Abstract
Changes in the normal nucleotide sequence of the human genome plays an integral part in the development and progression of all cancers. It is now recognized that certain genomic alterations can be used to determine prognosis and potential response to treatment for patients diagnosed with a range of cancers. Breast cancer is perhaps the best example of how gene expression analysis can influence treatment choices and potentially prevent patients from receiving adjuvant chemotherapy that would have a negligible benefit. It is becoming increasingly likely that most, if not all, tumors will be subjected to some form of genomic analysis in the future in an attempt to provide patients with a personalized treatment plan that maximizes efficacy and reduces toxicity. This paper gives a brief history of genomic sequencing followed by a description of clinically relevant oncogenomics for those working in the colorectal cancer field. The relevance of RAS, BRAF, mismatch repair, and microsatellite instability status are discussed in detail. Potential implications of PIK3CA mutations are briefly described. It finishes by providing a summary of more complex techniques such as whole exome and whole genome sequencing, some of which may be used in the clinical setting and some of which will be reserved solely for novel target and biomarker identification in the academic setting. An understanding of the molecular mechanisms that underlie the development and progression of colorectal cancer is necessary and will become more relevant as the practice of personalized medicine is more widely implemented.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The treatment of many malignancies has undergone significant change in recent decades with a shift towards minimally invasive and organ-preserving surgery in an attempt to reduce morbidity and improve quality of life without compromising survival [1,2,3]. While surgical resection of the tumor with clear margins remains one of the core principles of surgical oncology and a vital component in the treatment of patients diagnosed with a malignancy, newer treatment strategies such as targeted therapy and immunotherapy have begun to play an important role for many patients and will be key to reducing disease recurrence and progression as well as improving long-term survival for patients going forward [4, 5]. The traditional multidisciplinary team (MDT) consisting of surgeons, medical oncologists, radiation oncologists, pathologists, and radiologists has grown in recent years to include nurse specialists and clinical geneticists and in the future may include specialists in the field of bioinformatics [6, 7]. Personalized medicine, once an idea of the future, is already in practice for many patients who are undergoing treatment for a wide variety of malignancies [8]. The decision as to which treatment will be of benefit to which patient can be based on many factors, but increasingly these decisions will be made based on a single genomic mutation or panel of such mutations [9]. To what extent this is relevant to the surgeon is debatable; however, these genomic alterations are being discussed more frequently at MDT meetings and patients are increasingly educated about their prognoses and even the molecular intricacies of their cancers [10]. An understanding of the methods of determining these mutations, knowledge of the clinical utility of certain mutations, and what the future of sequencing is likely to hold are important for the twenty-first century surgeon in order to enable them to interact with other members of the MDT and their patients in a meaningful way [11]. This paper presents a brief history of molecular genomics and relevant techniques followed by a description of clinically relevant oncogenomics of colorectal cancer (CRC) for the surgeon. Finally, we outline novel oncogenomic and transcriptomic techniques that may become clinically relevant in the not so distant future.
Discussion
The History of Sequencing, Oncogenomics, and Personalized Medicine
The three-dimensional structure of DNA was discovered in 1953 by Watson and Crick; however, uncovering methods of accurately sequencing the nucleotides that make up deoxyribonucleic acid (DNA) was not discovered until many years later [12]. Successful sequencing of protein and RNA was achieved before sequencing of DNA, and this was pioneered by Fred Sanger [13]. Sanger determined the protein sequence of insulin in the 1950s; however, the sequencing of DNA proved to be cumbersome initially, and it was not until 1976 that two methods were developed that were capable of decoding hundreds of bases in an afternoon, and with this, the science of sequencing was transformed [14,15,16,17]. By the 1980s, automated Sanger sequencing machines were widely available and the spirit of data sharing saw the creation of multiple publicly available sequencing data repositories. The field continued to grow, and in 2004, the Human Genome Project (HGP) released the finished sequence of the entire human genome [18, 19]. The HGP cost approximately $2.7 billion and took over 13 years to complete; at present, an entire human genome can be sequenced in a matter of days for under $1000, and the time and cost reduction of whole genome sequencing (WGS) has been an important step in making personalized medicine more affordable. After the HGP, a technique called massively parallel next-generation DNA sequencing (NGS) superseded Sanger sequencing [20]. These sequencing technologies were largely used for academic purposes initially, but more recently the technology has been applied to the discovery of cancer driver genes, to detect chromosomal aneuploidies by noninvasive prenatal testing and to help discover the genes responsible for Mendelian disorders [21,22,23,24]. In more recent years, sequencing of tumor DNA to look for specific mutations has been shown to be a valid technique for determining potential response to targeted treatments [25].
Gene expression analysis differs from gene sequencing for specific mutations. Sequencing involves determining the exact nucleotides in a specific gene and comparing the sequence in DNA extracted from tissue to that of the known normal sequence. Gene expression analysis involves extracting ribonucleic acid (RNA) from tissue and then using a method known as reverse transcription polymerase chain reaction (RT-PCR), the RNA is reverse transcribed into complementary DNA (cDNA), and amplification of specific DNA targets is performed using PCR thus allowing one to calculate the quantity of a gene product that is being produced [26]. This technique can be used to generate multipanel transcript signatures that can be used to determine prognosis and predict response to treatment.
