Abstract
Aims
This research developed a prognostic model for OS patients based on the Mechanistic Target of Rapamycin Complex 1 (mTORC1) signature.
Background
The mTORC1 signaling pathway has a critical role in the maintenance of cellular homeostasis and tumorigenesis and development through the regulation of cell growth, metabolism and autophagy. However, the mechanism of action of this signaling pathway in Osteosarcoma (OS) remains unclear.
Objective
The datasets including the TARGET-OS and GSE39058, and 200 mTORC1 genes were collected.
Methods
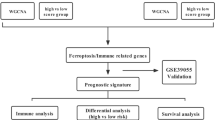
The mTORC1 signaling-related genes were obtained based on the Molecular Signatures Database (MSigDB) database, and the single sample gene set enrichment analysis (ssGSEA) algorithm was utilized in order to calculate the mTORC1 score. Then, the WGCNA were performed for the mTORC1-correlated gene module, the un/multivariate and lasso Cox regression analysis were conducted for the RiskScore model. The immune infiltration analysis was performed by using the ssGSEA method, ESTIMATE tool and MCP-Count algorithm. KM survival and Receiver Operating Characteristic (ROC) Curve analysis were performed by using the survival and timeROC package.
Results
The mTORC1 score and WGCNA with β = 5 screened the mTORC1 positively correlated skyblue2 module that included 67 genes, which are also associated with the metabolism and hypoxia pathways. Further narrowing of candidate genes and calculating the regression coefficient, we developed a useful and reliable RiskScore model, which can classify the patients in the training and validation set into high and low-risk groups based on the median value of RiskScore as an independent and robust prognostic factor. High-risk patients had a significantly poor prognosis, lower immune infiltration level of multiple immune cells and prone to cancer metastasis. Finally, we a nomogram model incorporating the metastasis features and RiskScore showed excellent prediction accuracy and clinical practicability.
Conclusion
We developed a useful and reliable risk prognosis model based on the mTORC1 signaling signature.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Osteosarcomas (OS) is the most popular primary malignancy that develops in bone and defined by the existing malignant mesenchymal cells generating immature or osteoid bone [1]. The incidence of OS in children and adolescence (especially 10–14 years old) [2] is quietly high that may be due to the rapid bone turnover and growth and its annual incidence peaks are at 8–11/million/year worldwide [3],. Osteosarcomas often occur in the long bones of extremities, near the metaphyseal growth plate including the humerus, femur and tibia, and less commonly in the pelvis, skull and jaw [4]. Following surgery, patients receive neo-adjuvant chemotherapy and chemotherapy treatment with a cocktail of chemotherapies [5], for instance, high-dose methotrexate (12 g/m2), ifosfamide and etoposide for younger patients in the sarcome-09 study [6], or combination of cisplatin, ifosfamide and doxorubicin with or without high-dose methotrexate [7]. This has been the standard of care for OS patients since the introduction of chemotherapies in the 1970s and the use of these therapeutic regimens has increased the overall 5-years survival rate of patients that had localized disease from 20 to 78% and the 10-year survival from 30–50% [8], but unfortunately, 20–30% patients are the recurrent and metastatic cases and their 5-year survival rate is less than 25% [9]. However, with the updated management strategies, such as several anti-carcinogens clinically application, the survival rate has not obviously improved for patients who has metastases or not over the last few decades [10]. A major reason of this poor prognosis is the high levels of tumor heterogeneity existing in the OS patients, and the complexity genetic and molecular mechanisms of osteosarcoma agenesis makes it quietly difficult to develop a singular effective therapeutic method in clinical practice. In addition, the stagnant survival rates of OS patients also indicate an urgent need for the better understanding of this disease, and the developing an effective approach of diagnosis and treatment by using the multiple combined, modern and interdisciplinary therapeutic regimens [11].
As a type of conserved serine/threonine kinase, mechanistic target of rapamycin kinase complex 1 (mTORC1) is mainly involved in the cell growth, proliferation, migration, immune responses, survival, autophagy, and metabolism regulation for maintaining cellular homeostasis [12]. MTORC1 forms two functionally distinct and structurally multi-subunit complexes of mTORC1 and mTORC2 that support mTOR signaling cascade [13]. mTORC1 consists of mLST8, Raptor, mTOR, PRAS40 and DEPTOR protein [14], and positively controls cell growth by stimulating protein and lipid synthesis, and its main downstream targets include the 4E binding protein 1 (4EBP1), insulin growth factor receptor (IGF-1R), transcription factor EB (TFEB), p70S6 kinase (S6K), protein kinase C and Unc-51-like autophagy-activating kinases [14, 15]. Dysregulations of mTORC1 signaling are associated with many disease, such as the neuronal disorders, diabetes, epilepsy and cancer [16]. Immunosuppressant rapamycin can inhibit the mTORC1 activity for tumor treatment, but is invalid for the mTORC1-dependent phosphorylation of 4EBP1 in some tumor cells [17]. The mTORC1 signaling can be activated by the EGFR/PI3K/Akt and IR/PI3K/Akt pathways [18] and phosphorylates S6K and 4EBP1 to initiate mRNA translation for cell growth and metabolism. Under physiological conditions, the mTORC1 signaling is tightly controlled, the loss of negative regulation caused the unrestrained cell growth in cancers [19], the recent clinical trials on tyrosine kinase inhibitors has revealed that the mTORC1 is a potential molecular target in osteosarcoma despite the single-agent of mTOR inhibitors is failure in osteosarcoma trials [20, 21]. Combinations of mTOR and other pathway inhibitors, such as the sorafenib with mTOR inhibition [22] suggested that molecular targets based on specific biomarkers are an advisable for the novel therapies developing.
