Abstract
Increased grain yield (GY) is the primary breeding target of wheat breeders. We performed the genome-wide association study (GWAS) on 168 elite winter wheat lines from an ongoing breeding program to identify the main determinants of grain yield. Sequencing of Diversity Array Technology fragments (DArTseq) resulted in 19,350 single-nucleotide polymorphism (SNP) and presence-absence variation (PAV) markers. We identified 15 main genomic regions located in ten wheat chromosomes (1B, 2B, 2D, 3A, 3D, 5A, 5B, 6A, 6B, and 7B) that explained from 7.9 to 20.3% of the variation in grain yield and 13.3% of the yield stability. Loci identified in the reduced genepool are important for wheat improvement using marker-assisted selection. We found marker-trait associations between three genes involved in starch biosynthesis and grain yield. Two starch synthase genes (TraesCS2B03G1238800 and TraesCS2D03G1048800) and a sucrose synthase gene (TraesCS3D03G0024300) were found in regions of QGy.rut-2B.2, QGy.rut-2D.1, and QGy.rut-3D, respectively. These loci and other significantly associated SNP markers found in this study can be used for pyramiding favorable alleles in high-yielding varieties or to improve the accuracy of prediction in genomic selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cultivating common wheat (Triticum aestivum L.) provides about 20% of the total calories used by the human population (Rasheed et al. 2018). Worldwide wheat harvest area exceeds 213 million ha, and about 28% of this area is located in Europe, including over 2.3 million ha in Poland (FAOSTAT 2022). However, there are limitations to the territorial expansion of wheat cultivation, and to meet the challenge of doubling the wheat yield by 2050 (Rasheed et al. 2018), significant yield increase per unit of area is required. To meet this challenge, increased genetic diversity deposited in landraces (Vikram et al. 2016), synthetic wheat varieties (Li et al. 2018), and wild relatives (Rasheed et al. 2018) needs to be identified and exploited in modern wheat cultivars, besides agronomical practices for yield improvement. The sequencing of the 17 Gb allohexaploid wheat (AABBDD) genome of Chinese Spring paved the way for genome-wide association studies (GWAS) and genomic selection in common wheat (Lukaszewski et al. 2014; Appels et al. 2018).
The wheat reference sequence provided a physical framework for mapping previously developed genetic markers with known sequences (Alaux et al. 2018), and marker sequences deposited in databases can be used to find regions with target genes (Tyrka et al. 2021b). Hybridization arrays or next-generation sequencing (NGS) are the most common ways to find single-nucleotide polymorphisms (SNPs) and presence–absence variations (PAVs). With the continuous development of new high-throughput NGS methods, the application of genotyping by sequencing (GBS) technologies (e.g., DArTseq) is considered to be the cost-efficient genotyping alternative (Jia et al. 2018) for genomics-based breeding (Poland et al. 2012). GBS gives the genetic information needed to determine economically significant marker-trait associations and develop new wheat cultivars.
Two main approaches to dissecting the genetic basis of complex quantitative traits in crop plants are genome-wide association studies (GWAS) and quantitative trait loci (QTL) mapping. Many QTLs associated with yield-related traits in bread wheat have been identified in biparental populations (Jin et al. 2020; Isham et al. 2021; Kang et al. 2021; Li et al. 2022). At present, GWAS has become more frequently used as it allows for the identification of parts of the complex, essential traits valid in a studied panel of genotypes (Neumann et al. 2011; Sukumaran et al. 2018; Liu et al. 2018; Garcia et al. 2019; Qaseem et al. 2019; Sheoran et al. 2019; Akram et al. 2021). Regions associated with grain yield and its component traits in wheat have been identified in drought and irrigated production conditions (Golabadi et al. 2011; Neumann et al. 2011; Assanga et al. 2017; Bhusal et al. 2017; Li et al. 2019; Khan et al. 2022). Haplotypes found in GWAS (Sehgal et al. 2020) linked to GY are also needed to map candidate genes (Nadolska-Orczyk et al. 2017).
Increased GY is the primary breeding purpose of wheat. The environment strongly influences GY and can be dissected into numerous traits related to phenology and kernel development. Major QTLs significantly associated with yield are currently the target for cloning, and comparative analysis of yield-related traits revealed 145 meta-QTLs and candidate genes (Yang et al. 2021). One of the leading environmental factors influencing yield is nitrogen availability. Nitrogen fertilizer, often used to increase production per unit area, can cause lodging. In wheat, lodging generally occurs after the flowering stage and can affect both the grain yield and the quality of the wheat. Lodging can also be caused by environmental factors, diseases, or pests affecting stems or roots (Keller et al. 1999). In wheat, stem characteristics such as material strength based on lignin concentration and stem thickness play a role (Berry et al. 2007; Berry and Berry 2015; Dreccer et al. 2020, 2022; Piñera-Chavez et al. 2016).
Depending on the population studied, GWAS can identify different genome regions responsible for shaping a trait (Yang et al. 2021). By introducing varieties with very different yield potential into the analyzed population, regions with major effects can be identified. Under long-term selection, polymorphism in these regions may have been lost, and other regions may contribute to yield. GWAS analysis of advanced breeding lines provides an opportunity to identify loci responsible for yield in a narrow gene pool and should indicate the main selection goals that can be achieved using marker-assisted selection. The present study aimed to identify the genomic region(s) associated with grain yield (GY) and component traits, i.e., coefficients of yield stability (STA), days to heading (DTH), plant height (PH), lodging (LDG), and thousand kernel weight (TKW) in a panel of elite wheat genotypes in a range of environments through the GWAS approach.
Material and methods
Phenotypic data collection and analysis
Plant material included 168 breeding lines of common winter wheat and three cultivars evaluated in pre-registration trials in the 2019/2020 season (Table S1). The lines were planted at ten research stations located at Dębina (DED, N54°7′40″, E19°2′7″), Kobierzyce (KBP, N50°58′34″, E16°55′53″), Kończewice (KOH, N53°11′5″, E18°33′15″), Krzemlin (KRZ, N53°4′30″, E14°52′48″), Modzurów (MOB, N50°9′21″, E18°7′38″), Nagradowice (NAD, N52°19′4″, E17°9′1.7″), Polanowice (POB, N50°12′25″, E20°5′5″), Radzików (RAH, N52°12′53″, E20°38′45″), Smolice (SMH, N51°41′58″, E17°10′29″), and Strzelce (STH, N52°18′52″, E19°24′20″) dispersed across Poland (Fig. 1). Weather data indicate low rainfalls in March and April in most of the sites (Figure S1). The experiments were set up in a split-block design in three sets of 56 with three reference cultivars (Artist, Patras, and RGT Kilimanjaro) and 21 incomplete blocks per set. Each block consisted of 8 or 9 randomly assigned genotypes, accounting for three repetitions per genotype. The yield was measured for a 10-m2 plot (8 rows, 12.5 cm apart, and 10 m long). Only the inner six rows were harvested to avoid edge effects. Two agrotechnical levels were used. At the standard level (A1), the way the plants were grown and fertilized was the same as what was done for production at the respective experimental stations. At the intensive level (A2), nitrogen fertilization was increased by 40 kg/ha compared to level A1, and the plants were protected from disease and lodging. Experiments on the A1 level were conducted at five stations: DED, NAD, POB, RAH, and STH. Yield at the A2 level was measured at KBP, KOH, KRZ, MOB, and SMH stations. Grain yield was recorded along with four traits (Table 1). Grain yield was compared with the average values of three high-yielding reference cultivars (Artist, Patras, and RGT Kilimanjaro) referred to as a base of 100% (GY%). For yield, coefficients of stability (STA) were also calculated and used on GWAS to determine loci responsible for reducing environment-specific effects (see the “Data analysis” section below).
