Abstract
Low-density polyethylene (LDPE) is the predominant single-use plastic and rarely decomposes after disposal. The primary objective of this study was to identify potential bacteria capable of degrading LDPE plastic and investigating the biochemical pathways of this process. Bacteria were isolated from soil samples collected from a local garbage dumping site in Thailand and tested on their capability to degrade LDPE plastic. Two of the bacteria isolated from the dumping site, Bacillus sp. AS3 and Sphingobacterium sp. AS8, demonstrated 3.06% and 2.01% (w/w) LDPE plastic weight loss over four weeks, respectively. Analysis by FTIR showed that both bacterial strains degraded the LDPE in the region of 3200–3400 cm−1, which represents the OH group in a commercial LDPE polymer. Bacillus sp. AS3 caused the formation of a new range in the carbonyl group (C=O stretch) and the alcohol, carboxylic acid, esters, and ethers group (–C–O stretch). GC–MS analysis revealed various depolymerized compounds, such as alkane, alcohol, and carboxylic compounds, during LDPE degradation by Bacillus sp. AS3. Bacillus sp. AS3 illustrated esterase activity as 0.608 ± 0.004 U/mL after incubation. The proposed schematic of the LDPE biodegrading pathway by Bacillus sp. AS3 relies on the identification of depolymerized molecules as evidence. This suggests that Bacillus sp. AS3 possesses extracellular enzymes that break down LDPE into smaller molecules through depolymerization. Moreover, the surface of LDPE degraded by Bacillus sp. AS3 and Spingobacterium sp. AS8 was marked by cavities and a rough texture when observed under SEM analysis. This study provides microbial applications to reduce plastic pollution by utilising microorganisms to assimilate plastic waste as a carbon source.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
The plastic pollution has emerged as one of the most critical environmental and economic challenges. Microplastic contamination in water, food ingredients, and agricultural products, has resulted in low-quality output [1]. Annually, a staggering 500 billion to 1.5 trillion single-use plastic bags are utilized worldwide, despite their minimal worth in terms of recycling [2]. Low-density polyethylene (LDPE) is frequently employed in the manufacturing of plastic bags and food containers [3]. These plastic bags are the most common type of plastic waste found in landfills, accounting for approximately 69% of the total weight [4]. However, single use plastic bags rarely decompose after disposal, polluting the environment [5], and contaminating the food chain in the form of microplastics [6]. Plastic degradation by microorganisms is a sustainable solution to the environmental problem of plastic waste. Researchers have identified LDPE-degrading bacteria in several genera, including Pseudomonas, Bacillus, and Rhodococcus [7]. This process involves the degradation of plastic into water (H2O) and carbon dioxide (CO2) [8, 9]. However, there are primary constraints facing biodegradation, including the excessive amount of plastic waste and the plastic characteristics that impede microbial attraction, such as surface hydrophobicity, long-chain polymers, high molecular weight (MW), and crystal structure [10]. Hence, the process of plastic biodegradation in an environment is highly time-consuming.
Microorganisms hydrolyse plastic polymers through complex biochemical reactions via extracellular and intracellular depolymerases through the four stages of biodeterioration, biofragmentation, assimilation, and mineralization [11]. Since LDPE is an inert polymer with a high molecular weight and hydrophobicity [12], it requires specific microorganisms to utilize it. Several bacterial strains have been isolated from open environments including the soil of dumping sites and mangrove ecosystem [12,13,14]. Although numerous bacteria can degrade LDPE, there is still a need to study enzymatic capabilities for more rapid and effective degradation. Esterase is an extracellular enzyme capable of hydrolysis of ester bonds present in plastic polymers, resulting in the formation of simpler molecules [15]. After the extracellular enzyme degrades the polymerized chemicals, the cell can assimilate them, and the intracellular enzymes can then convert the intermediate products into compounds that bacteria can use as a carbon source [16]. However, there is currently a scarcity of information regarding the enzymes that degrade LDPE.
Evaluating the biodegradability and potential microbial pathways represents an important advancement in the understanding of plastic waste degradation in nature. It is crucial to emphasize that modifications in plastic chemistry function as the result of interactions between microorganisms and the plastic surface. However, it is also necessary to include the identification of the intermediate products and the microbial enzyme involved in the biodegradation. In this study, bacterial strains were identified from a dumping site and assessed their capability to degrade LDPE. Bacterial activity on LDPE by conducting studies on bacterial survival rates, enzyme activity, changes in LDPE film properties (percent weight loss, surface modifications, and chemical structure change), and the detection of depolymerized products were measured. Further, the biodegradation pathway of LDPE by the isolated bacteria was described. The bacteria in this study exhibit enhanced degradation potential, promising to address environmental challenges associated with plastic waste.
