Abstract
Fungal-mediated biosynthesis of selenium nanoparticles (SeNPs) is one of the promising biological-based nanomanufacturing process. On the other hand, the use of endophytic fungi in this respect has emerged as a new approach for green and cost-effective production of several nanoparticles. In the present study, two endophytic fungal isolates, identified as Penicillium citrinum and Rhizopus arrhizus morphologically and genetically using the ITS rRNA- gene. These fungal strains exhibited tolerance up to 40 mM NaSeO3 accompanied with red coloration of the medium that suggested selenite reduction and formation of selenium nanoparticles (SeNPs). The reduced selenite was quantified using inductively coupled plasma mass spectrometry (ICP-MS), and the results revealed that these fungi under the optimum growth conditions are able to transform > 99.0% of 3.0 mM selenite into elemental selenium. The crystalline structure, particle-sized distribution, and morphology of the purified selenium particles were extracted and characterized by different techniques including UV–Vis, X-ray diffraction and high-resolution transmission electron microscopy. The results indicated the production of regular spherical shapes of SeNPs with a majority of average size between 50 and 80 nm. Fourier transform infrared spectroscopy (FT-IR) of the produced particles revealed the presence of different functional groups that would be implicated in the synthesis process as bioreducers and capping agents. The results of optimum growth conditions revealed that the higher fungal growth resulted in the higher selenite reduction activity. Sabouraud’s and yeast extract-peptone-glucose (YPG) broth media are the best media for maximum growth of P. citrinum and R. arrhizus, respectively, and synthesis of SeNPs. The best carbon sources are sucrose and starch while NH4NO3 and NH4Cl are the best nitrogen sources for growth and synthesis of SeNPs by the fungal strains. Selenite reduction and biosynthesis of SeNPs by the fungi seemed to increase with increasing pH and maximized at alkaline pH value (9.0) being 97.94 and 97.13% for P. citrinum and R. arrhizus respectively. Initial selenite concentration markedly influenced SeNP production and the maximum rates were 96.94 and 98.47% recorded at 3.0 mM selenite for both fungi. In conclusion, the results were discussed in relation to the potentiality of these endophytic fungi for biosynthesis of SeNPs, and the favorable nutritional conditions for maximum production.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Selenium (Se) is a double-faced element; it is considered as a fundamental element in the diet, needed for health and growth of human and animals. It has a vital role in the activity of various proteins and enzymes implicated in protection against the harmful effects of free radicals [1, 2]. Selenium nanoparticles are a novel selenium species with unique biological activities and low toxicity; therefore, considerable attention has recently been paid for production of this form of selenium. Biological synthesis of selenium nanoparticles by different living organisms has emerged as an attractive alternative to overcome the side effects of physical and chemical synthesis methods [3, 4]. Much attention has been recently given to exploring fungi as potent biofactories for the synthesis of nanoparticles due to their ability to accumulate metals by physicochemical mechanisms [5,6,7,8]. Fungi are easy to culture, scaling up manipulate, and they can grow in highly concentrated media with heavy metals cations. Survival and reproduction of fungi in high selenium concentration and transformation of selenium oxy-anions into elemental selenium by fungi were previously reported [9, 10]. In [11, 12], it is stated that fungi release reductive proteins and enzymes into the extracellular medium; these biomolecules reduce selenium ions into harmless selenium nanoparticles.
On the other hand, fungi have high diversity in nature and are found in different habitats, and endophytic fungi have gained great interest in different fields. Endophytes are microorganisms that reside in the living internal tissues of plant without showing any apparent symptoms of their presence [13, 14]. These microbes have attracted the attention worldwide and considered untapped sources for bioactive compounds [15]. The reason that makes endophytes overcome their free counterparts is that endophytes produce secondary metabolites much higher than their free counterpart [16]. Moreover, the tolerance of endophytes to metals is much higher than their free-living microorganisms [17]. Accordingly, the aim of this work was to investigate the biosynthesis of selenium nanoparticles by potential endophytic fungi, Penicillium citrinum and Rhizopus arrhizus, and finding out the optimum growth conditions required for maximum production.
