Abstract
Biochar and dung amendments have been extensively employed in soil remediation and fertilization of grasslands, which are the largest terrestrial sinks for methane. However, how these exogenous amendments regulate methane metabolisms at the molecular and community levels remains elusive. In this study, we investigated the functional genes and community assemblies of methanogens and methanotrophs using Geochip 5.0 and high-throughput sequencing to reveal the impacts of biochar and dung on soil methanogenesis and methane oxidation. The interactions between methane metabolic genes and other biogeochemical genes were also examined. According to Geochip microarrays, methanogenic gene mcrA decreased and increased with dung or biochar amendment, respectively; The methanotrophic gene pmoA showed a reverse but not significant tendency. Undominated processes contributed 65.51% to replace homogeneous selections as primary driving forces of methanogen assembly after dung amendment; the contribution of dispersal limitation increased to 46.13% in methanotroph assembly after biochar amendment. The diversity and association of co-occurrence networks for carbon–nitrogen cycling genes decreased after exogenous amendments. These results indicated that biochar and dung amendments prominently regulated the functional genes and community assembly involved in methane metabolisms. The co-existence patterns of methane metabolic genes and other related geochemical genes were also shaped by these amendments. This study provides the scientific reference for the development of grassland management in the context of global warming.
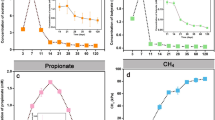
Graphical Abstract

Highlights
-
1.
Biochar amendment impeded methanogenesis and facilitated methane oxidation whereas dung amendment facilitated methanogenesis and impeded methane oxidation in grassland soils.
-
2.
Dung amendment generated undominated processes rather than homogeneous selection as the primary driving force in the community assembly of methanogen.
-
3.
Biochar amendment raised the contribution of dispersal limitation in the community assembly of methanotrophs.
-
4.
Diversity and association of the network among the functional genes involved in carbon and nitrogen cycling decreased after biochar and dung amendments.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Rising greenhouse gas (GHG) emissions caused by natural and anthropogenic activities are resulting in a warmer world (IPCC 2007; Ross et al. 2019). Although carbon dioxide (CO2) accounts for sizable proportion of GHG, methane (CH4) was regarded as another remarkable GHG with the potential to drive profound climate change because its molecular warming potential is 28 times greater than that of CO2 (IPCC 2007, 2021). As an important source and sink of methane, soil harbors methane generation and consumption processes to accomplish methane emissions (Conrad 2009; Zhao et al. 2021b). Methane generation is attributed to methanogenesis, which is mediated by methanogens. Substrates are consumed by methanogens via hydrogenotrophic, acetotrophic, and methylotrophic pathways using a nickel-containing methyl coenzyme M reductase (MCR), which is specific to methanogens (except methanotrophic archaea) (Conrad 2009, 2020). The functional gene mcrA that encodes MCR is commonly used in taxonomic studies of methanogens (Luton et al. 2002; Morris et al. 2002; Shima and Thauer 2005). Methane consumption is accomplished through the methanotroph-mediated oxidation processes. During methane oxidation, CH4 is oxidized to CO2 and H2O through the production of a range of intermediate metabolites that require methane monooxygenase (MMO) (Dedysh et al. 2002). Methane monooxygenases are divided into two structural forms: particulate methane monooxygenase (pMMO) and soluble methane monooxygenase (sMMO) (Dedysh et al. 2002; Liebner and Svenning 2013). Both pMMO and sMMO catalyze methane oxidation by breaking the C–H bond in methane, thereby oxidizing it to methanol and H2O. Most type I methanotrophs are detected by the pmoA gene, which encodes the β subunit of the pMMO protein. Meanwhile, the mmoX gene is selected as a biomarker for a few type II or type I methanotrophs since they utilize mmoX to encode sMMO rather than pMMO (Horz et al. 2001; Liebner and Svenning 2013).
Biochar and livestock dung amendments are applied in the grazed grassland as the critical management measures for soil remediation and fertilization. Recent studies have found that biochar and dung amendments significantly regulated soil methane emission. Biochar was the carbonaceous residue derived from the oxygen-limited pyrolysis of carbon (C)-rich biomass (Lehmann et al. 2011; Lehmann 2019; Zhao et al. 2021a). Biochar regulated both methanogenic and methanotrophic activities by controlling key soil properties (Jeffery et al. 2016; Zhao et al. 2021b). The elevation of soil porosity, aggregation, and water holding capacity due to biochar amendment shaped inappropriate microenvironments for methanogens while these responses of soil parameters promoted or inhibited methanotrophs in different cases (Atkinson et al. 2010; Jeffery et al. 2016; Rasa et al. 2018; Razzaghi et al. 2020). Despite these physical parameters, soil pH and N also increased by the liming effect and N mineralization was enhanced because of biochar amendment. Both methanogen and methanotroph growth depended on the threshold of soil pH and N (Cayuela et al. 2013; Gul and Whalen 2016; Karbin et al. 2016; Buss et al. 2018). The decrease in dissolved organic carbon after biochar amendment limited soil methanogenesis (Zimmerman et al. 2011; Nan et al. 2021). Methane fluxes increased significantly in the dung amended soils (Du et al. 2016; Cai et al. 2017; Cardoso et al. 2019). Fresh dung provided abundant methanogens to enhance soil methanogenic activity. The deposition of soluble C acted as an energy source for methanotrophs (Rastogi et al. 2008; Ho et al. 2013). The dung coverage and the biological crust on the mulch surface created anaerobic soil micro-environments, which were conducive to methanogens rather than methanotrophs (Ma et al. 2006; Singh et al. 2010; Wang et al. 2013). The high moisture of the fresh dung also hindered oxygen availability. Soil microbes decomposed the organic matter in dung to further depleted the oxygen required by methanotrophs (Malyan et al. 2016; Cai et al. 2017; Zhao et al. 2021b). The leaching of NH4+-N and dissolved nitrogen caused by the dung amendment supplied abundant N sources to promote the growthof methanogens and methanotrophs (Lin et al. 2009; Hartmann et al. 2013; Cai and Akiyama 2016; Additional file 1: 1.1). Conversely, methane oxidation was inhibited with sufficient ammonia because of the exclusive competition between amoA and pmoA genes (Bédard and Knowles 1989; Gulledge and Schimel 1998). Both biochar and dung amendments might regulate methane emission via interactions between methane metabolism and nitrogen (N) cycling because the processes involved in N cycling could either augment or moderate methanogenesis or methane oxidation (Additional file 1). However, despite previous studies reporting the response of the methane flux, specific mechanisms of the regulation on the soil methanogenic and methanotrophic activities by biochar and dung from the perspectives of functional gene and community assembly remain elusive. Additionally, the linkages and interactions between methane metabolic genes and other functional genes involved in geochemical cycling have not been well studied.
In this study, we aimed to investigate the functional gene composition and microbial community assembly of both methanogens and methanotrophs in the biochar and dung amended soils, and to identify the driving factors in these processes. We also discussed the co-occurrence and interactions between methane metabolic genes and other functional gene modules involved in C and N cycling to clarify the critical role of methane metabolic genes in geochemical cycling with biochar and dung amendments. Microcosm incubation, high-throughput sequencing, and Geochip 5.0 gene microarray were performed to explore these responses of functional genes and microbial community to the biochar and dung amendments. We hypothesized that biochar and dung amendments would reshape the functional gene structure and microbial community assembly related to methanogenesis and methane oxidation by altering the soil physicochemical characteristics and impairing the interactions between methane metabolic genes and other biogeochemical genes. This study provides scientific guidance for the implementation of sustainable soil managements to address the challenge of global warming.
