Abstract
At present, various blood-based biomarkers have found their applications in the field of sports medicine. This current opinion addresses biomarkers that warrant consideration in future research for monitoring the athlete training load. In this regard, we identified a variety of emerging load-sensitive biomarkers, e.g., cytokines (such as IL-6), chaperones (such as heat shock proteins) or enzymes (such as myeloperoxidase) that could improve future athlete load monitoring as they have shown meaningful increases in acute and chronic exercise settings. In some cases, they have even been linked to training status or performance characteristics. However, many of these markers have not been extensively studied and the cost and effort of measuring these parameters are still high, making them inconvenient for practitioners so far. We therefore outline strategies to improve knowledge of acute and chronic biomarker responses, including ideas for standardized study settings. In addition, we emphasize the need for methodological advances such as the development of minimally invasive point-of-care devices as well as statistical aspects related to the evaluation of these monitoring tools to make biomarkers suitable for regular load monitoring.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
A variety of novel blood-based biomarkers such as cytokines, chaperones, enzymes and other inflammatory signaling molecules are sensitive to acute and chronic exercise load and potentially useful for the monitoring of athlete training load. |
Omics, RNA and DNA approaches as well as genetic testing could further improve athlete workload management. |
Longitudinal study designs will shed light on acute and chronic training load responses to assess the suitability of emerging biomarkers for regular load monitoring. Methodological advances such as the development of point-of-care devices, as well as overcoming statistical challenges, are critical for the further progress of load monitoring with biomarkers. |
1 Introduction
Biomarkers (i.e., “indicators of normal biological processes, pathogenic processes, or responses to an exposure” [1]) have seen an upsurge in so-called personalized medicine, that is, procedures individually tailored to patients. They have great potential to be an objective complement to other screening methods in the diagnosis and prognosis of diseases, therapeutic decisions, as well as in the assessment of therapeutic success. For instance, concentrations of biomarkers have been shown to predict mortality or diagnose disease severity in certain conditions [2, 3]. Recent developments are moving from single biomarker measurement to multiple marker approaches to determine a whole range of biological measures. Artificial intelligence (AI) could be utilized to help analyze this data and assist physicians to make informed decisions about treatment options [4].
Transferring this perspective to the field of elite sports, ‘treatment options’ could be interpreted as the prescription of personalized training load and recovery. Blood-based biomarkers, which in principle are able to objectively reflect training load, fatigue, and recovery needs, are already being applied by practitioners to facilitate decision making and to ensure an individualized load management (i.e., prescription, monitoring, and adjustment of workload [5]) aimed at optimizing performance and avoiding injury [6, 7]. However, established biomarkers such as creatine kinase (CK) or lactate, while sensitive to training load and both convenient and quick to measure with point-of-care (POC) devices, primarily capture a specific physiological domain when measured solely in the absence of other markers. This means that practitioners in professional sports settings need to capture multiple markers to holistically assess the training response and consequently manage athlete workloads. High chronic training loads, body composition, as well as delayed peaks in concentrations may further lead to misinterpretations [8,9,10]. In the search for other reliable alternatives, a variety of novel biomarkers (e.g., CD163, heat shock proteins [HSP], cell-free DNA [cfDNA], blood cell ratios) have shown marked increases after standardized exercise settings [11,12,13,14], providing potential added value for load management.
To date, the cost and effort of measuring innovative parameters, such as cfDNA, HSP or cytokines, on a regular basis are still high, making them inconvenient for monitoring purposes. Due to evolving technology, these biomarkers may soon become relevant to sports medicine practitioners through POC devices. Many studies in this area also aim to determine a generalizable resilience or trainability profile of an athlete that may be relevant throughout their career, e.g., genetic markers for ligament injury [15]. Underlying all of these approaches in sports is a similar rationale to the clinical setting, which is the assessment of biomarkers to individualize treatment.
This current opinion article discusses innovative and load-sensitive biomarkers followed by advances in biotechnology, such as omics approaches, that enable the assessment of a variety of biomarkers and biosignatures. The section on practical applications focuses on methodological and statistical considerations, and ideas for future study designs to establish emerging biomarkers for load management in sports science.
2 Evidence and Usefulness of Emerging Biomarkers for Workload Management
Many researchers have increased their efforts to identify new robust surrogate biomarkers to help in the management of training load [16, 17]. The goal is to identify biomarkers that represent appropriate and reliable responses to training load, reflect recovery cycles and regeneration processes, and thus are expected to make an important contribution to the field of load management in professional sports settings.
A frequently used starting point in recent years has been the immune system. Physical activity induces a systemic immune response manifested by leukocytosis, a shift in the proportion of leukocyte subpopulations, and the release of numerous pro- and anti-inflammatory cytokines [18]. Some of these molecules have a close relationship to metabolic changes during exercise or belong to the class of myokines, which act at the immunological level in addition to metabolic signaling pathways [19]. These immunological markers are very sensitive to acute exercise, depending, among other factors, on the duration and intensity of the load, with partial regulation also observed depending on the type of exercise [20, 21]. Some of these markers are classified as chemokines, such as chemokine-ligands, while others are classified as enzymes, such as myeloperoxidase [22]. The potential use of these proteins as biomarkers in exercise settings would also be of interest, as immunological markers indicate differential disturbances in physiological homoeostasis or tissue integrity. For example, some markers are regulated depending on the level of muscle damage, while others have dependencies on, for example, neuroimmunological processes, energetic deficit, or heat production [21, 22].
We understand that blood reflects only a small fraction of immunological processes, but these are quite well studied. Accordingly, immunological markers can provide an estimation of the internal load at different physiological levels and indicate regeneration processes. Markers of oxidative stress show a close connection to the immune response. Physical activity induces an increased level of reactive oxygen species (ROS). Accordingly, more products of oxidative stress are released, which can also be detected directly or indirectly in the blood [23]. Here, some markers are currently in focus that reflect the level of oxidative stress in the context of athletic exertion and the subsequent recovery cycles. Markers of the stress proteome, such as HSP, have a relationship to both immune changes and oxidative stress. For example, blood levels of HSP70/72 and HSP90 respond very sensitively to physiological stressors, such as endurance exercise, and are also quickly regulated back after the end of the exercise. Interestingly, HSP70 increases were shown to depend on intensity and duration with evidence of changes in resting concentrations after intensified training periods [14]. In line with these findings, recent studies have demonstrated the importance of Ca2+-binding proteins of the S100 group as possibly useful biomarkers in sports. S100 proteins represent a class of calcium-binding proteins that are sensitively released depending on the exercise load [24], showing increases after long-lasting endurance exercise with S100 Calcium Binding Protein B increasing mainly after running but not cycling exercise [25, 26]. In addition, Irisin is released in response to exercise as a result of proteolytic cleavage of FNDC5 protein present in the membrane of myocytes. The exact physiological mechanism of release is not yet completely understood [27], however, chronic training may lead to decreased levels of circulating irisin [28]. Another important category represents the field of adenosine triphosphate (ATP) metabolism catabolites, such as ammonia, hypoxanthine, or xanthine. In this respect, the combined measurement of lactate, ammonium, and hypoxanthine was shown to indirectly reflect changes in energy status during exercise [29]. In addition, hypoxanthine is discussed as being a promising marker for training status and a predictor of sport performance in athletes [30].
Of note, there are many other metabolites being discussed as suitable biomarkers in sports. The significant strain on the metabolism during exercise induces significant blood concentration changes after exercise in the composition of the plasma metabolome. For example, lipids and lipid-like substances are mobilized into the blood during long-lasting endurance exercise [31]. Metabolites can also provide information about the training status, as the metabolome of trained athletes changes according to the training adaptation at the metabolic level. Accordingly, some metabolites associated with cardiopulmonary fitness have already been identified, such as several acyl-alkyl-phosphatidylcholine species [32], while others are more prevalent in strength athletes, such as phosphatidylcholines [33]. Table 1 outlines relevant exercise response biomarkers that could potentially help practitioners monitor and manage athlete loads in the future.
These markers were selected as they exhibit clear regulation from homeostasis by acute exercise and/or regular training, as well as a re-regulation to baseline during recovery. Initial conclusions about their usefulness are thus already possible, as temporal changes in the course of acute load-recovery cycles and also short-term cumulative training cycles have been shown (e.g., [11, 12, 20]). In the long term, the extent to which such markers reflect adaptive processes, for example by being associated with changes in cardiopulmonary fitness, is of particular interest [81]. Furthermore, for some markers, it is not understood to what extent these markers are differentially regulated during different types of exercise, such as strength or endurance training. Knowledge of sex-specific characteristics or classifications relevant to training status is not yet available and points to further limitations that need to be explored in the validation phases.
