Abstract
Objectives
In this systematic review we aimed at assessing how artificial intelligence (AI), including machine learning (ML) techniques have been deployed to predict, diagnose, and treat chronic kidney disease (CKD). We systematically reviewed the available evidence on these innovative techniques to improve CKD diagnosis and patient management.
Methods
We included English language studies retrieved from PubMed. The review is therefore to be classified as a “rapid review”, since it includes one database only, and has language restrictions; the novelty and importance of the issue make missing relevant papers unlikely. We extracted 16 variables, including: main aim, studied population, data source, sample size, problem type (regression, classification), predictors used, and performance metrics. We followed the Preferred Reporting Items for Systematic Reviews (PRISMA) approach; all main steps were done in duplicate.
Results
From a total of 648 studies initially retrieved, 68 articles met the inclusion criteria.
Models, as reported by authors, performed well, but the reported metrics were not homogeneous across articles and therefore direct comparison was not feasible. The most common aim was prediction of prognosis, followed by diagnosis of CKD. Algorithm generalizability, and testing on diverse populations was rarely taken into account. Furthermore, the clinical evaluation and validation of the models/algorithms was perused; only a fraction of the included studies, 6 out of 68, were performed in a clinical context.
Conclusions
Machine learning is a promising tool for the prediction of risk, diagnosis, and therapy management for CKD patients. Nonetheless, future work is needed to address the interpretability, generalizability, and fairness of the models to ensure the safe application of such technologies in routine clinical practice.
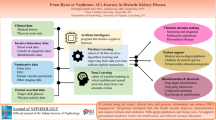
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Chronic Kidney Disease (CKD) is a state of progressive loss of kidney function ultimately resulting in the need for renal replacement therapy (dialysis or transplantation) [1]. It is defined as the presence of kidney damage or an estimated glomerular filtration rate less than 60 ml/min per 1.73 m2, persisting for 3 months or more [2]. CKD prevalence is growing worldwide, along with demographic and epidemiological transitions [3]. The implications of this disease are enormous for our society in terms of quality of life and the overall sustainability of national health systems. Worldwide, CKD accounted for 2,968,600 (1%) disability-adjusted life-years and 2,546,700 (1% to 3%) life-years lost in 2012 [4]. Therefore, it is of the utmost importance to assess how to promptly and adequately diagnose and treat patients with CKD.
The causes of CKD vary globally. The most common primary diseases causing CKD and ultimately kidney failure are diabetes mellitus, hypertension, and primary glomerulonephritis, representing 70–90% of the total primary causes [1, 2, 4]. Although these three causes are at the top of the CKD etiology charts, other features are involved in CKD pathophysiology (e.g., pollution, infections and autoimmune diseases) [5,6,7,8,9]. Similarly, there are numerous factors that play a role in CKD progression, namely non-modifiable risk factors (e.g., age, gender, ethnicity) and modifiable ones (e.g., systolic and diastolic blood pressure, proteinuria) [1, 2, 4,5,6,7,8,9].
Given how dauntingly vast the number of factors that can play a significant role in the etiology and progression of CKD is, it can be difficult to correctly assess the individual risk of CKD and its progression. Naturally, as with any complex problem, humans seek simplification, and therefore the question shifts to what to take into account when assessing CKD risk. Thanks to new methodological techniques, we now have the ability to improve our diagnostic and predictive capabilities.
Artificial Intelligence (AI) is the capacity of human-built machines to manifest complex decision-making or data analysis in a similar or augmented fashion in comparison to human intelligence [10]. Machine Learning (ML) is the collection of algorithms that empower models to learn from data, and therefore to undertake complex tasks through complex calculations [11,12,13,14,15]. In recent years AI and ML have offered enticing solutions to clinical problems, such as how to perform a diagnosis from sparse and seemingly contrasting data, or how to predict a prognosis [16]. Given the enormous potential of ML, and its capacity to learn from data, researchers have tried to apply its capacities to resolve complex problems, such as predicting CKD diagnosis and prognosis, and managing its treatment.
In this complex scenario, we aimed to systematically review the published studies that applied machine learning in the diagnosis and prediction, prognosis, and treatment of CKD patients. In doing so, the primary objective is to describe how ML models and variables have been used to predict, diagnose and treat CKD, as well as what results have been achieved in this field.
Methods
Search strategy and selection criteria
We conducted a systematic literature review, following the Preferred Reporting Items for Systematic Reviews (PRISMA) approach [17], including studies that applied ML algorithms to CKD forecasting, diagnosis, prognosis, and treatment. This systematic review’s outcomes of interest are machine learning models, features used, performances and uses regarding diagnosis, prognosis and treatment of CKD. The review itself and its protocol were not registered.
The initial search was implemented on October 20, 2021. The search query consisted of terms considered pertinent by the authors.
We searched for publications on PubMed using the following search string: “((artificial intelligence[Title/Abstract]) OR (machine learning[Title/Abstract]) OR (computational*[Title/Abstract]) OR (deep learning[Title/Abstract])) AND ((ckd) OR (chronic kidney disease) OR (chronic kidney injury) OR (chronic kidney) OR (chronic renal) OR (end stage renal) OR (end stage kidney) OR (ESKD) OR (ESRD) OR (CKJ) OR (CKI) OR (((renal) OR (kidney)) AND (failure)))”.
We included articles for review if they were in vivo studies (human-based), which applied AI & ML techniques in order to assess the diagnosis, prognosis, or therapy of CKD patients and reported original data. We did not limit our inclusion criteria to any specific study design, nor to any outcome of interest, as our main goal was to be as inclusive as possible, and we wanted to capture all available evidence from any study design and any outcome of interest.
We excluded studies that were not in English, those focusing on animals, reviews, systematic reviews, opinions, editorials, and case reports. We decided to exclude in vitro studies (conducted on cellular substrates) and studies focusing on animals, in order to summarize the current evidence on the application of ML models on humans.
Data extraction
Data were extracted by two independent reviewers (AC and FS). Disagreement on extracted data was discussed with an independent arbiter (DGol).
The following data were extracted from each included article (main text and/or supplementary material): author(s) name, date of publication, first author affiliation (country and region), main study objective, objective category (risk, diagnosis, prognosis, and treatment), prognosis category, study population, data source, sample size, problem type (regression, classification), machine learning algorithms examined in the study, predictor categories, number of predictors used, predictor list, performance metrics, final conclusions, use in clinical context and the 5 most important model features. When more than one model was considered in the study, the one the authors deemed best was extracted. Performance metrics always refer to the models’ performance on test sets.
