Abstract
Changes resulting from different tillage practices can affect the structure of microbial communities, thereby altering soil ecosystems and their functioning. The aim of this study was to explore and compare the physical, chemical properties and bacterial community composition of soils from different land use types (forest, grassland, vineyard, and arable field) in a small catchment. 16S rRNA gene-based amplicon sequencing was used to reveal the taxonomic diversity of summer and autumn soil samples taken from two different slope positions. The greater the anthropogenic impact was on the type of land use, the greater the change was in soil physical and chemical parameters. All sample types were dominated by the phyla Pseudomonadota, Acidobacteriota, Actinobacteriota, Bacteroidota and Verrucomicrobiota. Differences in the relative abundance of various bacterial taxa reflected the different land use types, the seasonality, and the topography. These diversity changes were consistent with the differences in soil properties.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Healthy soils are necessary to sustain plant growth and food production, store carbon, and thus help mitigate some of the negative effects of climate change. Changes in land-use or human interventions due to different management practices such as tillage or the addition of fertilizers can significantly affect the structure and biodiversity of the soil microbiota and the biogeochemical processes involving microorganisms.
In general, agricultural lands have different levels of human influences on soil ecosystems, with forests being the least and arable field the most anthropogenically influenced land-use types. Anthropogenic effects such as soil management practices (e.g. tillage, fertilizer additions, or lack thereof) can disrupt soil ecosystem services and soil physical, chemical, and biological properties. The type and frequency of soil management practices can also influence soil parameters. Tillage and fertilizer addition to soils can affect the soil surface chemistry, such as an increase in phosphorus accumulation (Obour et al. 2017) or soil organic matter content (Hussain et al. 1999; Obour et al. 2017) in no-tillage soils compared to conventional tillage. Soil carbon and nitrogen content, soil respiration, and soil enzyme activities can be significantly decreased in cultivated areas compared to forests (Meena and Rao 2021). Another important aspect of soil ecosystems is the moisture available in the soil. Soil can act as a filter for water and thus regulate the water cycle. Soils with low soil moisture retention can thus limit plant growth and affect soil microbial diversity and activity (Wan et al. 2021). Soil water supply in agricultural soils may be limited due to changes in climatic conditions (e.g. lower rainfall), increased plant water uptake (temporal increase in root growth and root density or increase in transpiration), or changes in soil physical properties (increase in evaporation). Soil chemistry can also influence soil water content, for example, organic matter can increase soil water retention (Lal 2020), thereby promoting healthy ecosystem functioning.
Land-use types can have a strong influence on soil microbial communities due to the changes in soil abiotic conditions (Ye and Wright 2010; Tian et al. 2017), even more than geographical conditions (Lee et al. 2020). In recent decades, several studies have addressed the relationship between different soil management practices and changes in the structure of soil microbial communities from local to regional scales in different climate zones (Acosta-Martínez et al. 2008; Lauber et al. 2013; Rampelotto et al. 2013; Hartmann et al. 2015; Thomson et al. 2015; Peng et al. 2017; Lynn et al. 2017; Szoboszlay et al. 2017; Plassart et al. 2019; Petersen et al. 2019; Barnett et al. 2020; Sengupta et al. 2020; Li et al. 2020; Lee et al. 2020; Xue et al. 2022). The main drivers of the shifts in the diversity of soil bacterial communities can be related to the different land-use intensity (e.g. from arable field to permanent grassland and forest) (Gschwend et al. 2022), the soil tillage or monocropping systems and the fertilization scheme (Hartmann et al. 2015; Barnett et al. 2020), as well as the soil properties (e.g. pH, resource availability and texture) (Thomson et al. 2015; Peng et al. 2017; Plassart et al. 2019; Barnett et al. 2020; Gschwend et al. 2022). The results of various studies performed up to now suggest that the diversity of soil microbial communities responds in different ways to the different land-use types, mainly depending on the variation of abiotic factors in soils.
To expand our knowledge on soil ecosystem health, it is therefore necessary to further study the effects of human interventions based on different land-use types on soil microbial community composition on a local scale, under similar soil and weather conditions. It was hypothesized that both the varying degree of anthropogenic impacts and the slope positions can significantly alter the soil physical and chemical parameters and consequently the composition of soil bacterial communities.
The aim of the present study was, therefore, to explore how the taxonomic composition of the soil bacterial communities of geographically close but different land-use types in a small catchment area is affected by the physical–chemical properties of the soil, the slope gradient and the seasonal variation.
Materials and methods
Site description and land-use types
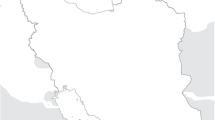
The Csorsza catchment is in the Balaton Uplands, Hungary, which is an agriculturally dominated area, with the most typical land-use types being the forest, grassland or pasture, vineyards, croplands, and orchards. It is in the Pannonian biogeographical zone, with a continental climate. Normally, it is characterized by hot summers and cold winters (Dövényi 2010). The soil in the area is generally Cambisols and Calcisols (Schad et al. 2016). The cropland is located at the catchment valley, where Endoleptic Cambisols are found (Horel et al. 2022), as the loose sediment is more than a meter thick. Part of the soil is formed from loess, and carbonates are present at deeper horizons of the soil profiles (Horel et al. 2022). The map of the catchment and the points indicating the sampling sites are shown in Fig. 1.
