Abstract
Purpose of Review
In this study, we illustrate recent findings regarding the genetics and epigenetics of alcohol use disorder (AUD). We further outline the future direction of genetic and epigenetic research in AUD.
Recent Findings
Recent genome- and epigenome-wide studies allow new insight into genetic and epigenetic variation associated with AUD. The largest EWAS of AUD so far/to date found evidence for altered glucocorticoid receptor regulation. Longitudinal studies provide insight into the dynamics of the disease. Analyses of postmortem brain tissue reveal the impact of chronic alcohol consumption on DNA methylation in the brain.
Summary
Genetic and environmental factors—mediated via epigenetic mechanisms—play an important role in AUD. Although knowledge of the biological underpinnings of AUD is still limited, ongoing research will ultimately lead to the development of biomarkers for disease classification, course of disease, and treatment response to support personalized medicine in the future.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The recent WHO global status report on alcohol and health estimates that alcohol is consumed by more than half of the population in both the USA and Europe. Harmful use of alcohol caused about 3 million deaths worldwide in 2016, which are 5.3% of all deaths [1]. In 2010, the costs of excessive drinking in the USA amounted to 249 billion US dollars, which is 46 billion dollars more than in 2006 [2]. Epidemiologic data on the DSM-5 classification of alcohol use disorder (AUD) in the years 2012 and 2013 in the USA showed that the 12-month and lifetime prevalences of AUD were 13.9% and 29.1%, respectively [3]. Compared with similar data from 2002, alcohol consumption among US adults has increased over the course of a decade [4]. Prevalence of AUD in countries of the European Union, Iceland, Norway, and Switzerland was estimated at 23 million affected people in 2010, equivalent to 3.4% among people 18–64 years of age [5]. A meta-analysis showed that mortality is significantly higher in persons with alcohol dependence, but a reduction in alcohol intake can decrease this risk [6]. The increased risk of mortality associated with AUD might be due to the genetic predisposition of individuals who develop AUD and a direct result of having AUD and/or increased alcohol intake associated with AUD. While genetic predisposition has a greater effect on mortality early in life, the direct results of having AUD become more important later in life and over the course of the disease [7].
AUD can be characterized by several criteria such as intake in spite of harmful consequences, loss of control over alcohol consumption, drug craving, tolerance development, and the manifestation of withdrawal symptoms [8]. On a neurobiological level, the process of developing AUD includes sensitization of dopamine release within the reward system, which has been shown in several animal models [9, 10], (for the sensitization concept [11]), the reorganization of striatal loops, which is supposedly associated with the habituation of drug intake [12, 13], and neuroadaptation within the limbic and stress systems [14, 15]. Studies have found that genetic variants associated with alcohol dependence are involved in alcohol metabolism as well as in dopaminergic and serotonergic transmission [16•]. While past research has mainly focused on the biological changes involved in the development of AUD, the complex role of environmental factors such as psychosocial stress is subject of more recent studies: In an adoption study, the authors found that AUD in offspring was associated with AUD and other diseases in not-lived-with biological parents as well as in stepparents, underlining that genetic and environmental risk factors act additively on the risk of developing AUD [17]. Effects of environmental factors may be mediated by epigenetic mechanisms [16•]. In the sections below, we present and discuss recent advances in research on the genetics and epigenetics of AUD. We will first give a short introduction to DNA methylation, the most extensively studied epigenetic mechanism. This introduction will be followed by an outline of epigenetic mechanisms investigated in association with alcohol use. We will then present recent findings of genetic studies in the field. Finally, we will discuss the prospects of research on genetic and epigenetic factors in AUD disease etiology.
Epigenetic Mechanisms
Environmental factors may have an effect on the development, course, and treatment of diseases via epigenetic mechanisms (e.g., [18, 19]). Epigenetic mechanisms describe a series of biochemical processes that can alter the phenotype of an individual without altering the DNA sequence. One of the best-studied epigenetic mechanisms is DNA methylation, which can alter gene function [20]. Methylation refers to the addition of a methyl group to the nucleobase cytosine at the 5′ position of the cytosine pyrimidine ring in cytosine-guanine (CG) dinucleotide sites. Genomic regions where CG dinucleotides occur at a high frequency of more than 200 base pairs in a row are referred to as CpG islands. As CpG islands usually occur near the promoter region or at the start site of genes, their methylation plays an important role in gene regulation.