Quality and risk assessment
Evaluation of the included studies was performed using both PROBAST [18] and the Guidelines for developing and reporting machine learning predictive models in biomedical research developed by Luo and colleagues [19].
Results
Included studies
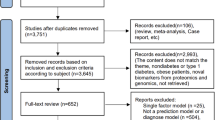
Of the 648 articles retrieved from PubMed, 421 were ruled out after title screening, and 140 were excluded after abstract screening; a total of 87 articles were selected for full-text screening (Fig. 1). Of these 87 studies, 68 were included in the final set of articles (Table 1) [20,21,22,23,24,25,26,27,28,29,30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87].
Most of the included articles (n = 51) were published from 2019 to 2021. Among the 68 articles selected for data extraction, the majority were published by authors from organizations based in Asia (n = 33; 48.5%). The remaining articles were published by authors from Europe (n = 17; 25%), North America (n = 12; 17.6%), Africa (n = 5; 7.35%) and South America (n = 1; 1.47%). The analyzed studies were classified as observational.
Study aim
A total of 28 studies focused on the use of ML algorithms in disease prognosis analysis, 21 investigated the use of ML techniques on diagnosis (4 evaluated both), 12 evaluated the risk of developing the disease, and 3 investigated the use of ML in CKD treatment. Among the articles focusing on prognosis, the majority studied the application of ML in evaluating CKD progression (n = 13) and mortality (n = 8).
Study populations and sample size
The most commonly investigated study population consisted of patients with CKD and healthy subjects (n = 26; 38.2%), followed by patients with CKD only (n = 16; 23.5%) and patients with CKD treated with hemodialysis (n = 12; 17.6%). The sample size investigated in the selected articles varied from a minimum of 30 individuals to a maximum of 550,000 (median = 776; IQR 400–12,020).
Data sources
The majority of the included articles analyzed data obtained from single-hospital registries (n = 33; 48.5%), datasets provided by universities (n = 15; 22.1%), and datasets collected in multi-center studies (n = 12, 17.6%). Five studies analyzed health insurance data (7.35%) and 3 studies used data provided by national health services (4.41%).
The most commonly used data were various combinations of demographic data along with individual clinical characteristics and laboratory data (n = 60; 82.24%), followed by data obtained by medical imaging technologies (n = 5; 7.35%) and genomic data (n = 3; 4.41%).
Models
The number of models tested and reported in each article varied from a minimum of 1 model to a maximum of 10 (mean = 3). The most frequently tested model class was tree algorithms (n = 58, 33.53%), such as random forest (n = 27, 15.61%), decision trees (n = 10, 5.78%) and extreme gradient boosting (n = 9, 5.20). Subsequently, neural networks (NNs) were often inspected (n = 44, 16.18%), especially the multilayer perceptron (MLP) (n = 28, 16.18%). Another popular choice of machine learning model class was Support Vector Machines (n = 25, 14.45%) and logistic regression (n = 18, 10.45%) with various regularizations. Another popular method that we did not classify into a larger model class was the non-parametric k-Nearest Neighbors algorithm (n = 8, 2.31%). The complete list of models can be found in Table 2.
All the articles implemented supervised learning algorithms, 57 (83.8%) of them addressed classification tasks and 11 (16.2%) regression tasks.
The majority of the included articles (n = 52) specified the total number of features used to train the models. These models used a highly variable number of features, ranging from 4 to 6624 (median = 24; IQR = 17—46). Of the 68 included studies, 55 specified the variables used in the models (n = 130). The most frequently used features are reported in Fig. 2.
Performance metrics
The most common performance metrics were accuracy (n = 30, 17.05%) and the area under the receiver operating characteristic curve (often also referred to as ROC-AUC, AUROC, AUC, or C-statistic) (n = 30, 17.05%). Subsequently, other classification metrics, such as sensitivity (n = 29, 16.48%), specificity (n = 24, 13.64%), precision (n = 16, 9.09%), and F1-score (n = 14, 7.95%) were often used to compare the machine learning models. Note that all the aforementioned metrics, except ROC AUC, were used for classification and required establishing a risk threshold as a decision boundary. ROC AUC conversely did not require setting a decision threshold as it was calculated by iterating over all the decision thresholds. In terms of regression, the most used metrics for comparison were mean absolute error (n = 6, 3.41%) and root mean squared error (n = 5, 2.84%). The full list of the metrics and how often they occurred can be found in Table 3.
Best performing models, and their performances
In the included articles, neural networks were the models that commonly performed best (n = 28, 41.18%) compared to the median performance of other models, such as MLP (n = 18, 26.47%) and convolutional neural networks (n = 7, 24.53%). Tree-based algorithms performed best (n = 24, 35.29%); these algorithms included Random Forest (n = 16, 23.53%) and Extreme Gradient Boosting (n = 5, 7.35%). The results for Support Vector Machines (n = 5, 7.35%) were also noteworthy. A complete list of the best performing models in the selected papers can be found in Table 4.
In terms of performance, we compared the metrics of prediction models, diagnostic models and risk prediction models separately. Of the 25 (36.76%) machine learning models for diagnosis, 19 papers reported accuracy. Three models reported the highest accuracy of 1.00 while the lowest reported accuracy is 0.80 (mean = 0.95, median = 0.98). Sensitivity was reported 15 times, with a maximum of 1.00, a minimum of 0.56, a mean of 0.95 and a median of 0.99. In addition, specificity was reported in 13 cases (max = 1.00, min = 0.79, mean = 0.96, median = 0.99). The ROC-AUC was reported in 6 papers (max = 0.99, min = 0.91, mean = 0.941, median = 0.94).
For the prediction models (n = 32, 47.06%), 15 papers reported the ROC-AUC with a maximum of 0.96 and a minimum of 0.69 (mean = 0.82, median = 0.82). Ten papers reported accuracy, ranging from 0.54 to 0.99, with a mean of 0.85 and a median of 0.87. Sensitivity was reported 8 times, ranging from 0.54 to 0.93 (mean = 0.765, median = 0.76), and specificity was reported 5 times (max = 0.99, min = 0.78, mean = 0.917, median = 0.96).