The forest sampling site is a mixture of sessile oak (Quercus petraea L.) and black locust (Robinia pseudoacacia L.) and is located on an approximately 8% slope. This study site is anthropogenically not disturbed, e.g., even the dead trees are kept on site.
On the grassland sampling site, the topsoil is shallow (about 40–45 cm deep) and usually very low in soil moisture due to a long-term soil erosion. This site is strongly affected by low rainfall.
The vineyard sampling site has not been tilled in previous years and there is grass between the rows of vines. The vines are weeded if necessary. A light topping is performed yearly. Organic fertilizer (sheep manure) is being applied approximately every 5-year intervals. The study slope has a white wine variety of grapes called furmint (Vitis vinifera L.).
The arable field (cropland) sampling site has normally inorganic fertilizer (carbamide of 100 kg N/ha) application annually and crop rotation is being practiced. During the studied period maize (Zea mays L.) was cultivated, while before that winter wheat (2019), sunflower (2018), or triticale (2017) were sown. It is in the valley of the catchment, with a slope of less than 5%.
The other studied areas (forest, grassland, and vineyard) are located on slopes and prone to erosion. There is no irrigation, the entire catchment is fed by rainwater.
Sampling
The geographical location of the sampling sites, i.e., the forest (F; 46.91285 N, 17.69751 E), the grassland (G; 46.912236 N, 17.697514 E), the vineyard (W; 46.91654 N, 17.689753 E) and the arable field (A; 46.92648 N, 17.68248 E) is shown in Fig. 1. Samples were taken from the upper 10 cm of topsoil in the four different land-use types of the small catchment. Sampling sites were designated at the upper (U) and lower (L) positions of the slopes. Samplings took place twice in 2020, in the middle and at the end of the vegetation period, in July (0720) and October (1020), respectively. In both periods, soil samples were taken in three replicates (A, B, C) in all four sampling areas. A total of 48 samples were included in the analyses.
Approximately 500 g of soil samples were collected in disposable plastic bags and stored at 6–8 °C until laboratory processing within 24 h. The samples were divided into two parts: approximately 10 g was put in sterile microtubes for subsequent environmental DNA isolation, and the other part was air-dried for soil physical and chemical analyses.
Soil physical and chemical analyses
Particle size distribution was determined using the sieve-pipette method. Briefly, the soil:water suspension was mixed in a sedimentation cylinder and sampled with a pipette. After sieveing the air-dried soil samples (< 2 mm), the soil NH4+–N, NO3−–N, total nitrogen (NTot) (determined using the Kjeldhal method; ISO 11261:1995), K2O (Al soluble), P2O5 (Al soluble) (Quotation ICP-OES, Ultima 2, Thermo Fischer Scientific, Waltham, MA, USA), soil organic carbon (SOC) content (wet oxidation method according to the Tyurin method), electrical conductivity, and pHH2O (WTW multi 350i multiline pH meter with 1:2.5 soil to water suspension ratio) were determined. Detailed description can be found in Horel and Zsigmond (2023).
An EGM-5 (PPSYSTEMS, U.S.) in situ IR gas-analyzer was used to determine CO2 emissions, where the measurements were based on two minutes of airtight measurements (Dencső et al. 2021). The volumetric soil water contents (SWC) of the upper 10 cm of the soil horizons for all measurement points were determined by Campbell Scientific Hydrosense II SWC probe with an accuracy of 3% and a resolution of < 0.05%. Detailed descriptions can be found in (Dencső et al. 2021).
Environmental DNA extraction and 16S rRNA gene-based amplicon sequencing
Environmental DNA from the soil samples was extracted using DNeasy powersoil kit (QIAGEN, Hilden, Germany) and the automated QIAcube system (Qiagen) according to the manufacturer’s instructions. The library construction and sequencing of 16S rRNA gene amplicons were performed at the RTSF Genomics Core at Michigan State University (East Lansing, USA). The V4 region of the 16S rRNA gene was amplified using dual-indexed primers (515F/806R) following a one-step PCR method developed by Kozich et al. (2013). The normalized, pooled, purified, and high-quality libraries were sequenced in a 2 × 250 bp paired-end format using a MiSeq v2 500 cycle reagent cartridge on the Illumina MiSeq platform (Illumina Inc., San Diego, USA).
Bioinformatic and statistical analysis
Raw sequencing data is available at sequence read archive (SRA) under the BioProject accession number PRJNA 782964. The reads were processed using mothur v1.44.3 (Schloss et al. 2009) following the developers’ pipeline (www.mothur.org/wiki/MiSeq_SOP accessed at 01.07.2021) (Kozich et al. 2013). Chimeras identified with VSEARCH (Rognes et al. 2016) and singletons were removed before further analysis (Kunin et al. 2010). Sequences were clustered into operational taxonomic units (OTU) at 97% identity threshold (Tindall et al. 2010), then taxonomy was assigned using a minimum bootstrap confidence value of 80% based on the ARB-SILVA v138 SSU reference database was used for taxonomic assignment (Quast et al. 2012). All non-bacterial sequences were excluded. Richness and diversity indices (i.e. ACE, Chao1, inverse Simpson) were calculated for subsampled datasets (n = 28,027) with mothur’s built-in calculators.