Genotyping and annotations
DArTseq technology (Diversity Arrays Technology Pty Ltd., Bruce, Australia) was used for genotyping 171 winter wheat lines. Markers with minor allele frequencies below 0.05 and over 25% of missing data were removed. The genotyping resulted in 11,117 dominant type silicoDArTs (identified by the presence or absence of the whole target marker sequence) and 8233 SNPs. Most of the genomic and marker data for wheat was annotated on Chinese Spring IWGSC v1.0, and DArT sequences were mapped to the updated reference IWGSC v2.1 at URGI. Based on the BLAST e-score values for the 1.0E-05 threshold, markers’ locations were labeled as unique, most likely, homologous, or missing. BLAST of selected DArTseq markers vs. winter (Julius, Jagger, Arina, Mattis, Mace, Norin61, Robigus, and Clair) and spring (Weebill, Lancer, Stanley, Paragon, Spelt, Cadenza, Landmark) wheat from the pangenome project (Walkowiak et al. 2020) was performed on the Galaxy platform (Afgan et al. 2018) accessible at IPK Gatersleben (https://galaxy-web.ipk-gatersleben.de/). Additionally, markers were mapped to recently sequenced wheat cultivars (Renan_2.1, Zhang1817, Attraktion, Kariega, Fielder) at NCBI (The National Center for Biotechnology Information).
Data analysis
Data were first processed using the Statistica 13.3 software (Tibco, CA, USA). The distribution of the data was checked with the Shapiro–Wilk test. The data were analyzed within a group of experiments conducted under the same agrotechnical conditions (A1 or A2) in Genstat version 21 (VSN International). Yield observations were analyzed using a linear model incorporating random effects of genotype, genotype × experiment interaction, and blocks within experiments. This model was used to calculate the yield score (BLUP) of the genotypes in each experiment, the average yield in the series, and heritability (Cullis et al. 2006). Also, for each set of experiments, an analysis was done using an additive main effect and multiplicative interaction (AMMI) model (Gauch 1992) to determine the coefficients of genotype stability (Purchase et al. 2000) which are the weighted distances of the genotypes from zero in the 2-dimensional plot of AMMI genotype scores.
An array of 8233 SNP markers was cut down to 7422 markers with known locations so that the structure of the population could be analyzed. These markers were further grouped into 705 linkage blocks based on a shared location within a 5-Mbp window (Tyrka et al. 2021a). In the same way, 8914 out of 11,117 silicoDArT markers were mapped on 21 CS wheat chromosomes, and 817 markers representing independent blocks of coupled markers were chosen. Markers with the lowest number of missing data in the blocks were used for the population structure analysis utilizing the STRUCTURE version 2.3.4 software (Pritchard et al. 2000). The admixture model was selected with 10,000 cycles and 1000 repetitions per cycle. The test was carried out over ten repetitions for ten possible subpopulations (K = 1–10). The K parameter was selected according to Evanno et al. (2005). The general (GLM) and mixed (MLM) linear models with PCA-based structure correction were used to determine the marker-trait associations using the TASSEL 5.0 (Ithaca, New York, NY, USA) (Bradbury et al. 2007). Benjamini-Hochberg (BH) method (Benjamini and Hochberg 1995) was used to adjust the P-values for allelic substitution effects for multiple tests. Associations were considered significant if the BH-corrected P-value was below 0.05, which usually meant that the original P-value was below 0.001. The Bonferroni corrected P-values for associations with silicoDArTs and SNPs were 0.0007 and 0.0006, respectively.
Results
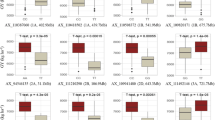
Significant variations and effects of the environment were found for all the traits studied (Table 2). The experimental design does not allow a direct comparison of the effects of applied nitrogen fertilization at A1 and A2 agrotechnical levels. The mean grain yields at A1 and A2 cultivation levels were 11.29 and 10.84 t·ha−1, respectively. At the A1 level, the average plant height was 102.2 cm, and the lodging score was 7.59. Retardant sprays were applied at the A2 level, resulting in a mean plant height of 97.6 cm and lodging of 7.07. Except for DTH, standard deviations from experiments at level A1 were lower than those at level A2 (Table 2). Experiments conducted on A1 showed higher heritability of GY compared to the A2 level (0.714 and 0.568, respectively); therefore, they may provide more stable data for GWAS and genomic prediction studies. Higher heritability values on the A1 cultivation level were also found for PH, LDG, and TKW (Table 2).
DArTseq analysis yielded two types of markers, i.e., SNPs and PAVs (silicoDArT). Due to the different characteristics of these markers, they were used separately in the analysis. SNP markers were identified as polymorphisms in 69-bp long nucleotide sequences of DArTseq markers. SilicoDArT markers, on the other hand, refer to the presence or absence of an entire marker sequence in individual genotypes. PAVs may result from mutations of a genetic or epigenetic nature in the site recognized by the restriction enzymes used to generate the marker fragments. The distribution of both marker types in the wheat genome is not even, and differences in marker saturation on the chromosomes and genomes can be noticed (Fig. 2, Table S2).
The distribution of DArTseq markers on wheat chromosomes is not random, and a higher density can be observed in the distal regions. The silicoDArT markers cover the genome better than the SNP markers, which is best seen in the proximal regions of chromosomes 4B, 4D, 6A, and 6D (Fig. 2). At the sub-genome scale, most markers were mapped to chromosomes from the B, A, and then D genomes.
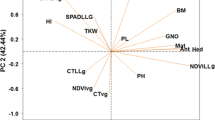
To compensate for the uneven representation of particular regions of the genome, 1706 SNP and 2383 silicoDArT markers, spaced every 5 Mbp, were selected for the analysis of population structure (Fig. 3). It was found that the genotypes could be allocated to two subpopulations, while the detailed allocation of genotypes to these subpopulations based on SNP and silicoDArT markers overlapped for only half of the lines tested (Table S1).