Next, the risk prediction models (n = 12, 17.65%) showed ROC-AUC 9 times (max = 0.96, min = 0.76, mean = 0.864, median = 0.86) and accuracy 4 times (max = 0.99, min = 0.82, mean = 0.901, median = 0.91).
Finally, 3 (4.41%) papers focused on therapy, one of which reported an accuracy of 0.95, while the other two focused on outcome differences (p-values).
Most common variables and most important ones
The total number of variables used in the included studies was 813. The five most common ones were: Blood Pressure (n = 62, 7.63%), Age (n = 45, 5.54%), Hemoglobin (n = 37, 4.55%), Creatinine (serum) (n = 31, 3.81%) and Sex (n = 31, 3.81%).
Nonetheless, to better capture how variables were used in the selected papers, we classified the variables into 4 subsets (CKD Prognosis, CKD Diagnosis, Risk of Developing CKD, CKD Treatment) based on the primary aim the authors stated their model would have attempted to achieve.
Regarding CKD Prognosis, 342 variables were used out of 813 total (42%). The most common ones were: Blood Pressure (n = 24, 7%), Age (n = 19, 5,56%), Cholesterol (serum) (n = 18, 5.26%), Sex (n = 14, 4%) and Hemoglobin (blood) (n = 13, 3.8%), with the most important variables being: Age, Hemoglobin and Proteinuria.
Concerning CKD Diagnosis, 311 variables were used out of 813 total (38.25%). The most common ones were: Blood Pressure (n = 22, 7%), Hemoglobin (blood) (n = 19, 6.1%), Pus Cell General—used to indicate the number of dead white cells in urine—(n = 18, 5.79%), Age (n = 14, 4.50%) and Glucose (serum) (n = 14, 4.50%). The most important variables in this case were Albumin, Creatinine, and Hemoglobin.
With regard to Risk of Developing CKD, 137 variables were used out of 813 total (16.85%). The most common ones were: Blood Pressure (n = 12, 8.75%), Age (n = 9, 6.57%), Sex (n = 7, 5.11%), History of Cardiovascular Disease (n = 6, 4.38%) and estimated Glomerular Filtration Rate (eGFR) (n = 6, 4.38%). The most important variables were Age, GFR and Blood Pressure.
Finally, regarding CKD Treatment, 23 variables were used out of 813 total (2.83%). The most common ones were: Blood Iron (n = 5, 21.74%), Hemoglobin (n = 3, 13%), Drugs Used (n = 2, 8.70%), MCV (n = 2, 8.70%) and White Blood Cells (blood) (n = 2, 8.70%). Regarding this aim, no weights were listed in the examined articles.
The complete spreadsheet with all variables and percentages can be found in Supplemental Material, together with the most important variables, divided per aim.
Fairness
Other than using PROBAST to assess risk of bias, we also assessed fairness based on how the authors explicitly used variables. In some studies, variables were not fully listed, and in such cases, if the variable (sex, or race/ethnicity) was not indexed, we considered the feature as not included in the general model.
Out of 68 studies, 43 included gender in the model and 12 included race/ethnicity. When Non-Hispanic Whites were part of the assessed cohort, they were the majority group, ranging from 87 to 31%. Ten out of 68 studies addressed both gender and race/ethnicity, and included these variables in the model.
Race/ethnicity was included in 4 out of 12 studies predicting risk, in 5 out of 28 studies predicting prognosis, and in 3 out of 21 studies classifying diagnosis. It was never included in models investigating prognosis and diagnosis combined, and therapeutics.
Clinical Deployment
Regarding Diagnosis, just one model was actually deployed in a clinical environment [60]. The authors applied a lasso regression with metabolites as features, achieving an accuracy of 99%; the authors used data from a real clinical context, and therefore they deployed and evaluated their model performance on a clinical context, nevertheless, they did not validate their model. Regarding Prognosis, just 3 studies were conducted in a clinical setting [49, 50, 62]. Komaru et al. [49] predicted 1-year mortality following the start of hemodialysis through hierarchical clustering and achieved an AUC of 0.8; the authors used data from a clinical prospective study to deploy and evaluate their model. Furthermore, they validated the used clusters. Kanda et al. [50] applied a support vector machine model onto a real population in an observational study to deploy and evaluate their model. The authors achieved an accuracy of 89% through 13 variables; unfortunately, they did not disclose the weights of the variables nor did they validate the model, and therefore we do not know which variables were the most important. Akbilgic et al. [62] used a model based on a Random Forest algorithm, and achieved an AUC of 0.69; the most important features were eGFR, Spontaneous Bacterial Peritonitis, Age, Diastolic Blood Pressure and BUN. The authors used data from a real clinical context to deploy and evaluate their model; furthermore, they validated their results and model internally. Regarding Risk of developing CKD, one study’s model was used in a clinical context [42]. The authors used a NN, achieving an AUC of 0.89, using retinal images as features from a clinical context to deploy, evaluate and validate their model. Finally, regarding CKD Treatment, one study’s model was used in a clinical environment [26]; they presented their results through differences in achieved values by their algorithms, and the best performance was achieved by a NN. They evaluated the model with clinical data, but did not validate it.
Quality assessment
According to the PROBAST assessment tool [18], most of the included articles showed an overall low risk of bias (n = 48; 67.6%), and 65 (91.5%) of the included articles showed low applicability. Moreover, only 8.5% of the included studies scored less than 70% in the reporting guidelines for machine learning predictive models in biomedical research developed by Luo and colleagues [19]. The complete quality assessment can be found in Supplemental Material.
Discussion
This systematic review describes how machine learning has been used for CKD. Six overarching themes were found, each of which underlines the need for further consideration by the scientific community.
First, despite the ever-growing number of studies focusing on the topic, a staggeringly low amount are being considered for actual clinical implementation. In this review, just 5 out of 68 articles tried to deploy their model in a real clinical setting. This might indicate either that the technology is not ready yet, or, considering 4 of these 5 articles were published in the last 3 years, that the technology is just starting to creep into real clinical settings. Recent evidence suggests that it is paramount to test newly developed algorithms in clinical settings before trying to deploy them [88]. Despite promising laboratory results, clinical translation is not always guaranteed. As an example, when studying the feasibility of providing an automated electronic alarm for acute kidney injury in different clinical settings, substantial heterogeneity in the findings among hospitals was described, with the worrying result of a significantly increased risk of death for some hospitals [89].