Most studies in this area were conducted in the context of acute exercise (Table 1), suggesting the use of biomarkers immediately after cessation of exercise to assess the acute physiological response in combination with parameters of external workload. In addition to evaluating training load, biomarker responses can provide additional insight, such as an estimation of the athlete's risk of disease as indicated by various cytokine responses after exercise (e.g., in IL1-Ra, IL-10) [82]. However, there is a need for easy-to-use instruments, such as POC devices, that allow practitioners to measure these biomarkers in a timely and simple manner. In addition, marker-specific obstacles need to be considered or overcome (e.g., low specificity, venous blood needed) before they can be regularly used for load monitoring. Section 4 will therefore address methodological aspects of biomarker development.
It should be noted that there are other physiological domains that we have thoroughly investigated. For example, proteomics approaches have identified fitness- or body fat-associated biomarkers such as leptin [83]. However, our focus has been on load-sensitive biomarkers of immunology rather than fitness or anthropometry, as we consider the latter to be less relevant for load management. Another interesting area of research could be cardiac biomarkers such as NT-proBNP or cardiac troponin, which have shown load-dependent patterns during exercise [84, 85]. As new biomarkers are identified, bioanalytical methods for measuring biomarkers continue to evolve, as discussed in Sect. 3.
3 Bioanalytical Approaches with a Focus on Protein, Free-Circulating Nucleic Acids and Genetic Biosignatures
3.1 Omics—Deciphering Bio-Signatures with a Multiple Marker Approach
Exercise induces a multilevel and complex physiological stimulus that is reflected at the level of gene expression, epigenetic processes, protein synthesis, energy metabolism, and the associated metabolome [86]. Sophisticated large-scale analytical methods to quantify gene expression (transcriptomics), proteins (proteomics), lipids (lipidomics) and metabolites (metabolomics) in different organs and tissues, in the context of exercise mainly in the muscle and blood, are now available to identify biomarkers that define different status of stress and recovery cycles or physiological adaptation processes. The so-called multi-omics profiling can even couple such technologies to analyze the interrelationships of the individual levels. Such technologies have also been used in sports and have significantly developed the choreographic interaction of different physiological levels [87].
Omics technologies need to be coupled with appropriate bioinformatic methods so that the truly relevant ones can be identified from the multitude of possible targets. Pathway and network analyses are an essential part of the bioinformatics analysis, which can strengthen the basic scientific understanding of the regulation of the markers. Such approaches have evolved in recent years to pathway and network-based biomarker analysis, which has a particular focus on discovering panels of markers that can serve as a biosignature rather than singular biomarkers. Bioinformatic databases have supported the functional analysis and interpretation of results [88]. Accordingly, today’s omics technologies can be valuable contributors to the identification of biomarkers, or biosignatures, and their physiological classification in the context of sports.
However, this approach should be understood as the first step in the complex process of biomarker research. Multi-omics profiling can, as a first step in identifying biomarkers, highlight those candidates within a network that have the greatest potential to be more robust to individual differences or to environmental factors other than training load. This idea can be illustrated by a study from Nieman et al. [57], who attempted to identify candidate biomarkers through a proteomics approach for functional overreaching over 3 days of extreme stress. Of nearly 600 proteins, over 70 proteins were identified that increased during the following recovery period at rest. Finally, the authors suggested the application of the identified biomarker panel in a more sophisticated training study with additional monitoring tools to examine the potential of these markers for predicting overtraining in athletes.
3.2 Specific Approaches for Protein and Free Nucleic Acid Detection
Proteins are considered key biomarkers that allow monitoring of the training process. Due to their multiple functional roles as enzymes, cellular signaling molecules, co-factors, and neurotransmitters, many of these factors reflect load and recovery as well as adaptation processes. High-throughput analyses, such as those possible using multiplex assays or mass spectrometry, can provide indications for the identification of potential candidate proteins as a first step [16, 89].
In analogy to proteins, DNA or RNA are complex macromolecules that can be studied inside or out of cells as cfDNA [90] or circulating RNA (cirRNA). As for protein analysis, mass spectrometry gains importance in DNA and RNA analysis since it is not only capable of identifying post-translational protein modifications [91] but also nucleotide modifications of functional and physiological relevance. For proteins in fluids, enzyme-linked immunosorbent assay (ELISA) are used in laboratories, while derived from this principle, lateral flow immunoassays are used at POC. Lateral flow assays can nowadays also be used to analyze DNA or RNA at POC and techniques were developed during the COVID-19 pandemic to do this semi-quantitatively using the cell phone [92]. For high-throughput medical diagnostics of several different proteins or nucleic acids to be analyzed simultaneously in one sample, the principles of ELISA have been combined with flow cytometry.
Together with proteins, nucleic acids belong to a group of circulating macromolecules that can be subjected to covalent modifications in addition to their amino acid or base sequence and can therefore contain essential qualitative information in addition to their quantity. In analogy to typical glycosylation, phosphorylation, or citrullination of amino acids, DNA-bases can be methylated carrying epigenetic information [93]. While amino acid modifications are mostly indicators of a different conformational state, functional status, or stability, methylation of DNA can for certain sequence parts be cell-type specific and may therefore reveal the origin of DNA [93]. At present, there is a rapidly evolving field of so-called liquid biopsy by analyzing circulating nucleic acids in blood [94].
At the DNA level, quantification of cfDNA is a putative forward approach for monitoring acute [11, 79] and chronic [78, 80] exercise load. Quantitative analysis in combination with qualitative information has just been able to confirm the origin of cfDNA from cells of the hematopoietic lineage [95]. Cell-type specific epigenetic analysis of cfDNA has revealed that most of the DNA released during exercise is released within minutes from neutrophils [77]. Thus, unlike versatile cirRNA, cfDNA is the lead molecule for a specific process that is initially triggered with the onset of movement, that is, neutrophil activation. Increases of cfDNA, therefore, fall into the category of markers attributed to neutrophil activation that have been described as a prominent pathway affected during acute exercise in several omics approaches and a recent multi-omics approach [87]. Analysis of DNA by a highly sensitive detection technology has furthermore confirmed that the majority of DNA released by neutrophils is indeed free-floating in the bloodstream and not associated with or incorporated in extracellular vesicles [96, 97], while the full complexity of mRNA, long non-coding RNA, or microRNA analysis is not yet well investigated or even understood. Of note, in the field of cirRNA analysis, small differences in the analytical procedures can lead to unpredictably different outcomes. CirRNA can be protected from rapid decay by extracellular vesicles, which in turn are subject to significant change upon the onset of exercise [97, 98].
3.3 Genetic Testing—The Promise of Once in a Lifetime Individualization
In contrast to previous approaches, genetic testing already plays a prominent role in load management, whereby a one-time measurement can already be relevant for the entire career. The determination of, for example, injury propensity [15, 99, 100] could be used to identify at-risk individuals and prescribe individualized exercise programs to counteract this injury propensity.
In recent years, the Genome-Wide Association Study approach has enabled a detailed understanding of the importance of specific genetic polymorphisms for sport-specific performance [101]. Basic research on gene polymorphisms and their association with phenotypic traits relevant in sports has generated evidence that specific genetic polymorphisms can affect training responses, the ability to regenerate, and the susceptibility to injury [99]. For example, IGF-1R 275124 A>C rs1464430 polymorphism was shown to be represented in endurance athletes and PPARGC1A polymorphism was also shown to be related to endurance performance. Furthermore, the RR genotype of ACTN3 R577X polymorphism, the C allele of IGF-1R polymorphism and the gene variant FTO T>A rs9939609 and/or their AA genotype showed a relation to muscle strength while gene variants of the MMP group (rs591058 and rs679620) and the COL5A1 rs13946 polymorphisms are associated with increased susceptibility to injury in athletes [99]. However, most of these studies are associative studies, which limits their practical value for designing training programs.