2 Materials and methods
2.1 Isolation of endophytic fungi
Endophytic fungi were isolated from roots and leaves of wild plants Catharanthus roseus (periwinkle) and Euphorbia milli (crown of thorns) as the method described by [18]. The plant samples were washed with tap water to remove dust and soil particles and then washed with sterile distilled water. Small pieces were immersed in 70% ethanol for 60 s and then washed with 4% sodium hypochlorite solution for 60–90 s. They were rinsed in sterile distilled water several times and pressed between sterile filter papers to remove excess water. Then they were cultured on potato dextrose agar (PDA) (peeled potato slice 200.00 g/l, dextrose/glucose 20.00 g/l, and agar–agar 20 g/l); the medium was supplemented with chloramphenicol (50 µg/ml) to prevent bacterial growth. Plates were incubated at 28 °C for 14 days and examined periodically for detection of growing fungal endophytes. The grown fungal colonies were purified and screened for selenite tolerance and reduction by growth on solid PDA medium containing 3.mM sodium selenite (Na2Se2O3) for 7 days. The higher fungal growth with red coloration suggested the higher selenite reducing strains that were selected for subsequent experimentations.
2.2 Identification of the fungal isolates
Two fungal isolates were selected, grown on (PDA) plates and examined microscopically for primarily morphological identification according to [19]. Molecular identification was also performed according to the method by [20] of using PCR-amplified ITS5 and ITS4 rRNA- gene. DNA of the fungal strains was extracted using QIAamp DNeasy Plant Mini kit (Qiagen, Germany, GmbH) and sequenced by Elim Biopharmaceuticals, USA. The compositions of the used primers were ITS1 (5'-TCC GTA GGT GAA CCT GCG G-3') and ITS4 (5'-TCC TCC GCT TAT TGA TAT GC-3'). PCR protocol for amplification was as follows: primary denaturation at 94 °C for 5 min, secondary denaturation at 94 °C for 30 s, annealing at 56 °C for 40 s, extension at 72 °C for 45 s, number of cycles 35, and final extension period at 72 °C for 10 min. DNA sequences were obtained by Applied Biosystems3130 genetic analyzer (HITACHI, Japan); a BLAST® analysis (Basic Local Alignment Search Tool) [21] was initially performed to establish sequence identity to Gen Bank accessions. The phylogenetic tree was created by the MegAlign module of LasergeneDNAStar version 12.1 [22], and Phylogenetic analyses was done using maximum likelihood, neighbor joining and maximum parsimony in MEGA6 [23].
2.3 Preparation of fungal mycelia for extracellular synthesis of selenium nanoparticles
Potato dextrose broth (PDB) medium in grams per liter (extract of peeled potato slice 200.0, and dextrose/glucose 20.0) was prepared, distributed in 250-ml conical flasks each containing 100 ml, and then sterilized in the autoclave. Each flask was inoculated with a disc (6 mm in diameter) of actively growing fungus (from the margin of 7-day-old cultures). The flasks were incubated in a shaker incubator at 28 °C, 150 rpm for 4 days, at which the fungus displayed higher selenite reduction activity as was indicated in a preliminary experiment (results not shown). After the incubation period, the fungal biomass were harvested, washed several times with double distilled water, and the dry weight was estimated Fresh biomass was placed between two filter papers and gently pressed to remove excess water, 5.0 g was inoculated into 250-ml conical flasks each containing 100 ml of 3.0 mM sodium selenite (Na2Se2O3) solution adjusted at pH 9.0, and then the flasks were incubated in a shaker 150 rpm at 28 °C for 4 days. All steps were performed in triplicate. The visible change in the color of the fungal colony to red was initially considered as a positive result for the bio-reduction of selenite and thereby SeNP synthesis.