Second, as expected, the most important features were profoundly related to the main aim the authors were pursuing. In this regard, there were no surprises in the studied topics as the most important features were related to conditions known to lead to CKD diagnosis, worsening of prognosis and risk of developing CKD (e.g., age, comorbidities, systolic and diastolic blood pressure and eGFR values).
Third, a lack of consistency in reporting results was found. Most of the studies chose to report accuracy, but this was not the norm. Furthermore, while accuracy provides information on model performance, it fails to consider class imbalance and data representation. This is extremely important as accuracy in highly unbalanced datasets can be very high by always predicting the same binary outcome because of a flawed model. For instance, considering a low prevalence disease, if the algorithm is flawed for it always predicts a negative event, the accuracy will be high, but the veracity of the model will not [90]. As a result, AUCs and ROCs better measure the model precision without requiring the definition of a risk threshold. Twenty-nine authors chose to express their results including AUCs and ROCs: the minimum value was 0.69 and the maximum was 0.99 (mean: 0.83, median: 0.84). These results best express how precise the algorithms were and confirm the overall high performance of the assessed models.
Fourth, a common conundrum regarding feature selection and output was found in studies assessing CKD diagnosis. The definition of CKD requires certain variables to be present in order to make a diagnosis, thus including those variables in the model might be considered mandatory. Nonetheless, including those variables forces the model to streamline its decision process to a simple match in altered values, effectively transforming a complex machine learning model into a linear decision flow-chart, the performance of which will always be stellar.
This phenomenon is especially clear in four of the studies this systematic review assessed [36, 39, 46, 47]. In these studies, the same database [91] is used, and accuracy, sensitivity, specificity, and ROC-AUC are never below 98%. We believe researchers should carefully assess the variables used in their machine learning models to make sure that no data leakage is present between features and results.
Fifth, model bias and fairness were almost never considered. This is critical, as both biased and unfair models will not achieve the same results in different demographics, and their societal impact could exasperate disparities in certain populations. These issues need to be further explored before any model can be implemented at point of care.
Finally, among the included studies, only 6 evaluated their models in a clinical setting [26, 42, 49, 50, 60, 62], and only 3 were validated [42, 49, 62]. These studies showed promising results and did not report any unintended consequences after evaluation and/or validation. Notwithstanding the robust results described by the authors, as discussed before, recent evidence suggests that it is paramount to test newly developed algorithms in clinical settings to avoid adverse or unintended consequences [88, 89]. Taking into account the pinnacle of importance of validating ones’ results in real clinical contexts and not just “in lab”, in reading their results, their generalizability has to be questioned, especially since no multi-center validations were described among the validated models.
This systematic review presents a few limitations: first, only one database (PubMed) was used to collect studies of interest. It should be noted that systematic reviews are usually exhorted to use at least two databases as stated by the PRISMA statement. Nonetheless, as PubMed has grown to be one of the most used search engines for medical sciences this limitation should be self-amending. Secondly, this systematic review assessed only papers written in English since English is the most widely adopted and commonly used language for the publication of medical papers.
In addition to these limitations, due to this review’s design, all in vitro studies (on cellular substrates) were excluded. Consequently, the evidence presented in this review is not to be interpreted as definitive for all things concerning CKD, since in vitro studies (on cellular substrates), the insight of which is critical in understanding pathogenetic as well as therapeutic mechanisms, were not assessed.
Lastly, the majority of included studies did not evaluate the integration of ML models in daily clinical practice, therefore the results and discussion have to be considered largely from an academic standpoint. Despite these limitations, we feel this review advances the knowledge on the current state of data-driven algorithms to advance CKD diagnosis, prognosis and treatment.
Despite the potential benefits, the application of machine learning for CKD diagnosis, prognosis, and treatment presents several issues, namely fairness, model and result interpretability [90], and the lack of validated models. Result interpretability concerns reflect the inability to explain which aspects of the dataset used in the training phase led to a predicted result in a particular case [92, 93]. Therefore, as the trend in machine learning techniques moves from traditional algorithms (e.g., lasso regressions, support vector machine, and decision trees), to more complex ones (e.g., ensemble algorithms and deep learning), the interpretability concerns become more pronounced [90]. Notably, researchers highlighted the need for explainability and for models that could have a significant impact on patients' health [94, 95]. These models should be reported using best practice reporting guidelines such as the Transparent Reporting of a Multivariate Prediction Model for Individual Prognosis or Diagnosis (TRIPOD) [94] or MINimum Information for Medical AI Reporting (MINIMAR) [97]. Transparent and accurate reports are also fundamental in advancing multi-center validations of the applied models, which in turn is an essential step to ensure that only safe and sound models are applied on a large scale.
Most of the studies failed to report on the ethical issues revolving around their model development; the impact on the patient's well-being can also be affected by algorithmic bias [98, 99] and this can be worse in certain underrepresented populations. This concern is closely related to the generalizability of the developed model [100,101,102]. Specifically, retrospective data that are usually used during the training phase often have significant biases towards subgroups of individuals that have been defined by factors such as age, gender, educational level, socioeconomic status, and location [98]. The issues of fairness and bias in algorithms should be evaluated by investigating the models’ performance within population subgroups.
This systematic review underlines the potential benefits and pitfalls of ML in the diagnosis, prognosis, and management of CKD. We found that most of the studies included in this systematic review reported that ML offers invaluable help to clinicians allowing them to make informed decisions and provide better care to their patients; nonetheless most of those articles were not actually piloted in real life settings, and therefore, notwithstanding the excellent model performance results reported by authors, the technology might not be ready for mass real-time adoption or implementation.
Although future work is needed to address the viability, interpretability, generalizability, and fairness issues, to allow a safer translation of these models for use in daily clinical practice, the implementation of these techniques could further enhance the effective management of hospital resources in a timely and efficient manner by potentially identifying patients at high risk for adverse events and the need for additional resources.
We hope the summarized evidence from this article will facilitate implementation of ML approaches in the clinical practice.
Data availability Statement
Data that support the findings of this study are available upon reasonable request from the corresponding author, AC.