2 Materials and methods
2.1 First bacterial screening and isolation for LDPE biodegradation
Soil and plastic rubbish around 0–5 cm depth were collected from the dumping area of the Asian Institute of Technology (AIT), Thailand, at latitude 14.0785 and longitude 100.6140. The samples were then kept at the Biotechnology Laboratory, Asian Institute of Technology (AIT), Thailand, for further experimental processing. Each sample (10 g) was added to 250 mL of mineral salt broth (MSM). The MSM (1 L) consists of 1 g NH4NO3, 0.2 g MgSO4.7H2O, 1 g K2HPO4, 0.1 g CaCl2.2H2O, 0.15 g KCl, 0.1 g yeast extract, 1 mL of the surfactant tween 80 and 1 mg of each of the following micronutrients: FeSO4.6H2O, ZnSO4.7H2O, MnSO4. The pH of the media was adjusted to neutral (pH 7). LDPE films (obtained from commercial market bags in Thailand) were cut into 10 pieces at a size of 15 × 15 mm, added to the mixture, and incubated in an orbital incubator shaker (Edmund Bühler, Netherland) at 120 rpm for 15 days at 30 °C. Following a 15-day incubation period, 0.1 mL of the enrichment culture was spread onto MSM agar, and nine pieces of LDPE films were then positioned on the plate. Subsequently, the plates were incubated at 30 °C for 7 days, after which the bacteria were isolated. The method was adapted from Sangeetha Devi et al.[17]. All the bacterial isolates were named using AS to stand for Asian Institute of Technology (AIT) strain and followed by the number of the strain, such as AIT strain 1: AS1.
2.2 Second bacterial screening for LDPE degradation
The strains were streaked onto MSM agar containing 0.1% LDPE powder as a source of carbon and then placed in an incubator (Edmund Bühler, Netherland) set at 30 °C for 48 h. Following that, the culture plates showed bacterial growth. The method was adapted from Montazer et al. [18]. Bacterial isolates that exhibited growth on MSM agar supplemented with 0.1% LDPE powder indicated a positive result. Conversely, a bacterial isolate that showed no growth or slow growth (microbial growth on MSM agar containing 0.1% LDPE powder after 48 h is referred to as slow growth) indicated a negative result.
2.3 Biodegradation assay on LDPE films
Three milliliters of each bacteria were inoculated in 300 mL of MSM broth containing 0.3 g of LDPE films (size 5 × 5 mm) and adjusted to a final concentration of 1.5 × 106 CFU/mL. The culture was then incubated at 30 °C in an orbital incubator shaker (Edmund Bühler, Netherland) at 120 rpm for four weeks. Every week, the cell count was determined by the spread plate technique, and cell turbidity (OD600) was observed by using a spectrophotometer (Shimadzu UV-1800, Japan). The control group consists of two samples: one containing MSM supplemented with plastic films but without bacterial inoculation, and MSM with bacterial inoculation but without plastic films. After four weeks of incubation, the microbial biofilms were removed from the LDPE films by immersing them in a 2% (w/v) aqueous sodium dodecyl sulfate (SDS) solution for four hours at 50 °C in water bath (Memmert, Germany) and drying them at 60 °C in an oven (UE700, Memmert, Germany) for six hours for further analysis. Finally, the percentage weight loss of the LDPE films was computed by using the following formula, which compares the initial weights of the preincubated plastic film samples to the inoculated sample. The biodegradation assay was adapted from Montazer et al. [18] and Auta et al. [19].
2.4 Molecular technique of identifying bacteria
Identification of LDPE-degrading bacteria was performed using the molecular technique of reverse transcriptase polymerase chain reaction (RT-PCR) to amplify the 16S rRNA genes [20]. The universal primers used were 27F (5′AGAGTTTGATCM TGGCTCAG 3′) and 1492R (5′TACGGYTACCTTGTTACGACTT 3′). InstaGene™ Matrix was used for DNA extraction, then the PCR reaction was performed using a 20 ng DNA template with 30 µl of EF-Taq (SolGent, Korea) mixture. The PCR amplification occurred under the following conditions: initial denaturation at 95 °C for 2 min, followed by 35 cycles of 95 °C for 1 min, 55 °C for 1 min, 72 °C for 1 min and final extension at 72 °C for 10 min. A multiscreen filter plate (Millipore Corp, USA) was used to purify the amplification products, and sequencing was done with an ABI Prism 3730XL DNA analyzer (Applied Biosystems, USA). The sequences were aligned and then reported in the Basic Local Alignment Search Tool (BLAST) of the National Center for Biotechnology Information (NCBI) GenBank database. Lastly, the phylogenetic tree was created by the MEGA 11 program after multiple sequence alignment.
2.5 Fourier transform infrared spectroscopy (FTIR) analysis
The LDPE films were taken out of the treatment after four weeks of incubation. The films were immersed with 2% (w/v) SDS for four hours at 50 °C in a water bath (Memmert, Germany) and dried (oven model UE700, Memmert, Germany) at 60 °C for six hours. An FTIR spectrometer (Nicolet iS50, Thermo Scientific, USA) was used to measure the formation of the functional groups of alkene, alcohol, aldehyde, ketone, ester, and carboxylic acid. Plastic films were scanned in the 400–4000 cm−1 range and resolution at 4 cm−1 to detect changes in polymer bonding [21].
