Abstract
Introduction
Persistent nasal carriage has been associated with Staphylococcus aureus infection. Previous S. aureus studies in Asia have primarily focused on clinical patients, providing limited information on persistent nasal carriage among the general adult population.
Methods
This study examined 143 healthy adults in a community in Jiangsu, China. Nasal swab samples were collected 10 times. The colonization status was identified using SPA typing. We also determined antimicrobial susceptibility, genotype, and genomic characteristics of S. aureus.
Results
The prevalence of S. aureus nasal carriage among the community individuals was on average 16.78%. The carriage rates of methicillin-resistant S. aureus and multidrug-resistant S. aureus were 6.29% and 7.69%, respectively. We identified 8.39% persistent carriers, 39.16% intermittent carriers, and 52.45% noncarriers. Furthermore, family members displayed concordance in terms of genotype and genomic characteristics.
Conclusion
Persistent nasal sampling captured intermittent carriers that were missed during short-term sampling, thus highlighting the necessity for regular community testing. SPA typing can serve as a rapid method for determining S. aureus colonization. The potential for intrafamilial transmission of S. aureus is evident, with persistent carriers being the most probable source of infection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Why carry out the study? |
The SPA typing method indeed serves as a rapid means of identifying carriers of Staphylococcus aureus. |
Longitudinal multiple nasal sampling provides a more precise representation of Staphylococcus aureus nasal carriage compared to cross-sectional sampling. |
What was learned from the study? |
The community population of Jiangsu province demonstrates a higher prevalence of ST59-MRSA strains, characterized by heightened virulence and drug resistance. Consequently, vigilant monitoring of ST59 colonization is warranted. |
Linezolid, teicoplanin, and vancomycin may serve as potential last-resort options for MRSA treatment. |
Introduction
Staphylococcus aureus, an opportunistic pathogen, is a major cause of bacteremia, pneumonia, endocarditis, and skin and soft tissue infections (SSTIs) [1]. Methicillin-resistant S. aureus (MRSA) is a prevalent multidrug-resistant S. aureus (MDRSA) which is resistant to β-lactam and other antibiotic classes [2]. Therefore, treating MRSA infections often proves more challenging and costly than methicillin-susceptible S. aureus (MSSA) infections. Community-associated MRSA (CA-MRSA) spreading in hospitals and communities increases MRSA infections, with nasal carriage of S. aureus as a major risk factor for both CA and nosocomial infections [3, 4]. Additionally, the majority of studies on S. aureus nasal carriage have employed a cross-sectional approach, relying on a single nasal sampling to determine whether the participant is a carrier. However, this leads to misclassifying the patient’s carriage state [5, 6]. Therefore, serial sampling of the nasal cavity within the community population can help in ascertaining nasal carriage prevalence and transmission mechanisms of S. aureus within these cohorts [7, 8].
Whole-genome sequencing (WGS) offers a high level of precision and can provide additional insights into resistance mechanisms, pathogenicity factors, and population structure [9]. Therefore, this study aimed to (i) report the distinct nasal carriage patterns of S. aureus among community-dwelling adults in Jiangsu, China, and (ii) analyze the antimicrobial resistance and genetic characteristics of these isolates to furnish insights into containment and intervention strategies against S. aureus infections.
Methods
Participants
In this study, we retrospectively studied surveillance samples from a community in Jiangsu Province from June 7, 2016 to June 14, 2017. We analyzed 10 nasal swab samples and demographic data (including age, sex, and body mass index (BMI)) from 144 healthy adult residents aged 18–65. The time points for nasal swab sampling were days 0, 3, 7, 10, 14, 17, 21, 42, 90, and 180. Individuals with a history of S. aureus infection, hospitalization, admission to a nursing facility, dialysis for renal failure, recent surgery, or the use of permanent indwelling catheters or medical devices within the last year were excluded from this study.
We utilized PASS v15.0 software to calculate the required sample size for the study. Based on domestic and foreign literature research, considering the high population density in China, the carriage rate of S. aureus in the community population was set at 35%, with α = 0.05, β = 0.10, and δ = 14%. Accounting for a potential loss to follow-up rate of approximately 10% during the study process, we determined that at least 127 samples were necessary. Ultimately, the first 144 surveillance samples that met the above exclusion requirements and sample sizes were included in this study.
Nasal Swabs
The participants were tested for S. aureus nasal carriage by streaking both anterior nares with sterile moistened cotton swabs to a depth of approximately 1 cm and rotating them for approximately 15 s. Nasal swabs were then placed in tubes containing sterile normal saline and refrigerated at 4 °C before being quickly sent to the microbiology laboratory within 4 h. The workflow of this study is illustrated in Fig. 1.