2 Materials and methods
2.1 Study area and sample collection
This study employed the soil samples in grazed grassland for the subsequent laboratory incubation. Samples were collected from Research Station of Animal Ecology (44° 18′ N, 116° 45′ E, 1079 m a.s.l) located in the Maodeng Pasture, Inner Mongolia Autonomous Region, China. The sampling area belongs to the typical temperate steppe with a continental temperate semi-arid climate. The details of the sampling grassland are described in Additional file 1: 2.1. Soil cores were collected at a depth of 0–20 cm by using a 3 cm diameter soil auger from 1 m × 1 m sample boxes in selected sample plots, with three replicates.
2.2 Biochar preparation
The biochar in this study employed straws of wheat (Triticum aestivum) as the feedstock. After oven dried at 60 ℃ for 24 h, the straws were pyrolyzed in an oxygen-free stainless-steel furnace at 400 ℃ for 2 h to prepare the biochar. The collected biochar was sieved by 0.2 mm sieves for the subsequent experiments.
2.3 Microcosm incubation
5 g of air-dried soil was adjusted to the soil with field moisture of 20% and pre-cultured in 100 mL culture flasks at room temperature without light for 7 days. The flasks were then closed at atmospheric pressure. The experiments were carried out in three replicates for each treatment group: CK control, BC, FC, and BF under the aerobic condition. 0.5 g of biochar was added to BC; 0.5 g of dung was added to FC; 0.5 g of biochar and 0.5 g of dung were amended to BF together. The incubation temperature and time were 25 °C and 18 days, respectively. When the oxygen in the incubation flasks was depleted, soil samples were removed by destructive sampling. The physicochemical properties were determined following previous protocols according to Zhao et al. (2021c) (Additional file 1: 2.2).
2.4 DNA extraction, purification, and sequencing
Soil DNA was extracted from the collected samples using the MoBio PowerSoil isolation kits according to the manufacturer's instructions (MoBio Laboratories, Carlsbad, CA, USA). Subsequently, a quality and quantification assessment of the purified DNA was conducted by Nano-Drop ND-1000 Spectrophotometer (NanoDrop Technologies Inc., Wilmington, DE). The final DNA samples obtained from three biotopes were diluted and stored at − 80 °C for further analysis. The DNA samples were sequenced using primers such as 189F/682R, 338F/806R, and 524F10extF/Arch958R to obtain the amplicon sequencing of methanotrophs, bacteria, and archaea on the Illumina Hiseq 2500 platform (details of primer sequence are presented in Additional file 1: 2.3).
2.5 Geochip 5.0 gene microarray
The extracted DNA samples were also analyzed by Geochip 5.0 gene microarray (Yang et al. 2013; Li et al. 2021). The purified DNA was labeled with the fluorescent dye Cy5 using random primers. The labeled DNA was purified using the QIAquick PCR purification kit (Qiagen, Valencia, CA, USA) and subsequently dried in a SpeedVac (DNA Speedvac, Model DNA 100, Savant) at 45 °C for 45 min. The dried DNA was suspended in the hybridization buffer to perform hybridization reactions in the MAUI Hybridization Station (BioMicro Systems, Salt Lake City, UT, USA) at 42 °C for 12 h. The hybridized microarrays were scanned using a Scan Array Express microarray scanner (PerkinElmer, Boston, MA, USA) at 90% laser power and 75% photomultiplier gain. The images were then analyzed using ImaGene 6.0 (Biodiscovery, EI Segundo, CA, USA) for the assessment of the signal intensity of each spot. Results with signal-to-noise ratio (SNR = signal mean-background mean/background standard deviation) < 2 were censored with ImaGene markers. Only results that were detected in at least two of the four replicates were applied for the subsequent processes. Signal intensities of all genes were normalized by relative abundance and then subjected to relevant statistical analysis (Yang et al. 2013; Tang et al. 2019).
2.6 Statistical analysis
The statistical analysis was performed in R library using packages of “vegan”, “Hmisc”, “ggtern”, “picante”, “icamp”, “spaa” and “ggplot2” except for the otherwise annotation. All the data of functional genes were obtained according to Geochip 5.0 and the community assembly processes were analyzed based on the Amplicon sequencing.
To examine the environmental impacts on methane metabolic genes, the canonical correlation analysis (CCA) was performed using the “CCA” function in the “vegan” package and embellished by “ggplot2” package with the environmental factors of “TOC”, “nitrate”, “ammonia”, “pH”, and “moisture” for visualization (Dixon 2003; Wickham 2009).
The assembly processes of methanotrophs and methanogens were investigated by “picante” and “icamp” packages based on neutral and null models among samples from four treatments (Kembel et al. 2010; Ning et al. 2020). Mean nearest taxon distance (MNTD) was calculated based on the methanogenic and methanotrophic phylogenetic trees. NTI was calculated by observed MNTD based on the null model. βNTI could be obtained by calculation the MNTD values for 999 randomizations (Webb 2000). When βNTI was > 2 or < − 2, deterministic processes dominated the assembly process while the stochasticity dominated when βNTI was < 2 and > − 2 (Vellend 2010; Stegen et al. 2013; Zhou and Ning 2017). Bray–Curtis-based Raup–Crick index (RCBray) was employed to further identify the stochastic process such as homogenizing dispersal, dispersal limitation, and undominated processes according to the threshold |0.95| (Stegen et al. 2013; Zhou and Ning 2017). The niche characteristics such as niche breadth and niche overlaps were calculated based on Levin method using the “spaa” package to quantify habitat specialization and elucidate the contribution of assembly processes (Smith and Levins 1970).
The co-occurrence network was performed based on the Spearman correlation matrix calculated by “Hmisc” package (Harrell 2008). After screening the significant genes by false discovery rate (FDR) procedure, the matrix data were visualized on the Cytoscape platform (Benjamini et al. 2006; Bastian et al. 2009). Detailed procedures of co-occurrence network analysis are presented in Additional file 1: 2.4.
3 Results
3.1 The abundance of methane metabolic genes
To assess the amendment effects of biochar and livestock dung on the soil functional genes related to methane metabolisms, we employed intensities of genes such as mcrA, pmoA, and mmoX obtained from Geochip 5.0 to monitor the relative gene abundance. mcrA gene was regarded as the typical methanogenic gene providing MCR. The mcrA gene intensity decreased by 1.47% in the BC group with the biochar amendment while it increased in the FC or BF group by 0.38% and 0.82% with the dung amendment or the dual amendments of biochar and dung compared to the CK control (Fig. 1A). Although the gene intensity of pmoA, which was the most typical methanotrophic gene encoding the pMMO, did not vary significantly between the CK and BC groups, the increasing tendency was still observed after the biochar amendment. The intensity of this pmoA gene also showed decreasing tendency in the FC group compared to the CK group (Fig. 1A; Additional file 1: Table S1). Another crucial gene mmoX produced sMMO to support methane oxidation processes mediated by the other type of methanotrophs. The gene intensity of mmoX dramatically increased by 2.48%, 1.96%, and 2.78% in all the BC, FC, and BF groups compared to the CK control (Fig. 1A). These gene intensity variations indicated that biochar amendment reduced the gene abundance of mcrA and induced the gene abundance of pmoA and mmoX. On the contrary, the dung amendment induced the gene abundance of mcrA and mmoX, and only pmoA abundance was reduced.
A Response of gene intensities of methane metabolic genes to the biochar and dung amendments. Methane metabolic genes were methanogic gene mcrA and methanotrophic genes pmoA and mmoX. Canonical correlation analysis of the relationships between environmental variables and soil methanogenic (B) or methanotrophic (C) genes with biochar and dung amendments. Environmental variables such as soil total organic carbon (TOC), pH, moisture, nitrate and ammonia were presented
3.2 Environmental factors driving methane metabolism
Canonical Correspondence Analysis (CCA) demonstrated the correlation between environmental factors and the methane metabolic genes with the biochar and dung amendments (Fig. 1B and C). Soil total organic carbon (TOC) and nitrate were the most influential environmental attributes to the methanogenic genes. Particularly, the methanogenic genes in the BC group were strongly positively correlated to soil moisture and negatively correlated to soil pH and ammonia. Methanogenic genes in the FC group were negatively correlated to soil TOC (Fig. 1B). As far as methanotrophic genes were concerned, soil pH, ammonia, moisture, and TOC contributed similarly to the exogenous amendments. Similar to the methanogenic genes, methanotrophic genes in the BC group were still strongly negatively correlated to soil pH and ammonia, and those in the FC group were negatively correlated to soil TOC. A positive correlation between soil moisture and methane oxidation genes in the FC group could be observed (Fig. 1C).
