Abstract
Reactive nitrogen (N) plays a pivotal role in supplying N to plants and soil microbes, but it has negative environmental impacts through influencing the quality of water and air, which in turn influences human health. Thus, there is an urgent necessity to maximize N benefits while reducing the negative impacts on the environment. Improving crop N use efficiency (NUE) is required for environmental conservation and agricultural sustainability. Thus, the pivotal objective of this article is to introduce the modern developments and imminent prospects of improving crops NUE using various complementary methods. Here, the approaches of site-specific N management, use of synthetic and biological nitrification inhibitors, microbial nitrate (NO3−) immobilization stimulation, and stimulation of the dissimilatory nitrate reduction to ammonium (DNRA), adopting agroforestry system, breeding techniques, quantitative trait loci (QTL) mapping, omics approaches, and potential new targets and overexpression of N-related genes were presented as effective approaches to improving NUE. Optimal rate, time, application methods, using specially formulated forms of fertilizer, and using nitrification inhibitors are the most agricultural practices linked with improving NUE. The fertilizer recommendations could be often justified across the field rather than a uniform application using spatial variability of nutrient content. Restoring soil NO3− retention capacity and adopting agroforestry system can also be promising ways to improve NUE. Furthermore, the use of genetic modification or the development of new cultivars that use N more efficiently is critical. In addition, omics data, including transcriptomics and metabolomics, not only advance our current understanding of N reactions but also help us move towards strategies, which are more effective in improving NUE and enhancing crop production. In conclusion, this article strongly advocates the use of integrated approaches with a private insight of genetics and agricultural management when managing N.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Reactive nitrogen (Nr; all species of nitrogen (N) except N2) plays a pivotal role in supplying N to plants and soil microbes, but it has negative environmental impacts through influencing the quality of water and air, which in turn influences human health (Galloway and Cowling 2021). Thus, there is an urgent necessity to maximize N benefits while reducing the negative impacts on the environment. However, managing N dynamics is a challenge, especially with the massive increase in the population followed by a huge increase in food demand, leading to intensification of agricultural practices and consequently excessive use of inorganic N fertilizers (Zhang et al. 2019). The global use of inorganic N fertilizers increased from 12.0 Tg (1012 g) year−1 in 1961 to 108 Tg year−1 in 2019 (IFA 2022). It is estimated that by 2050, 70–100% growth in overall agricultural production will be urgently needed to meet the food demand of the world’s nine billion people (Godfray et al. 2010). This will undoubtedly require the use of more inorganic N fertilizers. In Africa, for example, Elrys et al. (2020a) reported that increasing total N inputs from 35 to 181 kg N ha−1 year−1 during 2016–2050 is required to achieve food self-sufficiency. However, excessive N fertilization results in low N use efficiency (NUE). The hardly increase in crop yields under the drastic increment in N fertilizer demonstrates a vast reduction in NUE globally. In 2014, Lassaletta et al. (2014) found that only 47% of the fertilized N globally is transformed into agricultural commodities, compared to 68% in 1960. This was also consistent with the findings of Yan et al. (2020) in cereal cropping systems on a global scale. Table 1 introduces examples for different crops’ annual NUE from different countries (Norton et al. 2015). Moreover, Ti et al. (2015) reported that 19–31% of the annual N inputs (1.2–1.9 Tg N year−1) in vegetable production of China accumulated in soil. Recently, Elrys et al. (2020a) reported that NUE would decline from 63 to 50% in Africa during 2010–2050 if African countries followed the recommended rate of 181 kg N ha−1 year−1 to achieve food self-sufficiency. They also found that total N surplus would increase from 13 to 90 kg N ha−1 year−1 during the same period, while soil gaseous emissions would increase from 8.0 to 61 kg N ha−1 year−1. These increases in N losses and decreases in NUE may lead to adverse environmental effects like soil acidification, underground water pollution with leaching nitrate (NO3−), and greenhouse gas (GHG) emissions (Yu et al. 2015). These large N losses are believed to take place in terrestrial ecosystems that are saturated with N, that is, where the supply of N exceeds the actual requirement for microbes and plants (Butterbach-Bahl et al. 2013; Ramm et al. 2022). Decreased synchronization between N application and demand for crop plants leads to lower NUE (Abebe and Feyisa 2017).
At different spatial and temporal scales, NUE indicators are widely utilized by agronomists, biogeochemists, environmental scientists, and policymakers to estimate the efficiency and potential environmental implications of N use in crop production systems (EU Nitrogen Expert Panel 2015; Quan et al. 2021). The NUE is a useful indicator of the environmental and economic performance of production systems because it is an indicator of the balance between costs and benefits of primary production systems (Hutchings et al. 2020). Through adopting a mix of agricultural technologies and practices, enhancing NUE can be accomplished. This target includes using enhanced crop varieties, creating motivations for enhanced N management, shifting fertilizer practices and technologies, and following the 4Rs of N fertilizer application: right type, right rate, right timing, and right placement (Zhang et al. 2015; Houlton et al. 2019). Although these improving agricultural practices and developing appropriate environmental policies, which have been well shown in several previous review articles, have been revealed as effective strategies, the presentation of other solutions to control soil N transformation that increase or decrease N loss is required. For example, dissimilatory nitrate reduction to ammonium (DNRA) converts NO3− to NH4+ in soils providing additional NH4+ for assimilation and uptake by microbes and plants (Pandey et al. 2020). In Chilean temperate rainforests, DNRA consumes approximately all soil NO3−, conserving N in soil–plant system (Huygens et al. 2007). More recently, Cheng et al. (2022a, b) suggested that promoting soil N retention through DNRA by high C inputs and low N inputs may help to reducing fertilizer-N rates and N2O emissions, while improving NUE. Similarly, Wang et al. (2021a) reported that organic amendments with high carbon (C) to N ratio could stimulate NO3− immobilization without causing high N losses via denitrification in an upland soil. In addition, previous studies demonstrated that the deep trees rooting in agroforestry systems can catch and recycle subsoil inorganic N that have leaked below the rooting zone of associated croplands, resulting in a more efficient interception of N leached (Lang et al. 2019). Moreover, studies revealed that biological nitrification inhibitors are a low-cost and environmentally friendly compared to synthetic nitrification inhibitors. Molecular investigations also have provided joint validation of the physiological conceptual framework for NUE of crops. In addition, breeding more N-efficient crops can be a motivating alternative to increase crop production with low doses of inorganic N fertilizer (Raviv 2010; Hirel and Lea 2011). Therefore, these modern alternative cultivation techniques and genetic modification can also be an attractive way to improve NUE of crops. However, to our knowledge, review articles describing the potential use of these integrated approaches (e.g., N retention stimulation, agroforestry system, and use of biological nitrification inhibitors) to enhance NUE while shedding more light on the role of genetics remain limited.
This review article represents the recent alternative approaches for improving crops NUE using a variety of complementary approaches comprising molecular genetics, conventional breeding, and alternative farming performances (e.g., N retention stimulation, agroforestry system, and use of biological nitrification inhibitors), in addition to a brief presentation of the other measures used.
2 Global Patterns of Inorganic N Fertilizer Use
Agricultural fertilizer usage was considered the magic practice for alleviating N limitations of optimizing soil fertility and maximizing crop yield during the past century (Lu and Tian 2017). Mainly, in the early twentieth century, a massive amount of unreactive N has been converted to inorganic N fertilizer using Haber–Bosch process (Galloway et al. 2004), which caused a dramatic increment in agricultural production and alleviated global hunger (Erisman et al. 2008). Accelerated increment in inorganic N fertilizers usage was reported globally as shown in Fig. 1 as inorganic N fertilizer consumption increased 813% over the last five decades. However, this increase was 35% and 555% in developed and developing counties, respectively (Fig. 2). East and South Asia were the largest consuming regions, representing together 53.7% of the global inorganic N consumption, followed by North America (13.5%), Western and Central Europe (10.9%), Latin America (8.30), Eastern Europe and Central Asia (5.40), Africa (3.63%), and West Asia (3.05%), respectively (Fig. 1). Although the lowest inorganic N consumption was recorded in Oceania (1.74%), it recorded an increase in inorganic N consumption of 4293% in 2019 compared to 1961. The highest rate of increase (5493%) in inorganic N consumption over the past five decades has been recorded in South Asia. Whereas, the lowest rate of increase has been recorded in North America (364%) and Eastern Europe and Central Asia (576%) as a result of significant improvements in NUE and growing the manure-N recycling. Furthermore, total N fertilizer applied to cereals reached 57.5 Tg N, i.e., more than half of the world utilized N fertilizer (Heffer and Prud’homme 2016). The three main kinds of cereals including wheat, rice and maize accounted for 52.7% of the global use of N fertilizer, while all other cereals accounted for 4.80% (Fig. 3). Moreover, oilseeds recorded 7.80%. Roots and tubers (mostly potatoes), sugar crops (mostly sugar cane) and fiber crops (mostly cotton) recorded 2.10%, 3.60%, and 4.10%, respectively, of global N fertilizer usage, while vegetables and fruits recorded together 13.5% (Heffer and Prud’homme 2016).
The use of synthetic N fertilizers across the years throughout the world regions from 1960 to 2019 (data were extracted from IFA 2022)
Estimates of synthetic N fertilizer use by crop category at the global scale (data were extracted from Heffer and Prud’homme 2016)
3 NUE Indices
To estimate the efficiency of N use in crop production and its potential environmental influences, NUE indicators are widely utilized. The main approaches for identifying and measuring NUE include N balance, N difference, and 15 N tracer (Quan et al. 2021). In the N balance approach, the data of N input and N output are easily utilized to compute NUE according to the following formula: NUE = N output/N input (EU Nitrogen Expert Panel 2015; Quan et al. 2021). In the N difference approach, the NUE is calculated by dividing the difference between the harvested N in fertilized plots (treatment) and unfertilized plots (control) by total inputs of N fertilizer (Quan et al. 2021). In the 15 N tracing approach, the fertilizer is applied with 15 N tracer and then the 15 N proportion of the yield in the crop product is traced (Congreves et al. 2021). However, the results of these methods are often different, which may lead to confusion or misinterpretation by stakeholders and may hinder comparisons between studies across different regions (EU Nitrogen Expert Panel 2015). In China, for example, Quan et al. (2021) reported that NUE calculated by N difference and 15 N tracer approaches ranged between 0.27 and 0.37, while NUE values determined by N balance approach is around 0.68, indicating a very different picture of crop N management to various stakeholders. The three approaches are used to assess how much N input to the system is removed from the system as crop products, but in practice, they have different experimental settings and are likely to produce inconsistent results.
The mass balance of inputs and N outputs of agricultural systems is more complex because a large portion of N is dissipated into water bodies or the atmosphere or accumulates in the system (temporarily), while only a small portion of N input into agricultural systems ends up in the products (Congreves et al. 2021). The output of N is utilized in crops that are harvested and removed from the soil to express the NUE of crop system and also for livestock production systems (Quan et al. 2021). While, both livestock and crop products are involved in the mixed systems (EU Nitrogen Expert Panel 2015; Quan et al. 2021). By increasing the N output of harvested products and/or reducing the N input, the NUE will increase depending on the system and its management. NUE does not characterize interior N transformation pathways in a system (e.g., N nitrification or mineralization), nor quantitatively estimate N loss from the system directly, as the unremoved N of biomass might remain within the soil (Congreves et al. 2021). For long-run, however, changes in soil N storage are usually related weakly with inputs and outputs, and thus, lower NUE values over several years are probably incidental indicators for significant losses of N to the environment (Norton et al. 2015; Quan et al. 2021). Availability of data generally at both national and farm levels give this NUE definition an important advantage. On the farm level, the N content in harvested products and manure are not often reported, but they can be projected from regional literature reports. Nationally and locally, production data by commodity type (e.g., wheat, rice, maize, other crops, meat, and dairy products) and total apparent fertilizer consumed by country are interpreted by authorities and the FAO reports when actual data are not available (Norton et al. 2015). Efforts have been made by the scientific society to differentiate national data according to crop-specific application rates through agricultural areas (e.g., Potter et al. 2011), that could be used as reference or standards values. IFA has announced reports on using fertilizer by country and crop from collected data (Heffer 2013), but these do not include all countries. Even though there are some data uncertainties and limitations, outputs and inputs could be assessed locally and nationally, and thus, NUE can be obtained from those estimates. Recently, Quan et al. (2021) suggested that the averages of the NUE calculated by the N balance approach over several years tend to be stable under proportionate management practices, notwithstanding the fluctuation caused by year-to-year yield variation, because the changes of mean annual soil N stock over a long period can be neglected compared with the annual N fertilizer inputs (Yan et al. 2014).
The N difference approach considers background soil N contents by accounting for N uptake or production in unfertilized plots (control) and is thus suitable for research contexts. Although this is better than ignoring the effect of background N contents on the crop N response (as in the N balance approach), it can present misleading NUE values if soil N reserves are depleted in unfertilized plots over long-term periods (Congreves et al. 2021; Quan et al. 2021). In addition, this approach ignores the fact that plants mediate their response to N (Congreves et al. 2021) and have different strategies to deal with N limitation through the formation and recruitment of microbial communities of N-cycling (Moreau et al. 2019), manipulation of microbial functional groups (Blagodatskaya et al. 2009), and improving their competition with bacteria for N (Legay et al. 2020). Although the N difference approach is a simple and cost-effective method to evaluate the short-term response of harvested N to N fertilization under various management or environments conditions, it is often hard to conduct in practice because it is difficult to persuade farmers to allocate a part of their land as an unfertilized control plot because this will lead to a decrease in productivity and income (Quan et al. 2021).
The 15 N tracing approach is utilized to trace N flow derived from fertilizer, soil, or biologically fixed N within cropping systems. Using 15 N-labeled fertilizers, it is possible to determine the recovery of fertilizer into plant components, and to estimate the N quantity derived from fertilizer in one growing season (Taveira et al. 2020). By knowing the proportion of N derived from the fertilizer, the proportion of N attributable to the indigenous soil can also be determined (Congreves et al. 2021). Unlike the N balance and N difference approaches, the 15 N tracing approach offers a direct measure of recovery, rather than an apparent recovery (Congreves et al. 2021). Moreover, using this approach it is possible to study other 15 N labeled N sources such as crop residue-N and biological N fixation (Taveira et al. 2020). To improve our understanding of soil–plant N cycling, therefore, implementation of multi-year 15 N monitoring to provide data on long-term use and N recovery is required. However, the proportion of cumulative N derived from fertilizer will likely increase where more 15 N will be recovered as a result of the pulse application of 15 N tracer (Quan et al. 2021). Moreover, the high cost of 15 N materials and measurements make this approach usually applicable at relatively small space–time scales. The NUE estimates by the N difference and 15 N tracing approaches are likely to increase when the observation is made for extended periods. The NUE estimates from the N difference and 15 N tracing approaches tend to converge with that from the N balance method for long-term observations (Quan et al. 2021). Consequently, the N balance approach provides great advantages in assessing the N inputs performance in crop production, especially when the mean change in soil N stock is negligible compared with the annual N inputs.
4 How Can NUE Be Managed?
The major objective of agricultural production is to efficiently meet food security needs, ecosystem services, and sustainability (Foley et al. 2011; Houlton et al. 2019). This underscores the importance of integrated approaches, which offer powerful tools for controlling crop productivity in parallel with improving the efficiency of N use (Sinclair and Rufty 2012; Houlton et al. 2019). There is an urgent need for reliable environmentally acceptable agronomic management practices to improve NUE and crop productivity while reducing N environmental risks of using fertilizers. NUE can be improved by adopting a combination of agricultural practices and techniques as discussed below.