2.6 Evaluation of biofragmented compounds from plastics by gas chromatography coupled with mass spectroscopy (GC–MS)
GC–MS (GC Sampler 80, Agilent Technologies, USA) analysis of the LDPE films was used to identify the depolymerized products after a four-week incubation. The films were immersed in 2% (w/v) SDS for four hours at 50 °C in a water bath (Memmert, Germany) and dried (oven model UE700, Memmert, Germany) at 60 °C for six hours. Ten mL of chloroform was mixed with 0.5 g of the LDPE films. Then the mixture was ultrasonicated (Ultrasonic Water Bath, Ulangee, China) for two hours at 55 °C. The extract was evaporated at 25 °C before being mixed with 2 mL of chloroform and filtered through a 0.2-micron PTFE syringe filter [22]. With helium as the carrier gas, this filtered sample was examined using GC–MS. The GC was equipped with column HP-5MS 30 m × 0.25 mm × 0.25 µm (Agilent Technologies, USA). The extracted sample (1 µL) was injected at a flow rate of 1 mL/min and maintained at temperature of 300 °C. The oven temperature was raised at a rate of 10 °C/min from 40 °C (hold for 3 min) to 280 °C (hold for 10 min). The peak assignment compared the obtained spectra with the mass spectra libraries, specifically the NIST 14 library.
2.7 Esterase activity
The measurement of esterase activity was conducted after a four-week incubation period. The cell pellet was discarded, and the cell-free supernatant was collected after centrifugation at 5000 rpm for 15 min at 30 °C. The cell-free supernatant was continuously concentrated using an ultra-centrifugal protein concentrator equipped with a 10 K molecular weight cutoff (MWCO) membrane (Thermo Fisher Scientific, US). Esterase assay was performed in 96-well microtiter plates. P-nitrophenyl (PNP) butyrate substrate was dissolved in 1 mL of acetonitrile and the volume was adjusted to 100 mL by adding 0.1 M phosphate buffer at pH 7. The reaction mixture consisted of 0.8 mL of 0.1 M phosphate buffer pH 7.0, 0.134 mL of substrate, and 0.066 mL of the purified supernatant. The mixture was then incubated at 30 °C for 30 min. The control supernatant was used as a blank for non-enzymatic degradation. The 200 µl of the sample was transferred into the 96 well microtiter plate, and the change in colour was detected using a spectrophotometer at a wavelength of 410 nm. A unit was defined as the amount of enzyme required to release 1 µmol of p-nitrophenol per minute at a temperature of 30 °C [23].
2.8 Surface morphology analysis of plastics by field emission scanning electron microscope (FE-SEM)
The FE-SEM (JEOL JSM7800F, Japan) measured the erosion of the LDPE surface. The LDPE films were removed from the culture after four weeks and immersed in 2% (w/v) SDS for four hours at 50 °C in a water bath (Memmert, Germany) and drying (oven model UE700, Memmert, Germany) at 60 °C for six hours. Before SEM analysis, the dried films were coated with a thin gold layer for 45 s on the sample-loaded specimen stub, using a sputtering machine (QUORUM Q150R ES, UK) at a sputtering current of 23 mA. The SEM was used to examine the surface for plastic erosion by Bacillus sp. AS3 and Sphingobacterium sp. AS8 treatment as compared to the control sample (LDPE films without bacterial inoculation).
2.9 Statistical analysis
A completely randomized design (CRD) was used for the total experiments. The experimental data were collected in triplicates and presented as the mean ± standard deviation. SPSS software version 25.0 was used to interpret the statistical analysis. The significant differences (p ≤ 0.05) between the means were used to analyse the variances.
3 Results
3.1 Isolation and screening of LDPE degradable bacteria
In the primary bacterial screening twenty-two strains were identified, however, in the secondary screening only two isolates, strains AS3 and AS8, exhibited growth on MSM agar supplemented with 0.1% LDPE powder, as depicted in Fig. S1. Conversely, the remaining strains demonstrated negative outcomes, characterized by either slow growth or no growth. The morphological differences of the AS3 and AS8 strains were classified and documented in Table S1. AS3 exhibited a white colony with a rod-shaped gram-positive morphology, whereas AS8 exhibited a yellow colony with a rod-shaped gram-negative morphology.