Results were displayed using PAST 4.15 (PAleontological STatistics) software (Hammer et al. 2001). For the boxplot visualization, the predicted number of OTUs (Sobs), and the estimated richness (ACE, Chao1) and diversity (Inverse Simpson) indices of the bacterial communities by land-use type was used. For the hierarchical cluster analysis, the subsampled sequences of bacterial phyla were applied using the Bray–Curtis similarity index. The non-metric multidimensional scaling (NMDS) ordination was performed using OTUs occurring at a relative abundance above 1% in at least one sample.
The factors of slope position and land-use types on the investigated soil parameters were analyzed using nonparametric statistical analyses of the Wilcoxon signed rank test or the Wilcoxon rank sum and Kruskal–Wallis test. Spearman’s correlation coefficient (r) was used to calculate the linear correlation between the soils’ physical and chemical characteristics and the taxonomy data. The principal component analysis (PCA) was applied to explore the factor pattern of the selected soil parameters and taxonomy. All statistical calculations were performed using the software package R (R Core Team, Version 4.0.2). Statistical significance of the data sets was determined at p < 0.05.
Results
Soil physical and chemical characteristics
The particle size distribution, soil chemical parameters, the average water content of the soil samples, and the average soil CO2 fluxes are presented in Table 1. As the forest upper point had a high amount of plant tissues in the samples, the chemical parameters were measured, but not included in many analyses due to the uncertainty of its quality.
Among land-use types, significant differences were found in many physical and chemical parameters of the soil using Kruskal–Wallis test. The highest soil water content (SWC) was measured for the arable field site, along with the highest pH, P2O5, and the lowest soil organic carbon (SOC) and total nitrogen contents. Forest showed significantly higher soil organic carbon (SOC) compared to the other land-use types.
Differences in soil physical and chemical characteristics were also found among the different slope positions, but these differences were not as pronounced as between land-use types. The SWC, CO2 fluxes, and C/N data showed no significant differences among slope positions within a land-use type (p > 0.05, W = 14–26, U(df) = 1 for SWC, p > 0.05, W = 6–19, U(df) = 1, and p > 0.05, W = 13–21, U(df) = 1 for total N). The SOC and total nitrogen contents for vineyard and grassland showed significantly higher values for the lower sites compared to the higher ones, while had higher SOC contents along with grassland compared to vineyard or arable field (Table 1).
The PCA analyses revealed that the first principal component (PC1) accounted for 49.28% of the variation caused by the interaction, while PC2 accounted for 18.12% (Fig. 2). The objects showed a distribution primarily by land-use type, and secondarily by upper and lower slope positions, especially for forest and vineyard. The parallel samples from different times were not separated by soil physical and chemical properties. Based on the Spearman correlation analyses of all land-use physical and chemical data, we observed strong and significant correlations between SOC and sand contents (r = 0.88; p < 0.001; n = 48), negative correlations between SOC and clay contents (r = − 0.71; p < 0.001; n = 48). Negative correlations were observed between CO2 fluxes and SWC (r = − 0.56; p < 0.001; n = 48). When only July data was studied, soil temperature and CO2 fluxes showed strong negative correlations (r = − 0.73; p < 0.001; n = 18), while in October moderately strong positive connections were observed (r = 0.64; p = 0.006; n = 10).
Principal component analysis (PCA) ordination biplot for the different land-use types of forest (green), grassland (orange), vineyard (blue), and arable field (red) and lower (square) or upper (circle) slope positions, with environmental variables (selected soil physical and chemical parameters) represented as vectors, and the pairwise representation of the study sites (SWC: soil water content; SOC: soil organic carbon content; Total N: total nitrogen content; CN is the carbon to nitrogen ratio; (n = 48))
Diversity and taxonomic composition of soil bacterial communities
The amplicon sequencing resulted in a total of 2,875,629 bacterial sequences from the 47 out of 48 soil samples, which were assigned into 1163 OTUs. One of the soil samples (AU0720B) could not be processed for technical reasons. The sequences represented the bacterial communities well, as the Good’s coverage value was at least 0.96 in the studied samples (Supplementary Table 1). Both, the predicted number of OTUs (Sobs), and the estimated richness (ACE, Chao1) and diversity (Inverse Simpson) indices showed differences according to land-use types (Fig. 3). The highest median values, with the largest standard deviation, were found for all four parameters for the forest samples. This was followed by the grassland, then the vineyards and the arable field, with much smaller but very similar values. Higher average OTU numbers, as well as richness and diversity values were estimated in the July soil samples than in the October ones, except for vineyards. The average values of the indices were generally higher in the upper soil samples of the slopes than in the lower ones. The opposite trend was only observed in the forest samples.
