]. Similarly, AR-directed therapies suppress AR activity in patients with metastatic PCa, such that resistant tumors are less dependent on AR for growth and survival [39]. Thus, histologically and clinically aggressive tumors may reflect a causal link between increased MYC activity and decreased but persistent AR activity, due in part to a transcriptional pause-release effect that accounts for lower AR tumors having worse clinical outcomes [4, 40]. A logical future study will be to broadly assess the metabolic and lipogenic ramifications of differential AR/MYC status.
This current study has two important limitations. First, we observed a switch in the association between KMT2A or PLA2G4F expression and MYC activity, AR activity, and HOXB13 expression transcriptomically between primary localized PCa. Although we could recapitulate the inverse association using ChIP-seq from LuCaP PDX models for which matched transcriptional data was available, similar ChIP-seq assays from primary PCa with matched RNA-seq were not obtainable. Second, we demonstrated functional changes in lipid metabolism using [18F]ArA uptake as a surrogate for cellular metabolic needs. Measurements of actual cPLA2 protein levels would have been ideal, but after extensive testing, none of the four commercial antibodies with described reactivity against the Group IV zeta (PLA2G4F) isoform were specific.
Despite these limitations, we have demonstrated concordance between transcriptional and chromatin readouts of TF activity within the same panel of PDX models, with further qualification of potential lipid metabolic differences using the PDXs in an in vitro functional assay. Our findings indicate that ArA metabolism is specifically sensitive to AR and MYC activity during PCa progression.
4 Conclusions
We have reported that ArA metabolism and uptake are enriched in MYC-high prostate tumors. This finding has implications for the subset of treatment resistant prostate cancers that maintain high MYC activity and develop resistance to both hormonal and cytotoxic chemotherapies. Future studies are needed to validate these findings in larger cohorts, assess the viability of targeting ArA metabolism therapeutically, and determine whether ArA-based imaging agents demonstrate specific uptake in MYC-high/KMT2A-low prostate cancers.
5 Methods
5.1 Study design
The aim of this study was to assess changes in gene expression patterns and corresponding cellular phenotypes associated with repositioning of MYC in prostate cancer. This study utilized LNCaP cells engineered to have reduced levels of MYC expression as described in [22], LuCaP PDX organoids as described in [31], and integration of both novel and published datasets derived from these lines and patient cohorts [8, 22, 23, 29,30,31].
5.2 ChIP-seq analysis in LNCaP cells
ChIP-seq against MYC in LNCaP cells harboring a nontargeting control hairpin (NTC) or MYC-targeting hairpins (MYC shRNA) was described previously [22]. ChIP-seq performed in LNCaP cells or LuCaP patient-derived xenografts (PDXs) were previously described [23, 29] and were retrieved from GEO (https://www.ncbi.nlm.nih.gov/geo/) using accession numbers GSE94682 and GSE130408, respectively. Downloaded FASTQ files were reprocessed through the nextflow [41] nf-core/chipseq (v1.2.2) pipeline (https://github.com/nf-core/chipseq). De-multiplexed reads were aligned to build GRCh37 of the human genome using BWA-MEM [42]. MACS2 [43] was used to perform peak calling using a cutoff FDR q-value of 0.01 and the parameter --narrow_peak. HOMER [44] annotePeaks.pl was used to annotate called peaks relative to known genomic features.
5.3 RNA-seq analysis
Whole transcriptome profiling of laser capture microdissected (LCM) primary prostate cancer (PCa) was previously described [22]. Transcriptomes from the Stand Up To Cancer West Coast Dream Team-Prostate Cancer Foundation metastatic castration resistant prostate cancer (WCDT-PCF mCRPC) cohort [8] were downloaded from the NCI Genomic Data Commons (https://gdc.cancer.gov) via access to dbGaP phs001648. Transcriptomes from the LuCaP [23, 29,30,31] series of patient-derived xenografts (PDXs) were downloaded from GEO using accession numbers GSE113741, GSE156292, and GSE126078. Downloaded FASTQ files were processed through the nextflow nf-core/rnaseq (v3.5) pipeline (https://github.com/nf-core/rnaseq). Sample reads were mapped to the GRCh37 reference genome using STAR [45] and quantified using featureCounts [46] with default parameters. EdgeR (v3.32.1) was used to generate the log2 counts per million (CPM) values used for downstream analysis [47].