3.3 The community assembly process of methanogens and methanotrophs
The assembly process of methanogen and methanotroph communities based on null models was elucidated after the amplicon sequencing (Fig. 2A). The β-nearest taxon index (βNTI) scores were calculated across all the samples from four treatments. The vast majority and overall proportion of methanogenic and methanotrophic βNTI fell in the range of − 2 to 2, implying the dominantly neutral processes in the methanogen and methanotroph assemblies. Only a small proportion of methanogenic βNTI was higher than + 2, indicating an inferior driving effect of heterogeneous selection in the methanogen assembly process (Fig. 2A). To further identify the neutral process, the βNTI scores were combined with Bray–Curtis-based Raup–Crick (RCBray). The largest fraction of RCBray fell in the range of − 0.95 to + 0.95 representing the undominated processes such as weak selection, weak dispersal, diversification, and drift rather than probabilistic dispersal dominated the methanogen and methanotroph assembly process. Very few RCBray scores lower than − 0.95 occurred in both methanogen and methanotroph communities. This indicated the inferior effect of dispersal limitation (Fig. 2B). The RCBray scores of methanotroph higher than + 0.95 represented the homogenizing dispersal. Thus, it is the stochastic process, especially undominated processes, to lead the assembly of methanogen and methanotroph community across all the treatments with biochar and dung amendments.
A βNTI of methanogen and methanotroph community assembly. B Bray–Curtis-based Raup–Crick index (RCBray) of methanogen and methanotroph community assembly. C Contribution of various assembly processes to the methanogen and methanotroph assembly. A, B were based on samples across all treatments with biochar and dung amendments; and C was based on samples from CK, BC, and FC groups , respectively
When we focused on the methanogen and methanotroph assembly processes of samples from each particular group, quite different results were presented (Fig. 2C). Homogeneous selection rather than stochastic process accounted for 99.32% in the assembly of methanogen communities from the CK group. Similarly, the homogeneous selection also accounted for 99.23% in methanogen assembly in the BC group. In terms of the FC group, homogeneous selection only accounted for 32.60% and undominated processes contributed 65.51%. In the CK group, the assembly of the methanotroph community was mainly driven by undominated processes with 65.51% percentages. Homogeneous selection, homogeneous dispersal, and dispersal limitation contributed to 24.23%, 7.82%, and 6.87%, respectively. In the BC group, dispersal limitation became the main driven process with a contribution of 46.13% percentages. Undominated processes and homogeneous selection accounted for 34.05% and 19.03%. When it comes to the methanotrophs in the FC group, the contribution of undominated processes was 40.14%, also lower than that of the CK group. Homogeneous selection, homogeneous dispersal, and dispersal limitation contributed to 29.62%, 16.62%, and 13.43%, respectively (Fig. 2C). These results highlighted that dung amendment generated undominated processes rather than homogeneous selection as the main driving force in the methanogen assembly and biochar amendment led to more contribution of dispersal limitation in the methanotroph assembly.
3.4 Niche breadth and niche overlap of methanogens and methanotrophs
To deeply clarify the ecological role of methanogens and methanotrophs, the niche breadth and niche overlap were investigated as the important parameters of niche characterization (Fig. 3). Niche breadth described the capacity of species to obtain resources with various niches (Sexton et al. 2017). The niche breadth value of methanogen was maintained at a quite low level and even lower than the entire archaea community (Fig. 3A). This narrow niche breadth indicated poor environment adaptivity and niche-discriminatory growth of methanogens. The niche breadth value of methanotrophs is higher than that of methanogens but also lower than that of the entire bacteria community (Fig. 3A). This result indicated that methanotrophs were more adaptive compared to methanogens but still pone to be shaped by niche selection compared to the other bacteria.
A Niche breadth of the methanotrophs, bacteria, methanogens and archaea with biochar and dung amendments. All the niche breadths were calculated based on samples across all treatments. B Niche overslaps between microbes involved in methane metabolism and nitrogen cycling. Microbes involved in methane metabolism were divided into methanogens and methanotrophs; Microbes involved in nitrogen cycling were related to DNRA, nitrogen fixation, ammonification, nitrification, and denitrification
Since several processes in N cycling were profoundly influential to methanogenesis and methane oxidation, the niche overlaps based on the Levin method between microorganisms mediating these processes and methanogens or methanotrophs were also calculated. Figure 3B depicts the prominently high niche overlaps between microbes mediating dissimilatory nitrogen reduction (DNRA) or ammonification and methane metabolic microbes. The niche overlap analysis employed functional genes involved in methanogenesis, methane oxidation, DNRA, ammonification, nitrogen fixation, nitrification, and denitrification to reveal the niche competition between microbes related to these processes of methane metabolism or nitrogen cycling. This niche overlaps implied the methane metabolic microbes competed for more resource attributes with microbes mediating DNRA or ammonification than diazotrophs, denitrifiers, and nitrifiers. All these microbes related to the N cycle overlapped more with methanotrophs than with methanogens across all the treatments. This result further proved only methanotrophs rather than methanogens could compete resources with microbes related to N cycles in various niches.
3.5 Amendment effect of biochar and dung on N cycling genes
Significant variation of functional genes involved in the N cycling among the CK and BC, FC, or BF groups could be observed (Fig. 4). nifH gene representing N fixation, ureC gene representing ammonification, amoA gene representing nitrification, nirB gene representing DNRA, narG, nirS/K, nosZ genes representing denitrification all increased more in the FC than in the BC group. The higher value of gene intensity shift in FC than in BC reflected the stronger effect of dung amendment than that of biochar amendment. Referring to all these genes, the sums of variation in the BC and FC were all lower than those in the BF (Fig. 4).
Response of the functional genes involved in nitrogen cycling to biochar and dung amendments. The values along with the genes represent the intensity variation of genes in groups after biochar, dung and biochar-dung amendments. The three percentage values indicate the ratio of the gene intensity increase in BC, FC, or BF group to the gene intensity in CK group
3.6 Co-occurrence network of functional genes involved in C and N cycling
There were substantial and strong positive correlations (r > 0.9) among gene modules involved in C and N cycling such as methane metabolism, N cycling, C fixation, and C degradation in the CK group. A few strongly negative correlations (r > 0.9) could be observed between the C fixation module and the other three modules (Fig. 5A). After the amendments of biochar and dung, plenty of weak positive correlations (r < 0.9) occurred among the four modules. The strong negative correlations (r > 0.9) among the modules dramatically decreased. Only the negative correlation between modules of C fixation and C degradation rarely occurred (Fig. 5B). The topo-properties of this co-occurrence network depicted the decreasing cohesion parameters such as clustering coefficient, network density, and network centralization in the BF group compared to the CK group. Simultaneously, the homogenous parameters such as network heterogeneity increased in the BF group (Table 1). These results indicated that biochar and dung amendments both weakened the association and diversity of the functional genes involved in C and N cycling.