Change history
06 March 2023
A Correction to this paper has been published: https://doi.org/10.1007/s40620-023-01609-9
References
Webster AC, Nagler EV, Morton RL et al (2017) Chronic kidney disease. Lancet Lond Engl 389(10075):1238–1252. https://doi.org/10.1016/S0140-6736(16)32064-5
Chen TK, Knicely DH, Grams ME (2019) Chronic kidney disease diagnosis and management. JAMA 322(13):1294–1304. https://doi.org/10.1001/jama.2019.14745
Global, regional, and national burden of chronic kidney disease, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet Lond Engl. 2020;395(10225):709–733. doi:https://doi.org/10.1016/S0140-6736(20)30045-3
Vaidya SR, Aeddula NR. Chronic Renal Failure. In: StatPearls. StatPearls Publishing; 2022. Accessed July 28, 2022. http://www.ncbi.nlm.nih.gov/books/NBK535404/
Romagnani P, Remuzzi G, Glassock R et al (2017) Chronic kidney disease. Nat Rev Dis Primer 3:17088. https://doi.org/10.1038/nrdp.2017.88
Thomas R, Kanso A, Sedor JR (2008) Chronic kidney disease and its complications. Prim Care 35(2):329–vii. https://doi.org/10.1016/j.pop.2008.01.008
Fraser SD, Blakeman T (2016) Chronic kidney disease: identification and management in primary care. Pragmatic Obs Res 7:21–32. https://doi.org/10.2147/POR.S97310
Chronic Kidney Disease: Overview. Institute for Quality and Efficiency in Health Care (IQWiG); 2018. Accessed July 28, 2022. https://www.ncbi.nlm.nih.gov/books/NBK492977/
Kazancioğlu R (2013) Risk factors for chronic kidney disease: an update. Kidney Int Suppl 3(4):368–371. https://doi.org/10.1038/kisup.2013.79
Topol EJ (2019) High-performance medicine: the convergence of human and artificial intelligence. Nat Med 25(1):44–56. https://doi.org/10.1038/s41591-018-0300-7
Nichols JA, Herbert Chan HW, Baker MAB (2018) Machine learning: applications of artificial intelligence to imaging and diagnosis. Biophys Rev 11(1):111–118. https://doi.org/10.1007/s12551-018-0449-9
Sidey-Gibbons JAM, Sidey-Gibbons CJ (2019) Machine learning in medicine: a practical introduction. BMC Med Res Methodol 19:64. https://doi.org/10.1186/s12874-019-0681-4
Peterson DJ, Ostberg NP, Blayney DW et al (2021) Machine learning applied to electronic health records: identification of chemotherapy patients at high risk for preventable emergency department visits and hospital admissions. JCO Clin Cancer Inform 5:1106–1126. https://doi.org/10.1200/CCI.21.00116
Lenain R, Seneviratne MG, Bozkurt S et al (2019) Machine learning approaches for extracting stage from pathology reports in prostate cancer. Stud Health Technol Inform 264:1522–1523. https://doi.org/10.3233/SHTI190515
Cahan EM, Hernandez-Boussard T, Thadaney-Israni S et al (2019) Putting the data before the algorithm in big data addressing personalized healthcare. NPJ Digit Med 2:78. https://doi.org/10.1038/s41746-019-0157-2
Rajpurkar P, Chen E, Banerjee O et al (2022) AI in health and medicine. Nat Med 28(1):31–38. https://doi.org/10.1038/s41591-021-01614-0
Page MJ, McKenzie JE, Bossuyt PM et al (2021) The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ 372:71. https://doi.org/10.1136/bmj.n71
Wolff RF, Moons KGM, Riley RD et al (2019) PROBAST: a tool to assess the risk of bias and applicability of prediction model studies. Ann Intern Med 170(1):51–58. https://doi.org/10.7326/M18-1376
Luo W, Phung D, Tran T et al (2016) Guidelines for developing and reporting machine learning predictive models in biomedical research: a multidisciplinary view. J Med Internet Res 18(12):e323. https://doi.org/10.2196/jmir.5870
Goldstein BA, Pomann GM, Winkelmayer WC et al (2017) A comparison of risk prediction methods using repeated observations: an application to electronic health records for hemodialysis. Stat Med 36(17):2750–2763. https://doi.org/10.1002/sim.7308
Sabanayagam C, Xu D, Ting DSW et al (2020) A deep learning algorithm to detect chronic kidney disease from retinal photographs in community-based populations. Lancet Digit Health 2(6):e295–e302. https://doi.org/10.1016/S2589-7500(20)30063-7
Rodriguez M, Salmeron MD, Martin-Malo A et al (2016) A new data analysis system to quantify associations between biochemical parameters of chronic kidney disease-mineral bone disease. PLoS ONE 11(1):e0146801. https://doi.org/10.1371/journal.pone.0146801
Barbieri C, Mari F, Stopper A et al (2015) A new machine learning approach for predicting the response to anemia treatment in a large cohort of End Stage Renal Disease patients undergoing dialysis. Comput Biol Med 61:56–61. https://doi.org/10.1016/j.compbiomed.2015.03.019
Kumar A, Sinha N, Bhardwaj A (2020) A novel fitness function in genetic programming for medical data classification. J Biomed Inform 112:103623. https://doi.org/10.1016/j.jbi.2020.103623
Peng H, Zhu H, Ieong CWA et al (2021) A two-stage neural network prediction of chronic kidney disease. IET Syst Biol 15(5):163–171. https://doi.org/10.1049/syb2.12031
Barbieri C, Molina M, Ponce P et al (2016) An international observational study suggests that artificial intelligence for clinical decision support optimizes anemia management in hemodialysis patients. Kidney Int 90(2):422–429. https://doi.org/10.1016/j.kint.2016.03.036
Kanda E, Epureanu BI, Adachi T et al (2020) Application of explainable ensemble artificial intelligence model to categorization of hemodialysis-patient and treatment using nationwide-real-world data in Japan. PLoS ONE 15(5):e0233491. https://doi.org/10.1371/journal.pone.0233491
Yu H, Samuels DC, Zhao YY, Guo Y (2019) Architectures and accuracy of artificial neural network for disease classification from omics data. BMC Genom 20(1):167. https://doi.org/10.1186/s12864-019-5546-z
Lin SY, Hsieh MH, Lin CL et al (2019) Artificial intelligence prediction model for the cost and mortality of renal replacement therapy in aged and super-aged populations in Taiwan. J Clin Med. https://doi.org/10.3390/jcm8070995