3.2 The number of survive bacteria and the percentage weight loss of LDPE film biodegradation
The number of cells profiled per week is illustrated on a bar graph of the total cell counts and a line graph of absorbance for each bacterial strain AS3 and AS8 is shown in Fig. 1a. The AS3 and AS8 treatments were compared to two control samples consisting of MSM supplemented LDPE without bacterial inoculation and MSM with bacterial inoculation. All the bacterial strains exhibited a progressive increase in both cell count and absorbance when cultivated in the LDPE-supplemented medium for 7 days consecutively. The highest cell count was found (7.86 log CFU/mL, OD600 value 0.59) for AS3 in MSM containing LDPE on the 7th day. This was followed by 7.75, 7.50, and 7.37 log CFU/mL for AS8 in MSM with LDPE, control AS8 in MSM, and control AS3 in MSM, respectively. However, the AS3, AS8 and the control treatment exhibited distinct variations in their cell development patterns between 14 to 28 days of incubation. At 14 to 28 days of incubation, the AS3 and AS8 treatment illustrated 7.79 to 7.38 log CFU/mL and 7.72 to 7.09 log CFU/mL, respectively. In contrast, the control treatment of AS3 and AS8 strains exhibited a decline in total cell count, with a decrease in cell count from 6.86 to 5.98 log CFU/mL and from 6.70 to 5.83 log CFU/mL, respectively. Moreover, the weight loss of LDPE films after four weeks of incubation, as depicted in Fig. S2, demonstrated that the microorganisms utilize LDPE films. The AS3 strain exhibited the most effective reduction in the weight of LDPE films which was recorded as 3.06 ± 0.52%, followed by 2.01 ± 0.20% for AS8.
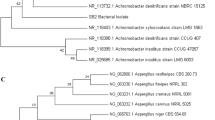
a The bar graph shows total cell counts and cell turbidity (OD600) of the bacterial strains AS3 and AS8 over four weeks in MSM medium containing LDPE films compared to the control sample of MSM containing LDPE without inoculation, the control sample of AS3 in MSM without LDPE, and the control sample of AS8 in MSM without LDPE. Total cell counts and cell turbidity were determined at 0, 7, 14, 21 and 28 days. b Phylogenetic analysis of strains Bacillus sp. AS3 (PP316647) and Sphingobacterium sp. AS8 (PP316667) based on 16S rDNA. A phylogenetic tree was constructed using the MEGA neighbor joining method. The bacterial isolates AS3 and AS8 showed high similarity to Bacillus sp. and Sphigobacterium sp., respectively
3.3 Bacterial identification
The bacterial strains AS3 and AS8 were identified using 16S rRNA gene sequencing in conjunction with phylogenetic tree analysis to determine that AS3 (GenBank accession no. PP316647) and AS8 (GenBank accession no. PP316667) exhibit high similarity to Bacillus toyonensis 97.31% (MG561349.1) and Sphingobacterium sp. 97.85% (KJ411920.1), respectively (Fig. 1b).
3.4 Fourier transform infrared spectrometer (FT-IR) analysis of LDPE biodegradation
After a four-week incubation, the AS3, AS8 and control (without bacterial inoculation) treatments had peaks of C–H stretching and C–H blending (in the range of 2800–3000, 1400–1550, and 650–750 cm−1) which indicates functional groups of alkanes in the LDPE backbone (Fig. 2) [24]. The control samples of LDPE treatment exhibited the range of 3200–3400 cm−1which relates to the peak for OH group, whereas the samples of inoculated bacterial strains AS3 and AS8 exhibited a reduction in the peak intensity in the same range of 3200–3400 cm−1. The AS3 treatment generated new functional groups in the nitro compound (–N=O stretch) range of 1550–1500 cm−1 and the alcohol, carboxylic acid, esters, ethers compound (–C–O stretch) range of 1320–1000 cm−1 which was not observed in the control treatment (Fig. 2a) [25]. Our results indicate that Bacillus sp. AS3 engages in the degrading process of LDPE polymers. This involvement was confirmed through the identification of intermediate compounds using GC–MS analysis.
FTIR spectra of LDPE films after biodegradation over 4 weeks; a represents control treatment (LDPE films without bacterial inoculation) versus the LDPE film inoculated with Bacillus sp. AS3, and b represents the control treatment (LDPE films without bacterial inoculation) versus the LDPE film inoculated with Sphingobacterium sp. AS8
3.5 Depolymerized product analysis by GC–MS and esterase activity
GC–MS was used to detect the depolymerized compounds by Bacillus sp. AS3 compared to a control treatment after a four-week incubation. Both treatments illustrated C8 to C22 alkane compounds including dodecane, pentadecane, hexadecane, 2,6,11,15-tetramethyl, heneicosane, hexadecane, 2,6,10,14-tetramethyl, and docosane (Fig. 3). Different intermediate compounds from Bacillus sp. AS3 treatment, which consisted of different types of compounds such as alcohol compounds (1-dodecanol, 1-Eicosanol, and 1-hexadecanol, 2-methyl), and carboxylic acids (Acetic acid, 3,7,11,15-tetramethyl-hexadecyl ester, hexadecanoic acid, octadecanoic acid, and 7-Z-tetradecen-1-ol acetate) as Fig. 3 and Fig. S3. The percentage area is determined of various intermediate chemicals, as shown in Table 1. Furthermore, our research revealed that following four weeks of incubation, the AS3 treatment exhibited an esterase activity of 0.608 ± 0.004 U/mL as illustrated in Table S2.