Epigenetic modulation of gene transcription and consecutive long-term changes in gene function have been suggested to account for a large part of phenotypic variation. Although it was initially assumed that epigenetic modifications are “fixed” very early in development, evidence now suggests that this is not strictly the case. For example, in animals, individual responses to stress in adulthood can be altered by adversity in early life [21]. Maternal rats’ caring behavior in the first week after birth has been shown to alter the offspring’s DNA methylation at the glucocorticoid reporter gene in the hippocampus, a brain region mainly involved in learning and habituation processes. Animals with caring mothers showed higher glucocorticoid receptor levels, had reduced corticosterone, and were less anxious compared with the offspring of less-caring mothers [22]. In a translation from previous findings in animal studies [23, 24], it has been shown in humans that childhood abuse [25, 26] and parental separation [27] have an effect on the epigenetic regulation of genes involved in stress-regulation [25, 26], immune response, and cellular signaling systems relevant for neural communication and brain development and functioning [27]. In another animal study, rats exposed to alcohol during fetal development (n = 6) showed altered methylation levels of proopiomelanocortin gene (POMC) in the arcuate nucleus, which is relevant for both stress and fear management [28].
Methylation levels can also be altered by interventions such as alcohol withdrawal [29] and psychotherapy for stress prevention [19], thus giving insight into the molecular epigenetic mechanisms underlying therapy response.
In summary, epigenetic modifications such as DNA methylation regulate gene expression, ultimately leading to alterations in phenotype. Epigenetic mechanisms can explain some aspects of the role of environmental factors in disease etiology and shed light on the underlying mechanisms of therapy response.
Epigenetic Factors
In the following paragraphs, we will outline recent advances in epigenetic studies for different aspects of AUD. The findings are presented in order of the type of study: peripheral blood sampling, longitudinal sampling, and finally brain tissue sampling. The advantages and limitations of each method for studying epigenetic markers are discussed.
The most common approach to identify the epigenetic mechanisms involved in AUD is to compare methylation profiles of cases to those of healthy controls in peripheral blood samples. To date, the largest epigenome-wide association study (EWAS) of alcohol consumption was conducted in over 13,000 subjects from different cohorts and found 328 differentially methylated CpG sites in European- and 165 in African-ancestry samples. The methylation levels of 144 differentially methylated CpG sites provided substantial discrimination between current heavy drinkers and non-drinkers of European ancestry [30]. Further, in a subset of the European ancestry sample, the amount of alcohol exposure was associated with methylation patterns in the γ-aminobutyric acid-A receptor (GABA) A and B genes that are related to the development of addiction [30, 31]. Differentially methylated CpG sites were also identified in promoter regions of genes relevant to immune functioning [30]. An EWAS in 1,135 subjects analyzing the association between methylation levels and the amount of alcohol consumed found 64 CpG sites associated with alcohol intake [32]. Six of the 64 CpG sites that have been previously reported to be associated with AUD [30], liver function [33], body mass index, and lipid metabolism [34] could be replicated [32].
An EWAS of 18 twins discordant for AUD showed an association between AUD and methylation levels within the promoter region of 3′-protein-phosphatase-1G gene (PPM1G), a gene related to cell stress response [35]. This finding was validated in epigenome-wide data on binge drinking in a cohort of 499 adolescents [35]. Additionally, in this cohort, methylation levels were associated with the intensity of a reward signal from a functional magnetic resonance imaging (fMRI) scan, demonstrating the relevance of epigenetic factors for neurobiological function [35]. In another fMRI study, the authors tested 383 heavy drinkers with an fMRI alcohol reward paradigm and found an association of dopamine D2 receptor gene (DRD2) promoter methylation level with signal change in the striatum during the presentation of alcohol cues as well as the severity of AUD [36]. These findings suggest an effect of methylation patterns associated with addiction-relevant behavior on brain areas functionally relevant for the development and maintenance of addiction-relevant behavior.
Longitudinal studies can shed further light on the biological mechanisms underlying the course of illness [for AUD] and the effectiveness of interventions. In a longitudinal study of 165 female participants, different degrees of alcohol consumption over the past 6 months were found to be associated with two loci reaching epigenome-wide significance [37]. Another EWAS compared methylation patterns in 24 patients before and after a 3-week detoxification program and 23 controls [38]. Here, 59 CpG sites were differentially methylated between patients and controls before entering the treatment program and 48 CpG sites were differentially methylated comparing patients before and after treatment [38].