2.4 Purification, characterization, and quantification of selenium nanoparticles
After 4 days of incubation, the fungal biomass was separated and then the suspension of SeNPs was filtered through filter paper Whatman No 1. The filtrate was centrifuged at 14,000 rpm, for 30 min; the collected precipitate was washed twice in ethanol then in double distilled H2O and dried in an oven at 60 °C for 2 h. The obtained particulates were characterized by (1) UV–Vis spectroscopy (UVS-2700/UVS-2800 Spectrophotometer, Labomed, Inc. USA), (2) Fourier transform infrared spectroscopy recorded at 400–4000 cm−1 by ATR technique (FT-IR- MODEL: ALPHA II spectrophotometer, BRUKER, Germany), (3) X-ray diffraction (XRD) patterns recorded in the range 10° ≤ 2ϴ ≤ 70° (Cu-Kα radiation, wavelength of 1.5406°A, at 30 kV, and 10 mA) using an AXS Bruker X-ray Diffractometer, Serial No: 209450, USA), and (4) transmission electron microscopy (JOEL Transmission Electron Microscope, model 1400 plus, Japan). For quantification of the selenium particulates produced ater the incubation period, 20 ml of the obtained suspension was centrifuged at 14,000 rpm, 15 °C for 20 min. The unreduced Se ions (selenite concentration in the supernatant) were quantified using ICP-MS (ICP MS Analyzer ID Code: GEBRI /01/06/00), and the amount of Se NPS was calculated as the following equation:
The amount of reduced SeNPs = Ci– Cf; where Ci is the initial concentration of sodium selenite solution, and Cf is the concentration of unreduced Se. The reduction rate of selenite into elemental selenium was calculated as the following equation:
2.5 Growth conditions for maximum production of SeNPs
The influence of growth medium on the activity of fungal mycelia for biosynthesis of SeNPs was carried out by using five growth media: malt yeast peptone glucose (MYPG) in g/l (malt extract 3.00, yeast extract 3.00, peptone 5.00, and glucose 10.00); potato dextrose broth (PDB) as mentioned above, Sabouraud’s dextrose broth (glucose 40.00, peptone 10.00), yeast peptone glucose (YPG) (yeast extract 2.00, peptone 10.00, and glucose 20.00), and modified Czapek’s Dox broth (CDB) (glucose 31.57, NH4Cl 1.22, K2HPO 41.00, KCl 0.5, MgSO4 0.5, FeSO4 0.01). To study the influence of carbon and nitrogen sources, CDA medium was used in which glucose or ammonium chloride was replaced with equivalent carbon or nitrogen respectively. For preparation of fungal mycelia for the effects of initial selenite concentration and pH values on the synthesis of SeNPs, P. citrinum and R. arrhizus were grown on YPG and Sabouraud’s media, respectively. Triplicates of 250-ml conical flasks each containing 100 ml of each medium were sterilized by autoclaving; the pH was adjusted to 6.5. Each flask was inoculated with one fungal disc from the margin of 7-day-old cultures and flasks were incubated at 28 °C, with shaking 120 rpm for 4 days. After that the obtained fungal biomass were harvested, washed 2 × with double distilled H2O, dried, and weighed. To evaluate the biosynthetic activity of fungal mycelia that had been subjected to different growth conditions, triplicates of 5.0 g fresh biomass in 100 ml of 3.0 mM selenite were used in the experimentation carried out as described above..
2.6 Statistics
The recorded results were expressed as the mean taken from triplicate measurements ± the standard deviation. Significant differences were determined using the analysis of (one-way ANOVA) test followed by Duncan’s test at P < 0.05 (SPSS software version 27, IBM, NY).