Culture and Identification
S. aureus colonies were detected according to the National Standard of China GB/T 4789.37–2008. After incubation at 37 °C for 45–48 h, 2–4 typical colonies were selected from each Baird-Parker agar medium (BD, Beijing, China) for subsequent identification. The S. aureus strains were identified using plasma coagulase and VITEK-2 biochemical tests.
Definitions for Colonization Status
The S. aureus nasal carriage state was assessed on the basis of the results of the nasal swab culture as follows [10, 11]:
-
Persistent carriers: Individuals showing ≥ 80% positive results for S. aureus in their consecutively tested samples.
-
Intermittent carriers: Individuals who tested positive for S. aureus in < 80% of their samples.
-
Non-carriers: Individuals with no detectable S. aureus nasal carriage.
SPA Typing
Polymorphic regions of the spa gene were amplified and sequenced for all isolates [12]. The resulting SPA types were used to categorize the carriage state of each individual. For volunteers with intermittent carriage, antimicrobial susceptibility testing and WGS were performed on strains initially isolated from the same SPA type. In cases of persistent carriage, these analyses encompassed the first strain, last strain, and strain marking a change in SPA type. Isolates that could not be classified as having any known SPA type were defined as nontypable (NT).
Drug Sensitivity Identification
The antimicrobial susceptibility of the isolates was determined using the BD Phoenix M50 Automated Microbiology System, and evaluated 23 antibiotics across seven classes (Supplementary Table 1). Staphylococcus aureus ATCC29213 was used as a quality control strain. The results were interpreted on the basis of SIR (S: Sensitive; I: Intermediate; R: Resistant) according to the Clinical and Laboratory Standards Institute (CLSI M100-S32) guidelines. MRSA was defined as S. aureus expressing the mecA gene and/or Phenocillin®, while MDRSA strains were classified as MRSA and any isolates resistant to at least three antimicrobial classes [13].
WGS
The genome was sequenced using the MGISEQ-2000 platform. The raw data were filtered and analyzed using CLC Genomics Workbench 22.0.4 software (Qiagen, Hilden, Germany) and underwent de novo assembly to generate contig sequence files for each strain. The sequence types (STs) were determined by comparing them with corresponding allelic profiles in the Multilocus sequence typing (MLST) database (http://www.pubmlst.org). BioNumerics software was utilized to create minimum spanning trees (MST), while the genetic distances in the single nucleotide polymorphisms (SNPs) between every pair of genome were computed using the CLC software. Phylogenetic trees were constructed using iTOL v6.
COL(NC_002951.2/USA/human/ST25) and 12 S. aureus strains sourced from the National Center for Biotechnology Information (NCBI) were used as references (Supplementary Table 2) [14]. All sequence data were submitted to the NCBI. Accession numbers will be provided during the review process.
Genetic Characterization Analysis
The ResFinder database served as a reference for analyzing antibiotic resistance genes (ARs), whereas the Virulence Factor Database (VFDB) was utilized to screen virulence factors (VFs). The results from these databases were directly obtained through the CLC Genomics Workbench using FASTA format data as input. To identify and compare the Integrative Conjugative Elements/Integrative Mobile Elements (ICEs/IMEs), the online tool ICEfinder (sjtu.edu.cn) was employed to support the use of both FASTA and GenBank format files.
Statistical Analysis
The data were analyzed using SPSS 26.0 to assess possible associations between S. aureus nasal carriage and age, sex, or BMI using ordinal logistic regression analysis. Statistical significance was set at p < 0.05.
Ethical Approval
This study used samples from previously in routine monitoring, and the Ethics Review Committee at the Jiangsu Provincial Center for Disease Control and Prevention decided that it does not need a special ethical review. This study was performed in accordance with the Helsinki Declaration of 1964 and its later amendments. We made sure to protect the privacy of the people involved and followed all the ethical rules when publishing the article.
Results
Participants
One participant dropped out, leaving 143 volunteers who consecutively provided nasal swabs over the 10 time periods. From each nasal swab sample, we selected 2–4 colonies for analysis, and 47.55% (68/143) of the individuals exhibited S. aureus nasal carriage. Across 10 cross-sectional samplings, the nasal carriage rates of S. aureus were displayed in Fig. 2. The average nasal carriage rate of S. aureus in the 10 samplings was 16.78%.
