Abstract
Bayesian optimization is incorporated into genomic prediction to identify the best genotype from a candidate population. Several expected improvement (EI) criteria are proposed for the Bayesian optimization. The iterative search process of the optimization consists of two main steps. First, a genomic BLUP (GBLUP) prediction model is constructed using the phenotype and genotype data of a training set. Second, an EI criterion, estimated from the resulting GBLUP model, is employed to select the individuals that are phenotyped and added to the current training set to update the GBLUP model until the sequential observed EI values are less than a stopping tolerance. Three real datasets are analyzed to illustrate the proposed approach. Furthermore, a detailed simulation study is conducted to compare the performance of the EI criteria. The simulation results show that one augmented version derived from the distribution of predicted genotypic values is able to identify the best genotype from a large candidate population with an economical training set, and it can therefore be recommended for practical use. Supplementary materials accompanying this paper appear on-line.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Accelerating the genetic improvement of crops through innovative breeding techniques and strategies is an urgent and important task when dealing with the problem of food security (2010). Owing to the long history of domestication and selection in agriculture, elite breeding populations or commercial cultivars have lost their genetic diversity (Reif et al. 2005). Thus, introgression of rich variation from wild, exotic, or indigenous germplasm has become essential to promote genetic diversity and to enhance the efficiency of plant breeding programs (McCouch et al. 2013). Plant breeders will therefore be faced with the need to identify superior accessions from germplasm collections. In this study, we attempt to provide a solution to this problem via genomic prediction (GP).
GP, which has become an increasingly popular approach in plant or animal breeding, takes advantage of dense DNA markers over an entire genome to predict genotypic values and then applies these to select superior individuals from genetic resources (Meuwissen et al. 2001). Typically, a training set with known genotype and phenotype data is employed to construct a GP model. The trained GP model is then used to predict genomic estimated breeding values (GEBVs) for the individuals of a breeding population with known genotype data. A linear regression model fitting the phenotypic values by marker effects is often implemented for GP. However, it is challenging to estimate all the regression coefficients in such a GP model, because the number of marker effects is usually much greater than the number of observed phenotypic values (the large-p-with-small-n problem). Therefore, shrinkage methods such as ridge regression, the least absolute shrinkage and selector operator (LASSO) (Tibshirani 1996), and elastic nets have been applied to estimate marker effects in the whole-genome regression model. Also, there are two kinds of linear mixed effects models for GP: (i) best linear unbiased prediction (BLUP) based on markers and (ii) BLUP based on a relationship matrix. In a marker-based model, marker effects are treated as random effects and GEBVs of individuals are obtained by multiplying their marker scores by these BLUP estimates. The ridge regression BLUP (rr-BLUP) model (Meuwissen et al. 2001) adopts this approach. In a relationship matrix-based model, genotypic values of individuals are treated directly as random effects and estimated through a genomic relationship matrix. The genomic BLUP (GBLUP) model (Habier et al. 2007) is in the spirit. Tempelman (2015) provided an in-depth review and discussion of the above GP models and noted that great care should be taken in correctly inferring hyperparameters using hierarchical Bayesian models in GP.
In the genomic era, genotyping costs have fallen dramatically, while phenotyping costs have remained relatively constant (Akdemir and Sanchez 2019). It would therefore be advantageous to sample individuals for selective phenotyping in more than one stage, because a sequential phenotyping approach can reduce the size of the required training set. Recently, Tanaka and Iwata (2018) proposed such an approach to discover the best genotype from a candidate population. They developed their procedure by incorporating Bayesian optimization into GP. Bayesian optimization is a powerful tool for solving black-box global optimization problems with computationally expensive function evaluation (Jones et al. 1998). The classical Bayesian optimization problem can be formulated as follows. The goal of the Bayesian optimization is to maximize or minimize an objective function \(f(\varvec{x})\) within a feasible space \(\Omega \). That is, let \(\varvec{x}^*\) be the desired optimum; then,
Also, let y denote the response of an experiment conducted to evaluate the objective function; then
where e denotes the random error. The experiment is said to be noisy if the response is measured with random error. Typically, the evaluation function is deterministic (fixed effect), and the feasible space is a continuous and compact region.
The optimization usually begins by evaluating a small number of randomly selected points and fitting a Gaussian process to the response. The Gaussian posterior provides an estimate of the object function at each point in the feasible space, as well as the uncertainty of this estimate (Letham et al. 2019). An acquisition function, such as the expected improvement (EI), is constructed based on the posterior estimation for determining new query points to evaluate the object function. The choice of the new query points should be a trade-off between exploration and exploitation so that one can optimize the objective function using as few query points as possible (Gong et al. 2019). For the classical Bayesian optimization problem of a continuous and compact feasible region, except in a few atypical cases, the EI has been proved to generally converge to the optimum for both noisy experiments (Vazquez et al. 2008; Letham et al. 2019) and noiseless function evaluation (Bull 2011). Additionally, the EI-based approach has been well developed to tackle various practical problems (Jordan et al. 2011; Shahriari et al. 2016).