3 Results
3.1 Identification of fungal isolates
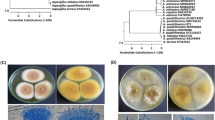
Identification of the most active two fungal strains in respect to production of SeNPs was accomplished by colony morphology and growth characteristics. The colony morphology of the first fungus grown on PDA plates for 7 days is olive green with irregular margins, and the reverse view of the colony showed yellowish pigmentation. Microscopic examination showed septated and hyaline hyphae; conidiophores are septated and biverticillate. Medullae were longer than phialides. From phialides a chain of small and globose conidia arises giving a brush-like shape which is a typical characteristic of Penicillium species [19]. The colony morphology of the second fungus grown on PDA plates is a gray-brownish mycelium with wool appearance in the right view of 7-day-old cultures, while the reverse view showed the gray color of mycelium. Microscopic examination showed non-septated hyphae; the organism is characterized by sporangiophores that arise from nodes at the point where the rhizoids are formed. Sporangiophores are in groups of 2–5, end with hemispherical columella and lemon-shaped sporangia from which hemispherical sporangiospores are released.
Molecular characterization was performed according to the method by [20] using PCR-amplified ITS5 and ITS4 rRNA- gene. The two potent strains were identified as Penicillium citrinum and Rhizopus arrhizus as shown in (Fig. 1). Sequences obtained from molecular characterization of Penicillium citrinum were deposited in GenBank under number OQ443236, then analyzed, and a phylogenetic tree was created. The obtained data confirmed the high homogeneity of the fungus with P. citrinum LT558885, P. citrinum LT558884, P. citrinum MH864240, P. citrinum LT558886, and P. citrinum LT558886 where the percent of identity was 100%. Sequences obtained from molecular characterization of Rhizopus arrhizus were deposited in GenBank under number OQ448832, then analyzed, and a phylogenetic tree was created. The obtained data (Fig. 1) confirmed the high homogeneity of the fungus with R. arrhizus MH865587, R. arrhizus MN547407, R. arrhizus MH707054, R. arrhizus MH707053, and R. arrhizus MF685318, where the percent of identity was 100%. The two strains were deposited in culture collection under number CCASU-2023-F8 and CCASU-2023-F7 for Penicillium citrinum and Rhizopus arrhizus, respectively.
3.2 UV–visible spectra, XRD, TEM, and FTIR analysis of SeNPs
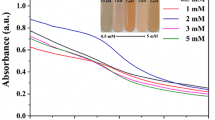
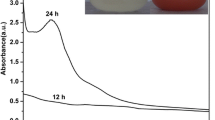
Five grams of biomass fresh weight of each fungal isolate weas inoculated into 250-ml conical flasks containing 100 ml of sodium selenite solution (3 mM) and incubated in a shaker incubator at 28 °C, at 150 rpm for 3 days. The bio-reduction of selenite to SeNPs was observed. The colorless solution of sodium selenite was turned to a red color as shown in Fig. 2A–B (insets). The surface plasmon vibrations of SeNPs were confirmed by UV–visible spectra (Fig. 2A–B, insets) that showed that the maximum absorbtion peak was observed at 257 nm for P. citrinum and at 260 nm for R. arrhizus which confirms the formation of SeNPs. The UV spectra centered between 200 and 300 nm were due to the formation and surface plasmon vibration of SeNPs [16]. The crystallinity and crystalline size of mycosynthesized SeNPs were studied by XRD analysis (Fig. 3). The diffraction peaks for the SeNPs synthesized by P. citrinum were 23.28, 26.37, 29.49, 41.06, 43.47, 45.15, 51.51, 55.57, and 61.89 corresponding to the (100), (101), (110), (102), (111), (201), (112), (202), and (210) planes respectively, and for the SeNPs synthesized by R. arrhizus were 23.35, 29.58, 41.18, 43. 55, 45.13, 51. 52, 55.71, 61.32, and 65. 01 corresponding to the (100), (101), (110), (102), (111), (201), (112), (202), and (210) planes respectively The two types of SeNPs were fitted with the card of pure hexagonal Se crystals (R050656). Additionally, the crystalline size was calculated using Scherrer’s equation. The average calculated crystalline size was 17 nm for SeNPs synthesized by P. citrinum and 14 nm for SeNPs synthesized by R. arrhizus.
Figure 4 revealed the TEM micrographs of SeNPs synthesized by P. citrinum (Fig. 4A, B) and R. arrhizus (Fig. 4C, D). They indicated the regular spherical shape of SeNPs with the majority of average size between 50 and 80 nm.