3.6 Morphological studies by scanning electron microscope (SEM)
The surface modifications of the LDPE films were examined after undergoing a washing process using a 2% SDS solution. The treatment of Bacillus sp. AS3, Sphingobacterium sp. AS8 and the control treatment (LDPE film without bacterial inoculation) were observed under SEM after four weeks of incubation (Fig. 4). LDPE films inoculated with Bacillus sp. AS3 (Fig. 4b) and Sphingobacterium sp. AS8 LDPE films (Fig. 4c) presented a hole and rougher surfaces than the control (Fig. 4a). Additionally, the presence of bio polysaccharides were discovered on the surface of LDPE, which could have been formed by a biofilm. In contrast, the control sample exhibited a uniformly smooth surface.
4 Discussion
In dumping sites characterized by substantial plastic contamination, the microbial community exhibits the ability to utilize both natural and synthetic substances as a carbon source [26]. Therefore, a local dumping site which contains a substantial amount of plastic waste was invesigated and two bacterial strains that degrade LDPE, Bacillus sp. AS3 and Sphingobacterium sp. AS8, were identified. Studies have recorded the degradation of LDPE by different types of Bacillus species, including Bacillus toyonensis [27]. However, there is limited information available about Sphingobacterium sp. Biodegradation is affected by a range of factors, such as the rate of microbial growth and the carbon source that microbes use [28]. Further, due to its hydrophobic properties and chemical inertness, LDPE has an elevated level of resistance to degradation by preventing water absorption, which restricts microbial activity [29]. Screening of LDPE degrading bacteria in this study found that AS3 and AS8 strains grew on the MSM containing 0.1% LDPE powder medium. Similarly, when comparing the growth rates of different Bacillus species, B. thuringiensis JNU01 showed the highest growth rate on 0.05% polyethylene [30]. This suggests that AS3 and AS8 strains are tolerant to hydrophobic environments. As such, the AS3 and AS8 strains were utilized in the biodegradation assay to further assess the effectiveness of LDPE degradation. After seven days of incubation in MSM broth containing LDPE films, it was observed that the cell counts, and absorbance of all bacterial strains had increased. However, between 14 and 28 days, differences were seen in the cell growth patterns of the AS3, AS8, and control treatment groups. The control treatment exhibited a decreased cell count compared to the AS3 and AS8 treatment groups. The observed disparity could be attributed to the presence of yeast extract within the medium. which enhances bacterial proliferation during the initial week, but afterwards leads to a decline in cell counts within the control treatment group after 14 to 28 days due to a reduction of residual carbon. When exposed to a low-nutrient environment, only active bacteria can grow and display biodegradation by releasing extracellular enzymes that depolymerize the plastic backbone to derive the carbon source [31]. Our study found that Bacillus sp. AS3 had an esterase activity of 0.608 ± 0.004 U/mL after 30 days of incubation. Marine bacterial strains, specifically Marinobacter sp. H-244, Marinobacter sp. H-246, and Bacillus subtilis H-248, also identified the esterase enzyme in the degradation of LDPE [32]. Moreover, AS3 strain demonstrated the most effective reduction in the weight of LDPE films at 3.06%, followed by 2.01% for AS8 within 28 days. The AS3 and AS8 strains exhibited comparable LDPE degradation to previously studied organisms when considering the parameter of total weight loss. After 30 days of degradation, B. subtilis ATCC6051 and B. licheniformis ATCC14580 showed 3.49% and 2.83% of LDPE weight loss, respectively [33]. Rhodococcus ruber exhibited an LDPE weight decrease of 2.5% [34]. Pseudomonas aeruginosa (PAO1) strain (B1), Pseudomonas aeruginosa (ATCC) strain (B2), Pseudomonas putida strain (B3), and Pseudomonas syringae strain (B4) also showed 20%, 11%, 9%, and 11.3% weight loss after 120 days [35]. The cavities and roughness was observed on the LDPE surfaces under electron microscopy after incubation with AS3 and AS8.
FTIR and GC–MS are employed as analytical methods for the identification of intermediates in LDPE biodegradation. The bacterial strains AS3 and AS8 exhibited a reduction in the peak of the OH group in LDPE films within the range of 3200 – 3400 cm−1. Similar changes were found to disappear in the peak of CHO stretch (2,660 cm−1) on LDPE after incubation with Bacillus amyloliquefaciens BSM-2 and BSM-1 [36]. Hence, AS3 created a new range of nitro compounds (–N=O stretch) within the spectral range of 1550–1500 cm−1, and alcohol, carboxylic acid, esters, and ether compounds (–C–O stretch) within the range of 1320–1000 cm−1, which were not observed in the control treatment. Similarly, Bacillus licheniformis SARR1, which is capable of degrading LDPE, formed new functional groups of C=C stretching, O–H stretching, CHO stretching, and C=O stretching at peaks in the range of 1500–1540 cm−1 and 1000–1100 cm−1 [37]. This suggests that AS3 and AS8 strains are involved in the change of chemical structure of the LDPE film during degradation. The biodegradation process of LDPE by microorganisms involves a series of intricate reactions which produce several intermediate products. Our study found the presence of C8 to C22 alkane compounds such as dodecane, pentadecane, hexadecane, 2,6,11,15-tetramethyl, heneicosane, hexadecane, 2,6,10,14-tetramethyl, and docosane which alkane compounds from C19 to C30 can be used by microorganisms as a sole carbon source [38, 39]. Moreover, alcohol compounds (1-dodecanol, 1-eicosanol, 1-hexadecanol, and 2-methyl) and carboxylic acid compounds (acetic acid, 3,7,11,15-tetramethyl-hexadecyl ester, hexadecenoic acid, octadecanoic acid, and 7-Z-tetradecen-1-ol acetate) were found in the AS3 treatment. Marinobacter sp. (H-244 and H-246) and Bacillus subtilis (H-248) also produced intermediate compounds such as alkane, alkene, carboxylic acid, and ester during the degradation of LDPE [32]. Fatty acids, including tri-decanoic and octa-decanoic acids, were discovered after LDPE incubation with consortia of Brevibacillus sp. and Aneurinibacillus sp. for 140 days [40]. This implies that AS3 facilitates alkane degradation to carboxylic acid.