Tanaka and Iwata (2018) considered the following GBLUP model:
where \(\varvec{y}\) denotes the vector of phenotypic values, \(\mu \) the general mean, \(\varvec{1}_n\) the unit vector of length n, \(\varvec{g}\) the vector of genotypic values, and \(\varvec{e}\) the vector of random errors. It is assumed that \(\varvec{g}\) follows a multivariate normal distribution, denoted by \(\varvec{g}\sim MVN(\varvec{0},\sigma _g^2\varvec{K})\), where \(\varvec{0}\) is the zero vector, \(\sigma _g^2\) is the additive genetic variance, and \(\varvec{K}\) is a known genomic relationship matrix. Also, the vector of random errors \(\varvec{e}\sim MVN(\varvec{0},\sigma _e^2\varvec{I}_n)\), where \(\sigma _e^2\) is the random error variance and \(\varvec{I}_n\) is the identity matrix of order n. Moreover, \(\varvec{g}\) and \(\varvec{e}\) are assumed to be mutually independent. The evaluation function here is the unknown genotypic value for an individual, which is evaluated through its DNA marker scores (genotype). Interestingly, the evaluation function of genotypic value is stochastic (random effect) and the feasible space is a finite candidate population set in the GBLUP model, which are different from those in the classic Bayesian optimization problem. Tanaka and Iwata (2018) proposed an EI criterion and showed their Bayesian optimization approach to have advantage compared with the selection method based on posterior means and with a random sampling method. The aim of our study is to investigate potential EI-based Bayesian optimization approaches for identifying the best genotype from a large candidate population.
The rest of this article is organized as follows. In Sect. 2, we derive some modified versions of the EI criterion proposed by Tanaka and Iwata (2018). In Sect. 3, we present the search process of the Bayesian optimization approach and a stopping rule for real data analysis. We illustrate the proposed approach through analysis of two rice (Oryza sativa L.) and one wheat (Triticum aestivum L.) genome datasets in Sect. 4. We further conduct a detailed simulation study to compare the performance of the EI criteria in Sect. 5. Some concluding remarks are provided in Sect. 6.
2 The Expected Improvement Criteria
It is known that the genomic relationship matrix \(\varvec{K}\) plays an important role in building the GBLUP model (1). We considered the case where \(\varvec{K}=\varvec{M}\varvec{M}^\top /p\), where \(\varvec{M}\) is the standardized marker score matrix and p is the number of DNA markers. Let \(m_{ij}\) denote the (ij)th element of \(\varvec{M}\); then \(m_{ij}= (a_{ij}-\bar{a}_j)/s_j\), where \(a_{ij}\) is the (ij)th element of the original marker score matrix, and \(\bar{a}_j\) and \(s_j\) are, respectively, the sample mean and the sample standard deviation of the elements in column j. We first present the distributions of predicted phenotypic values (PPVs) and predicted genotypic values (PGVs), and then derive several EI criteria.
2.1 Distributions of Predicted Phenotypic and Genotypic Values
Let \(\varvec{y}_1\) and \(\varvec{g}_1\), respectively, denote the vectors of phenotypic and genotypic values for the training set, both of which are of size \(n_1\). Likewise, let \(\varvec{y}_2\) and \(\varvec{g}_2\) denote the corresponding vectors for the remaining \(n_2\) individuals not chosen in the training set (the non-phenotyped set), where \(n_1+ n_2=n\). Therefore, the GBLUP model (1) can be partitioned as
where
We use the R package BGLR (Perez and de los Campos, G. 2014) to perform the Bayesian reproducing kernel Hilbert space (RKHS) method for estimating \(\mu \), \(\sigma _g^2\) and \(\sigma _e^2\), and the BLUP of \(\varvec{g}_1\) using the training set. More precisely, the estimation is based only on the model
where
The resulting estimated values are denoted by \(\hat{\mu }\), \(\hat{\sigma }_g^2\), \(\hat{\sigma }_e^2\), and \(\hat{\varvec{g}}_1\). We refer to Xavier et al. (2016) for more details regarding the Bayesian RKHS method.
Under the condition that \(\hat{\mu }\), \(\hat{\sigma }_g^2\), \(\hat{\sigma }_e^2\), and \(\hat{\varvec{g}}_1\) are all assumed to be fixed and known values, the distribution of PPVs for the non-phenotyped set, i.e., the distribution of \({\varvec{y}}_2\) conditioned by \({\varvec{y}}_1=\hat{\varvec{y}}_1\), is given by
where \(\hat{\varvec{\mu }}_p=\hat{\mu }\varvec{1}_{n_2}+\varvec{K}_{21}\varvec{K}_{11}^{-1}\hat{\varvec{g}}_1\) and \(\hat{\varvec{\Sigma }}_p=\hat{\varvec{\Sigma }}_{22}-\hat{\varvec{\Sigma }}_{21}\hat{\varvec{\Sigma }}_{11}^{-1}\hat{\varvec{\Sigma }}_{12}\). Here, \(\hat{\varvec{\Sigma }}_{22}=\hat{\sigma }_g^2\varvec{K}_{22}+\hat{\sigma }_e^2\varvec{I}_{n_2}\), \(\hat{\varvec{\Sigma }}_{21}=\hat{\sigma }_g^2\varvec{K}_{21}\), \(\hat{\varvec{\Sigma }}_{11}=\hat{\sigma }_g^2\varvec{K}_{11}+\hat{\sigma }_e^2\varvec{I}_{n_1}\), and \(\hat{\varvec{\Sigma }}_{12}=\hat{\sigma }_g^2\varvec{K}_{12}\). Likewise, the distribution of PGVs for the non-phenotyped set, i.e., the distribution of \({\varvec{g}}_2\) conditioned by \({\varvec{g}}_1=\hat{\varvec{g}}_1\), is given by