Massive alterations of methylation patterns were found between 99 male AUD patients and 95 controls at the time of acute withdrawal as well as after 14 days of withdrawal [39]. Especially pathways related to immune function were implicated, which is in line with previous studies linking immune function to alcohol consumption and withdrawal (e.g., [40]). Interestingly, in this study, differences between patients and controls were less pronounced after withdrawal, suggesting that methylation levels may have reverted back to normal during treatment. Those differences that remained between patients and controls after withdrawal may indicate genes and pathways implicated in alcohol addiction itself or the vulnerability to become addicted. In a sample of 69 detoxified patients followed up for 12 months, no difference in methylation levels between patients who relapsed and those who abstained was found [41]; moreover, no change over time was found.
In human studies, DNA methylation changes in peripheral blood are most commonly investigated. Samples from other tissues, which are likely to be altered by long-lasting alcohol consumption (e.g., brain, liver) are rare and difficult to obtain. There is evidence for some concordance between peripheral blood and brain tissue concerning their DNA methylation patterns, with correlation coefficients estimated to range between 0.33 and 0.40 [42•]. Still, the analysis of DNA methylation patterns in human brain tissue is of particular interest, as the brain is likely involved in the development and maintenance of AUD.
One of the first EWAS in human postmortem brain tissue investigated DNA methylation differences in the prefrontal cortex (PFC; Brodmann Area 9) of 23 subjects with AUD (7 female) and 23 age- and sex-matched controls [43]. Analyses were conducted separately for male and female subjects. A total of 1,812 differentially methylated CpG sites were found to be associated with AUD case/control status in males. No observed differences in methylation between cases and controls were present in female subjects, indicating a possible effect of sex. Furthermore, 22 co-methylation modules (i.e., groups of co-methylated CpGs, clustered by pairwise correlations) were associated with AUD in males. The largest module was enriched for the biological processes: cell projection organization, cell projection morphogenesis, and neuron development. Of note, genes identified to be associated with substance use phenotypes in genome-wide association studies (GWAS) were overrepresented in the two largest modules. A more recent investigation of DNA methylation levels in tissue from Brodmann Area 9 found two differentially methylated regions, one annotated to the gene that codes for the discs large-associated protein 2 (DLGAP2) [44]. DLGAP2 encodes membrane-associated guanylate kinases relevant for the organization of synapses involved in neuronal cell signaling. In another study, over 400 differentially methylated CpG sites were identified in precuneus brain samples of 49 subjects (11 female) and 47 controls (12 female) [45].
In a recent EWAS, DNA methylation was investigated in the PFC (Brodmann Area 10) of25 pairs of AUD cases and controls [46]. While the authors identified a large number of nominally significant differentially methylated sites, they focused on genes related to stress adaptation, including the nuclear receptor subfamily 3 group C member 1 gene (NR3C1). They found chronic alcohol consumption to be associated with increased methylation levels of NR3C1 and decreased expression of the related protein as well as alterations in the expression of several other stress-responsive genes in the PFC of AUD cases. These findings underline the notion that environmental factors including stress exposure are one of the key factors explaining the development and continuation of behavior related to alcohol use disorders [35, 46, 47].
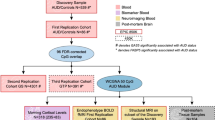
Moreover, in a cross-tissue and cross-phenotype study, the authors demonstrated an association between DNA methylation levels in the promoter region of the proprotein convertase subtilisin/kexin 9 gene (PCSK9), a gene involved in cholesterol metabolism, and AUD [48]. Findings from the three discovery set analyses in human brain tissue were replicated in blood and liver tissue as well as in animal models [48]. In the largest EWAS of AUD so far, evidence for an altered glucocorticoid regulation in AUD emerged [49••]. The EWAS was conducted in six independent cohorts on subjects with AUD, and results were replicated across multiple tissues. AUD was associated with DNA methylation in genes associated with immune response and glucocorticoid signaling. A marker that has been consistently associated was the growth arrest–specific five gene (GAS5), which has been implicated in the regulation of glucocorticoid receptors [49••].