FTIR spectrum of synthetic SeNPs by the two strains displayed vibrational bands of different functional groups as shown in Fig. 5. The bands at 3263 and 3087 cm−1 characterized to -OH and -NH groups respectively. The two peaks at 2923 and 2857 cm−1 correspond to C-H in CH2. Moreover, the two bands at 1628 and 1528 cm−1 correspond to amide I and amide II in proteins respectively. The absorption band at 1451 cm−1 can be attributed to -CH2 and -CH3 in proteins, lipids, and polyesters. COO− group characterized by the absorption band at 1394 cm−1 and the peak at 1228 cm−1 corresponded to P = O. Absorption band at 1053 cm−1 corresponded to C–O–C in polyester. The vibrational bands in the region of finger print at 570 and 530 cm−1 in the spectrum of P. citrinum and R. arrhizus respectively can be attributed to the elemental selenium (Se0).
3.3 Growth conditions for maximum production of SeNPs
Table 1 shows that Sabouraud’s and YPG broth media were the best media for the fungal growth (the mycelia dry weight recorded 8.77 ± 0.02 and 6.59 ± 0.03 g/l for P. citrinum and R. arrhizus respectively). Coincidentally, synthesis of SeNPs by the fresh mycelia that had been grown on these media containing 3.0 mM selenite displayed the highest rate (99.31 and 99.19% respectively). The best carbon sources were starch and sucrose with the same order while NH4NO3 and NH4Cl were the best nitrogen sources for fungal growth and synthesis of SeNPs by both fungal strains (Tables 2 and 3). The present study hereby exhibits the necessity of weakly alkaline pH for reducing the selenite to SeNPs by action of fungi. The rate of selenite reduction increased with increasing the pH to record its optimum value at pH 9.0 being 97.94 and 97.13% for P. citrinum and R. arrhizus respectively (Table 4). Then this activity showed a decrease by increasing the pH value to record its minimum at pH 12.0. The results of the present study revealed an apparent direct relationship between fungal growth and selenite reduction activity as the condition favors higher fungal growth which resulted in the higher selenite transformation rate. The influence of initial selenite concentration on SeNP production is indicated in Table 5 that shows maximum rates 96.94 and 98.47% for both fungi detected at 3.0 mM selenite. However, with increasing selenite concentration, these rates declined down to 84.88 and 76.98%, at 40 mM selenite for both fungi.
4 Discussion
Several studies reported the biosynthesis of selenium nanoparticles by intracellular and extracellular methods [8,9,10,11]. The biosynthesis of SeNPs extracellularly by both fungi was firstly distinguished by the appearance of red color (Fig. 2), then it was confirmed by an absorption peak in UV–Vis spectrum as was reported [16]. The crystallinity and crystalline size of mycosynthesized SeNPs were also studied by XRD analysis (Fig. 3). The diffraction peaks for the SeNPs synthesized by both fungal strains confirm the hexagonal crystalline structure. Additionally, the results of TEM micrograph indicated the regular spherical shape of SeNPs is in agreement with that reported by [1,2,3,4,5].
FTIR spectra of the two types of mycosynthetic SeNPs revealed vibrational bands of different functional groups in the range of (4000–400 cm−1) suggesting a mechanism for the process of SeNP synthesis by both fungi. The vibrational bands in the region of fingerprint at 570 and 530 cm−1 in the spectrum of P. citrinum and R. arrhizus, respectively, can be attributed to the elemental selenium (Se0). Similar observations were mentioned by [16] that recorded the appearance of new bands in the spectrum of SeNPs. Interestingly, this analysis manifested the presence of biomolecules and functional groups such as hydroxyl, amine, and lipids with SeNPs that have a role as reducing agents to convert selenite to elemental selenium, and as capping agents to avoid their aggregation. Previous studies reported that the enzyme nitrate reductase is involved in bioreducing metal ions to form nanoparticles in the process of extracellular production of nanoparticles [24]. Additionally, implication of protein capping during the synthesis of selenium nanoparticles was also reported [23,24,25,26].