Single-sample gene set enrichment (ssGSEA) was performed using the R package GSVA (v1.38.2) with the following parameters: tau: 0.75; ssgsea.norm: FALSE [48]. For each sample, ssGSEA projection values were obtained for a 54-gene MYC activity score [49] or a 266-gene AR activity score [50].
5.4 Identification of core and redistributed ChIP-seq peaks in LNCaP cells
Bed files for LNCaP ChIP-seq peaks created by MACS2 were used as input for BEDTools (v2.30.0) [51] to identify core and redistributed peaks. Each independent MYC ChIP-seq (NTC and MYC shRNA) was compared to AR or HOXB13 ChIP-seq data. Here, core (overlapping) peaks were defined as co-occupied binding sites enriched in both NTC- and MYC shRNA-harboring LNCaP cells. Redistributed (unique) peaks were defined as co-occupied binding sites enriched in MYC shRNA cells. New peak files representing this intersection were collated where appropriate and annotated to the nearest gene body (GB) or transcription start site (TSS) within ± 3 kb, and redundant gene symbols were removed.
5.5 ChIP-seq analysis in LuCaP xenografts
LuCaP xenografts for which ChIP-seq data were available were classified as KMT2A-high (N = 9) or KMT2A-low (N = 6) based on corresponding average KMT2A log2 CPM expression values across the entire LuCaP cohort, including cases where ChIP-seq was not performed. DESeq2 (v1.30.1) was used to identify differential peaks between KMT2A-high and KMT2A-low samples selecting peaks with adjusted P < 0.05 [52]. BigWig files were merged to create a consensus wiggle file based on KMT2A status for each ChIP-seq using WiggleTools (v1.2) [53] and converted into bigWig files using the UCSC wigToBigWig tool [54]. Generated bigWig files were visualized with the integrative genomics viewer. ChIP-seq heatmaps were prepared using deepTools (v3.5.1) computeMatrix and plotHeatmap tools [55].
5.6 Pathway analysis
Core analyses were performed with Ingenuity Pathway Analysis (IPA) tools [56]. For deduplicated core and redistributed peak lists derived from LNCaP ChIP-seq data, Upstream Regulator Analysis (URA) was used to identify factors upstream of the genes in the peak lists. URA was limited to transcription regulators. Enriched gene sets selected by GSEA (FDR q < 0.05) were used as input for Canonical Pathway analysis and normalized gene counts for shared core enrichment genes identified for primary PCa or CRPC were used. Counts were normalized as follows: (MYC counts + KMT2A counts)/total shared gene sets. Consensus differential peaks for LuCaP ChIP-seq data were also used for Canonical Pathways analysis, with overlapping pathways selected at a cutoff of P < 0.1.
5.7 Fluorine-18 labeled arachidonic acid ([18F]ArA) uptake in LuCaP xenografts
LuCaP xenografts were processed and cultured as organoids as described previously [31]. Organoid identity was validated every 6 months by STR profiling (Laragen). Organoids were cultured for 7 days in 2% Matrigel, collected, washed with PBS, then resuspended in 1 mL of Beshiri’s Modified Clevers Media [31] in duplicate tubes for in vitro [18F]ArA uptake assays.
[18F]ArA was prepared following as previously described with minor modifications [57]. Briefly, the tosylate precursor was heated with [18F]TBAF in acetonitrile at 120 °C for 20 min followed by hydrolysis with potassium hydroxide (120 °C for 10 min) to produce 20-[18F]ArA. The overall radiochemical yields were 15–24% (uncorrected, n > 6) with a molar activity of 59–163 GBq/µmol (n > 6).