The study cohort consisted of 65 men and 78 women. The median age and BMI were 47 and 24, respectively. Women, younger individuals, and those with a higher BMI exhibited a greater propensity for colonization by S. aureus. However, none of these differences were statistically significant (Table 1).
SPA and Carrier State Analysis
A total of 240 S. aureus strains were isolated and subjected to SPA typing analysis. The SPA typing results revealed 44 distinct SPA types and eight NTs in 225 and 15 strains, respectively. The prominent SPA types included t437, t015, and t164.
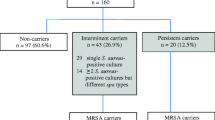
On the basis of SPA results, we found that approximately 8.39% (12/143) of the participants could be categorized as persistent carriers, 39.16% (56/143) as intermittent carriers, and 52.45% (75/143) as noncarriers. Two discernible patterns of S. aureus carriage emerged: continuous carriage with a single SPA type, and transitions between SPA types (Table 2, Supplementary Table 3). The majority of persistent carriers (66.67%, 8/12) exclusively presented with one SPA type throughout the study period. In total, 107 strains were selected for antimicrobial susceptibility testing and WGS.
MLST Typing
A total of 25 STs were identified among the 107 strains (Table 2, Fig. 3a and Supplementary Table 3). ST59 was the most prevalent ST, the second and third most common clusters were ST398 and ST1281. Of the 15 STs belonging to six different clonal complex (CC), CC1 emerged as the most prevalent, followed by CC8, CC15, CC5, and CC45. Among the nasal carriage isolates from persistent carriers, three CCs including seven STs were detected: CC1, CC5, and CC45 (Fig. 3b). CC15 were exclusively detected in intermittent carriers, indicating a heightened prevalence of intermittent carriage among individuals colonized by this particular lineage. Among the MRSA strains analyzed in this study, the three predominant types were ST59, ST398, and ST508 (Fig. 3c).
a Genetic correlation of S. aureus based on MLST profile. The minimal spanning tree allows comparison of STs for MRSA and MSSA from various sources, including hospital-acquired, community-acquired, and livestock-associated settings. The size of each circle corresponds to the number of isolates sharing the same ST. Circles are color-coded to distinguish between MRSA (red) and MSSA (green) strains. b CCs of S. aureus from different colonization states. The inner concentric sector of the pie chart delineates the frequency distribution of CCs pertaining to persistent colonizers, while the outer concentric sector illustrates the frequency distribution of CCs corresponding to intermittent colonizers. c Distribution of the STs in MRSA strains. The prominent pie chart displayed on the left side illustrates the three STs categories harboring the highest abundance of MRSA strains
Antimicrobial Susceptibility
Resistance analysis revealed that 107 strains were susceptible to nine antibiotics (Table 3). Evaluation of oxacillin resistance demonstrated that MRSA accounted for approximately 24.30% (26/107) of the isolates. Furthermore, MDRSA constituted 38.32% (41/107), with 68.29% (28/41) originating from persistent carriers (Supplementary Table 1). Drug resistance of MRSA strains are shown in the Supplementary Table 1.
The samples from the initial cross-sectional sampling (day 0) of 143 individuals were subjected to drug sensitivity tests. The prevalence of MRSA and MDRSA carriers was approximately 6.29% (9/143) and 7.69% (11/143), respectively. Among the 10 nasal samplings, 14.69% (21/143) were positive for MRSA at least once. Nearly 19.05% (4/21) exhibited persistent carriage among the 21 MRSA carriers, while the remaining 80.95% (17/21) displayed intermittent carriage.
SNP Analysis
The SNP analysis for S. aureus was conducted on the basis of the alignment of 94,963 core genome SNPs (0–54411). When comparing later-stage isolates with early-stage isolates from the same SPA/ST strains of the 12 persistent individuals, SNP distances were highly homologous (SNP < 10).
Using SNP analysis, we identified close relatedness among volunteers 66, 67, 68, 82, 83, 99, 101, 102, 123, and 125. Epidemiological investigations revealed that they belonged to four different families. The carriers within the same family exhibited identical SPA and MLST values (Fig. 4). In Family 1 and 2, ST398-MRSA and ST6-MSSA were isolated from the family members (66, 67, and 68; 82, 83) in the day 90 sampling, respectively, without any significant SNP differences between among the strains. In Family 3, a small number of SNP distances (1–7 SNPs) were observed between the five ST508-MRSA strains isolated from the family members.