Breast cancer has without doubt been the pioneer of personalized medicine in oncology [27]. Immunohistochemical (IHC) testing is used to determine estrogen and progesterone receptor status [28]. Patients who have breast cancers that are hormone receptor positive on IHC are potential candidates for treatment with selective estrogen receptor modulators such as tamoxifen or with aromatase inhibitors such as anastrozole [29]. IHC or fluorescent in situ hybridization (FISH) can be used to determine human epidermal growth factor 2 (HER2) receptor expression or amplification, respectively [30]. HER2-targeted therapy in the form of trastuzumab or one of the other related compounds can be offered to the majority of patients with HER2-positive breast cancer [31]. Along with helping to decide on potential treatment strategies, these markers are used to help determine prognosis [28]. Molecular signature analysis using commercially available kits such as Oncotype DX® is clinically validated and is now used to determine the risk of recurrence and the likely benefit of chemotherapy for patients with early-stage estrogen receptor-positive, HER2-negative breast cancer [32, 33]. This test uses RT-PCR to measure the expression of 16 cancer-related genes and five reference genes. Each patient is given a score between 0 and 100; those with a score less than 18 can be spared chemotherapy due to their low risk of recurrence [34,35,36]. The stratification of patients based on the results of these molecular tests has resulted in personalized and precision medicine for those diagnosed with breast cancer. This approach has created the ideal scenario where those patients at high risk of disease recurrence are offered systemic chemotherapy while those with a very low risk are spared chemotherapy and its potential side effects.
Current Applications of DNA Sequencing for Colorectal Cancer—Where We Started
Until relatively recently, studies into the oncogenomics of CRC were concerned predominantly with tumor development and progression [37]. These studies demonstrated that activation of oncogenes and inhibition of tumor suppression genes were responsible for the development of polyps which gradually progressed to CRC. As the polyp progressed through increasing grades of dysplasia and eventually into invasive disease, it gradually accumulated a greater number of mutations [38]. The adenomatous polyposis coli (APC) tumor suppressor gene found on chromosome 5q21 was found to be mutated very early in the adenoma-carcinoma sequence [39, 40]. Germline mutations in this gene are responsible for causing familial adenomatous polyposis (FAP) [41]. Since the discovery of the genomic pathway underlying the adenoma-carcinoma sequence, more focused research into drug targets and biomarkers of sensitivity and resistance to treatment has been carried out.
Current Applications of DNA Sequencing for Colorectal Cancer—Mutations in the RAS Gene Family
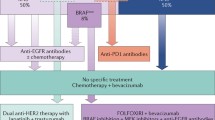
Sequencing of tumor DNA has proven to be useful for determining which patients with metastatic CRC should be considered for adjuvant treatment with epidermal growth factor receptor (EGFR) inhibitors such as cetuximab and panitumumab [42]. At present, the RAS and RAF status are analyzed in patients being considered for targeted therapies; these proteins are involved in the RAS/RAF/MEK/ERK signaling pathway. Mutations in these proteins result in continuous activation of the pathway, and this leads to progression of the cell cycle and eventually uncontrolled cellular growth [43, 44]. In particular, patients with RAS mutations respond poorly to anti-EGFR agents and only those who are RAS wild type should be commenced on one of these medications [45]. It is now recommended that patients should undergo extended KRAS analysis which looks for mutations on codons 12 and 13 of exon 2, codon 61 of exon 3, and codons 117 and 146 of exon 4. Exons 2, 3, and 4 of NRAS should also be analyzed for mutations [46,47,48]. Despite the recommendations for extended RAS testing in clinical practice, there is evidence to suggest that implementation of extended testing has been slow and is still not widely performed [49]. RAS mutations are present in approximately 45–56% of metastatic CRCs, with the vast majority of mutations occurring at KRAS exon 2 [50, 51].
Current Applications of DNA Sequencing for Colorectal Cancer—Mutations in the BRAF Gene
RAS and BRAF mutations are usually mutually exclusive [52]. The BRAF mutation results in a constitutively active protein which continuously activates the mitogen-activated protein kinase (MAPK) pathway; this in turn has effects on cell growth, proliferation, survival, and migration. The BRAFV600E mutation accounts for 80% of all BRAF mutations and is found in 8–12% of metastatic CRCs; furthermore, there is accumulating evidence to suggest that this mutation is also associated with resistance to anti-EGFR therapy alone or in combination with cytotoxics [50, 52,53,54,55]. Patients with BRAF-mutant metastatic CRC are known to have a poorer prognosis; in particular, this mutation is associated with shorter progression free and overall survival [56,57,58]. While BRAF inhibitors have shown some success in the setting of melanoma, the same results have not been found in CRC where only about 5% of patients are found to respond [59]. There is a strong association between BRAF-mutant and microsatellite instability high (MSI-H) CRC, and patients with this genomic subtype may get some response to the immune-oncology drugs used for metastatic MSI-H CRC [60]. More recently, there have been positive results with the combination of an EGFR inhibitor, a BRAF inhibitor, and a MEK inhibitor for patients with BRAFV600E-mutant CRC [61].
Current Applications of DNA Sequencing for Colorectal Cancer—PIK3CA Mutations and HER2 Amplification
PIK3CA is the catalytic subunit of phosphatidylinositol-3-kinase (PI3K) and is involved in the promotion of various cellular processes, including proliferation, survival, apoptosis, migration, and metabolism [62]. PIK3CA mutations can be found in 10–20% of CRCs with the majority of mutations occurring in exons 9 and 20 [63,64,65,66,67,68,69]. These mutations are often found in association with KRAS mutations [70]. Patients with PIK3CA mutations who are KRAS wild type have been shown to have a poorer prognosis compared to those with KRAS mutations [52, 67]. PIK3CA mutations have also been associated with resistance to anti-EGFR therapies, and much of this resistance may be due to exon 20 mutations with exon 9 mutations potentially playing no part in the resistance pattern [52, 71,72,73]. Patients with tumors harboring PIK3CA mutations who are taking aspirin have been shown to have reduced recurrence and improved survival compared to those not taking aspirin [64, 74]. KRAS, NRAS, BRAF, and PIK3CA are clearly linked by the fact that certain somatic mutations in these genes can result in resistance to anti-EGFR therapies. It has been suggested that only patients who are quadruple wild type in these four genes will derive a benefit from anti-EGFR therapy, and this is biologically plausible when one considers the actions of the protein products derived from these genes [52]. Even within the quadruple wild-type cohort, there will be some patients that will not achieve a meaningful benefit from treatment with anti-EGFR therapy. It is known that a proportion of patients in this group have amplification of the HER2 gene resulting in an increase in the HER2 gene copy number which has been shown to be associated with resistance to anti-EGFR therapy [75].