Thus, it seems particularly reasonable to proceed to interventional studies using genetic information. One of the first studies that used genetic testing to differentiate a training program was conducted by Jones et al. [102]. In this study, an algorithm of 15 different single nucleotide polymorphisms (SNPs) was developed to determine a power/endurance score ratio. The SNPs used included variants of the genes ACE, ACTN3, ADRB2, AGT, BDKRB2, COL5A1, CRP, GABPB1, IL6, PPARA, PPARGC1A, TRHR, VDR, and VEGFA. One group of athletes performed an 8-week strength training program aligned by genetic profile to be either more intense or of greater volume, while the control group performed exactly the training that did not match their genotype. Although the study can be critiqued methodologically in some aspects, the initial evidence generated showed that pre-intervention genetic profiling and assignment to a training program leads to a favorable outcome in terms of explosive power and aerobic fitness.
In relation to the release of circulating markers, genetic testing has a potentially growing importance. For some circulating markers, such as CK, there are high responders and low responders, which is most likely due to genetic polymorphisms. Accordingly, testing of such SNPs prior to the actual measurement of the markers may help to classify athletes with respect to their responder assignment in the future. For this purpose, causal links between the presence of individual or multiple SNPs still need to be established in future studies [103].
However, the extent to which genetic testing is finding its way into sports could also be a cause for concern. There has been a recent surge in commercial direct-to-consumer genetic testing without the involvement of a physician. Athletes and coaches are naturally focused on implementing, for example, effective training strategies to optimize performance, which may make this cohort particularly susceptible to such testing, believing that those results will contribute to improved performance outcomes [104]. Pickering and Kiely [105] found in 110 athletes and 133 practitioners that about 10% of both groups had already made use of such tests. However, when genetic testing is not conducted by experienced personnel, this may lead to misinterpretation and potentially serious data security issues and raise questions about whether athletes undergoing such procedures are actually fully informed in terms of purpose, possible results, and ramifications [104, 106]. In comparison with other tests, genetic testing may accidentally deliver an outcome that is as a stand-alone not valid enough for diagnostics but provides an outcome that should prompt further medical diagnostics. If the tests are not initiated by the athletes themselves, but by a sports authority, and are thus not exclusively voluntary, they could undoubtedly be classified as unethical [104, 107]. The fact that most genetic studies have been conducted in people of European ancestry [108] also appears to be problematic as the predictive performance may not be relevant for people of other ancestries.
In summary, genetic testing in sports has yielded interesting findings but, unlike disease-related genetic testing, it is still in its infancy. To date, neither tests for talent identification [107, 109] nor tests for exercise prescription or injury prevention [106] seem to have sufficient predictive value.
4 Considerations for Researchers in Upcoming Studies
In this section, we highlight topics to further develop load management: (i) biomarker development, (ii) methodological advances, and (iii) statistical considerations.
4.1 Biomarker Development
Biomarker development typically proceeds in three phases, that is, discovery, verification, and validation [110]. Biomarker discovery phase is primarily concerned with identifying a set of potential marker candidates in a given experimental design, using approaches such as omics as described above [40, 111, 112] to select potentially suitable load-sensitive markers for future studies. Verification is regularly performed by repeated measurements. In proteomics, for example, candidate biomarkers are subjected to additional analysis to verify their identity and expression in the same samples. The remaining identified markers can then be tested for sensitivity, specificity, and repeatability in a manner similar to the discovery phase (analytical validation). Finally, in the clinical validation (i.e., ‘qualification’ [110]), the identified reliable and robust markers [12] can be applied to similar interventions [110]. At each stage, the markers of interest are further narrowed, leading to an intensified testing of biomarker performance for clinical use [110].
The potential for biomarker discovery also arises from the primary use of markers in clinical settings and their transfer to sports. Historically, cfDNA has been introduced as a marker for cancer screening [113] but also showed a pronounced acute response after strenuous exercise [11] and first evidence of indicating overtraining status [78]. In addition, well-known ratios such as the neutrophil-to-lymphocyte or testosterone-to-cortisol ratios were also introduced as promising tools for load monitoring. In this respect, associations with overtraining or anabolic adaptation to training have already been observed [13, 114, 115].
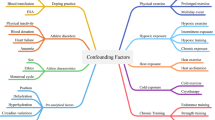
Regarding further evaluation of biomarkers, standardized test–retest settings seem reasonable for assessing the reliability of a biomarker and its response to an acute load [12]. An adequate post-exercise observation period provides information on the reregulation to baseline and the biological half-life time of these biomarkers. Amateur athletes or youth squads of elite teams [7] can be used as a study population, with a subsequent transfer to the domain of professional athletes. Accurate characterization of participants taking into account high and low responders, the documentation of potential confounding variables due to the effects of environmental factors, and the determination of the external load can provide insights into dose–response patterns [14, 116, 117].
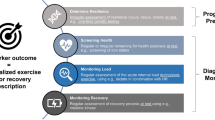
Finally, longitudinal study designs are efficient in observing the potential of biomarkers to ‘forecast’ an event such as an injury or illness, as well as to reveal the acute and chronic response to an unusual intervention, such as training camps. Repeated blood sampling at rest and post-exercise—in combination with established monitoring tools—can help to assess the training load and the corresponding biomarker responses while establishing individualized reference values [118] with a conclusion on the principal suitability of the biomarker (panel) (Fig. 1). Furthermore, insights can be gained about gender specifics, and effects of training volume and training status [119].
4.2 Methodological Considerations
Since repeated blood sampling is necessary to generate reliable data, the development of convenient and efficient measurement methods is important to avoid excessive burden on athletes and to keep the amount of blood collected low [120]. Various cytokines in capillary blood from the finger are regulated similarly to those in venous blood. Regarding blood plasma from the earlobe, there is less data on biomarker regulation [121, 122] although the measurement is possible in principle [123], which would greatly improve the accessibility of the biomarker and would not require specialized personnel. Such a continuous transition to capillary or even noninvasive saliva or urine collection has also been demonstrated for other markers such as cfDNA [124,125,126]. Concurrently, the development and establishment of measurement methods is challenging, as concentrations in different fluids are usually difficult to compare.
Another methodological aspect is that results of biomarker analyses are often available only with a delay due to the sophisticated technology involved. This provides only a retrospective view of the training session, which limits the possibilities of load management for practitioners [127]. To fulfill the requirement for rapidly available results, POC devices, mobile tools, or microarray-based screenings are necessary but may have limitations in terms of limit of detection and sensitivity. Therefore, it is of great importance to introduce innovative technologies with higher sensitivity and in miniaturized form to the market, such as the already presented protein microarrays for evaluation on a picogram level [128] for validated, reliable, load-, injury-, or overtraining-associated marker panels. Currently, various manufacturers are developing POC devices that can measure biomarkers, such as HSP, or various cytokines, such as IL-6, IL-8 or IL-10, highly efficiently in small amounts of plasma [129,130,131]. They work in the form of miniaturized protein microarray-based assays or as molecular processors and have made technological progress, particularly because of the COVID-19 pandemic [132].
4.3 Data Processing and Statistical Concerns
To obtain a comprehensive picture of the athlete, a holistic marker panel that covers various aspects of performance, muscle status, and even nutritional aspects is recommended [16], with the downside of large data sets occurring for monitoring purposes. Biomarker, performance, and questionnaire data lead to myriad ways to analyze these data. In the case of biomarkers, the literature has shown that the simple cause-effect model is not sufficient to understand the biology of an athlete [133]. The response after exercise is complex and vast, with interconnected processes and pathways.
It is generally recognized that AI, and especially machine learning (ML), can supplement basic statistical approaches when it comes to (i) data processing and visualization but also for (ii) planning and decision making [134]. ML algorithms can make predictive capability both more accessible and more accurate using existing registries and big databases together with new variables. These algorithms increase the prediction power by calibrating the equations from the different distributions of the analyzed variables, which hold the potential to change decision making dramatically and optimize individual outcomes [135, 136]. These approaches are making great progress in personalized medicine, where computer-aided analysis of biomarker data and/or imaging techniques are used to improve, for instance, a cancer patient's therapy [137]. Biomarker data have proven helpful, although decision making is still delayed due to the increasing complexity of novel biomarkers [138]. Thus, high throughput computational approaches are fundamental to create accurate prediction tools with clinical applicability and translation that hold highly impactful potential.