However, there are several challenges concerning postmortem studies of brain tissue: due to cell type heterogeneity, comparability of data retrieved from a given tissue might not be given. Also, in the context of AUD, there are far more male postmortem brain samples available (e.g., New South Wales Tissue Resource Centre (NSW TRC)). Therefore, it is difficult to identify underlying biological mechanisms that differ considerably between males and females. Future studies on the single-cell level will help to analyze methylation patterns in more detail. Because of the rarity of this type of tissue and the high number of samples needed to reach adequate statistical power, we conclude that available datasets need to be combined with in silico brain banks and meta-analytical approaches.
In summary, there is accumulating evidence for the association of epigenetic markers with AUD. However, a causal relationship has yet to be established.
Genetic Factors
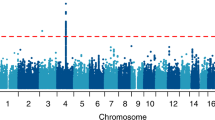
Apart from epigenetic factors, genetic factors contribute to AUD. The heritability of AUD is typically estimated to be around 40% [50]. Genetic factors also play a role in the course and severity of addiction [51]. Large GWAS and meta-analyses have identified genes and pathways implicated in AUD [52, 53, 54•, 55]. The most recent GWAS identified 29 risk variants associated with problematic alcohol use [56••]. These studies could also corroborate known candidate genes, e.g., in neurotransmitter systems such as the dopaminergic, cholinergic, serotonergic, noradrenergic, glutamatergic, and GABAergic systems. These insights facilitate a better understanding of the molecular mechanisms underlying AUD.
Polygenic methods can be used to analyze the aggregated genetic effects of all markers associated with an outcome. A polygenic risk score (PRS) is the weighted sum of all known risk alleles for a specific outcome calculated using the information of large GWAS [57]. It is a single value that reflects an individual’s overall genetic risk for, or inherited susceptibility to, a disease. Because polygenic risk scores are calculated based on the summary statistics of GWAS, the features of the GWAS (e.g., sample size, ancestry, explained SNP-heritability) also influence the PRS. PRS are associated with disease severity and the course of illness for a variety of psychiatric disorders (e.g., [58]). With respect to AUD, past studies have shown that aggregated genetic risk measured by PRS is associated with alcohol consumption as well as alcohol dependence [59, 60]. Using polygenic methods to analyze genetic correlations, relations were detected between problematic drinking and other substance use and psychiatric traits, as well as between alcohol-related traits measured longitudinally by using Alcohol Use Disorder Identification Test-Consumption (AUDIT-C) scores [61] and AUD.
Furthermore, there are efforts to investigate these associations in a more refined way by examining the influence of a specific genetic predisposition, e.g., scores only including genes from dopaminergic, serotonergic, and glutamatergic pathways and/or PRS for related domains such as cognition, stress reactivity, and sensation seeking on disease mechanisms involved in substance use and addiction [62]. An area of particular interest that has not yet been fully exploited is the association of these PRS with dimensions of observable behavior and neurobiological measures as measured with research domain criteria (RDoCs, e.g., impulsivity, negative affect, cognitive control) [63].
In other areas of medicine such as cardiology, PRS have been used to successfully identify high-risk subgroups of patients who benefit more from specific therapies [64]. The reliable use of PRS requires sufficient GWAS summary statistics, and larger sample sizes are necessary. It is common that large GWAS only include European samples [56••]. While there have been substantial efforts to perform cross-ancestral GWAS [52, 54•, 61], especially in the field of substance use disorders, prediction of the genetic risk for AUD remains less accurate for non-European populations.
Prospects of Genetic and Epigenetic Factors in AUD
Recent advances in genetic and epigenetic research of AUD provide insight into possible mechanistic underpinnings and potential biomarkers for disease classification, course of disease, and treatment response. To date, however, knowledge of the molecular basis, i.e., the genomic and epigenomic processes underlying addiction, remains fragmentary. Large-scale and/or longitudinal studies with an intensive characterization of participants from multiple populations are necessary to further identify the underlying biological processes that contribute to the risk of alcohol addiction and recovery. Previous efforts to investigate the genetic markers of AUD for clinical use have been largely limited to studies of single markers in pharmacological applications. So far, there is a lack of evidence regarding genetic predictors of medication efficacy (e.g., [65]). Also, the prediction accuracy of aggregated genetic markers, such as PRS, is still limited. Larger GWAS are necessary to enhance the power of the discovery sets.
