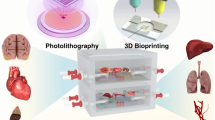
Abstract
Three-dimensional cancer-on-a-chip tissue models aim to replicate the key hallmarks of the tumour microenvironment and allow for the study of dynamic interactions that occur during tumour progression. Recently, complex cancer-on-a-chip models incorporating multiple cell types and biomimetic extracellular matrices have been developed. These models have generated new research directions in engineering and medicine by allowing for the real-time observation of cancer-host cell interactions in a physiologically relevant microenvironment. However, these cancer-on-a-chip models have yet to overcome limitations including the complexity of device manufacturing, the selection of optimal materials for preclinical drug screening studies, long-term microfluidic cell culture as well as associated challenges, and the technical robustness or difficulty in the use of these microfluidic platforms. In this review, an overview of the tumour microenvironment, its unique characteristics, and the recent advances of cancer-on-a-chip models that recapitulate native features of the tumour microenvironment are presented. The current challenges that cancer-on-a-chip models face and the future directions of research that are expected to be seen are also discussed.
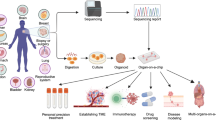
Graphical Abstract

Highlights
• Recent advanced cancer-on-a-chip models for studying the tumour microenvironment are reviewed.
• Models examining interactions between tumours and multiple components of the tumour microenvironment are discussed.
• Advanced cancer-on-a-chip models can provide a controlled environment for studying complex aspects of cancer biology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cancer is an adaptive and dynamic disease mainly driven by the genetic mutation(s) of cells, resulting in uncontrolled cell proliferation and eventual invasion into other organs [1,2,3]. Cancer cells can undergo several mutations over the course of their lifetime, thereby making them genetically unstable [3, 4]. As a result, this genomic instability promotes diversity within the tumour and ultimately fosters the development of tumour heterogeneity [4]. Advances in genomic sequencing technologies have revealed that bulk primary or secondary tumours that develop over the course of the disease can contain numerous cancer cell populations with distinct phenotypic and morphological profiles [3]. Tumour heterogeneity creates a significance clinical issue, as small subpopulations of cancer cells can be intrinsically resistant to anti-cancer therapies, or acquire the necessary molecular, metabolic, and/or physical changes that enable them to adapt to treatment [3, 4].
In the past, cancer was viewed as simply a group of tumour cells, and chemotherapies were developed in the second half of the twentieth century to target these cancer cells [4,5,6]. More recently, the view on cancer has shifted. Tumours are now thought to more closely resemble organs with abnormal structure and function to create the tumour microenvironment (TME) [7, 8]. The TME and the structures surrounding it consist of both cellular and acellular components [7]. Tumours can interact, recruit, and transform the anti-cancer functions of various cells close to their surroundings to promote tumour growth [9]. Such cells include fibroblasts [10], macrophages [11], and endothelial cells [12, 13]. Moreover, primary tumours have been shown to secrete various factors including extracellular vesicles (EVs) [14, 15] carrying various cargoes including proteins, lipids, growth factors, and microRNAs (miRNAs) [16] to form the pre-metastatic niche at distant sites to create a permissive soil for the metastasis and colonization of the tumour [17]. The TME plays an active role in the survival, growth and metastasis of cancer [7, 8]. It also plays a key role in modulating the progression of tumour growth and resistance to chemotherapies [7, 8].
The adaptive response of the TME in addition to the global healthcare burden of cancer highlights the need for the discovery and development of novel cancer therapies. Currently, the resistance of cancers to chemotherapies remains one of the most challenging issues facing cancer researchers to date [18, 19]. While many promising drug candidates have been identified for cancer treatments over the last 50 years, the success rate of anticancer therapies is very low due to poor efficacy or unpredicted adverse side effects [20]. This can be attributed to the use of conventional preclinical drug screening models using in vitro cancer models (two-dimensional (2D) monolayers or three-dimensional (3D) culture models or in vivo animal testing [20, 21]. While cost-efficient and simple to perform or operate, these methods fail to recapitulate the unique physicochemical profile (for example, oxygen [22] or nutrient [23] gradients) and the cellular interactions that occur within the TME [24]. Therefore, the research and development of an optimal in vitro tumour model that can accurately mimic the TME is of great interest in the field of oncology. Such a platform would allow for the study of cancer biology and for the precise screening of novel anti-cancer drug candidates [20, 21, 25, 26].