Mismatch Repair Deficiency and Microsatellite Instability—How Is It Determined and What Is the Relevance
Determining whether a tumor is mismatch repair (MMR) deficient or MSI-H has become increasingly important for several reasons. Firstly, recognizing those patients with germline mutations in one of the mismatch repair genes allows a diagnosis of Lynch syndrome to be made, and these patients can be offered appropriate screening for Lynch syndrome-associated cancers [76, 77]. Secondly, there is evidence that some patients who are MMR deficient or MSI-H respond poorly to 5-flourouracil (5-FU) chemotherapy although the importance of this is somewhat questionable given that most patients are currently offered a combined chemotherapy regimen in the adjuvant setting [78,79,80]. Despite the poor response to 5-FU, there is accumulating evidence that patients with MMR-deficient or MSI-H tumors have a better prognosis when compared to those with MMR-proficient or microsatellite stable tumors (MSS) [81,82,83,84,85]. The importance of determining the MMR/MSI status has gained increasing importance recently for those patients with metastatic CRC. Patients with unresectable or metastatic MMR-deficient or MSI-H tumors are now eligible for treatment with pembrolizumab or nivolumab, both programmed cell death-1 (PD-1) monoclonal antibodies [60, 86,87,88,89,90]. MMR status is determined using immunohistochemistry (IHC). Normal tissue and tumor are stained for the MMR proteins: MLH1, MSH2, MSH6, and PMS2. If there is loss of staining for one or more of these proteins, then the tumor is deemed to be MMR deficient. MSI status may be determined as a first line technique or may be used in cases where the IHC result is equivocal. PCR is the technique used to determine MSI status. With this technique, DNA is extracted from the normal epithelium and from the tumor and then amplified using specific PCR primers. The amplified material is then analyzed by fragment analysis. DNA from both the normal epithelium and tumor epithelium are compared. The number, type, and identity of the microsatellites that should be used for MSI assessment remain unclear, and the criteria used to diagnose MSI differ among studies [91]. The microsatellites analyzed in the assay used in our institution are the DNA mononucleotide repeat sequences NR-21, NR-24, NR 27, Bat-25, and Bat26. Microsatellite status can be divided into three categories based on the following criteria: MSI-high (MSI-H), indicating instability at two or more loci (or > 30% of loci if a larger panel of markers is used); MSI-low (MSI-L), indicating instability at one locus (or in 10–30% of loci in larger panels); and MSS, indicating no loci with instability (or < 10% of loci in larger panels) [92].
Current Applications of DNA Sequencing for Colorectal Cancer—When Will It Be Used
The implementation of oncogenomics could have the potential to improve outcomes for our patients. There is no doubt that the use of oncogenomics will result in a change of practice for oncologists as they begin to personalize treatment for patients based on the molecular characteristics of their tumors as opposed to treating them based on tumor morphology and disease stage. There is still a lot of uncertainty as to how the widespread implementation of oncogenomics will change surgical practice. Sequencing results from biopsy specimens in the case of rectal cancer may be taken into consideration when considering which patients should receive neoadjuvant chemoradiotherapy and what regimens they should receive. The use of genomic data to help make treatment decisions when targeting advanced and metastatic tumors may allow some patients to be considered for surgical resection and potentially curative treatment in situations where that may not have been possible in the past. More recently, there has been evidence to show that there are genomic differences between MSS tumors arising from the right colon compared to MSS tumors arising from the left colon. Right-sided MSS tumors have a significant enrichment of oncogenic alterations in KRAS, BRAF, PIK3CA, SMAD2, and SMAD4, whereas left-sided MSS tumors are enriched for oncogenic alterations in TP53 and APC. Understanding the mutational pattern of right- and left-sided CRC is important as right-sided tumors appear to have a worse prognosis, and it seems logical that this is in part driven by the pattern of genomic alterations found in these tumors. Identifying actionable mutations in diagnostic biopsies may allow patients to commence on appropriate targeted therapy from the time of diagnosis, and this may ultimately improve their long-term outcomes [93]. For these reasons, it is important for surgeons to have an understanding of the potential use of genomic medicine in the setting of CRC.