Figure 5 illustrates the proposed biodegradation pathway of LDPE by Bacillus sp. AS3 in which extracellular enzymes facilitate the oxidation of alkane compounds C8 to C22 (hexadecane and octadecane) by introducing a hydroxyl (–OH) group to alkane compounds. This oxidation reaction results in the production of alcohols such as 1-hexadecanol. The primary alcohols are further oxidized to an aldehyde and a carboxylic acid (hexadecanoic acid and octadecanoic acid). The final step is the assimilation of the carboxylic acid compounds into the cells, which can then be utilized in cellular respiration via the TCA cycle, hence facilitating energy generation for bacterial cells. Therefore, AS3 was found to induce the release of these intermediate chemicals during the degradation process of biodegradation and utilize these compounds to generate energy through cellular respiration.
5 Conclusion
Plastic degradation by microorganisms is a time-consuming process. Therefore, it is essential to locate microorganisms with great efficiency in plastic degradation. Moreover, comprehending the function of microbial enzymes and the LDPE degradation pathway can contribute to bridging existing gaps in knowledge. The present study isolated Bacillus sp. AS3 and Sphingobacterium sp. AS8 from the dumping site. Bacillus sp. AS3 shows the greatest potential as an LDPE degrader based on the results of cell growth, the high percentage of LDPE weight loss, microscopic damage to LDPE film, and the observed intermediates of LDPE degradation. Moreover, the intermediate products, which revealed alkanes, alcohols, and carboxylic acid compounds, were used to propose the biodegradation mechanisms of LDPE by Bacillus sp. AS3. Bacillus sp. AS3 has the potential to be upscaled for the degradation of LDPE, offering sustainable and an affordable method for the future development of plastic waste bioremediation.
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information. Bacillus sp. AS3 sequences data in the paper was deposited into the National Center for Biotechnology Information (NCBI) GenBank database accession number PP316647 at the following URL: https://www.ncbi.nlm.nih.gov/nuccore/PP316647. Sphingobacterium sp. AS8 sequences data in the paper was deposited into the National Center for Biotechnology Information (NCBI) GenBank under accession number PP316667 at the following URL: https://www.ncbi.nlm.nih.gov/nuccore/PP316667.
References
Rossatto A, Arlindo MZF, de Morais MS, de Souza TD, Ogrodowski CS. Microplastics in aquatic systems: a review of occurrence, monitoring and potential environmental risks. Environ Adv. 2023. https://doi.org/10.1016/J.ENVADV.2023.100396.
Vassanadumrongdee S, Hoontrakool D, Marks D. Perception and behavioral changes of Thai youths towards the plastic bag charging program. Appl Environm Res. 2020. https://doi.org/10.35762/AER.2020.42.2.3.
Li P, Wang X, Su M, Zou X, Duan L, Zhang H. Characteristics of plastic pollution in the environment: a review. Bull Environ Contam Toxicol. 2021. https://doi.org/10.1007/s00128-020-02820-1.
Mohanan N, Montazer Z, Sharma PK, Levin DB. Microbial and enzymatic degradation of synthetic plastics. Front Microbiol. 2020. https://doi.org/10.3389/fmicb.2020.580709.
Ghatge S, Yang Y, Ahn JH, Hur HG. Biodegradation of polyethylene: a brief review. Appl Biol Chem. 2020. https://doi.org/10.1186/s13765-020-00511-3.
Uwamungu JY, Wang Y, Shi G, Pan S, Wang Z, Wang L, Yang S. Microplastic contamination in soil agro-ecosystems: a review. Environ Adv. 2022. https://doi.org/10.1016/J.ENVADV.2022.100273.
Montazer Z, Najafi MBH, Levin DB. Challenges with verifying microbial degradation of polyethylene. Polymers. 2020. https://doi.org/10.3390/polym12010123.