In general, developing and establishing robust biomarkers and algorithms to predict diagnoses and treatment response necessitates the comprehensive integration of intensive genetic, epigenetic, transcriptomic, metabolomic, microbiomic, and phenotypic longitudinal data [66]. As sample sizes increase, machine learning approaches can be used as a promising data-driven way to analyze large-scale multi-omics data. New developments such as massively parallelizing applied algorithms in combination with the development of more powerful computing hardware make it possible to process such data within a reasonable timeframe and will eventually lead to applicability for individualized interventions and prevention.
Moreover, clinicians and patients must be well informed about this extremely complex subject involving the importance of genetics and environmental factors. The participation of patient representatives, clinicians, geneticists, and bioinformaticians is essential in this process.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
World Health Organization, Management of Substance Abuse Team, World Health Organization (2018) Global status report on alcohol and health 2018.
Sacks JJ, Gonzales KR, Bouchery EE, Tomedi LE, Brewer RD. 2010 national and state costs of excessive alcohol consumption. Am J Prev Med. 2015;49:e73–9.
Grant BF, Goldstein RB, Saha TD, Chou SP, Jung J, Zhang H, et al. Epidemiology of DSM-5 alcohol use disorder: results from the National Epidemiologic Survey on Alcohol and Related Conditions III. JAMA Psychiatry. 2015;72:757–66.
Dawson DA, Goldstein RB, Saha TD, Grant BF. Changes in alcohol consumption: United States, 2001-2002 to 2012-2013. Drug Alcohol Depend. 2015;148:56–61.
Rehm J, Anderson P, Barry J, Dimitrov P, Elekes Z, Feijão F, et al. Prevalence of and potential influencing factors for alcohol dependence in Europe. Eur Addict Res. 2015;21:6–18.
Laramée P, Leonard S, Buchanan-Hughes A, Warnakula S, Daeppen J-B, Rehm J. Risk of all-cause mortality in alcohol-dependent individuals: a systematic literature review and meta-analysis. EBioMedicine. 2015;2:1394–404.
Kendler KS, Ohlsson H, Sundquist J, Sundquist K. Alcohol use disorder and mortality across the lifespan: a longitudinal cohort and co-relative analysis. JAMA Psychiatry. 2016;73:575–81.
American Psychiatric Association (ed) (2013) Diagnostic and statistical manual of mental disorders (DSM-5®). American Psychiatric Pub.
Volkow ND, Morales M. The brain on drugs: from reward to addiction. Cell. 2015;162:712–25.
Heinz A, Higley JD, Gorey JG, Saunders RC, Jones DW, Hommer D, et al. In vivo association between alcohol intoxication, aggression, and serotonin transporter availability in nonhuman primates. Am J Psychiatry. 1998;155:1023–8.
Robinson TE, Berridge KC. The neural basis of drug craving: an incentive-sensitization theory of addiction. Brain Res Rev. 1993;18:247–91.
Vollstädt-Klein S, Loeber S, Richter A, Kirsch M, Bach P, von der Goltz C, et al. Validating incentive salience with functional magnetic resonance imaging: association between mesolimbic cue reactivity and attentional bias in alcohol-dependent patients: neural cue reactivity triggers attention. Addict Biol. 2012;17:807–16.
Everitt BJ, Robbins TW. Drug addiction: updating actions to habits to compulsions ten years on. Annu Rev Psychol. 2016;67:23–50.
Heinz AJ, Beck A, Meyer-Lindenberg A, Sterzer P, Heinz A. Cognitive and neurobiological mechanisms of alcohol-related aggression. Nat Rev Neurosci. 2011;12:400–13.
Volkow ND, Morales M. 2. Cell. 2015;162:712–25.
• Hamilton PJ, Nestler EJ. Epigenetics and addiction. Curr Opin Neurobiol. 2019;59:128–36 Recent review on challenges in the field of research on epigenetics of alcohol use disorder. The authors also discuss the emerging field of locus-specific neuro-epigenetic editing.
Kendler KS, Ji J, Edwards AC, Ohlsson H, Sundquist J, Sundquist K. An extended Swedish national adoption study of alcohol use disorder. JAMA Psychiatry. 2015;72:211–8.