Bioinformatics is a new interdisciplinary subject with huge development potential and can help to reveal the laws and mysteries of complex biological process through the comprehensive application of computer science, molecular biology and information technology. In this study, we developed a useful and reliable prognosis model for the risk stratification and personalized treatment decision of OS patients based on the mTORC1 signaling features. By WGCNA screening, we analyzed the gene module that associated with the mTORC1 score, these genes are enriched in the Glycolysis, Central carbon metabolism in cancer and response to hypoxia pathway. Patients were grouped into high and low risk groups using the RiskScore system, with high-risk patients having significantly poor prognosis. In addition, the higher RiskScore is prone to indicate the cancer metastasis and the lower immune infiltration of the Natural killer cell, Macrophage, activated CD8 T cell, Effector memory CD8 T cell, activated B cell, and Type 1 T helper cell. Finally, we developed a nomogram model combination of the RiskScore and metastatic feature for the precise prediction of patients’ probability of surviving. Our work provides a useful tool to optimize the treatment strategies.
2 Materials and methods
2.1 Data acquisition
The clinical information and RNA-seq of human osteosarcoma (OS) specimens were collected from the TARGET (https://ocg.cancer.gov/programs/target) database and it include 84 OS samples (TARGET-OS) with the encoding protein genes. Another dataset GSE39058 was downloaded from the Gene Expression Omnibus database (GEO, https:/ww.ncbinlm.nih.gov/geo/) and a total of filtering 42 OS samples were included in this study [23]. We obtained 200 mTORC1 signaling related genes from the Molecular Signatures Database (MSigDB, https://www.gsea-msigdb.org/gsea/msigdb).
2.2 The ssGSEA for MTORC1 score
The single sample gene set enrichment analysis (ssGSEA) is able to analyze each single tumor sample to be able to generate a score of the activity of a specific signaling pathway in that sample, which in turn helps to identify the specific activation of the signaling pathway in different samples [24]. The HALLMARK_MTORC1_SIGNALING.v2023.2 and h.all.v2023.2.Hs.symbols pathway score of patients were calculated by performing ssGSEA using the GSVA R package in the TARGET-OS cohort [25].
2.3 Weighted Gene Co-expression Network Analysis (WGCNA)
Gene module related to the mTORC1 singling signature was sectioned using WGCNA [26]. The pickSoftThreshold function was used to determine the soft threshold (β) ensuring the scale-free network, after that the hierarchical clustering was performed to screen the gene module that include at 60 genes. The correlation analysis between gene module and mTORC1 signature (score) were performed for the most correlated gene module.
2.4 Enrichment analysis
The function enrichment analysis of module genes was analyzed using the Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) with clusterProfiler R package [25].
2.5 Construction of RiskScore model and validation of prognostic value
To shrink the number of candidate gene, Least Absolute Shrinkage and Selection Operator (Lasso) Cox regression analysis was performed with the glmnet R package [25]. Subsequently, the multivariate Cox regression analysis was performed to determine the key risk and the regression coefficient for RiskScore construction, according to the formula: RiskScore =\(\:{\Sigma\:}{\upbeta\:}\text{i}\times\:\text{E}\text{x}\text{p}\text{i}\). (βi is the regression coefficient, Expi is the expression level of risk gene). The RiskScore of patients was calculated by using the RiskScore system and divided the patients into high- and low-risk groups based on the median value, then the Kaplan-Meier (KM) survival analysis was conducted by using the survival R package, the ROC was plotted by the timeROC R package for the classifier efficiency evaluation [27].
2.6 Tumor microenvironment (TME) analysis among various risk groups
The ssGSEA method of GSVA R package was used to analyze the immune infiltration score of 28 immune cells [28]. Subsequently, we used the ESTIMATE algorithm for use in assessing the composition of immune and stromal cells in the tumor microenvironment of OS patients. Specifically, by analyzing the gene expression data of the samples, immune scores, stromal scores and ESTIMATE scores were generated, which in turn helped to quantify the characteristics of the tumor microenvironment [29]. Meanwhile, the MCP-Count algorithm was used to analyze immune infiltration score of 10 immune cells. In addition, the progeny R package was used to calculate the tumorigenesis-related hallmark pathway score and the correlation between the RiskScore and these pathways was analyzed [30].
2.7 Identifying the independent prognostic factor and developing a nomogram
The decision-making tree model usually used for a nonparametric supervised learning algorithm for classification task and was constructed by using the rpart R package [31]. To select significant and independent prognostic factors, we performed univariate and multivariate Cox regression analysis. The rms R package was used to construct a developing nomogram model that combined various clinical factors, then the predictive accuracy of nomogram model was assessed by the calibration curve [32], and the decision curve was performed for the practical benefit evaluation in clinical practice.