Potential Applications of Sequencing in the Future—Whole Genome and Whole Exome Sequencing
Whole exome sequencing and whole genome sequencing are two modalities which have the potential to become integrated into clinical practice in the near future. Exons account for approximately 1% of the entire genome and contain the coding sequences that provide instructions for building proteins. All exons in the genome are known as the exome, and the method of sequencing them is called whole exome sequencing (WES). Variations in the protein coding region of most genes can be identified with this method, and given that the majority of known disease-causing mutations occur in exons, it is felt to be a practical method [94]. Whole genome sequencing (WGS) can determine the nucleotide sequence of the entire human genome which includes the exons, introns, and other noncoding DNA and hence can detect variations in promoter, enhancer, and regulatory regions of the genome as well as in protein coding regions [95]. WES and WGS are predominantly used in the research setting, but they have been used successfully in the clinical setting to identify the causative genetic variation in a number of Mendelian disorders. The sequencing can be carried out on a piece of fresh frozen (FF) tissue or on formalin-fixed paraffin-embedded (FFPE) tissue. A sample of normal tissue, often normal colonic mucosa adjacent to the tumor, from the patient is sent with the tumor sample to be sequenced at the same time, and this is used as a comparison. As the price of WES and WGS continues to drop and the amount of time it takes to carry out the sequencing gets shorter, it is likely that one of these methods may be used routinely for patients diagnosed with CRC instead of sequencing being performed for a small number of targets such as KRAS, NRAS, and BRAF. At present, one of the biggest limiting factors of this technique is the availability of bioinformaticians to accurately analyze the vast quantity of data that is returned with each case after sequencing [96]. Currently, there are a number of laboratories around the world that can perform WES or WGS at the request of patients. The laboratories will make direct contact with the hospital and arrange transport of the sample either FF or FFPE, and once the sequencing and bioinformatic analysis have been performed, the patient and their clinicians will receive a detailed sequencing report. The reports typically contain information on the quality of sequencing that was undertaken, the genomic variations that were identified, and details on potentially suitable treatments. The recommendations regarding treatment options are based on published literature, and the reports will contain the level of evidence the recommendation is based upon. The clinician is provided details on treatments the patient is sensitive to, treatments they may be resistant to, and also treatments that the patient is more likely to experience toxicity with. At present, there is still uncertainty as to where this type of technology fits into clinical practice and to what extent if any the results of WES or WGS should be used when making decisions about a patient’s treatment; however, there are emerging and encouraging reports of how this technology is being used in the clinical setting specifically for patients with CRC [97]. WES and WGS interpretation and reporting are still in its infancy, but we are beginning to see a push from informatic associations towards standardized reporting of WES and WGS much like a radiology report so that they can be easily interpreted and used in the clinical setting [98].
Potential Applications of Sequencing in the Future—Liquid Biopsy, Cell-Free DNA, and Circulating Tumor DNA
Liquid biopsy facilitates identification and analysis of fragments of cancer DNA from a fluid sample; to date, this has predominantly been through analysis of blood. Circulating tumor DNA (ctDNA) is derived from cell-free DNA (cfDNA) which represents DNA present in the noncellular portion of blood that originated from either normal tissue or tumor sources. Circulating tumor cells (CTCs) are cells that have shed from the primary tumor into the blood or lymphatic system. CTCs will be found in the cellular portion of blood and can be differentiated from normal cells by the distinct physical properties of the cells or by their differential expression of cell surface proteins. Studies have demonstrated that the frequency of genomic alterations identified in cfDNA is similar to that found when direct sequencing of the primary tumor is performed. Further advantages in the setting of CRC include the identification of novel EGFR mutations which confer resistance to anti-EGFR therapies and providing a greater understanding of tumor heterogeneity [99]. It appears that the main advantages of liquid biopsy, ctDNA, and CTCs will be in detecting minimal residual disease; this typically reflects the presence of tumor cells disseminated from the primary lesion to distant organs in patients who lack any clinical or radiological signs of metastasis or residual tumor cells left behind after local therapy that eventually lead to local recurrence [100]. It seems likely that liquid biopsy will be used for early detection of residual disease and also to help personalize treatment of residual disease based on the genomic alterations identified.
Potential Applications of Sequencing in the Future—Transcriptomics, from DNA to RNA
The use of transcriptomics in the setting of CRC also looks promising. Molecular signature analysis for colon cancer using Oncotype DX® and other tests is available but is yet to be implemented into routine clinical practice. This test uses RT-PCR to analyze the expression of seven cancer-related genes and five reference genes for patients with stage II (mismatch repair proficient) and stage IIIA/B colon cancer; the 12 gene panel was chosen from a selection of 761 candidate genes. The test is aimed at offering an individualized quantifiable risk of recurrence which can be used when deciding the need for adjuvant treatment. The recurrence score derived from Oncotype DX® has been shown to be accurate at predicting recurrence in stage II and stage III colon cancer; however, it is still unclear if it is able to accurately predict response to chemotherapy [101,102,103,104].
Guinney et al. have used unbiased transcriptome analysis data from 4151 patients to propose four consensus molecular subtypes (CMS) of CRC that are assumed to have a common biology while Isella et al. used patient-derived xenografts from 244 patients to define five CRC intrinsic subtypes (CRIS) [105, 106]. CMS1 represents 14% of cases, and these tumors tend to be hypermutated and microsatellite unstable with strong immune activation. CMS2 accounts for 37% of cases, and its features include marked activation of signaling pathways involved in cancer cell proliferation and survival such as the WNT and MYC signaling pathways. CMS3 includes 13% of cases and is known as the metabolic type; there is epithelial and evident metabolic dysregulation. CMS4 is a mesenchymal type accounting for 23% of cases; this subtype has prominent transforming growth factor-β activation, stromal invasion, and angiogenesis. The 13% of samples with mixed features possibly represent a transition phenotype or intratumoral heterogeneity. The subtypes correlate with clinicopathological variables and outcomes. For example, CMS1 tumors tend to be found most frequently in females that have right-sided tumors with higher histopathological grades. CMS4 tumors tend to have the worst overall survival and relapse-free survival while CMS2 patients have superior survival after relapse [105]. This is just one example of how gene expression data might be used to help determine prognosis and need for adjuvant treatment for patients with CRC in the future. A NanoString-based assay that uses molecular barcodes and microscopic imaging to detect and count up to several hundred unique transcripts in one hybridization reaction could be a useful way to use transcriptomics routinely in the clinical setting; such assays have already been described in the literature and are likely to be commercially available once validated [107].