ML is expected to be increasingly used in professional sport settings in the future [134], with the aim that ‘artificial trainers’ will support coaches in their decision making in workload management. This is already common in the automated analysis of tools such as wearable heart rate monitors, which record daily activity, visualize data, and provide personalized training recommendations [134]. In the case of athlete monitoring, first steps to predict injuries have been taken in both individual [139] and team sports [140]. ML approaches were shown to be principally capable of predicting the internal load (RPE) in Australian Football players with an artificial-neural-network analysis (ANN) with session distance as a predictor [141]. Furthermore, ANN and least absolute shrinkage and selection operator revealed decelerations as an important variable to predict the RPE in soccer players [142]. A meta-analysis on ML approaches for injury prediction showed that research has already identified several predictor variables ranging from sleep quality and genetic variables to external load variables, such as the distance covered [143]. However, the quality of the included studies was moderate to low.
Accurate prediction of athlete responses to a planned training session and knowledge of injury risk factors allows practitioners to effectively prescribe individualized training loads [141]. There has been a focus on external load variables, which have been integrated into the ML models as predictor variables. The integration of biomarker data could further improve models, such as for injury prediction [144]. Finally, the use of multi-marker approaches and their evaluation with ML can also help to identify the suitability of biomarkers [145] for load management, as large data sets and aggregation of biomarkers with all other data can reveal previously unrecognized patterns.
Finally, the use of additional monitoring tools not only results in additional data, but also requires financial and human resources that must be carefully weighed against the associated benefits of a serially measured biomarker to ultimately decide on its use [127]. If a sports organization is able to perform, analyze, and interpret the biomarker in a nonobstructive and frequent manner, this may represent an additional benefit. If these additional resources are not available, organizations and practitioners are well advised to use easy-to-use tools such as questionnaires for the determination of the psychophysiological exercise response.
5 Conclusion and Outlook
Promising methodological approaches could soon transfer robust and valid biomarkers from the medical field to practical application in sports science. Many biomarkers have been shown to be capable of reflecting different aspects of exercise load (Table 1). Ideally, these biomarkers will be accessible in a short time using minimally invasive blood collection methods and a miniaturized POC device, although this is still in the future for some of the mentioned biomarkers. In addition, several markers have yet to prove their full potential compared with established markers such as CK or urea, for which POC devices are already available.
The collected biomarker data should be analyzed in combination with athlete characteristics, training data, and further monitoring tools using appropriate statistical approaches, to create a personalized athlete profile or at least an athlete cluster leading to individualized load management approaches. Ideally, these approaches may help to estimate the individual training responses, adverse events such as injuries, and resilience factors in athletes. Algorithms that incorporate training load recommendations can serve as a solid foundation for practitioner decision making. The ultimate goal is to combine ML with practitioner expertise to take load management to a new level [142].
References
Group F-NBW. BEST (Biomarkers, EndpointS, and other Tools) resource [Internet]. Silver Spring: 2016. Available from: https://www.ncbi.nlm.nih.gov/pubmed/27010052.
Zhang ZH, Xu X. Lactate clearance is a useful biomarker for the prediction of all-cause mortality in critically ill patients: a systematic review and meta-analysis. Crit Care Med. 2014;42(9):2118–25. https://doi.org/10.1097/Ccm.0000000000000405.
Abers MS, Delmonte OM, Ricotta EE, Fintzi J, Fink DL, de Jesus AAA, et al. An immune-based biomarker signature is associated with mortality in COVID-19 patients. Jci Insight. 2021. https://doi.org/10.1172/jci.insight.144455.
Echle A, Rindtorff NT, Brinker TJ, Luedde T, Pearson AT, Kather JN. Deep learning in cancer pathology: a new generation of clinical biomarkers. Br J Cancer. 2021;124(4):686–96. https://doi.org/10.1038/s41416-020-01122-x.
Soligard T, Schwellnus M, Alonso JM, Bahr R, Clarsen B, Dijkstra HP, et al. How much is too much? (Part 1) International Olympic Committee consensus statement on load in sport and risk of injury. Br J Sports Med. 2016;50(17):1030–41. https://doi.org/10.1136/bjsports-2016-096581.
Halson SL. Monitoring training load to understand fatigue in athletes. Sports Med. 2014;44(Suppl 2):S139–47. https://doi.org/10.1007/s40279-014-0253-z.
Akenhead R, Nassis GP. Training load and player monitoring in high-level football: current practice and perceptions. Int J Sports Physiol Perform. 2016;11(5):587–93. https://doi.org/10.1123/ijspp.2015-0331.
Baird MF, Graham SM, Baker JS, Bickerstaff GF. Creatine-kinase- and exercise-related muscle damage implications for muscle performance and recovery. J Nutr Metab. 2012;2012: 960363. https://doi.org/10.1155/2012/960363.
Brancaccio P, Maffulli N, Limongelli FM. Creatine kinase monitoring in sport medicine. Br Med Bull. 2007;81–82:209–30. https://doi.org/10.1093/bmb/ldm014.
Brewster LM, Coronel CM, Sluiter W, Clark JF, van Montfrans GA. Ethnic differences in tissue creatine kinase activity: an observational study. PLoS ONE. 2012;7(3): e32471. https://doi.org/10.1371/journal.pone.0032471.
Haller N, Helmig S, Taenny P, Petry J, Schmidt S, Simon P. Circulating, cell-free DNA as a marker for exercise load in intermittent sports. PLoS ONE. 2018;13(1): e0191915. https://doi.org/10.1371/journal.pone.0191915.
Reichel T, Bosslau TK, Palmowski J, Eder K, Ringseis R, Mooren FC, et al. Reliability and suitability of physiological exercise response and recovery markers. Sci Rep. 2020;10(1):11924. https://doi.org/10.1038/s41598-020-69280-9.
Walzik D, Joisten N, Zacher J, Zimmer P. Transferring clinically established immune inflammation markers into exercise physiology: focus on neutrophil-to-lymphocyte ratio, platelet-to-lymphocyte ratio and systemic immune-inflammation index. Eur J Appl Physiol. 2021;121(7):1803–14. https://doi.org/10.1007/s00421-021-04668-7.
Kruger K, Reichel T, Zeilinger C. Role of heat shock proteins 70/90 in exercise physiology and exercise immunology and their diagnostic potential in sports. J Appl Physiol. 2019;126(4):916–27. https://doi.org/10.1152/japplphysiol.01052.2018.
Mannion S, Mtintsilana A, Posthumus M, van der Merwe W, Hobbs H, Collins M, et al. Genes encoding proteoglycans are associated with the risk of anterior cruciate ligament ruptures. Br J Sports Med. 2014;48(22):1640–6. https://doi.org/10.1136/bjsports-2013-093201.
Lee EC, Fragala MS, Kavouras SA, Queen RM, Pryor JL, Casa DJ. Biomarkers in sports and exercise: tracking health, performance, and recovery in athletes. J Strength Cond Res. 2017;31(10):2920–37. https://doi.org/10.1519/JSC.0000000000002122.
Wiewelhove T, Raeder C, Meyer T, Kellmann M, Pfeiffer M, Ferrauti A. Markers for routine assessment of fatigue and recovery in male and female team sport athletes during high-intensity interval training. PLoS ONE. 2015;10(10): e0139801. https://doi.org/10.1371/journal.pone.0139801.
Walsh NP, Gleeson M, Shephard RJ, Gleeson M, Woods JA, Bishop NC, et al. Position statement. Part one: immune function and exercise. Exerc Immunol Rev. 2011;17:6–63.
Peake JM, Della Gatta P, Suzuki K, Nieman DC. Cytokine expression and secretion by skeletal muscle cells: regulatory mechanisms and exercise effects. Exerc Immunol Rev. 2015;21:8–25.
Hacker S, Reichel T, Hecksteden A, Weyh C, Gebhardt K, Pfeiffer M, et al. Recovery-stress response of blood-based biomarkers. Int J Environ Res Public Health. 2021. https://doi.org/10.3390/ijerph18115776.
Proschinger S, Freese J. Neuroimmunological and neuroenergetic aspects in exercise-induced fatigue. Exerc Immunol Rev. 2019;25:8–19.
Kanda K, Sugama K, Hayashida H, Sakuma J, Kawakami Y, Miura S, et al. Eccentric exercise-induced delayed-onset muscle soreness and changes in markers of muscle damage and inflammation. Exerc Immunol Rev. 2013;19:72–85.
Nikolaidis MG, Jamurtas AZ. Blood as a reactive species generator and redox status regulator during exercise. Arch Biochem Biophys. 2009;490(2):77–84. https://doi.org/10.1016/j.abb.2009.08.015.