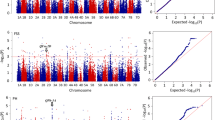
Genotypic and phenotypic data were used to identify markers associated with grain yield and the other traits studied (Tables S3 and S4, Figs. 4 and 5). Yield analysis used BLUP values, yield relative to the standard (GY%), and stability results. No MTA was found for yield data at two locations (KOH and SMH) and respective BLUP values on the A2 fertilization level. In total, 95 and 422 MTAs with GY data were identified for SNPs and silicoDArTs, respectively (Table S5). MTAs for GY%, GY_BLUP, and site-specific yield that were mapped on common linkage blocks were used to choose the main loci responsible for GY variation in the selected panel of genotypes.
Distribution of MTA for grain yield (GY) for BLUP values, relative to standard (GY%), lines yield at selected locations at A1 and A2 level (GY_LOC_A1 and GY_LOC_A2, respectively), days to heading (DTH), thousand kernel weight (TKW), lodging (LDG), plant height (PH), and stability (GY_STA) on chromosomes covered with SNP markers
Distribution of MTA for grain yield (GY) for BLUP values, relative to standard (GY%), lines yield at selected locations at A1 and A2 level (GY_LOC_A1 and GY_LOC_A2, respectively), days to heading (DTH), thousand kernel weight (TKW), lodging (LDG), and plant height (PH) on chromosomes covered by with silicoDArT markers
The universal loci significant for yield improvement were selected to better understand the main genetic factors influencing GY in an ongoing wheat breeding program. The main regions were selected when at least four independent MTAs for grain yield or stability coincided in a single linkage block. In total, 15 main regions were identified using the combined MTAs obtained for SNP and DArTseq markers (Table S5). The variation in GY_BLUP explained by the selected loci varied from 20.3% for QGy.rut-3A to 7.9% for QGy.rut-6A. Loci QGy.rut-3D, QGy.rut-5B, and QGy.rut-6B had pleiotropic effects. QGy.rut-3D shaped additionally other traits such as PH (9%), TKW (9.4%), and LDG (11.7%). QGy.rut-5B was simultaneously responsible for 15.9% of the variation in DTH, and QGy.rut-6B had a pleiotropic effect on PH (10% of variation). A single MTA (QGy.rut-5A) for stability was identified (Table 3).
MTAs for 183 SNP and 198 PAV loci were identified for heading time, lodging resistance, plant height, or thousand kernel weight (Table S6). These data were used to select 23 central regions corresponding with variation, mainly in PH, DTH, TKW, and LGD, with 18, 8, 6, and 2 linkage blocks, respectively (Table 4). Selected loci like QPhen.rut-3D and QPhen.rut-2A accumulated 76 and 44 MTAs and explained 15.8% and 10.3% of the variation in PH, respectively. In addition, two main loci (QPhen.rut-5B.1 and QPhen.rut-6A.1) were found for variation in lodging.
Discussion
Genomic regions harboring selection signatures were different by over 80% between the European and Asian germplasm, suggesting independent improvement targets from the two geographic origins (Pont et al. 2019). Therefore, the selection of genotypes for association analyses depends on the research objective. Genetically diverse or segregating populations can be used to identify major loci determining complex quantitative traits. However, not all loci determining wide variation may be relevant for ongoing breeding programs. In the genetic background uniform for selected main genes, other genes are becoming more important. GWAS on elite lines from pre-registration experiments enables the identification of regions significant for yield improvement.
We identified a set of SNP and PAV markers for 15 main regions for yield improvement in ongoing winter wheat breeding programs. We used meta-analyses (Yang et al. 2021) to find four yield-related regions overlapping with meta-QTLs (Table 5). QTL 2B-5 was reported to affect the number of grains. Regions 3A-4 and 6A-8, with the genes MOC2 and OSGA20ox1, determine variation in seed number, weight, and yield. Another QTL 7B-8 with the Brd2 gene conditions seed number and weight (Yang et al. 2021). The location of QGy.rut-5A was consistent with the position of haplotype H20271, which is associated with variation in yield (Li et al. 2018), and QYld.aww-5A explained 2.3% of the variance (Garcia et al. 2019). SNP S2B_692461029 (TraesCS2B01G495700) affecting the number of grains was localized in the region corresponding to QGy.rut-2B.4 (Pradhan et al. 2019).
Wheat yield strongly depends on the efficient accumulation of starch in grains. Starch contributes to 60–75% of the total dry weight of the wheat grain (Sawaya et al. 1984). Starch biosynthesis involves enzymes necessary to produce sucrose in the photosynthesis process. Then, sucrose is transported to amyloplasts and metabolized to hexose phosphate. Hexose phosphate is a substrate for the biosynthesis of oil, protein, and starch. During endosperm development, most of the phosphate is used to produce starch. In amyloplasts, hexose phosphate is metabolized to ADP-glucose (Shewry 2009; Thitisaksakul et al. 2012). The activities of four key enzymes involved in sucrose-to-starch conversion, sucrose synthase (SuSase), adenosine diphosphate-glucose pyrophosphorylase (AGPase), starch synthase (StSase), and starch branching enzyme (SBE), were significantly correlated with the grain-filling rate (Zhang et al. 2011). The wheat sucrose synthase 2 gene (TaSus2-2B) affecting grain weight has also been identified (Jiang et al. 2011) on chromosome 2B at 179 Mbp. We found probable sucrose-phosphate synthase 4 (LOC123076775; 3D: 4,469,233.0.4477157) is close to QGy.rut-3D. Two loci coding starch synthase 3 (LOC100136992 2B: 698,067,030.0.698075303; LOC123054641 2D: 577,064,215.0.577073489) are localized in the regions of QGy.rut-2B.2 and QGy.rut-2D.1, respectively.
Comparative analysis of yield-related traits revealed 145 meta-QTLs and candidate genes (Yang et al. 2021). About 40 genes associated with GY and related traits have been cloned (Liu et al. 2012; Rasheed et al. 2016; Nadolska-Orczyk et al. 2017), and functional markers have been converted to competitive allele-specific PCR (KASP) (Rasheed et al. 2016). However, some of these genes have already been established in modern lines. For example, no genetic differentiation was detected around the photoperiod regulation genes Ppd-B1, Vrn-2, and Vrn-3 (Cavanagh et al. 2013). Most accessions carrying the favorable haplotype at these QTLs came from CIMMYT, with 95% of them also carrying the dwarfing allele at Rht-B1 (Garcia et al. 2019). Other genes, TaNMR-1B and TaCOL5-7B, associated with yield increase in biparental populations have been cloned (Kan et al. 2020; Zhang et al. 2022) but not introduced to Polish breeding programs, and no significant MTAs were found in the respective regions.