The development of microfluidic devices has provided advantages for biological studies and led to the generation of new research directions in engineering and medicine [25, 26]. Microfluidics is well suited for the control or manipulation of cells and biomolecules such as nucleic acids [27] proteins [28] and antibodies or antigens [29,30,31]. The flexibility in device design allows for the customization of microfluidic device architecture and/or patterning for the requirements of individual cell types or co-cultures and for monitoring cellular response to various chemical, mechanical, or biological stimuli [31]. Microfluidic systems are able to more closely recreate the physiological environment through, for example, the continuous perfusion of medium [32], the creation of biochemical gradients [33] or through recreating flow regimes [34] which naturally occur in vivo [30, 35].
Biological microfluidic systems referred to as “organ-on-a-chip” models aim to mimic the in vivo physiological environment of organs and tissues [35]. Models of human organs-on-a-chip for the heart [36], lung [37], liver [38], kidney [39] and brain [40] (to name a few) have been reported [21, 35, 41]. The principal purpose of organ-on-a-chip systems is to improve the quality of in vitro testing conditions and results by more accurately recapitulating in vivo conditions for biomedical research [31, 35]. Organ-on-a-chip models contain three defining characteristics: the use of multiple cell types, the 3D arrangement of cell and tissues in the microsystem, and the addition of biomechanical features such as fluid flow shear stress and mechanical strain to construct more physiologically relevant models [35]. These tools allow for a better understanding of biological processes or phenomena, such as mechanisms of the onset and progression of disease and for reduced animal testing, a practice whose ethics and morals can, at times, be controversial [31].
Cancer-on-a-chip models aim to replicate several hallmarks of cancer and the TME by allowing for the study of dynamic interactions between cancer cells and the 3D microenvironment [21]. These studies, performed on a biomimetic platform, have the potential to further elucidate biological mechanisms and processes for cancer research. Beneficial features of using cancer-on-a-chip models compared to conventional in vitro models are shown in Table 1. Several cancer-on-a-chip models have been developed over the last 15 years [21, 25]. Various cancer models have been designed and tested to study intravasation [42] extravasation [43] and angiogenesis [44]. Orthotopic models of breast and lung cancer have also been developed [21]. Patient-derived tumour cells have been integrated into these platforms to closely mimic native conditions for personalized medicine and the assessment of individualized responses towards cancer therapies [21, 25, 26].
In this review, recent advances from the last five years in 3D cancer-on-a-chip models incorporating multiple cell types are examined. Different from existing reviews on cancer-on-a-chip models that focused on design aspects of cancer-on-a-chip devices [20], the use of microfluidic devices to model cellular responses of cancer cells [21], the extravasation of immune and cancer cells on microfluidic devices [45], the migration of cancer cells in microfluidic 3D environments [46], the comparison between 2D cultures or in vivo murine models versus advanced microfluidic models [47,48,49], or general discussions of models [26, 50], this review uniquely focuses on cancer tissue models that are able to simulate multiple organ and system interactions within the TME, the biological interactions between cancer cells and various critical components of the TME, including endothelial cells, stromal cells and immune cells, and emphasizes the novel fundamental biological discoveries made using these advanced tissue models.
With the recent development of cancer immunotherapies as well as the emphasis of the role of the immune system in the development, progression and metastasis of cancer, this review discusses the involvement of immune cells in these microfluidic models. A brief background of the TME is presented first to provide information on hallmark characteristics of the TME that should be incorporated into 3D cancer tissue models. Recent advances of microfluidic cancer models that closely examine interactions with nearby cells within the TME that contribute to the growth and progression of the primary tumour, namely, tumour-blood vessel, tumour-stroma, tumour-immune cell interactions are discussed. Then, recently developed microfluidic tissue models that examine cancer cell behaviours and interactions with secondary tumour sites, namely the formation of the pre-metastatic niche and metastasis are examined. The fundamental biological discoveries made from using these platforms are also highlighted. Finally, current challenges and the future directions of research in 3D cancer-on-a-chip models are discussed.