Ohara T, Ikeda H, Sugitani Y et al (2021) Artificial intelligence supported anemia control system (AISACS) to prevent anemia in maintenance hemodialysis patients. Int J Med Sci 18(8):1831–1839. https://doi.org/10.7150/ijms.53298
Akl AI, Sobh MA, Enab YM et al (2001) Artificial intelligence: a new approach for prescription and monitoring of hemodialysis therapy. Am J Kidney Dis Off J Natl Kidney Found 38(6):1277–1283. https://doi.org/10.1053/ajkd.2001.29225
Kolachalama VB, Singh P, Lin CQ et al (2018) Association of pathological fibrosis with renal survival using deep neural networks. Kidney Int Rep 3(2):464–475. https://doi.org/10.1016/j.ekir.2017.11.002
Daniel AJ, Buchanan CE, Allcock T et al (2021) Automated renal segmentation in healthy and chronic kidney disease subjects using a convolutional neural network. Magn Reson Med 86(2):1125–1136. https://doi.org/10.1002/mrm.28768
Kuo CC, Chang CM, Liu KT et al (2019) Automation of the kidney function prediction and classification through ultrasound-based kidney imaging using deep learning. NPJ Digit Med 2:29. https://doi.org/10.1038/s41746-019-0104-2
Parab J, Sequeira M, Lanjewar M et al (2021) Backpropagation neural network-based machine learning model for prediction of blood urea and glucose in CKD patients. IEEE J Transl Eng Health Med 9:4900608. https://doi.org/10.1109/JTEHM.2021.3079714
Chen Z, Zhang X, Zhang Z (2016) Clinical risk assessment of patients with chronic kidney disease by using clinical data and multivariate models. Int Urol Nephrol 48(12):2069–2075. https://doi.org/10.1007/s11255-016-1346-4
Rashed-Al-Mahfuz M, Haque A, Azad A et al (2021) Clinically applicable machine learning approaches to identify attributes of chronic kidney disease (CKD) for use in low-cost diagnostic screening. IEEE J Transl Eng Health Med 9:4900511. https://doi.org/10.1109/JTEHM.2021.3073629
Roth JA, Radevski G, Marzolini C et al (2021) Cohort-derived machine learning models for individual prediction of chronic kidney disease in people living with human immunodeficiency virus: a prospective multicenter cohort study. J Infect Dis 224(7):1198–1208. https://doi.org/10.1093/infdis/jiaa236
Huang ML, Chou YC (2019) Combining a gravitational search algorithm, particle swarm optimization, and fuzzy rules to improve the classification performance of a feed-forward neural network. Comput Methods Programs Biomed. 180:105016. https://doi.org/10.1016/j.cmpb.2019.105016
Jeong B, Cho H, Kim J et al (2020) Comparison between statistical models and machine learning methods on classification for highly imbalanced multiclass kidney data. Diagn Basel Switz. https://doi.org/10.3390/diagnostics10060415
Xin G, Zhou G, Zhang W et al (2020) Construction and validation of predictive model to identify critical genes associated with advanced kidney disease. Int J Genomics 2020:7524057. https://doi.org/10.1155/2020/7524057
Zhang K, Liu X, Xu J et al (2021) Deep-learning models for the detection and incidence prediction of chronic kidney disease and type 2 diabetes from retinal fundus images. Nat Biomed Eng 5(6):533–545. https://doi.org/10.1038/s41551-021-00745-6
Schena FP, Anelli VW, Trotta J et al (2021) Development and testing of an artificial intelligence tool for predicting end-stage kidney disease in patients with immunoglobulin A nephropathy. Kidney Int 99(5):1179–1188. https://doi.org/10.1016/j.kint.2020.07.046
Galloway CD, Valys AV, Shreibati JB et al (2019) Development and validation of a deep-learning model to screen for hyperkalemia from the electrocardiogram. JAMA Cardiol 4(5):428–436. https://doi.org/10.1001/jamacardio.2019.0640
Yuan Q, Zhang H, Xie Y et al (2020) Development of prognostic model for patients at CKD stage 3a and 3b in South Central China using computational intelligence. Clin Exp Nephrol 24(10):865–875. https://doi.org/10.1007/s10157-020-01909-5
Polat H, Danaei Mehr H, Cetin A (2017) Diagnosis of chronic kidney disease based on support vector machine by feature selection methods. J Med Syst 41(4):55. https://doi.org/10.1007/s10916-017-0703-x
Senan EM, Al-Adhaileh MH, Alsaade FW et al (2021) Diagnosis of chronic kidney disease using effective classification algorithms and recursive feature elimination techniques. J Healthc Eng 2021:1004767. https://doi.org/10.1155/2021/1004767
Pellicer-Valero OJ, Cattinelli I, Neri L et al (2020) Enhanced prediction of hemoglobin concentration in a very large cohort of hemodialysis patients by means of deep recurrent neural networks. Artif Intell Med. 107:101898. https://doi.org/10.1016/j.artmed.2020.101898
Komaru Y, Yoshida T, Hamasaki Y et al (2020) Hierarchical clustering analysis for predicting 1-year mortality after starting hemodialysis. Kidney Int Rep 5(8):1188–1195. https://doi.org/10.1016/j.ekir.2020.05.007
Kanda E, Kanno Y, Katsukawa F (2019) Identifying progressive CKD from healthy population using Bayesian network and artificial intelligence: a worksite-based cohort study. Sci Rep 9(1):5082. https://doi.org/10.1038/s41598-019-41663-7
Singh A, Nadkarni G, Gottesman O et al (2015) Incorporating temporal EHR data in predictive models for risk stratification of renal function deterioration. J Biomed Inform 53:220–228. https://doi.org/10.1016/j.jbi.2014.11.005
Inaguma D, Kitagawa A, Yanagiya R et al (2020) Increasing tendency of urine protein is a risk factor for rapid eGFR decline in patients with CKD: a machine learning-based prediction model by using a big database. PLoS ONE 15(9):e0239262. https://doi.org/10.1371/journal.pone.0239262
Elhoseny M, Shankar K, Uthayakumar J (2019) Intelligent diagnostic prediction and classification system for chronic kidney disease. Sci Rep 9(1):9583. https://doi.org/10.1038/s41598-019-46074-2