The present study indicated that Sabouraud’s and YPG broth media were the best growth media for both growth and synthesis of SeNPs by P. citrinum and R. arrhizus respectively. Both media contain glucose which is absorbed directly by fungal cells and involved in the metabolic pathway of the fungal cell. Moreover, these media contain various organic compounds such as free amino acids, peptides/polypeptides, carbohydrates, different vitamins, organic acids, and others [27], and the high nitrogen content increases the synthesis of enzymes and proteins. In [2,3,4,5,6,7,8, 27,28,29,30,31], it is reported that fungi release reductive proteins and enzymes into the extracellular medium, and these biomolecules would be implicated in reduction of selenite ions and transformation into harmless elemental selenium and subsequent formation of SeNps. In [27], it is demonstrated the potential use of yeast extract (YE) to enhance the generation of metal NPs such as silver and gold NPs. Besides containing rich nutrients, it also constitutes a capping agent on the surface of NPs [32]. The results also confirmed the highest growth and selenite reduction in the presence of ammonium nitrogen (Table 3). This is because ammonium ions are readily utilized by fungal cells and can be directly assimilated into the amino acids that serve as precursors for the biosynthesis proteins in addition to facilitating the synthesis of many important cellular macromolecules. Our results obviously indicated that the nutritional conditions that favor the best fungal growth are the best for selenite reduction and synthesis of SeNPs.
The present study also indicated that pH is considered a crucial factor in the process of selenite reduction and formation of selenium nanoparticles. Previous studies reported that the pH of the reaction medium has a significant effect on the size, shape, and composition of nanoparticles produced by either biological or chemical methods [32, 33]. Our results show that the two fungal strains exhibited optimum selenite reduction activity and formation of SeNPs at pH 9.0 (Table 4). This finding is in agreement with [34] who reported that pH 9 had the maximum abundance of SeNPs produced from SeO2 by the secretion of Bacillus sp. EKT1. In [1], it is also concluded that pH could play a role in the expression of specific proteins of the fungus Mariannaea sp. HJ which might affect the formation of SeNPs indirectly. This result as well as FTIR spectra analysis might suggest increasing the chance of involvement the functional groups of fungal cells in selenite reduction as electron donors due to decreasing the availability of H+ concentration. The physicochemical interactions between selenite ions and different functional groups such as carboxyl, hydroxyl, amine, and phosphoryl on the surface of biomolecules are related to hydrogen concentration. Therefore, [35] suggested the possibility of the nanoparticle manipulation of size and shape by controlling the pH of the medium. Table 5 revealed that the optimum initial selenite concentration was 3.0 mM to gain the highest rate of SeNP production. However, quantitatively, the produced selenium nanoparticles increased with increasing the selenite concentration. This result could be attributed to toxicity of selenium to the cellular biomolecules implicated in reduction process [1], and fungal tolerance to selenite [9].
In conclusion, this study confirms the potentiality of two endophytic fungal strains, P. citrinum and R. arrhizus as promising fungi for green synthesis of selenium nanoparticles in a spherical shape with a diameter ranging between 50 and 80 nm. Additionally, this work included the optimum growth conditions favorable for maximum production that has a significant importance in different fields of applications. These findings encourage working on scaling up the process of SeNP biosynthesis by these promising fungal strains and exploring the efficiency of these nanoparticles in several fields including medicine, food, and agriculture; we are currently studying their biological activities.
Data availability
By using manuscript elements (figures, tables, methods).