Co-occurrence network of functional genes involved in carbon and nitrogen cycling without (A) or with (B) biochar and dung amendments. Nodes represent individual genes; edges represent significant spearman correlations (P < 0.01); strong positive correlations (ρ > 0.9), weak positive correlations (ρ < 0.9), and strong negative correlations (ρ > 0.9) were presented with colours of light blue, grey and blue, respectively; genes are divided into 4 modules representing methane metabolism, nitrogen cycling, carbon fixation and carbon degradation with different colours
4 Discussion
As important measures for grassland management, biochar and dung amendments showed enormous potential to shape the soil functional genes and microbial community. Because grasslands are the greatest terrestrial sink for the atmospheric methane, the microbiomes involved in the methane metabolism in grazed grassland soils with biochar and dung amendments have been thoroughly investigated (Gul et al. 2015; Palansooriya et al. 2019). We examined the soil functional gene structure and microbial community assembly using Geochip 5.0 and high throughput sequencing, respectively.
4.1 Biochar effects on methane metabolic genes
The abundance of the methanogenesis gene mcrA and methane oxidation gene pmoA decreased and increased, respectively, with biochar amendment (Fig. 1A). This might be mainly attributed to the alteration of soil porosity by biochar amendment. Biochar could improve soil aeration by providing the numerous internal particle pores and interparticle voids to the surrounding soils (Atkinson et al. 2010; Sohi et al. 2010; Hardie et al. 2013). The soil porosity might also be improved by the modification of the inherent pore size distribution because biochar created accommodation macropores in the surrounding soil in addition to directly contributing its own pores (Chia et al. 2012; Hardie et al. 2013; Guo et al. 2015). These accommodation macropores also decreased the soil bulk density (Devereux et al. 2013; Peake et al. 2014; Nan et al. 2021). Oxygen fluxes correspondingly increased in soils with high porosity and low density. Methanogenic activities were inhibited because this oxygen availability was harmful to Methanosarcina, which acted as the primary methanogenic taxa in this study (Kim et al. 2017; Nan et al. 2021). Instead, the dominant methanotrophs belong to USCγ with an extremely high preference for O2 and high affinity for CH4. Methanotrophic activities were enhanced because of the higher substrates supply such as CH4 and O2 in the biochar amended soil (Scheutz et al. 2009; Bohn et al. 2011; Cong et al. 2018).
pH might be another vital parameter for biochar regulation on the methanogenic and methanotrophic gene abundance. The pH of the biochar in this study reached 9.6. After the biochar amendment, the pH of the slightly alkaline soil in this study further increased due to the liming effect of biochar (Jeffery et al. 2016; Si et al. 2018). Methanogens were inhibited since the optimum pH range of Methanosarcina fell in 6.5–7.1 (Fig. 1B). Methanotrophic activities were also limited because the optimum pH of methanotrophs ranged from 5.0 to 6.5 (Jeffery et al. 2016; Nan et al. 2021). Incidentally, pH elevation mitigated the Al3+ availability to protect methanotrophs from the Al3+ toxicity (Tamai et al. 2007). Biochar amendment resulted in NH4+-N input by enhancing N mineralization. When soil ammonia was sufficient, either growth of methanogens or methanotrophs might be limited (Bédard and Knowles 1989; Gulledge and Schimel 1998; Zhao et al. 2021b). In addition, biochar amendment might diminish methanogen growth by reducing substrates for methanogens such as soil dissolved organic carbons (DOCs) (Zimmerman et al. 2011). Decreases in soil DOC and methane emissions from biochar amended soil have been observed in several previous studies (Han et al. 2016; Zheng et al. 2016). Altogether, the high porosity in the biochar amended soil resulted in lower methanogenic and higher methanotrophic gene abundance, and the altered pH and DOCs further diminished the methanogenic gene abundance.
4.2 Dung effects on methane metabolic genes
Soil porosity was also the primary factor controlled by the dung effect to regulate methanogenesis and methane oxidation (Fig. 1A). The dung amendment harbored biological crusts in the dung-soil patch to create an anoxic interface. The high moisture of the fresh dung also hindered the oxygen flux in the dung amended soil. The microbial degradation of the abundant organic matters from the dung further depleted oxygen. Thus, more anoxic compartments occurred after the dung amendment. These anoxic compartments created microenvironments that promote anaerobic methanogenesis and inhibit aerobic methanotrophic activities (Chadwick et al. 2000; Ma et al. 2006; Cai et al. 2017). Leaching of ammonia nitrogen (NH4+-N) and dissolved organic N (DON) simulated by dung allowed methanotrophs to obtain sufficient N (Cai and Akiyama 2016). The N-enriched environments could mitigate the limiting effect of C on microorganisms through higher litter inputs, thereby increasing the activity of methanogens (Banger et al. 2012; Siciliano et al. 2013). Under conditions of adequate N, the methanotrophic gene pmoA compete with the amoA gene, leading to further mitigation of methane oxidation (Bédard and Knowles 1989; Gulledge and Schimel 1998). Nitrite and nitrate oxidized from ammonia were also toxic to methanotrophs (Bédard and Knowles 1989; Gulledge and Schimel 1998; Bykova et al. 2006; Nazaries et al. 2013). More importantly, the methanogens in fresh dung from the intestine of livestock further enhanced the methanogenic activity (Zhao et al. 2021c). Notably, the abundance of another methanotrophic gene mmoX increased after the dung amendment. Methylosinus with mmoX were found in the incubated soils. These methanotrophs utilizing either pMMO or sMMO favored methane-enriched and oxygen-impoverished environment (Tentori and Richardson 2020). The anaerobic compartments in soils with dung amendment produced considerable endogenous CH4. The abundance of mmoX gene increased correspondingly with Methylosinus. Notably sMMO was inactive with the presence of copper ion and pMMO activity was facilitated in such environment. This was also consistent with the divergent abundance change of pmoA and mmoX genes after the dung amendment with copper input in this study. Unfortunately, how these reciprocally regulations of Cu on sMMO and pMMO formed is not confirmed (Semrau et al. 2010). Methanotrophs could synthesize a copper chelate called methanobactin to uptake copper (Semrau et al. 2020). How this copper uptaking interact in the methane oxidation was controversial until pMMO was proposed to employ a mono-copper as the only center to catalyze the methane oxidation, so the uptaken copper might provide extra binding sites for methane consumption (Ross et al. 2019). On the contrary, sMMO is a multicomponent soluble di-iron monooxygenase (Semrau et al. 2020). This might explain the divergent response of pmoA and mmoX genes to the copper input. Therefore, the disruption of aeration conditions, ascending N and Cu input, accounted for the increase and decrease in methanogenic and methanotrophic gene abundance, respectively.
4.3 Regulation of exogenous amendment on methanogenic and methanotrophic community assembly
The narrow niche breadth of methanogen made methanogens niche-discriminatory specialists rather than generalists in various environments (Fig. 3A) (Sexton et al. 2017; Xu et al. 2021). The poor environment adaptability led the methanogen community to hardly exist in various niches but be a strong competitor in a limited niche. The methanogen assembly process was therefore shaped by homogeneous selection without exogenous amendment (Fischbach and Segre 2016; Delgado-Baquerizo et al. 2017; Zhou and Ning 2017). Nonetheless, the undominated processes replaced homogeneous selection to primarily drive methanogen assembly with dung amendment (Fig. 2B). This change might be attributed to the exogenous methanogen input by fresh dung from the livestock intestine. Livestock intestine formed an extreme environment to harbor enormous methanogens which contributed a considerable proportion of the methanogen community in dung amended soils (Zhao et al. 2021c). When these methanogens were excreted outside the intestinal tracts with fresh dung, environmental conditions such as pH, moisture, temperature stress, and aeration drastically fluctuated. This environmental randomness poses difficulty in shaping methanogen populations by environment selection. Instead, undominated processes such as weak selection and dispersal dominated the assembly process.