Conclusion
DNA analysis of tumor specimens has become increasingly common and has begun to influence decision-making, particularly with regard to adjuvant treatment. Breast cancer is the best example of how personalized medicine can be implemented into clinical practice and how it can help inform decision-making. At present, a small number of gene markers have proven to be useful in the setting of CRC, but for the most part, these are limited to RAS, BRAF, PIK3CA, and MMR/MSI status. There are many exciting and promising applications in the pipeline that have the potential to be used when determining prognosis and likely response to targeted therapies for patients with CRC. While it is important for clinicians to be open minded about new technologies, caution should be taken before implementing these into routine clinical practice. Even as the price of sequencing technologies continues to fall, they still remain quite expensive and one must ensure that these technologies serve a useful purpose and are cost-effective before they are implemented into routine clinical practice. The practical issues of implementing these technologies into routine practice must not be forgotten and include issues such as tissue availability, sequencing technology expertise, and most importantly the availability of clinical bioinformaticians to inform the cancer MDT. Surgeons will also have to play an active role if genomic and transcriptomic data is to be successfully utilized when considering treatment options for patients with cancer of the colon and rectum.
References
Kanehira E, Tanida T, Kamei A, Takahashi K. Minimally invasive, organ-preserving surgery for large submucosal tumors in the abdominal esophagus. Surg Laparosc Endosc Percutan Tech. 2017;27(3):189–93.
Caudle AS, Kuerer HM. Breast conservation therapy after neoadjuvant chemotherapy: optimization of a multimodality approach. J Surg Oncol. 2014;110(1):32–6.
Stijns RCH, de Graaf EJR, Punt CJA, et al. Long-term oncological and functional outcomes of chemoradiotherapy followed by organ-sparing transanal endoscopic microsurgery for distal rectal cancer: the CARTS Study. JAMA Surg. 2019;154(1):47–54.
Gotwals P, Cameron S, Cipolletta D, et al. Prospects for combining targeted and conventional cancer therapy with immunotherapy. Nat Rev Cancer. 2017;17(5):286–301.
Baudino TA. Targeted cancer therapy: the next generation of cancer treatment. Curr Drug Discov Technol. 2015;12(1):3–20.
Apweiler R, Beissbarth T, Berthold MR, et al. Whither systems medicine? Exp Mol Med. 2018;50(3):e453.
Koole T, van Burgsteden L, Harms P, et al. Participation in interdisciplinary meetings on genetic diagnostics (NGS). Eur J Hum Genet. 2017;25(10):1099–105.
Jackson SE, Chester JD. Personalised cancer medicine. Int J Cancer. 2015;137(2):262–6.
Zhang X, Cha IH, Kim KY. Use of a combined gene expression profile in implementing a drug sensitivity predictive model for breast cancer. Cancer Res Treat. 2017;49(1):116–28.
Cartwright LA, Dumenci L, Siminoff LA, Matsuyama RK. Cancer patients’ understanding of prognostic information. J Cancer Educ. 2014;29(2):311–7.
Kant R, Ravi B. The surgeon and molecular knowledge. Indian J Surg. 2013;75(3):167–9.
Watson JD, Crick FH. Molecular structure of nucleic acids; a structure for deoxyribose nucleic acid. Nature. 1953;171(4356):737–8.
Shendure J, Balasubramanian S, Church GM, et al. DNA sequencing at 40: past, present and future. Nature. 2017;550(7676):345–53.
Sanger F, Thompson EO. The amino-acid sequence in the glycyl chain of insulin. I. The identification of lower peptides from partial hydrolysates. Biochem J. 1953;53(3):353–66.
Sanger F, Thompson EO. The amino-acid sequence in the glycyl chain of insulin. II. The investigation of peptides from enzymic hydrolysates. Biochem J. 1953;53(3):366–74.
Sanger F, Nicklen S, Coulson AR. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci USA. 1977;74(12):5463–7.
Maxam AM, Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci USA. 1977;74(2):560–4.
Lander ES, Linton LM, Birren B, et al. Initial sequencing and analysis of the human genome. Nature. 2001;409(6822):860–921.
Finishing the euchromatic sequence of the human genome (2004). Nature 431 (7011):931–945.
Gould Rothberg BE, Rothberg JM. Massively parallel (“next-generation”) DNA sequencing. Clin Chem. 2015;61(7):997–8.
Ng SB, Turner EH, Robertson PD, et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature. 2009;461(7261):272–6.
Choi M, Scholl UI, Ji W, et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci USA. 2009;106(45):19096–101.
Wood LD, Parsons DW, Jones S, et al. The genomic landscapes of human breast and colorectal cancers. Science (New York, NY). 2007;318(5853):1108–13.
Chiu RW, Chan KC, Gao Y, et al. Noninvasive prenatal diagnosis of fetal chromosomal aneuploidy by massively parallel genomic sequencing of DNA in maternal plasma. Proc Natl Acad Sci USA. 2008;105(51):20458–63.
Johnson DB, Frampton GM, Rioth MJ, et al. Targeted next generation sequencing identifies markers of response to PD-1 blockade. Cancer Immunol Res. 2016;4(11):959–67.
Segundo-Val IS, Sanz-Lozano CS. Introduction to the gene expression analysis. Methods Mol Biol. 2016;1434:29–43.
Chan CWH, Law BMH, So WKW et al. Novel strategies on personalized medicine for breast cancer treatment: an update. Int J Mol Sci 2017;18 (11).
Senkus E, Kyriakides S, Ohno S, et al. Primary breast cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Annals of oncology : official journal of the European Society for Medical Oncology. 2015;26(Suppl 5):v8-30.
Burstein HJ, Lacchetti C, Anderson H, et al. Adjuvant endocrine therapy for women with hormone receptor-positive breast cancer: ASCO Clinical Practice Guideline Focused Update. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2019;37(5):423–38.