2.8 Statistical tests
Statistical analysis was performed in the R software (version3.6.0). Difference between two sets of continuous variables was analyzed using the wilcoxon rank-sum test. The spearman method and log-rank test were used for correlation analysis and survival difference comparison among various risk patients. A p < 0.05 was statistically significant.
3 Results
3.1 Skyblue2 is the most correlated module with the mTORC1 signature
The mTORC1 score of each patient was computed based on the 200 mTORC1 related genes expression, and the WGCNA identified the optimal soft thresholdβ is 5 (Fig. 1A) and obtained 57 co-expression modules after hierarchical clustering and module merging, in which the grey is an invalid module that is not aggregated to other modules (Fig. 1B). The correlation analysis showed that the skyblue2 is the most correlated module with the mTORC1 score (p < 0.05, Fig. 1C) and contained 67 genes (Fig. 1D). Function enrichment analysis revealed that these 67 module genes were closely related to the Glycolysis/Gluconeogenesis, Fructose and mannose metabolism, Galactose metabolism pathway, Central carbon metabolism in cancer, HIF-1 signaling pathway, and Carbon metabolism in KEGG (Fig. 1E) and closely associated with the metabolism and hypoxia biological process (BP) including the the response to decreased oxygen levels process, glycolytic process through fructose-6-phosphate, canonical glycolysis, response to hypoxia, pyruvate metabolic process (Fig. 1F). These findings suggested that the mTORC1 signaling pathway involved in the cancer progression through altering the metabolism- and hypoxia-related pathways.
WGCNA for the mTORC1 signature related gene module. A Scale-free fitting index analysis of various soft threshold power (β). B Gene tree map based on dissimilarity measure (1-TOM) clustering. C The correlation heatmap between gene module and mTORC1 score. D The number of genes in each module. E KEGG enrichment analysis of skyblue2 module genes. F Biological process of GO enrichment analysis of skyblue2 module genes
3.2 The established RiskScore is a reliable prognostic classification model
The univariate Cox regression analysis was used to determine the significant prognostic genes (p < 0.05) in 67 module genes, the lasso Cox regression analysis was used to reduce the numbers of candidate gene (Fig. 2A) and multivariate Cox regression analysis was used for RiskScore construction (Fig. 2B), Riskscore=\(\:\left(0.157\text{*}\text{A}\text{N}\text{K}\text{R}\text{D}37\right)+\left(0.226\text{*}\text{B}\text{N}\text{I}\text{P}3\right)+\left(0.556\text{*}\text{P}\text{D}\text{E}4\text{C}\right)+\left(0.162\text{*}\text{P}\text{D}\text{K}1\right)+\left(0.343\text{*}\text{S}\text{E}\text{R}\text{P}\text{I}\text{N}\text{E}2\right).\) The patients were classified into low- and high-risk groups, with the high-risk patients having noticeably poor prognosis (p < 0.05, Fig. 2C), the ROC analysis revealed that the AUC values of 1-, 3- and 5-years survival rate is high, which are 0.75, 0.77 and 0.76 respectively (Fig. 2D), suggesting the RiskScore is a good classifier in long- and short-term prognosis. In addition, the Principal Component Analysis (PCA) also exhibited that the high- and low-risk patients had clearly boundary (Fig. 2E), we visualized the expression of the model genes and found that these genes as risk factors were significantly overexpressed in the high-risk groups (p < 0.05, Fig. 2F). To verify the model robustness, we analyzed the prognosis and classifier efficiency of RiskScore in the validation set (GSE39058), which is important in the analysis biomarker [33]. The results showed that the patients in the high-risk groups had significantly poor prognosis (p < 0.05, Fig. 2G) and the AUC of 1-, 3- and 5- years survival rate is higher (more than 0.7), which are 0.74, 0.8 and 0.77 respectively (Fig. 2H), revealing the highly reliable of RiskScore model.
Establishing and validating prognostic risk models A The trajectory of each independent variable changing with lambda and the confidence interval under lambda. B Distribution of coefficients of the prognostic gene signatures. C KM survival analysis of patients in TARGET-OS cohort. D ROC analysis of patients in TARGET-OS cohort. E Principal Component Analysis of patients in TARGET-OS cohort. F The expression difference of model genes in various risk groups. G KM survival analysis of patients in GSE39058 cohort. H ROC analysis of patients in GSE39058 cohort
3.3 The higher RiskScore may be a cancer metastasis-related risk factor
We compared the distributional difference of RiskScore in various clinicopathologic features and found that the patients with ≤ 15 years old had higher ratio (Fig. 3A), the male low-risk patients had higher ratio (Fig. 3B) and most patients in low-risk are no metastatic (Fig. 3C). After that, further analysis revealed that the RiskScore is not affected the age (Fig. 3D) and gender distribution (Fig. 3E) of patients, but affected the metastatic distribution of patients, in which the patients with higher RiskScore are significantly prone to cancer metastasis (Fig. 3F). These results further suggest that the RiskScore may be an important indicator of poor prognosis in patients with OS.