a Genetic tree analysis based on SNPs of 107 S. aureus. CC1, CC8, CC5, CC15, CC45, and CC22 clones are represented by red, green, blue, orange, purple, and brown branches, respectively. Strains from volunteers belonging to the same family with similar SNPs are labeled as Family 1, Family 2, Family 3, and Family 4. The green boxes represent MSSA, while the red boxes represent MRSA. Continuous red-green outer ring indicates strains collected from persistent/intermittent carriers. Red circles denote whether the strains are MDR, while the blue circles represent strains carrying ICEs/IMEs with inserted ARs, and empty blue circles indicate strains carrying ICEs/IMEs without ARs. b 107 strains of S. aureus were compared with the retrieved 13 sequences. Gray branches indicate the retrieved S. aureus, which is distinguished from the S. aureus in this study. Different colors of the outer ring represent different STs. Stars and their colors represent different sources (community: red, hospital: blue, livestock: green)
VFs, ARs, and ICEs/IMEs
Clustering analysis was performed using SNPs, and the heat map in Fig. 5 illustrates the distribution of VFs, ARs, and ICEs/IMEs. Interestingly, the ST59 strains possessed the most VFs. Moreover, seb-selk-selq was exclusively found in the ST59 strains. Panton-Valentine leukocidin (PVL) genes (lukF and lukS) were detected in 70.09% (75/107) of the strains, including 38.46% (10/26) of MRSA isolates (Table 4). The frequencies of VFs carried by strains from the same family members showed a similar pattern. tsst-1 was present only in volunteers from Family 3. No significant difference was observed in the VFs between persistent and intermittent carriers.
Complex heatmap illustrating the distribution of VFs and ARs in all isolated S. aureus strains. The tree colors indicate strains from 4 families and ST59 strains. The right side of the dendrogram indicates whether the strain is MRSA or MSSA, carrier type, and a heatmap. Circles on the right side of the heatmap denote the carriage status of MDR or ICEs/IMEs. The left side of the heatmap shows the presence (red) or absence (blue) of VF in 107 S. aureus strains, indicating their carriage status. In contrast, the right side of the heatmap shows the presence (yellow) or absence (green) of AR genes, indicating their carriage status. Strains originating from volunteers within the same family are labeled as Family 1, Family 2, Family 3, and Family 4
The ResFinder analysis predicted a total of 23 ARs (Table 5). The most prevalent ARs were blaZ, followed by mecA and erm(C). These ARs are associated with β-lactam, macrolides/lincosamides/streptomycins (MLS), and aminoglycoside antibiotics. All MRSA isolates harbored mecA. TetK was detected in 84.62% (11/13) of the non-sensitive (R/I) tetracycline isolates. Erm(C) was found in 60% (21/35) of the erythromycin-R isolates, whereas erm(B) was detected in 83.33% (10/12) of the ST59 strains. Notably, ARs were relatively concentrated within the ST59 strains, with 91.7%(11/12) of the ST59 strains being MDRSA. 38 out of the 68 individuals carrying 47 ICEs and 10 IMEs. 13 strains (from 11 intermittent and 2 persistent carriers) possessed ARs in the ICEs (Supplementary Table 4).
Discussion
To our knowledge, this is the first longitudinal study equivalent to 10 cross-sectional studies that provides a comprehensive insight into the prevalence of nasal carriage state, resistance phenotype, and genetic characteristics of S. aureus among the community population in China. The average nasal carriage rate of S. aureus was 16.78% among the 10 samplings. This finding coincides with the prevalence rates observed in the cross-sectional studies conducted in Northern China (16.5%) and Guangzhou (23.4%) [15, 16]. Cross-sectional studies conducted outside China found S. aureus prevalence ranging from 21.9% to 47.6% among different populations, with MRSA identified in 0.7–8.81% [17,18,19,20]. The prevalence rates of MRSA and MDRSA carriage in our first cross-sectional study were approximately 6.29% and 7.69%, respectively. Therefore, compared with other countries, the general Chinese population exhibits a lower rate of nasal carriage, but a higher prevalence of MRSA nasal carriage.
In our longitudinal nasal sampling, 14.69% of individuals harbored MRSA at least once, which is twice as the rate perceived in our single cross-sectional MRSA study, and significantly surpassed those reported in both domestic and international cross-sectional studies. This suggested that relying solely on cross-sectional data to determine carrier status may underestimate true prevalence, thereby emphasizing the importance of collecting multiple consecutive samples to determine a more accurate carrier rate. However, accurately determining the persistent carriage rate of MRSA in a healthy community remains challenging because of the absence of comprehensive antibiotic susceptibility testing for all S. aureus isolates.