Wolff AC, Hammond MEH, Allison KH, et al. Human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists Clinical Practice Guideline Focused Update. J Clin Oncol. 2018;36(20):2105–22.
Giordano SH, Temin S, Chandarlapaty S, et al. Systemic therapy for patients with advanced human epidermal growth factor receptor 2-positive breast cancer: ASCO Clinical Practice Guideline Update. Journal of clinical oncology : official journal of the American Society of Clinical Oncology. 2018;36(26):2736–40.
Paik S, Shak S, Tang G, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004;351(27):2817–26.
Habel LA, Shak S, Jacobs MK, et al. A population-based study of tumor gene expression and risk of breast cancer death among lymph node-negative patients. Breast Cancer Res. 2006;8(3):R25.
Toi M, Iwata H, Yamanaka T, et al. Clinical significance of the 21-gene signature (Oncotype DX) in hormone receptor-positive early stage primary breast cancer in the Japanese population. Cancer. 2010;116(13):3112–8.
Dowsett M, Cuzick J, Wale C, et al. Prediction of risk of distant recurrence using the 21-gene recurrence score in node-negative and node-positive postmenopausal patients with breast cancer treated with anastrozole or tamoxifen: a TransATAC study. J Clin Oncol. 2010;28(11):1829–34.
Albain KS, Barlow WE, Shak S, et al. Prognostic and predictive value of the 21-gene recurrence score assay in postmenopausal women with node-positive, oestrogen-receptor-positive breast cancer on chemotherapy: a retrospective analysis of a randomised trial. Lancet Oncol. 2010;11(1):55–65.
Vogelstein B, Fearon ER, Hamilton SR, et al. Genetic alterations during colorectal-tumor development. N Engl J Med. 1988;319(9):525–32.
Fearon ER, Vogelstein B. A genetic model for colorectal tumorigenesis. Cell. 1990;61(5):759–67.
Bodmer WF, Bailey CJ, Bodmer J, et al. Localization of the gene for familial adenomatous polyposis on chromosome 5. Nature. 1987;328(6131):614–6.
Groden J, Thliveris A, Samowitz W, et al. Identification and characterization of the familial adenomatous polyposis coli gene. Cell. 1991;66(3):589–600.
Bodmer W. The somatic evolution of cancer The Harveian Oration of 1996. J R Coll Physicians Lond. 1997;31(1):82–9.
Miyamoto Y, Suyama K, Baba H (2017) Recent advances in targeting the EGFR signaling pathway for the treatment of metastatic colorectal cancer. Int J Mol Sci 2017;18 (4).
McCormick F. K-Ras protein as a drug target. J Mol Med (Berl). 2016;94(3):253–8.
Arrington AK, Heinrich EL, Lee W, et al. Prognostic and predictive roles of KRAS mutation in colorectal cancer. Int J Mol Sci. 2012;13(10):12153–68.
Van Cutsem E, Cervantes A, Nordlinger B, Arnold D (2014) Metastatic colorectal cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 25 Suppl 3:iii1–9.
Douillard JY, Siena S, Cassidy J, et al. Randomized, phase III trial of panitumumab with infusional fluorouracil, leucovorin, and oxaliplatin (FOLFOX4) versus FOLFOX4 alone as first-line treatment in patients with previously untreated metastatic colorectal cancer: the PRIME study. J Clin Oncol. 2010;28(31):4697–705.
Grothey A. EGFR antibodies in colorectal cancer: where do they belong? J Clin Oncol. 2010;28(31):4668–70.
Douillard JY, Oliner KS, Siena S, et al. Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N Engl J Med. 2013;369(11):1023–34.
Richman SD, Fairley J, Butler R, Deans ZC. How close are we to standardised extended RAS gene mutation testing? The UK NEQAS evaluation. J Clin Pathol. 2017;70(1):58–62.
Porru M, Pompili L, Caruso C, et al. Targeting KRAS in metastatic colorectal cancer: current strategies and emerging opportunities. J Exp Clin Cancer Res. 2018;37(1):57.
Peeters M, Kafatos G, Taylor A et al. Prevalence of RAS mutations and individual variation patterns among patients with metastatic colorectal cancer: a pooled analysis of randomised controlled trials. European journal of cancer (Oxford, England : 1990) 2015;51 (13):1704–1713.
De Roock W, Claes B, Bernasconi D, et al. Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol. 2010;11(8):753–62.
Tol J, Nagtegaal ID, Punt CJ. BRAF mutation in metastatic colorectal cancer. N Engl J Med. 2009;361(1):98–9.
Di Nicolantonio F, Martini M, Molinari F, et al. Wild-type BRAF is required for response to panitumumab or cetuximab in metastatic colorectal cancer. J Clin Oncol. 2008;26(35):5705–12.
Seymour MT, Brown SR, Middleton G, et al. Panitumumab and irinotecan versus irinotecan alone for patients with KRAS wild-type, fluorouracil-resistant advanced colorectal cancer (PICCOLO): a prospectively stratified randomised trial. Lancet Oncol. 2013;14(8):749–59.
Van Cutsem E, Kohne CH, Lang I, et al. Cetuximab plus irinotecan, fluorouracil, and leucovorin as first-line treatment for metastatic colorectal cancer: updated analysis of overall survival according to tumor KRAS and BRAF mutation status. J Clin Oncol. 2011;29(15):2011–9.
Farina-Sarasqueta A, van Lijnschoten G, Moerland E, et al. The BRAF V600E mutation is an independent prognostic factor for survival in stage II and stage III colon cancer patients. Ann Oncol. 2010;21(12):2396–402.