Compared to the methanogens, the wider niche breadth of methanotrophs indicated that they were more environmentally adaptive and tended to be generalists in various biotopes (Fig. 3A). The higher niche overlaps between methanotrophs and microbes involved in N cycling also proved that methanotrophs could compete for resources with other species in much more extensive niches (Sexton et al. 2017). This environment tolerance resulted in stochastic processes rather than deterministic processes dominated the methanotroph assembly process (Fig. 2B). Stochastic processes increased the community redundancy of methanotrophs because other microbes overlapping methanotrophic niches could serve the same functions in the ecosystem (Fig. 3B) (De Vrieze et al. 2020). Although the passive dispersal of microorganisms was always random due to small sizes, microbial dispersal sometimes showed strong environmental patterns owing to dispersal limitation (Martiny et al. 2006; Hanson et al. 2012; Nemergut et al. 2013; Peay et al. 2016). When grassland soil was amended with biochar, the significant modification of the internal pores and interparticle void structure might be the limitation to eliminate the randomness of microbial dispersal (Zhou and Ning 2017; Ning et al. 2020). Thus, dispersal limitation became the primary driving force for the methanotroph assembly process in the biochar amended soils.
4.4 Interactions between methane metabolic genes and other functional genes involved in C and N cycling
Both biochar and dung amendments enhanced ecological processes such as N fixation, ammonification, nitrification, denitrification, and DNRA involved in N cycling (Fig. 4). Dung amendment showed stronger effects on these N cycling processes. Dung amendment increased N sequestration within the soil to participate in long-term N cycling. Dung deposition enhances soil organic C availability and microbial activity, thereby stimulating the mineralization of pre-existing N and microbial N fixation in the soil (Cai and Akiyama 2016; Cai et al. 2017). At the same time, invertebrates such as dung beetles, earthworms, termites, and maggots degraded the fresh dung to mineralize the organic N and residual organic N, which were also aminated by invertebrates (Mendez et al. 2013; Evans et al. 2019). Nitrification converted NH4+ to NO3− in soils and the ammoxidation gene amoA completes the first step of nitrification by oxidizing NH4+ to NH2OH. The ammoxidation microorganisms with amoA typically include ammonia-oxidizing bacteria (AOB) and ammonia-oxidizing archaea (AOA). Dung deposition induced soil NH4+, which stimulates ammoxidation (Hartmann et al. 2013; Cardenas et al. 2016). AOB in grassland soils were reported to increase with the dung amendment (Schauss et al. 2009; Yamamoto et al. 2012; Xue et al. 2016). AOA abundance also increased following dung amendment, utilizing additional labile C (Cai et al. 2017). The aforementioned N fixation, N mineralization, and nitrification prominently altered soil NH4+. Sufficient NH4+ inhibited methanotrophic activities through exclusive competition with CH4 (Bédard and Knowles 1989; Gulledge and Schimel 1998). Significant niche overlaps were found between methanotrophs and microbes involved in these processes (Fig. 3B). Sums of these gene variations in BC and FC were lower than those in BF. This implied synergy effects of biochar and dung on N cycling (Fig. 4).
After biochar and dung amendments, the occurrence of numerous weak positive correlations in the co-occurrence network of C and N cycling genes suggested less common preference for environmental conditions or niche-overlapping among species with these functional genes. The disappearance of strong negative correlations indicated competition or niche-partitioning among these species involved in C and N cycling (Fig. 5) (Faust and Raes 2012; He et al. 2021). These results were consistent with the decreasing association parameters in the topological parameters of the co-occurrence network. The increasing network heterogeneity depicted the decreasing diversity among these functional genes (Table 1). The decreasing complexity and connectivity of the network implied that the species with these genes were less interconnected after applying biochar and dung amendments and the efficiency of resource and information transfer was lessened (Morrien et al. 2017; He et al. 2021). Hence, biochar and dung amendments might result in lower stability of the microbial community that participated in C and N cycling (Mougi and Kondoh 2012).
5 Conclusion
Dung amendment promoted methanogenic genes and inhibited methanotrophic genes while biochar amendment inhibited methanogenic genes and promoted methanotrophic genes via controlling soil porosity, pH and ammonia. Dung amendment led undominated processes rather than homogeneous selection to dominate the methanogen assembly and biochar amendment raised the contribution of dispersal limitation in methanotroph assembly. The two exogenous amendments weakened the interconnection between methane metabolic genes and other functional genes involved in C and N cycling. This study provided an integrated understanding of the microbial coexistence patterns and community assembly related to methane emission with typical management in grasslands. Deeper investigation on the interactions among the functional modules involved in biogeochemical cycling would better optimize soil utilization in the context of global climate change.
Data availability statement
The data that support the findings of this study are available in Sequence Read Archive (SRA) database on National Center for Biotechnology Information (NCBI) platform with a bioproject Accession number PRJNA783127 (https://dataview.ncbi.nlm.nih.gov/object/PRJNA783127).
References
Atkinson CJ, Fitzgerald JD, Hipps NA (2010) Potential mechanisms for achieving agricultural benefits from biochar application to temperate soils: a review. Plant Soil 337(1–2):1–18. https://doi.org/10.1007/s11104-010-0464-5
Banger K, Tian H, Lu C (2012) Do nitrogen fertilizers stimulate or inhibit methane emissions from rice fields? Glob Chang Biol 18(10):3259–3267. https://doi.org/10.1111/j.1365-2486.2012.02762.x
Bastian M, Heymann S, Jacomy M (2009) Gephi: an open source software for exploring and manipulating networks. In: International AAAI conference on weblogs & social media; third international AAAI conference on weblogs & social media
Bédard C, Knowles R (1989) Physiology, biochemistry, and specific inhibitors of Ch4, Nh4+, and co oxidation by methanotrophs and nitrifiers. Microbiol Rev 53(1):68–84
Benjamini Y, Krieger AM, Yekutieli D (2006) Adaptive linear step-up procedures that control the false discovery rate. Biometrika 93(3):491–507. https://doi.org/10.1093/biomet/93.3.491
Bohn S, Brunke P, Gebert J, Jager J (2011) Improving the aeration of critical fine-grained landfill top cover material by vegetation to increase the microbial methane oxidation efficiency. Waste Manag 31(5):854–863. https://doi.org/10.1016/j.wasman.2010.11.009