Nine QTLs colocalized with regions identified in meta-analysis (Table 6) by Yang et al. (2021), including six (2A-2, 2B-2, 2D-2, 4A-2, 5A-3, and 6A-1) associated with kernel number, width, and yield. In addition, region QPhen.rut-3A corresponded to the IWA94 marker (3A 727.9–741.1) of a pleiotropic locus significantly associated with GY and six other yield-related traits (Li et al. 2019).
Some loci with known genes such as Rht-B1, Rht-D1, Ppd-D1, Ppd-B1, Ppd-A1, Vrn-A1, Vrn-D1, and Vrn-B1 have been routinely employed in marker-assisted selection (Garcia et al. 2019). For this set of genes, only the QPhen.rut-5A.3 locus is located at the position of the Vrn-A1 gene (NC_057806.1, 5A:589,259,335.0.589271309), while no significant effects were found for the remaining genes, which may indicate the fixation of these alleles in modern breeding lines. For example, we found no significant effect of Rht24 localized on the 6A chromosome at position 413.7 Mbp (Würschum et al. 2017).
Markers connected with plant height may also be significant for grain yield. For example, TaRht12 increases the grain number per spike and the effective tiller number and decreases thousand-grain weight (Chen et al. 2013). This gene significantly improved the elite winter wheat lines investigated (QGy.rut-5A, Table 5). Furthermore, markers Ex_c3405_203 (6B: 0.9 Mbp) and Excalibur_rep_c102984_157 (2D: 641.1 Mbp) associated with the lodging score corresponded to QGy.rut-6B (6B: 4.9–9.1 Mbp) and QGy.rut-2D.2 (2D: 633–635 Mbp), respectively (Dreccer et al. 2022).
Achieving optimal plant height is of prime importance for the cultivars’ stability, productivity, and yield potential (Griffiths et al. 2012). Improvement in wheat yield during the Green Revolution was achieved through the introduction of reduced-height (Rht) dwarfing genes. More than 50 loci and 25 height-reducing genes have been detected for wheat (Yang et al. 2021; Muhammad et al. 2021; Mokrzycka et al. 2022). Lodging may contribute to a reduction in grain yields of up to 50% (Stapper and Fischer 1990) and a loss of bread-making quality (Berry et al. 2004). The unpredictable occurrence of lodging has made it difficult for breeders to select for lodging tolerance. Ultimately, diagnostic genetic markers would help improve standability in a breeding program (Dreccer et al. 2022). By adding the semi-dwarfing genes Rht-B1b and Rht-D1b to modern wheat cultivars (Wilhelm et al. 2013; Berry and Berry 2015), the risk of lodging has been cut down. We found no effects from loci in the region of these genes. However, the TaCM (triacetin 3′,4′,5′-O-trimethyltransferase-like) gene responsible for lodging tolerance (Ma 2009) was mapped to chromosome 3B in a position consistent with QPhen.rut-3B.2.
The loci and significant SNP markers from this study can be used to create high-yield varieties by pyramiding the advantageous alleles. The introduction of a few major genes/QTL as fixed effects in GS models increases the accuracy of genomic selection for quantitative traits (Bernardo 2014) if each gene contributes to ≥ 10% of the variance (Sehgal et al. 2020). However, such significant effects of QTLs are rarely identified for complex traits such as GY in a typical GWAS study (Sehgal et al. 2016; 2017). The significant MTAs found in this study show a change in the genetic variation of the tested elite germplasm. To improve yield gains, an optimized set of markers should be used.
Data Availability
All data underlying the findings described in the manuscript are fully available without restriction from corresponding author.
Change history
13 May 2023
A Correction to this paper has been published: https://doi.org/10.1007/s13353-023-00760-0
References
Afgan E, Baker D, Batut B, van den Beek M, Bouvier D, Ech M, Chilton J, Clements D, Coraor N, Grüning BA, Guerler A, Hillman-Jackson J, Hiltemann S, Jalili V, Rasche H, Soranzo N, Goecks J, Taylor J, Nekrutenko A, Blankenberg D (2018) The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res 46:W537–W544. https://doi.org/10.1093/NAR/GKY379
Akram S, Arif MAR, Hameed A (2021) A GBS-based GWAS analysis of adaptability and yield traits in bread wheat (Triticum aestivum L.). J Appl Genet 62:27–41. https://doi.org/10.1007/S13353-020-00593-1
Alaux M, Rogers J, Letellier T, Flores R, Alfama F, Pommier C, Mohellibi N, Durand S, Kimmel E, Michotey C, Guerche C, Loaec M, Lainé M, Steinbach D, Choulet F, Rimbert H, Leroy P, Guilhot N, Salse J, Feuillet C, Paux E, Eversole K, Adam-Blondon AF, Quesneville H (2018) Linking the international wheat genome sequencing consortium bread wheat reference genome sequence to wheat genetic and phenomic data. Genome Biol 19:111. https://doi.org/10.1186/s13059-018-1491-4
Appels R, Eversole K, Feuillet C, Keller B, Rogers J, Stein N, Pozniak CJ, Choulet F, Distelfeld A, Poland J, Ronen G, Barad O, Baruch K, Keeble-Gagnère G, Mascher M, Ben-Zvi G, Josselin AA, Himmelbach A, Balfourier F, Gutierrez-Gonzalez J, Hayden M, Koh CS, Muehlbauer G, Pasam RK, Paux E, Rigault P, Tibbits J, Tiwari V, Spannagl M, Lang D, Gundlach H, Haberer G, Mayer KFX, Ormanbekova D, Prade V, Wicker T, Swarbreck D, Rimbert H, Felder M, Guilhot N, Kaithakottil G, Keilwagen J, Leroy P, Lux T, Twardziok S, Venturini L, Juhasz A, Abrouk M, Fischer I, Uauy C, Borrill P, Ramirez-Gonzalez RH, Arnaud D, Chalabi S, Chalhoub B, Cory A, Datla R, Davey MW, Jacobs J, Robinson SJ, Steuernagel B, van Ex F, Wulff BBH, Benhamed M, Bendahmane A, Concia L, Latrasse D, Alaux M, Bartoš J, Bellec A, Berges H, Doležel J, Frenkel Z, Gill B, Korol A, Letellier T, Olsen OA, Šimková H, Singh K, Valárik M, van der