where \(\hat{\varvec{\mu }}_g=\varvec{K}_{21}\varvec{K}_{11}^{-1}\hat{\varvec{g}}_1\) and \(\hat{\varvec{\Sigma }}_g=\hat{\sigma }_g^2(\varvec{K}_{22}-\varvec{K}_{21}\varvec{K}_{11}^{-1}\varvec{K}_{12})\). Note that \(\tilde{\varvec{g}}_2=\varvec{K}_{21}\varvec{K}_{11}^{-1}\tilde{\varvec{g}}_1\), where \(\tilde{\varvec{g}}_1\) is the BLUP of \(\varvec{g}_1\). From Henderson (1977), it follows that \(\tilde{\varvec{g}}_2\) is the BLUP of \(\varvec{g}_2\). Obviously, \(\tilde{\varvec{g}}_2\) differs from \(\tilde{\varvec{y}}_2\) in distribution by the term \(\hat{\mu }\) in the mean vector and the term \(\hat{\sigma }_e^2\) in the covariance matrix. That is, \(\hat{\varvec{\mu }}_p=\hat{\varvec{\mu }}_g\) and \(\hat{\varvec{\Sigma }}_p=\hat{\varvec{\Sigma }}_g\) if \(\hat{\mu }=\hat{\sigma }_e^2=0\). In other words, the PPVs are affected by the random error, but the PGVs are free of this variation. The best genotype for a target trait is defined as the individual with the maximal or minimal genotypic value among the candidate population. Intuitively, an EI criterion derived from the distribution of PGVs rather than PPVs should be more efficient in identifying the desired best genotype. A construction method for the statistical tolerance interval for a distribution free of measurement errors like that of PGVs can be found in Lin et al. (2008).
2.2 Plug-In Expected Improvement Criteria
Tanaka and Iwata (2018) considered the EI criterion based on the distribution of PPVs in (2). Let \(\hat{\varvec{y}}_1=\hat{\mu }\varvec{1}_{n_1}+\hat{\varvec{g}}_1\), representing the vector of estimated phenotyped values for the training set, and \(\hat{f}_{Mp}\) be the maximal value among \(\hat{\varvec{y}}_1\). Also, let the marginal distribution for each individual in (2) be denoted by
The improvement function for \(\tilde{y}_{2i}\) is defined as
Here, \(Im (\tilde{y}_{2i})\) is a random variable associated with the distribution of \(\tilde{y}_{2i}\). From Jones et al. (1998), the expected improvement function has an elegant closed form given by
where \(Z_{pi}=(\hat{\mu }_{pi}-\hat{f}_{Mp})/\hat{\sigma }_{pi}\), \(\Phi (\cdot )\) is the cumulative density function of the standard normal distribution, and \(\phi (\cdot )\) is the probability density function of the standard normal distribution. The EI in (5) was referred as a “plug-in” strategy by Picheny et al. (2013) because \(\hat{f}_{MP}\) is used for a noisy experiment. We simply call it EI-PPV. Alternatively, we consider the plug-in EI criterion based on the distribution of PGVs in (3). Likewise, we have the EI-PGV given by
where \(\tilde{g}_{2i}\sim N(\hat{\mu }_{gi},\hat{\sigma }_{gi}^2)\) (the corresponding marginal distribution in (3)) and \(\hat{f}_{Mg}\) is the maximal value among \(\hat{\varvec{g}}_1\) and \(Z_{gi}=(\hat{\mu }_{gi}-\hat{f}_{Mg})/\hat{\sigma }_{gi}\).
2.3 Forward Expected Improvement Criteria
Clearly, both of the above EI-PPV and EI-PGV ignore the genomic correlation among the candidate individuals. Thus, we take into account in the criterion the property that selection of future individuals could be associated with the current selected individual. We rewrite the distribution of PPVs as
where \(\tilde{y}_*\) represents the PPV of the individual with the largest \(EI(\tilde{y}_{2i})\) from (5), i.e., the individual with \(\tilde{y}_*\) is the first chosen from the non-phenotyped set. Subsequently, we search for the next individual with the largest \(EI(\tilde{y}_{2i})\) among the remaining candidate individuals whose PPVs follow the conditional distribution
where \(\hat{\varvec{\mu }}_{cp}=\hat{\varvec{\mu }}_{*c}\) and \(\hat{\varvec{\Sigma }}_{cp}=\hat{\varvec{\Sigma }}_{p22}-\hat{\varvec{\Sigma }}_{p21}(\hat{\sigma }_*^2)^{-1}\hat{\varvec{\Sigma }}_{p12}\). We here adjust the distribution of \(\tilde{\varvec{y}}_c\) for the mean value of \(\tilde{y}_*\). Let
representing a marginal distribution for individual i in (7); then, the corresponding EI can be derived as
where \(Z_{cpi}=(\hat{\mu }_{cpi}-\hat{f}_{Mp})/\hat{\sigma }_{cpi}\). Similarly, we have the corresponding EI for the distribution of the PGVs as
where \(Z_{cgi}=(\hat{\mu }_{cgi}-\hat{f}_{Mg})/\hat{\sigma }_{cgi}\). We abbreviate the EI in (8) as EI-PPV-fwd, and the EI in (9) as EI-PGV-fwd. Note that “fwd” here means forward selection. For a specific candidate individual i, \(\hat{\mu }_{cpi}=\hat{\mu }_{pi}\) and \(\hat{\sigma }_{cpi}\le \hat{\sigma }_{pi}\). That is to say, EI-PPV-fwd has a smaller prediction standard error than EI-PPV for a candidate individual. Similarly, the prediction standard error in EI-PGV-fwd is smaller than that in EI-PGV.