Emisha L, Wilfred N, Kavitha S, Halder G, Haldar D, Patel AK, Singhania RR, Pandey A. Biodegradation of microplastics: advancement in the strategic approaches towards prevention of its accumulation and harmful effects. Chemosphere. 2023;9:140661. https://doi.org/10.1016/J.CHEMOSPHERE.2023.140661.
Akinsemolu AA. Principles of green microbiology: the microbial blueprint for sustainable development. Environ Adv. 2023. https://doi.org/10.1016/J.ENVADV.2023.100440.
Restrepo-Flórez JM, Bassi A, Thompson MR. Microbial degradation and deterioration of polyethylene—a review. Int Biodeterior Biodegrad. 2014. https://doi.org/10.1016/J.IBIOD.2013.12.014.
Kumar Sen S, Raut S. Microbial degradation of low density polyethylene (LDPE): a review. J Environ Chem Eng. 2015. https://doi.org/10.1016/J.JECE.2015.01.003.
Mathur G, Mathur A, Prasad R. Colonization and degradation of thermally oxidized high-density polyethylene by Aspergillus Niger (ITCC No. 6052) isolated from plastic waste dumpsite. Bioremed J. 2011;15(2):69–76. https://doi.org/10.1080/10889868.2011.570281.
Oluwole OA, Oluyege JO, Olowomofe TO. Biodegradation of polyethylene based films used in water packaging by dumpsite bacteria. Bioremediat. 2022. https://doi.org/10.1080/10889868.2022.2087591.
Singh P, Nagarajan A, Chua KO, Ting ASY. Biodegradation of low-density polyethylene by novel halophilic bacteria from mangrove ecosystem. Bioremediat. 2023. https://doi.org/10.1080/10889868.2023.2297181.
Buchholz PCF, Feuerriegel G, Zhang H, Perez-Garcia P, Nover L, Chow J, Streit WR, Pleiss J. Plastics degradation by hydrolytic enzymes: the plastics-active enzymes database—PAZy. Proteins Struct Funct Bioinform. 2022. https://doi.org/10.1002/prot.26325.
Amobonye A, Bhagwat P, Singh S, Pillai S. Plastic biodegradation: frontline microbes and their enzymes. Sci Total Environ. 2021. https://doi.org/10.1016/j.scitotenv.2020.143536.
Sangeetha Devi R, Ramya R, Kannan K, Robert Antony A, Rajesh KV. Investigation of biodegradation potentials of high density polyethylene degrading marine bacteria isolated from the coastal regions of Tamil Nadu India. Mar Pollut Bull. 2019. https://doi.org/10.1016/j.marpolbul.2018.12.001.
Montazer Z, Habibi-Najafi MB, Mohebbi M, Oromiehei A. Microbial degradation of UV-pretreated low-density polyethylene films by novel polyethylene-degrading bacteria isolated from plastic-dump soil. J Polym Environ. 2018. https://doi.org/10.1007/s10924-018-1245-0.
Auta HS, Emenike CU, Fauziah SH. Screening of Bacillus strains isolated from mangrove ecosystems in Peninsular Malaysia for microplastic degradation. Environ Pollut. 2017. https://doi.org/10.1016/J.ENVPOL.2017.09.043.
Church DL, Cerutti L, Gürtler A, Griener T, Zelazny A, Emler S. Performance and application of 16S rRNA gene cycle sequencing for routine identification of bacteria in the clinical microbiology laboratory. Clin Microbiol Rev. 2020. https://doi.org/10.1128/CMR.00053-19.
Gajendiran A, Krishnamoorthy S, Abraham J. Microbial degradation of low-density polyethylene (LDPE) by Aspergillus clavatus strain JASK1 isolated from landfill soil. 3 Biotech. 2016. https://doi.org/10.1007/s13205-016-0394-x.
Kyaw BM, Champakalakshmi R, Sakharkar MK, Lim CS, Sakharkar KR. Biodegradation of low density polythene (LDPE) by Pseudomonas species. Indian J Microbiol. 2012. https://doi.org/10.1007/s12088-012-0250-6.
Khandare SD, Chaudhary DR, Jha B. Marine bacterial biodegradation of low-density polyethylene (LDPE) plastic. Biodegrad. 2021. https://doi.org/10.1007/s10532-021-09927-0.
Jung MR, Horgen FD, Orski SV, Rodriguez CV, Beers KL, Balazs GH, Jones TT, Work TM, Brignac KC, Royer SJ, Hyrenbach KD, Jensen BA, Lynch JM. Validation of ATR FT-IR to identify polymers of plastic marine debris, including those ingested by marine organisms. Mar Pollut Bull. 2018. https://doi.org/10.1016/j.marpolbul.2017.12.061.
Soleimani Z, Gharavi S, Soudi M, Moosavi-Nejad Z. A survey of intact low-density polyethylene film biodegradation by terrestrial actinobacterial species. Int Microbiol. 2021;24:65–73. https://doi.org/10.1007/s10123-020-00142-0/Published.