Boschen KE, Keller SM, Roth TL, Klintsova AY. Epigenetic mechanisms in alcohol- and adversity-induced developmental origins of neurobehavioral functioning. Neurotoxicol Teratol. 2018;66:63–79.
Stoffel M, Aguilar-Raab C, Rahn S, Steinhilber B, Witt SH, Alexander N, et al. Effects of mindfulness-based stress prevention on serotonin transporter gene methylation. Psychother Psychosom. 2019;88:317–9.
Moore LD, Le T, Fan G. DNA methylation and its basic function. Neuropsychopharmacol Off Publ Am Coll Neuropsychopharmacol. 2013;38:23–38.
Meaney MJ, Szyf M. Environmental programming of stress responses through DNA methylation: life at the interface between a dynamic environment and a fixed genome. Dialogues Clin Neurosci. 2005;7:103–23.
Weaver ICG, Cervoni N, Champagne FA, D’Alessio AC, Sharma S, Seckl JR, et al. Epigenetic programming by maternal behavior. Nat Neurosci. 2004;7:847–54.
Heinz A, Higley JD, Gorey JG, Saunders RC, Jones DW, Hommer D, et al. In vivo association between alcohol intoxication, aggression, and serotonin transporter availability in nonhuman primates. Am J Psychiatry. 1998;155:1023–8.
Shannon C, Schwandt ML, Champoux M, Shoaf SE, Suomi SJ, Linnoila M, et al. Maternal absence and stability of individual differences in CSF 5-HIAA concentrations in rhesus monkey infants. Am J Psychiatry. 2005;162:1658–64.
McGowan PO, Sasaki A, D’Alessio AC, Dymov S, Labonté B, Szyf M, et al. Epigenetic regulation of the glucocorticoid receptor in human brain associates with childhood abuse. Nat Neurosci. 2009;12:342–8.
Labonte B, Yerko V, Gross J, Mechawar N, Meaney MJ, Szyf M, et al. Differential glucocorticoid receptor exon 1(B), 1(C), and 1(H) expression and methylation in suicide completers with a history of childhood abuse. Biol Psychiatry. 2012;72:41–8.
Naumova OY, Lee M, Koposov R, Szyf M, Dozier M, Grigorenko EL. Differential patterns of whole-genome DNA methylation in institutionalized children and children raised by their biological parents. Dev Psychopathol. 2012;24:143–55.
Gangisetty O, Bekdash R, Maglakelidze G, Sarkar DK. Fetal alcohol exposure alters Proopiomelanocortin gene expression and hypothalamic-pituitary-adrenal axis function via increasing MeCP2 expression in the hypothalamus. PLoS One. 2014;9:e113228.
Ovenden ES, McGregor NW, Emsley RA, Warnich L. DNA methylation and antipsychotic treatment mechanisms in schizophrenia: Progress and future directions. Prog Neuro-Psychopharmacol Biol Psychiatry. 2018;81:38–49.
Liu C, Marioni RE, Hedman \AA K, et al (2018) A DNA methylation biomarker of alcohol consumption. Mol Psychiatry 23:422–433.
Krystal JH, Staley J, Mason G, Petrakis IL, Kaufman J, Harris RA, et al. γ-Aminobutyric acid type a receptors and alcoholism: intoxication, dependence, vulnerability, and treatment. Arch Gen Psychiatry. 2006;63:957–68.
Xu K, Montalvo-Ortiz JL, Zhang X, Southwick SM, Krystal JH, Pietrzak RH, et al. Epigenome-wide DNA methylation association analysis identified novel loci in peripheral cells for alcohol consumption among European American male veterans. Alcohol Clin Exp Res. 2019;43:2111–21.
Nano J, Ghanbari M, Wang W, et al. Epigenome-wide association study identifies methylation sites associated with liver enzymes and hepatic Steatosis. Gastroenterology. 2017;153:1096–1106.e2.
Wahl S, Drong A, Lehne B, Loh M, Scott WR, Kunze S, et al. Epigenome-wide association study of body mass index, and the adverse outcomes of adiposity. Nature. 2017;541:81–6.
Ruggeri B, Nymberg C, Vuoksimaa E, Lourdusamy A, Wong CP, Carvalho FM, et al. Association of protein phosphatase PPM1G with alcohol use disorder and brain activity during behavioral control in a genome-wide methylation analysis. Am J Psychiatry. 2015;172:543–52.