Organoid cultures were radiolabeled with 2 μCi [18F]ArA in BMCM for 120 min at 37 °C/5% CO2, washed twice with PBS to remove residual unincorporated [18F]ArA, and resuspended in 1 mL trypsin as a single-cell suspension. Cells were counted using acridine orange/propidium iodide cell counting dye. [18F]ArA uptake was determined by PerkinElmer 2480 Wizard3 Gamma Counter. For each sample, percent uptake per 106 live cells was calculated as follows:
5.8 Statistical analysis
Statistical analyses were performed with GraphPad Prism version 9 (GraphPad Software) for Mac. Associations between factors were measured using Pearson or Spearman correlations. Comparisons of single factors between dichotomized samples was performed using Mann–Whitney U tests.
Data availability
The data underlying this article have been deposited in Database of Genotypes and Phenotypes (dbGaP) and Gene Expression Omnibus (GEO) at https://www.ncbi.nlm.nih.gov/gap/ and https://www.ncbi.nlm.nih.gov/geo/, respectively, and can be accessed with phs001813.v2.p1 (dbGaP), phs001938.v3.p1 (dbGaP) and GSE135942 (GEO).
Abbreviations
- [18F]ArA:
-
Fluorine-18 labeled arachidonic acid
- AR:
-
Androgen receptor
- ArA:
-
Arachidonic acid
- CRPC:
-
Castration-resistant prostate cancer
- mCRPC:
-
Metastatic castration-resistant prostate cancer
- PCa:
-
Prostate cancer
- PDX:
-
Patient-derived xenograft
- TF:
-
Transcription factor
References
Westaby D, Fenor de La Maza MLD, Paschalis A, Jimenez-Vacas JM, Welti J, de Bono J, Sharp A. A new old target: androgen receptor signaling and advanced prostate cancer. Annu Rev Pharmacol Toxicol. 2022;62:131–53.
Hendriksen PJ, Dits NF, Kokame K, Veldhoven A, van Weerden WM, Bangma CH, Trapman J, Jenster G. Evolution of the androgen receptor pathway during progression of prostate cancer. Cancer Res. 2006;66(10):5012–20.
Bai S, Cao S, Jin L, Kobelski M, Schouest B, Wang X, Ungerleider N, Baddoo M, Zhang W, Corey E, et al. A positive role of c-Myc in regulating androgen receptor and its splice variants in prostate cancer. Oncogene. 2019;38(25):4977–89.
Barfeld SJ, Urbanucci A, Itkonen HM, Fazli L, Hicks JL, Thiede B, Rennie PS, Yegnasubramanian S, DeMarzo AM, Mills IG. c-Myc antagonises the transcriptional activity of the androgen receptor in prostate cancer affecting key gene networks. EBioMedicine. 2017;18:83–93.
Sato K, Qian J, Slezak JM, Lieber MM, Bostwick DG, Bergstralh EJ, Jenkins RB. Clinical significance of alterations of chromosome 8 in high-grade, advanced, nonmetastatic prostate carcinoma. J Natl Cancer Inst. 1999;91(18):1574–80.
Stopsack KH, Whittaker CA, Gerke TA, Loda M, Kantoff PW, Mucci LA, Amon A. Aneuploidy drives lethal progression in prostate cancer. Proc Natl Acad Sci USA. 2019;116(23):11390–5.
Gurel B, Iwata T, Koh CM, Jenkins RB, Lan F, Van Dang C, Hicks JL, Morgan J, Cornish TC, Sutcliffe S, et al. Nuclear MYC protein overexpression is an early alteration in human prostate carcinogenesis. Mod Pathol. 2008;21(9):1156–67.
Quigley DA, Dang HX, Zhao SG, Lloyd P, Aggarwal R, Alumkal JJ, Foye A, Kothari V, Perry MD, Bailey AM, et al. Genomic hallmarks and structural variation in metastatic prostate cancer. Cell. 2018;174(3):758–69.
Robinson D, Van Allen EM, Wu Y-M, Schultz N, Lonigro Robert J, Mosquera J-M, Montgomery B, Taplin M-E, Pritchard Colin C, Attard G, et al. Integrative clinical genomics of advanced prostate cancer. Cell. 2015;161(5):1215–28.