Riskscore distribution characteristics among various clinicopathologic features A The age distribution of patients in high and low risk group. B The gender distribution of patients in high and low risk group. C The metastatic distribution of patients in high and low risk group. D The RiskScore difference among various age groups. E The RiskScore difference among various gender groups. F The RiskScore difference among various metastatic groups
3.4 High-risk patients are related to a suppressive TME
Differences of the tumor microenvironment (TME) among high and low risk groups were evaluated, the ESTIMATE results demonstrated poor immune infiltration including the overall significantly lower ESTIMATE, stroma, immune scores in high-risk group (p < 0.05, Fig. 4A). The immune cells, such as the CD8 T cells, T cells, Monocytic lineage and endothelial cells are significantly high infiltration levels in the low-risk group (p < 0.05, Fig. 4B). In addition, the activated B and CD8 T cells, effector memory CD8 T cells, immature B cells, memory B cells, Type 1 T helper cells, macrophage, central memory CD8 T cell, MDSC, natural killer cells of others 28 immune cells also had significantly higher infiltration in the low-risk group (p < 0.05, Fig. 4C), suggested that the high-risk patients are associated with the suppressive TME.
Immune infiltration difference among high- and low-risk groups A The ESTIMATE of immune infiltration among high- and low-risk groups. B MCP-counter of 10 immune cell infiltration in the high- and low-risk groups. C ssGSEA of 28 immune cell infiltration in the high- and low-risk groups. (* p < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001)
3.5 Hypoxia and mTORC1 signaling are crucial for promoting OS progression
We calculated the tumorigenesis-related hallmark pathway score and their correlation with the RiskScore, and found that the Hypoxia pathway is significant positively correlated with the RiskScore, and the JAK-STAT pathway is significant negatively correlated with the RiskScore (Fig. 5A). The Wilcoxon rank sum test identified that the pathways of glycolysis, hypoxia and mTORC1 signaling had significant higher activity in the high-risk group (p < 0.05, Fig. 5B), indicating these pathways including the mTORC1 signaling are crucial for promoting OS progression.
3.6 RiskScore is key independent prognostic factor and supported a reliable nomogram model
Finally, we constructed a decision-making tree model based on the age, gender, metastatic and Risktype features in the TARGET-OS cohort, the model showed that the classification ability of RiskType is more effective than other clinical features, followed by the metastatic feature, and four risk subgroups were obtained (Fig. 6A), in which the C1 subgroup is the significant good prognosis (p < 0.05, Fig. 6B), the C1 and C2 group were categorized into the high-risk group (Fig. 6C). Subsequently, the univariate Cox regression analysis showed that the metastatic and RiskScore features are the significant prognostic factors (p < 0.05, Fig. 6D) and the multivariate Cox regression analysis showed that they are independent prognostic factors (p < 0.05, Fig. 6E). To quantify the survival probability of patients, we combined the metastatic and RiskScore features to develop a nomogram model (Fig. 6F), which indicated that the RiskScore has the greatest influence on survival prediction. The calibration curve showed that the prediction curve of 1-, 3- and 5-years calibration points is closely coincided with the standard curve (Fig. 6G), suggesting that the nomogram has excellent prediction accuracy. The decision curve revealed that the net benefit of RiskScore and nomogram model is obvious higher than the extreme curve (Fig. 6H), indicating that our model had excellent clinical practicability. ROC analysis also demonstrated that the nomogram model is an excellent short- and long-term prognostic classifier.
Analysis of independent prognostic factors and a nomogram developing A Decision-making tree including Risktype, Metastatic, and age for risk stratification. B KM survival analysis among four risk subgroups. C The risk distribution among four risk subgroups. D Univariate Cox regression analysis for significant prognostic factors. E Multivariate Cox regression analysis for independent prognostic factors. F A developing nomogram model was constructed. G The 1-, 3- and 5-year calibration curves of the nomogram. H The decision curve of nomogram model. I ROC analysis of nomogram model
4 Discussion
Osteosarcoma (OS) is a popular primary malignant bone tumor with early metastasis, high-grade aggressive and poor prognosis features in adolescents [34], meanwhile its complex heterogeneity generated a various of patients with different subtypes that exhibited unique immune response pattern, tumor microenvironment, genotypes and phenotypes at the macroscopic and microscopic aspects [35]. At present, the surgery with adjuvant chemotherapy is the primary means for osteosarcoma, but not benefit to all patients that highlighted the fact of many patients do not respond to the standard therapies [36]. In addition, several chemo-resistant cell models have been well constructed and many differentially expressed markers and noncoding RNAs were identified [37, 38]. Hashimoto et al. NY-ESO-1 and MAGE-A4 expression may be associated with the immune status in the tumor microenvironment and highlighted that identification of validated biomarkers in OS has a key role in patient prognostic assessment and treatment stratification [39]. This study analyzed the actual variation among osteosarcoma patients with differing intrinsic mechanisms and outcomes to develop novel therapeutic targets and strategy for those with a dismal prognosis. This study constructed a useful and reliable prognostic model to screen the high-risk patients and offer the tailoring supplementary treatments for improving prognosis based on the mTORC1-related signature.