4.1 Site-Specific N Management
Site-specific N management is one of a number of strategies that used to improve NUE and minimize N losses to the environment (Argento et al. 2021; Pilar et al. 2018). It is a concept that includes field N management strategies plus metric knowledge of field-specific variability of expected soil N supplying power and crop N requirement (Giller et al. 2004). The use of site-specific N management increased corn NUE by 43%-50% (Pilar et al. 2018). Site-specific N management improved NUE in winter wheat by 10% (Argento et al. 2021). Improving NUE can be accomplished by following the 4Rs of N fertilizer application: right rate, right type, right placement, and right timing (Zhang et al. 2015; Houlton et al. 2019). These 4Rs are the most problematic agricultural practices linked with N management, but they are also easily relative resolvable issues. Recently, Liang et al. (2021) reported that the application of an appropriate N rate (210–260 kg N ha−1) remarkably increased the NUE of rice compared to the high rates (315 and 420 kg N ha−1). In the North China Plain, it was suggested that N addition rate of more than 240 kg N ha−1 was not beneficial for wheat growth and production (Si et al. 2020). The meta-analysis of Ma et al. (2020) suggested that the optimum N application rate for winter wheat was 188 kg N ha−1 in areas with mean annual temperature (MAT) of > 10 ℃, and it was 225 kg N ha−1 when MAT is ≤ 10 ℃, whereas the optimum N application rate was 250 kg N ha−1 in the area with mean annual precipitation (MAP) of > 600 mm, and it was 235 kg N ha−1 when MAP is ≤ 600 mm. They also reported that the optimum N application rate was 226 kg N ha−1 under the topsoil organic matter content of ≤ 12 g kg−1, and it was 163 kg N ha−1 when the topsoil organic matter content is > 12 g kg−1. Belete et al. (2018) found that the NUE of wheat plants reduced with increasing N rate (120–360 kg N ha−1), manifesting poor use of N, and that the split N application (¼ at sowing, ½ at tillering, and ¼ at booting) resulted in higher NUE. The source of applied N also has a vital role in monitoring diverse N losses, affecting N recovery and availability (Elrys et al. 2018). Nitrate fertilizers, for example, are subject to be leached as compared with NH4+ and amide-containing N fertilizers. In contrast, amide and NH4+ fertilizers are more liable to volatilization losses comparing with NO3−-containing N fertilizers (Yadav et al. 2017). Recently, Li et al. (2021) found that ammonium sulfate contributed to higher NUE of wheat plants compared to urea, ammonium nitrate, and calcium nitrate, while urea contributed to higher NUE in maize. Furthermore, it has been pointed out that placing N fertilizer near the plant roots can dramatically grow yield up, and at the same time allows the farmer to minimize N rates (Vossenkemper 2010). The applied N in the middle of the row is more readily for weeds consumption against crops use (Blackshaw and Brandt 2008) so it is preferred to banding N near the row for increasing crop uptake and thus increasing the crop a competitive advantage against the weeds (Blackshaw et al. 2002). Urea-N should place beneath the soil not on its surface to reduce N loss by volatilization (Wessels and Coetzee 2017). If urea is band placed into wetted soil and the soil is then sealed, the risk of volatilization is reduced. The splitting of N fertilizer at three equal doses (before planting, during planting, and after planting) has been proven to give the best yield (Nemati and Sharifi 2012). By redistributing and decreasing the quantity of N fertilizer applied, NUE was enhanced around 10% (Argento et al. 2021). Ammonia (NH3) volatilization was reduced by 12.5% with each additional cm of incorporation depth, but was negligible when incorporating at depths greater than 7.5 cm (Rochette et al. 2013). Similarly, Nasielski et al. (2020) found that NH3 volatilization was 4 times less when injected at 7 cm depth compared to broadcast. In an irrigated maize system, the surface-banded N caused much greater N2O emissions than the surface broadcast N (Halvorson and Del Grosso 2013). Shallower placement (< 5 cm) of N fertilizer caused a significant increase in N2O emissions relative to deep placement (> 5 cm) (van Kessel et al. 2013).
Some promising options include widespread adoption of slow-release N fertilizers, created by coating fertilizer pellets in a polymer film, that provide N in proportion to crop requirements (Houlton et al. 2019). The use of slow-release N fertilizer has the potential to further enhance NUE and crop production, while reducing N loss (Dong et al. 2020; Gui et al. 2022). For example, Ye et al. (2013) found that polymer-coated urea significantly reduces soil N surplus as well as N contents in surface water. Deep addition of slow-release N fertilizer can effectively enhance N transfer from vegetative tissues to grains, increasing plant NUE and production (Zhu et al. 2019). Moreover, Zhu et al. (2019) and Hou et al. (2021) point out that slow-release N fertilizer addition can effectively increase NUE in paddy soils. More recently, Gui et al. (2022) found that the deep application (at a depth of 10 cm) of slow-release N fertilizer increased N recovery efficiency by 171–232% compared to adding the same rate by the manual fertilizer broadcasting method. Unfortunately, high production costs and non-availability are two important reasons for limiting these compounds’ usage by farmers in developing countries (Yadav et al. 2017). However, for example, the production and use of slow-release N fertilizers in China, which has the lowest NUE, have increased rapidly in recent years, representing half of global consumption (Liu et al. 2020). Similarly, recent advances in sensing techniques that directly monitor fertilizer N in the plant rooting zone could greatly enhance NUE (Houlton et al. 2019). Since N is a function of multiple soil factors, there are opportunities to use precision agriculture diagnostics to measure and optimize NUE at a field scale that targets soil traits. Precision agriculture is using typical practices such as sensors, remote sensing, data management, geographical information systems, spectral imaging, and the global positioning system (Zhang and Kovacs 2012) which help to classify field patterns of variability to control plant and soil sampling for rational and optimal spatial input use (Franzen et al. 2011). For optimal crop N management, particularly with non-legumes (e.g., wheat and corn), precision agriculture technologies have been extensively applied (Yost et al. 2017). The in situ measurements of available soil nutrients and soil pH are needed to permit a variable input rate for optimizing field-level NUE and support site-specific management of soil chemical properties (Thilakarathna and Raizada 2018).
Although site-specific management N measures already mentioned aim to increase the NUE and reduce N loss into the environment, farmers do not follow these strategies and NUE has not significantly increased over the past few decades, especially in the developing countries. There is often a gap between what is practiced at farms and scientific studies (Rütting et al. 2018). The uncertainty of almost all farmers about the efficacy of new technological tools is still an obstacle. Lower implementation costs and reduction of economic and social barriers limiting the application of 4R related measures can help achieve scalable impacts and encourage farmers (Houlton et al. 2019). Furthermore, the interaction between scientists and stakeholders should be improved to acquire innovative concepts based on sound socio-economic, environmental, and production-ecological research to expedite the transition into food systems which will optimize N use to the end-users from the farm level (Van Grinsven et al. 2013).
4.2 Using of Synthetic and Biological Nitrification Inhibitors
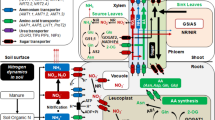
Soil NH4+ is easily oxidized to highly mobile NO3– within 2–3 weeks through nitrification process, which is vulnerable to loss via runoff, leaching and N2O emission during denitrification (Elrys et al. 2020b). Thus, controlling soil nitrification rates is a prerequisite to reduce N losses and improve NUE. One of the highly effective mitigation technologies in controlling nitrification process and improving NUE and yields is using nitrification inhibitors (NIs) (Moir et al. 2012). NIs greatly suppress the oxidation of NH4+ to NO3− (nitrification process) in the soil (Fig. 4) and subsequent denitrification of NO3− as compared with conventional fertilizers (Akiyama et al. 2010). Specifically, the synthetic NIs inhibit the ammonia monooxygenase (AMO) pathway (Fig. 4) within nitrification (Subbarao et al. 2008). The application of NIs improves NUE and agronomic yield benefit as well as the environmental and economic footprint of food production (Li et al. 2018; Raza et al. 2018). Only a few among hundreds of NIs have gained commercial importance, such as dicyandiamide (DCD), which is commonly the most used synthetic NIs (Ball et al. 2012; Elrys et al. 2020b) with high efficiency in improving NUE and yield and reducing N losses through emissions (N2O) and leaching (NO3−) (Moir et al. 2012; Raza et al. 2021). Soil that received DCD had higher NH4+ and lower NO3− contents than the control (soil without DCD) (Ma et al. 2021). Similarly, Raza et al. (2021) found that DCD effectively suppressed NH4+ oxidation and N2O emissions. Ammonia-oxidizing bacteria (AOB), which is responsible for soil NH4+ oxidation, increased significantly with urea addition, while this influence was suppressed in soils when DCD was added (Elrys et al. 2020b). The 3,4-dymethylpyrazole phosphate (DMPP) is also a commonly used synthetic NI and has high efficiency, especially when mixed with ammonium sulfate, ammonium sulfate nitrate (ASN), or urea (Trenkel 2010). Former studies have indicated that use of DMPP may decrease NO3− leaching losses, as well as NO and N2O emissions, and thus increasing NUE (Liu et al. 2013; Ruser and Schulz 2015). Martinez et al. (2015) noticed improvements in crop yield and N uptake after application of DMPP with urea. In three tropical soils, de Paulo et al. (2022) found that DMPP reduces N leaching and enhances the agronomic efficiency of N fertilizers added to cotton. Yang et al. (2016) reported in a meta-analysis that DCD and DMPP had the same effect on altering soil inorganic N content, N2O emissions, and N leaching. Meanwhile, DCD was more effective than DMPP in improving plant productivity. Additionally, Cheng et al. (2022a, b) in a wheat-rice rotation system reported that the combined use of urea and urease inhibitor N-(n-butyl) thiophosphorictriamide (NBPT) or the new nitrification inhibitor 3,4-dimethylpyrazole succinic acid (DMPSA) resulted in comparable grain yield and NUE but with lesser NH3 and N2O emissions. Drulis et al. (2022) also suggested that urease inhibitors can decrease dependence on N fertilizers and enhance maize production. Although synthetic NIs are effective in increasing NUE and reducing N loss, they have not been widely adopted in production agriculture due to the lack of cost-effectiveness and their inconsistent performance across diverse agro-climatic and soil environments (Wang et al. 2021b). The effectiveness of synthetic NIs depends on the soil or environment conditions, management strategies, and cropping system (Elrys et al. 2020b; Raza et al. 2021; Wu et al. 2007). By a meta-analysis study, Abalos et al. (2014) reported that the synthetic NIs efficiency in acidic soils and medium or coarse texture were higher than that in alkaline soils and fine texture with decidedly variable effects on certain experiments. Experimentally, Raza et al. (2021) found that DCD degraded quickly at 25 °C and 60% water filled pore space, that is why it was least effective in suppressing nitrification process and driving gaseous fluxes. In addition, Elrys et al. (2020b) reported that soil pH, soil organic matter, and soil texture have significant influences on the performance of DCD. In alkaline soils, for example, NH3 volatilization was significantly increased when DMPP (Xu et al. 2019) or DCD (Elrys et al. 2020b) was added to the soil compared with urea addition. Moreover, plants can pick up synthetic NIs (e.g., DCD) (Marsden et al. 2015), and then find their way into the human food chain, leading to negative health effects (Lucas 2013).
Alternatively, biological NIs are a low-cost and environmentally friendly compared to the synthetic NIs (Subbarao et al. 2009a). Functionally similar compounds released from plant tissues and roots influence biogeochemical processes, including the N cycle (Fig. 4) (Subbarao et al. 2013). Certain plants can suppress soil nitrification by releasing NIs from plant roots such as that released by sorghum roots which distinguished it among all other cereals (Subbarao et al. 2013). Therefore, utilizing natural NIs could become a powerful strategy with beneficial returns both agriculturally and environmentally. For example, pastures covered by Urochloa humidicola, which has the capacity for biological NI, have been known for their low soil potential nitrification rate, which may be due to the release of brachialactone from plant roots (Subbarao et al. 2009b; Teutscherova et al. 2019). The study of Villegas et al. (2020) provides evidence of the capacity of Megathyrsus maximus to inhibit soil nitrification and N2O emissions. Yao et al. (2020) found that the biological nitrification inhibitor methyl 3-(4-hydroxyphenyl) propionate significantly decreased soil N2O emissions due to the inhibition of nitrification. Moreover, Lu et al. (2018) reported that biological nitrification inhibitor 1,9-decanediol suppresses ammonia-oxidizing archaea (AOA). Using stable isotopes techniques; however, Vázquez et al. (2020) did not emphasize lower gross nitrification rates under the high-biological NI genotypes, notwithstanding reduced potential net nitrification rates and AOB abundance, which could be explained by increased NH4+ and NO3− immobilization under high-BNI genotypes. This was in line with the finding of Ma et al. (2021), who suggested that linoleic and linolenic acids addition may cause apparent nitrification inhibition by stimulating soil microbial NH4+ and/or NO3− immobilization. Furthermore, the role of essential oils, along with the role of oil by-products in retarding the nitrification process, has been studied (Arora and Srivastava 2014; Elrys et al. 2021a). For instance, Patra et al. (2009) concluded that the apparent N recovery in the cropping system of mint-wheat-rice can be improved by coating the urea with mint (Mentha arvensis) oil by-products. Elrys et al. (2019) suggested that using of moringa (Moringa oleifera) seed extracted to inhibit AOB abundance in the soil is a significant strategy to decrease nitrification rate and N loss, and therefore improve NUE. The most important particular distinct of nitrification inhibitors based on a by-product of an essential oil is that besides regulating nitrification, it helps reduce urea-N volatilization loss (Arora and Srivastava 2014). Moringa oil coated urea, for example, significantly decreased the abundance of AOB in the soil and the accumulative of N2O and NH3 emissions, while enhancing potato NUE (Elrys et al. 2021a). In a Typic Haplustert under paddy, neem cake when used for urea encapsulation reduced the loss of NH3-N to only 1.7–6.7% at different stages of plant growth (Purakayastha et al. 1997).
4.3 Increased Insight into NO 3 − Retention in Croplands
Long-term use of high levels of synthetic N fertilizers induces nitrification and NO3− accumulation in croplands (Bai et al. 2021). In China, for example, 19–31% of the annual N inputs in vegetable production accumulated in soil (Ti et al. 2015). Soil NO3− accumulation is susceptible to losses through leaching, surface runoff, and denitrification. After land-use change from croplands (cereals) to apple orchards on the Loess Plateau of China, Zhu et al. (2021) found that the mean NO3− accumulation in the soil profile (0–200 cm) was 2635 kg N ha−1, and that 67% of the total accumulated N surplus in the soil profile was NO3−. Thus, the fact that decreasing NO3− accumulation in the soil has an effective role in enhancing NUE and reducing N loss indicates the need to realize the relative magnitude of global NO3− consumption processes (microbial NO3− immobilization and DNRA) and their controlling factors. Accordingly, in addition to the traditional use of site-specific N management and NIs, restoring NO3− retention capacity can be another promising way to decrease the excessive NO3− accumulation in croplands, and ultimately improve NUE.
4.3.1 DNRA
Soil NO3− can be reduced via DNRA by soil microbes (bacteria and/or fungi) to release NH4+ in soil, as an additional path to the mineralization process (Silver et al. 2001). Therefore, the DNRA process reduces the NO3− pool and enhances the availability of NH4+, which can conserve soil N (Lai et al. 2021). More recently, Cheng et al. (2022a, b) found a global average DNRA rate of 0.31 ± 0.05 mg N kg−1 day−1, being significantly greater in paddy soils (1.30 ± 0.59 mg N kg−1 day−1) than in grasslands (0.52 ± 0.15 mg N kg−1 day−1), forests (0.24 ± 0.03 mg N kg−1 day−1), and croplands (0.18 ± 0.04 mg N kg−1 day−1). Former studies revealed that DNRA was sometimes the main fate of NO3− in soils. In Chilean temperate rainforest soils, for example, DNRA process consumes almost all soil NO3–, conserving N in the soil–plant system (Rütting et al. 2008). In addition to enhancing the availability of NH4+ in the soil, DNRA process can reduce N2O emissions (Cheng et al. (2022a, b; Kraft et al. 2014). In the absence of oxygen, NO3− and nitrite (NO2−) are used by soil microbes as alternative electron acceptors within denitrification (Tiedje 1982), thus emitting N2O and N2. Whilst, DNRA process also used the same substrates (NO3− or NO2−) and thus it can effectively compete with denitrification, decreasing N2O emission (Rütting et al. 2011; Pandey et al. 2020). In rhizosphere and non-rhizosphere soils in arid croplands, Pan et al. (2020) reported that DNRA process dominated NO3− consumption process against denitrification. A recent meta-analysis of Cheng et al. (2022a, b) revealed that precipitation and its effect on soil oxidation state is the master factor affecting soil DNRA. They also suggested that the DNRA process may play a key role in improving NUE and decreasing N2O emissions in paddy soils and thus should be taken into account when studying N transformations in paddy soils. Experimentally, Nojiri et al. (2020) found that DNRA decreases N loss through NO3− leaching and denitrification and supplies NH4+ to rice roots. Thus, the N mineralization model of Schimel and Bennett (2004) must be modified, at least for paddy soils, to comprise DNRA process as an alternative producing process for NH4+. Previous studies also reported that soil DNRA process increased significantly with increasing soil C, a condition that restricts soil nitrification (Cheng et al. 2022a, b; Rütting et al. 2011; Pandey et al. 2020). Soil C supplies electrons through respiration or fermentation, which reduces NO3− to NH4+ and thus provides energy to DNRA microbes (Van Den Berg et al. 2016). Moreover, Pandey et al. (2018) demonstrated that DNRA plays a key role in N retention in paddy soil, as it represents up to 55% of the total NO3-consumption, but this is decreased by N addition. This was in line with the finding of Cheng et al. (2022a, b) on a global scale, who found that soil DNRA rate decreased significantly with increasing soil N availability. Thus, we can suggest that the exogenous organic C additions aligned with low rates of N application could stimulate DNRA process, decreasing soil NO3− accumulation and enhancing NUE, especially in paddy soils. As denitrification process caused N loss, whilst DNRA process results in N retention, the relative contributions of DNRA versus denitrification activities should be appropriately taken into account when evaluating soil N transformations and N fertilizer management in croplands (Pan et al. 2020).