References
Zhang H, Zhou H, Bai J, Li Y, Yang J, Ma Q, Qu Y (2019) Biosynthesis of selenium nanoparticles mediated by fungus Mariannaea sp. HJ and their characterization. Colloidal Surf A: Physicochem Eng Asp 571:9–16
Alghuthaymi MA, Diab AM, ELzahy AF, Mazro UKE, Tayel AA, Moussa SH (2021) Green biosynthesized selenium nanoparticles by cinnamon extract and their antimicrobial activity and application as edible coatings with nano-chitosan. J Food Qual 2021:1–10
Sandhu SS, Shukla H, Shukla S (2017) Biosynthesis of silver nanoparticles by endophytic fungi: its mechanism, characterization techniques and antimicrobial potential. Afr J Biotech 16:683–698
Wadhwani SA, Gorain M, Banerjee P, Shedbalkar UU, Singh R, Kundu GC, Chopade BA (2017) Green synthesis of selenium nanoparticles using Acinetobacter sp. SW30: optimization, characterization and its anticancer activity in breast cancer cells. Int J Nanomedicine 12:6841
Srivastava N, Mukhopadhyay M (2013) Biosynthesis and structural characterization of selenium nanoparticles mediated by Zooglea ramigera. Powder Technol 244:26–29
Srivastava N, Mukhopadhyay M (2015) Biosynthesis and structural characterization of selenium nanoparticles using Gliocladium roseum. J Cluster Sci 26:1473–1482
Zambonino MC, Quizhpe EM, Jaramillo FE, Rahman A, Santiago Vispo N, Jeffrye SC, Dahoumane SA (2021) Green synthesis of selenium and tellurium nanoparticles: current trends, biological properties and biomedical applications. Int J Mol Sci 22:989
Sarkar J, Dey P, Saha S, AcharyA K (2011) Mycosynthesis of selenium nanoparticles. Micro Nano Lett 6:599–602
Gharieb MM, Wilkinson S, Gadd GM (1995) Reduction of selenium oxyanions by unicellular, polymorphic and filamentous fungi: cellular location of reduced selenium and implications for tolerance. J Ind Microbiol 14:300–311
Gharieb MM, Gadd GM (1998) Evidence for the involvement of vacuolar activity in metal(loid) tolerance: vacuolar-lacking and-defective mutants of Saccharomyces cerevisiae display higher sensitivity to chromate, tellurite and selenite. Biometals 11:101–106
Zare B, Babaie S, Setayesh N, ShahverdI AR (2013) Isolation and characterization of a fungus for extracellular synthesis of small selenium nanoparticles. Nanomedicine J 1:13–19
Liang X, Perez MAM, Nwoko KC, Egbers P, Feldmann J, Csetenyi L, Gadd GM (2019) Fungal formation of selenium and tellurium nanoparticles. Appl Microbiol Biotechnol 103:7241–7259
Amin MA, Ismail MA, Badawy AA, Awad MA, Hamza MF, Awad MF, Fouda A (2021) The Potency of fungal-fabricated selenium nanoparticles to improve the growth performance of Helianthus annuus L. and control of cutworm Agrotis ipsilon. Catalysts 11:1551
Patil RH, Maheshwari VL (2021) Endophytes: potential source of compounds of commercial and therapeutic applications. Springer Nature
EL-sayed ESR, Hazaa MA, Shebl MM, Amer MM, Mahmoud SR, Khattab AA (2022) Bioprospecting endophytic fungi for bioactive metabolites and use of irradiation to improve their bioactivities. AMB Express 12:46
Hussein HG, El-sayed ESR, Younis NA, Hamdy AE, Easa SM (2022) Harnessing endophytic fungi for biosynthesis of selenium nanoparticles and exploring their bioactivities. AMB Express 12:68
Shahid M, Zeyad MT, Syed A, Singh UB, Mohamed A, BahkalI AH, ELgorban AM, Pichtel J (2022) Stress-tolerant endophytic isolate Priestia aryabhattai BPR-9 modulates physio-biochemical mechanisms in wheat (Triticum aestivum L.) for enhanced salt olerance. Int J Environ Res Public Health 19:10883
GaryalI S, Kumar A, Reddy MS (2013) Taxol production by an endophytic fungus, Fusarium redolens, isolated from Himalayan yew. J Microbiol Biotechnol 23:1372–1380
Gilman J (1957) A manual of soil fungi. Soil Sci 84:183