In this study 8.39% of individuals were identified as persistent carriers, 39.16% as intermittent carriers, and 52.45% as non-carriers. This contrasts with the findings of several longitudinal studies, which indicate 20% persistent carriers, 30% intermittent carriers, and 50% non-carriers [21]. This discrepancy may stem from the use of diverse culture techniques and intervals, different study populations, and varying colonization state definitions. Our research confirmed that the rates of persistent carriers and non-carriers decrease with longer follow-up periods and fewer culture intervals, suggesting potential misclassification of intermittent carriers as either persistent carriers or non-carriers if the follow-up period is brief or if only a few samples are cultured [22]. Previous colonization studies simplified carriage phenotypes into just two categories: persistent carriers and intermittent/non-carriers; however, given the intricate nature of the disparities between persistent and intermittent carriers, we concur that the colonization status should be divided into three categories [23, 24].
Although not statistically significant (p > 0.05), there was a trend for higher S. aureus colonization in the nasal cavity in women, younger individuals, and those with higher BMI. This result was consistent with the findings of Olsen et al., who observed a positive correlation between BMI and colonization in younger women [25]. This discrepancy highlights the need for further research to elucidate variations in carriage rates and patterns of S. aureus and MRSA among different populations.
Our study further demonstrated that persistent carriers are inclined to carry the same S. aureus genotype, while intermittent carriers may carry varying genotypes, owing to decolonization and recolonization over time [21, 22]. This dynamic nature underscores the complexity of S. aureus nasal carriage [26]. Once colonized by stubborn S. aureus for an extended period, decolonization becomes more difficult. Therefore, monitoring and treating nasal carriages is crucial to reduce nosocomial infection rates, thus emphasizing the need for expanded prehospitalization nasal swab testing.
We observed a correlation between the genotype, SNP distance, and SPA results, which confirmed that SPA typing can be used as a quick method for identifying S. aureus carriers. The correlation between SPA typing and MLST was previously assessed, and it was determined that SPA typing could effectively predict MLST clonal complexes, as defined by eBURST [27, 28]. Similarly, all t084 in our study corresponded to CC15. An intriguing discovery was made in both the aforementioned study and ours, indicating a connection between intermittent carriage and t084.
However, our study uncovered widespread resistance to penicillin and ampicillin in all strains, likely attributed to their extensive use. Furthermore, 24.30% of the strains demonstrated oxacillin resistance and all strains displayed sensitivity to ceftaroline. Oxacillin and ceftaroline were the most suitable choices for treating CA-MSSA. Linezolid, teicoplanin, and vancomycin were effective against all strains, indicating their potential as last-resort options for MRSA treatment. Clindamycin and erythromycin are commonly utilized to treat SSTIs [29]; but, our CA-MRSA strains exhibited substantial resistance to clindamycin and erythromycin, underscoring the critical need for judicious antibiotic selection. Furthermore, we identified 41 MDR strains, of which approximately 68.29% of the MDRSA strains originated from the persistent carriers. Prolonged colonization by MDRSA poses an increased potential risk for carriers as it renders treatment more challenging once infected. This finding underscores the need for regular testing of healthy individuals and timely eradication of staphylococcal carriage to prevent S. aureus infection.
Our study found that ARs are consistent with resistance phenotypes, such as tet(L) and erm(C) genes, which are associated with tetracycline and MLS antibiotic resistance, respectively, and consistent with previous research [30]. Besides, all 26 MRSA strains carried mecA and were resistant to oxacillin, which aligns with the definition of MRSA. β-lactam resistance profiles are generally biased by MRSA status. In addition, our findings suggest that the presence of ARs in ICEs/IMEs may contribute to the spread of antibiotic resistance among bacterial populations. Overall, our study emphasizes the need for a more comprehensive approach to address antibiotic resistance, considering the role of ICEs/IMEs [31].
ST59 was identified as the most prevalent ST, closely followed by ST398. Notably, 91.7% of the ST59 strains were identified as MRSA. ST59-MRSA has long been acknowledged as one of the most successful and enduring CA-MRSA clones in Asia, including China [32]. Furthermore, ST59 strains frequently harbor ARs to macrolides and β-lactams (ermB and mecA, respectively), and present significant challenges for treating infections caused by these strains [32]. All ST59 strains belonged to MDRSA, which is consistent with previous findings. Additionally, the ST59 strains carried the highest number of VFs. ST59-t437, a prominent strain in our study, has been extensively documented in China and has been associated with severe inflammatory reactions, ultimately leading to patient mortality [33]. Therefore, our findings underscore the importance of maintaining heightened vigilance regarding ST59 colonization in our healthy population. Our study revealed that the proportion of ST398-MRSA strains was comparable to that of ST398-MSSA strains, indicating an increasing prevalence of ST398-MRSA among healthy adults in Jiangsu. Furthermore, we noted a predominance of ST398 strains among intermittent colonizers, with a significant proportion of these strains carrying ICEs, thus reinforcing our hypothesis regarding the association between intermittent colonization and ICE carriage. We noted an interesting phenomenon among the intermittent colonizers (67, 137, and 140), wherein ST59 and ST398 were observed as alternating strains carried by these individuals. This observation prompts further investigations into the potential connection between these two STs.