2.4 Augmented Expected Improvement Criteria
Huang et al. (2006) proposed an augmented EI criterion to handle noisy experiments. We adopt such an augmented version for the above EI-PGV and EI-PGV-fwd. The augmented EI-PGV is modified from (6) and is given by
and the augmented EI-PGV-fwd is modified from (9) and is given by
where \(\hat{f}_{Mg}^*\) denotes the current “effective best solution,” \(Z_{gi}^*=(\hat{\mu }_{gi}-\hat{f}_{Mg}^*)/\hat{\sigma }_{gi}\), and \(Z_{cgi}^*=(\hat{\mu }_{cgi}-\hat{f}_{Mg}^*)/\hat{\sigma }_{cgi}\). The current effective best solution \(\hat{f}_{Mg}^*\) is obtained from \(\hat{\varvec{g}}_1\) and maximizes the utility function
where \(\gamma \) is a constant that reflects the degree of risk aversion. That is,
We specify \(\gamma =1\), which implies that 1 unit of \(\hat{g}_{1i}\) is traded for 1 unit of \(\hat{\sigma }_{1gi}\). Following Huang et al. (2006), \(\hat{f}_{Mg}^*\) is used to reduce prediction uncertainty, and the heuristic multiplier of the EI accounts for the diminishing return of additional replicates as the prediction becomes more accurate. The EIs in (10) and (11) are abbreviated as aug-EI-PGV and aug-EI-PGV-fwd, respectively.
The above EI criteria are all developed for the situation in which the best genotype is assumed to give the maximal genotypic value for some target trait such as grain yield of a crop. The corresponding EI criteria for the situation in which the best genotype is desired to give the minimal genotypic value, such as plant height or flowering time of a crop, are provided in the Appendix.
3 Search Process
For a given candidate population in which all the individuals have been genotyped but not phenotyped, the iterative search process to identify the best genotype from the candidate population can be described as follows.
- Step 0::
-
Randomly choose \(n_0\) individuals from the candidate population to be phenotyped for a target trait. The chosen individuals serve as the starting training set, denoted by \(\varvec{S}_0\). Initialize \(n_{tr} \leftarrow n_0\) and \(\varvec{S}_{tr} \leftarrow \varvec{S}_0\), where \(\varvec{S}_{tr}\) denotes the current training set and \(n_{tr}\) is its sample size.
- Step 1::
-
Perform the Bayesian RKHS method using the phenotype and genotype data of \(\varvec{S}_{tr}\) to construct the GBLUP model, and obtain the estimated values of \(\hat{\mu }\), \(\hat{\sigma }_g^2\), \(\hat{\sigma }_e^2\), and \(\hat{\varvec{g}}_1\).
- Step 2::
-
Select \(n_{sel}\) individuals from the non-phenotyped set using each of the above EI criteria.
- Step 3::
-
If the average of the realized EI values among the \(n_{sel}\) selected individuals is less than a stopping tolerance \(\delta \), then STOP the search and GO TO Step 4. Otherwise, phenotype the selected \(n_{sel}\) individuals, denoted by \(\varvec{S}_{sel}\), and add them to the current training set. That is, the union of \(\varvec{S}_{tr}\) and \(\varvec{S}_{sel}\) forms the new training set, expressed as \(\varvec{S}_{tr} \leftarrow \varvec{S}_{tr}\cup \varvec{S}_{sel}\). Similarly, \(n_{tr} \leftarrow n_{tr}+n_{sel}\). GO TO Step 1.
- Step 4::
-
Identify the individual with the maximal (or minimal) estimated genotypic value among the current selected individuals as the best genotype.
Since the candidate population is a finite set, it is easy to argue that all the EI criteria converge to 0. For example, from the definition of the improvement function in (4), the EI value becomes equal to 0 if all \(\tilde{y}_{2i}<\hat{f}_{Mp}\), and this surely happens for a finite candidate set. In other words, \(\hat{f}_{Mp}\) definitely can be found after all candidate individuals have been evaluated.
4 Illustrative Examples
We use the following three genome datasets to illustrate the search process.
4.1 Example 1: 44k Rice Genome Dataset
This dataset was presented in Zhao et al. (2011) and contains 413 accessions and 44,100 single nucleotide polymorphism (SNP) markers. This dataset consists of five subpopulations and one admixed group. SNPs with missing rate \(> 0.05\) and minor allele frequency \(< 0.05\) were removed from the dataset, with the result that 36,901 out of the 44,100 SNPs were retained. The SNP genotype at each locus was coded as \(-1\), 0, or 1, where 1 indicates the homogeneous genotype of the major allele, \(-1\) the homogeneous genotype of the minor allele, and 0 the heterogeneous genotype. After SNP coding, any missing locus was taken as 1. Only 300 out of the 413 accessions with no missing phenotypic values on the flowering time in all three locations—Arkansas (FT-Ark), Faridpur (FT-Far), and Aberdeen (FT-Abe)—were used for the demonstration. We fixed the stopping tolerance as \(\delta =0.1\) for the above three target traits.