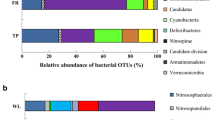
Of the 77 phyla or phylum-level taxonomic lineages detected in the soil samples, Pseudomonadota (formerly Proteobacteria), Acidobacteriota (formerly Acidobacteria), Actinobacteriota (formerly Actinobacteria), Bacteroidota (formerly Bacteroidetes), Verrucomicrobiota (formerly Verrucomicrobia), Gemmatimonadota (formerly Gemmatimonadetes), Myxococcota, Planctomycetota (formerly Planctomycetes), Chloroflexota (formerly Chloroflexi), Bdellovibrionota (formerly within the Deltaproteobacteria) and Elusimicrobiota (formerly Termite Group 1) had a relative abundance of 1% or more in at least one sample. All soil samples were dominated by representatives of phyla Pseudomonadota, Acidobacteriota, Actinobacteriota, Bacteroidota and Verrucomicrobiota (Fig. 4). The primary separation of the samples was observed according to the sampling times. The mean relative abundance of phyla Actinobacteriota, Bacteroidota, and Verrucomicrobiota was higher in July, while that of phyla Pseudomonadota, Acidobacteriota, and Gemmatimonadota was higher in October. The mean relative abundance of the four dominant phyla was different by land-use types, such as:, F-W-G-A (29%-25%-24%-21%) for Pseudomonadota, A-F-G-W (21%-19%-18%-18%) for Acidobacteriota, G-F-W-A (18%-13%-13%-13%) for Actinobacteriota, W-G-F-A (15%-13%-11%-10%) for Bacteroidota and F-G-A-W (10%-8%-8%-7%) for Verrucomicrobiota. Based on the taxonomic distribution at the phylum level, samples from the same land-use were generally clustered in pairs in the different sampling times. There were also no significant differences in the mean relative abundance values of the phyla according to the slope positions.
Percentage distribution of 16S rRNA gene amplicon sequences of bacterial phyla detected from soil samples of different land-use types together with the results of Bray–Curtis similarity index-based UPGMA cluster analysis (sample signs for land-use types: F, forest; G, grassland; W, vineyard; A, arable field; for slope positions: U, upper; L, lower; for sampling times: 0720, 2020 July; 1020, 2020 October.)
Effects of land-use types on the bacterial communities
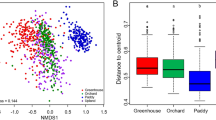
The impact of land-use type on the composition of bacterial communities can be observed at OTU level (Fig. 5). Based on the nonmetric multidimensional scaling (NMDS) ordination, the samples from the cultivated areas were clearly separated from that of the non-cultivated areas along the first coordinate. The arable field (A) and vineyard (W) samples as well as the forest (F) and grassland (G) samples were separated along the second coordinate. Only the arable field samples showed clear separation by sampling time (July versus October). Differences in topography led to the clear separation of sampling sites (i.e. upper and lower sites of the slopes) only in the forest land-use.
NMDS ordination based on Bray–Curtis similarity index of the bacterial OTUs, from the soil samples of different land-use types (Stress = 0.2678. Vectors indicate the most abundant 15 OTUs. Color codes for land use types: green is the forest, orange is the grassland, blue is the vineyard, and red is the arable field.)
The most abundant and identified genera were Sphingomonas (Pseudomonadota), Chthoniobacter (Verrucomicrobiota), Udaeobacter (Verrucomicrobiota) and Gemmatimonas (Gemmatimonadota). The majority of OTUs, however, could only be identified at order or higher taxonomic level. The OTUs responsible for the separation of the soil samples according to land-use types were: Comamonadaceae (Pseudomonadota) and 67–14 (Actinobacteriota) for grassland, Udaeobacter and Gemmatimonas for forest, Vicinamibacteraceae (Acidobacteriota), WD2101 (Planctomycetota) and an uncultured Gemmatimonadota for arable field, and Chthoniobacter, Chitinophagaceae (Bacteroidota), Pedosphaeraceae (Verrucomicrobiota) and an uncultured Acidobacteriota for vineyard (Fig. 5).
Discussion
In the present study, the different land-use types on the diversity of the soil bacterial communities in a small, sloping catchment area were investigated. To determine the factors most influencing the structure of the microbial communities, the physical and chemical parameters of the soil, as well as the CO2 emissions and water content of the soil were measured. As many areas in a given catchment are positioned on different angled slopes, it was also investigated how the soil redistribution processes affect the distribution of microorganisms in the soil. As the events of the vegetation period are also important factors in terms of the diversity of soil microbial communities, the examinations were performed at two sampling times, during the summer flowering and after the autumn harvest before leaf fall.
The distribution of dominant phyla Pseudomonadota, Acidobacteria, Actinobacteria, Bacteroidetes, and Verrucomicrobia of the soil bacterial communities from different land-use areas in the small catchment were similar to previous studies (Peng et al. 2017; Plassart et al. 2019; Sengupta et al. 2020; Li et al. 2020; Lee et al. 2020; Kim et al. 2021). These phyla were also found as core bacterial taxa in maize fields under different long-term fertilization regimes and in a nearby fallow land in Hungary (Megyes et al. 2021).
The mean relative abundance of the phylum Pseudomonadota was the highest for all land-use types of the examined small catchment area. It is in line with the results of a European transect study of forests, grasslands, and arable fields (Plassart et al. 2019), and long-term land-use practices of maize fields and adjacent grassland and forest (Sengupta et al. 2020), as well as saline-alkaline soil with different land-uses such as agricultural land, forest land, and grassland (Peng et al. 2017).
The highest relative abundance of the phylum Pseudomonadota was detected in the sample collected from the forest in October from the upper position of the slope. Here, the long-term undisturbed soil condition, the high organic matter content, and the highest amount of nitrogen forms, as well as the stable soil structure, may have contributed to the highest taxonomic diversity of Pseudomonadota. The influence of the topography in the small catchment area was also mainly observed in the undisturbed forest, based on the differences in the physical and chemical properties of the samples from the upper and lower positions of the slope. These differences were also reflected to a lesser extent in the bacterial community structures of the forest soil, based on the OTU-level analysis.