4.3.2 Microbial NO 3 − Immobilization
Soil microbial NO3− immobilization converts NO3− into microbial biomass N temporarily, which can posteriorly be converted to stable organic N or re-mineralized, thus increasing soil NO3− residence time and thus decreasing its risk of loss (Davidson et al. 1992; Elrys et al. 2022; Li et al. 2019). In forest soils, NO3− immobilization and rapid microbial turnover play a key role in driving NO3− pool size and thus shape a main mechanism for soil N retention (Elrys et al. 2022; Li et al. 2020; Stark and Hart 1997; Zhang et al. 2013). However, the conversion of forests to croplands significantly suppresses soil microbial NO3− immobilization (Zhang et al. 2013). Thus, in addition to long-term excessive inorganic N fertilizer inputs and nitrification promoting (Bai et al. 2021), impairment of microbial NO3− immobilization is another major cause of NO3− accumulation in croplands (Li et al. 2020; Zhang et al. 2013). Thus, besides the traditional use of NIs and site-specific N management, stimulation of NO3− immobilization in croplands can be another good way to enhance NUE (Cheng et al. 2017; Cao et al. 2021). The impairment of microbial NO3− immobilization in croplands is likely due to the limitation of heterotrophic microorganisms' C (Shi and Norton 2000). In croplands, available C is probably not sufficient to supply the requirements of energy for microbial NO3− immobilization (Li et al. 2019). In croplands, microbial NO3− immobilization was negligible when potassium nitrate was applied alone, however it occurred when glucose-C was applied along with the same amount of potassium nitrate (Recous et al. 1990). Sufficient addition of C-glucose leads to a rapid depletion of NH4+, and for the microorganisms to maintain their growth they shift toward NO3− immobilization, ultimately improving NO3− immobilization (Cheng et al. 2015). In an upland soil, Wang et al. (2021a) suggested that organic material with high C to N ratio could stimulate microbial NO3− immobilization. Thus, exogenous organic C applications with a high C to N ratio or at a high rate could stimulate microbial NO3− immobilization and decrease soil NO3 accumulation, and ultimately improve NUE. Furthermore, fungi exhibit a hyphal growth form, which allows them to translocate NO3− across wide spatial scales from micro-sites with high N production to micro-sites with high C availability (Frey et al. 2003). Fungi favor relatively undisturbed soils with high organic C and C to N ratio, and low nutrient availability and pH (Högberg et al. 2007). Li et al. (2019) found that land conversion from forests to croplands significantly reduces fungi NO3− immobilization capacity by 81%, manifesting that improving fungi living conditions would be a good way to recover microbial NO3− immobilization capacity in croplands, thus enhancing NUE.
4.4 Importance of Agroforestry System in the N Cycling
Croplands had a lower potential to conserve N, increasing a higher risk of N loss. In the forested land use, however, the processes of gross N mineralization and gross N immobilization were tightly coupled, resulting in reduced substrate (NH4+) for nitrification and ultimately reduced N loss to the environment (Elrys et al. 2022; Lang et al. 2019). Agroforestry systems are multifunctional land use systems with trees incorporated into agricultural crops. This incorporation benefits these systems ecologically, environmentally, and economically (Jose 2009). Although the agroforestry system is still not common, the establishment of this system could minimize total N surplus of farming systems because (1) the N requirements of short-rotation trees are generally less than that of agricultural crops (Sevel et al. 2014), thus the external N fertilizer inputs and the N surplus of the whole agroforestry system can be decreased, (2) it has the potential to enhance the NUE of the system due to its capacity to improve nutrient uptake and soil quality (Ilany et al. 2010; Patra 2013), (3) the deep rooting of trees in agroforestry systems can capture and recycle subsoil inorganic N that have leaked below the rooting zone of linked agricultural crops, causing more efficient interception of leaked N (Cannavo et al. 2013), and (4) there is greater microbial N immobilization in tree-based systems as compared to monocropping systems, thus reducing N loss (Kaur et al. 2000; Lang et al. 2019). Lin et al. (2016) found that the agroforestry system can enhance NUE and reduce N surplus. Accordingly, we suggest that the global N cycle in the context of preserving the environment requires a shift towards agroforestry systems with a gradual increase in the ratio of forests to croplands (Elrys et al. 2021b, c; Elrys et al. 2022).
5 The Molecular and Genetic Approaches of NUE in Crop Plants
Genetic improvement of NUE is considered to be a paramount aspect and crucial for eco-friendly crop production and sustainable agriculture (Yu et al. 2015). Unraveling the molecular basis of how plants respond to changes in N availability would enable the improvement of new strategies to increase NUE (Liu et al. 2015). Genetic approaches involved in, using classical genetic methods focused on the use of known allelic variations for NUE traits, phenotyping problems, mapping of quantitative trait loci (QTLs), and the selection of candidate genes for NUE enhancement. Improving NUE could be achieved by improving and/or modifying genes concerned in the structure of roots, N transportation, primary N metabolism, N remobilization genes, N developmental and regulatory genes, and others (Table 2; Han et al. 2015; Reddy and Ulaganathan 2015).
NUE is a complex trait affected by a wide range of various genes. Many gene targets have been selected for bioengineering trials depending on the identification of either; established N metabolism functional genes, QTL genetic surveys, various up-regulated genes in transcriptome analysis of N-limitation responsive genes, or N-efficient plants. Gene identification by QTL and transcriptomic studies are important for NUE and encode integral proteins for N metabolism, for instance, NO3− transport (NRT), NH4+ transport (AMT), glutamate synthase, glutamine oxoglutarate aminotransferase (GOGAT), and glutamine synthetase (GS) (Li et al. 2017; Xu et al. 2017). Over-expression of these genes led to an increase in amino acid levels, N levels, as well as an increase in the biomass or number of seeds. N-related genes could be classified into two large gene groups, N metabolism pathways and N developmental and growth pathways. These groups can also be classified into six gene families (Table 2). First, the N metabolism pathways class encompasses transportation-related genes, amino acid biosynthesis, assimilation, and a collection of other genes (Han et al. 2015). Second, the growth and development class contains genes linked to transcription factors, small RNA, senescence, and signalling processes (Fischer et al. 2013; Han et al. 2015).
5.1 Using Classical Genetics
Classical genetics is one of the genetic approaches used for improving NUE of crops. This includes traditional breeding and QTL, as well as a set of marker-assisted selection (MAS) to monitor key regions of the NUE-segregated chromosomes. One of the primary steps to achieve this objective is to describe the NUE-related genes and then use specific gene profiles to link genetics with plant physiology, resulted in enhancing plant performance (Beatty et al. 2010; Hawkesford 2011; Liu et al. 2012).
5.1.1 Breeding Techniques
Improving the NUE in crop production has been a long-lasting global challenge. One of the genetic approaches is breeding plants for some traits that enable plants not only to minimize the use of N-fertilizers and hence N-leaching in production but also to increase plant tolerance to several stresses (Merwad et al. 2018). Plant-bred cultivars have a high NUE relative to older varieties. For instance, some modern varieties of wheat, maize, rice, and barley that showed better growth and yield production than the older ones, under N-limited conditions (Chen et al. 2013; de Carvalho et al. 2016). Genetic screens have been conducted to detect NUE-trait-related genes on some of these NUE varieties. In some cases, it was found that primary N metabolism appears to be important for the NUE trait (Quraishi et al. 2011). However, till now, the molecular mechanisms that lead to high NUE are still not known in such varieties (Hawkesford 2011). The GmCBS21 gene has been identified and overexpressed as an NUE-candidate gene in soybean, which has shown greater low N stress tolerance than wild-type (WT) plants, which indicates its function in decreasing N stress tolerance and can be an excellent candidate for breeding plants with increased NUE and yield (Hao et al. 2016). TaGS1 and TaGS2 were highly expressed in post-anthesis and pre-anthesis, respectively, in N-efficient winter wheat cultivars compared with N-inefficient ones (Zhang et al. 2017). In the winter wheat, four metabolic regulatory points were described to be implicated with GS, through various types of tissue. However, an analysis of the barley NUE genes showed that some NUE-related genes co-segregated with QTLs assessed by the field for the NUE (Han et al. 2016a, b). Interestingly, Stahl et al. (2017) studied recent genetic gains in NUE in winter oilseed rape varieties under two divergent N fertilization levels (NFLs) and the results showed a highly significant genetics-driven increase in NUE at both NFLs and seed yield per se. On average, modern varieties displayed an increase in seed yield and oil concentration and a decrease in protein concentration comparing to old varieties.
In addition, another contemporary breeding technique is to use microspore mutagenesis and field screening which will produce a mutant line with high N uptake efficiency. In vitro mutagenesis using microspore, culture isolation affords an efficient way for producing multiple dual haploid (DH) lines with mutation and homozygosis stabilization, which could be utilized for direct screening (Gao et al. 2018). For instance, field screening and microspore mutagenesis have been studied for the rapid generation of barley mutant lines with a higher N uptake efficiency (Gao et al. 2018), and the results showed that the N uptake efficiency (NUP) of two mutant lines, A1-84 and A9-29, improved under low N, whereas four lines, A1-84, A1-226, A9-29, and A16-11, have been greatly improved under high N, meaning that their NUE had improved depending on the enhancement of N uptake. These results implied the importance of field screening and microspore mutagenesis in the NUE enhancement in barley.
5.1.2 Quantitative trait loci (QTL) mapping
QTL mapping may reveal unknown genes’ chromosomal locations that affect the quantitative variation of complex traits such as NUE and NUP, providing valuable information about the genetic basis for such traits and their impact on yield production. Numerous studies were carried out to map the QTL for NUE in plants (Table 3). For rice, QTLs associated with NUE were determined for the first time in and the analysis results identified six loci that cosegregated with NADH-GOGAT activity and seven loci that cosegregated with GS1 activity (Obara et al. 2001). On chromosome 6, one QTL regulating rice NUE was identified, qNUEP-6, in a recombinant inbred line (RIL) population on two lines, Minghui 63 and Zhenshan 97 (Shan et al. 2005). Under low N conditions, one QTL for rice NUE named, pnue9, was identified on chromosome 9 using a RIL population of two lines, TR22183 and TR22183 (Cho et al. 2007). A major QTL associated with rice NUE was detected on chromosome 3 under three N levels, in a DH population from Azucena and Azucena (Senthilvel et al. 2008). Recently, a set of thirteen QTL for rice NUE and NUP have been identified, six of them for NUE while seven for NUP, in plants under hydroponic culture with all-sufficient quantities nutrients supplied using high-throughput genotyped chromosome segment substitution lines (CSSLs); meanwhile, several QTLs for grain yield and biomass yield were identified displaying that grain yield was substantially positively associated with NUE and NUP (Zhou et al. 2017).
For maize, a meta-analysis study was conducted for QTLs for yield components and resulted in detecting a total of 22 of meta-QTLs under low N conditions (Liu et al. 2012). Several QTLs were identified for GY-related traits on maize chromosomes, which will also provide targets for gene cloning and QTL pyramiding (Tuberosa and Salvi 2009). On the other hand, using QTL analysis in maize enabled Li et al. (2015) to reveal the genetic correlation between NUE and root system architecture (RSA) using a recombination inbred line population, and several 331 QTLs were identified, including 147 and 184 QTLs for RSA- and NUE-related traits, respectively. This study uncovered the most promising RSA marker-assisted genomic regions that resulted in enhancing NUE in maize (Li et al. 2015).
For barley, many QTLs maps have shown an association with barley yield (Backes et al. 1995). An association mapping approach was used to search for genes incorporate with a set of traits in barley and several QTLs were found to be involved in grain and biomass yields traits that are associated with NUE (Varshney et al. 2012). Also, 329 genotypes of winter barley were mapped for markers associated with NUE and two QTLs were identified (Berger et al. 2013). A detailed analysis described three QTLs associated with N storage and remobilization on chromosomes 5H and 6H, explaining 37.4% of the genetic variation (Mickelson et al. 2003). QTLs were analyzed in field trials toward barely NUE, and findings provide a novel insight into identifying NUE genes that contribute to a profound understanding of the physiology of the different gene families that may be involved in the NUE of barley (Han et al. 2016a, b).
In wheat, fifty-four genomic regions linked to GY and their components have been identified in 196 wheat accessions through the whole-genome mapping. In addition, identifying a total of 23 N responsive genomic regions, which could be utilized by programs of wheat breeding (Bordes et al. 2013). Also, it was found that N fertilization influenced QTLs of a wheat RIL population and has an impact on N concentration in shoots, grain, and NUE traits. Further evaluation of newly identified QTLs, including QTLs on chromosomes 7A and 4B, may be useful for the characteristics of the yield component traits (Xu et al. 2014).
Many QTLs correlated with N use and yield have been mapped to the chromosome regions of glutamine synthetase 2 (GS2) in wheat and rice (Fontaine et al. 2009; Habash et al. 2007), indicating that the genomic region around GS2 may be useful for breeding programs of rice and wheat varieties with higher NUE and yield (Yang et al. 2007). In wheat and barley, some QTLs shared physiological traits such as N metabolites and N metabolic enzyme activities (Heidlebaugh et al. 2008; Fontaine et al. 2009). Though such traits are particularly vulnerable to experimental and environmental factors (e.g., glasshouse versus field), there are possible outcomes such as metabolic QTLs (mQTLs) which introduce direct proof for linking agronomic traits to potential QTLs genes (Hill et al. 2013).
5.2 Omics Approaches, Metabolomics, and Transcriptomics
Omics approaches including transcriptomics and metabolomics have become a powerful and promising way for investigating complex network interactions of the N metabolism, including N uptake and regulation, translocation, remobilization, and amino acid metabolism (Miller et al. 2008). A schematic figure was drawn for a better understanding of the relationships between the genome, transcriptome, proteome, and metabolome (Fig. 5). Owing to the high-throughput technologies, deep sequencing of transcriptome and gene expression profiles using microarray techniques enables the investigation of gene regulation networks in N signaling and metabolism (Fukushima and Kusano 2014). Moreover, immense omics datasets that describe cellular components, including transcripts and metabolites, help to realize complex genotype–phenotype relationships by facilitating the evaluation of interactions between these components and networks, for example, protein–protein interactions, transcriptional regulatory networks, and metabolic pathways (Barabási and Oltvai 2004). This analysis highlights recent advances in omics, including transcript profiling, deep sequencing, and use of microarrays, besides introduces advanced technical developments in the N metabolomics research. Thus, omics data not only extend our knowledge of N responses but also lead to successful strategies that enhance NUE and improve crop production (Schwanhäusser et al. 2011).
5.2.1 Transcriptomics Using Microarray and High-Throughput RNA Sequencing
A transcriptome is a collection of total RNA molecules in a single cell or a cell population. It is also occasionally used to mean either all RNAs or only mRNA, particularly based on the experiment (Wang et al. 2009). Transcriptomic technology includes DNA microarrays and next-generation sequencing (NGS) technologies termed RNA-Seq or high-throughput sequencing (HTS) (Ogorodnikov et al. 2016). To analyze RNA-seq data, some software used for this purpose such as Gene Pattern (Reich et al. 2006), RobiNA (Lohse et al. 2012), metaseqR (Moulos and Hatzis 2015), globalSeq (Rauschenberger et al. 2016), and MAPTest (Cao et al. 2019).
Global transcription profiling using microarrays has been an effective approach to study the molecular features of stress responses and nutrients. Microarray analysis in rice revealed an expression profile of 10,422 unique genes, whereas no major differences were detected in the transcriptome of leaf tissues; however, a set of 471 genes displayed differential expression in root tissues under low-N stress (Lian et al. 2006). On the other hand, microarray analysis in Arabidopsis has been conducted to investigate gene expression variations under various concentrations of NO3− for both long-term and short-term treatments, resulting in the detection of several genes associated with N response (Peng et al. 2007). In maize, numerous N-responsive genes have been detected after examining the global gene expression variations using the Affymetrix maize genome array in two maize inbred lines, using leaf tissues, under N-stress conditions compared to N-stress tolerance (Chen et al. 2011). Many candidate genes related to N utilization also have been detected in two soybean genotypes exposed to N-stress via the transcriptional profiling using Illumina RNA-sequencing (Hao et al. 2016). High-throughput RNA sequencing of maize hybrids and their parents revealed various mechanisms of response to N limitations in shoots and roots, for example, the metabolism of the cellular N compound, amino acids metabolism, and expression patterns, including up/down-regulation cytokinin degradation pathway (Bi et al. 2007, 2009).
The entire genome sequencing demonstrates potential novel targets for enhancing N utilization and uptake in cereals (Masclaux-Daubresse et al. 2010). A set of 230 genes possibly involved in N utilization and uptake were selected and analyzed for genetic diversity and signatures in various panels of sorghum lines, revealing a total of 128 genes targets to increase NUE in sorghum and cereals alike. Besides, some main candidate genes have been detected and further utilized in breeding programs to improve NUE in cereal crops (Massel et al. 2016). Moreover, another genetic approach is genome-wide transcriptional profiling used to identify differentially expressed genes among genotypes with variation in N stress tolerance (Gelli et al. 2014). A transcriptome study using Illumina RNA-sequencing was performed for seven sorghum genotypes root tissues with different responses to low N stress, revealing that four low-N-tolerant genotypes share many common differentially expressed genes with three sensitive genotypes. While the higher expression in tolerant sorghum genotypes would be utilized for over-expression in sensitive genotypes and cereals like crops for improving N-stress tolerance, thus increase NUE for sustainable agriculture (Gelli et al. 2014; Elrys et al. 2020c).
5.2.2 Metabolomics
A metabolome is the whole set of low-molecular-weight of the organism’s metabolites, whereas metabolomics means the biochemical study of all biological metabolites (Saito and Matsuda 2010). Recent development in the field of metabolomics has led to monitoring of a broad range of primary metabolite products, including amino acids, fatty acids, sugars, organic acids, and secondary metabolites such as carotenoids and phenylpropanoids, which means that metabolomics offers basic information regarding plant responses to N-induced environmental or physiological changes (Tschoep et al. 2009). Profiling of metabolites needs an analytical pipeline involving the sampling, extraction of samples, separation, analysis, data collection, and analysis of data using bioinformatics (Sugimoto et al. 2012).