Tumour microenvironment
Formation and hallmark characteristics
Normal tissue architecture and homeostasis restrain the growth and invasion of tumours [7, 8]. Therefore, the survival of tumours depends on the conversion of the physiological microenvironment into a pro-tumorigenic state [41, 51,52,53,54,55]. Both tumours and the TME are highly heterogenous with complex genetic profiles [7]. Tumours recruit a number of other cells such as fibroblasts and immune cells to promote growth [54]. These recruited cells can then transform into a cancer-associated phenotype to further drive progression. The TME is a complex system composed of the solid tumour and the structures surrounding it, including cellular and acellular components [8, 51, 55]. As shown in Fig. 1, components of the TME include cancer-associated fibroblasts (CAFs), inflammatory and immune cells (macrophages, T cells, B cells), adipocytes, mesenchymal stem cells (MSCs), endothelial cells, lymph and blood vessels [55]. Acellular components include the extracellular matrix (ECM) and secreted factors such as cytokines, chemokines, growth factors, metabolites, and EVs [7, 8, 55].
Schematic of the TME with associated cell types such as fibroblasts and immune cells, acellular components such as the ECM and various signaling factors, and established chemical gradients of oxygen, pH and cell metabolites unique to the TME [53]
The TME is a specialized entity. It is dynamic and responsive to external stimuli. Cancer cells in turn respond to various cues provided by the TME, therefore reflecting the dynamic and interdependent interactions present in the local microenvironment [53]. The types of interactions between cancer cells and TME components include direct cell-to-cell interactions through the formation of tight junctions [56], gap junctions or electrical coupling via direct cell–cell contact [57], indirect communications via autocrine and paracrine signalling [58], as well as cell-to-ECM interactions [59], all of which are crucial for tumour development [8, 54].
The TME also forms carefully established mechanical and chemical gradients of matrix stiffness, metabolites, oxygen, pH and nutrients that further drive tumour progression [53, 60]. More specifically, to overcome an acidic and hypoxic microenvironment, the TME is able to promote angiogenesis to restore oxygen and remove accumulated metabolic waste [53, 60]. Recent studies have attested to the existence of vicious feed-forward paracrine loops, where cancer-associated cells secrete growth factors that stimulate tumour cells [61]. In turn, the tumour cells secrete growth factors that stimulate cells in the TME, ensuring persistent growth and migration of cancer cells towards the vasculature [60, 62]. Local reciprocal gradients of growth factors such as transforming growth factor β (TGF-β), vascular endothelial growth factor (VEGF), and fibroblast growth factor (FGF) have been shown to exist, although the molecular mechanisms behind these signalling cascades remain to be elucidated [60, 62]. Tumour cells can also sense, process and respond to mechanical stimuli through complex mechanotransduction signalling cascades [60].
The TME is associated with increased ECM stiffness as cancer cells recruit, transform stromal cells, and subsequently remodel the surrounding ECM [62, 63]. Dynamic mechanical interactions between tumour cells and the tumour stroma (and its components) regulate a wide range of cellular processes critical to tumorigenesis as well as the initiation of metastasis [62]. Properties of the ECM such as stiffness, pore size, cross-linking proteins, density, ECM fiber network configuration and cancer cell stiffness all contribute to such dynamic interactions [62, 64, 65].
The increased stiffness of tumours, particularly in breast and colorectal cancers, is highly correlated with cancer progression and metastasis [60, 62]. The stiffening of the ECM and tumour stroma is mainly attributed to the increased deposition and remodelling of the ECM in the TME [66, 67]. CAFs and tumour-associated macrophages (TAMs) coordinate to modify the ECM within the TME through the excessive deposition of collagen fibers and fibronectin [68], cross-linking glycoproteins and proteoglycans [64, 69], the secretion and regulation of growth factors and ECM-transforming enzymes, [70] and the reconfiguration of the stroma through the realignment of ECM fibers [62].