4.1.1 Example 2: Tropical Rice Genome Dataset
This dataset was presented in Spindle et al. (2015) and contains 73,147 SNP markers and 363 elite breeding lines belonging to indica or indica-admixed group. Phenotypic observations included 4 years (2009–2012, with two seasons (dry and wet) per year) of data on grain yield (GY), flowering time (FT), and plant height (PH), although PH data were not available for the wet season of 2009. Phenotypic values of 35 out of the 363 individuals were missing; therefore, adjusted means of only 328 individuals were used for this example. Additionally, only 10,772 out of the 73,147 SNPs were used in this study. One SNP marker was selected per 0.1 cM interval over each chromosome, because the chosen subset of the full marker set has been shown to be sufficient for genomic selection in this collection of rice germplasm (Spindle et al. 2015). The SNP coding was the same as in Example 1. We set the stopping tolerance as \(\delta = 0.01\) for this dataset.
4.1.2 Example 3: Wheat Genome Dataset
This dataset was used in Crossa et al. (2010) and contains 599 historical wheat lines from the International Maize and Wheat Improvement Center (CIMMYT ) and 1279 diversity arrays technology (DArT) markers. The wheat dataset is from CIMMYT’s Global Wheat Program, which conducted numerous international trials across a variety of producing environments. These environments were grouped into four mega-environments (E1 to E4) comprising four main agro-climatic regions. The phenotypic value was the average grain yield of the 599 wheat lines evaluated in each of these four mega-environments. The marker score for each DArT marker was coded as 1 for presence and 0 for absence. We set the stopping tolerance as \(\delta =0.001\) for all the target traits E1 to E4.
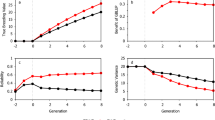
We set \(n_0=10\) and \(n_{sel}=20\) in the search process for all three of the above datasets. The sample size of the training set when the search was stopped and the identified best genotype according to each of the EI criteria are displayed in Table 1, from which it can be seen that most of the EI criteria found a common best genotype for a specific target trait. However, the final sample sizes of the training set may differ from different EI criteria, and even from different traits within the same dataset. Certainly, the specification of the stopping tolerance and the convergence rate of the EI criteria can have a significant impact on the final sample size. The convergence process of the EI criteria for the three datasets is displayed in Figures S1 to S3 of the Supplementary Materials. The figures reveal that those EI criteria considering the random error variance, EI-PPV, EI-PPV-fwd, aug-EI-PGV, and aug-EI-PGV-fwd, have relatively quick convergence rates and smooth curves compared with EI-PGV, and EI-PGV-fwd, both of which don’t take \(\hat{\sigma }_e^2\) into account. In particular, the convergence process of aug-EI-PGV appears to be the fastest and stablest for most of the cases. An R code for the analysis of the tropical rice dataset by the aug-EI-PGV criterion is provided in the Supplementary Materials.
5 Simulation Study
We further conducted a simulation study to compare the EI criteria using the above three datasets as templates. The genomic relationship matrix \(\varvec{K}\) was first calculated from all available genotype data for each dataset, and then the GBLUP model (1) was employed to generate the simulated datasets. Genotype data of 373, 328, and 599 individuals for the 44k rice, tropical rice, and wheat datasets, respectively, were used in this simulation study. The procedure of the simulation study can be described as follows.
- Step 1::
-
Generate the vector of genotypic values \(\varvec{g}\) from \(MVN(\varvec{0},\sigma _g^2\varvec{K})\), where \(\sigma _g^2\) is set to be 25. The simulated \(\varvec{g}\) is treated as the vector of true genotypic values, and the individual with the largest genotypic value is assumed to be the best genotype.
- Step 2::
-
Generate the vector of random errors \(\varvec{e}\) from \(MVN(\varvec{0},\sigma _e^2\varvec{I}_n)\), where \(\sigma _e^2=\sigma _g^2(1-h^2)/h^2\). Here, \(h^2\) represents the trait heritability fixed at 0.2, 0.5, and 0.8 for low, intermediate, and high heritability, respectively.
- Step 3::
-
Repeat the above two steps 5,000 times to generate 5,000 simulated datasets of phenotypic values for each of the fixed values of \(h^2\). That is, \(\varvec{y}=\mu \varvec{1}+\varvec{g}+\varvec{e}\), where \(\mu \) is set to 100.
- Step 4::
-
Conduct steps 1 to 3 of the search process presented in Sect. 3 for each simulated dataset. Instead of the rule for stopping tolerance that was used in the real data analysis, we stop the search once the best genotype has been identified from a simulated dataset, and then record the number of the individuals that have been selected at this point. We set \(n_0=10\) and \(n_{sel}=10,20,40\) for this simulation study.