Nusinovici S, Tham YC, Chak Yan MY et al (2020) Logistic regression was as good as machine learning for predicting major chronic diseases. J Clin Epidemiol 122:56–69. https://doi.org/10.1016/j.jclinepi.2020.03.002
Song X, Waitman LR, Yu AS et al (2020) Longitudinal risk prediction of chronic kidney disease in diabetic patients using a temporal-enhanced gradient boosting machine: retrospective cohort study. JMIR Med Inform 8(1):e15510. https://doi.org/10.2196/15510
Tang Y, Zhang W, Zhu M et al (2018) Lupus nephritis pathology prediction with clinical indices. Sci Rep 8(1):10231. https://doi.org/10.1038/s41598-018-28611-7
Segal Z, Kalifa D, Radinsky K et al (2020) Machine learning algorithm for early detection of end-stage renal disease. BMC Nephrol 21(1):518. https://doi.org/10.1186/s12882-020-02093-0
Forné C, Cambray S, Bermudez-Lopez M et al (2020) Machine learning analysis of serum biomarkers for cardiovascular risk assessment in chronic kidney disease. Clin Kidney J 13(4):631–639. https://doi.org/10.1093/ckj/sfz094
Huang J, Huth C, Covic M et al (2020) Machine learning approaches reveal metabolic signatures of incident chronic kidney disease in individuals with prediabetes and type 2 diabetes. Diabetes 69(12):2756–2765. https://doi.org/10.2337/db20-0586
Guo Y, Yu H, Chen D et al (2019) Machine learning distilled metabolite biomarkers for early stage renal injury. Metabolomics Off J Metabolomic Soc 16(1):4. https://doi.org/10.1007/s11306-019-1624-0
Krishnamurthy S, Ks K, Dovgan E et al (2021) Machine learning prediction models for chronic kidney disease using national health insurance claim data in Taiwan. Healthc Basel Switz. https://doi.org/10.3390/healthcare9050546
Akbilgic O, Obi Y, Potukuchi PK et al (2019) Machine learning to identify dialysis patients at high death risk. Kidney Int Rep 4(9):1219–1229. https://doi.org/10.1016/j.ekir.2019.06.009
Belur Nagaraj S, Pena MJ, Ju W et al (2020) Machine-learning-based early prediction of end-stage renal disease in patients with diabetic kidney disease using clinical trials data. Diabetes Obes Metab 22(12):2479–2486. https://doi.org/10.1111/dom.14178
Vitsios D, Petrovski S (2020) Mantis-ml: disease-agnostic gene prioritization from high-throughput genomic screens by stochastic semi-supervised learning. Am J Hum Genet 106(5):659–678. https://doi.org/10.1016/j.ajhg.2020.03.012
Shang N, Khan A, Polubriaginof F et al (2021) Medical records-based chronic kidney disease phenotype for clinical care and “big data” observational and genetic studies. NPJ Digit Med 4(1):70. https://doi.org/10.1038/s41746-021-00428-1
Luo L, Small D, Stewart WF et al (2013) Methods for estimating kidney disease stage transition probabilities using electronic medical records. EGEMS Wash DC 1(3):1040. https://doi.org/10.13063/2327-9214.1040
Almansour NA, Syed HF, Khayat NR et al (2019) Neural network and support vector machine for the prediction of chronic kidney disease: a comparative study. Comput Biol Med 109:101–111. https://doi.org/10.1016/j.compbiomed.2019.04.017
Chen CA, Li YC, Lin YF et al (2007) Neuro-fuzzy technology as a predictor of parathyroid hormone level in hemodialysis patients. Tohoku J Exp Med 211(1):81–87. https://doi.org/10.1620/tjem.211.81
Escandell-Montero P, Chermisi M, Martínez-Martínez JM et al (2014) Optimization of anemia treatment in hemodialysis patients via reinforcement learning. Artif Intell Med 62(1):47–60. https://doi.org/10.1016/j.artmed.2014.07.004
Weber C, Röschke L, Modersohn L et al (2020) Optimized identification of advanced chronic kidney disease and absence of kidney disease by combining different electronic health data resources and by applying machine learning strategies. J Clin Med. https://doi.org/10.3390/jcm9092955
Garcia-Montemayor V, Martin-Malo A, Barbieri C et al (2021) Predicting mortality in hemodialysis patients using machine learning analysis. Clin Kidney J 14(5):1388–1395. https://doi.org/10.1093/ckj/sfaa126
Norouzi J, Yadollahpour A, Mirbagheri SA et al (2016) Predicting renal failure progression in chronic kidney disease using integrated intelligent fuzzy expert system. Comput Math Methods Med 2016:6080814. https://doi.org/10.1155/2016/6080814
Kusiak A, Dixon B, Shah S (2005) Predicting survival time for kidney dialysis patients: a data mining approach. Comput Biol Med 35(4):311–327. https://doi.org/10.1016/j.compbiomed.2004.02.004
Jeong YS, Kim J, Kim D et al (2021) Prediction of postoperative complications for patients of end stage renal disease. Sensors. https://doi.org/10.3390/s21020544
Martínez-Martínez JM, Escandell-Montero P, Barbieri C et al (2014) Prediction of the hemoglobin level in hemodialysis patients using machine learning techniques. Comput Methods Programs Biomed 117(2):208–217. https://doi.org/10.1016/j.cmpb.2014.07.001
Noh J, Yoo KD, Bae W et al (2020) Prediction of the mortality risk in peritoneal dialysis patients using machine learning models: a nation-wide prospective cohort in Korea. Sci Rep 10(1):7470. https://doi.org/10.1038/s41598-020-64184-0
Glazyrin YE, Veprintsev DV, Ler IA et al (2020) Proteomics-based machine learning approach as an alternative to conventional biomarkers for differential diagnosis of chronic kidney diseases. Int J Mol Sci. https://doi.org/10.3390/ijms21134802
Navaneeth B, Suchetha M (2019) PSO optimized 1-D CNN-SVM architecture for real-time detection and classification applications. Comput Biol Med 108:85–92. https://doi.org/10.1016/j.compbiomed.2019.03.017
Chen C, Yang L, Li H et al (2020) Raman spectroscopy combined with multiple algorithms for analysis and rapid screening of chronic renal failure. Photodiagnosis Photodyn Ther. 30:101792. https://doi.org/10.1016/j.pdpdt.2020.101792
Han X, Zheng X, Wang Y et al (2019) Random forest can accurately predict the development of end-stage renal disease in immunoglobulin a nephropathy patients. Ann Transl Med 7(11):234. https://doi.org/10.21037/atm.2018.12.11