Among the genera identified within the phylum Pseudomonadota, Sphingomonas was one of the most common in all land use types, especially in grassland. Members of this genus have a broad catabolic potential for the degradation of recalcitrant organic pollutants, including aromatic and polyaromatic hydrocarbons, and pesticides (White et al. 1996), which is especially important in non-organic farming. The genus Sphingomonas can be considered as a common member of bacterial communities in different land-use types and reacts sensitively to changes in land use (Peng et al. 2017; Szoboszlay et al. 2017; Plassart et al. 2019; Wu et al. 2019; Sengupta et al. 2020; Li et al. 2020). Szoboszlay et al. (2017) found that the conversion of cropland to forest caused a decrease, while the conversion of grassland to cropland caused a significant increase in the relative abundance of Sphingomonas. The relative abundance of Sphingomonas was reported the highest in a maize field, compared to different land use types (natural secondary Quercus mongolica forest, shrubland, Larix gmelinii and Pinus koraiensis land) and agricultural land (Z. mays) in northeast China (Wu et al. 2019).
The proportion of the phylum, Acidobacteriota (formerly Acidobacteria), was almost the same in all land-use types examined. The pH was previously ascribed to be the main environmental factor determining the distribution of the phylum Acidobacteria, widespread in soils but with a largely unknown taxonomic composition (Jones et al. 2009; Kuramae et al. 2012; Kim et al. 2021). Although significant differences were detected in soil pH values among the different land use types of the small catchment, they all fell in the neutral range, favoring only the neutrophilic phylotypes of Acidobacteria. The separation of the highest pH (7.6) of the arable field from other land use types was mainly due to its close positive correlation with Vicinamibacteraceae. Members of the Vicinamibacteraceae fam. nov. described by Huber and Overmann (2018) are aerobic chemoorgano-heterotrophs, psychrotolerant to mesophilic and prevail in soils with neutral pH (Neal et al. 2021). Consistent with our results, significant negative correlation of temperature with relative abundance of Vicinamibacteraceae was detected by Wang et al. (2023) in newly formed wetlands in coal mining subsidence areas in Jining (Shandong Province, China). In addition to the physical–chemical properties of the soil, changes in the phenological stages of the maize plant may also have played an important role in the seasonal separation of the arable field samples. The effect of maize growth stages on soil bacterial community structure and activity has been demonstrated previously (Baudoin et al. 2002; Li et al. 2014; Bonkowski et al. 2021).
The relative abundance of the phylum Actinobacteriota was almost a third higher in the grassland than in the other areas. The high relative abundance of this phylum in different land-use types is consistent with the results of other studies (Peng et al. 2017; Szoboszlay et al. 2017; Plassart et al. 2019; Sengupta et al. 2020; Megyes et al. 2021). The phylum Actinobacteria is not only taxonomically but also metabolically diverse and plays a major role in the soil carbon cycle through the transformation and/or degradation of a wide variety of organic matter. These so-called K-strategist bacteria are mostly slow-growing and very efficient in the use of resources.
Of the land use types studied, the shallow topsoil of grassland had the lowest moisture content, where rainwater infiltration was also limited due to dense vegetation. In this area of the catchment, drought-tolerant plant species such as Helianthemum nummularium L., Salvia pratensis L. or Thymus sp. form part of the vegetation (Horel et al. 2022). The high relative abundance of the phylum Actinobacteriota, especially during the warm and dry season in July, could be related to the fact that many of its members can survive harsh environmental conditions (e.g. drought and/or shortage of resources) for a long time. Jenkins et al. (2009) found that the distribution of Micrococcus, Nocardioides, Streptomyces, Micromonospora and Mycobacterium was influenced by soil water content in the Northumberland (UK) long term managed grassland.
The most abundant OTUs of Actinobacteria primarily responsible for the separation of grassland samples could not be identified at genus level. Felske et al. (1997) previously reported that novel uncultured Actinobacteria may play a physiologically active role in native microbial communities of grassland soils. Nevertheless, the high taxonomic diversity of this phylum was indicated by the identification of more than a hundred genera that were generally present in the samples of the small catchment at relative abundances of less than 1%. These included genera that are well known and widespread on a wide range of soils (e.g. Corynebacterium, Mycobacterium, Gordonia, Nocardia, Rhodococcus, Frankia, Cellulomonas, Agromyces, Nesterenkonia, Pseudarthrobacter, Micromonospora, Nocardioides, Streptomyces, Gaiella, Solirubrobacter), but also genera described from soils in recent years (e.g. Pseudactinotalea, Actinocatenispora, Stackebrandtia, Mumia, Tenggerimyces). In studying the relationship between Actinobacteria and different land uses, Szoboszlay et al. (2017) designated as indicative taxa Nocardiaceae for cropland soils, Streptomyces and Solirubrobacterales for grassland, and Mycobacterium for forest. In another study, sequences assigned to the genus Gaiella were found in much higher abundance in fallow than in different long-term fertilization regimes of maize fields (Megyes et al. 2021).