Experimental results indicate that the induction of N uptake is reduced when urea and NO3− are present in the external solution, whereas N utilization and plant growth is promoted (Zanin et al. 2015). Such physiological behavior may reflect the relationship between the acquisition processes in which various N pathways are assimilated, such as plastidic and cytosolic pathways, which contribute to increased nutrient uptake management. Based on physiological and molecular evidence, plants could increase N metabolism and successfully enhance N-uptake assimilation (Pinton et al. 2016). NUE in maize were discussed from metabolic modelling studies providing an overview of NUE improvement in the yield and highlighted the metabolome and proteome datasets, leading to a better understanding of N regulation. It is difficult to analyze “omics” data in the absence of metabolic flow information in genome-scale models. Such models thus offer a basis to understand metabolic processes that are the cornerstone for the optimal usage of N-based fertilizers (Simons et al. 2014; Peng-Fei 2019).
Our lack of knowledge of the complicated metabolic networks, signalling mechanisms, and factors that physiologically affect NUE is one of the main obstacles to a full understanding of plant metabolism (Krapp et al. 2014). Moreover, understanding metabolic reprogramming in response to N-limited conditions is critical to achieving continuous progress in the production of NUE improved varieties (Shachar-Hill 2013). To decipher metabolic and molecular information for N availability, metabolic profiling and co-expression networking analysis were conducted in rice showing the relationship between signal transduction components and the regulatory framework for N mobilization upon N-limiting conditions (Sun et al. 2014; Coneva et al. 2014).
Notably, these observations sustained by metabolic profiling demonstrated that several compounds which involved directly in N assimilation, transport, and storage such as allantoin, glutamate, glutamine, asparagine, and most amino acids in leaves were changed according to nitrate level and respond with higher levels under nitrate reduction levels (Beatty et al. 2013; Coneva et al. 2014). Moreover, a model revealing the complex flux of N through plants was innovated which describes how experimental data on metabolomics could be utilized to improve such models, such as nutrient uptake analyses (Beatty et al. 2016). In short, the metabolomic approaches can offer effective mechanisms to understand NUE that may eventually provide engineered plants with higher yields and more effective crop management (Beatty et al. 2013).
5.3 Detection of N-Related Genes and Novel Target Genes for NUE Improvement
The NUE can generally be divided into three components: N uptake, N assimilation, and N remobilization or N utilization efficiency. To improve the NUE crop, it is important to understand the mechanisms regulating this process. Consequently, gene function regulation may be necessary to enhance the NUE in all processes of N metabolism including N uptake, assimilation, translocation, and remobilization (Huang et al. 2018).
Nitrogen Uptake. Four families were known for NO3− carriers, including 1/peptide transporter (NPF or family NRT1/PTR), the chloride canal (CLC), 2-family NO3− transport (NRT2), and channel slow-anion slow-moving homologues (SLAC/SLAH) families. Four families were known for NO3− transporters, including NO3− transporter 1/peptide transporter family (NPF, or PTR/NRT1family), NO3− transporter 2 (NRT2), the chloride channel family (CLC), and slow anion channel-associated homologues (SLAH/SLAC) (Krapp et al. 2014). At least 80 genes have been identified in the rice genome of the NPF family (Tsay et al. 2007). NRT2 and NPF are two identified transporter families for NO3− translocation and uptake (Table 4) (Xu et al. 2012; Léran et al. 2013). The majority of NPF family members have low-affinity NO3− transport activity, but the OsNPF6.5 (NRT1.1b) exhibited dual-affinity nitrate behaviors linked to improving NO3− uptake and root-to-shoot transport (Hu et al. 2015). In addition, NH4+ uptake is mainly regulated by the superfamily transport's methylammonium (MEP)/ NH3 transport protein (AMT)/ rhesus (RH) (Khademi et al. 2004). The AMT members' activities may have a major role in NUP in rice that prefers NH4+ rather than NO3−-preferring crops. It was reported that at least 10 putative OsAMT-like genes within the rice genome are assembled in 4 subfamilies, including a single gene for OsAMT4 and three genes for each of OsAMT1, OsAMT2, and OsAMT3 (Table 4) (Suenaga et al. 2003).
Nitrogen Assimilation. after N uptake, NO3− is assimilated in roots and transferred to shoots. Nitrogen assimilation is accomplished in three steps, firstly by catalyzing into nitrite by the cytoplasmic NO3− reductase (NR) enzyme, second to NH4+ in the plastids by nitrite reductase (NiR) enzyme, third assimilating the NH4+ into amino acids (aa) through the GS/GOGAT cycle (Fig. 6) (Xu et al. 2012; Neeraja et al. 2016). The transition of amide group of glutamines formed by GS to 2-oxoglutarate (2-OG) has been catalyzed by GOGAT. This process produces two glutamate (Glu) molecules, one of which can be recycled as a substrate for the GS reaction, and the second GLu molecule can be utilized for several reactions (Forde and Lea 2007). Rice contains a chloroplastic gene (OsGS2) and three homologous genes (OsGS1-1, OsGS1-2, and OsGS1-3); having some differences, for the cyto-glutamine synthetase. Both OsGS1-1 and OsGS1-2 can be catalyzed by NH4+ via the central cylinder of the elongating zone in rice since these genes have a high affinity for ammonium (Ishiyama et al. 2004). Besides, a tiny family of GOGAT genes, which constitutes one ferredoxin (Fd)-dependent type and two NADH-dependent types, OsFd-GOGAT, is abundantly present in light-regulating cells like mesophyll cells and other chloroplast-containing cells (Tamura et al. 2010).
Overview of the expression and multilevel regulation of the cytosolic enzyme glutamine synthetase 1 (GS1). While, carbon metabolites contain sugar substances like sucrose; nitrogen metabolites include NH4+, glutamate (Glu), and glutamine (Gln); both of Dof. and MYB are transcription factors; refer to the positive regulation is referred by open arrows whereas negative regulation is indicated by the T bars
Nitrogen Remobilization and Reassimilation. Leaves are a sink for N in the vegetative phase; later, in the senescence stage, N is remobilized for reuse in the developing seeds, mostly as amino acid (Okumoto and Pilot 2011). Asparagine (Asn) and Gln are major forms of the total amino acids in rice phloem and xylem sap (Xu et al. 2012; Ohashi et al. 2015). During senescence, both Gln and Asn concentrations are increased within phloem sap, suggesting their important role in providing the available N for remobilization from the senescence leaves (Huang et al. 2018). The GS/GOGAT activities were previously emphasized as essential in N re-assimilation, remobilization, growth rates, and grain filling or yield (Yamaya and Kusano 2014). GS1; 2 are also essential in developing the active tillers during the NH4+ assimilation which are developed through lignin synthesis (Ohashi et al. 2015). Along with GS, Asn synthetase is supposed to play a primary role in N metabolism, by catalyzing the Glu and Asn formation from Gln and aspartic acid (Xu et al. 2012; Ohashi et al. 2015).
The overexpression of GS1 was investigated in many cases to increase the NUE of crops (Fig. 6) (Swarbreck et al. 2011; Orsel et al. 2014; Thomsen et al. 2014). A key function of GS1 might be related to maintaining the internal N sensing and N flows throughout the vital developmental stages (Gadaleta et al. 2014). Increasing GS1 activity will accelerate vegetative growth due to increased whole-plant N uptake either during N deficiency conditions or senescence phase, besides increased GS1 activity will improve N remobilization from vegetative tissues into the grains (Oliveira et al. 2002; Goodall et al. 2013). For example, over-expression of AlaAT in rice under N-deficient conditions showed an increased biomass/yield, total-plant N, and key metabolites (Shrawat et al. 2008). Importantly, four AlaAT genes have been identified in the poplar genome which was expressed in seedlings by the exogenous N, though PnAlaAT3 was mostly expressed in roots and regulated by Gln and Gln- associated metabolites (Xu et al. 2017).
6 Conclusions and the Future Prospects
In current systems of crop production, efficient N management is crucial to improve long-term sustainability. Integrated N management and balanced fertilization improve plant performance as well as NUE of crop production systems. In this review we also suggested that restoring NO3− retention capacity and adopting agroforestry systems can be promising ways to improve NUE. The use of innovative science interventions along with recent technologies has a significant effect on NUE. NUE is a complicated process that is defined by quantitative trait loci and influenced by changes in the environment and the efficient integration of N uptake and N utilization. There are complex regulations of N uptake, assimilation, and remobilization. Genetically, modified crops and integrated management practices are the most effective biotechnological methods for enhancing NUE. This could be obtained by overexpressing the N uptake-related genes or the transporters genes of NH4+ and NO3−, in addition by management of main genes regulating the balanced metabolism between N and C. Moreover, identification of novel NUE-related genes will continue along with the candidate NUE genes related to N uptake, assimilation, amino acids biosynthesis, C/N metabolism and storage, and signal transduction pathways regulating N metabolism, translocation, remobilization, and senescence. A major objective in the future is to develop integrating research programs combining these approaches which primarily depend on plant physiology, molecular genetic approaches including, quantitative genetics, reverse and forward genetics, and agronomy approaches to increase NUE of plants. Besides genetics, the interplay between some other factors should be considered, including the interactions between water availability and N uptake, micronutrient–macronutrient interactions, and the interaction between N utilization and C metabolism. On the other hand, scientists should collaborate with stakeholders to develop creative ideas that are focused on environmental, sound production-ecological, and socio-economic studies to enable a transition to food systems that maximize N usage between the farmer and end-consumers. In short, we need to focus on the improved integrated agricultural management approaches since NUE was successfully increased and N inputs were decreased while yield was maintained.
References
Abalos D, Jeffery S, Sanz-Cobena A, Guardia G, Vallejo A (2014) Meta-analysis of the effect of urease and nitrification inhibitors on crop productivity and nitrogen use efficiency. Agric Ecosyst Environ 189:136–144. https://doi.org/10.1016/j.agee.2014.03.036
Abebe Z, Feyisa H (2017) Effects of nitrogen rates and time of application on yield of maize: rainfall variability influenced time of N application. Int J Agron 2017:1545280. https://doi.org/10.1155/2017/1545280
Abiko T, Wakayama M, Kawakami A, Obara M, Kisaka H, Miwa T, Aoki N, Ohsugi R (2010) Changes in nitrogen assimilation, metabolism, and growth in transgenic rice plants expressing a fungal NADP (H)-dependent glutamate dehydrogenase (gdhA). Planta 232:299–311. https://doi.org/10.1007/s00425-010-1172-3
Akiyama H, Yan XY, Yagi K (2010) Evaluation of effectiveness of enhanced efficiency fertilizers as mitigation options for N2O and NO emissions from agricultural soils: meta-analysis. Global Change Biol 16:1837–1846. https://doi.org/10.1111/j.1365-2486.2009.02031.x
Araki R, Hasegawa H (2006) Expression of rice (Oryza sativa L.) genes involved in high-affinity nitrate transport during the period of nitrate induction. Breeding Sci 56:295–302. https://doi.org/10.1270/jsbbs.56.295
Argento F, Anken T, Abt F, Vogelsanger E, Walter A, Liebisch F (2021) Site-specific nitrogen management in winter wheat supported by low-altitude remote sensing and soil data. Precision Agric 22:364–386. https://doi.org/10.1007/s11119-020-09733-3
Arora K, Srivastava A (2014) Variation in nitrification inhibition activity of neem leaves collected from different locations of Lucknow (India). Int J Plant Soil Sci 3:457–466. https://doi.org/10.9734/IJPSS/2014/8518
Ashikari M, Sakakibara H, Lin S, Yamamoto T, Takashi T, Nishimura A, Angeles ER, Qian Q, Kitano H, Matsuoka M (2005) Cytokinin oxidase regulates rice grain production. Agric Scie 309:741–745. https://doi.org/10.1126/science.1113373
Backes G, Graner A, Foroughi-Wehr B, Fischbeck G, Wenzel G, Jahoor A (1995) Localization of quantitative trait loci (QTL) for agronomic important characters by the use of an RFLP map in barley (Hordeum vulgare L.). Theor Appl Genet 90:294–302. https://springerlink.fh-diploma.de/article/10.1007/BF00222217
Bai X, Jiang Y, Miao H, Xue S, Chen Z, and Zhou J (2021). Intensive vegetable production results in high nitrate accumulation in deep soil profiles in China. Environ Pollut 117598. https://doi.org/10.1016/j.envpol.2021.117598
Ball BC, Cameron KC, Di HJ, Moore S (2012) Effects of trampling of a wet dairy pasture soil on soil porosity and on mitigation of nitrous oxide emissions by a nitrification inhibitor, dicyandiamide. Soil Use Manage 28:194–201. https://doi.org/10.1111/j.1475-2743.2012.00389.x
Barabási AL, Oltvai ZN (2004) Network biology: understanding the cell’s functional organization. Nat Rev Genet 5:101–114. https://doi.org/10.1038/nrg1272
Beatty PH, Anbessa Y, Juskiw P, Carroll RT, Wang J, Good AG (2010) Nitrogen use efficiencies of spring barley grown under varying nitrogen conditions in the field and growth chamber. Ann Bot 105:1171–1182. https://doi.org/10.1093/aob/mcq025
Beatty PH, Carroll RT, Shrawat AK, Guevara D, Good AG (2013) Physiological analysis of nitrogen-efficient rice overexpressing alanine aminotransferase under different N regimes. Botany 91:866–883. https://doi.org/10.1139/cjb-2013-0171
Beatty PH, Klein MS, Fischer JJ, Lewis IA, Muench DG, Good AG (2016) Understanding plant nitrogen metabolism through metabolomics and computational approaches. Plants (basel) 5:1–16. https://doi.org/10.3390/plants5040039
Belete F, Dechassa N, Molla A, Tana T (2018) Effect of split application of different N rates on productivity and nitrogen use efficiency of bread wheat (Triticum aestivum L.). Agric Food Security 7:1–10. https://doi.org/10.1186/s40066-018-0242-9
Berger GL, Liu S, Hall MD, Brooks WS, Chao S et al (2013) Marker-trait associations in Virginia Tech winter barley identified using genome-wide mapping. Theor Appl Genet 126:693–710. https://doi.org/10.1007/s00122-012-2011-7
Bi YM, Wang RL, Zhu T, Rothstein SJ (2007) Global transcription profiling reveals differential responses to chronic nitrogen stress and putative nitrogen regulatory components in Arabidopsis. BMC Genomics 8:281. https://doi.org/10.1186/1471-2164-8-281
Bi YM, Kant S, Clark J, Gidda S, Ming F, Xu J, Rochon A, Shelp BJ, Hao L, Zhao R, Mullen RT, Zhu T, Rothstein SJ (2009) Increased nitrogen-use efficiency in transgenic rice plants over-expressing a nitrogen-responsive early nodulin gene identified from rice expression profiling. Plant Cell Environ 32:1749–1760. https://doi.org/10.1111/j.1365-3040.2009.02032.x
Blackshaw RE, Brandt RN (2008) Nitrogen fertilizer rate effects on weed competitiveness is species dependent. Weed Sci 56:743–747. https://doi.org/10.1614/WS-08-065.1
Blackshaw RE, Semach G, Janzen HH (2002) Fertilizer application method affects nitrogen uptake in weeds and wheat. Weed Scie 50:634–641. https://doi.org/10.1614/0043-1745(2002)050[0634:FAMANU]2.0.CO;2
Blagodatskaya EV, Blagodatsky SA, Anderson T-H, Kuzyakov Y (2009) Contrasting effects of glucose, living roots and maize straw on microbial growth kinetics and substrate availability in soil. Eur J Soil Sci 60:186–197. https://doi.org/10.1111/j.1365-2389.2008.01103.x