Bone sarcomas, especially the osteosarcoma develop in a highly dynamic bone microenvironment consisting of bone cells (osteocytes, osteoblasts, and osteoclasts), stromal cells (mesenchymal stem/stromal cells (MSCs), fibroblasts), vascular cells (pericytes and endothelial cells), a mineralized extracellular matrix (ECM), and immune cells (lymphocytes, macrophages) [40]. A fine-tuned orchestrated activity of stromal, vascular and bone cells maintain the bone homeostasis through the cellular communications (autocrine and paracrine). However, the chromosomal aneuploidy and gene mutation initiate tumorigenesis, the tumor cells will highjack bone physiological pathways for their survive and grow [41]. Osteosarcoma also is highly vascularized bone tumors and was surrounded by an acidic and hypoxic bone microenvironment [42], the RiskScore and the genes in the skyblue2 module are closely associated with the Hypoxia signaling and the hypoxia-induced factor-1 (HIF-1) pathway, which may be further promote the angiogenesis [43], thus the drugs of anti-angiogenesis may benefit to the high-risk OS patients, such as the sorafenib alone [44], or combined with everolimus (an mTOR2 inhibitor) [22]. In addition, the tumor-associated macrophages (TAMs) is another major component of OS microenvironment, with a number of myeloid, dendritic and lymphoid cells [45]. Increased infiltration of TAMs is related to less active metastasis and longer survival as compared to epithelial tumors [46, 47]. The mechanism of TAMs inhibiting metastasis in OS is unclear, different teams in precise heterogeneous population case found and suggested that M2 macrophages exert an anti-metastatic effect rather anti-inflammatory [47], the macrophage-activating agent mifamurtide significantly improved the 6-year overall survival in a clinical trial [48]. MSCs and MSC-derived osteoblasts are sensors of their microenvironment, they not only express multiple signaling receptors, but also secrete ECM components, such as the cytokines and metalloproteinases (MMPs) to modulate tumor microenvironment [49]. The tumor infiltration of immune cells is strictly dependent on the proteolysis and plasticity of ECM [50], Nicolas-Boluda et al. demonstrated that T cells can cross the blood vessel but was trapped by the tumor nodules [51]. T cells do not release the MMPs to lyse the matrix and cannot progress through the dense and tight fibers [52], this may be explained that the T cells are significantly decrease in the high-risk patients, thus induced proteolysis may be a key event to achieving immune-cell therapy success in osteosarcoma.
These model genes were defined as risk factors in the multivariate Cox regression analysis. Ankyrin repeat domain protein 37 (ANKRD37) is a hypoxia-associated protein and was up-regulated in colon cancer indicating poorer survival rate [53]. BNIP3 is another hypoxia-associated mitophagy protein, Vara-Pérez el. reported that the elevated BNIP3 levels as pro-tumorigenic regulator in melanoma is associated with poorer patient’s survival, the depletion of BNIP3 can compromise tumor growth in vivo [54]. However, the deletion or silencing of BNIP3 in the chronic kidney disease can significantly increase the mitochondrial damage, the production of mitochondrial ROS and activation of the NLRP3 inflammasome, demonstrating that the BNIP3- mediated mitophagy played a crucial protective role against hypoxia-induced cell injury [55], these evidences indicated that the protective role of BNIP3 is specific to some disease. Phosphodiesterase type 4 C (PDE4C) is a key messenger that specifically hydrolyze cAMP in cell signaling systems, Wright el. reviewed the deleterious or protective role in different cancer, where PDE activity can facilitate cancer via avoiding cAMP-triggered cell cycle arrest or the regulation of cAMP prevent cancer initiation [56], and it can be a potential cancer treatment target. The PDK1 was thought as master kinase with autophosphorylation to involve in the cell growth, invasion, metastasis and apoptosis regulation through activating the PI3K-AKT signaling pathways, the overexpression of PDK1 was observed in a plethora of cancers, such as, while the silenced PDK1 can inhibit tumor growth and proliferation through modulating tumor microenvironments and affecting tumor immunotherapies [57]. SERPINE2 can promote the radio-resistance of lung cancer cells and knockdown of SERPINE2 improve the tumor radio-sensitivity in vitro and in vivo [58], Chen el. reported that SERPINE2 could be acted as a target for metastasis in advanced renal cell carcinoma [59]. These results suggest that the key genes we identified may be aberrantly expressed in OS, which in turn may promote aggressiveness and drug resistance in OS and worsen patient prognosis by affecting hypoxia response, metabolic regulation and signaling pathways. Meanwhile, the regulatory effects of these genes on the tumor microenvironment may further support the progression of OS, suggesting their importance as potential therapeutic targets in OS research and treatment.
There also had some limitations in our study. First, the small sample size of this study may not be sufficient to represent the diversity of the entire OS patient population; in the future, we will add data from multiple centers and different populations to improve the reliability and generalizability of the results. In addition, this study lacks in-depth experimental validation to explore the specific mechanism of action of these genes in OS. Therefore, we will further combine functional experiments, including gene knockout and animal models, to be able to provide a potential molecular basis for OS-targeted therapy.