The pathogenesis of toxic shock syndrome (TSS) is closely linked to S. aureus, which produces tsst-1. All the ST508-t015-CC45 isolates in our study carried this toxin and were classified as MDRSA. However, previous studies conducted in China have primarily associated CC5 and CC398 genotypes with tsst-1 [34]. A strong association between PVL and CA-MRSA has already been demonstrated [35, 36]. However, contrary to these findings, our results revealed that only 38.46% of CA-MRSA strains carried the PVL toxin. Similarly, during the same period as our research in Germany, 40.4% of CA-MRSA isolates harbored the virulence factor PVL [37], indicating a probability similar to ours. Therefore, we believe that solely relying on the presence of PVL cannot definitively determine MRSA strain as CA-MRSA. MRSA strains possessing both lukS and lukF genes are considered more virulent and have been associated with the development of severe necrotizing pneumonia [38]. Notably, all MRSA strains carrying both lukS and lukF genes in this study belonged to the ST59-MRSA lineage and also carried the seb-selk-selq genes, highlighting their pathogenic potential within the ST59 lineage.
Notably, S. aureus collected from individuals within the same household exhibited close genetic relatedness, suggesting the possibility of intra-household transmission, and therefore, aligning with findings from previous research [5]. Within these households, we identified both persistent and intermittent carriers, with persistent colonizers potentially serving as sources of infection, thereby facilitating transmission within the household setting. Consequently, cohabitation with persistent carriers represents a significant risk factor for healthy individuals in communities, which increases the likelihood of S. aureus infection.
Conclusion
On the basis of our findings, longitudinal multiple nasal sampling offers a more accurate depiction of S. aureus nasal carriage than cross-sectional sampling. The high carriage rates among intermittent carriers underscore the importance of regular community testing. Continuous monitoring is critical to understand and mitigate the prevalence of S. aureus, particularly the widespread ST59-MRSA strain, in healthy individuals in the Jiangsu community. SPA typing presents itself as a rapid method to determine S. aureus colonization. The potential for intrafamilial transmission of S. aureus is evident, with persistent carriers being the primary sources of infection. Oxacillin and ceftaroline are preferred treatment options for CA-MSSA. Linezolid, teicoplanin, and vancomycin demonstrated efficacy against all the strains, indicating their potential as 11th-hour treatment options for CA-MRSA. Our study revealed a notable prevalence of antimicrobial resistance in healthy individuals. This underscores the need for vigilant monitoring of S. aureus colonization and infection, along with timely alerts regarding the evolving trend of MSSA resistance to β-lactam antibiotics.
Data Availability
All sequence data were submitted to the NCBI. Accession numbers were obtained at the time of submission and will be provided during review.
References
Turner NA, Sharma-Kuinkel BK, Maskarinec SA, et al. Methicillin-resistant Staphylococcus aureus: an overview of basic and clinical research. Nat Rev Microbiol. 2019;17(4):203–18.
Gajdács M. The continuing threat of methicillin-resistant Staphylococcus aureus. Antibiotics (Basel). 2019;8(2):52.
Chuang YY, Huang YC. Molecular epidemiology of community-associated meticillin-resistant Staphylococcus aureus in Asia. Lancet Infect Dis. 2013;13(8):698–708.
Song JH, Hsueh PR, Chung DR, et al. Spread of methicillin-resistant Staphylococcus aureus between the community and the hospitals in Asian countries: an ANSORP study. J Antimicrob Chemother. 2011;66(5):1061–9.
Wertheim HF, Melles DC, Vos MC, et al. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect Dis. 2005;5(12):751–62.
Nouwen JL, Fieren MW, Snijders S, et al. Persistent (not intermittent) nasal carriage of Staphylococcus aureus is the determinant of CPD-related infections. Kidney Int. 2005;67(3):1084–92.