Maclean J, Mayanna S, Benning LG, Horn F, Bartholomäus A, Wiesner Y, Wagner D, Liebner S. The terrestrial plastisphere: diversity and polymer-colonizing potential of plastic-associated microbial communities in soil. Microorg. 2021. https://doi.org/10.3390/microorganisms9091876.
Ndahebwa Muhonja C, Magoma G, Imbuga M, Makonde HM. Molecular characterization of low-density polyethene (LDPE) degrading bacteria and fungi from Dandora dumpsite, Nairobi Kenya. Int J Microbiol. 2018. https://doi.org/10.1155/2018/4167845.
Cai Z, Li M, Zhu Z, Wang X, Huang Y, Li T, Gong H, Yan M. Biological degradation of plastics and microplastics: a recent perspective on associated mechanisms and influencing factors. Microorg. 2023. https://doi.org/10.3390/microorganisms11071661.
Elahi A, Bukhari DA, Shamim S, Rehman A. Plastics degradation by microbes: a sustainable approach. J King Saud Univ-Sci. 2021;33(6):101538. https://doi.org/10.1016/j.jksus.2021.101538.
Do YS, Lee CO, Kim HW, An SJ, Kim S, Seo MJ, Park C, Yun CH, Chi WS, Yeom SJ. Exploring a new biocatalyst from Bacillus Thuringiensis JNU01 for polyethylene biodegradation. Environ Sci Technol Lett. 2023. https://doi.org/10.1021/acs.estlett.3c00189.
Lens-Pechakova LS. Recent studies on enzyme-catalysed recycling and biodegradation of synthetic polymers. Adv Ind and Eng Polym Res. 2021. https://doi.org/10.1016/J.AIEPR.2021.06.005.
Khandare SD, Agrawal D, Mehru N, Chaudhary DR. Marine bacterial based enzymatic degradation of low-density polyethylene (LDPE) plastic. J Environ Chem Eng. 2022. https://doi.org/10.1016/j.jece.2022.107437.
Yao Z, Seong HJ, Jang YS. Degradation of low density polyethylene by Bacillus species. Appl Biol Chem. 2022;65(1):84. https://doi.org/10.1186/s13765-022-00753-3.
Santo M, Weitsman R, Sivan A. The role of the copper-binding enzyme—laccase—in the biodegradation of polyethylene by the actinomycete Rhodococcus ruber. Int Biodeterior Biodegrad. 2013. https://doi.org/10.1016/j.ibiod.2012.03.001.
Kyaw BM, Champakalakshmi R, Sakharkar MK, Lim CS. Biodegradation of low density polythene (LDPE) by Pseudomonas species. Indian J Microbiol. 2012. https://doi.org/10.1007/s12088-012-0250-6.
Das MP, Kumar S. An approach to low-density polyethylene biodegradation by Bacillus amyloliquefaciens. 3 Biotech. 2015. https://doi.org/10.1007/s13205-014-0205-1.
Rani R, Rathee J, Kumari P, Singh NP, Santal AR. Biodegradation and detoxification of low-density polyethylene by an indigenous strain Bacillus licheniformis SARR1. J Appl Biol Biotechnol. 2022;10:9–21. https://doi.org/10.7324/JABB.2021.100102.
Al-Kaabi N, Al Disi Z, Al-Ghouti MA, Solling TI, Zouari N. Interaction between indigenous hydrocarbon-degrading bacteria in reconstituted mixtures for remediation of weathered oil in soil. Biotechnol Rep. 2022;1(36):e00767. https://doi.org/10.1016/J.BTRE.2022.E00767.
Niu L, Chen Y, Li Y, Wang Y, Shen J, Wang L, Zhang W, Zhang H, Zhao B. Diversity, abundance and distribution characteristics of potential polyethylene and polypropylene microplastic degradation bacterial communities in the urban river. Water Res. 2023. https://doi.org/10.1016/J.WATRES.2023.119704.
Skariyachan S, Patil AA, Shankar A, Manjunath M, Bachappanavar N, Kiran S. Enhanced polymer degradation of polyethylene and polypropylene by novel thermophilic consortia of Brevibacillus sps and Aneurinibacillus sp screened from waste management landfills and sewage treatment plants. Polym Degrad Stab. 2018. https://doi.org/10.1016/j.polymdegradstab.2018.01.018.
Funding
No funding was received for conducting this study.
Author information
Authors and Affiliations
Contributions
Pornya Khampratueng: Experiment, edit the manuscript, statistic, graphs and tables Daniel Lee Rice: Review the manuscript, Experiment Anil Kumar Anal: Review the manuscript, Experiment
Corresponding author
Ethics declarations
Competing interests
All authors certify that they have no affiliations with or involvement in any organization or entity with any financial interest or non-financial interest in the subject matter or materials discussed in this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Khampratueng, P., Rice, D. & Anal, A.K. Biodegradation of low-density polyethylene by the bacterial strains isolated from the dumping site community. Discov Appl Sci 6, 348 (2024). https://doi.org/10.1007/s42452-024-06052-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42452-024-06052-4