Koh SXT, Lee JKW. S100B as a marker for brain damage and blood–brain barrier disruption following exercise. Sports Med. 2014;44(3):369–85. https://doi.org/10.1007/s40279-013-0119-9.
Stocchero CM, Oses JP, Cunha GS, Martins JB, Brum LM, Zimmer ER, et al. Serum S100B level increases after running but not cycling exercise. Appl Physiol Nutr Metab. 2014;39(3):340–4. https://doi.org/10.1139/apnm-2013-0308.
Schulte S, Schiffer T, Sperlich B, Kleinoder H, Holmberg HC. Serum concentrations of S100B are not affected by cycling to exhaustion with or without vibration. J Hum Kinet. 2011;30:59–63. https://doi.org/10.2478/v10078-011-0073-2.
Korta P, Pochec E, Mazur-Bialy A. Irisin as a multifunctional protein: implications for health and certain diseases. Medicina-Lithuania. 2019. https://doi.org/10.3390/medicina55080485.
Qiu S, Cai X, Sun Z, Schumann U, Zugel M, Steinacker JM. Chronic exercise training and circulating irisin in adults: a meta-analysis. Sports Med. 2015;45(11):1577–88. https://doi.org/10.1007/s40279-014-0293-4.
Wlodarczyk M, Kusy K, Slominska E, Krasinski Z, Zielinski J. Change in lactate, ammonia, and hypoxanthine concentrations in a 1-year training cycle in highly trained athletes: applying biomarkers as tools to assess training status. J Strength Cond Res. 2020;34(2):355–64. https://doi.org/10.1519/JSC.0000000000003375.
Zielinski J, Kusy K. Hypoxanthine: a universal metabolic indicator of training status in competitive sports. Exerc Sport Sci Rev. 2015;43(4):214–21. https://doi.org/10.1249/JES.0000000000000055.
Pla R, Pujos-Guillot E, Durand S, Brandolini-Bunlon M, Centeno D, Pyne DB, et al. Non-targeted metabolomics analyses by mass spectrometry to explore metabolic stress after six training weeks in high level swimmers. J Sports Sci. 2021;39(9):969–78. https://doi.org/10.1080/02640414.2020.1851933.
Kistner S, Doring M, Kruger R, Rist MJ, Weinert CH, Bunzel D, et al. Sex-specific relationship between the cardiorespiratory fitness and plasma metabolite patterns in healthy humans-results of the KarMeN study. Metabolites. 2021. https://doi.org/10.3390/metabo11070463.
Schranner D, Schonfelder M, Romisch-Margl W, Scherr J, Schlegel J, Zelger O, et al. Physiological extremes of the human blood metabolome: a metabolomics analysis of highly glycolytic, oxidative, and anabolic athletes. Physiol Rep. 2021;9(12): e14885. https://doi.org/10.14814/phy2.14885.
Arend WP, Malyak M, Guthridge CJ, Gabay C. Interleukin-1 receptor antagonist: role in biology. Annu Rev Immunol. 1998;16:27–55. https://doi.org/10.1146/annurev.immunol.16.1.27.
Sugama K, Suzuki K, Yoshitani K, Shiraishi K, Kometani T. Urinary excretion of cytokines versus their plasma levels after endurance exercise. Exerc Immunol Rev. 2013;19:29–48.
Muders K, Pilat C, Deuster V, Frech T, Kruger K, Pons-Kuhnemann J, et al. Effects of Traumeel (Tr14) on exercise-induced muscle damage response in healthy subjects: a double-blind RCT. Mediators Inflamm. 2016;2016:1693918. https://doi.org/10.1155/2016/1693918.
Gill SK, Teixeira A, Rama L, Prestes J, Rosado F, Hankey J, et al. Circulatory endotoxin concentration and cytokine profile in response to exertional-heat stress during a multi-stage ultra-marathon competition. Exerc Immunol Rev. 2015;21:114–28.
Ng QY, Lee KW, Byrne C, Ho TF, Lim CL. Plasma endotoxin and immune responses during a 21-km road race under a warm and humid environment. Ann Acad Med Singap. 2008;37(4):307–14.
Dinarello CA. Overview of the IL-1 family in innate inflammation and acquired immunity. Immunol Rev. 2018;281(1):8–27. https://doi.org/10.1111/imr.12621.
Schild M, Eichner G, Beiter T, Zugel M, Krumholz-Wagner I, Hudemann J, et al. Effects of acute endurance exercise on plasma protein profiles of endurance-trained and untrained individuals over time. Mediators Inflamm. 2016;2016:4851935. https://doi.org/10.1155/2016/4851935.
Duramad P, Tager IB, Holland NT. Cytokines and other immunological biomarkers in children’s environmental health studies. Toxicol Lett. 2007;172(1–2):48–59. https://doi.org/10.1016/j.toxlet.2007.05.017.
Finsterer J. Biomarkers of peripheral muscle fatigue during exercise. BMC Musculoskelet Disord. 2012;13:218. https://doi.org/10.1186/1471-2474-13-218.
Catoire M, Kersten S. The search for exercise factors in humans. FASEB J. 2015;29(5):1615–28. https://doi.org/10.1096/fj.14-263699.
JanssenDuijghuijsen LM, Keijer J, Mensink M, Lenaerts K, Ridder L, Nierkens S, et al. Adaptation of exercise-induced stress in well-trained healthy young men. Exp Physiol. 2017;102(1):86–99. https://doi.org/10.1113/EP086025.
Schnyder S, Handschin C. Skeletal muscle as an endocrine organ: PGC-1 alpha, myokines and exercise. Bone. 2015;80:115–25. https://doi.org/10.1016/j.bone.2015.02.008.
Islam H, Neudorf H, Mui AL, Little JP. Interpreting “anti-inflammatory” cytokine responses to exercise: focus on interleukin-10. J Physiol. 2021;599(23):5163–77. https://doi.org/10.1113/JP281356.
Petersen AM, Pedersen BK. The anti-inflammatory effect of exercise. J Appl Physiol (1985). 2005;98(4):1154–62. https://doi.org/10.1152/japplphysiol.00164.2004.
Cabral-Santos C, Castrillon CI, Miranda RA, Monteiro PA, Inoue DS, Campos EZ, et al. Inflammatory cytokines and BDNF response to high-intensity intermittent exercise: effect the exercise volume. Front Physiol. 2016;7:509. https://doi.org/10.3389/fphys.2016.00509.
Perera PY, Lichy JH, Waldmann TA, Perera LP. The role of interleukin-15 in inflammation and immune responses to infection: implications for its therapeutic use. Microbes Infect. 2012;14(3):247–61. https://doi.org/10.1016/j.micinf.2011.10.006.
Tamura Y, Watanabe K, Kantani T, Hayashi J, Ishida N, Kaneki M. Upregulation of circulating IL-15 by treadmill running in healthy individuals: is IL-15 an endocrine mediator of the beneficial effects of endurance exercise? Endocr J. 2011;58(3):211–5. https://doi.org/10.1507/endocrj.k10e-400.
Araujo NC, Neto AMM, Fujimori M, Bortolini MS, Justino AB, Honorio-Franca AC, et al. Immune and hormonal response to high-intensity exercise during orienteering. Int J Sports Med. 2019;40(12):768–73. https://doi.org/10.1055/a-0970-9064.
Fatouros IG. Is irisin the new player in exercise-induced adaptations or not? A 2017 update. Clin Chem Lab Med. 2018;56(4):525–48. https://doi.org/10.1515/cclm-2017-0674.
Norheim F, Langleite TM, Hjorth M, Holen T, Kielland A, Stadheim HK, et al. The effects of acute and chronic exercise on PGC-1alpha, irisin and browning of subcutaneous adipose tissue in humans. FEBS J. 2014;281(3):739–49. https://doi.org/10.1111/febs.12619.
Hubal MJ, Chen TC, Thompson PD, Clarkson PM. Inflammatory gene changes associated with the repeated-bout effect. Am J Physiol Regul Integr Comp Physiol. 2008;294(5):R1628–37. https://doi.org/10.1152/ajpregu.00853.2007.
Odobasic D, Kitching AR, Holdsworth SR. Neutrophil-mediated regulation of innate and adaptive immunity: the role of myeloperoxidase. J Immunol Res. 2016;2016:2349817. https://doi.org/10.1155/2016/2349817.