5 Conclusion
The inappropriate diagnosis and sub-optimal therapy can be irrevocably reduce the survival probability, the prospective clinical trials and prognostic tool will perfect the standard practice of OS treatment. This study developed a mTORC1 signaling-related risk prognostic model that acted as an independent prognostic factor for osteosarcoma. Our study offers a useful tool to predict the prognosis and indicate the immune infiltration, and may be provide potential treatment targets for immunotherapy.
Data availability
The datasets generated and/or analyzed during the current study are available in the [GSE39058] repository, [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi? acc= GSE39058].
Abbreviations
- OS:
-
Osteosarcoma
- WGCNA:
-
Weighted Gene Co-expression Network Analysis
- ROC:
-
Receiver operating characteristic curve
- TME:
-
Tumor microenvironment
- mTORC1:
-
Mechanistic target of rapamycin kinase complex 1
- TARGET:
-
Therapeutically Applicable Research to Generate Effective Treatments
- GEO:
-
Gene Expression Omnibus database
- MSigDB:
-
Molecular Signatures Database
- ssGSEA:
-
Single sample gene set enrichment analysis
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- Lasso:
-
Least Absolute Shrinkage and Selection Operator
- MSCs:
-
Mesenchymal stem/stromal cells
- ECM:
-
Extracellular matrix
- HIF-1:
-
Hypoxia-induced factor-1
- TAMs:
-
Tumor-associated macrophages
- MMPs:
-
Metalloproteinases
References
Sbaraglia M, Bellan E, Dei Tos AP. The 2020 WHO classification of soft tissue tumours: news and perspectives. Pathologica. 2021;113:70–84.
Pritchard-Jones K, Kaatsch P, Steliarova-Foucher E, Stiller CA, Coebergh JW. Cancer in children and adolescents in Europe: developments over 20 years and future challenges. Eur J Cancer. 2006;42:2183–90.
Ritter J, Bielack SS. Osteosarcoma. Ann Oncol. 2010;21(Suppl 7):vii320-325.
Kansara M, Teng MW, Smyth MJ, Thomas DM. Translational biology of osteosarcoma. Nat Rev Cancer. 2014;14:722–35.
Thanindratarn P, Dean DC, Nelson SD, Hornicek FJ, Duan Z. Advances in immune checkpoint inhibitors for bone sarcoma therapy. J Bone Oncol. 2019;15:100221.
Gaspar N, Occean BV, Pacquement H, Bompas E, Bouvier C, Brisse HJ, et al. Results of methotrexate-etoposide-ifosfamide based regimen (M-EI) in osteosarcoma patients included in the French OS2006/sarcome-09 study. Eur J Cancer. 2018;88:57–66.
Smrke A, Anderson PM, Gulia A, Gennatas S, Huang PH, Jones RL. Future directions in the treatment of osteosarcoma. Cells. 2021;10:172.
Berner K, Johannesen TB, Berner A, Haugland HK, Bjerkehagen B, Bøhler PJ, et al. Time-trends on incidence and survival in a nationwide and unselected cohort of patients with skeletal osteosarcoma. Acta Oncol. 2015;54:25–33.
Wedekind MF, Wagner LM, Cripe TP. Immunotherapy for osteosarcoma: where do we go from here? Pediatr Blood Cancer. 2018;65:e27227.
Lussier DM, Johnson JL, Hingorani P, Blattman JN. Combination immunotherapy with α-CTLA-4 and α-PD-L1 antibody blockade prevents immune escape and leads to complete control of metastatic osteosarcoma. J Immunother Cancer. 2015;3:21.
Mirabello L, Troisi RJ, Savage SA. International osteosarcoma incidence patterns in children and adolescents, middle ages and elderly persons. Int J Cancer. 2009;125:229–34.
Inoki K, Ouyang H, Li Y, Guan KL. Signaling by target of rapamycin proteins in cell growth control. Microbiol Mol Biol Rev. 2005;69:79–100.
Szwed A, Kim E, Jacinto E. Regulation and metabolic functions of mTORC1 and mTORC2. Physiol Rev. 2021;101:1371–426.
Panwar V, Singh A, Bhatt M, Tonk RK, Azizov S, Raza AS, et al. Multifaceted role of mTOR (mammalian target of rapamycin) signaling pathway in human health and disease. Signal Transduct Target Ther. 2023;8:375.
Ma XM, Blenis J. Molecular mechanisms of mTOR-mediated translational control. Nat Rev Mol Cell Biol. 2009;10:307–18.
Saxton RA, Sabatini DM. mTOR signaling in growth, metabolism, and disease. Cell. 2017;169:361–71.
Thoreen CC, Kang SA, Chang JW, Liu Q, Zhang J, Gao Y, et al. An ATP-competitive mammalian target of rapamycin inhibitor reveals rapamycin-resistant functions of mTORC1. J Biol Chem. 2009;284:8023–32.
Zhu M, Wang XQ. Regulation of mTORC1 by small GTPases in response to nutrients. J Nutr. 2020;150:1004–11.