Bordes J, Ravel C, Jaubertie JP, Duperrier B, Garde O et al (2013) Genomic regions associated with the nitrogen limitation response revealed in a global wheat core collection. Theor Appl Genet 126:805. https://doi.org/10.1007/s00122-012-2019-z
Brauer EK, Rochon A, Bi YM, Bozzo GG, Rothstein SJ, Shelp BJ (2011) Reappraisal of nitrogen use efficiency in rice overexpressing glutamine synthetase1. Physiol Plant 141:361–372. https://doi.org/10.1111/j.1399-3054.2011.01443.x
Butterbach-Bahl K, Baggs EM, Dannenmann M, Kiese R, Zechmeister-Boltenstern S (2013) Nitrous oxide emissions from soils: how well do we understand the processes and their controls? Phil Trans R Soc B 368:20130122. https://doi.org/10.1098/rstb.2013.0122
Cai C, Wang JY, Zhu YG (2008) Gene structure and expression of the high-affinity nitrate transport system in rice roots. J Integr Plant Biol 50:443–451. https://doi.org/10.1111/j.1744-7909.2008.00642.x
Cannavo P, Harmand J-M, Zeller B, Vaast P, Ramírez J, Dambrine E (2013) Low nitrogen use efficiency and high nitrate leaching in a highly fertilized Coffea arabica–Inga densiflora agroforestry system: a 15N labeled fertilizer study. Nutr Cycl Agroecosyst 95:377–394. https://doi.org/10.1007/s10705-013-9571-z
Cao M, Zhou W, Breidt FJ, Peers G (2019) Large Scale Maximum Average Power Multiple Inference on Time-Course Count Data with Application to RNA-Seq Analysis. Biometrics to Appear. https://doi.org/10.1111/biom.13144
Cao Y, He Z, Zhu T, Zhao F (2021) Organic-C quality as a key driver of microbial nitrogen immobilization in soil: a meta-analysis. Geoderma 383:114784. https://doi.org/10.1016/j.geoderma.2020.114784
De Carvalho EV, Afférri FS, Peluzio JM, Rotili EA, Dotto MA, de Faria LA (2016) Genetics parameters and association of NUE methods in maize under different nitrogen levels/ Parâmetros genéticos e associação de metodologias de EUN no milho sob diferentes doses de nitrogênio 03: 36–41. https://doi.org/10.18067/jbfs.v3i1.61
Chen R, Tian M, Wu X, Huang Y (2011) Differential global gene expression changes in response to low nitrogen stress in two maize inbred lines with contrasting low nitrogen tolerance. Genes Genomics 33:491–497. https://doi.org/10.1007/s13258-010-0163-x
Chen X, Chen F, Chen Y, Gao Q, Yang X, Yuan L, Zhang F, Mi G (2013) Modern maize hybrids in Northeast China exhibit increased yield potential and resource use efficiency despite adverse climate change. Glob Chang Biol 19:923–936. https://doi.org/10.1111/gcb.12093
Chen M, Zhao Y, Zhuo C, Lu S, Guo Z (2015) Overexpression of a NF-YC transcription factor from Bermuda grass confers tolerance to drought and salinity in transgenic rice. Plant Biotechnol J 13:482–491. https://doi.org/10.1111/pbi.12270
Cheng Y, Zhang J-B, Müller C, Wang SQ (2015) N tracing study to understand the N supply associated with organic amendments in a vineyard soil. Biol Fertil Soils 51:983–993. https://doi.org/10.1007/s00374-015-1044-x
Cheng Y, Wang J, Wang J, Chang SX, Wang S (2017) The quality and quantity of exogenous organic carbon input control microbial NO3− immobilization: a meta-analysis. Soil Biol Biochem 115:357–363. https://doi.org/10.1016/j.soilbio.2017.09.006
Cheng Y, Elrys AS, Merwad ARM, Zhang H, Chen Z, Zhang J et al (2022a) Global patterns and drivers of soil dissimilatory nitrate reduction to ammonium. Environ Sci Technol. https://doi.org/10.1021/acs.est.1c07997
Cheng Y, Elrys AS, Wang J, Xu C, Ni K, Zhang J, Pacholski A (2022b) Application of enhanced-efficiency nitrogen fertilizers reduces mineral nitrogen usage and emissions of both N2O and NH3 while sustaining yields in a wheat-rice rotation system. Agr Ecosyst Environ 324:107720. https://doi.org/10.1016/j.agee.2021.107720
Chiasson DM, Loughlin PC, Mazurkiewicz D, Mohammadidehcheshmeh M, Fedorova EE, Okamoto M, McLean E, Glass ADM, Smith SE, Bisseling T, Tyerman SD, Day DA, Kaiser BN (2014) Soybean SAT1 (symbiotic ammonium transporter 1) encodes a bHLH transcription factor involved in nodule growth and NH4+ transport. Proc Nat Acad Sci 111:4814–4819. https://doi.org/10.1073/pnas.1312801111
Cho YI, Jiang WZ, Chin JH, Piao ZZ, Cho YG, McCouch SR et al (2007) Identification of QTLs associated with physiological nitrogen use efficiency in rice. Mol Cells 23: 72–79. https://pubmed.ncbi.nlm.nih.gov/17464214/
Christiansen MW, Matthewman C, Podzimska-Sroka D, O’Shea C, Lindemose S, Mollegaard NE, Holme IB, Hebelstrup K, Skriver K, Gregersen PL (2016) Barley plants over-expressing the NAC transcription factor gene HvNAC005 show stunting and delay in development combined with early senescence. J Exp Bot 67:5259–5273. https://doi.org/10.1093/jxb/erw286
Coneva V, Simopoulos C, Casaretto JA, El-kereamy A, Guevara DR, Cohn J, Zhu T, Guo L, Alexander DC, Yong-Mei B, McNicholas PD, Rothstein SJ (2014) Metabolic and co-expression network-based analyses associated with nitrate response in rice. Genomics 15: 1056. http://www.biomedcentral.com/1471-2164/15/1056
Congreves KA, Otchere O, Ferland D, Farzadfar S, Williams S, Arcand MM (2021) Nitrogen use efficiency definitions of today and tomorrow. Front Plant Sci 12:637108. https://doi.org/10.3389/fpls.2021.637108
Davidson EA, Hart SC, Firestone MK (1992) Internal cycling of nitrate in soils of a mature coniferous forest. Ecology 73:1148–1156. https://doi.org/10.1016/j.soilbio.2019.03.023
De Paulo EN, Galindo FS, Rabêlo FHS et al (2022) 3,4-Dimethylpyrazole phosphate (DMPP) reduces nitrogen leaching in three tropical soils and improves the agronomic efficiency of nitrogen fertilizers applied to cotton. J Soil Sci Plant Nutr. https://doi.org/10.1007/s42729-022-00824-w
Del Pilar M-P, Cipriotti PA, Urricariet S, Peralta NR, Niborski M (2018) Using site-specific nitrogen management in rainfed corn to reduce the risk of nitrate leaching. Agric Water Manag 199:61–70. https://doi.org/10.1016/j.agwat.2017.12.002
Dong D, Wang C, Zwieten LV, Wang HL, Jiang PK, Zhou MM, Wu WX (2020) An effective biochar-based slow-release fertilizer for reducing nitrogen loss in paddy fields. J Soil Sediment 20:3027–3040. https://doi.org/10.1007/s11368-019-02401-8
Drulis P, Kriaučiūnienė Z, Liakas V (2022) The influence of different nitrogen fertilizer rates, urease inhibitors and biological preparations on maize grain yield and yield structure elements. Agronomy 12:741. https://doi.org/10.3390/agronomy12030741
Duan P, Rao Y, Zeng D, Yang Y, Xu R, Zhang B, Dong G, Qian Q, Li Y (2014) SMALL GRAIN1, which encodes a mitogen-activated protein kinase kinase 4, influences grain size in rice. The Plant J 77:547–557. https://doi.org/10.1111/tpj.12405
Elrys AS, Abdo AIE, Desoky EM (2018) Potato tubers contamination with nitrate under the influence of nitrogen fertilizers and spray with molybdenum and salicylic acid. Environ Sci Pollut Res 25:7076–7089. https://doi.org/10.1007/s11356-017-1075-y
Elrys AS, Desoky ESM, El-Maati MFA, Elnahal AS, Abdo AI, Raza S, Zhou J (2019) Can secondary metabolites extracted from Moringa seeds suppress ammonia oxidizers to increase nitrogen use efficiency and reduce nitrate contamination in potato tubers? Ecotoxicol Environ Saf 185:109689. https://doi.org/10.1016/j.ecoenv.2019.109689
Elrys AS, Metwally MS, Raza S, Alnaimy MA, Shaheen SM, Chen Z, Zhou J (2020a) How much nitrogen does Africa need to feed itself by 2050? J Environ Manage 268:110488. https://doi.org/10.1016/j.jenvman.2020.110488
Elrys AS, Raza S, Elnahal ASM, Na M, Ahmed M, Zhou J, Chen Z (2020b) Do soil property variations affect dicyandiamide efficiency in inhibiting nitrification and minimizing carbon dioxide emissions? Ecotoxicol Environ Saf 202:110875. https://doi.org/10.1016/j.ecoenv.2020.110875
Elrys AS, Abdo AIE, Abdel-Hamed Enas MW, Desoky EM (2020c) Integrative application of licorice root extract or lipoic acid with fulvic acid improves wheat production and defenses under salt stress conditions. Ecotoxicol Environ Saf 190:110144. https://doi.org/10.1016/j.ecoenv.2019.110144
Elrys AS, El-Maati MFA, Abdel-Hamed EMW, Arnaout SM, El-Tarabily KA, Desoky ESM (2021a) Mitigate nitrate contamination in potato tubers and increase nitrogen recovery by combining dicyandiamide, moringa oil and zeolite with nitrogen fertilizer. Ecotoxicol Environ Saf 209:111839. https://doi.org/10.1016/j.ecoenv.2020.111839
Elrys AS, Ali A, Zhang H, Cheng Y, Zhang J, Cai Z, Müller C, Chang SX (2021b) Patterns and drivers of global gross nitrogen mineralization in soils. Global Change Biol 27:5950. https://doi.org/10.1111/gcb.15851
Elrys AS, Wang J, Metwally MA, Cheng Y, Zhang J, Cai Z, Chang SX, Müller C (2021c) Global gross nitrification rates are dominantly driven by soil carbon-to-nitrogen stoichiometry and total nitrogen. Global Change Biol 27:6512. https://doi.org/10.1111/gcb.15883
Elrys AS, Chen Z, Wang J, Uwiragiye Y, Helmy AM, Desoky EM, Cheng Y, Zhang J, Cai Z, Müller C (2022) Global patterns of soil gross immobilization of ammonium and nitrate in terrestrial ecosystems. Global Change Biol 00:1–17. https://doi.org/10.1111/gcb.16202
Erisman JW, Sutton MA, Galloway J, Klimont Z, Winiwarter W (2008) How a century of ammonia synthesis changed the world. Nat Geosci 1: 636–639. https://www.nature.com/articles/ngeo325
EU Nitrogen Expert Panel (2015) Nitrogen Use Efficiency - an indicator for the utilization of nitrogen in agriculture and food systems. Wageningen University, Alterra, PO Box 47, NL-6700 Wageningen, Netherlands. www.eunep.com
Fan XR, Xie D, Chen JG (2014) Over-expression of OsPTR6, in rice increased plant growth at different nitrogen supplies but decreased nitrogen use efficiency at high ammonium supply. Plant Sci 227:1–11. https://doi.org/10.1016/j.plantsci.2014.05.013
Fan XR, Feng HM, Tan YW (2016a) A putative 6-transmembrane nitrate transporter OsNRT1.1b plays a key role in rice under low nitrogen. J Integr Plant Biol 58:590–599. https://doi.org/10.1111/jipb.12382
Fan XR, Tang Z, Tan YW (2016b) Overexpression of a pH-sensitive nitrate transporter in rice increases crop yields. Proc Natl Acad Sci 113:71187123. https://doi.org/10.1073/pnas.1525184113
Feng HM, Yan M, Fan XR (2011) Spatial expression and regulation of rice high- affinity nitrate transporters by nitrogen and carbon status. J Exp Bot 62:2319–2332. https://doi.org/10.1093/jxb/erq403
Fischer JJ, Beatty PH, Good AG, Muench DG (2013) Manipulation of microRNA expression to improve nitrogen use efficiency. Plant Sci 210:70–81. https://doi.org/10.1016/j.plantsci.2013.05.009
Foley JA, Ramankutty N, Brauman KA, Cassidy ES, Gerber JS et al (2011) Solutions for a cultivated planet. Nature 478:337–342. https://doi.org/10.1038/nature10452
Fontaine JX, Ravel C, Pageau K, Heumez E, Dubois F et al (2009) A quantitative genetic study for elucidating the contribution of glutamine synthetase, glutamate dehydrogenase and other nitrogen-related physiological traits to the agronomic performance of common wheat. Theor Appl Genet 119:645–662. https://doi.org/10.1007/s00122-009-1076-4
Forde BG, Lea PJ (2007) Glutamate in plants: Metabolism, regulation, and signalling. J Exp Bot 58:2339–2358. https://doi.org/10.1093/jxb/erm121
Franzen D, Long D, Sims A, Lamb J, Casey F, Staricka J, Halvorson M, Hofman V (2011) Evaluation of methods to determine residual soil nitrate zones across the northern Great Plains of the USA. Precis Agric 12:594–606. https://doi.org/10.1007/s11119-010-9207-0
Frey S, Six J, Elliott E (2003) Reciprocal transfer of carbon and nitrogen by decomposer fungi at the soil–litter interface. Soil Biol Biochem 35:1001–1004. https://doi.org/10.1016/S0038-0717(03)00155-X
Fukushima A, Kusano M (2014) A network perspective on nitrogen metabolism from model to crop plants using integrated ‘omics’ approaches. J Exp Bot 65:5619–5630. https://doi.org/10.1093/jxb/eru322
Gadaleta A et al (2014) Isolation and characterization of cytosolic glutamine synthetase (GSe) genes and association with grain protein content in durum wheat. Crop Pasture Sci 65:38–45. https://doi.org/10.1071/CP13140
Galloway JN, Cowling EB (2021) Reflections on 200 years of nitrogen, 20 years later. Ambio 50:745–749. https://doi.org/10.1007/s13280-020-01464-z
Galloway JN, Dentener FJ, Capone DG, Boyer EW, Howarth RW, Seitzinger SP, Asner GP, Cleveland CC, Green PA, Holland EA, Karl DM (2004) Nitrogen cycles: past, present and future. Biogeochem 70: 153–226. https://springerlink.fh-diploma.de/article/10.1007/s10533-004-0370-0
Gao R, Guo G, Fang C, Huang S, Chen J, Lu R, Huang J, Fan X, Liu C (2018) Rapid generation of barley mutant lines with high nitrogen uptake efficiency by microspore mutagenesis and field screening. Front Plant Sci 9:450. https://doi.org/10.3389/fpls.2018.00450
Gelli M, Duo Y, Anji RK, Zhang C, Holding D, Dweikat I (2014) Identification of differentially expressed genes between sorghum genotypes with contrasting nitrogen stress tolerance by genome-wide transcriptional profiling. Genomics 15:179. http://www.biomedcentral.com/1471-2164/15/179
Giller KE, Chalk PM, Dobermann A, Hammond L, Hever P, Ladha JK, Maene L, Nyamudeza P, Ssali H, Freney JR (2004) Emerging technologies to increase the efficiency of use of fertilizer nitrogen. In “Agriculture and the nitrogen cycle: assessing the impacts of fertilizer use on food production and the environment”. AR Mosier, JK Syers, and JR Freney (eds). Paris, France 65:35–51
Godfray HC, Beddington JR, Crute IR, Haddad L (2010) Food security: the challenge of feeding 9 billion people. Science 327:812–818. https://doi.org/10.1126/science.1185383
Goodall AJ et al (2013) Identification and expression analyses of cytosolic glutamine synthetase genes in barley (Hordeum vulgare L.). Plant Cell Physiol 54:494–505. https://doi.org/10.1093/pcp/pct006
Van Grinsven HJM, Spiertz JHJ, Westhoek HJ, Bouwman AF, Erisman JW (2013) Nitrogen workshop special issue paper nitrogen use and food production in European regions from a global perspective. J Agric Sci 9-19.https://doi.org/10.1017/S0021859613000853
Gui R, Mo Z, Zeng S, Wen Z, Long W (2022) Effects of mechanized, deep application of slow-release fertilizer on yield and nitrogen, phosphorus, and potassium utilization of direct-seeded rice. J Plant Growth Regul 1–10. https://doi.org/10.1007/s00344-022-10645-x
Habash DZ, Bernard S, Schondelmaier J, Weyen J, Quarrie SA (2007) The genetics of nitrogen use inhexaploid wheat: N utilization, development and yield. Theor Appl Genet 114:403–419. https://doi.org/10.1007/s00122-006-0429-5
Halvorson AD, Del Grosso SJ (2013) Nitrogen placement and source effects on nitrous oxide emissions and yields of irrigated corn. J Environ Qual 42:312–322. https://doi.org/10.2134/jeq2012.0315
Han M, Okamoto M, Beatty PH, Rothstein SJ, Good AG (2015) The genetics of nitrogen use efficiency in crop plants. Ann Rev Genet 49:269–289. https://doi.org/10.1146/annurev-genet-112414-055037
Han M, Wong J, Su T, Beatty PH, Good AG (2016a) Identification of nitrogen use efficiency genes in barley: searching for QTLs controlling complex physiological traits. Front Plant Sci 7:1587. https://doi.org/10.3389/fpls.2016.01587
Han P, Zhang W, Wang G, Sun W, Huang Y (2016b) Changes in soil organic carbon in croplands subjected to fertilizer management: a global meta-analysis. Scientific Reports 6: 27199. https://www.nature.com/articles/srep27199?origin=ppub
Hanke GT, Endo T, Satoh F, Hase T (2008) Altered photosynthetic electron channeling into cyclic electron flow and nitrite assimilation in a mutant of ferredoxin: NADP(H) reductase. Plant Cell Environ 31:1017–1028. https://doi.org/10.14841/jspp.2008.0.0112.0
Hao G, Stover E, Gupta G (2016) Overexpression of a modified plant thionin enhances disease resistance to citrus canker and huanglongbing (HLB). Front Plant Sci 7:1078. https://doi.org/10.3389/fpls.2016.01078
Hawkesford MJ (2011) An overview of nutrient use efficiency and strategies for crop improvement. In: The molecular and physiological basis of nutrient use efficiency in crops. Wiley-Blackwell 3–19. https://doi.org/10.1002/9780470960707.ch1
Heffer P (2013) Assessment of fertilizer use by crops at the global level 2010–2010/11. International Fertilizer Industry Association. Paris France 9
Heffer P, Prud’homme M (2016) Global nitrogen fertilizer demand and supply: trend, current level and outlook. Proceedings of the 2016 International Nitrogen Initiative Conference, "Solutions to improve nitrogen use efficiency for the world", 4 – 8 December 2016, Melbourne, Australia. www.ini2016.com.