Shih CC, Lu CJ, Chen GD et al (2020) Risk prediction for early chronic kidney disease: results from an adult health examination program of 19,270 individuals. Int J Environ Res Public Health. https://doi.org/10.3390/ijerph17144973
Kannan S, Morgan LA, Liang B et al (2019) Segmentation of glomeruli within trichrome images using deep learning. Kidney Int Rep 4(7):955–962. https://doi.org/10.1016/j.ekir.2019.04.008
Aldhyani THH, Alshebami AS, Alzahrani MY (2020) Soft clustering for enhancing the diagnosis of chronic diseases over machine learning algorithms. J Healthc Eng 2020:4984967. https://doi.org/10.1155/2020/4984967
Kleiman RS, LaRose ER, Badger JC et al (2018) Using machine learning algorithms to predict risk for development of calciphylaxis in patients with chronic kidney disease. AMIA Jt Summits Transl Sci Proc AMIA Jt Summits Transl Sci 2017:139–146
Dovgan E, Gradišek A, Luštrek M et al (2020) Using machine learning models to predict the initiation of renal replacement therapy among chronic kidney disease patients. PLoS ONE 15(6):e0233976. https://doi.org/10.1371/journal.pone.0233976
Wu X, Yuan X, Wang W et al (2020) Value of a machine learning approach for predicting clinical outcomes in young patients with hypertension. Hypertens Dallas Tex. 75(5):1271–1278. https://doi.org/10.1161/HYPERTENSIONAHA.119.13404
Ogunleye A, Wang QG (2020) XGBoost model for chronic kidney disease diagnosis. IEEE/ACM Trans Comput Biol Bioinform 17(6):2131–2140. https://doi.org/10.1109/TCBB.2019.2911071
Connell A, Black G, Montgomery H et al (2019) Implementation of a digitally enabled care pathway (part 2): qualitative analysis of experiences of health care professionals. J Med Internet Res 21(7):e13143. https://doi.org/10.2196/13143
Wilson FP, Martin M, Yamamoto Y et al (2021) Electronic health record alerts for acute kidney injury: multicenter, randomized clinical trial. BMJ 372:4786. https://doi.org/10.1136/bmj.m4786
Röösli E, Bozkurt S, Hernandez-Boussard T (2022) Peeking into a black box, the fairness and generalizability of a MIMIC-III benchmarking model. Sci Data 9(1):24. https://doi.org/10.1038/s41597-021-01110-7
UCI Machine Learning Repository. Accessed July 28, 2022. https://archive.ics.uci.edu/ml/index.php
Linardatos P, Papastefanopoulos V, Kotsiantis S (2020) Explainable AI: a review of machine learning interpretability methods. Entropy 23(1):18. https://doi.org/10.3390/e23010018
Murdoch WJ, Singh C, Kumbier K et al (2019) Definitions, methods, and applications in interpretable machine learning. Proc Natl Acad Sci U S A 116(44):22071–22080. https://doi.org/10.1073/pnas.1900654116
Amann J, Blasimme A, Vayena E et al (2020) Explainability for artificial intelligence in healthcare: a multidisciplinary perspective. BMC Med Inform Decis Mak 20:310. https://doi.org/10.1186/s12911-020-01332-6
Payrovnaziri SN, Chen Z, Rengifo-Moreno P et al (2020) Explainable artificial intelligence models using real-world electronic health record data: a systematic scoping review. J Am Med Inform Assoc JAMIA 27(7):1173–1185. https://doi.org/10.1093/jamia/ocaa053
Collins GS, Reitsma JB, Altman DG et al (2015) Transparent reporting of a multivariable prediction model for individual prognosis or diagnosis (TRIPOD): the TRIPOD Statement. BMC Med 13(1):1. https://doi.org/10.1186/s12916-014-0241-z
Hernandez-Boussard T, Bozkurt S, Ioannidis JPA et al (2020) MINIMAR (MINimum Information for Medical AI Reporting): developing reporting standards for artificial intelligence in health care. J Am Med Inform Assoc JAMIA 27(12):2011–2015. https://doi.org/10.1093/jamia/ocaa088
Gianfrancesco MA, Tamang S, Yazdany J et al (2018) Potential biases in machine learning algorithms using electronic health record data. JAMA Intern Med 178(11):1544–1547. https://doi.org/10.1001/jamainternmed.2018.3763
Panch T, Mattie H, Atun R (2002) Artificial intelligence and algorithmic bias: implications for health systems. J Glob Health 9(2):20318. https://doi.org/10.7189/jogh.09.020318
Ramspek CL, Jager KJ, Dekker FW et al (2020) External validation of prognostic models: what, why, how, when and where? Clin Kidney J 14(1):49–58. https://doi.org/10.1093/ckj/sfaa188
Steyerberg EW, Bleeker SE, Moll HA et al (2003) Internal and external validation of predictive models: a simulation study of bias and precision in small samples. J Clin Epidemiol 56(5):441–447. https://doi.org/10.1016/s0895-4356(03)00047-7
Riley RD, Ensor J, Snell KIE et al (2016) External validation of clinical prediction models using big datasets from e-health records or IPD meta-analysis: opportunities and challenges. BMJ 353:3140. https://doi.org/10.1136/bmj.i3140
Acknowledgements
None.
Funding
Open access funding provided by Alma Mater Studiorum - Università di Bologna within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Contributions
FS and AC had the idea, extracted, and analyzed the data and wrote the manuscript. CF analyzed the data and wrote the manuscript. DGol, DGor, helped in results interpretation. THB revised the manuscript and helped in results interpretation. AC supervised the entire process.
Corresponding author
Ethics declarations
Disclosure statement
The authors did not receive support from any organization for the submitted work. The authors do not have any conflicts of interest to report.
Ethics approval
Not Applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original article has been updated: Due to Abstract changes.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sanmarchi, F., Fanconi, C., Golinelli, D. et al. Predict, diagnose, and treat chronic kidney disease with machine learning: a systematic literature review. J Nephrol 36, 1101–1117 (2023). https://doi.org/10.1007/s40620-023-01573-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40620-023-01573-4