Buss W, Shepherd JG, Heal KV, Mašek O (2018) Spatial and temporal microscale Ph change at the soil-biochar interface. Geoderma 331:50–52. https://doi.org/10.1016/j.geoderma.2018.06.016
Bykova S, Boeckx P, Kravchenko I, Galchenko V, Van Cleemput O (2006) Response of Ch4 oxidation and methanotrophic diversity to NH4+ and CH4 mixing ratios. Biol Fertil Soil 43(3):341–348. https://doi.org/10.1007/s00374-006-0114-5
Cai Y, Akiyama H (2016) Nitrogen loss factors of nitrogen trace gas emissions and leaching from excreta patches in grassland ecosystems: a summary of available data. Sci Total Environ 572:185–195. https://doi.org/10.1016/j.scitotenv.2016.07.222
Cai YJ, Chang SX, Cheng Y (2017) Greenhouse gas emissions from excreta patches of grazing animals and their mitigation strategies. Earth Sci Rev 171:44–57. https://doi.org/10.1016/j.earscirev.2017.05.013
Cardenas LM, Misselbrook TM, Hodgson C, Donovan N, Gilhespy S, Smith KA, Dhanoa MS, Chadwick D (2016) Effect of the application of cattle urine with or without the nitrification inhibitor Dcd, and dung on greenhouse gas emissions from a Uk Grassland Soil. Agric Ecosyst Environ 235:229–241. https://doi.org/10.1016/j.agee.2016.10.025
Cardoso AD, Oliveira SC, Janusckiewicz ER, Brito LF, Morgado ED, Reis RA, Ruggieri AC (2019) Seasonal effects on ammonia, nitrous oxide, and methane emissions for beef cattle excreta and urea fertilizer applied to a tropical pasture. Soil till Res 194:104341. https://doi.org/10.1016/j.still.2019.104341
Cayuela ML, Sanchez-Monedero MA, Roig A, Hanley K, Enders A, Lehmann J (2013) Biochar and denitrification in soils: when, how much and why does biochar reduce N2O emissions? Sci Rep 3:1732. https://doi.org/10.1038/srep01732
Chadwick DR, Pain BF, Brookman SKE (2000) Nitrous oxide and methane emissions following application of animal manures to Grassland. J Environ Qual 29:277–287
Chia CH, Gong B, Joseph SD, Marjo CE, Munroe P, Rich AM (2012) Imaging of mineral-enriched biochar by Ftir, Raman and Sem–Edx. Vibrat Spectro 62:248–257. https://doi.org/10.1016/j.vibspec.2012.06.006
Cong W, Meng J, Ying SC (2018) Impact of soil properties on the soil methane flux response to biochar addition: a meta-analysis. Environ Sci Process Impacts 20(9):1202–1209. https://doi.org/10.1039/c8em00278a
Conrad R (2009) The global methane cycle: recent advances in understanding the microbial processes involved. Environ Microbiol Rep 1(5):285–292. https://doi.org/10.1111/j.1758-2229.2009.00038.x
Conrad R (2020) Importance of hydrogenotrophic, aceticlastic and methylotrophic methanogenesis for methane production in terrestrial, aquatic and other anoxic environments: a mini review. Pedosphere 30(1):25–39. https://doi.org/10.1016/s1002-0160(18)60052-9
De Vrieze J, De Mulder T, Matassa S, Zhou J, Angenent LT, Boon N, Verstraete W (2020) Stochasticity in microbiology: managing unpredictability to reach the sustainable development goals. Microb Biotechnol 13(4):829–843. https://doi.org/10.1111/1751-7915.13575
Dedysh SN, Khmelenina VN, Suzina NE, Trotsenko YA, Semrau JD, Liesack W, Tiedje JM (2002) Methylocapsa acidiphila gen nov., sp. nov., a novel methane-oxidizing and dinitrogen-fixing acidophilic bacterium from sphagnum bog. Int J Syst Evol Microbiol 52(1):251–261. https://doi.org/10.1099/00207713-52-1-251
Delgado-Baquerizo M, Reich PB, Khachane AN, Campbell CD, Thomas N, Freitag TE, Abu Al-Soud W, Sorensen S, Bardgett RD, Singh BK (2017) It is elemental: soil nutrient stoichiometry drives bacterial diversity. Environ Microbiol 19(3):1176–1188. https://doi.org/10.1111/1462-2920.13642
Devereux RC, Sturrock CJ, Mooney SJ (2013) The effects of biochar on soil physical properties and winter wheat growth. Earth Environ Sci Trans 103(1):13–18. https://doi.org/10.1017/s1755691012000011
Dixon P (2003) Vegan, a package of R functions for community ecology. J Veg Sci 14(6):927–930. https://doi.org/10.1658/1100-9233(2003)014[0927:Vaporf]2.0.Co;2
Du ZY, Wang XD, Liu XP, Cai YJ (2016) Effects of rock fragments on yak dung greenhouse gas emissions on the Qinghai-Tibetan Plateau. J Mount Sci 13(11):2006–2014. https://doi.org/10.1007/s11629-015-3798-x
Evans KS, Mamo M, Wingeyer A, Schacht WH, Eskridge KM, Bradshaw J, Ginting D (2019) Dung beetles increase greenhouse gas fluxes from dung pats in a North Temperate Grassland. J Environ Qual 48(3):537–548. https://doi.org/10.2134/jeq2018.03.0111
Faust K, Raes J (2012) Microbial interactions: from networks to models. Nat Rev Microbiol 10(8):538–550. https://doi.org/10.1038/nrmicro2832
Fischbach MA, Segre JA (2016) Signaling in host-associated microbial communities. Cell 164(6):1288–1300. https://doi.org/10.1016/j.cell.2016.02.037
Gul S, Whalen JK (2016) Biochemical cycling of nitrogen and phosphorus in biochar-amended soils. Soil Biol Biochem 103:1–15. https://doi.org/10.1016/j.soilbio.2016.08.001
Gul S, Whalen JK, Thomas BW, Sachdeva V, Deng H (2015) Physico-chemical properties and microbial responses in biochar-amended soils: mechanisms and future directions. Agric Ecosyst Environ 206:46–59. https://doi.org/10.1016/j.agee.2015.03.015
Gulledge J, Schimel JP (1998) Low-concentration kinetics of atmospheric CH4 oxidation in soil and mechanism of NH4+ inhibition. Appl Environ Microbiol 64:4291–4298
Guo G, Kong W, Liu J, Zhao J, Du H, Zhang X, Xia P (2015) Diversity and distribution of autotrophic microbial community along environmental gradients in grassland soils on the Tibetan Plateau. Appl Microbiol Biotechnol 99(20):8765–8776. https://doi.org/10.1007/s00253-015-6723-x
Han X, Sun X, Wang C, Wu M, Dong D, Zhong T, Thies JE, Wu W (2016) Mitigating methane emission from paddy soil with rice-straw biochar amendment under projected climate change. Sci Rep 6:24731. https://doi.org/10.1038/srep24731
Hanson CA, Fuhrman JA, Horner-Devine MC, Martiny JB (2012) Beyond biogeographic patterns: processes shaping the microbial landscape. Nat Rev Microbiol 10(7):497–506. https://doi.org/10.1038/nrmicro2795
Hardie M, Clothier B, Bound S, Oliver G, Close D (2013) Does biochar influence soil physical properties and soil water availability? Plant Soil 376(1–2):347–361. https://doi.org/10.1007/s11104-013-1980-x
Harrell FE (2008) Hmisc: Harrell miscellaneous 3:4-4
Hartmann AA, Barnard RL, Marhan S, Niklaus PA (2013) Effects of drought and N-fertilization on N cycling in two grassland soils. Oecologia 171(3):705–717. https://doi.org/10.1007/s00442-012-2578-3
He Q, Wang S, Hou W, Feng K, Li F, Hai W, Zhang Y, Sun Y, Deng Y (2021) Temperature and microbial interactions drive the deterministic assembly processes in sediments of hot springs. Sci Total Environ 772:145465. https://doi.org/10.1016/j.scitotenv.2021.145465
Ho A, Kerckhof FM, Luke C, Reim A, Krause S, Boon N, Bodelier PL (2013) Conceptualizing functional traits and ecological characteristics of methane-oxidizing bacteria as life strategies. Environ Microbiol Rep 5(3):335–345. https://doi.org/10.1111/j.1758-2229.2012.00370.x
Horz HP, Yimga MT, Liesack W (2001) Detection of methanotroph diversity on roots of submerged rice plants by molecular retrieval of Pmoa, Mmox, Mxaf, and 16s Rrna and Ribosomal DNA, including Pmoa-based terminal restriction fragment length polymorphism profiling. Appl Environ Microbiol 67(9):4177–4185. https://doi.org/10.1128/aem.67.9.4177-4185.2001