Heidlebaugh NM, Trethewey BR, Jukanti AK, Parrott DL, Martin JM, Fischer AM (2008) Effects of a barley (Hordeum vulgare L.) chromosome 6 grain protein content locus on whole-plant nitrogen reallocation under two different fertilization regimes. Funct Plant Biol 35:619–632. https://doi.org/10.1071/FP07247
Hill C, Taylor J, Edwards J, Matherm D (2013) Whole-genome mapping of agronomic and metabolic traits to identify novel quantitative trait loci in bread wheat grown in a water-limited environment. Plant Physiol 162:1266–1281. https://doi.org/10.1104/pp.113.217851
Hirel B, Lea PJ (2011) The molecular genetics of nitrogen use efficiency in crops. In: Hawkesford MJ, Barraclough PB, eds. The molecular and physiological basis of nutrient use effciency in crops. Wiley-Blackwell, Chichester 139–164
Hirner A, Ladwig F, Stransky H, Okumoto S, Keinath M, Harms A, Frommer WB, Koch W (2006) Arabidopsis LHT1 is a high-affinity transporter for cellular amino acid uptake in both root epidermis and leaf mesophyll. Plant Cell 18:1931–1946. https://doi.org/10.1105/tpc.106.041012
Högberg MN, Högberg P, Myrold DD (2007) Is microbial community composition in boreal forest soils determined by pH, C-to-N ratio, the trees, or all three? Oecologia 150:590–601. https://doi.org/10.1007/s00442-006-0562-5
Hou PF, Yuan WS, Li GH, Petropoulos E, Xue LX, Feng YY, Xue LH, Yang LZ, Ding YF (2021) Deep fertilization with controlled release fertilizer for higher cereal yield and N utilization in paddies: the optimal fertilization depth. Agron J 113:5027–5039. https://doi.org/10.1002/agj2.20772
Houlton BZ, Almaraz M, Aneja V, Austin AT, Bai E, Cassman KG, Compton JE, Davidson EA, Erisman JW, Galloway JN, Gu B (2019) A world of cobenefits: solving the global nitrogen challenge. Earth’s Future 7:865–872. https://doi.org/10.1029/2019EF001222
Hsieh MH, Lam HM, van de Loo FJ, Coruzzi G (1998) A PII-like protein in 1207 Arabidopsis: putative role in nitrogen sensing. Proc Nat Acad Sci 95(13965–13970):1208. https://doi.org/10.1073/pnas.95.23.13965
Hu B, Wang W, Ou SJ (2015) Variation in NRT1.1B contributes to nitrate-use divergence between rice subspecies. Nat Genet 47:834–838. https://doi.org/10.1038/ng.3337
Hu R, Qiu DY, Chen Y (2016) Knock-down of a tonoplast localized low-affinity nitrate transporter OsNPF7.2 affects rice growth under high nitrate supply. Frontiers in Plant Sci 7:1529. https://doi.org/10.3389/fpls.2016.01529
Huang R, Han M, Meng L, Chen X (2018) Transcriptome-wide discovery of coding and noncoding RNA-binding proteins. PNAS April 24:3879-E3887. https://doi.org/10.1073/pnas.1718406115
Hutchings NJ, Sørensen P, Cordovil CM, Leip A, Amon B (2020) Measures to increase the nitrogen use efficiency of European agricultural production. Glob Food Sec 26:100381. https://doi.org/10.1016/j.gfs.2020.100381
Huygens D, Rütting T, Boeckx P, Van Cleemput O, Godoy R, Müller C (2007) Soil nitrogen conservation mechanisms in a pristine south Chilean Nothofagus forest ecosystem. Soil Biol Biochem 39:2448–2458. https://doi.org/10.1016/j.soilbio.2007.04.013
IFA (2022) Nitrogen statistics from 1961–2019. IFADATA Statistics, France http://ifadata.fertilizer.org/ucSearch.aspx
Ilany T, Ashton MS, Montagnini F, Martinez C (2010) Using agroforestry to improve soil fertility: effects of intercropping on Ilex paraguariensis (yerba mate) plantations with Araucaria angustifolia. Agroforest Syst 80:399–409. https://doi.org/10.1007/s10457-010-9317-8
Ishiyama K, Inoue E, Tabuchi M (2004) Biochemical background and compartmentalized functions of cytosolic glutamine synthetase for active ammonium assimilation in rice roots. Plant Cell Physiol 45:1640–1647. https://doi.org/10.1093/pcp/pch190
Ivanov A, Kameka A, Pajak A, Bruneau L, Beyaert R, Hernández-Sebastià C, Marsolais F (2012) Arabidopsis mutants lacking asparaginases develop normally but exhibit enhanced root inhibition by exogenous asparagine. Amino Acids 42:2307–2318. https://doi.org/10.1007/s00726-011-0973-4
Jose S (2009) Agroforestry for ecosystem services and environmental benefits: an overview. Agroforest Syst 76:1–10. https://doi.org/10.1007/s10457-009-9229-7
Kaur B, Gupta S, Singh G (2000) Soil carbon, microbial activity and nitrogen availability in agroforestry systems on moderately alkaline soils in northern India. Appl Soil Ecol 15:283–294. https://doi.org/10.1016/S0929-1393(00)00079-2
Khademi S, O’Connell J, Remism J (2004) Mechanism of ammonia transport by amt/mep/rh: Structure of amtb at 1.35 A. Sci 305:1587–1594. https://doi.org/10.1126/science.1101952
Kindu GA, Tang J, Yin X, Struik PC (2014) Quantitative trait locus analysis of nitrogen 1254 use efficiencyin barley (Hordeum vulgare L.). Euphytica 199(207–21):1255. https://doi.org/10.1007/s10681-014-1138-9
Kraft B, Tegetmeyer HE, Sharma R, Klotz MG, Ferdelman TG, Hettich RL, Geelhoed JS, Strous M (2014) The environmental controls that govern the end product of bacterial nitrate respiration. Science 345:676–679. https://doi.org/10.1126/science.1254070
Krapp A, David LC, Chardin C, Girin T, Marmagne A, Leprince AS, Chaillou S, Ferrario-Mery S, Meyer C, Daniel-Vedele F (2014) Nitrate transport and signalling in Arabidopsis. J ExpBot 65:789–798. https://doi.org/10.1093/jxb/eru001
Kusano M, Tabuchi M, Fukushima A (2011) Metabolomics data reveal a crucial role of cytosolic glutamine synthetase 1; 1 in coordinating metabolic balance in rice. Plant J 66:456–466. https://doi.org/10.1111/j.1365-313X.2011.04506.x
Lai TV, Ryder MH, Rathjen JR, Bolan NS, Croxford AE, Denton MD (2021) Dissimilatory nitrate reduction to ammonium increased with rising temperature. Biol Fertil Soils 57:363–372. https://doi.org/10.1007/s00374-020-01529-x
Lam HM, Wong P, Chan HK, Yam KM, Chen L, Chow CM, Coruzzi GM (2003) Overexpression of the ASN1 gene enhances nitrogen status in seeds of Arabidopsis. Plant Physiol 132:926–935. https://doi.org/10.1104/pp.103.020123
Lang M, Li P, Ti C, Zhu S, Yan X, Chang SX (2019) Soil gross nitrogen transformations are related to land-uses in two agroforestry systems. Ecol Eng 127:431–439. https://doi.org/10.1016/j.ecoleng.2018.12.022
Laperche A, Brancourt-Hulmel M, Heumez E, Gardet O, Hanocq E et al (2007) Using genotype x nitrogen interaction variables to evaluate the QTL involved in wheat tolerance to nitrogen constraints. Theor Appl Genet 115:399–415. https://doi.org/10.1007/s00122-007-0575-4
Lassaletta L, Billen G, Grizzetti B, Anglade J, Garnier J (2014) 50-year trends in nitrogen use efficiency of world cropping systems: the relationship between yield and nitrogen input to cropland Environ. Res Lett 9:105011. https://doi.org/10.1088/1748-9326/9/10/105011
Lea US, Leydecker MT, Quilleré I, Meyer C, Lillo C (2006) Posttranslational regulation of nitrate reductase strongly affects the levels of free amino acids and nitrate, 1282 whereas transcriptional regulation has only minor influence. Plant Physiol 140:1085–1094. https://doi.org/10.1104/pp.105.074633
Legay N, Clément JC, Grassein F, Lavorel S, Lemauviel-Lavenant S, Personeni E et al (2020) Plant growth drives soil nitrogen cycling and N-related microbial activity through changing root traits. Fungal Ecol 44:100910. https://doi.org/10.1016/J.FUNECO.2019.100910
Léran S, Varala K, Boyer J (2013) A unified nomenclature of nitrate transporter 1/peptide transporter family members in plants. Trends in Plant Scie 19:5–9. https://doi.org/10.1016/j.tplants.2013.08.008
Li S, Zhao B, Yuan D, Duan M, Qian Q, Tang L, Wang B, Liu X, Zhang J, Wang J, Sun J, Liu Z, Feng YQ, Yuan L, Li C (2013) Rice zinc finger protein DST enhances grain production through controlling Gn1a/OsCKX2 expression. Proc Nat Acad Sci 110:3167–3172. https://doi.org/10.1073/pnas.1300359110
Li P, Chen F, Cai H, Liu J, Pan Q, Liu Z, Gu R, Mi G, Zhang F, Yuan L (2015) A genetic relationship between nitrogen use efficiency and seedling root traits in maize as revealed by QTL analysis. J Exp Bot 66:3175–3188. https://doi.org/10.1093/jxb/erv127
Li H, Hu B, Chu C (2017) Nitrogen use efficiency in crops: lessons from Arabidopsis and rice. J Exp Bot 68:2477–2488. https://doi.org/10.1093/jxb/erx101
Li T, Zhang W, Yin J, Chadwick D, Norse D, Lu Y et al (2018) Enhanced-efficiency fertilizers are not a panacea for resolving the nitrogen problem. Glob Change Biol 24:511–521. https://doi.org/10.1111/gcb.13918
Li X, He H, Zhang X, Yan X, Six J, Cai Z et al (2019) Distinct responses of soil fungal and bacterial nitrate immobilization to land conversion from forest to agriculture. Soil Biol Biochem 134:81–89. https://doi.org/10.1016/j.soilbio.2019.03.023
Li X, Li Z, Zhang X, Xia L, Zhang W, Ma Q, He H (2020) Disentangling immobilization of nitrate by fungi and bacteria in soil to plant residue amendment. Geoderma 374:114450. https://doi.org/10.1016/j.geoderma.2020.114450
Li H, Mei X, Nangia V, Guo R, Liu Y, Hao W, Wang J (2021) Effects of different nitrogen fertilizers on the yield, water-and nitrogen-use efficiencies of drip-fertigated wheat and maize in the North China Plain. Agric Water Manag 243:106474. https://doi.org/10.1016/j.agwat.2020.106474
Li YF, Li MM, Cao GL, Han LZ (2010) Effects of genetic background on expression of QTL for nitrogen efficiency in irrigated rice and upland rice. Sci Agric Sin 43: 4331–4340. https://www.cabdirect.org/cabdirect/abstract/20113004850
Lian X, Wang S, Zhang J, Feng Q, Zhang L, Fan D, Li X, Yuan D, Han B, Zhang Q (2006) Expression profiles of 10, 422 genes at early stage of low nitrogen stress in rice assayed using a cDNA microarray. Plant Mol Biol 60:617–631. https://doi.org/10.1007/s11103-005-5441-7
Liang HL, Gao SY, Ma JX, Zhang T, Wang TY, Zhang S, Wu Z (2021) Effect of nitrogen application rates on the nitrogen utilization, yield and quality of rice. Food Nutr Sci 12:13–27. https://doi.org/10.4236/fns.2021.121002
Lin HC, Huber JA, Gerl G, Hülsbergen KJ (2016) Nitrogen balances and nitrogen-use efficiency of different organic and conventional farming systems. Nutr Cycl Agroecosyst 105:1–23. https://doi.org/10.1007/s10705-016-9770-5
Liu C, Wang K, Zheng X (2013) Effects of nitrification inhibitors (DCD and DMPP) on nitrous oxide emission: crop yield and nitrogen uptake in a wheat-maize cropping system. Biogeosciences 10:2427–2437. https://doi.org/10.5194/bg-10-2427-2013
Liu R, Zhang H, Zhao P, Zhang Z, Liang W et al (2012) Mining of candidate maize genes for nitrogen use efficiency by integrating gene expression and QTL data. Plant Mol Biol Rep 30:297–308. https://doi.org/10.1007/s11105-011-0346-x
Liu X, Xu W, Pan Y, Du ELiu, et al (2015) suspect that Zhu et al. (2015) may have underestimated dissolved organic nitrogen (N) but overestimated total particulate N in wet deposition in China. Sci Total Environ 1:300–301. https://doi.org/10.1016/j.scitotenv
Liu XD, Chen LY, Hua ZL, Mei SC, Wang P, Wang SW (2020) Comparing ammonia volatilization between conventional and slow-release nitrogen fertilizers in paddy fields in the Taihu Lake region. Environ Sci Pollut R 27:8386–8394. https://doi.org/10.1007/s11356-019-07536-2
Lohse T, Riedel N, Spengel C (2012) The increasing importance of transfer pricing regulations? A worldwide overview, Oxford University Centre for Business Taxation Working Paper 12/27. https://kluwerlawonline.com/journalarticle/Intertax/42.6/TAXI2014038
Lu C, Tian H (2017) Global nitrogen and phosphorus fertilizer use for agriculture production in the past half century: shifted hot spots and nutrient imbalance. Earth Syst Sci Data 9:181–192. https://doi.org/10.5194/essd-9-181-2017
Lu Y, Zhang X, Jiang J et al (2018) Effects of the biological nitrification inhibitor 1,9-decanediol on nitrification and ammonia oxidizers in three agricultural soils. Soil Biol Biochem 129:48–59. https://doi.org/10.1016/j.soilbio.2018.11.008
Lucas GN (2013) Dicyandiamide contamination of milk powders. Sri Lanka Journal of Child Health 42:63–64. https://doi.org/10.4038/sljch.v42i2.5624
Ma D, Yin L, Liu Y, Yang W, Deng X, Wang S (2020) A Meta-analysis of the effects of nitrogen application rates on yield and water use efficiency of winter wheat in dryland of Loess Plateau[J]. Sci Agric Sin 53:486–499. https://doi.org/10.3864/j.issn.0578-1752.2020.03.003
Ma Y, Jones DL, Wang J, Cardenas LM, Chadwick DR (2021) Relative efficacy and stability of biological and synthetic nitrification inhibitors in a highly nitrifying soil: Evidence of apparent nitrification inhibition by linoleic acid and linolenic acid. Eur J Soil Sci 72:2356–2371. https://doi.org/10.1111/ejss.13096
Marsden KA, Scowen M, Hill, et al (2015) Plant acquisition and metabolism of the synthetic nitrification inhibitor dicyandiamide and naturally-occurring guanidine from agricultural soils. Plant Soil 395:201–214. https://doi.org/10.1007/s11104-015-2549-7
Martinez F, Palencia P, Weiland CM, Alonso D, Oliveira JA (2015) Influence ofnitrification inhibitor DMPP on yield: fruit quality and SPAD values ofstrawberry plants. Sci Hortic 185:233–239. https://doi.org/10.1016/j.scienta.2015.02.004
Masclaux-Daubresse C, Daniel-Vedele F, Dechorgnat J, Chardon F, Gaufichon L, Suzuki A (2010) Nitrogen uptake, assimilation and remobilization in plants: challenges for sustainable and productive agriculture. Ann Bot 105:1141–1157. https://doi.org/10.1093/aob/mcq028
Masle J, Hudson GS, Badger MR (1993) Effects of ambient CO2 concentration on growth and nitrogen use in tobacco (Nicotiana tabacum) Plants transformed with an antisense gene to the small subunit of Ribulose-1, 5-Bisphosphate Carboxylase/Oxygenase. Plant Physiol 103:1075–1088. https://doi.org/10.1104/pp.103.4.1075
Massel K, Campbell BC, Mace ES, Tai S, Tao Y, Worland BG, Jordan DR, Botella JR, Godwin ID (2016) Whole Genome Sequencing Reveals Potential New Targets for Improving Nitrogen Uptake and Utilization in Sorghum bicolor. Front Plant Sci 7:1544. https://doi.org/10.3389/fpls.2016.01544
Merwad AMA, Desoky EM, Rady MM (2018) Response of water deficit-stressed Vigna unguiculata performances to silicon, proline or methionine foliar application. Sci Hortic 228:132–144
Mickelson S, See D, Meyer FD, Garner JP, Foster CR et al (2003) Mapping of QTL associated with nitrogen storage and remobilization in barley (Hordeum vulgare L.) leaves. J Exp Bot 54:801–812. https://doi.org/10.1093/jxb/erg084
Miller AC, Seymour H, King C, Herman TG (2008) Loss of seven-up from Drosophila R1/R6 photoreceptors reveals a stochastic fate choice that is normally biased by Notch. Development 135:707–715. https://doi.org/10.1242/dev.016386
Moir JL, Malcolm BJ, Cameron KC, Di HJ (2012) The effect of dicyandiamide on pasture nitrate concentration, yield and N off take under high N loading in winter and spring. Grass Forage Sci 67:391–402. https://doi.org/10.1111/j.1365-2494.2012.00857.x
Moreau D, Bardgett RD, Finlay RD, Jones DL, Philippot L (2019) A plant perspective on nitrogen cycling in the rhizosphere. Funct Ecol 33:540–552. https://doi.org/10.1111/1365-2435.13303