Bidwell LC, Karoly HC, Thayer RE, Claus ED, Bryan AD, Weiland BJ, et al. DRD2 promoter methylation and measures of alcohol reward: functional activation of reward circuits and clinical severity. Addict Biol. 2018;24:539–48. https://doi.org/10.1111/adb.12614.
Philibert RA, Plume JM, Gibbons FX, Brody GH, Beach SRH. The impact of recent alcohol use on genome wide DNA methylation signatures. Front Genet. 2012;3. https://doi.org/10.3389/fgene.2012.00054.
Brückmann C, Islam SA, MacIsaac JL, et al. DNA methylation signatures of chronic alcohol dependence in purified CD3+ T-cells of patients undergoing alcohol treatment. Sci Rep. 2017;7:6605.
Witt SH, Frank J, Frischknecht U, Treutlein J, Streit F, Foo JC, et al. Acute alcohol withdrawal and recovery in men lead to profound changes in DNA methylation profiles: a longitudinal clinical study. Addiction. 2020. https://doi.org/10.1111/add.15020.
Coleman LG, Crews FT. Innate immune signaling and alcohol use disorders. Handb Exp Pharmacol. 2018;248:369–96.
Friedel E, Walter H, Veer IM, Zimmermann US, Heinz A, Frieling H, et al. Impact of long-term alcohol consumption and relapse on genome-wide DNA methylation changes in alcohol-dependent subjects: a longitudinal study. Alcohol Clin Exp Res. 2020;44:1356–65. https://doi.org/10.1111/acer.14354.
• Edgar RD, Jones MJ, Meaney MJ, Turecki G, Kobor MS. BECon: a tool for interpreting DNA methylation findings from blood in the context of brain. Transl Psychiatry. 2017;7:1–18 This is a detailed description for a useful online tool for evaluating the relationship between peripheral methylation count and its association with methylation count in other tissues (e.g., central nervous system) derived from tissue banks including post mortem studies.
Wang F, Xu H, Zhao H, Gelernter J, Zhang H. DNA co-methylation modules in postmortem prefrontal cortex tissues of European Australians with alcohol use disorders. Sci Rep. 2016;6:19430.
IMAGEN Consortium, Meng W, Sjöholm LK, et al. Genotype-dependent epigenetic regulation of DLGAP2 in alcohol use and dependence. Mol Psychiatry. 2019. https://doi.org/10.1038/s41380-019-0588-9.
Hagerty SL, YorkWilliams SL, Bidwell LC, Weiland BJ, Sabbineni A, Blaine SK, et al. DRD2 methylation is associated with executive control network connectivity and severity of alcohol problems among a sample of polysubstance users. Addict Biol. 2018;25. https://doi.org/10.1111/adb.12684.
Gatta E, Grayson DR, Auta J, Saudagar V, Dong E, Chen Y, et al. Genome-wide methylation in alcohol use disorder subjects: implications for an epigenetic regulation of the cortico-limbic glucocorticoid receptors (NR3C1). Mol Psychiatry. 2019:1–13.
Lupien SJ, McEwen BS, Gunnar MR, Heim C. Effects of stress throughout the lifespan on the brain, behaviour and cognition. Nat Rev Neurosci. 2009;10:434–45.
Lohoff FW, Sorcher JL, Rosen AD, Mauro KL, Fanelli RR, Momenan R, et al. Methylomic profiling and replication implicates deregulation of PCSK9 in alcohol use disorder. Mol Psychiatry. 2018;23:1900–10.
•• Lohoff FW, Roy A, Jung J, et al. Epigenome-wide association study and multi-tissue replication of individuals with alcohol use disorder: evidence for abnormal glucocorticoid signaling pathway gene regulation. Mol Psychiatry. 2020:1–14 This is the largest EWAS of AUD to date. The authors of this study used epigenome-wide data from six independent cohorts and replicated these results across multiple tissues. A top marker was the growth-arrest specific five gene (GAS5), which is implicated in the regulation of glucocorticoid receptors.
Goldman D, Oroszi G, Ducci F. The genetics of addictions: uncovering the genes. Nat Rev Genet. 2005;6:521–32.
Liu I-C, Blacker DL, Xu R, Fitzmaurice G, Lyons MJ, Tsuang MT. Genetic and environmental contributions to the development of alcohol dependence in male twins. Arch Gen Psychiatry. 2004;61:897–903.