The sample size of the training set required to discover the best genotype for each EI criterion obtained from Step 4 was used as a measure of the performance for the EI criterion. The mean and the standard deviation over the training set sizes from the 5,000 simulated datasets are reported in Table 2. As expected, the performance generally improves with increasing heritability \(h^2\) and with decreasing size \(n_{sel}\) of the training set selected at each batch. Also, the EIs based on PGVs slightly outperform their counterparts based on PPVs. Interestingly, the forward EIs of EI-PPV-fwd, EI-PGV-fwd, and aug-EI-PGV-fwd have relatively unsatisfactory performance, which could be because their smaller prediction variances reduce the exploration capacity and hence increase the size of the required training set. The aug-EI-PGV performs quite similarly to EI-PGV, probably because the prediction standard errors are almost the same for all candidate individuals (the diagonal elements of the genomic relationship matrix are all equal to or close to 1). Thus, the current effective best solution \(\hat{f}_{Mg}^*\) and the heuristic multiplier in the aug-EI-PGV have little impact on the identification of the best genotype. We thus made a more detailed comparison between EI-PPV, EI-PGV, and aug-EI-PGV. Let n(PPV), n(PGV), and n(aPGV) denote the numbers of individuals required to identify the true best genotype from one repetition of simulation using EI-PPV, EI-PGV, and aug-EI-PGV, respectively. Then, the proportions over the 5,000 repetitions that \(n(PPV) < n(PGV)\), \(n(PPV) < n(aPGV)\), and \(n(PGV) < n(aPGV)\) under the setting of \(n_0=10\) and \(n_{sel}=10\) are displayed in Table 3. From the table, both EI-PGV and aug-EI-PGV have better performance than EI-PPV because they often require less individuals to identify the true best genotype. Moreover, aug-EI-PGV slightly outperforms EI-PGV based on both the 44k rice and wheat templates, but shows a significant advantage over EI-PGV based on the tropical rice template. The corresponding scattered plots, displayed in Figures S4 to S6 of the Supplementary Materials, reflect the consistent results. In consequence, aug-EI-PGV can be recommended for practical use.
In real data analysis, we propose a feasible stopping rule for the search process according to the fact that the empirical EI values are expected to converge to 0 as the number of selected individuals increases. However, the specification of stopping tolerance \(\delta \) can significantly affect the final sample size of the required training set. It is therefore desirable to provide some tool to determine the stopping tolerance before performing the real data analysis. For a dataset with known genotype data, following the above simulation steps, we could similarly obtain the smallest sample size of the training set required to meet the standard set by a fixed \(\delta \). That is, the empirical EI value with the training set size is less than the fixed \(\delta \). Let \(\bar{n}_{tr}(\delta )\) and \(\bar{n}_{tr}\) denote the sample size averages over the 5,000 simulation repetitions of meeting the standard set by the fixed \(\delta \) and of identifying the true best genotype, respectively. Then, the largest stopping tolerance \(\delta ^*\) among those under consideration such that \(\bar{n}_{tr}(\delta ^*) \ge \bar{n}_{tr}\) is recommended for the dataset. Let \(\delta \) be evaluated over \(10^{-k}\), for \(k=1,2, \ldots , 6\), and the trait heritability \(h^2\) be set from 0.1 to 0.9 by increment of 0.1, then the resulting \(\delta ^*\) based on the three templates using aug-EI-PGV under the setting of \(n_0=10\) and \(n_{sel}=10\) is displayed in Table 4. From Table 4, the stopping tolerance \(\delta \) can be pre-specified as \(10^{-3}\), \(10^{-2}\), and \(10^{-3}\) for the datasets of 44k rice, tropical rice, and wheat, respectively. An R code is provided in the Supplementary Materials for users to determine the stopping tolerance for their own datasets.
As mentioned earlier, the genomic relationship matrix \(\varvec{K}\) plays an important role in construction of the GBLUP model, and might also affect the result of searching the best candidate in the current study. Again, following the above simulation steps, we could compare different relationship matrices in identifying the true best genotype for each simulated dataset generated according to the original relationship matrix described in Sect. 2. For the sake of clarity, let this original relationship matrix be denoted as \(\varvec{K}_0\). Another three possible formulas for calculating the genomic correlation discussed in Rincent et al. (2012) are summarized in Eq. (A1)–(A3) of the Appendix. Let the relationship matrices corresponding to Eq. (A1)–(A3) be denoted by \(\varvec{K}_1\), \(\varvec{K}_2\), and \(\varvec{K}_3\), respectively. The mean of the training set sizes required to discover the true best genotype from the 5,000 simulated datasets using the different relationship matrices by aug-EI-PGV under the setting of \(n_0=10\) and \(n_{sel}=10\) is displayed in Table 5. From the table, it is interesting to find that \(\varvec{K}_1\), \(\varvec{K}_2\), and \(\varvec{K}_3\) perform almost equally well in identifying the true best genotype. Notice that this study somewhat favors the original relationship matrix \(\varvec{K}_0\) because the datasets were simulated based on it.
6 Concluding Remarks
The EI-based Bayesian optimization approach allows a trade-off between exploration and exploitation. From the definition of the improvement function, when there are genotypes better than the best identified at the current stage, the EI-based approach tends to select the genotypes with the large predicted means (exploitation). On the other hand, when there is no genotype better than the best, it tends to select the genotypes with large estimated standard deviations (exploration). The exploration here can enhance the genomic diversity of the training set and thereby increase the chance of discovering the true best genotype in the next search. The problem of interest in this study is actually a batch Bayesian optimization problem, i.e., one in which multiple query points are to be selected simultaneously. Some advanced approaches to such batch Bayesian optimization discussed in Gong et al. (2019) may be adapted for our current study.
To the best of our knowledge, there are no reports in the literature of sequential phenotyping experiments for GP studies in plant breeding. This could be due to the practical constraint that plant breeders are usually able to collect phenotypic data such as grain yield only after a whole growing season of a crop. This can make sequential phenotyping experiment unfeasible if the growing season is too long. From the simulation results, even though a scenario with a small batch size (\(n_{sel}=10\)) at a specific heritability can be more efficient than a large one (\(n_{sel}=40\)), the difference is not remarkable in practice. This indicates that breeders can trade a little efficiency to phenotype a large batch of individuals in one stage instead of small batches in several stages. It is known that phenotype is affected by the genotype (G), environment (E), and G\(\times \)E interaction. In reality, environment can have a significant impact on the performance of individuals selected at each batch. Thus, the proposed multi-stage strategy may need more extensive evaluation to make sure the method works.