Dhama K, Latheef SK, Dadar M, Samad HA, Munjal A, Khandia R, et al. Biomarkers in stress related diseases/disorders: diagnostic, prognostic, and therapeutic values. Front Mol Biosci. 2019;6:91. https://doi.org/10.3389/fmolb.2019.00091.
Nieman DC, Groen AJ, Pugachev A, Vacca G. Detection of functional overreaching in endurance athletes using proteomics. Proteomes. 2018. https://doi.org/10.3390/proteomes6030033.
Maltseva DV, Ryabenko EA, Sizova SV, Yashin DV, Khaustova SA, Shkurnikov MY. Effect of exercise on the expression of HSPBP1, PGLYRP1, and HSPA1A genes in human leukocytes. Bull Exp Biol Med. 2012;153(6):866–8. https://doi.org/10.1007/s10517-012-1846-x.
Noble EG, Shen GX. Impact of exercise and metabolic disorders on heat shock proteins and vascular inflammation. Autoimmune Dis. 2012;2012: 836519. https://doi.org/10.1155/2012/836519.
Milne KJ, Noble EG. Exercise-induced elevation of HSP70 is intensity dependent. J Appl Physiol (1985). 2002;93(2):561–8. https://doi.org/10.1152/japplphysiol.00528.2001.
van Oosten-Hawle P. Organismal roles of Hsp90. Biomolecules. 2023. https://doi.org/10.3390/biom13020251.
Tuttle JA, Castle PC, Metcalfe AJ, Midgley AW, Taylor L, Lewis MP. Downhill running and exercise in hot environments increase leukocyte Hsp72 (HSPA1A) and Hsp90 alpha (HSPC1) gene transcripts. J Appl Physiol. 2015;118(8):996–1005. https://doi.org/10.1152/japplphysiol.00387.2014.
Harris MB, Mitchell BM, Sood SG, Webb RC, Venema RC. Increased nitric oxide synthase activity and Hsp90 association in skeletal muscle following chronic exercise. Eur J Appl Physiol. 2008;104(5):795–802. https://doi.org/10.1007/s00421-008-0833-4.
Mooren FC, Lechtermann A, Fobker M, Brandt B, Sorg C, Volker K, et al. The response of the novel pro-inflammatory molecules S100A8/A9 to exercise. Int J Sports Med. 2006;27(9):751–8. https://doi.org/10.1055/s-2005-872909.
Peake J, Peiffer JJ, Abbiss CR, Nosaka K, Okutsu M, Laursen PB, et al. Body temperature and its effect on leukocyte mobilization, cytokines and markers of neutrophil activation during and after exercise. Eur J Appl Physiol. 2008;102(4):391–401. https://doi.org/10.1007/s00421-007-0598-1.
Fagerhol MK, Nielsen HG, Vetlesen A, Sandvik K, Lyberg T. Increase in plasma calprotectin during long-distance running. Scand J Clin Lab Inv. 2005;65(3):211–20. https://doi.org/10.1080/00365510510013587.
Mortensen OH, Andersen K, Fischer C, Nielsen AR, Nielsen S, Akerstrom T, et al. Calprotectin is released from human skeletal muscle tissue during exercise. J Physiol-Lond. 2008;586(14):3551–62. https://doi.org/10.1113/jphysiol.2008.153551.
Niemela M, Kangastupa P, Niemela O, Bloigu R, Juvonen T. Acute changes in inflammatory biomarker levels in recreational runners participating in a marathon or half-marathon. Sports Med Open. 2016;2(1):21. https://doi.org/10.1186/s40798-016-0045-0.
Jansakova K, Belica I, Rajcaniova E, Rajcani J, Kyselicova K, Celusakova H, et al. The acute effect of psychosocial stress on the level of oxidative stress in children. Int J Psychophysiol. 2021;161:86–90. https://doi.org/10.1016/j.ijpsycho.2021.01.007.
Margonis K, Fatouros IG, Jamurtas AZ, Nikolaidis MG, Douroudos I, Chatzinikolaou A, et al. Oxidative stress biomarkers responses to physical overtraining: implications for diagnosis. Free Radic Biol Med. 2007;43(6):901–10. https://doi.org/10.1016/j.freeradbiomed.2007.05.022.
Kruger K, Alack K, Ringseis R, Mink L, Pfeifer E, Schinle M, et al. Apoptosis of T-cell subsets after acute high-intensity interval exercise. Med Sci Sports Exerc. 2016;48(10):2021–9. https://doi.org/10.1249/MSS.0000000000000979.
Groussard C, Rannou-Bekono F, Machefer G, Chevanne M, Vincent S, Sergent O, et al. Changes in blood lipid peroxidation markers and antioxidants after a single sprint anaerobic exercise. Eur J Appl Physiol. 2003;89(1):14–20. https://doi.org/10.1007/s00421-002-0767-1.
Mahanty A, Xi L. Utility of cardiac biomarkers in sports medicine: focusing on troponin, natriuretic peptides, and hypoxanthine. Sports Med Health Sci. 2020;2(2):65–71. https://doi.org/10.1016/j.smhs.2020.05.003.
Balsom PD, Seger JY, Sjodin B, Ekblom B. Physiological responses to maximal intensity intermittent exercise. Eur J Appl Physiol Occup Physiol. 1992;65(2):144–9. https://doi.org/10.1007/BF00705072.
Hellsten-Westing Y, Sollevi A, Sjodin B. Plasma accumulation of hypoxanthine, uric acid and creatine kinase following exhausting runs of differing durations in man. Eur J Appl Physiol Occup Physiol. 1991;62(5):380–4. https://doi.org/10.1007/BF00634977.
Sjodin B, Westing YH. Changes in plasma-concentration of hypoxanthine and uric-acid in man with short-distance running at various intensities. Int J Sports Med. 1990;11(6):493–5. https://doi.org/10.1055/s-2007-1024844.
Neuberger EWI, Sontag S, Brahmer A, Philippi KFA, Radsak MP, Wagner W, et al. Physical activity specifically evokes release of cell-free DNA from granulocytes thereby affecting liquid biopsy. Clin Epigenet. 2022;14(1):29. https://doi.org/10.1186/s13148-022-01245-3.
Fatouros IG, Destouni A, Margonis K, Jamurtas AZ, Vrettou C, Kouretas D, et al. Cell-free plasma DNA as a novel marker of aseptic inflammation severity related to exercise overtraining. Clin Chem. 2006;52(9):1820–4. https://doi.org/10.1373/clinchem.2006.070417.
Haller N, Tug S, Breitbach S, Jorgensen A, Simon P. Increases in circulating, cell-free DNA during aerobic running depend on intensity and duration. Int J Sports Physiol Perform. 2016. https://doi.org/10.1123/ijspp.2015-0540.
Haller N, Ehlert T, Schmidt S, Ochmann D, Sterzing B, Grus F, et al. Circulating, cell-free DNA for monitoring player load in professional football. Int J Sports Physiol Perform. 2018. https://doi.org/10.1123/ijspp.2018-0756.
Meinecke A, Mitzka S, Just A, Cushman S, Stojanovic SD, Xiao K, et al. Cardiac endurance training alters plasma profiles of circular RNA MBOAT2. Am J Physiol Heart Circ Physiol. 2020;319(1):H13–21. https://doi.org/10.1152/ajpheart.00067.2020.
Cox AJ, Pyne DB, Saunders PU, Callister R, Gleeson M. Cytokine responses to treadmill running in healthy and illness-prone athletes. Med Sci Sports Exerc. 2007;39(11):1918–26. https://doi.org/10.1249/mss.0b013e318149f2aa.
Williams SA, Kivimaki M, Langenberg C, Hingorani AD, Casas JP, Bouchard C, et al. Plasma protein patterns as comprehensive indicators of health. Nat Med. 2019;25(12):1851. https://doi.org/10.1038/s41591-019-0665-2.
Serrano-Ostariz E, Terreros-Blanco JL, Legaz-Arrese A, George K, Shave R, Bocos-Terraz P, et al. The impact of exercise duration and intensity on the release of cardiac biomarkers. Scand J Med Sci Sports. 2011;21(2):244–9. https://doi.org/10.1111/j.1600-0838.2009.01042.x.