Vossen E, Vautrin S, Weining S, Fahima T, Glikson V, Raats D, Toegelová H, Vrána J, Sourdille P, Darrier B, Barabaschi D, Cattivelli L, Hernandez P, Galvez S, Budak H, Jones JDG, Witek K, Yu G, Small I, Melonek J, Zhou R, Belova T, Kanyuka K, King R, Nilsen K, Walkowiak S, Cuthbert R, Knox R, Wiebe K, Xiang D, Rohde A, Golds T, Čížkova J, Akpinar BA, Biyiklioglu S, Gao L, N’Daiye A, Číhalíková J, Kubaláková M, Šafář J, Alfama F, Adam-Blondon AF, Flores R, Guerche C, Loaec M, Quesneville H, Sharpe AG, Condie J, Ens J, Maclachlan R, Tan Y, Alberti A, Aury JM, Barbe V, Couloux A, Cruaud C, Labadie K, Mangenot S, Wincker P, Kaur G, Luo M, Sehgal S, Chhuneja P, Gupta OP, Jindal S, Kaur P, Malik P, Sharma P, Yadav B, Singh NK, Khurana JP, Chaudhary C, Khurana P, Kumar V, Mahato A, Mathur S, Sevanthi A, Sharma N, Tomar RS, Holušová K, Plíhal O, Clark MD, Heavens D, Kettleborough G, Wright J, Balcárková B, Hu Y, Ravin N, Skryabin K, Beletsky A, Kadnikov V, Mardanov A, Nesterov M, Rakitin A, Sergeeva E, Kanamori H, Katagiri S, Kobayashi F, Nasuda S, Tanaka T, Wu J, Cattonaro F, Jiumeng M, Kugler K, Pfeifer M, Sandve S, Xun X, Zhan B, Batley J, Bayer PE, Edwards D, Hayashi S, Tulpová Z, Visendi P, Cui L, Du X, Feng K, Nie X, Tong W, Wang L (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:eaar7191. https://doi.org/10.1126/SCIENCE.AAR7191
Assanga SO, Fuentealba M, Zhang G, Tan CT, Dhakal S, Rudd JC, Ibrahim AMH, Xue Q, Haley S, Chen J, Chao S, Baker J, Jessup K, Liu S (2017) Mapping of quantitative trait loci for grain yield and its components in a US popular winter wheat TAM 111 using 90K SNPs. PLoS One 12:e0189669. https://doi.org/10.1371/journal.pone.0189669
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J Roy Stat Soc: Ser B (methodol) 57:289–300. https://doi.org/10.1111/J.2517-6161.1995.TB02031.X
Bernardo R (2014) Genome wide selection when major genes are known. Crop Sci 54:68–75. https://doi.org/10.2135/cropsci2013.05.0315
Berry PM, Berry ST (2015) Understanding the genetic control of lodging-associated plant characters in winter wheat (Triticum aestivum L.). Euphytica 205:671–689. https://doi.org/10.1007/S10681-015-1387-2
Berry PM, Sterling M, Spink JH, Baker CJ, Sylvester-Bradley R, Mooney S, Tams A, Ennos AR (2004) Understanding and reducing lodging in cereals. Adv Agron 84:215–269
Berry PM, Sylvester-Bradley R, Berry S (2007) Ideotype design for lodging-proof wheat. Euphytica 154:165–179
Bhusal N, Sarial AK, Sharma P, Sareen S (2017) Mapping QTLs for grain yield components in wheat under heat stress. PLoS One 12:e0189594. https://doi.org/10.1371/JOURNAL.PONE.0189594
Bradbury PJ, Zhang Z, Kroon DE, Casstevens TM, Ramdoss Y, Buckler ES (2007) TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23:2633–2635. https://doi.org/10.1093/BIOINFORMATICS/BTM308
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, Forrest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D, Bai G, Pumphrey M, Tomar L, Wong D, Kong S, Reynolds M, da Silva ML, Bockelman H, Talbert L, Anderson JA, Dreisigacker S, Baenziger S, Carter A, Korzun V, Morrell PL, Dubcovsky J, Morell MK, Sorrells ME, Hayden MJ, Akhunov E (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Natl Acad Sci U S A 110:8057–8062. https://doi.org/10.1073/PNAS.1217133110
Chen L, Andrew LP, Anthony GC, Martin AJP, Hu YG (2013) GA-responsive dwarfing gene Rht12 affects the developmental and agronomic traits in common bread wheat. PLoS ONE 8:e62285
Cullis BR, Smith AB, Coombes NE (2006) On the design of early generation variety trials with correlated data. Journal of Agricultural, Biological, and Environmental Statistics 11(4):381–393. https://doi.org/10.1198/108571106X154443
Dreccer MF, Condon AG, Macdonald B, Rebetzke GJ, Awasi M-A, Borgognone MG, Peake A, Piñera-Chavez FJ, Hundt A, Jackway P, McIntyre CL (2020) Genotypic variation for lodging tolerance in spring wheat: wider and deeper root plates, a feature of low lodging, high yielding germplasm. Field Crop Res 258:107942
Dreccer MF, Macdonald B, Farnsworth CA, Paccapelo MV, Awasi MA, Condon AG, Forrest K, Lee Long I, McIntyre CL (2022) Multi-donor × elite-based populations reveal QTL for low-lodging wheat. Theor Appl Genet 135:1685–1703. https://doi.org/10.1007/S00122-022-04063-6
Ellis MH, Rebetzke GJ, Azanza F, Richards RA, Spielmeyer W (2005) Molecular mapping of gibberellin-responsive dwarfing genes in bread wheat. Theor Appl Genet 111:423–430. https://doi.org/10.1007/S00122-005-2008-6
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620. https://doi.org/10.1111/J.1365-294X.2005.02553.X
FAOSTAT. Food and Agriculture Organization of the United Nations (2022) Production domain. In: Crops. FAO, Rome, Italy. https://www.fao.org/faostat/. Accessed 02 Nov 2022
Garcia M, Eckermann P, Haefele S, Satija S, Sznajder B, Timmins A, Baumann U, Wolters P, Mather DE, Fleury D (2019) Genome-wide association mapping of grain yield in a diverse collection of spring wheat (Triticum aestivum L.) evaluated in southern Australia. PLoS One 14:e0211730. https://doi.org/10.1371/JOURNAL.PONE.0211730
Gauch HG Jr (1992) Statistical analysis of regional yield trials: AMMI analysis of factorial designs. Elsevier, Amsterdam
Golabadi M, Arzani A, Mirmohammadi Maibody SAM, Tabatabaei BES, Mohammadi SA (2011) Identification of microsatellite markers linked with yield components under drought stress at terminal growth stages in durum wheat. Euphytica 177:207–221. https://doi.org/10.1007/S10681-010-0242-8
Griffiths S, Simmonds J, Leverington M, Wang Y, Fish L, Sayers L, Alibert L, Orford S, Wingen L, Snape J (2012) Meta-QTL analysis of the genetic control of crop height in elite European winter wheat germplasm. Mol Breeding 29:159–171. https://doi.org/10.1007/S11032-010-9534-X