Phipps AI, Buchanan DD, Makar KW, et al. BRAF mutation status and survival after colorectal cancer diagnosis according to patient and tumor characteristics. Cancer Epidemiol Biomarkers Prev. 2012;21(10):1792–8.
Barras D. BRAF mutation in colorectal cancer: an update. Biomarkers in cancer. 2015;7(Suppl 1):9–12.
Overman MJ, McDermott R, Leach JL, et al. Nivolumab in patients with metastatic DNA mismatch repair-deficient or microsatellite instability-high colorectal cancer (CheckMate 142): an open-label, multicentre, phase 2 study. Lancet Oncol. 2017;18(9):1182–91.
Corcoran RB, André T, Atreya CE, et al. Combined BRAF, EGFR, and MEK inhibition in patients with BRAF(V600E)-mutant colorectal cancer. Cancer Discov. 2018;8(4):428–43.
Vivanco I, Sawyers CL. The phosphatidylinositol 3-kinase AKT pathway in human cancer. Nat Rev Cancer. 2002;2(7):489–501.
Ogino S, Lochhead P, Giovannucci E, et al. Discovery of colorectal cancer PIK3CA mutation as potential predictive biomarker: power and promise of molecular pathological epidemiology. Oncogene. 2014;33(23):2949–55.
Liao X, Lochhead P, Nishihara R, et al. Aspirin use, tumor PIK3CA mutation, and colorectal-cancer survival. N Engl J Med. 2012;367(17):1596–606.
Ogino S, Nosho K, Kirkner GJ, et al. PIK3CA mutation is associated with poor prognosis among patients with curatively resected colon cancer. J Clin Oncol. 2009;27(9):1477–84.
Nosho K, Kawasaki T, Ohnishi M, et al. PIK3CA mutation in colorectal cancer: relationship with genetic and epigenetic alterations. Neoplasia (New York, NY). 2008;10(6):534–41.
Iida S, Kato S, Ishiguro M, et al. PIK3CA mutation and methylation influences the outcome of colorectal cancer. Oncol Lett. 2012;3(3):565–70.
Hsieh LL, Er TK, Chen CC et al. Characteristics and prevalence of KRAS, BRAF, and PIK3CA mutations in colorectal cancer by high-resolution melting analysis in Taiwanese population. Clin Chim Acta. 2012;413(19–20):1605–1611.
Mouradov D, Domingo E, Gibbs P, et al. Survival in stage II/III colorectal cancer is independently predicted by chromosomal and microsatellite instability, but not by specific driver mutations. Am J Gastroenterol. 2013;108(11):1785–93.
Rosty C, Young JP, Walsh MD, et al. PIK3CA activating mutation in colorectal carcinoma: associations with molecular features and survival. PloS one. 2013;8(6):e65479.
Sartore-Bianchi A, Martini M, Molinari F, et al. PIK3CA mutations in colorectal cancer are associated with clinical resistance to EGFR-targeted monoclonal antibodies. Can Res. 2009;69(5):1851–7.
Sood A, McClain D, Maitra R, et al. PTEN gene expression and mutations in the PIK3CA gene as predictors of clinical benefit to anti-epidermal growth factor receptor antibody therapy in patients with KRAS wild-type metastatic colorectal cancer. Clin Colorectal Cancer. 2012;11(2):143–50.
Perrone F, Lampis A, Orsenigo M, et al. PI3KCA/PTEN deregulation contributes to impaired responses to cetuximab in metastatic colorectal cancer patients. Ann Oncol. 2009;20(1):84–90.
Domingo E, Church DN, Sieber O, et al. Evaluation of PIK3CA mutation as a predictor of benefit from nonsteroidal anti-inflammatory drug therapy in colorectal cancer. Am J Clin Oncol. 2013;31(34):4297–305.
Martin V, Landi L, Molinari F, et al. HER2 gene copy number status may influence clinical efficacy to anti-EGFR monoclonal antibodies in metastatic colorectal cancer patients. Br J Cancer. 2013;108(3):668–75.
Carethers JM, Stoffel EM. Lynch syndrome and Lynch syndrome mimics: the growing complex landscape of hereditary colon cancer. World J Gastroenterol. 2015;21(31):9253–61.
Vasen HF, Blanco I, Aktan-Collan K, et al. Revised guidelines for the clinical management of Lynch syndrome (HNPCC): recommendations by a group of European experts. Gut. 2013;62(6):812–23.
Jover R, Zapater P, Castells A, et al. Mismatch repair status in the prediction of benefit from adjuvant fluorouracil chemotherapy in colorectal cancer. Gut. 2006;55(6):848–55.
Carethers JM, Smith EJ, Behling CA, et al. Use of 5-fluorouracil and survival in patients with microsatellite-unstable colorectal cancer. Gastroenterology. 2004;126(2):394–401.
Ribic CM, Sargent DJ, Moore MJ, et al. Tumor microsatellite-instability status as a predictor of benefit from fluorouracil-based adjuvant chemotherapy for colon cancer. N Engl J Med. 2003;349(3):247–57.
Thibodeau SN, Bren G, Schaid D. Microsatellite instability in cancer of the proximal colon. Science (New York, NY). 1993;260(5109):816–9.
Gryfe R, Kim H, Hsieh ET, et al. Tumor microsatellite instability and clinical outcome in young patients with colorectal cancer. N Engl J Med. 2000;342(2):69–77.
Parc Y, Gueroult S, Mourra N, et al. Prognostic significance of microsatellite instability determined by immunohistochemical staining of MSH2 and MLH1 in sporadic T3N0M0 colon cancer. Gut. 2004;53(3):371–5.