The relative abundance of the phylum Bacteroidota was the most abundant in the vineyard soil samples from the lower position of the slope. Of the land use types studied, this one had the highest clay content and the highest concentration of available phosphate. Its organic carbon and nitrogen content was low, as was that of arable field. Although grass strips were planted between the grape rows to reduce soil erosion, there was a significant difference between the upper and lower positions of the slope in terms of several soil parameters. Erosion processes resulting from land use and topography affected soil bacterial communities, as well. Changes at the lower position of the slope favored the increase in the phylum Bacteroidota. These so-called r-strategist bacteria exploit the sudden increase in resources resulting from changing and/or unpredictable environmental conditions by growing rapidly. Diversity shifts in soil bacterial communities from K-to r-strategist (e.g. Bacteroidetes) were also reported by Peng et al. (2022) as an adaptation to environmental disturbance caused by grassland degradation.
The family Chitinophagaceae (Bacteroidota), which was responsible for the separation of the vineyard, is widely distributed in different soils, including vineyards (Gupta et al. 2019; Nerva et al. 2021). Strains of Chitinophagaceae can be isolated or recovered from cow manure and compost (Storey et al. 2015; Ren et al. 2016). Its dominant presence in the soil of the small catchment vineyard may therefore be related to the organic fertilizer applied to this area. In black soil, the addition of cow dung significantly increased the relative abundance of e.g. the family Chitinophagaceae compared to untreated soil, thus promoting the degradation of organic pollutants (Cheng et al. 2023).
Although the phylum Verrucomicrobiota represents globally distributed soil bacteria, it still has only a few described species (Floyd et al. 2005). Among the sequences of the phylum Verrucomicrobiota, those related to the genus Udaeobacter were the most distinguishable for the forest, while those of the genus Chthoniobacter and the family Pedosphaeraceae for the vineyard. Willms et al. (2020) reported that “Candidatus” Udaeobacter may have advantages over other soil bacteria in grassland and forest ecosystems through its antibiotic release and multidrug resistance, and by using hydrogen as an alternative electron donor. Studying the metabolically active microbiome in vineyard soil, Rosado-Porto et al. (2023) found that the relative abundance of amplicon sequence variants of both Chthoniobacter and Pedosphaeraceae was positively stimulated under elevated CO2 concentrations.
Conclusions for future biology
Anthropogenic disturbances can have significant effects on the structure of soil bacterial communities. The results showed that both soil physical–chemical properties and bacterial taxonomic diversity resulted in a primary separation between untilled (forest and grassland) and tilled (vineyard and arable field) soil samples from the small catchment. The undisturbed forest soils with the highest organic carbon content had the highest bacterial taxonomic diversity. Regarding the bacterial community structures, the effect of the slope gradient was pronounced for the forest soil, while the effect of seasonality was observed for the arable field. This highlights the importance of studying soil–plant-microbe systems as holobionts in the future.
References
Acosta-Martínez V, Dowd S, Sun Y, Allen V (2008) Tag-encoded pyrosequencing analysis of bacterial diversity in a single soil type as affected by management and land use. Soil Biol Biochem 40:2762–2770. https://doi.org/10.1016/j.soilbio.2008.07.022
Barnett SE, Youngblut ND, Buckley DH (2020) Soil characteristics and land-use drive bacterial community assembly patterns. FEMS Microbiol Ecol 96:fiz194. https://doi.org/10.1093/femsec/fiz194
Baudoin E, Benizri E, Guckert A (2002) Impact of growth stage on the bacterial community structure along maize roots, as determined by metabolic and genetic fingerprinting. Appl Soil Ecol 19:135–145. https://doi.org/10.1016/S0929-1393(01)00185-8
Bonkowski M, Tarkka M, Razavi BS et al (2021) Spatiotemporal dynamics of maize (Zea mays L.) root growth and its potential consequences for the assembly of the rhizosphere microbiota. Front Microbiol. https://doi.org/10.3389/fmicb.2021.619499
Cheng L, Wang L, Wang X et al (2023) The various effect of cow manure compost on the degradation of imazethapyr in different soil types. Chemosphere 337:139325. https://doi.org/10.1016/j.chemosphere.2023.139325
Dencső M, Horel Á, Bogunovic I, Tóth E (2021) Effects of environmental drivers and agricultural management on soil CO2 and N2O emissions. Agronomy 11:54. https://doi.org/10.3390/agronomy11010054
Dövényi Z (2010) Magyarország kistájainak katasztere (in Hungarian) MTA Földrajztudományi Kutatóintézet. Budapest, Hungary
Felske A, Rheims H, Wolterink A et al (1997) Ribosome analysis reveals prominent activity of an uncultured member of the class Actinobacteria in grassland soils. Microbiology (NY) 143:2983–2989. https://doi.org/10.1099/00221287-143-9-2983
Floyd MM, Tang J, Kane M, Emerson D (2005) Captured diversity in a culture collection: case study of the geographic and habitat distributions of environmental isolates held at the American type culture collection. Appl Environ Microbiol 71:2813–2823. https://doi.org/10.1128/AEM.71.6.2813-2823.2005