Isham K, Wang R, Zhao W, Wheeler J, Klassen N, Akhunov E, Chen J (2021) QTL mapping for grain yield and three yield components in a population derived from two high-yielding spring wheat cultivars. Theor Appl Genet 134:2079–2095. https://doi.org/10.1007/S00122-021-03806-1
Jia M, Guan J, Zhai Z, Geng S, Zhang X, Mao L, Li A (2018) Wheat functional genomics in the era of next generation sequencing: an update. Crop J 6:7–14. https://doi.org/10.1016/J.CJ.2017.09.003
Jiang Q, Hou J, Hao C, Wang L, Ge H, Dong Y, Zhang X (2011) The wheat (T. aestivum) sucrose synthase 2 gene (TaSus2) active in endosperm development is associated with yield traits. Funct Integr Genomics 11:49–61. https://doi.org/10.1007/S10142-010-0188-X
Jin J, Liu D, Qi Y, Ma J, Zhen W (2020) Major QTL for seven yield-related traits in common wheat (Triticum aestivum L.). Front Genet 11:1012. https://doi.org/10.3389/FGENE.2020.01012
Kan CC, Jia H, Powers C, Carver BF, Yan L (2020) Genetic characterization and deployment of a major gene for grain yield on chromosome arm 1BS in winter wheat. Mol Breeding 40:1–12. https://doi.org/10.1007/S11032-020-1106-0
Kang CS, Mo YJ, Kim KM, Kim KH, Chun JB, Park CS, Cho SW (2021) Mapping quantitative trait loci for yield potential traits in wheat recombinant inbred lines. Agronomy 11:22. https://doi.org/10.3390/AGRONOMY11010022
Keller M, Karutz C, Schmid JE, Stamp P, Winzeler M, Keller B, Messmer MM (1999) Quantitative trait loci for lodging resistance in a segregating wheat spelt population. Theoretical and Applied Genetics 98(6):1171–1182. https://doi.org/10.1007/S001220051182
Khan H, Krishnappa G, Kumar S, Mishra CN, Krishna H, Devate NB, Rathan ND, Parkash O, Yadav SS, Srivastava P, Biradar S, Kumar M, Singh GP (2022) Genome-wide association study for grain yield and component traits in bread wheat (Triticum aestivum L.). Front Genet 13:2156. https://doi.org/10.3389/FGENE.2022.982589
Korzun V, Röder MS, Ganal MW, Worland AJ, Law CN (1998) Genetic analysis of the dwarfing gene (Rht8) in wheat. Part I. Molecular mapping of Rht8 on the short arm of chromosome 2D of bread wheat (Triticum aestivum L.). Theoretical and Applied Genetics 96(8):1104–1109. https://doi.org/10.1007/S001220050845
Li A, Liu D, Yang W, Kishii M, Mao L (2018) Synthetic hexaploid wheat: yesterday, today, and tomorrow. Engineering 4:552–558. https://doi.org/10.1016/J.ENG.2018.07.001
Li F, Wen W, Liu J, Zhang Y, Cao S, He Z, Rasheed A, Jin H, Zhang C, Yan J, Zhang P, Wan Y, Xia X (2019) Genetic architecture of grain yield in bread wheat based on genome-wide association studies. BMC Plant Biol 19:1–19. https://doi.org/10.1186/S12870-019-1781-3
Li T, Li Q, Wang J, Yang Z, Tang Y, Su Y, Zhang J, Qiu X, Pu X, Pan Z, Zhang H, Liang J, Liu Z, Li J, Yan W, Yu M, Long H, Wei Y, Deng G (2022) High-resolution detection of quantitative trait loci for seven important yield-related traits in wheat (Triticum aestivum L.) using a high-density SLAF-seq genetic map. BMC Genome Data 23:1–16. https://doi.org/10.1186/S12863-022-01050-0
Liu Y, He Z, Appels R, Xia X (2012) Functional markers in wheat: current status and future prospects. Theor Appl Genet 125:1–10. https://doi.org/10.1007/S00122-012-1829-3
Liu J, Feng B, Xu Z, Fan X, Jiang F, Jin X, Cao J, Wang F, Liu Q, Yang L, Wang T (2018) A genome-wide association study of wheat yield and quality-related traits in southwest China. Molecular Breeding 38:1. https://doi.org/10.1007/S11032-017-0759-9
Lukaszewski AJ, Alberti A, Sharpe A, Kilian A, Stanca AM, Keller B, Clavijo BJ, Friebe B, Gill B, Wulff B, Chapman B, Steuernagel B, Feuillet C, Viseux C, Pozniak C, Rokhsar DS, Klassen D, Edwards D, Akhunov E, Paux E, Alfama F, Choulet F, Kobayashi F, Muehlbauer GJ, Quesneville H, Šimková H, Rimbert H, Gundlach H, Budak H, Sakai H, Handa H, Kanamori H, Batley J, Vrána J, Rogers J, Číhalíková J, Doležel J, Chapman J, Poland JA, Wu J, Khurana J, Wright J, Bader KC, Eversole K, Barry K, McLay K, Mayer KFX, Singh K, Clissold L, Pingault L, Couderc L, Cattivelli L, Spannagl M, Kubaláková M, Caccamo M, Mascher M, Bellgard M, Pfeifer M, Zytnicki M, Febrer M, Alaux M, Martis MM, Loaec M, Colaiacovo M, Singh NK, Glover N, Guilhot N, Stein N, Olsen OA, Maclachlan PR, Chhuneja P, Wincker P, Sourdille P, Faccioli P, Ramirez-Gonzalez RH, Waugh R, Šperková R, Knox R, Appels R, Sharma S, Ayling S, Praud S, Wang S, Lien S, Sandve SR, Matsumoto T, Endo TR, Itoh T, Nussbaumer T, Wicker T, Tanaka T, Scholz U, Barbe V, Jamilloux V, Ogihara Y, Dubská Z (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788. https://doi.org/10.1126/SCIENCE.1251788
Ma QH (2009) The expression of caffeic acid 3-O-methyltransferase in two wheat genotypes differing in lodging resistance. J Exp Bot 60:2763. https://doi.org/10.1093/JXB/ERP132
Mangini G, Gadaleta A, Colasuonno P, Marcotuli I, Signorile AM, Simeone R, de Vita P, Mastrangelo AM, Laidò G, Pecchioni N, Blanco A (2018) Genetic dissection of the relationships between grain yield components by genome-wide association mapping in a collection of tetraploid wheats. PLoS One 13:e0190162. https://doi.org/10.1371/JOURNAL.PONE.0190162
Mokrzycka M, Stojałowski S, Tyrka M, Matysik P, Żmijewska B, Marcinkowski R, Woźna-Pawlak U, Martofel R, Rokicki M, Rakoczy-Trojanowska M, Krajewski P (2022) Genome-wide association analysis for hybrid breeding in wheat. Int J Mol Sci 23:15321. https://doi.org/10.3390/IJMS232315321