Cancer metabolism and tumour microenvironment
To promote the growth and survival of tumours, both cancer and surrounding host cells undergo metabolic reprogramming [71]. Recently, this reprogramming has been shown to change the metabolic characteristics of cancer cells during progression from primary tumours and metastatic cancers even within the same patient(s) [71]. Therefore, metabolic reprogramming in the TME promotes malignancy and resistance to anti-cancer therapies [71,72,73,74]. It also creates a challenge to develop therapies that specifically target the metabolic vulnerabilities of cancer cells and/or other cell types in the TME [71, 72, 74].
In the nutrient-deficient and hypoxic TME, oncogenic alterations of cancer cells as well as host cell factors primarily regulate the core metabolic functions of both anabolic and redox homeostasis [71, 73]. Recent studies on oncogenic signalling from cancer cells as well as factors produced by host stromal cells, immune cells and the endocrine system have provided insight into the specific mechanisms that drive metabolic reprogramming and adaptation of cancer cells through cancer progression [71, 75]. There is increasing evidence demonstrating that host cell signals allow for the metabolic adaptation of cancer cells. The activation of the phosphatidylinositol-3 kinase/protein kinase B/mammalian target of rapamycin (Pl3K/AKT/mTOR) signalling pathway in cancer is typically enabled through multiple factors, including direct activation through the mutation of cancer cells, indirect activation through upstream oncogenic mutations, and through extrinsic activation through host-derived signalling factors [71]. All three contributions are required for optimal metabolic activity and adaptation in cancer cells, demonstrating the need for the addition of host cells when studying cancer using in vitro models [71].
Tumor interactions at distant locations: formation of the pre-metastatic niche
Common secondary sites for cancers to metastasize include the lungs, brain, liver, lymph nodes, and bone [76]. Much recent work has shown that these organs are remotely primed for the colonization of circulating tumour cells through the formation of the pre-metastatic niche [15, 77, 78]. Primary tumours secrete soluble factors such as receptor activator of nuclear factor kappa-Β ligand (RANKL) (bone metastasis) [14], tumour necrosis factor alpha (TNF-α), VEGF-A, EVs, enzymes such as lysyl oxidase (LOX), and miRNAs to modulate the functions of host cells at the secondary location, and ultimately change the microenvironment at the secondary site to permit the survival of circulating cancer cells upon their arrival at the secondary site [77, 78]. Changes to the secondary site can include increased vascular permeability [79], remodelling of the ECM [80], modulation of nutrient uptake to provide increased availability of certain nutrients [81], and recruitment of pro-tumorigenic immune cells to support metastatic cancer cell seeding and subsequent colonization [82]. As a result of these changes, typical characteristic markers of the pre-metastatic niche include increased concentrations of fibronectin secreted by perivascular cells and fibroblasts, increased LOX activity linked to ECM remodelling, as well as formation of pre-metastatic lesions in bone [77, 78]. The pre-metastatic niche is also linked to an upregulated expression of molecules such as matrix metalloproteinases (MMPs) -2 and -9, as well as pro-inflammatory S100 calcium-binding proteins A8 and A9 (S100A8, S100A9), all of which are considered as markers of poor prognosis in cancer patients [78].
In summary, the constantly evolving environment and heterogeneity of the TME both contribute to its adaptive response and subsequent resistance to chemotherapies. Many novel cancer treatments now target the TME, or more specifically aim to inhibit specific interactions (direct or indirect) occurring in the TME which are crucial for tumour progression and metastasis [51]. However, it is thought that the role or function of TME components from early tumour development to metastasis changes over the course of tumour growth, making the development of novel cancer therapies rather difficult [51].