IPCC (2007) Contribution of working group I to the fourth assessment report of the Intergovernmental Panel on Climate Change. Climate Change 2007: The Physical Science Basis
IPCC (2021) Contribution of working group I to the sixth assessment report of the Intergovernmental Panel on Climate Change. In Climate Change 2021: The Physical Science Basis
Jeffery S, Verheijen FGA, Kammann C, Abalos D (2016) Biochar effects on methane emissions from soils: a meta-analysis. Soil Biol Biochem 101:251–258. https://doi.org/10.1016/j.soilbio.2016.07.021
Karbin S, Hagedorn F, Hiltbrunner D, Zimmermann S, Niklaus PA (2016) Spatial micro-distribution of methanotrophic activity along a 120-year afforestation chronosequence. Plant Soil 415(1–2):13–23. https://doi.org/10.1007/s11104-016-3141-5
Kembel SW, Cowan PD, Helmus MR, Cornwell WK, Morlon H, Ackerly DD, Blomberg SP, Webb CO (2010) Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26(11):1463–1464. https://doi.org/10.1093/bioinformatics/btq166
Kim J, Yoo G, Kim D, Ding W, Kang H (2017) Combined application of biochar and slow-release fertilizer reduces methane emission but enhances rice yield by different mechanisms. Appl Soil Ecol 117–118:57–62. https://doi.org/10.1016/j.apsoil.2017.05.006
Lehmann J (2019) Science-to-action through global and regional biochar networks. Biochar 1(4):337–337. https://doi.org/10.1007/s42773-019-00029-y
Lehmann J, Rillig MC, Thies J, Masiello CA, Hockaday WC, Crowley D (2011) Biochar effects on soil biota—a review. Soil Biol Biochem 43(9):1812–1836. https://doi.org/10.1016/j.soilbio.2011.04.022
Li D, Ni H, Jiao S, Lu Y, Zhou J, Sun B, Liang Y (2021) Coexistence patterns of soil methanogens are closely tied to methane generation and community assembly in rice paddies. Microbiome 9(1):20. https://doi.org/10.1186/s40168-020-00978-8
Liebner S, Svenning MM (2013) Environmental transcription of Mmox by methane-oxidizing proteobacteria in a subarctic Palsa Peatland. Appl Environ Microbiol 79(2):701–706. https://doi.org/10.1128/AEM.02292-12
Lin XW, Wang SP, Ma XZ, Xu GP, Luo CY, Li YN, Jiang GM, Xie ZB (2009) Fluxes of Co2, Ch4, and N2o in an alpine meadow affected by yak excreta on the Qinghai-Tibetan Plateau during summer grazing periods. Soil Biol Biochem 41(4):718–725. https://doi.org/10.1016/j.soilbio.2009.01.007
Luton PE, Wayne JM, Sharp RJ, Riley PW (2002) The Mcra Gene as an alternative to 16s Rrna in the phylogenetic analysis of methanogen populations in landfill. Microbiology (reading) 148(Pt 11):3521–3530. https://doi.org/10.1099/00221287-148-11-3521
Ma XZ, Wang SP, Wang YF, Jiang GM, Nyren P (2006) Short-term effects of sheep excrement on carbon dioxide, nitrous oxide and methane fluxes in typical grassland of Inner Mongolia. N Z J Agric Res 49(3):285–297. https://doi.org/10.1080/00288233.2006.9513719
Malyan SK, Bhatia A, Kumar A, Gupta DK, Singh R, Kumar SS, Tomer R, Kumar O, Jain N (2016) Methane production, oxidation and mitigation: a mechanistic understanding and comprehensive evaluation of influencing factors. Sci Total Environ 572:874–896. https://doi.org/10.1016/j.scitotenv.2016.07.182
Martiny JB, Bohannan BJ, Brown JH, Colwell RK, Fuhrman JA, Green JL, Horner-Devine MC, Kane M, Krumins JA, Kuske CR, Morin PJ, Naeem S, Ovreas L, Reysenbach AL, Smith VH, Staley JT (2006) Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4(2):102–112. https://doi.org/10.1038/nrmicro1341
Mendez A, Tarquis AM, Saa-Requejo A, Guerrero F, Gasco G (2013) Influence of pyrolysis temperature on composted sewage sludge biochar priming effect in a loamy soil. Chemosphere 93(4):668–676. https://doi.org/10.1016/j.chemosphere.2013.06.004
Morrien E, Hannula SE, Snoek LB, Helmsing NR, Zweers H, de Hollander M, Soto RL, Bouffaud ML, Buee M, Dimmers W, Duyts H, Geisen S, Girlanda M, Griffiths RI, Jorgensen HB, Jensen J, Plassart P, Redecker D, Schmelz RM, Schmidt O, Thomson BC, Tisserant E, Uroz S, Winding A, Bailey MJ, Bonkowski M, Faber JH, Martin F, Lemanceau P, de Boer W, van Veen JA, van der Putten WH (2017) Soil networks become more connected and take up more carbon as nature restoration progresses. Nat Commun 8:14349. https://doi.org/10.1038/ncomms14349
Morris SA, Radajewski S, Willison TW, Murrell JC (2002) Identification of the functionally active methanotroph population in a peat soil microcosm by stable-isotope probing. Appl Environ Microbiol 68(3):1446–1453. https://doi.org/10.1128/aem.68.3.1446-1453.2002
Mougi A, Kondoh M (2012) Diversity of interaction types and ecological community stability. Science 337(6092):349–351. https://doi.org/10.1126/science.1220529
Nan Q, Xin L, Qin Y, Waqas M, Wu W (2021) Exploring long-term effects of biochar on mitigating methane emissions from paddy soil: a review. Biochar 3(2):125–134. https://doi.org/10.1007/s42773-021-00096-0
Nazaries L, Murrell JC, Millard P, Baggs L, Singh BK (2013) Methane, microbes and models: fundamental understanding of the soil methane cycle for future predictions. Environ Microbiol 15(9):2395–2417. https://doi.org/10.1111/1462-2920.12149
Nemergut DR, Schmidt SK, Fukami T, O’Neill SP, Bilinski TM, Stanish LF, Knelman JE, Darcy JL, Lynch RC, Wickey P, Ferrenberg S (2013) Patterns and processes of microbial community assembly. Microbiol Mol Biol Rev 77(3):342–356. https://doi.org/10.1128/MMBR.00051-12
Ning D, Yuan M, Wu L, Zhang Y, Guo X, Zhou X, Yang Y, Arkin AP, Firestone MK, Zhou J (2020) A quantitative framework reveals ecological drivers of grassland microbial community assembly in response to warming. Nat Commun 11(1):4717. https://doi.org/10.1038/s41467-020-18560-z
Palansooriya KN, Wong JTF, Hashimoto Y, Huang L, Rinklebe J, Chang SX, Bolan N, Wang H, Ok YS (2019) Response of microbial communities to biochar-amended soils: a critical review. Biochar 1(1):3–22. https://doi.org/10.1007/s42773-019-00009-2
Peake LR, Reid BJ, Tang X (2014) Quantifying the influence of biochar on the physical and hydrological properties of dissimilar soils. Geoderma 235–236:182–190. https://doi.org/10.1016/j.geoderma.2014.07.002
Peay KG, Kennedy PG, Talbot JM (2016) Dimensions of biodiversity in the earth mycobiome. Nat Rev Microbiol 14(7):434–447. https://doi.org/10.1038/nrmicro.2016.59
Rasa K, Heikkinen J, Hannula M, Arstila K, Kulju S, Hyväluoma J (2018) How and why does willow biochar increase a clay soil water retention capacity? Biomass Bioenergy 119:346–353. https://doi.org/10.1016/j.biombioe.2018.10.004
Rastogi G, Ranade DR, Yeole TY, Gupta AK, Patole MS, Shouche YS (2008) Molecular analyses of methanogen diversity associated with cattle dung. World J Microbiol Biotechnol 24(12):2973–2979. https://doi.org/10.1007/s11274-008-9840-1
Razzaghi F, Obour PB, Arthur E (2020) Does biochar improve soil water retention? A systematic review and meta-analysis. Geoderma 361:114055. https://doi.org/10.1016/j.geoderma.2019.114055
Ross MO, MacMillan F, Wang J, Nisthal A, Lawton TJ, Olafson BD, Mayo SL, Rosenzweig AC, Hoffman BM (2019) Particulate methane monooxygenase contains only mononuclear copper centers. Science 364(6440):566–570. https://doi.org/10.1126/science.aav2572