Guo H, Wu Y, Nouri M, Spisak S, Russo JW, Sowalsky AG, Pomerantz MM, Wei Z, Korthauer K, Seo JH, et al. Androgen receptor and MYC equilibration centralizes on developmental super-enhancer. Nat Commun. 2021;12(1):7308.
Hankey W, Chen Z, Wang Q. Shaping chromatin states in prostate cancer by pioneer transcription factors. Cancer Res. 2020;80(12):2427–36.
Malik R, Khan AP, Asangani IA, Cieslik M, Prensner JR, Wang X, Iyer MK, Jiang X, Borkin D, Escara-Wilke J, et al. Targeting the MLL complex in castration-resistant prostate cancer. Nat Med. 2015;21(4):344–52.
Rao RC, Dou Y. Hijacked in cancer: the KMT2 (MLL) family of methyltransferases. Nat Rev Cancer. 2015;15(6):334–46.
Costello LC, Franklin RB. Citrate metabolism of normal and malignant prostate epithelial cells. Urology. 1997;50(1):3–12.
Ahmad F, Cherukuri MK, Choyke PL. Metabolic reprogramming in prostate cancer. Br J Cancer. 2021;125(9):1185–96.
Zadra G, Ribeiro CF, Chetta P, Ho Y, Cacciatore S, Gao X, Syamala S, Bango C, Photopoulos C, Huang Y, et al. Inhibition of de novo lipogenesis targets androgen receptor signaling in castration-resistant prostate cancer. Proc Natl Acad Sci USA. 2019;116(2):631–40.
Priolo C, Pyne S, Rose J, Regan ER, Zadra G, Photopoulos C, Cacciatore S, Schultz D, Scaglia N, McDunn J, et al. AKT1 and MYC induce distinctive metabolic fingerprints in human prostate cancer. Cancer Res. 2014;74(24):7198–204.
Munir R, Lisec J, Swinnen JV, Zaidi N. Lipid metabolism in cancer cells under metabolic stress. Br J Cancer. 2019;120(12):1090–8.
Tallima H, El Ridi R. Arachidonic acid: Physiological roles and potential health benefits—a review. J Adv Res. 2018;11:33–41.
Dhall S, Wijesinghe DS, Karim ZA, Castro A, Vemana HP, Khasawneh FT, Chalfant CE, Martins-Green M. Arachidonic acid-derived signaling lipids and functions in impaired healing. Wound Repair Regen. 2015;23(5):644–56.
Ghosh M, Tucker DE, Burchett SA, Leslie CC. Properties of the Group IV phospholipase A2 family. Prog Lipid Res. 2006;45(6):487–510.
Whitlock NC, Trostel SY, Wilkinson S, Terrigino NT, Hennigan ST, Lake R, Carrabba NV, Atway R, Walton ED, Gryder BE, et al. MEIS1 down-regulation by MYC mediates prostate cancer development through elevated HOXB13 expression and AR activity. Oncogene. 2020;39(34):5663–74.
Stelloo S, Nevedomskaya E, Kim Y, Hoekman L, Bleijerveld OB, Mirza T, Wessels LFA, van Weerden WM, Altelaar AFM, Bergman AM, et al. Endogenous androgen receptor proteomic profiling reveals genomic subcomplex involved in prostate tumorigenesis. Oncogene. 2018;37(3):313–22.
Astolfi A, Fiore M, Melchionda F, Indio V, Bertuccio SN, Pession A. BCOR involvement in cancer. Epigenomics. 2019;11(7):835–55.
Zhu S, Zhao D, Li C, Li Q, Jiang W, Liu Q, Wang R, Fazli L, Li Y, Zhang L, et al. BMI1 is directly regulated by androgen receptor to promote castration-resistance in prostate cancer. Oncogene. 2020;39(1):17–29.
Vieira FQ, Costa-Pinheiro P, Ramalho-Carvalho J, Pereira A, Menezes FD, Antunes L, Carneiro I, Oliveira J, Henrique R, Jerónimo C. Deregulated expression of selected histone methylases and demethylases in prostate carcinoma. Endocr Relat Cancer. 2014;21(1):51–61.