Muhammad A, Li J, Hu W, Yu J, Khan SU, Khan MHU, Xie G, Wang J, Wang L (2021) Uncovering genomic regions controlling plant architectural traits in hexaploid wheat using different GWAS models. Sci Rep 11:6767. https://doi.org/10.1038/S41598-021-86127-Z
Nadolska-Orczyk A, Rajchel IK, Orczyk W, Gasparis S (2017) Major genes determining yield-related traits in wheat and barley. Theor Appl Genetics 130(6):1081–1098. https://doi.org/10.1007/S00122-017-2880-X
Neumann K, Kobiljski B, Denčić S, Varshney RK, Börner A (2011) Genome-wide association mapping: a case study in bread wheat (Triticum aestivum L.). Mol Breeding 27:37–58. https://doi.org/10.1007/S11032-010-9411-7
Peng J, Ronin Y, Fahima T, Röder MS, Li Y, Nevo E, Korol A (2003) Domestication quantitative trait loci in Triticum dicoccoides, the progenitor of wheat. Proc Natl Acad Sci U S A 100:2489. https://doi.org/10.1073/PNAS.252763199
Peng ZS, Li X, Yang ZJ, Liao ML (2011) A new reduced height gene found in the tetraploid semi-dwarf wheat landrace Aiganfanmai. Genet Mol Res 10:2349–2357. https://doi.org/10.4238/2011
Piñera-Chavez FJ, Berry PM, Foulkes MJ, Molero G, Reynolds MP (2016) Avoiding lodging in irrigated spring wheat: II: genetic variation of stem and root structural properties. Field Crop Res 196:64–74
Purchase JL, Hatting H, van Deventer CS (2000) Genotype × environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. South African Journal of Plant and Soil 17:101–107. https://doi.org/10.1080/02571862.2000.10634878
Poland JA, Brown PJ, Sorrells ME, Jannink JL (2012) Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS One 7:e32253
Pont C, Leroy T, Seidel M, Tondelli A, Duchemin W, Armisen D, Lang D, Bustos-Korts D, Goué N, Balfourier F, Molnár-Láng M, Lage J, Kilian B, Özkan H, Waite D, Dyer S, Letellier T, Alaux M, Russell J, Keller B, van Eeuwijk F, Spannagl M, Mayer KFX, Waugh R, Stein N, Cattivelli L, Haberer G, Charmet G, Salse J, Saintenac C, Lasserre-Zuber P, Perretant MR, Didier A, Bouchet S, Boudet J, Bancel E, Merlino M, Grand-Ravel C, Langin T, Bayer M, Booth A, Dawson I, Schweizer P, Neumann K, Kema G, Bink M, Molnar-Lang M, Megyeri M, Miko P, Linc G, Wright J, Clissold L, Krasileva K, de Vega J, Bailey P, Goody V, Wilbraham S, Anissi M, Moore J, Swan D, Watkins C, Spannagl DLM, Korol A, Krugman T, Fahima T, Rossini L, Jones H, Morris N, Costanzo A, Wicker T, Muller T, Martelli M, Ravaglia S, Bonard C, Crépieux S, Saranga J, Çakır E (2019) Tracing the ancestry of modern bread wheats. Nat Genet 51:905–911. https://doi.org/10.1038/s41588-019-0393-z
Pradhan S, Babar MA, Robbins K, Bai G, Mason RE, Khan J, Shahi D, Avci M, Guo J, Maksud Hossain M, Bhatta M, Mergoum M, Asseng S, Amand PS, Gezan S, Baik BK, Blount A, Bernardo A (2019) Understanding the genetic basis of spike fertility to improve grain number, harvest index, and grain yield in wheat under high temperature stress environments. Front Plant Sci 10:1481. https://doi.org/10.3389/FPLS.2019.01481
Pritchard JK, Stephens M, Rosenberg NA, Donnelly P (2000) Association mapping in structured populations. Am J Hum Genet 67:170. https://doi.org/10.1086/302959
Qaseem MF, Qureshi R, Shaheen H, Shafqat N (2019) Genome-wide association analyses for yield and yield-related traits in bread wheat (Triticum aestivum L) under pre-anthesis combined heat and drought stress in field conditions. PLoS One 14:e0213407. https://doi.org/10.1371/JOURNAL.PONE.0213407
Rasheed A, Wen W, Gao F, Zhai S, Jin H, Liu J, Guo Q, Zhang Y, Dreisigacker S, Xia X, He Z (2016) Development and validation of KASP assays for genes underpinning key economic traits in bread wheat. Theor Appl Genet 129:1843–1860. https://doi.org/10.1007/S00122-016-2743-X
Rasheed A, Mujeeb-Kazi A, Ogbonnaya FC, He Z, Rajaram S (2018) Wheat genetic resources in the post-genomics era: promise and challenges. Ann Bot 121:603. https://doi.org/10.1093/AOB/MCX148
Sawaya WN, Khalil JK, Safi WJ (1984) Chemical composition and nutritional quality of date seeds. J Food Sci 49:617–619. https://doi.org/10.1111/J.1365-2621.1984.TB12482.X
Sehgal D, Dreisigacker S, Belen S, Küçüközdemir Ü, Mert Z, Özer E, Morgounov A (2016) Mining centuries old in situ conserved Turkish wheat landraces for grain yield and stripe rust resistance genes. Front Genet 7:201. https://doi.org/10.3389/fgene.2016.00201
Sehgal D, Autrique E, Singh R, Ellis M, Singh S, Dreisigacker S (2017) Identification of genomic regions for grain yield and yield stability and their epistatic interactions. Sci Rep 7:41578. https://doi.org/10.1038/srep41578
Sehgal D, Rosyara U, Mondal S, Singh R, Poland J, Dreisigacker S (2020) Incorporating genome-wide association mapping results into genomic prediction models for grain yield and yield stability in CIMMYT spring bread wheat. Front Plant Sci 11:197. https://doi.org/10.3389/FPLS.2020.00197
Sheoran S, Jaiswal S, Kumar D, Raghav N, Sharma R, Pawar S, Paul S, Iquebal MA, Jaiswar A, Sharma P, Singh R, Singh CP, Gupta A, Kumar N, Angadi UB, Rai A, Singh GP, Kumar D, Tiwari R (2019) Uncovering genomic regions associated with 36 agro-morphological traits in Indian spring wheat using GWAS. Front Plant Sci 10:527. https://doi.org/10.3389/FPLS.2019.00527
Shewry PR (2009) The HEALTHGRAIN programme opens new opportunities for improving wheat for nutrition and health. Nutr Bull 34:225–231. https://doi.org/10.1111/J.1467-3010.2009.01747.X
Stapper M, Fischer RA (1990) Genotype, sowing date and plant spacing influence on high-yielding irrigated wheat in southern New South Wales. II. Growth, yield and nitrogen use. Aust J Agric Res 41:1021–1041. https://doi.org/10.1071/AR9901021