Moulos P, Hatzis P (2015) Systematic integration of RNA-Seq statistical algorithms for accurate detection of differential gene expression patterns. Nucleic Acids Res 43(4):e25. https://doi.org/10.1093/nar/gku1273
Nasielski J, Grant B, Smith W, Niemeyer C, Janovicek K, Deen B (2020) Effect of nitrogen source, placement and timing on the environmental performance of economically optimum nitrogen rates in maize. Field Crop Res 246:107686. https://doi.org/10.1016/j.fcr.2019.107686
Neeraja CN, Subramanyam D, Surekha K (2016) Advances in genetic basis of nitrogen use efficiency of rice. Indian J Plant Physiol 21:504–513. https://doi.org/10.1007/s40502-016-0254-z
Nemati AR, Sharifi RS (2012) Effects of rates and nitrogen application timing on yield, agronomic characteristics and nitrogen use efficiency in corn. Intl. J Agri Crop Sci 4: 534–539. https://www.cabdirect.org/cabdirect/abstract/20123364478
Nojiri Y, Kaneko Y, Azegami Y, Shiratori Y, Ohte N, Senoo K et al (2020) Dissimilatory nitrate reduction to ammonium and responsible microbes in japanese rice paddy soil. Microbes Environ 35:ME20069. https://doi.org/10.1264/jsme2.ME20069
Norton R, Davidson E, Roberts T (2015) Nitrogen use efficiency and nutrient performance indicators. Global Partnership on Nutrient Management (GPNM). https://stg-wedocs.unep.org/handle/20.500.11822/10750
Obara M, Kajiura M, Fukuta Y, Yano M, Hayashi M et al (2001) Mapping of QTLs associated with cytosolic glutamine synthetase and NADH-glutamate synthase in rice (Oryza sativa L.). J Exp Bot 52:1209–1217. https://doi.org/10.1093/jexbot/52.359.1209
Ogorodnikov A, Kargapolova Y, Danckwardt S (2016) Processing and transcriptome expansion at the mRNA 3′ end in health and disease: finding the right end. Pflugers Arch – Eur. J Physiol 468:993–1012. https://doi.org/10.1007/s00424-016-1828-3
Ohashi M, Ishiyama K, Kojima S (2015) Asparagine synthetase1, but not asparagine synthetase 2, is responsible for the biosynthesis of asparagine following the supply of ammonium to rice roots. Plant Cell Physiol 56:769–778. https://doi.org/10.1093/pcp/pcv005
Okumoto S, Pilot G (2011) Amino acid export in plants: a missing link in nitrogen cycling. Mol Plant 4:453–463. https://doi.org/10.1093/mp/ssr003
Oliveira IC et al (2002) Overexpression of cytosolic glutamine synthetase Relation to nitrogen, light, and photorespiration. Plant Physiol 129:1170–1180. https://doi.org/10.1104/pp.020013
Orsel M et al (2014) Sixteen cytosolic glutamine synthetase genes identified in the Brassica napus L. genome are differentially regulated depending on nitrogen regimes and leaf senescence. J Exp Bot 65:3927–3947. https://doi.org/10.1093/jxb/eru041
Ouyang J, Cai ZY, Xia KF (2010) Identification and analysis of eight peptide transporter homologs in rice. Plant Scie 179:374–382. https://doi.org/10.1016/j.plantsci.2010.06.013
Pan H, Qin Y, Wang Y, Liu S, Yu B, Song Y et al (2020) Dissimilatory nitrate/nitrite reduction to ammonium (DNRA) pathway dominates nitrate reduction processes in rhizosphere and non-rhizosphere of four fertilized farmland soil. Environ Res 186:109612. https://doi.org/10.1016/j.envres.2020.109612
Pandey A, Suter H, He JZ, Hu HW, Chen D (2018) Nitrogen addition decreases dissimilatory nitrate reduction to ammonium in rice paddies. Appl Environ Microbiol 84:e00870-e918. https://doi.org/10.1128/AEM.00870-18
Pandey C, Kumar U, Kaviraj M, Minick K, Mishra A, Singh J (2020) DNRA: A short-circuit in biological N-cycling to conserve nitrogen in terrestrial ecosystems. Sci Total Environ 738:139710. https://doi.org/10.1016/j.scitotenv.2020.139710
Park S-Y, Yu J-W, Park J-S, Li J, Yoo S-C, Lee N-Y, Lee S-K, Jeong S-W, Seo HS, Koh H-J, Jeon J-S, Park Y-I, Paek N-C (2007) The senescence-induced staygreen protein regulates chlorophyll degradation. Plant Cell 19:1649–1664. https://doi.org/10.1105/tpc.106.044891
Patra AK (2013) Tree–crop interaction in agroforestry. Sci Horiz 4:12–17
Patra DD, Chand S, Anwar M, et al (2009) Use of urea coated with natural products to inhibit urea hydrolysis and nitrification in soil. Patra DD, Kiran U, Chand S, et al Biol Fertil Soils 45: 617. https://doi.org/10.1007/s00374-009-0372-0
Peng M, Bi YM, Zhu T, Rothstein SJ (2007) Genome-wide analysis of Arabidopsis responsive transcriptome to nitrogen limitation and its regulation by the ubiquitin ligase gene NLA. Plant Mol Biol 65:775–797. https://doi.org/10.1007/s11103-007-9241-0
Peng-Fei D (2019) Response of maize genotypes with different nitrogen use efficiency to low nitrogen stresses. Acta Ecol Sin 39:77–80. https://doi.org/10.1016/j.chnaes.2018.05.005
Pinton R, Tomasi N, Zanin L (2016) Molecular and physiological interactions of urea and nitrate uptake in plants. Plant Signal Behav 11:e1076603. https://doi.org/10.1080/15592324.2015.1076603
Potter P, Ramankutty N, Bennett EM, Donner SD (2011) Global fertilizer and manure, version 1: nitrogen fertilizer application. http://sedac.ciesin.columbia.edu/data/set/ferman-v1-nitrogen-fertilizer-application/metadata Accessed 16 Feb. 2015
Purakayastha TJ, Katyal JC, Goswami NN et al (1997) Evaluation of some modified urea fertilizers applied to rice. Fertilizer News 42:53–56
Quan Z, Zhang X, Fang Y, Davidson EA (2021) Different quantification approaches for nitrogen use efficiency lead to divergent estimates with varying advantages. Nature Food 2:241–245. https://doi.org/10.1038/s43016-021-00263-3
Quraishi UM, Abrouk M, Murat F, Pont C, Foucrier S, Desmaizieres G, Confolent C, Riviere N, Charmet G, Paux E, Murigneux A, Guerreiro L, Lafarge S, Le Gouis J, Feuillet C, Salse J (2011) Cross-genome map based dissection of a nitrogen use efficiency ortho-meta QTL in bread wheat unravels concerted cereal genome evolution. Plant J 65:745–756. https://doi.org/10.1111/j.1365-313X.2010.04461.x
Ramm E, Liu C, Ambus P, Butterbach-Bahl K, Hu B, Martikainen PJ, Marushchak ME, Mueller CW, Rennenberg H, Schloter M, Siljanen HM (2022) A review of the importance of mineral nitrogen cycling in the plant-soil-microbe system of permafrost-affected soils–changing the paradigm. Environ Res Lett. EGU22. https://doi.org/10.1088/1748-9326/ac417e
Rauschenberger A, Jonker MA, van de Wiel MA, Menezes RX (2016) Testing for association between RNA-Seq and high-dimensional data. BMC Bioinformatics 17:118. https://doi.org/10.1186/s12859-016-0961-5
Raviv M (2010) The use of mycorrhiza in organically-grown crops under semi-arid conditions: A review of benefits, constraints and future challenges. Symbiosis 52:65–74. https://doi.org/10.1007/s13199-010-0089-8
Raza S, Chen Z, Ahmed M et al (2018) Dicyandiamide application improved nitrogen use efficiency and decreased nitrogen losses in wheat-maize crop rotation in Loess Plateau. Arch Agron Soil Sci 65:4. https://doi.org/10.1080/03650340.2018.1506584
Raza S, Jiang Y, Elrys AS, Tao J, Liu Z, Li Z et al (2021) Dicyandiamide efficacy of inhibiting nitrification and carbon dioxide emission from calcareous soil depends on temperature and moisture contents. Archives Agron Soil Sci 1–17. https://doi.org/10.1080/03650340.2021.1895432
Recous S, Mary B, Faurie G (1990) Microbial immobilization of ammonium and nitrate in cultivated soils. Soil Biol Biochem 22:913–922. https://doi.org/10.1016/0038-0717(90)90129-N
Reddy MM, Ulaganathan K (2015) Nitrogen nutrition, its regulation and biotechnological approaches to improve crop productivity. Am J Plant Sci 06:2745–2798. https://doi.org/10.4236/ajps.2015.618275
Reich PB, Hobbie SE, Lee T, Ellsworth DS, West JB, Tilman D, Trost J (2006) Nitrogen limitation constrains sustainability of ecosystem response to CO2. Nature 440:922–925. https://doi.org/10.1038/nature04486
Rochette P, Angers DA, Chantigny MH, Gasser MO, MacDonald JD, Pelster DE, Bertrand N (2013) Ammonia volatilization and nitrogen retention: how deep to incorporate urea? J Environ Qual 42:1635–1642. https://doi.org/10.2134/jeq2013.05.0192
Rolletschek H, Hosein F, Miranda M, Heim U, Gotz KP, Schlereth A, Borisjuk L, Saalbach I, Wobus U, Weber H (2005) Ectopic expression of an amino acid transporter (VfAAP1) in seeds of Vicia narbonensis and pea increases storage proteins. Plant Physiol 137:1236–1249. https://doi.org/10.1104/pp.104.056523
Rubio-Wilhelmi, del MM, Sanchez-Rodriguez E, Rosales MA, Blasco B, Rios JJ, Romero L, Ruiz JM (2011) Cytokinin-dependent improvement in transgenic psark::ipt tobacco under nitrogen deficiency. J Agric Food Chem 59(19):10491–10495. https://doi.org/10.1021/jf202604k
Ruser R, Schulz R (2015) The effect of nitrification inhibitors on the nitrous oxide (N2O) release from agricultural soils: a review. J Plant Nutr Soil Sci 178:171–188
Rütting T, Huygens D, Müller C, Van Cleemput O, Godoy R, Boeckx P (2008) Functional role of DNRA and nitrite reduction in a pristine south Chilean Nothofagus forest. Biogeochemistry 90:243–258. https://doi.org/10.1007/s10533-008-9250-3
Rütting T, Aronsson H, Delin S (2018) Efficient use of nitrogen in agriculture. Nutr Cycl Agroecosyst 110:1–5. https://doi.org/10.1007/s10705-017-9900-8
Rütting T, Boeckx P, Müller C, Klemedtsson L (2011) Assessment of the importance of dissimilatory nitrate reduction to ammonium for the terrestrial nitrogen cycle. Biogeosciences 8:1779–1791. https://doi.org/10.5194/bg-8-1779-2011
Saito K, Matsuda F (2010) Metabolomics for function algenomics, systems biology, and biotechnology. Annu Rev Plant Biol 61:463–489. https://doi.org/10.1146/annurev.arplant.043008.092035
Sato T, Maekawa S, Yasuda S, Domeki Y, Sueyoshi K, Fujiwara M, Fukao Y, Goto DB, Yamaguchi J (2011) Identification of 14–3-3 proteins as a target of ATL31 40 ubiquitin ligase, a regulator of the C/N response in Arabidopsis. Plant J 68:137–146. https://doi.org/10.1111/j.1365-313X.2011.04673.x
Schimel JP, Bennett J (2004) Nitrogen mineralization: challenges of a changing paradigm. Ecology 85:591–602. https://doi.org/10.1890/03-8002
Schofield RA, Bi YM, Kant S, Rothstein SJ (2009) Over-expression of STP13, a hexose transporter, improves plant growth and nitrogen use in Arabidopsis thaliana seedlings. Plant, Cell Environ 32:271–285. https://doi.org/10.1111/j.1365-3040.2008.01919.x
Schwanhäusser B et al (2011) Global quantification of mammalian gene expression control. Nature 473:337–342. https://doi.org/10.1038/nature10098.PMID21593866
Senthilvel S, Vinod KK, Malarvizhi P, Maheswaran M (2008) QTL and QTL x environment effects on agronomic and nitrogen acquisition traits in rice. J Integr Plant Biol 50:1108–1117. https://doi.org/10.1111/j.1744-7909.2008.00713.x
Sevel L, Nord-Larsen T, Ingerslev M, Jørgensen U, RaulundRasmussen K (2014) Fertilization of SRC willow, I: biomass production response. Bioenergy Res 7:319–328. https://doi.org/10.1007/s12155-013-9371-y
Shachar-Hill Y (2013) Metabolic network flux analysis for engineering plant systems. Curr Opin Biotechnol 24:247–255. https://doi.org/10.1016/j.copbio.2013.01.004
Shan YH, Wang YL, Pan XB (2005) Mapping of QTLs for nitrogen use efficiency and related traits in rice (Oryza sativa L.). Agric Sci Chin 4:721–727. https://doi.org/10.1007/s10142-016-0486-z
Shi W, Norton JM (2000) Microbial control of nitrate concentrations in an agricultural soil treated with dairy waste compost or ammonium fertilizer. Soil Biol Biochem 32(10):1453–1457. https://doi.org/10.1016/S0038-0717(00)00050-X
Shrawat AK, Carroll RT, DePauw M, Taylor GJ, Good AG (2008) Genetic engineering of improved nitrogen use efficiency in rice by the tissue-specific expression of alanine aminotransferase. Plant Biotechnol J 6:722–732. https://doi.org/10.1111/j.1467-7652.2008.00351.x
Si Z, Zain M, Mehmood F, Wang G, Gao Y, Duan A (2020) Effects of nitrogen application rate and irrigation regime on growth, yield, and water-nitrogen use efficiency of drip-irrigated winter wheat in the North China Plain. Agric Water Manag 231:106002. https://doi.org/10.1016/j.agwat.2020.106002
Silver WL, Herman DJ, Firestone MK (2001) Dissimilatory nitrate reduction to ammonium in upland tropical forest soils. Ecology 82:2410–2416. https://doi.org/10.1890/0012-9658(2001)082[2410:DNRTAI]2.0.CO;2
Simons M, Saha R, Guillard L, Clément G, Armengaud P, Cañas R, Maranas CD, Lea PJ, Hirel B (2014) Nitrogen-use efficiency in maize (Zea mays L.): from ‘omics’ studies to metabolic modelling. J Exp Bot 65:5657–5671. https://doi.org/10.1093/jxb/eru227
Sinclair TR, Rufty TW (2012) Nitrogen and water resources commonly limit crop yield increases, not necessarily plant genetics. Glob Food Sec 1:94–98. https://doi.org/10.1016/j.gfs.2012.07.001
Stahl A, Pfeifer M, Frisch M, Wittkop B, Snowdon RJ (2017) Recent genetic gains in nitrogen use efficiency in oilseed rape. Front Plant Sci 8:963. https://doi.org/10.3389/fpls.2017.00963
Stark JM, Hart SC (1997) High rates of nitrification and nitrate turnover in undisturbed coniferous forests. Nature 385:61–64. https://doi.org/10.1038/385061a0
Subbarao GV, Nakahara K, Ishikawa T, Yoshihashi T, Ito O, Ono H et al (2008) Free fatty acids from the pasture grass Brachiaria humidicola and one of their methyl esters as inhibitors of nitrification. Plant Soil 313:89–99. https://doi.org/10.1007/s11104-008-9682-5
Subbarao GV, Kishii M, Nakahara K, Ishikawa T, Ban T, Tsujimoto H, Ito O (2009a) Biological nitrification inhibition (BNI)-Is there potential for genetic interventions in the Triticeae? Breed Sci 59:529–545
Subbarao GV, Nakahara K, Hurtado MP, Ono H, Moreta DE, Salcedo AF, Yoshihashi AT, Ishikawa T, Ishitani M, Ohnishi-Kameyama M, Yoshida M, Rondon M, Rao IM, Lascano CE, Berry WL, Ito O (2009b) Evidence for biological nitrification inhibition in Brachiaria pastures. Proc Natl Acad Sci U S A 106:17302–17307. https://doi.org/10.1073/pnas.0903694106
Subbarao GV, Nakahara K, Ishikawa T, Ono H, Yoshida M, Yoshihashi T, Sahrawat KL (2013) Biological nitrification inhibition (BNI) activity in sorghum and its characterization. Plant Soil 366:243–259. https://doi.org/10.1007/s11104-012-1419-9
Suenaga A, Moriya K, Sonoda Y (2003) Constitutive expression of a novel-type ammonium transporter OsAMT2 in rice plants. Plant Cell Physiol 44:206–211. https://doi.org/10.1093/pcp/pcg017
Sugimoto M et al (2012) Molecular identification of t (w5): Vps52 promotes pluripotential cell differentiation through cell-cell interactions. Cell Rep 2:1363–1374. https://doi.org/10.1016/j.celrep.2012.10.004
Sun H, Qian Q, Wu K, Luo J, Wang S, Zhang C, Ma Y, Liu Q, Huang X, Yuan Q, Han R, Zhao M, Dong G, Guo L, Zhu X, Gou Z, Wang W, Wu Y, Lin H, Fu X (2014) Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat Genet 46:652–656. https://doi.org/10.1038/ng.2958
Swarbreck SM et al (2011) New perspectives on glutamine synthetase in grasses. J Exp Bot 62:1511–1522. https://doi.org/10.1093/jxb/erq356