Popat S, Hubner R, Houlston RS. Systematic review of microsatellite instability and colorectal cancer prognosis. J Clin Oncol. 2005;23(3):609–18.
Samowitz WS, Curtin K, Ma KN, et al. Microsatellite instability in sporadic colon cancer is associated with an improved prognosis at the population level. Cancer Epidemiol Biomarkers Prev. 2001;10(9):917–23.
Yan L, Zhang W. Precision medicine becomes reality-tumor type-agnostic therapy. Cancer Commun (Lond). 2018;38(1):6.
Brahmer JR, Drake CG, Wollner I, et al. Phase I study of single-agent anti-programmed death-1 (MDX-1106) in refractory solid tumors: safety, clinical activity, pharmacodynamics, and immunologic correlates. J Clin Oncol. 2010;28(19):3167–75.
Le DT, Uram JN, Wang H, et al. PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med. 2015;372(26):2509–20.
Le DT, Durham JN, Smith KN, et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science (New York, NY). 2017;357(6349):409–13.
Overman MJ, Lonardi S, Wong KYM, et al. Durable clinical benefit with nivolumab plus ipilimumab in DNA mismatch repair-deficient/microsatellite instability-high metastatic colorectal cancer. J Clin Oncol. 2018;36(8):773–9.
Pawlik TM, Raut CP, Rodriguez-Bigas MA. Colorectal carcinogenesis: MSI-H versus MSI-L. Dis Markers. 2004;20(4–5):199–206.
Zhang X, Li J. Era of universal testing of microsatellite instability in colorectal cancer. World J Gastrointest Oncol. 2013;5(2):12–9.
Yaeger R, Chatila WK, Lipsyc MD, et al. Clinical sequencing defines the genomic landscape of metastatic colorectal cancer. Cancer Cell. 2018;33(1):125-136.e123.
Warr A, Robert C, Hume D et al. (2015) Exome sequencing: current and future perspectives. G3 (Bethesda, Md) 5 (8):1543–1550.
Ng PC, Kirkness EF. Whole genome sequencing. Methods Mol Biol. 2010;628:215–26.
Scholz MB, Lo CC, Chain PS. Next generation sequencing and bioinformatic bottlenecks: the current state of metagenomic data analysis. Curr Opin Biotechnol. 2012;23(1):9–15.
Stodolna A, He M, Vasipalli M, et al. Clinical-grade whole-genome sequencing and 3′ transcriptome analysis of colorectal cancer patients. Genome Med. 2021;13(1):33.
Swaminathan R, Huang Y, Astbury C, et al. Clinical exome sequencing reports: current informatics practice and future opportunities. J Am Med Inform Assoc. 2017;24(6):1184–91.
Strickler JH, Loree JM, Ahronian LG, et al. Genomic landscape of cell-free DNA in patients with colorectal cancer. Cancer Discov. 2018;8(2):164–73.
Pantel K, Alix-Panabières C. Liquid biopsy and minimal residual disease - latest advances and implications for cure. Nat Rev Clin Oncol. 2019;16(7):409–24.
Yothers G, O’Connell MJ, Lee M, et al. Validation of the 12-gene colon cancer recurrence score in NSABP C-07 as a predictor of recurrence in patients with stage II and III colon cancer treated with fluorouracil and leucovorin (FU/LV) and FU/LV plus oxaliplatin. J Clin Oncol. 2013;31(36):4512–9.
Venook AP, Niedzwiecki D, Lopatin M, et al. Biologic determinants of tumor recurrence in stage II colon cancer: validation study of the 12-gene recurrence score in cancer and leukemia group B (CALGB) 9581. J Clin Oncol. 2013;31(14):1775–81.
Gray RG, Quirke P, Handley K, et al. Validation study of a quantitative multigene reverse transcriptase-polymerase chain reaction assay for assessment of recurrence risk in patients with stage II colon cancer. J Clin Oncol. 2011;29(35):4611–9.
Yamanaka T, Oki E, Yamazaki K, et al. 12-gene recurrence score assay stratifies the recurrence risk in stage II/III colon cancer with surgery alone: the SUNRISE Study. J Clin Oncol. 2016;34(24):2906–13.
Guinney J, Dienstmann R, Wang X, et al. The consensus molecular subtypes of colorectal cancer. Nat Med. 2015;21(11):1350–6.
Isella C, Brundu F, Bellomo SE, et al. Selective analysis of cancer-cell intrinsic transcriptional traits defines novel clinically relevant subtypes of colorectal cancer. Nat Commun. 2017;8:15107.
Ragulan C, Eason K, Fontana E, et al. Analytical validation of multiplex biomarker assay to stratify colorectal cancer into molecular subtypes. Sci Rep. 2019;9(1):7665.
Funding
Open Access funding provided by the IReL Consortium. Funding for this project was provided by the Beaumont Hospital Colorectal Research Fund.
Author information
Authors and Affiliations
Contributions
Study conceptualization and design: J.P. Burke, J.H.M. Prehn, and S.J. Furney
Study methodology: D.A. McNamara, J.P. Burke, and I.S. Reynolds,
Writing (original draft): I.S. Reynolds
Writing (review and editing): all authors
Supervision: J.P. Burke, S.J. Furney, and J.H.M. Prehn
Project administration: I.S. Reynolds
Funding acquisition: D.A. McNamara
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Reynolds, I.S., O’Connell, E., McNamara, D.A. et al. Clinical Oncogenomics and Personalized Medicine in Colorectal Cancer for the Surgeon: What We Need to Know and What the Future Holds. SN Compr. Clin. Med. 4, 70 (2022). https://doi.org/10.1007/s42399-022-01144-x
Accepted:
Published:
DOI: https://doi.org/10.1007/s42399-022-01144-x