Scharhag J, George K, Shave R, Urhausen A, Kindermann W. Exercise-associated increases in cardiac biomarkers. Med Sci Sports Exerc. 2008;40(8):1408–15. https://doi.org/10.1249/MSS.0b013e318172cf22.
Wheelock CE, Goss VM, Balgoma D, Nicholas B, Brandsma J, Skipp PJ, et al. Application of ’omics technologies to biomarker discovery in inflammatory lung diseases. Eur Respir J. 2013;42(3):802–25. https://doi.org/10.1183/09031936.00078812.
Contrepois K, Wu S, Moneghetti KJ, Hornburg D, Ahadi S, Tsai MS, et al. Molecular choreography of acute exercise. Cell. 2020;181(5):1112-30 e16. https://doi.org/10.1016/j.cell.2020.04.043.
Hu ZZ, Huang H, Wu CH, Jung M, Dritschilo A, Riegel AT, et al. Omics-based molecular target and biomarker identification. Methods Mol Biol. 2011;719:547–71. https://doi.org/10.1007/978-1-61779-027-0_26.
Van Gool A, Corrales F, Colovic M, Krstic D, Oliver-Martos B, Martinez-Caceres E, et al. Analytical techniques for multiplex analysis of protein biomarkers. Expert Rev Proteomics. 2020;17(4):257–73. https://doi.org/10.1080/14789450.2020.1763174.
Breitbach S, Tug S, Simon P. Circulating cell-free DNA: an up-coming molecular marker in exercise physiology. Sports Med. 2012;42(7):565–86. https://doi.org/10.2165/11631380-000000000-00000.
Aebersold R, Mann M. Mass-spectrometric exploration of proteome structure and function. Nature. 2016;537(7620):347–55. https://doi.org/10.1038/nature19949.
Ning B, Yu T, Zhang SW, Huang Z, Tian D, Lin Z, et al. A smartphone-read ultrasensitive and quantitative saliva test for COVID-19. Sci Adv. 2021. https://doi.org/10.1126/sciadv.abe3703.
Moss J, Magenheim J, Neiman D, Zemmour H, Loyfer N, Korach A, et al. Comprehensive human cell-type methylation atlas reveals origins of circulating cell-free DNA in health and disease. Nat Commun. 2018;9(1):5068. https://doi.org/10.1038/s41467-018-07466-6.
Szilagyi M, Pos O, Marton E, Buglyo G, Soltesz B, Keseru J, et al. Circulating cell-free nucleic acids: main characteristics and clinical application. Int J Mol Sci. 2020. https://doi.org/10.3390/ijms21186827.
Tug S, Helmig S, Deichmann ER, Schmeier-Jurchott A, Wagner E, Zimmermann T, et al. Exercise-induced increases in cell free DNA in human plasma originate predominantly from cells of the haematopoietic lineage. Exerc Immunol Rev. 2015;21:164–73.
Neuberger EWI, Hillen B, Mayr K, Simon P, Kramer-Albers EM, Brahmer A. Kinetics and topology of DNA associated with circulating extracellular vesicles released during exercise. Genes (Basel). 2021. https://doi.org/10.3390/genes12040522.
Helmig S, Fruhbeis C, Kramer-Albers EM, Simon P, Tug S. Release of bulk cell free DNA during physical exercise occurs independent of extracellular vesicles. Eur J Appl Physiol. 2015;115(11):2271–80. https://doi.org/10.1007/s00421-015-3207-8.
Brahmer A, Neuberger E, Esch-Heisser L, Haller N, Jorgensen MM, Baek R, et al. Platelets, endothelial cells and leukocytes contribute to the exercise-triggered release of extracellular vesicles into the circulation. J Extracell Vesicles. 2019;8(1):1615820. https://doi.org/10.1080/20013078.2019.1615820.
Appel M, Zentgraf K, Kruger K, Alack K. Effects of genetic variation on endurance performance, muscle strength, and injury susceptibility in sports: a systematic review. Front Physiol. 2021;12: 694411. https://doi.org/10.3389/fphys.2021.694411.
Pickering C, Kiely J. Understanding personalized training responses: can genetic assessment help? Open Sports Sci J. 2017;10(1):191–213. https://doi.org/10.2174/1875399X01710010191
Bouchard C. Exercise genomics—a paradigm shift is needed: a commentary. Br J Sports Med. 2015;49(23):1492–6. https://doi.org/10.1136/bjsports-2015-095294.
Jones N, Kiely J, Suraci B, Collins DJ, de Lorenzo D, Pickering C, et al. A genetic-based algorithm for personalized resistance training. Biol Sport. 2016;33(2):117–26. https://doi.org/10.5604/20831862.1198210.
Baumert P, Lake MJ, Stewart CE, Drust B, Erskine RM. Genetic variation and exercise-induced muscle damage: implications for athletic performance, injury and ageing. Eur J Appl Physiol. 2016;116(9):1595–625. https://doi.org/10.1007/s00421-016-3411-1.
Vlahovich N, Fricker PA, Brown MA, Hughes D. Ethics of genetic testing and research in sport: a position statement from the Australian Institute of Sport. Br J Sports Med. 2017;51(1):5–11. https://doi.org/10.1136/bjsports-2016-096661.
Pickering C, Kiely J. The frequency of, and attitudes towards, genetic testing amongst athletes and support staff. Perform Enhanc Health. 2021;8(4): 100184.
Vlahovich N, Hughes DC, Griffiths LR, Wang G, Pitsiladis YP, Pigozzi F, et al. Genetic testing for exercise prescription and injury prevention: AIS-Athlome consortium-FIMS joint statement. BMC Genomics. 2017;18(Suppl 8):818. https://doi.org/10.1186/s12864-017-4185-5.
Pitsiladis Y, Wang G, Wolfarth B, Scott R, Fuku N, Mikami E, et al. Genomics of elite sporting performance: what little we know and necessary advances. Brit J Sports Med. 2013;47(9):550. https://doi.org/10.1136/bjsports-2013-092400.
Morales J, Welter D, Bowler EH, Cerezo M, Harris LW, McMahon AC, et al. A standardized framework for representation of ancestry data in genomics studies, with application to the NHGRI-EBI GWAS Catalog. Genome Biol. 2018;19(1):21. https://doi.org/10.1186/s13059-018-1396-2.
Webborn N, Williams A, McNamee M, Bouchard C, Pitsiladis Y, Ahmetov I, et al. Direct-to-consumer genetic testing for predicting sports performance and talent identification: consensus statement. Br J Sports Med. 2015;49(23):1486–91. https://doi.org/10.1136/bjsports-2015-095343.
Nakayasu ES, Gritsenko M, Piehowski PD, Gao Y, Orton DJ, Schepmoes AA, et al. Tutorial: best practices and considerations for mass-spectrometry-based protein biomarker discovery and validation. Nat Protoc. 2021;16(8):3737–60. https://doi.org/10.1038/s41596-021-00566-6.
Knab AM, Nieman DC, Zingaretti LM, Groen AJ, Pugachev A. Proteomic profiling and monitoring of training distress and illness in university swimmers during a 25-week competitive season. Front Physiol. 2020;11:373. https://doi.org/10.3389/fphys.2020.00373.
Merritt EK, Nieman DC, Toone BR, Groen A, Pugachev A. Proteomic markers of non-functional overreaching during the race across America (RAAM): a case study. Front Physiol. 2019;10:1410. https://doi.org/10.3389/fphys.2019.01410.
Stroun M, Anker P, Lyautey J, Lederrey C, Maurice PA. Isolation and characterization of DNA from the plasma of cancer patients. Eur J Cancer Clin Oncol. 1987;23(6):707–12.
Banfi G, Dolci A. Free testosterone/cortisol ratio in soccer: usefulness of a categorization of values. J Sport Med Phys Fit. 2006;46(4):611–6.
Papacosta E, Nassis GP. Saliva as a tool for monitoring steroid, peptide and immune markers in sport and exercise science. J Sci Med Sport. 2011;14(5):424–34. https://doi.org/10.1016/j.jsams.2011.03.004.
Hecksteden A, Kraushaar J, Scharhag-Rosenberger F, Theisen D, Senn S, Meyer T. Individual response to exercise training—a statistical perspective. J Appl Physiol (1985). 2015;118(12):1450–9. https://doi.org/10.1152/japplphysiol.00714.2014.
Brancaccio P, Lippi G, Maffulli N. Biochemical markers of muscular damage. Clin Chem Lab Med. 2010;48(6):757–67. https://doi.org/10.1515/CCLM.2010.179.