Self WH, Wunderink RG, Williams DJ, et al. Staphylococcus aureus community-acquired pneumonia: prevalence, clinical characteristics, and outcomes. Clin Infect Dis. 2016;63(3):300–9.
Bischoff WE, Wallis ML, Tucker KB, et al. Staphylococcus aureus nasal carriage in a student community: prevalence, clonal relationships, and risk factors. Infect Control Hosp Epidemiol. 2004;25(6):485–91.
Von Eiff C, Becker K, Machka K, et al. Nasal carriage as a source of Staphylococcus aureus bacteremia. Study Group. N Engl J Med. 2001;344(1):11–6.
Nouwen JL, Ott A, Kluytmans-Vandenbergh MF, et al. Predicting the Staphylococcus aureus nasal carrier state: derivation and validation of a “culture rule”. Clin Infect Dis. 2004;39(6):806–11.
Verkaik NJ, de Vogel CP, Boelens HA, et al. Anti-staphylococcal humoral immune response in persistent nasal carriers and noncarriers of Staphylococcus aureus. J Infect Dis. 2009;199(5):625–32.
Strommenger B, Kettlitz C, Weniger T, et al. Assignment of Staphylococcus isolates to groups by spa typing, SmaI macrorestriction analysis, and multilocus sequence typing. J Clin Microbiol. 2006;44(7):2533–40.
Zhao Y, Chen D, Ji B, et al. Whole-genome sequencing reveals high-risk clones of Pseudomonas aeruginosa in Guangdong, China. Front Microbiol. 2023;14:1117017.
Wang Z, Zhou H, Wang H, et al. Comparative genomics of methicillin-resistant Staphylococcus aureus ST239: distinct geographical variants in Beijing and Hong Kong. BMC Genom. 2014;15(1):529.
Yan X, Song Y, Yu X, et al. Factors associated with Staphylococcus aureus nasal carriage among healthy people in Northern China. Clin Microbiol Infect. 2015;21(2):157–62.
Chen B, Dai X, He B, et al. Differences in Staphylococcus aureus nasal carriage and molecular characteristics among community residents and healthcare workers at Sun Yat-Sen University, Guangzhou, Southern China. BMC Infect Dis. 2015;15:303.
Scerri J, Monecke S, Borg MA. Prevalence and characteristics of community carriage of methicillin-resistant Staphylococcus aureus in Malta. J Epidemiol Glob Health. 2013;3(3):165–73.
Mehraj J, Akmatov MK, Strömpl J, et al. Methicillin-sensitive and methicillin-resistant Staphylococcus aureus nasal carriage in a random sample of non-hospitalized adult population in northern Germany. PLoS One. 2014;9(9):e107937.
Abraão LM, Fortaleza C, Camargo CH, et al. Staphylococcus aureus and CA-MRSA carriage among Brazilian Indians living in peri-urban areas and remote communities. Antibiotics (Basel). 2023;12(5):862.
Munckhof WJ, Nimmo GR, Schooneveldt JM, et al. Nasal carriage of Staphylococcus aureus, including community-associated methicillin-resistant strains, in Queensland adults. Clin Microbiol Infect. 2009;15(2):149–55.
Chen CJ, Wang SC, Chang HY, et al. Longitudinal analysis of methicillin-resistant and methicillin-susceptible Staphylococcus aureus carriage in healthy adolescents. J Clin Microbiol. 2013;51(8):2508–14.
Vandenbergh MF, Yzerman EP, van Belkum A, et al. Follow-up of Staphylococcus aureus nasal carriage after 8 years: redefining the persistent carrier state. J Clin Microbiol. 1999;37(10):3133–40.
Brown EL, Below JE, Fischer RS, et al. Genome-wide association study of Staphylococcus aureus carriage in a community-based sample of Mexican-Americans in Starr County, texas. PLoS One. 2015;10(11):e0142130.
van Belkum A, Verkaik NJ, de Vogel CP, et al. Reclassification of Staphylococcus aureus nasal carriage types. J Infect Dis. 2009;199(12):1820–6.
Olsen K, Danielsen K, Wilsgaard T, et al. Obesity and Staphylococcus aureus nasal colonization among women and men in a general population. PLoS One. 2013;8(5):e63716.
Ritchie SR, Isdale E, Priest P, et al. The turnover of strains in intermittent and persistent nasal carriers of Staphylococcus aureus. J Infect. 2016;72(3):295–301.
Faria NA, Carrico JA, Oliveira DC, et al. Analysis of typing methods for epidemiological surveillance of both methicillin-resistant and methicillin-susceptible Staphylococcus aureus strains. J Clin Microbiol. 2008;46(1):136–44.