Gelernter J, Kranzler HR, Sherva R, Almasy L, Koesterer R, Smith AH, et al. Genome-wide association study of alcohol dependence: significant findings in African- and European-Americans including novel risk loci. Mol Psychiatry. 2014;19:41–9.
Zuo L, Tan Y, Zhang X, Wang X, Krystal J, Tabakoff B, et al. A new genome-wide association meta-analysis of alcohol dependence. Alcohol Clin Exp Res. 2015;39:1388–95.
• Walters RK, Polimanti R, Johnson EC, et al. Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat Neurosci. 2018;21:1656–69 Meta-analysis of 14,904 individuals with AD and 37,944 controls from 28 case-control and family-based studies to characterize the relationship between AD and other psychiatric and behavioral outcomes.
Liu M, Jiang Y, Wedow R, et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet. 2019;51:237–44.
•• Zhou H, Sealock JM, Sanchez-Roige S, et al. Genome-wide meta-analysis of problematic alcohol use in 435,563 individuals yields insights into biology and relationships with other traits. Nat Neurosci. 2020;23:809–18 The largest GWAS on AUD and Problematic Alcohol Use (PAU) so far (n = 435,563). Data from the latest GWAS of Alcohol Dependence from the Psychiatric Genomics Consortium Substance Use Disorder Working Group, of AUD from the Million Veteran Program (MVP) and of an AUDIT-P problematic drinking proxy from the UK Biobank was meta-analyzed. 29 risk loci associated with PAU were found, 19 of them were novel findings.
Wray NR, Goddard ME, Visscher PM. Prediction of individual genetic risk to disease from genome-wide association studies. Genome Res. 2007;17:1520–8.
Frank J, Lang M, Witt SH, Strohmaier J, Rujescu D, Cichon S, et al. Identification of increased genetic risk scores for schizophrenia in treatment-resistant patients. Mol Psychiatry. 2015;20:150–1.
Taylor M, Simpkin AJ, Haycock PC, Dudbridge F, Zuccolo L. Exploration of a polygenic risk score for alcohol consumption: a longitudinal analysis from the ALSPAC cohort. PLoS One. 2016;11:e0167360.
Savage JE, Salvatore JE, Aliev F, Edwards AC, Hickman M, Kendler KS, et al. Polygenic risk score prediction of alcohol dependence symptoms across population-based and clinically ascertained samples. Alcohol Clin Exp Res. 2018;42:520–30.
Kranzler HR, Zhou H, Kember RL, Vickers Smith R, Justice AC, Damrauer S, et al. Genome-wide association study of alcohol consumption and use disorder in 274,424 individuals from multiple populations. Nat Commun. 2019;10:1499.
Groenman AP, Greven CU, van Donkelaar MMJ, Schellekens A, van Hulzen KJE, Rommelse N, et al. Dopamine and serotonin genetic risk scores predicting substance and nicotine use in attention-deficit/hyperactivity disorder. Addict Biol. 2016;21:915–23.
Garbusow M, Nebe S, Sommer C, Kuitunen-Paul S, Sebold M, Schad DJ, et al. Pavlovian-to-instrumental transfer and alcohol consumption in Young male social drinkers: behavioral, neural and polygenic correlates. J Clin Med. 2019;8:1188.
Natarajan P, Young R, Stitziel NO, Padmanabhan S, Baber U, Mehran R, et al. Polygenic risk score identifies subgroup with higher burden of atherosclerosis and greater relative benefit from statin therapy in the primary prevention setting. Circulation. 2017;135:2091–101.
Hartwell EE, Kranzler HR. Pharmacogenetics of alcohol use disorder treatments: an update. Expert Opin Drug Metab Toxicol. 2019;15:553–64.
Heinz A, Kiefer F, Smolka MN, et al. Addiction research consortium: losing and regaining control over drug intake (ReCoDe)-from trajectories to mechanisms and interventions. Addict Biol. 2020;25:e12866.
Funding
Open Access funding provided by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the Topical Collection on Alcohol
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wedemeyer, F., Kaminski, J.A., Zillich, L. et al. Prospects of Genetics and Epigenetics of Alcohol Use Disorder. Curr Addict Rep 7, 446–452 (2020). https://doi.org/10.1007/s40429-020-00331-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40429-020-00331-x