Sperlich B, Achtzehn S, de Marees M, von Papen H, Mester J. Load management in elite German distance runners during 3-weeks of high-altitude training. Physiol Rep. 2016. https://doi.org/10.14814/phy2.12845.
Hunter DJ, Nevitt M, Losina E, Kraus V. Biomarkers for osteoarthritis: current position and steps towards further validation. Best Pract Res Cl Rh. 2014;28(1):61–71. https://doi.org/10.1016/j.berh.2014.01.007.
Carling C, Lacome M, McCall A, Dupont G, Le Gall F, Simpson B, et al. Monitoring of post-match fatigue in professional soccer: welcome to the real world. Sports Med. 2018;48(12):2695–702. https://doi.org/10.1007/s40279-018-0935-z.
Tang R, Yang H, Choi JR, Gong Y, You M, Wen T, et al. Capillary blood for point-of-care testing. Crit Rev Clin Lab Sci. 2017;54(5):294–308. https://doi.org/10.1080/10408363.2017.1343796.
Parkitny L, McAuley JH, Kelly PJ, Di Pietro F, Cameron B, Moseley GL. Multiplex cytokine concentration measurement: how much do the medium and handling matter? Mediators inflamm. 2013;2013:890706. https://doi.org/10.1155/2013/890706.
Siart B, de Oliveira FMS, Shen QJ, Bjorkesten J, Pekar T, Steinborn R, et al. Protein measurements in venous plasma, earlobe capillary plasma and in plasma stored on filter paper. Anal Biochem. 2019;566:146–50. https://doi.org/10.1016/j.ab.2018.11.016.
Shishikura Y, Tokinoya K, Aita Y, Sekine N, Sugasawa T, Yoshida Y, et al. The dynamics of cell-free DNA from urine and blood after a full marathon. bioRxiv. 2021.
Breitbach S, Sterzing B, Magallanes C, Tug S, Simon P. Direct measurement of cell-free DNA from serially collected capillary plasma during incremental exercise. J Appl Physiol (1985). 2014;117(2):119–30. https://doi.org/10.1152/japplphysiol.00002.2014.
Haller N, Tomaskovic A, Stoggl T, Simon P, Neuberger E. Feasibility of cell-free DNA measurement from the earlobe during physiological exercise testing. Diagnostics (Basel). 2022. https://doi.org/10.3390/diagnostics12061379.
Pedlar CR, Newell J, Lewis NA. Blood biomarker profiling and monitoring for high-performance physiology and nutrition: current perspectives, limitations and recommendations. Sports Med. 2019;49(Suppl 2):185–98. https://doi.org/10.1007/s40279-019-01158-x.
Sauer U. Analytical protein microarrays: advancements towards clinical applications. Sensors (Basel). 2017. https://doi.org/10.3390/s17020256.
Schax E, Walter JG, Marzhauser H, Stahl F, Scheper T, Agard DA, et al. Microarray-based screening of heat shock protein inhibitors. J Biotechnol. 2014;180:1–9. https://doi.org/10.1016/j.jbiotec.2014.03.006.
Becker T, Hitzmann B, Muffler K, Portner R, Reardon KF, Stahl F, et al. Future aspects of bioprocess monitoring. Adv Biochem Eng Biotechnol. 2007;105:249–93. https://doi.org/10.1007/10_2006_036.
Tanak AS, Muthukumar S, Krishnan S, Schully KL, Clark DV, Prasad S. Multiplexed cytokine detection using electrochemical point-of-care sensing device towards rapid sepsis endotyping. Biosens Bioelectron. 2021. https://doi.org/10.1016/j.bios.2020.112726.
Liu G, Jiang C, Lin X, Yang Y. Point-of-care detection of cytokines in cytokine storm management and beyond: significance and challenges. View (Beijing). 2021;2(4):20210003. https://doi.org/10.1002/VIW.20210003.
Anto-Ocrah M, Jones CMC, Diacovo D, Bazarian JJ. Blood-based biomarkers for the identification of sports-related concussion. Neurol Clin. 2017;35(3):473–85. https://doi.org/10.1016/j.ncl.2017.03.008.
Fister I, Ljubic K, Suganthan PN, Perc M, Fister I. Computational intelligence in sports: challenges and opportunities within a new research domain. Appl Math Comput. 2015;262:178–86. https://doi.org/10.1016/j.amc.2015.04.004.
Basile AO, Ritchie MD. Informatics and machine learning to define the phenotype. Expert Rev Mol Diagn. 2018;18(3):219–26. https://doi.org/10.1080/14737159.2018.1439380.
Helm JM, Swiergosz AM, Haeberle HS, Karnuta JM, Schaffer JL, Krebs VE, et al. Machine learning and artificial intelligence: definitions, applications, and future directions. Curr Rev Musculoskelet Med. 2020;13(1):69–76. https://doi.org/10.1007/s12178-020-09600-8.
Ngiam KY, Khor IW. Big data and machine learning algorithms for health-care delivery. Lancet Oncol. 2019;20(5):e262–73. https://doi.org/10.1016/S1470-2045(19)30149-4.
Fitzgerald J, Higgins D, Mazo Vargas C, Watson W, Mooney C, Rahman A, et al. Future of biomarker evaluation in the realm of artificial intelligence algorithms: application in improved therapeutic stratification of patients with breast and prostate cancer. J Clin Pathol. 2021;74(7):429–34. https://doi.org/10.1136/jclinpath-2020-207351.
Lovdal SS, Den Hartigh RJR, Azzopardi G. Injury prediction in competitive runners with machine learning. Int J Sports Physiol Perform. 2021;16(10):1522–31. https://doi.org/10.1123/ijspp.2020-0518.
Claudino JG, Capanema DD, de Souza TV, Serrao JC, Pereira ACM, Nassis GP. Current approaches to the use of artificial intelligence for injury risk assessment and performance prediction in team sports: a systematic review. Sports Med-Open. 2019. https://doi.org/10.1186/s40798-019-0202-3.
Bartlett JD, O’Connor F, Pitchford N, Torres-Ronda L, Robertson SJ. Relationships between internal and external training load in team-sport athletes: evidence for an individualized approach. Int J Sports Physiol Perform. 2017;12(2):230–4. https://doi.org/10.1123/ijspp.2015-0791.
Jaspers A, De Beeck TO, Brink MS, Frencken WGP, Staes F, Davis JJ, et al. Relationships between the external and internal training load in professional soccer: what can we learn from machine learning? Int J Sport Physiol. 2018;13(5):625–30. https://doi.org/10.1123/ijspp.2017-0299.
Van Eetvelde H, Mendonca LD, Ley C, Seil R, Tischer T. Machine learning methods in sport injury prediction and prevention: a systematic review. J Exp Orthop. 2021;8(1):27. https://doi.org/10.1186/s40634-021-00346-x.
Rossi A, Pappalardo L, Filetti C, Cintia P. Blood sample profile helps to injury forecasting in elite soccer players. Sport Sci Health. 2022. https://doi.org/10.1007/s11332-022-00932-1.
Swan AL, Stekel DJ, Hodgman C, Allaway D, Alqahtani MH, Mobasheri A, et al. A machine learning heuristic to identify biologically relevant and minimal biomarker panels from omics data. BMC Genomics. 2015;16(Suppl 1):S2. https://doi.org/10.1186/1471-2164-16-S1-S2.
Acknowledgements
We would like to thank Natalia Nunes for her contribution in the ‘Statistical Considerations’ section.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Open Access funding enabled and organized by Projekt DEAL. No financial assistance was used for the writing and/or preparation of the manuscript.
Conflict of interest
Nils Haller, Michael Behringer, Thomas Reichel, Patrick Wahl, Perikles Simon, Karsten Krüger, Philipp Zimmer, and Thomas Stöggl declare that they have no conflict of interest.
Author contributions
All authors, MB, PS, TR, TS, KK, PZ, PW, and NH contributed substantially to the literature search and to writing and revising the manuscript.
Availability of Data and Material
Not applicable.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Haller, N., Reichel, T., Zimmer, P. et al. Blood-Based Biomarkers for Managing Workload in Athletes: Perspectives for Research on Emerging Biomarkers. Sports Med 53, 2039–2053 (2023). https://doi.org/10.1007/s40279-023-01866-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40279-023-01866-5