Gschwend F, Hartmann M, Mayerhofer J et al (2022) Site and land-use associations of soil bacteria and fungi define core and indicative taxa. FEMS Microbiol Ecol 97:fiab165. https://doi.org/10.1093/femsec/fiab165
Gupta VVSR, Bramley RGV, Greenfield P et al (2019) Vineyard soil microbiome composition related to rotundone concentration in Australian cool climate ‘peppery’ shiraz grapes. Front Microbiol. https://doi.org/10.3389/fmicb.2019.01607
Hammer Ø, Harper DATDA, Ryan PDPD (2001) PAST: paleontological statistics software package for education and data analysis. Paleontologica Electronica 4:1–9
Hartmann M, Frey B, Mayer J et al (2015) Distinct soil microbial diversity under long-term organic and conventional farming. ISME J 9:1177–1194. https://doi.org/10.1038/ismej.2014.210
Horel Á, Zsigmond T (2023) Plant growth and soil water content changes under different inter-row soil management methods in a sloping vineyard. Plants 12:1549. https://doi.org/10.3390/plants12071549
Horel Á, Zsigmond T, Molnár S et al (2022) Long-term soil water content dynamics under different land uses in a small agricultural catchment. J Hydrol Hydromech 70:284–294. https://doi.org/10.2478/johh-2022-0015
Huber KJ, Overmann J (2018) vicinamibacteraceae fam. nov., the first described family within the subdivision 6 acidobacteria. Int J Syst Evol Microbiol 68:2331–2334. https://doi.org/10.1099/ijsem.0.002841
Hussain I, Olson KR, Ebelhar SA (1999) Long-term tillage effects on soil chemical properties and organic matter fractions. Soil Sci Soc Am J 63:1335–1341. https://doi.org/10.2136/sssaj1999.6351335x
Jenkins SN, Waite IS, Blackburn A et al (2009) Actinobacterial community dynamics in long term managed grasslands. Antonie Van Leeuwenhoek 95:319–334. https://doi.org/10.1007/s10482-009-9317-8
Jones RT, Robeson MS, Lauber CL et al (2009) A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J 3:442–453. https://doi.org/10.1038/ismej.2008.127
Kim H-S, Lee S-H, Jo HY et al (2021) Diversity and composition of soil acidobacteria and proteobacteria communities as a bacterial indicator of past land-use change from forest to farmland. Sci Total Environ 797:148944. https://doi.org/10.1016/j.scitotenv.2021.148944
Kozich JJ, Westcott SL, Baxter NT et al (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq illumina sequencing platform. Appl Environ Microbiol 79:5112–5120. https://doi.org/10.1128/AEM.01043-13
Kunin V, Engelbrektson A, Ochman H, Hugenholtz P (2010) Wrinkles in the rare biosphere: pyrosequencing errors can lead to artificial inflation of diversity estimates. Environ Microbiol 12:118–123. https://doi.org/10.1111/j.1462-2920.2009.02051.x
Kuramae EE, Yergeau E, Wong LC et al (2012) Soil characteristics more strongly influence soil bacterial communities than land-use type. FEMS Microbiol Ecol 79:12–24. https://doi.org/10.1111/j.1574-6941.2011.01192.x
Lal R (2020) Soil organic matter and water retention. Agron J 112:3265–3277. https://doi.org/10.1002/agj2.20282
Lauber CL, Ramirez KS, Aanderud Z et al (2013) Temporal variability in soil microbial communities across land-use types. ISME J 7:1641–1650. https://doi.org/10.1038/ismej.2013.50
Lee SA, Kim JM, Kim Y et al (2020) Different types of agricultural land use drive distinct soil bacterial communities. Sci Rep 10:17418. https://doi.org/10.1038/s41598-020-74193-8
Li X, Rui J, Mao Y et al (2014) Dynamics of the bacterial community structure in the rhizosphere of a maize cultivar. Soil Biol Biochem 68:392–401. https://doi.org/10.1016/j.soilbio.2013.10.017
Li Y, Zeng C, Long M (2020) Variation of soil nutrients and bacterial community diversity of different land utilization types in Yangtze River Basin. Chongqing Municipality Peerj 8:e9386. https://doi.org/10.7717/peerj.9386
Lynn TM, Liu Q, Hu Y et al (2017) Influence of land use on bacterial and archaeal diversity and community structures in three natural ecosystems and one agricultural soil. Arch Microbiol 199:711–721. https://doi.org/10.1007/s00203-017-1347-4
Meena A, Rao KS (2021) Assessment of soil microbial and enzyme activity in the rhizosphere zone under different land use/cover of a semiarid region. India Ecol Process 10:16. https://doi.org/10.1186/s13717-021-00288-3
Megyes M, Borsodi AK, Árendás T, Márialigeti K (2021) Variations in the diversity of soil bacterial and archaeal communities in response to different long-term fertilization regimes in maize fields. Appl Soil Ecol 168:104120. https://doi.org/10.1016/j.apsoil.2021.104120
Neal AL, Hughes D, Clark IM et al (2021) Microbiome aggregated traits and assembly are more sensitive to soil management than diversity. Msystems. https://doi.org/10.1128/mSystems.01056-20
Nerva L, Moffa L, Giudice G et al (2021) Microscale analysis of soil characteristics and microbiomes reveals potential impacts on plants and fruit: vineyard as a model case study. Plant Soil 462:525–541. https://doi.org/10.1007/s11104-021-04884-2
Obour AK, Mikha MM, Holman JD, Stahlman PW (2017) Changes in soil surface chemistry after fifty years of tillage and nitrogen fertilization. Geoderma 308:46–53. https://doi.org/10.1016/j.geoderma.2017.08.020