Yuskaitis CJ, Jones BM, Wolfson RL, Super CE, Dhamne SC, Rotenberg A, et al. A mouse model of DEPDC5-related epilepsy: neuronal loss of Depdc5 causes dysplastic and ectopic neurons, increased mTOR signaling, and seizure susceptibility. Neurobiol Dis. 2018;111:91–101.
Ding L, Congwei L, Bei Q, Tao Y, Ruiguo W, Heze Y, et al. mTOR: an attractive therapeutic target for osteosarcoma? Oncotarget. 2016;7:50805–13.
Italiano A, Mir O, Mathoulin-Pelissier S, Penel N, Piperno-Neumann S, Bompas E, et al. Cabozantinib in patients with advanced ewing sarcoma or osteosarcoma (CABONE): a multicentre, single-arm, phase 2 trial. Lancet Oncol. 2020;21:446–55.
Grignani G, Palmerini E, Ferraresi V, D’Ambrosio L, Bertulli R, Asaftei SD, et al. Sorafenib and everolimus for patients with unresectable high-grade osteosarcoma progressing after standard treatment: a non-randomised phase 2 clinical trial. Lancet Oncol. 2015;16:98–107.
Kelly AD, Haibe-Kains B, Janeway KA, Hill KE, Howe E, Goldsmith J, et al. MicroRNA paraffin-based studies in osteosarcoma reveal reproducible independent prognostic profiles at 14q32. Genome Med. 2013;5:2.
Yi M, Nissley DV, McCormick F, Stephens RM. ssGSEA score-based Ras dependency indexes derived from gene expression data reveal potential Ras addiction mechanisms with possible clinical implications. Sci Rep. 2020;10:10258.
Zhao P, Zhen H, Zhao H, Huang Y, Cao B. Identification of hub genes and potential molecular mechanisms related to radiotherapy sensitivity in rectal cancer based on multiple datasets. J Transl Med. 2023;21:176.
Song Z, Yu J, Wang M, Shen W, Wang C, Lu T, et al. CHDTEPDB: transcriptome expression profile database and interactive analysis platform for congenital heart disease. Congenit Heart Dis. 2023;18:693–701.
Deng X, Thompson JA. An R package for survival-based gene set enrichment analysis. Res Sq. 2023. https://doi.org/10.21203/rs.3.rs-3367968/v1.
Hänzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 2013;14:7.
Yoshihara K, Shahmoradgoli M, Martínez E, Vegesna R, Kim H, Torres-Garcia W, et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat Commun. 2013;4:2612.
Zhao S, Liu Q, Li J, Hu C, Cao F, Ma W, et al. Construction and validation of prognostic regulation network based on RNA-binding protein genes in lung squamous cell carcinoma. DNA Cell Biol. 2021;40:1563–83.
Robinson M, Johnson AM, Fischer LK, MacKenzie HM. Two symptoms to triage acute concussions: using decision tree modeling to predict prolonged recovery after a concussion. Am J Phys Med Rehabil. 2022;101:135–8.
Tian S, Jiang Y, Wang Y, Song J. Effects of dexmedetomidine combined with remifentanil anesthesia and propofol combined with remifentanil anesthesia on perioperative inflammatory response and pulmonary function in patients with lung cancer. Oncologie. 2020;22:13–21.
Behrouz S, Abolghasem J. Importance of Method Validation in the Analysis of Biomarker Current Pharmaceutical Analysis 2022;18(6):567–569. https://doi.org/10.2174/1573412918666211213142638.
Morrow JJ, Bayles I, Funnell APW, Miller TE, Saiakhova A, Lizardo MM, et al. Positively selected enhancer elements endow osteosarcoma cells with metastatic competence. Nat Med. 2018;24:176–85.
Pinder M, CharafEddine A, Parnell AS, DiBardino DJ, Knudson JD. Osteosarcoma with cardiac metastasis in a 22-year-old man: a case report and review of cardiac tumors. Congenit Heart Dis. 2014;9:E147-52.
Smith MA, Seibel NL, Altekruse SF, Ries LA, Melbert DL, O’Leary M, et al. Outcomes for children and adolescents with cancer: challenges for the twenty-first century. J Clin Oncol. 2010;28:2625–34.
Wang Y, Zeng L, Liang C, Zan R, Ji W, Zhang Z, et al. Integrated analysis of transcriptome-wide m(6)a methylome of osteosarcoma stem cells enriched by chemotherapy. Epigenomics. 2019;11:1693–715.
Zhu KP, Zhang CL, Ma XL, Hu JP, Cai T, Zhang L. Analyzing the interactions of mRNAs and ncRNAs to predict competing endogenous RNA networks in osteosarcoma chemo-resistance. Mol Ther. 2019;27:518–30.
Hashimoto K, Nishimura S, Ito T, Oka N, Kakinoki R, Akagi M. Clinicopathological assessment of cancer/testis antigens NY–ESO–1 and MAGE–A4 in osteosarcoma. Eur J Histochem. 2022;66:3377.
Molina ER, Chim LK, Barrios S, Ludwig JA, Mikos AG. Modeling the tumor microenvironment and pathogenic signaling in bone sarcoma Tissue Eng Part B Rev. 2020;26:249–71.
Paget S. The distribution of secondary growths in cancer of the breast. Cancer Metastasis Rev. 1989;8:98–101.