Tarini NMA, Wahid MH, Ibrahim F, Yasmon A, Djauzi S (2010) Development of multiplex-PCR assay for rapid detection of Candida spp. Med J Indones 19:83–7
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Thompson JD, Higgins DG, Gibson TJ (1994) Clustal W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Tamura K, Stecher G, Peterson D, FilipskI A, Kumar S (2013) MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30:2725–2729
Salunke BK, Sawant SS, Lee SI, Kim BS (2016) Microorganisms as efficient biosystem for the synthesis of metal nanoparticles: current scenario and future possibilities. World J Microbiol Biotechnol 32:88
Tugarova AV, Mamchenkova PV, Dyatlova YA, Kamnev AA (2018) FTIR and Raman spectroscopic studies of selenium nanoparticles synthesised by the bacterium Azospirillum thiophilum. Spectrochimica Acta Part A. Mol Biomol Spectrosc 192:458–463
Kamnev AA, Mamchenkova PV, Dyatiova YA, Tugarova AV (2017) FTIR spectroscopic studies of selenite reduction by cells of the rhizobacterium Azospirillum brasilense Sp7 and the formation of selenium nanoparticles. J Mol Struct 1140:106–112
Yamal G, Sharmila P, Rao KS, Pardha-Saradhi P (2013) Yeast extract mannitol medium and its constituents promote synthesis of Au nanoparticles. Process Biochem 48:532–538
Ren L, Wu Z, Ma Y, Jian W, Xiong H, Zhou L (2021) Preparation and growth-promoting effect of selenium nanoparticles capped by polysaccharide-protein complexes on tilapia. J Sci Food Agric 101:476–485
Andrews AT (1978) The composition, structure and origin of proteose peptone component 5 of bovine milk. Eur J Biochem 90:59–65
Dobias J, Suvorova EI, Bernier-Latmani R (2011) Role of proteins in controlling selenium nanoparticle size. Nanotechnology 22:195605
Shu M, He F, Li Z, Zhu X, Ma Y, Zhou Z, Zeng M (2020) Biosynthesis and antibacterial activity of silver nanoparticles using yeast extract as reducing and capping agents. Nanoscale Res Lett 15:14
Alqadi M, Noqtah OA, Alzoubi F, Alzouby J, Aljarrah K (2014) pH effect on the aggregation of silver nanoparticles synthesized by chemical reduction. Mater Sci-Pol 32:107–111
Javed S, Sarwar A, Tassawar M, Faisal M (2016) Conversion of selenite to elemental selenium by indigenous bacteria isolated from polluted areas. Chem Speciat Bioavailab 27:162–168
Akçay F, Avcı A (2020) Effects of process conditions and yeast extract on the synthesis of selenium nanoparticles by a novel indigenous isolate Bacillus sp. EKT1 and characterization of nanoparticles. Arch Microbiol 202:2233–2243
Castro L, Blázquez ML, González F, Muñoz JA, Ballester A (2010) Extracellular biosynthesis of gold nanoparticles using sugar beet pulp. Chem Eng J 164:92–97
Acknowledgements
The authors gratefully acknowledge Prof. Ahmed Elhamalawy (Physics Department) and Dr Fatma Abu-Zayied (Chemistry Department), Faculty of Science Menoufia University, for assistance in physical and chemical analysis of the obtained results.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
The first author Mohamed Medhat Gharieb suggested the topic, designed research plan, result explanation, revising the manuscript, and data analysis. The second author Azza Mahmoud Sliman assist in suggesting the growth conditions and some result explanation, and the third author Mohamed sayed Omara carried out the research experiments, writing the draft of the manuscript and some results explanation. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gharieb, M.M., Soliman, A.M. & Omara, M.S. Biosynthesis of selenium nanoparticles by potential endophytic fungi Penicillium citrinum and Rhizopus arrhizus: characterization and maximization. Biomass Conv. Bioref. (2023). https://doi.org/10.1007/s13399-023-05084-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13399-023-05084-x