Peng J, Liu H, Hu Y et al (2022) Shift in soil bacterial communities from K to r-strategists facilitates adaptation to grassland degradation. Land Degrad Dev 33:2076–2091. https://doi.org/10.1002/ldr.4304
Peng M, Jia H, Wang Q (2017) The effect of land use on bacterial communities in saline–alkali soil. Curr Microbiol 74:325–333. https://doi.org/10.1007/s00284-017-1195-0
Petersen IAB, Meyer KM, Bohannan BJM (2019) Meta-analysis reveals consistent bacterial responses to land use change across the tropics. Front Ecol Evol 7:00391. https://doi.org/10.3389/fevo.2019.00391
Plassart P, Prévost-Bouré NC, Uroz S et al (2019) Soil parameters, land use, and geographical distance drive soil bacterial communities along a European transect. Sci Rep 9:605. https://doi.org/10.1038/s41598-018-36867-2
Quast C, Pruesse E, Yilmaz P et al (2012) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
Rampelotto PH, de Siqueira FA, Barboza ADM, Roesch LFW (2013) Changes in diversity, abundance, and structure of soil bacterial communities in Brazilian Savanna under different land use systems. Microb Ecol 66:593–607. https://doi.org/10.1007/s00248-013-0235-y
Ren G, Xu X, Qu J et al (2016) Evaluation of microbial population dynamics in the co-composting of cow manure and rice straw using high throughput sequencing analysis. World J Microbiol Biotechnol 32:101. https://doi.org/10.1007/s11274-016-2059-7
Rognes T, Flouri T, Nichols B et al (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Rosado-Porto D, Ratering S, Wohlfahrt Y et al (2023) Elevated atmospheric CO2 concentrations caused a shift of the metabolically active microbiome in vineyard soil. BMC Microbiol 23:46. https://doi.org/10.1186/s12866-023-02781-5
Schad P, van Huyssteen C, Micheli E (eds) (2016) IUSS working group WRB (2014) world reference base for soil resources 2014. FAO, Rome
Schloss PD, Westcott SL, Ryabin T et al (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Sengupta A, Hariharan J, Grewal PS, Dick WA (2020) Bacterial community dissimilarity in soils is driven by long-term land-use practices. Agrosyst Geosci Environ 3:e20031. https://doi.org/10.1002/agg2.20031
Storey S, Chualain DN, Doyle O et al (2015) Comparison of bacterial succession in green waste composts amended with inorganic fertiliser and wastewater treatment plant sludge. Bioresour Technol 179:71–77. https://doi.org/10.1016/j.biortech.2014.11.107
Szoboszlay M, Dohrmann AB, Poeplau C et al (2017) Impact of land-use change and soil organic carbon quality on microbial diversity in soils across Europe. FEMS Microbiol Ecol 93:fix146. https://doi.org/10.1093/femsec/fix146
Thomson BC, Tisserant E, Plassart P et al (2015) Soil conditions and land use intensification effects on soil microbial communities across a range of European field sites. Soil Biol Biochem 88:403–413. https://doi.org/10.1016/j.soilbio.2015.06.012
Tian Q, Taniguchi T, Shi W-Y et al (2017) Land-use types and soil chemical properties influence soil microbial communities in the semiarid loess plateau region in China. Sci Rep 7:45289. https://doi.org/10.1038/srep45289
Tindall BJ, Rosselló-Móra R, Busse HJ et al (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266. https://doi.org/10.1099/ijs.0.016949-0
Wan X, Chen X, Huang Z, Chen HYH (2021) Global soil microbial biomass decreases with aridity and land-use intensification. Glob Ecol Biogeogr 30:1056–1069. https://doi.org/10.1111/geb.13282
Wang M, Sun M, Zhao Y et al (2023) Seasonal changes of soil microbiota and its association with environmental factors in coal mining subsidence area. AMB Express 13(1):147. https://doi.org/10.1186/s13568-023-01653-5
White DC, Sutton SD, Ringelberg DB (1996) The genus Sphingomonas: physiology and ecology. Curr Opin Biotechnol 7:301–306. https://doi.org/10.1016/S0958-1669(96)80034-6
Willms IM, Rudolph AY, Göschel I et al (2020) Globally abundant Candidatus Udaeobacter benefits from release of antibiotics in soil and potentially performs trace gas scavenging. Msphere. https://doi.org/10.1128/mSphere.00186-20
Wu S-J, Deng J-J, Yin Y et al (2019) Bacterial community changes associated with land use type in the forest montane region of Northeast China. Forests 11:40. https://doi.org/10.3390/f11010040
Xue P, Minasny B, McBratney AB (2022) Land-use affects soil microbial co-occurrence networks and their putative functions. Appl Soil Ecol 169:104184. https://doi.org/10.1016/j.apsoil.2021.104184
Ye R, Wright AL (2010) Multivariate analysis of chemical and microbial properties in histosols as influenced by land-use types. Soil Tillage Res 110:94–100. https://doi.org/10.1016/j.still.2010.06.013
Funding
Open access funding provided by Eötvös Loránd University. The research is based upon work supported by the Hungarian National Research Fund (OT-KA/NKFI) project OTKA FK-131792 and by the Sustainable Development and Technologies National Program of the Hungarian Academy of Sciences (FFT NP FTA). This research was performed within the framework of the Széchenyi Plan Plus program with the support of the RRF 2.3.1 21 2022 00008 project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Borsodi, A.K., Megyes, M., Zsigmond, T. et al. Soil bacterial communities affected by land-use types in a small catchment area of the Balaton Uplands (Hungary). BIOLOGIA FUTURA (2024). https://doi.org/10.1007/s42977-024-00233-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42977-024-00233-3