Huang X, Wang L, Guo H, Zhang W, Shao Z. Single-cell transcriptomics reveals the regulative roles of cancer associated fibroblasts in tumor immune microenvironment of recurrent osteosarcoma. Theranostics. 2022;12:5877–87.
Ramasamy SK. Structure and functions of blood vessels and vascular niches in bone. Stem Cells Int. 2017;2017:5046953.
Grignani G, Palmerini E, Dileo P, Asaftei SD, D’Ambrosio L, Pignochino Y, et al. A phase II trial of sorafenib in relapsed and unresectable high-grade osteosarcoma after failure of standard multimodal therapy: an Italian Sarcoma Group study. Ann Oncol. 2012;23:508–16.
Inagaki Y, Hookway E, Williams KA, Hassan AB, Oppermann U, Tanaka Y, et al. Dendritic and mast cell involvement in the inflammatory response to primary malignant bone tumours. Clin Sarcoma Res. 2016;6:13.
Dumars C, Ngyuen JM, Gaultier A, Lanel R, Corradini N, Gouin F, et al. Dysregulation of macrophage polarization is associated with the metastatic process in osteosarcoma. Oncotarget. 2016;7:78343–54.
Gomez-Brouchet A, Illac C, Gilhodes J, Bouvier C, Aubert S, Guinebretiere JM, et al. CD163-positive tumor-associated macrophages and CD8-positive cytotoxic lymphocytes are powerful diagnostic markers for the therapeutic stratification of osteosarcoma patients: an immunohistochemical analysis of the biopsies fromthe French OS2006 phase 3 trial. Oncoimmunology. 2017;6:e1331193.
Meyers PA. Muramyl tripeptide (mifamurtide) for the treatment of osteosarcoma. Expert Rev Anticancer Ther. 2009;9:1035–49.
Ponte AL, Marais E, Gallay N, Langonné A, Delorme B, Hérault O, et al. The in vitro migration capacity of human bone marrow mesenchymal stem cells: comparison of chemokine and growth factor chemotactic activities. Stem Cells. 2007;25:1737–45.
Friedl P, Alexander S. Cancer invasion and the microenvironment: plasticity and reciprocity. Cell. 2011;147:992–1009.
Nicolas-Boluda A, Donnadieu E. Obstacles to T cell migration in the tumor microenvironment. Comp Immunol Microbiol Infect Dis. 2019;63:22–30.
Watanabe R, Maeda T, Zhang H, Berry GJ, Zeisbrich M, Brockett R, et al. MMP (Matrix Metalloprotease)-9-Producing monocytes enable T cells to invade the Vessel Wall and cause Vasculitis. Circ Res. 2018;123:700–15.
Deng M, Zhang W, Yuan L, Tan J, Chen Z. HIF-1a regulates hypoxia-induced autophagy via translocation of ANKRD37 in colon cancer. Exp Cell Res. 2020;395:112175.
Vara-Pérez M, Rossi M, Van den Haute C, Maes H, Sassano ML, Venkataramani V, et al. BNIP3 promotes HIF-1α-driven melanoma growth by curbing intracellular iron homeostasis. Embo j. 2021;40:e106214.
Li J, Lin Q, Shao X, Li S, Zhu X, Wu J, et al. HIF1α-BNIP3-mediated mitophagy protects against renal fibrosis by decreasing ROS and inhibiting activation of the NLRP3 inflammasome. Cell Death Dis. 2023;14:200.
Wright TA, Gemmell AO, Tejeda GS, Blair CM, Baillie GS, Cancer. Phosphodiesterase type 4 C (PDE4C), the forgotten subfamily as a therapeutic target. Int J Biochem Cell Biol. 2023;162:106453.
Zheng N, Wei J, Wu D, Xu Y, Guo J. Master kinase PDK1 in tumorigenesis. Biochim Biophys Acta Rev Cancer. 2023;1878:188971.
Zhang J, Wu Q, Zhu L, Xie S, Tu L, Yang Y, et al. SERPINE2/PN-1 regulates the DNA damage response and radioresistance by activating ATM in lung cancer. Cancer Lett. 2022;524:268–83.
Chen WJ, Dong KQ, Pan XW, Gan SS, Xu D, Chen JX, et al. Single-cell RNA-seq integrated with multi-omics reveals SERPINE2 as a target for metastasis in advanced renal cell carcinoma. Cell Death Dis. 2023;14:30.
Acknowledgements
Not applicable.
Funding
This study was supported by Natural Science Foundation of Chongqing (No. cstc2020jcyj-msxmX0966).
Author information
Authors and Affiliations
Contributions
All authors contributed to this present work: [HXC] and [WW] designed the study, [SCC] acquired the data. [XPH] and [WW] improved the figure quality. [NW] and [HXC] drafted the manuscript, [HXC] and [SCC] revised the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethical approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, H., Wang, W., Chang, S. et al. A useful mTORC1 signaling-related RiskScore model for the personalized treatment of osteosarcoma patients by using the bulk RNA-seq analysis. Discov Onc 15, 418 (2024). https://doi.org/10.1007/s12672-024-01301-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12672-024-01301-9