Sierra J, Yoshida T, Joazeiro CA, Jones KA. The APC tumor suppressor counteracts beta-catenin activation and H3K4 methylation at Wnt target genes. Genes Dev. 2006;20(5):586–600.
Sowalsky AG, Kissick HT, Gerrin SJ, Schaefer RJ, Xia Z, Russo JW, Arredouani MS, Bubley GJ, Sanda MG, Li W, et al. Gleason score 7 prostate cancers emerge through branched evolution of clonal Gleason pattern 3 and 4. Clin Cancer Res. 2017;23(14):3823–33.
Pomerantz MM, Qiu X, Zhu Y, Takeda DY, Pan W, Baca SC, Gusev A, Korthauer KD, Severson TM, Ha G, et al. Prostate cancer reactivates developmental epigenomic programs during metastatic progression. Nat Genet. 2020;52(8):790–9.
Cejas P, Xie Y, Font-Tello A, Lim K, Syamala S, Qiu X, Tewari AK, Shah N, Nguyen HM, Patel RA, et al. Subtype heterogeneity and epigenetic convergence in neuroendocrine prostate cancer. Nat Commun. 2021;12(1):5775.
Beshiri ML, Tice CM, Tran C, Nguyen HM, Sowalsky AG, Agarwal S, Jansson KH, Yang Q, McGowen KM, Yin J, et al. A PDX/organoid biobank of advanced prostate cancers captures genomic and phenotypic heterogeneity for disease modeling and therapeutic screening. Clin Cancer Res. 2018;24(17):4332–45.
Nguyen HM, Vessella RL, Morrissey C, Brown LG, Coleman IM, Higano CS, Mostaghel EA, Zhang X, True LD, Lam HM, et al. LuCaP prostate cancer patient-derived xenografts reflect the molecular heterogeneity of advanced disease and serve as models for evaluating cancer therapeutics. Prostate. 2017;77(6):654–71.
Biswas NK, Das S, Maitra A, Sarin R, Majumder PP. Somatic mutations in arachidonic acid metabolism pathway genes enhance oral cancer post-treatment disease-free survival. Nat Commun. 2014;5:5835.
Truica MI, Burns MC, Han H, Abdulkadir SA. Turning up the heat on MYC: progress in small-molecule inhibitors. Cancer Res. 2021;81(2):248–53.
Zirath H, Frenzel A, Oliynyk G, Segerstrom L, Westermark UK, Larsson K, Munksgaard Persson M, Hultenby K, Lehtio J, Einvik C, et al. MYC inhibition induces metabolic changes leading to accumulation of lipid droplets in tumor cells. Proc Natl Acad Sci USA. 2013;110(25):10258–63.
Carracedo A, Cantley LC, Pandolfi PP. Cancer metabolism: fatty acid oxidation in the limelight. Nat Rev Cancer. 2013;13(4):227–32.
Tawadros T, Brown MD, Hart CA, Clarke NW. Ligand-independent activation of EphA2 by arachidonic acid induces metastasis-like behaviour in prostate cancer cells. Br J Cancer. 2012;107(10):1737–44.
Zhao R, Lv Y, Feng T, Zhang R, Ge L, Pan J, Han B, Song G, Wang L. ATF6alpha promotes prostate cancer progression by enhancing PLA2G4A-mediated arachidonic acid metabolism and protecting tumor cells against ferroptosis. Prostate. 2022;82(5):617–29.
Labrecque MP, Alumkal JJ, Coleman IM, Nelson PS, Morrissey C. The heterogeneity of prostate cancers lacking AR activity will require diverse treatment approaches. Endocr Relat Cancer. 2021;28(8):T51–66.
Qiu X, Boufaied N, Hallal T, Feit A, de Polo A, Luoma AM, Alahmadi W, Larocque J, Zadra G, Xie Y, et al. MYC drives aggressive prostate cancer by disrupting transcriptional pause release at androgen receptor targets. Nat Commun. 2022;13(1):2559.