Clearly, the application of Bayesian optimization in this study is not to build training and testing sets for GP, but only to identify the best candidate that may be used to serve as a commercial cultivar or to form the parents for the next breeding generation. The EI-based approach could be particularly useful when the phenotyping process is expensive or not very time-consuming, and in addition, the experimenter is able to phenotype individuals sequentially in the order that our proposed approach suggests. For example, for a stress-resistance breeding program, plant breeders need to identify superior accessions with the relevant resistance genes from a germplasm collection. They can then transfer the genes of the identified accessions to a cultivar, thereby enabling it to perform to a desired degree in spite of being subjected to stress (Acquaah 2007). Suppose that the phenotyping cost for a large number of candidate accessions is not affordable to breeders. In such a situation, our proposed sequential phenotyping strategy is a potentially useful choice. Recently, Wu et al. (2019) proposed a GP procedure to assess pumpkin hybrid performance and then identified potential \(F_1\) hybrids with large fruit weight. There were 10,011 possible \(F_1\) hybrids that needed to be evaluated. In this case, our proposed sequential phenotyping strategy might be recommended to the breeder if they are unable to phenotype a large number of the \(F_1\) hybrids during a growing season.
Even though the vector of phenotypic values is assumed to follow a multivariate normal distribution in the GBLUP model (1), it seems not possible to assess the multivariate normality assumption based on such data with no replication on the phenotypic value vector. The multivariate normality assumption is actually imposed on the GBLUP model due to the genomic correlation between individuals, i.e., it is assumed that \(\varvec{g} \sim MVN(\varvec{0}, \sigma ^2_g \varvec{K})\). This single-trait GBLUP model is in essence a derivation from the univariate Gaussian regression model (Tempelman 2015), so it might be sufficient to make a univariate normality assessment using the phenotypic values of a single trait. Several graphic approaches or statistical tests can be applied for such normality assumption assessment (Garson 2012). For the proposed Bayesian optimization approach, the search process begins with a training set randomly selected from the candidate population. It would be beneficial to construct a better starting training set from some optimization algorithms (Ou and Liao 2019; Heslot and Feoktistov 2020). Also, from the results of the simulation study, it is reasonable to conclude that the selection efficiency generally improves with increasing trait heritability. The impact of other factors, such as trait genetic architecture, marker density, and linkage disequilibrium, needs further study. Another interesting issue for future study is the investigation of Bayesian optimization approaches with the aim of discovering a set of superior genotypes.
References
Acquaah G (2007) Principles of plant genetics and breeding. Blackwell Publishing, Malden
Akdemir D, Sanchez JI (2019) Design of training population for selective phenotyping in genomic prediction. Sci Rep 9:1446
Bull AD (2011) Convergence rates of efficient global optimization algorithms. J Mach Learn Res 12:2879–2904
Crossa J, Campos G, de los Pérez P (2010) Prediction of genetic values of quantitative traits in plant breeding using pedigree and molecular markers. Genetics 186:713–724
Garson GD (2012) Testing statistical assumptions. Statistical Publishing Associate, Asheboro
Gong C, Peng J, Liu Q (2019) Quantile Stein variational gradient descent for batch Bayesian optimization. In: Proceedings of the 36th international conference on machine learning, PMLR 97, pp 2347–2356
Habier D, Fernando RL, Dekkers JCM (2007) The impact of genetic relationship information on genomic-assisted breeding values. Genetics 177:2389–2397
Henderson CR (1977) Best linear unbiased prediction of breeding values not in the model for records. J Dairy Sci 60:783–787
Heslot N, Feoktistov V (2020) Optimization of selective phenotyping and population design for genomic prediction. J Agric Biol Environ Stat 25:601–616
Huang D, Allen TT, Notz WI, Zeng N (2006) Global optimization of stochastic black-box systems via sequential kriging meta-models. J Global Optim 34:441–466
Jones DR, Schonlau M, Welch WJ (1998) Efficient global optimization of expensive black-box functions. J Global Optim 13:455–492
Jordan DR, Mace ES, Cruickshank AW, Hunt CH, Henzell RG (2011) Exploring and exploiting genetic variation from unadapted sorghum germplasm in a breeding program. Crop Sci 51:1444–1457
Letham B, Karrer B, Ottoni G, Bakshy E (2019) Constrained Bayesian optimization with noisy experiments. Bayesian Anal 14:495–519
Lin TY, Liao CT, Iyer HK (2008) Tolerance intervals for unbalanced one-way random effects models with covariates and heterogeneous variances. J Agric Biol Environ Stat 13:221–241
McCouch S, Baute GJ, Bradeen J, Bramel P, Bretting PK, Buckler E et al (2013) Agriculture: feeding the future. Nature 499:23–24