Sangvik M, Olsen RS, Olsen K, et al. Age- and gender-associated Staphylococcus aureus spa types found among nasal carriers in a general population: the Tromso Staph and Skin Study. J Clin Microbiol. 2011;49(12):4213–8.
Methicillin-resistant Staphylococcus aureus. Nat Rev Dis Primers. 2018;4:18034.
Wang J, Qi K, Bai X, et al. Characterization of integrative and conjugative elements carrying antibiotic resistance genes of Streptococcus suis isolated in China. Front Microbiol. 2022;13:1074844.
Guédon G, Libante V, Coluzzi C, et al. The obscure world of integrative and mobilizable elements, highly widespread elements that pirate bacterial conjugative systems. Genes (Basel). 2017;8(11):337.
Ward MJ, Goncheva M, Richardson E, et al. Identification of source and sink populations for the emergence and global spread of the East-Asia clone of community-associated MRSA. Genome Biol. 2016;17(1):160.
Liao F, Gu W, Fu X, et al. Comparison of virulence-related determinants between the ST59-t437 and ST239-t030 genotypes of methicillin-resistant Staphylococcus aureus. BMC Microbiol. 2021;21(1):264.
Wang M, Zheng Y, Mediavilla JR, et al. Hospital dissemination of tst-1-positive clonal complex 5 (CC5) methicillin-resistant Staphylococcus aureus. Front Cell Infect Microbiol. 2017;7:101.
Najafi Olya Z, Najar-Peerayeh S, Yadegar A, et al. Clonal diversity and genomic characterization of Panton-Valentine leukocidin (PVL)-positive Staphylococcus aureus in Tehran, Iran. BMC Infect Dis. 2021;21(1):372.
Bhatta DR, Cavaco LM, Nath G, et al. Association of Panton Valentine leukocidin (PVL) genes with methicillin resistant Staphylococcus aureus (MRSA) in Western Nepal: a matter of concern for community infections (a hospital based prospective study). BMC Infect Dis. 2016;16:199.
Klein S, Menz MD, Zanger P, et al. Increase in the prevalence of Panton-Valentine leukocidin and clonal shift in community-onset methicillin-resistant Staphylococcus aureus causing skin and soft-tissue infections in the Rhine-Neckar Region, Germany, 2012–2016. Int J Antimicrob Agents. 2019;53(3):261–7.
Gillet Y, Issartel B, Vanhems P, et al. Association between Staphylococcus aureus strains carrying gene for Panton-Valentine leukocidin and highly lethal necrotising pneumonia in young immunocompetent patients. Lancet. 2002;359(9308):753–9.
Acknowledgements
We would like to express our sincere gratitude to the Taixing Center for Disease Control and Prevention for their support with organization and sampling.
Funding
The study was supported by National Natural Science Foundation of China (82222062), and Jiangsu Provincial Science Fund for Distinguished Young Scholars (BK20220064). The Rapid Service Fee was funded by the authors.
Author information
Authors and Affiliations
Contributions
Wenjing Hu, Yang Wang, Lu Zhou, Kai Chu, Pengfei Jin, Qi Liang, Jingxin Li, Zhongming Tan, and Fengzai Zhu made significant contributions to the work reported, whether in the conception, study design, execution, data acquisition, analysis, and interpretation, or in all these areas. They participated in drafting, revising, or critically reviewing the article and agreed on the journal to which the article has been submitted. They also gave final approval of the version to be published. and have committed to be accountable for contents of the article.
Corresponding authors
Ethics declarations
Conflict of Interest
Wenjing Hu, Yang Wang, Lu Zhou, Kai Chu, Pengfei Jin, Qi Liang, Jingxin Li, Zhongming Tan, and Fengzai Zhu report no conflict of interest in this work.
Ethics Statement
This study used samples from routine monitoring conducted previously, and the Ethics Review Committee of the Jiangsu Provincial Center for Disease Control and Prevention determined that a special ethical review was not necessary. The study was conducted in accordance with the principles of the Declaration of Helsinki. We ensured to protect the privacy of the participants involved and followed all the ethical guidelines during the publication of the article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Hu, W., Wang, Y., Zhou, L. et al. Nasal Staphylococcus aureus Carriage and Antimicrobial Resistance Profiles Among Community-Dwelling Adults in Jiangsu, China. Infect Dis Ther 13, 1215–1233 (2024). https://doi.org/10.1007/s40121-024-00969-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40121-024-00969-4