Di Tommaso P, Chatzou M, Floden EW, Barja PP, Palumbo E, Notredame C. Nextflow enables reproducible computational workflows. Nat Biotechnol. 2017;35(4):316–9.
Li H, Durbin R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics. 2010;26(5):589–95.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nusbaum C, Myers RM, Brown M, Li W, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9(9):R137.
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, Glass CK. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell. 2010;38(4):576–89.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29(1):15–21.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30(7):923–30.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26(1):139–40.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-Seq data. BMC Bioinformatics. 2013;14(1):7.
Shaffer AL, Wright G, Yang L, Powell J, Ngo V, Lamy L, Lam LT, Davis RE, Staudt LM. A library of gene expression signatures to illuminate normal and pathological lymphoid biology. Immunol Rev. 2006;210:67–85.
Mendiratta P, Mostaghel E, Guinney J, Tewari AK, Porrello A, Barry WT, Nelson PS, Febbo PG. Genomic strategy for targeting therapy in castration-resistant prostate cancer. J Clin Oncol. 2009;27(12):2022–9.
Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26(6):841–2.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15(12):550.
Zerbino DR, Johnson N, Juettemann T, Wilder SP, Flicek P. WiggleTools: parallel processing of large collections of genome-wide datasets for visualization and statistical analysis. Bioinformatics. 2014;30(7):1008–9.
Kent WJ, Zweig AS, Barber G, Hinrichs AS, Karolchik D. BigWig and BigBed: enabling browsing of large distributed datasets. Bioinformatics. 2010;26(17):2204–7.
Ramírez F, Ryan DP, Grüning B, Bhardwaj V, Kilpert F, Richter AS, Heyne S, Dündar F, Manke T. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 2016;44(W1):W160-165.
Krämer A, Green J, Pollard J Jr, Tugendreich S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics. 2014;30(4):523–30.
Nagatsugi F, Hokazono J, Sasaki S, Maeda M. Synthesis of 20-[18F]fluoroarachidonic acid: a potential phospholipid metabolic agent. J Labelled Compd Radiopharm. 1994;34(12):1121–7.
Acknowledgements
The authors gratefully acknowledge the patients and the families of patients who contributed to this study. Portions of this work utilized the computational resources of the NIH HPC Biowulf cluster.
Funding
Open Access funding provided by the National Institutes of Health (NIH). This work was supported by the Prostate Cancer Foundation (Young Investigator Awards to A.T.K and S.W.) the Department of Defense Prostate Cancer Research Program (W81XWH-19-1-0712 to S.W., W81XWH-16-1-0433 to A.G.S.), and the Intramural Research Program of the NIH, National Cancer Institute. These funding bodies had no role in the design of the study and collection, analysis, and interpretation of data, nor in writing the manuscript.
Author information
Authors and Affiliations
Contributions
Data acquisition: NCW, MEW, SA, LF, SW, SYT, KW, EMJ. Methodology: NCW, MEW, BJC, ATK, EMJ. Reagents: SA, LF, Z-DS, FB. Analysis: NCW, MEW, BJC, ATK, EMJ, AGS. Manuscript: NCW, MEW, AGS. Supervision: EMJ, KK, PLC, AGS. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The collection and analysis of tissue from patients with localized prostate cancer was approved by the Institutional Review Boards of the National Institutes of Health (protocol 15-c-0124), Beth Israel Deaconess Medical Center (protocol 2010-P-000254/0), and Dana-Farber/Harvard Cancer Center (protocols 11-250, 15-008, and 15-492). All patients provided informed consent before participating. This research was conducted in accordance with the principles of the Declaration of Helsinki. The propagation of patient-derived xenografts was approved by National Cancer Institute Animal Care and Use Committee.
Consent for publication
Not applicable.
Competing interests
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Whitlock, N.C., White, M.E., Capaldo, B.J. et al. Progression of prostate cancer reprograms MYC-mediated lipid metabolism via lysine methyltransferase 2A. Discov Onc 13, 97 (2022). https://doi.org/10.1007/s12672-022-00565-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-022-00565-3