Schauss K, Focks A, Leininger S, Kotzerke A, Heuer H, Thiele-Bruhn S, Sharma S, Wilke BM, Matthies M, Smalla K, Munch JC, Amelung W, Kaupenjohann M, Schloter M, Schleper C (2009) Dynamics and functional relevance of ammonia-oxidizing archaea in two agricultural soils. Environ Microbiol 11(2):446–456. https://doi.org/10.1111/j.1462-2920.2008.01783.x
Scheutz C, Kjeldsen P, Bogner JE, De Visscher A, Gebert J, Hilger HA, Huber-Humer M, Spokas K (2009) Microbial methane oxidation processes and technologies for mitigation of landfill gas emissions. Waste Manag Res 27(5):409–455. https://doi.org/10.1177/0734242X09339325
Semrau JD, DiSpirito AA, Yoon S (2010) Methanotrophs and copper. FEMS Microbiol Rev 34:496–531. https://doi.org/10.1111/j.1574-6976.2010.00212.x
Semrau JD, DiSpirito AA, Obulisamy PK, Kang-Yun CS (2020) Methanobactin from methanotrophs: genetics, structure, function and potential applications. FEMS Microbiol Lett 367:fnaa045. https://doi.org/10.1093/femsle/fnaa045
Sexton JP, Montiel J, Shay JE, Stephens MR, Slatyer RA (2017) Evolution of ecological niche breadth. Annu Rev Ecol Evol Syst 48(1):183–206. https://doi.org/10.1146/annurev-ecolsys-110316-023003
Shima S, Thauer RK (2005) Methyl-coenzyme M reductase and the anaerobic oxidation of methane in methanotrophic archaea. Curr Opin Microbiol 8(6):643–648. https://doi.org/10.1016/j.mib.2005.10.002
Si L, Xie Y, Ma Q, Wu L (2018) The short-term effects of rice straw biochar, nitrogen and phosphorus fertilizer on rice yield and soil properties in a cold waterlogged paddy field. Sustainability 10(2):537. https://doi.org/10.3390/su10020537
Siciliano A, Ruggiero C, De Rosa S (2013) A new integrated treatment for the reduction of organic and nitrogen loads in methanogenic landfill leachates. Proc Saf Environ Prot 91(4):311–320. https://doi.org/10.1016/j.psep.2012.06.008
Singh JS, Pandey VC, Singh DP, Singh RP (2010) Influence of pyrite and farmyard manure on population dynamics of soil methanotroph and rice yield in saline rain-fed paddy field. Agric Ecosyst Environ 139(1–2):74–79. https://doi.org/10.1016/j.agee.2010.07.003
Smith JM, Levins R (1970) Evolution in changing environments. Popul Stud 24(1):127
Sohi SP, Krull E, Lopez-Capel E, Bol R (2010) A review of biochar and its use and function in soil. Adv Agron 105:47–82
Stegen JC, Lin X, Fredrickson JK, Chen X, Kennedy DW, Murray CJ, Rockhold ML, Konopka A (2013) Quantifying community assembly processes and identifying features that impose them. ISME J 7(11):2069–2079. https://doi.org/10.1038/ismej.2013.93
Tamai N, Takenaka C, Ishizuka S (2007) Water-soluble Al inhibits methane oxidation at atmospheric concentration levels in Japanese Forest Soil. Soil Biol Biochem 39(7):1730–1736. https://doi.org/10.1016/j.soilbio.2007.01.029
Tang L, Zhong L, Xue K, Wang S, Xu Z, Lin Q, Luo C, Rui Y, Li X, Li M, Liu W-t, Yang Y, Zhou J, Wang Y (2019) Warming counteracts grazing effects on the functional structure of the soil microbial community in a Tibetan Grassland. Soil Biol Biochem 134:113–121. https://doi.org/10.1016/j.soilbio.2019.02.018
Tentori EF, Richardson RE (2020) Methane monooxygenase gene transcripts as quantitative biomarkers of methanotrophic activity in Methylosinus trichosporium Ob3b. Appl Environ Microbiol 86(23):e01048-20. https://doi.org/10.1128/AEM.01048-20
Vellend M (2010) Conceptual synthesis in community ecology. Q Rev Biol 85(2):183–206. https://doi.org/10.1086/652373
Wang XY, Huang D, Zhang YJ, Chen WQ, Wang CJ, Yang XM, Luo W (2013) Dynamic changes of Ch4 and Co2 emission from grazing sheep urine and dung patches in typical steppe. Atmos Environ 79:576–581. https://doi.org/10.1016/j.atmosenv.2013.07.003
Webb CO (2000) Exploring the phylogenetic structure of ecological communities: an example for rain forest trees. Am Nat 156:145–155
Wickham H (2009) Ggplot2: elegant graphics for data analysis. Springer Science & Business Media, New York
Xu QC, Luo GW, Guo JJ, Xiao Y, Zhang FG, Guo SW, Ling N, Shen QR (2021) Microbial generalist or specialist: intraspecific variation and dormancy potential matter. Mol Ecol 31:1–13. https://doi.org/10.1111/mec.16217
Xue C, Zhang X, Zhu C, Zhao J, Zhu P, Peng C, Ling N, Shen Q (2016) Quantitative and compositional responses of ammonia-oxidizing archaea and bacteria to long-term field fertilization. Sci Rep 6:28981. https://doi.org/10.1038/srep28981
Yamamoto N, Oishi R, Suyama Y, Tada C, Nakai Y (2012) Ammonia-oxidizing bacteria rather than ammonia-oxidizing archaea were widely distributed in animal manure composts from field-scale facilities. Microbes Environ 27(4):519–524. https://doi.org/10.1264/jsme2.me12053
Yang Y, Wu L, Lin Q, Yuan M, Xu D, Yu H, Hu Y, Duan J, Li X, He Z, Xue K, van Nostrand J, Wang S, Zhou J (2013) Responses of the functional structure of soil microbial community to livestock grazing in the Tibetan Alpine Grassland. Glob Chang Biol 19(2):637–648. https://doi.org/10.1111/gcb.12065
Zhao Q, Wang Y, Ayele G, Xu Z, Yu Z (2021a) Only mass migration of fungi runs through the biotopes of soil, phyllosphere, and feces. J Soil Sediment 21(2):1151–1164. https://doi.org/10.1007/s11368-020-02873-z
Zhao Q, Wang Y, Xu Z, Yu Z (2021b) How does biochar amendment affect soil methane oxidation? A Review. J Soil Sediment 1:1. https://doi.org/10.1007/s11368-021-02889-z
Zhao Q, Xu Z, Yu Z (2021c) Straw-derived biochar as the potential adsorbent for U(Vi) and Th(Iv) removal in aqueous solutions. Biomass Convers Biorefin. https://doi.org/10.1007/s13399-021-01810-5
Zheng J, Chen J, Pan G, Liu X, Zhang X, Li L, Bian R, Cheng K, Jinwei Z (2016) Biochar decreased microbial metabolic quotient and shifted community composition four years after a single incorporation in a slightly acid rice paddy from Southwest China. Sci Total Environ 571:206–217. https://doi.org/10.1016/j.scitotenv.2016.07.135
Zhou JZ, Ning DL (2017) Stochastic community assembly: does it matter in microbial ecology? Microbiol Mol Biol Rev 81:e00002-17
Zimmerman AR, Gao B, Ahn M-Y (2011) Positive and negative carbon mineralization priming effects among a variety of biochar-amended soils. Soil Biol Biochem 43(6):1169–1179. https://doi.org/10.1016/j.soilbio.2011.02.005
Acknowledgements
The authors thank Mr. Jianjun Chen and his team members for their help with the sample collection.
Funding
This work was supported by the National Key Research and Development Program of China (No. 2018YFD0800403).
Author information
Authors and Affiliations
Contributions
QZ: Conceptualization, methodology, formal analysis, investigation, visualization, writing—original draft. YW and JY: writing—review and editing. ZX and ZY: supervision and writing—review and editing. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Supplementary Information
Additional file 1.
Supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, Q., Wang, Y., Xu, Z. et al. Unravelling how biochar and dung amendments determine the functional structure and community assembly related to methane metabolisms in grassland soils. Biochar 4, 49 (2022). https://doi.org/10.1007/s42773-022-00167-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42773-022-00167-w