Meuwissen THE, Hayes BJ, Goddard ME (2001) Prediction of total genetic value using genome-wide dense marker maps. Genetics 157:1819–1829
Ou JH, Liao CT (2019) Training set determination for genomic selection. Theor Appl Genet 132:2781–2792
Perez P, de Los Campos G (2014) Genome-wide regression and prediction with the BGLR statistical package. Genetics 198:483–495
Picheny V, Ginsbourger D, Richet Y, Caplin G (2013) Quantile-based optimization of noise computer experiments with tunable precision. Technometrics 55:2–13
Reif JC, Zhang P, Dreisigacker S, Warburton ML, van Ginkel M et al (2005) Wheat genetic diversity trends during domestication and breeding. Theor Appl Genet 110:859–864
Rincent R, Laloe D, Nicolas S, Altmann T, Brunel D et al (2012) Maximizing the reliability of genomic selection by optimizing the calibration set of reference individuals: comparison of methods in two diverse groups of maize inbreds (Zea mays L.). Genetics 192:715–728
Shahriari B, Swersky K, Wang Z, Adams RP, De Freitas N (2016) Taking the human out of the loop: a review of Bayesian optimization. Proc IEEE 104:148–175
Spindel J, Begum H, Akdemir D, Virk P, Collard B et al (2015) Genomic selection and association mapping in rice (Oryza sativa): Effect of trait genetic architecture, training population composition, marker number and statistical model on accuracy of rice genomic selection in elite, tropical rice breeding lines. PLOS Genet 11:e1005350
Tanaka R, Iwata H (2018) Bayesian optimization for genomic selection: a method for discovering the best genotype among a large number of candidates. Theor Appl Genet 131:93–105
Tempelman RJ (2015) Statistical and computational challenges in whole genome prediction and genome-wide association analyses for plant and animal breeding. J Agric Biol Environ Stat 20:442–466
Tester M, Langridge P (2010) Breeding technologies to increase crop production in a changing world. Science 327:818–822
Tibshirani R (1996) Regression shrinkage and selection via the LASSO. J Royal Stat Soc Ser B Stati Methodol 58:267–288
Vazquez E, Villemonteix J, Sidorkiewicz M, Walter E (2008) Global optimization based on noisy evaluations: an empirical study of two statistical approaches. J Global Optim 43:373–389
Wu PY, Tung CW, Lee CY, Liao CT (2019) Genomic prediction of pumpkin hybrid performance. Plant Genome 12:180082
Xavier A, Muir WM, Craig B, Rainey KM (2016) Walking through the statistical black boxes of plant breeding. Theor Appl Genet 129:1933–1949
Zhao K, Tung CW, Eizenga GC, Wright MH, Ali ML et al (2011) Genome-wide association mapping reveals a rich genetic architecture of complex traits in Oryza sativa. Nat Commun 2:467
Acknowledgements
The authors thank the editor, an associate editor, and three reviewers for providing constructive comments, which helped us to improve both the content and presentation of the manuscript. This research was supported by the Ministry of Science and Technology, Taiwan (Grant No. MOST 108-2118-M-002-002-MY2).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Appendix
Appendix
1.1 The EI Criteria with the Minimal Genotypic Value
The EI criteria for the situation in which the best genotype is assumed to give the minimal genotypic value are summarized below.
1.1.1 Plug-in Expected Improvement Criteria
where \(Z_{pi}=(\hat{f}_{mp}-\hat{\mu }_{pi})/\hat{\sigma }_{pi}\) and \(\hat{f}_{mp}\) is the minimal value of \(\hat{\varvec{y}}_1\);
where \(Z_{gi}=(\hat{f}_{mg}-\hat{\mu }_{gi})/\hat{\sigma }_{gi}\) and \(\hat{f}_{mg}\) is the minimal value of \(\hat{\varvec{g}}_1\).
1.1.2 Forward Expected Improvement Criteria
where \(Z_{cpi}=(\hat{f}_{mp}-\hat{\mu }_{cpi})/\hat{\sigma }_{cpi}\);
where \(Z_{gi}=(\hat{f}_{mg}-\hat{\mu }_{cgi})/\hat{\sigma }_{cgi}\).
1.1.3 Augmented Expected Improvement Criteria
where \(Z_{gi}^*=(\hat{f}_{mg}^*-\hat{\mu }_{gi})/\hat{\sigma }_{gi}\), \(Z_{cgi}^*=(\hat{f}_{mg}^*-\hat{\mu }_{cgi})/\hat{\sigma }_{cgi}\), and \(\hat{f}_{mg}^*\) is obtained from \(\hat{\varvec{g}}_1\) and minimizes the utility function, i.e.,
where \(\gamma =1\).
1.2 The Formulas for Calculating the Genomic Correlation
Let \((a_1, a_2, \cdots , a_p)\) and \((b_1, b_2, \cdots , b_p)\) be the marker score vectors for individuals A and B, respectively. Then, the genomic correlation between these two individuals can be calculated by the following three formulas.
where the marker score is coded as 0, 0.5, or 1 for the genotype of \(M_1M_1\), \(M_1M_2\), or \(M_2M_2\). Here, \(M_1M_1\) and \(M_2M_2\) denote the two homogenous genotypes, and \(M_1M_2\) denotes the heterogenous genotype.
where the marker score is coded as 0, 1, or 2 for the genotype of \(M_1M_1\), \(M_1M_2\), or \(M_2M_2\).
where the marker score is coded as 0, 0.5, or 1 for the genotype of \(M_1M_1\), \(M_1M_2\), or \(M_2M_2\), and \(q_i\) is the proportion among the n individuals that the marker score is coded as 1 at marker i, for \(i=1, 2, \ldots , p\).
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tsai, SF., Shen, CC. & Liao, CT. Bayesian Optimization Approaches for Identifying the Best Genotype from a Candidate Population. JABES 26, 519–537 (2021). https://doi.org/10.1007/s13253-021-00454-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13253-021-00454-2