Sukumaran S, Reynolds MP, Sansaloni C (2018) Genome-wide association analyses identify QTL hotspots for yield and component traits in durum wheat grown under yield potential, drought, and heat stress environments. Front Plant Sci 9:81. https://doi.org/10.3389/FPLS.2018.00081
Sun L, Yang W, Li Y, Shan Q, Ye X, Wang D, Yu K, Lu W, Xin P, Pei Z, Guo X, Liu D, Sun J, Zhan K, Chu J, Zhang A (2019) A wheat dominant dwarfing line with Rht12, which reduces stem cell length and affects gibberellic acid synthesis, is a 5AL terminal deletion line. Plant J 97:887–900. https://doi.org/10.1111/TPJ.14168
Thitisaksakul M, Jiménez RC, Arias MC, Beckles DM (2012) Effects of environmental factors on cereal starch biosynthesis and composition. J Cereal Sci 56:67–80. https://doi.org/10.1016/J.JCS.2012.04.002
Tyrka M, Bakera B, Szeliga M, Święcicka M, Krajewski P, Mokrzycka M, Rakoczy-Trojanowska M (2021a) Identification of rf genes in hexaploid wheat (Triticum aestivum L.) by RNA-seq and paralog analyses. Int J Mol Sci 22:9146. https://doi.org/10.3390/IJMS22179146
Tyrka M, Mokrzycka M, Bakera B, Tyrka D, Szeliga M, Stojałowski S, Matysik P, Rokicki M, Rakoczy-Trojanowska M, Krajewski P (2021b) Evaluation of genetic structure in European wheat cultivars and advanced breeding lines using high-density genotyping-by-sequencing approach. BMC Genomics 22:81. https://doi.org/10.1186/S12864-020-07351-X
Vikram P, Franco J, Burgueño-Ferreira J, Li H, Sehgal D, Saint Pierre C, Ortiz C, Sneller C, Tattaris M, Guzman C, Sansaloni CP, Fuentes-Davila G, Reynolds M, Sonders K, Singh P, Payne T, Wenzl P, Sharma A, Bains NS, Singh GP, Crossa J, Singh S (2016) Unlocking the genetic diversity of Creole wheats. Sci Rep 6:23092–23092. https://doi.org/10.1038/SREP23092
Walkowiak S, Gao L, Monat C, Haberer G, Kassa MT, Brinton J, Ramirez-Gonzalez RH, Kolodziej MC, Delorean E, Thambugala D, Klymiuk V, Byrns B, Gundlach H, Bandi V, Siri JN, Nilsen K, Aquino C, Himmelbach A, Copetti D, Ban T, Venturini L, Bevan M, Clavijo B, Koo DH, Ens J, Wiebe K, N’Diaye A, Fritz AK, Gutwin C, Fiebig A, Fosker C, Fu BX, Accinelli GG, Gardner KA, Fradgley N, Gutierrez-Gonzalez J, Halstead-Nussloch G, Hatakeyama M, Koh CS, Deek J, Costamagna AC, Fobert P, Heavens D, Kanamori H, Kawaura K, Kobayashi F, Krasileva K, Kuo T, McKenzie N, Murata K, Nabeka Y, Paape T, Padmarasu S, Percival-Alwyn L, Kagale S, Scholz U, Sese J, Juliana P, Singh R, Shimizu-Inatsugi R, Swarbreck D, Cockram J, Budak H, Tameshige T, Tanaka T, Tsuji H, Wright J, Wu J, Steuernagel B, Small I, Cloutier S, Keeble-Gagnère G, Muehlbauer G, Tibbets J, Nasuda S, Melonek J, Hucl PJ, Sharpe AG, Clark M, Legg E, Bharti A, Langridge P, Hall A, Uauy C, Mascher M, Krattinger SG, Handa H, Shimizu KK, Distelfeld A, Chalmers K, Keller B, Mayer KFX, Poland J, Stein N, McCartney CA, Spannagl M, Wicker T, Pozniak CJ (2020) Multiple wheat genomes reveal global variation in modern breeding. Nature 588:277–283. https://doi.org/10.1038/s41586-020-2961-x
Wilhelm EP, Boulton MI, Barber TES, Greenland AJ, Powell W (2013) Genotype analysis of the wheat semi-dwarf Rht-B1b and Rht-D1b ancestral lineage. Plant Breed 132:539–545
Würschum T, Langer SM, Longin CFH, Tucker MR, Leiser WL (2017) A modern Green Revolution gene for reduced height in wheat. Plant J 92:892–903. https://doi.org/10.1111/TPJ.13726
Yang Y, Amo A, Wei D, Chai Y, Zheng J, Qiao P, Cui C, Lu S, Chen L, Hu YG (2021) Large-scale integration of meta-QTL and genome-wide association study discovers the genomic regions and candidate genes for yield and yield-related traits in bread wheat. Theor Appl Genet 134:3083–3109. https://doi.org/10.1007/S00122-021-03881-4
Zanke CD, Ling J, Plieske J, Kollers S, Ebmeyer E, Korzun V, Argillier O, Stiewe G, Hinze M, Neumann K, Ganal MW, Röder MS (2014) Whole genome association mapping of plant height in winter wheat (Triticum aestivum L.). PLoS One 9:e113287. https://doi.org/10.1371/JOURNAL.PONE.0113287
Zhang X, Jia H, Li T, Wu J, Nagarajan R, Lei L, Powers C, Kan CC, Hua W, Liu Z, Chen C, Carver BF, Yan L (2022) TaCol-B5 modifies spike architecture and enhances grain yield in wheat. Science 376:180–183. https://doi.org/10.1126/SCIENCE.ABM0717
Zhang LY, Liu DC, Guo XL, Yang WL, Sun JZ, Wang DW, Sourdille P, Zhang AM (2011) Investigation of genetic diversity and population structure of common wheat cultivars in northern China using DArT markers. BMC Genet 12:1–11. https://doi.org/10.1186/1471-2156-12-42
Funding
This project was supported by the grant “Genomic selection of common wheat” (DHR.hn.802.7.2022) from the Minister for Agriculture and Rural Development.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. The first draft of the manuscript was written by M. Tyrka and all authors commented on previous versions of the manuscript. Tyrka M., Krajewski P., Bednarek P.T, and Rączka K made the data analyses; Drzazga T, Matysik P, Martofel R, Woźna-Pawlak U, Jasińska D, Niewińska M, Ługowska B, Ratajczak D, Sikora T, Witkowski E, and Dorczyk A conducted field experiments and made phenotypic observations; Tyrka D conducted laboratory experiments. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Communicated by Izabela Pawłowicz.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: The original article published with inverted author names. Family name was captured first instead of given name. This is now correctly presented here.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tyrka, M., Krajewski, P., Bednarek, P.T. et al. Genome-wide association mapping in elite winter wheat breeding for yield improvement. J Appl Genetics 64, 377–391 (2023). https://doi.org/10.1007/s13353-023-00758-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13353-023-00758-8