Tabuchi M, Sugiyama K, Ishiyama K (2005) Severe reduction in growth rate and grain filling of rice mutants lacking OsGS1; 1, a cytosolic glutamine synthetase1; 1. Plant J Cell Mol Biol 42:641–651. https://doi.org/10.1111/j.1365-313X.2005.02406.x
Takahashi M, Sasaki Y, Ida S, Morikawa H (2001) Nitrite reductase gene enrichment improves assimilation of NO2 in Arabidopsis. Plant Physiol 126:731–741. https://doi.org/10.1104/pp.126.2.731
Tamura W, Hidaka Y, Tabuchi M (2010) Reverse genetics approach to characterize a function of NADH-glutamate synthase1 in rice plants. Amino Acids 39:1003–1012. https://doi.org/10.1007/s00726-010-0531-5
Tamura W, Kojima S, Toyokawa A, Watanabe H, Tabuchi-Kobayashi M, Hayakawa T, Yamaya T (2011) Disruption of a novel NADH-glutamate synthase 2 gene caused marked reduction in spikelet number of rice. Front Plant Sci 2:1–9. https://doi.org/10.3389/fpls.2011.00057
Tang Z, Fan XR, Li Q (2012) Knockdown of a rice stelar nitrate transporter alters long- distance translocation but not root influx. Plant Physiol 160:2052–2063. https://doi.org/10.1104/pp.112.204461
Taveira CJ, Farrell RE, Wagner-Riddle C, Machado PVF, Deen B, Congreves KA (2020) Tracing crop residue N into subsequent crops: Insight from long-term crop rotations that vary in diversity. Field Crop Res 255:107904. https://doi.org/10.1016/j.fcr.2020.107904
Terao T, Nagata K, Morino K, Hirose T (2010) A gene controlling the number of primary rachis branches also controls the vascular bundle formation and hence is responsible to increase the harvest index and grain yield in rice. Theor Appl Genet 120:875–893. https://doi.org/10.1007/s00122-009-1218-8
Teutscherova N, Vazquez E, Arevalo A, Pulleman M, Rao I, Arango J (2019) Differences in arbuscular mycorrhizal colonization and P acquisition between genotypes of the tropical Brachiaria grasses: is there a relation with BNI activity? Biol Fertil Soils 55:325–337. https://doi.org/10.1007/s00374-019-01353-y
Thilakarathna MS, Raizada MN (2018) Challenges in using precision agriculture to optimize symbiotic nitrogen fixation in legumes: progress, limitations, and future improvements needed in diagnostic testing. Agronomy 8:78. https://doi.org/10.3390/agronomy8050078
Thomsen HC, Eriksson D, Møller IS, Schjoerring JK (2014) Cytosolic glutamine synthetase: a target for improvement of crop nitrogen use efficiency? Trends in Plant Sci 19:10. https://doi.org/10.1016/j.tplants.2014.06.002
Ti C, Luo Y, Yan X (2015) Characteristics of nitrogen balance in open-air and greenhouse vegetable cropping systems of China. Environ Sci Pollut Res 22:18508–18518. https://doi.org/10.1007/s11356-015-5277-x
Tiedje JM (1982) Denitrification: ecological niches, competition, and survival. Antonie Leeuwenhoek 48:569. https://doi.org/10.1007/BF00399542
Trenkel ME (2010) Slow- and controlled-release and stabilized fertilizers: an option for enhancing nutrient efficiency in agriculture. IFA, Paris
Tsay YF, Schroeder JI, Feldmann KA, Crawford NM (1993) The herbicide sensitivity gene CHL1 of Arabidopsis encodes a nitrate-inducible nitrate transporter. Cell 72:705–713. https://doi.org/10.1016/0092-8674(93)90399-B
Tsay YF, Chiu CC, Tsai CB (2007) Nitrate transporters and peptide transporters. FEBS Lett 581:2290–2300. https://doi.org/10.1016/j.febslet.2007.04.047
Tschoep H, Gibon Y, Carillo P, Armengaud P, Szecowka M, Nunes-Nesi A, Fernie AR, Koehl K, Stitt M (2009) Adjustment of growth and central metabolism to a mild but sustained nitrogen limitation in Arabidopsis. Plant Cell Environ 32:300–318. https://doi.org/10.1111/j.1365-3040.2008.01921.x
Tuberosa R, Salvi S (2009) QTL for agronomic traits in maize production. In: Bennetzen J, Hake S (eds) Handbook of maize: its biology. Springer LLC 501–542. https://doi.org/10.1007/978-0-387-79418-1_26
Van Den Berg EM, Boleij M, Kuenen JG, Kleerebezem R, van Loosdrecht MCM (2016) DNRA and denitrification coexist over a broad range of acetate/N-NO3− ratios, in a chemostat enrichment culture. Front Microbiol 7:1842. https://doi.org/10.3389/fmicb.2016.01842
Van Kessel C, Venterea R, Six J, Adviento-Borbe MA, Linquist B, van Groenigen KJ (2013) Climate, duration, and N placement determine N2O emissions in reduced tillage systems: a meta-analysis. Glob Chang Biol 19:33–44. https://doi.org/10.1111/j.1365-2486.2012.02779.x
Varshney RK, Ribaut JM, Buckler ES, Tuberosa R, Rafalski JA, Langridge P (2012) Can genomics boost productivity of orphan crops? Nat Biotechnol 30:1172–1176. https://doi.org/10.1038/nbt.2440
Vázquez E, Teutscherova N, Dannenmann M, Töchterle P, Butterbach- Bahl K, Pulleman M, Arango J (2020) Gross nitrogen transformations in tropical pasture soils as affected by Urochloa genotypes differing in biological nitrification inhibition (BNI) capacity. Soil Biol Biochem 151:108058. https://doi.org/10.1016/j.soilbio.2020.108058
Villegas D, Arevalo A, Nuñez J, Mazabel J, Subbarao G, Rao I, Arango J (2020) Biological nitrification inhibition (BNI): phenotyping of a core germplasm collection of the tropical forage grass Megathyrsus maximus under greenhouse conditions. Front Plant Sci 11:820. https://doi.org/10.3389/fpls.2020.00820
Vossenkemper JP (2010) Maize (Zea-maize L.) grain yield response to nitrogen applied at different distances away from the row. Unpublished Master's Thesis, Oklahoma State University.
Wang E, Wang J, Zhu X, Hao W, Wang L, Li Q, Zhang L, He W, Lu B, Lin H, Ma H, Zhang G, He Z (2008) Control of rice grain-filling and yield by a gene with a potential signature of domestication. Nat Genet 40:1370–1374. https://doi.org/10.1038/ng.220
Wang J, Chen Z, Xu C, Elrys AS, Shen F, Cheng Y, Chang SX (2021a) Organic amendment enhanced microbial nitrate immobilization with negligible denitrification nitrogen loss in an upland soil. Environ Pollut 288:117721.117721. https://doi.org/10.1016/j.envpol.2021.117721
Wang X, Bai J, Xie T, Wang W, Zhang G, Yin S, Wang D (2021b) Effects of biological nitrification inhibitors on nitrogen use efficiency and greenhouse gas emissions in agricultural soils: A review. Ecotoxicol Environ Saf 220:112338. https://doi.org/10.1016/j.ecoenv.2021.112338
Wang X, Peng F, Li M, Yang L, Li G (2012) Expression of a heterologous SnRK1 in tomato increases carbon assimilation, nitrogen uptake and modifies fruit development. J Plant Physiol 169:1173–1182. https://doi.org/10.1016/j.jplph.2012.04.013
Wang Z, Gerstein M, Snyder M (2009) RNA-Seq: a revolutionary tool for transcriptomics. Nature Rev Genetics 10:57–63. https://doi.org/10.1038/nrg2484
Wei D, Cui KH, Ye GY, Pan JF, Xiang J, Huang JL et al (2012) QTL mapping for nitrogen-use efficiency and nitrogen-deficiency tolerance traits in rice. Plant Soil 359:281–295. https://doi.org/10.1007/s11104-012-1142-6
Wessels D, Coetzee K (2017) Fertilizer placement and optimal time of application. Omnia Nutriology. http://fertilizer.co.za/public-relations/news/2017/233-fertilizer-placement-and-optimal-time-of-application
Wu S, Wu L, Shi Q, Wang Z, Chen X, Li Y (2007) Effects of a new nitrification inhibitor 3, 4-dimethylpyrazole phosphate (DMPP) on nitrate and potassium leaching in two soils. J Environ Sci 19:841–847. https://doi.org/10.1016/S1001-0742(07)60140-5
Xu GH, Fan XR, Miller AJ (2012) Plant nitrogen assimilation and use efficiency. Annu Rev Plant Biol 63:153–182. https://doi.org/10.1146/annurev-arplant-042811-105532
Xu Y, Wang R, Tong Y, Zhao H, Xie Q et al (2014) Mapping QTLs for yield and nitrogen-related traits in wheat: influence of nitrogen and phosphorus fertilization on QTL expression. Theor Appl Genet 127:59–72. https://doi.org/10.1007/s00122-013-2201-y
Xu Z, Ma J, Qu C, Hu Y, Hao B, Sun Y, Liu Z, Yang H, Yang C, Wang H, Li Y, Liu G (2017) Identification and expression analyses of the alanine aminotransferase (AlaAT) gene family in poplar seedlings. Sci Rep 7:45933. https://doi.org/10.1038/srep45933
Xu J, Zhu T, Xue W, Ni D, Sun Y, Yang J, Liu M (2019) Influences of nitrification inhibitor 3, 4-dimethylpyrazole phosphate (DMPP) and application method on nitrogen dynamics at the centimeter-scale. Eur J Soil Biol 90:44–50. https://doi.org/10.1016/j.ejsobi.2018.12.004
Yadav MR, Kumar R, Parihar CM, Yadav RK, Jat SL, Ram H, Meena RK, Singh M, Birbal AP, Verma U, Kumar AG, Jat ML (2017) Strategies for improving nitrogen use efficiency: a review. Agricultural Reviews 38:29–40. https://doi.org/10.18805/ag.v0iOF.7306
Yamaya T, Kusano M (2014) Evidence supporting distinct functions of three cytosolic glutamine synthetases and two NADH-glutamate synthases in rice. J Exp Bot 65:5519–5525. https://doi.org/10.1093/jxb/eru103
Yan M, Fan XR, Feng HM (2011) Rice OsNAR2.1 interacts with OsNRT2.1, OsNRT2.2 and OsNRT2.3 a nitrate transporter to provide uptake over high and low concentration ranges. Plant, Cell Environ 34:1360–1372. https://doi.org/10.1111/j.1365-3040.2011.02335.x
Yan X et al (2014) Fertilizer nitrogen recovery efficiencies in crop production systems of China with and without consideration of the residual effect of nitrogen. Environ Res Lett 9:095002. https://doi.org/10.1088/1748-9326/9/9/095002
Yan M, Pan G, Lavallee JM, Conant RT (2020) Rethinking sources of nitrogen to cereal crops. Glob Change Biol. 26:191–199. https://doi.org/10.1111/gcb.15463
Yang DL, Jing RL, Chang XP, Li W (2007) Identification of quantitative trait loci and environmental interactions for accumulation and remobilization of water-soluble carbohydrates in wheat (Triticum aestivum L.) stems. Genetics 176:571–584. https://doi.org/10.1534/genetics.106.068361
Yang J, Worley E, Torres-Jerez I, Miller R, Wang M, Fu C, Wang ZY, Tang Y, Udvardi M (2015) PvNAC1 and PvNAC2 Are associated with leaf senescence and nitrogen use efficiency in switch grass. Bio Energ Res 8:868–880. https://doi.org/10.1007/s12155-014-9566-x
Yang M, Yunting F, Sun D, Shi Y (2016) Efficiency of two nitrification inhibitors (dicyandiamide and 3, 4-dimethypyrazole phosphate) on soil nitrogen transformations and plant productivity: a meta-analysis. Sci Rep 6:22075. https://doi.org/10.1038/srep22075
Yao SG, Sonoda Y, Tsutsui T (2008) Promoter analysis of OsAMT1; 2 and OsAMT1;3 implies their distinct roles in nitrogen utilization in rice. Breed Sci 58:201–207. https://doi.org/10.1270/jsbbs.58.201
Yao Y, Zeng K, Song Y (2020) Biological nitrification inhibitor for reducing N2O and NH3 emissions simultaneously under root zone fertilization in a Chinese rice field. Environ Pollut 264:114821. https://doi.org/10.1016/j.envpol.2020.114821
Ye YS, Liang XQ, Chen YX, Liu J, Gu JT, Guo R, Li L (2013) Alternate wetting and drying irrigation and controlled-release nitrogen fertilizer in late-season rice. Effects on dry matter accumulation, yield, water and nitrogen use. Field Crop Res 144:212–224. https://doi.org/10.1016/j.fcr.2012.12.003
Yost MA, Kitchen NR, Sudduth KA, Sadler EJ, Drummond ST, Volkmann MR (2017) Long-term impact of a precision agriculture system on grain crop production. Precis Agric 18:823–842. https://doi.org/10.1007/s11119-016-9490-5
Yu P, Li X, White PJ, Li C (2015) A large and deep root system underlies high nitrogen-use efficiency in maize production. PLoS ONE 10:e0126293. https://doi.org/10.1371/journal.pone.0126293
Yuan L, Loqué D, YeF FWB, vonWirén N (2007) Nitrogen dependent posttranscriptional regulation of the ammonium transporter AtAMT1. Plant Physiol 143:732–744. https://doi.org/10.1104/pp.106.093237
Zanin L, Zamboni A, Monte R, Tomasi N, Varanini Z, Cesco S, Pinton R (2015) Transcriptomic analysis highlights reciprocal interactions of urea and nitrate for nitrogen acquisition by maize roots. Plant Cell Physiol 56:532–548. https://doi.org/10.1093/pcp/pcu202
Zhang C, Kovacs JM (2012) The application of small unmanned aerial systems for precision agriculture: A review. Precis Agric 13:693–712. https://doi.org/10.1007/s11119-012-9274-5
Zhang J, Zhu T, Meng T, Zhang Y, Yang J, Yang W, Müller C, Cai Z (2013) Agricultural land use affects nitrate production and conservation in humid subtropical soils in China. Soil Biol Biochem 62:107–114. https://doi.org/10.1016/j.soilbio.2013.03.006
Zhang X, Davidson EA, Mauzerall DL, Searchinger TD, Dumas P, Shen Y (2015) Managing nitrogen for sustainable development. Nature 528:51. https://doi.org/10.1038/nature15743
Zhang Z, Xiong S, Wei Y, Meng X, Wang X, Ma X (2017) The role of glutamine synthetase isozymes in enhancing nitrogen use efficiency of N-efficient winter wheat. Sci Rep 7:1000. https://doi.org/10.1038/s41598-017-01071-1
Zhang C, Ju X, Powlson DS, Oenema O, Smith P (2019) Nitrogen surplus benchmarks for controlling N pollution in the main cropping systems of China. Environ Sci Technol 53(12):6678–6687. https://doi.org/10.1021/acs.est.8b06383
Zhang H, Forde BG (1998) An Arabidopsis MADS box gene that controls nutrient- induced changes in root architecture. Science 279:407–409. https://doi.org/10.1126/science.279.5349.407
Zhou Y, Cai H, Xiao J, Li X, Zhang Q, Lian X (2009) Over-expression of aspartate aminotransferase genes in rice resulted in altered nitrogen metabolism and increased amino acid content in seeds. Theor Appl Genet 118:1381–1390. https://doi.org/10.1007/s00122-009-0988-3
Zhou Y, Tao Y, Tang D, Wang J, Zhong J, Wang Y, Yuan Q, Yu X, Zhang Y, Wang Y, Liang G, Dong G (2017) Identification of QTL associated with nitrogen uptake and nitrogen use efficiency using high throughput genotyped CSSLS in rice (Oryza sativa L.). Frontiers in plant science 8:1166. https://doi.org/10.3389/fpls.2017.01166
Zhu C, Xiang J, Zhang Y, Zhang Y, Zhu D, Chen H (2019) Mechanized transplanting with side deep fertilization increases yield and nitrogen use efficiency of rice in Eastern China. Sci Rep 9:5653. https://doi.org/10.1038/s41598-019-42039-7
Zhu X, Fu W, Kong X, Chen C, Liu Z, Chen Z, Zhou J (2021) Nitrate accumulation in the soil profile is the main fate of surplus nitrogen after land-use change from cereal cultivation to apple orchards on the Loess Plateau. Agr Ecosyst Environ 319:107574. https://doi.org/10.1016/j.agee.2021.107574
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB). This work was supported by the National Natural Science Foundation of China (42150410380).
Author information
Authors and Affiliations
Contributions
Conceived and designed the review: ASE, ASE, and ESMD. Analyzed the data: ASE, ASE, AIA, ESMD, and MMR. Contributed reagents//analysis tools: ASE, ASE, AIAand ESMD. Wrote the paper: ASE, ASE, and ESMD. Revised the paper: AIA, ASE, and MMR. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Elrys, A.S., Elnahal, A.S., Abdo, A.I. et al. Traditional, Modern, and Molecular Strategies for Improving the Efficiency of Nitrogen Use in Crops for Sustainable Agriculture: a Fresh Look at an Old Issue. J Soil Sci Plant Nutr 22, 3130–3156 (2022). https://doi.org/10.1007/s42729-022-00873-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42729-022-00873-1