Cancer-on-a-chip models
Cancer-on-a-chip models for studying tumour-blood vessel interactions
Tumours require oxygen and nutrients from blood vessels to survive, proliferate and eventually metastasize [83]. The vascularization of solid tumours occurs through several distinct biological processes [83]. However, in rapidly growing tumours, the rate of tumour growth is often greater than the supply provided by the tumour vasculature, resulting in intratumoral hypoxia [53, 83]. Tumour vasculature differs greatly from normal vasculature in that these blood vessels exhibit abnormal structural dynamics and are highly permeable, immature, and dilated, resulting in “leaky” blood vessels [83]. Briefly, hypoxic conditions in the TME activate the angiogenic master switch, hypoxia inducible factor 1 (HIF-1), which in turn upregulates VEGF as shown in Fig. 2A [60]. VEGF then promotes tumour angiogenesis to deliver nutrients and oxygen to the tumour [60].
A Angiogenic signalling from tumour cells. B Vascularized lung cancer-on-a-chip (VLCC). Lung tumour spheroids were cultured in the decellularized extracellular matrix to mimic the native TME. The two cylindrical channels were seeded with HUVECs to form the perfusable vessel structure [84]. C Perfusable tumour mimetic chip with high and low physiological perfusion capabilities [85] D (Top left) Co-culture configuration of PDAC cells with biomimetic blood vessels and ablation of endothelial cells in the biomimetic pancreatic cancer model by Nguyen et al. [86], (Top right) Apoptosis of endothelial cells (red) indicated by cleaved caspoase-3 staining (white) due to ablation by PD7591 pancreatic cancer cells (green) (Bottom left) Close-up view of endothelial cells (Bottom right) Close-up of PD7591 cells [86]
The incorporation of a vascular network is critical for the formation of biomimetic in vitro cancer-on-a-chip systems [87,88,89]. 3D tissue models as in vitro drug screening tools for novel anti-angiogenic therapies can help to improve the understanding of mechanism of action and efficacy of these treatments [89]. Figure 2B shows a 3D microfluidic tissue model by Park et al. with perfusable vessels to perform drug screening. The rate at which perfusion of media containing anti-cancer therapies also plays an important role in preclinical drug screening models. Figure 2C shows a microfluidic platform developed by Pradhan et al. that permits the variation of perfusion rates. Microfluidic tissue models also allow for the observation of key interactions between cancer cells and endothelial cells. Figure 2D shows how pancreatic cancer cells can replace cells lining the endothelial lumen during the process of endothelial ablation. In this section, the most recently developed 3D tumour-vasculature models with perfusable blood vessels allowing for delivery of cancer therapies are examined with a particular emphasis on cancer-endothelial cell interactions.
Lung cancer
Park et al. developed a 3D vascularized lung-cancer-on-a-chip (VLCC) model, as shown in Fig. 2B. They used a lung decellularized ECM which resulted in a pro-angiogenic tissue mimetic hybrid mimicking the natural lung environment [84]. Tumour spheroids were formed using A549 lung cancer cells, human umbilical vein endothelial cells (HUVECs), and human lung fibroblasts (HLFs) to mimic a native solid tumour and induce angiogenesis [84]. The system also had perfusable large vessel-like structures to simulate arteries, veins and capillaries [84]. Two macrochannels were seeded with HUVECs inside the matrix to form the perfusable blood vessel system [84]. The authors then tested high and low concentrations of doxorubicin (DOX) and quantified caspase-3 and tumour protein p53 expression to determine apoptosis levels [84]. Drug screening revealed that the VLCC could demonstrate dose-dependent effects of DOX, whereas 2D and 3D spheroid cultures did not [84]. Compared to conventional in vitro hydrogels such as collagen, the use of the lung decellularized ECM provides the system with the mechanical properties and complex protein and cytokine composition of that found in native lung tissue. Moreover, the formation of perfusable vessels and use of multicellular spheroids contributed to the strength of this model, as demonstrated by the drug screening test [84].
Breast cancer
Pradhan et al. developed a microvascularized tumour-on-chip platform to examine the efficacy of anti-cancer therapies [85]. Figure 2C shows the platform mimicked the pathophysiological environment of in vivo breast tumours and facilitated the formation of mature, perfusable endothelium under physiological fluid flow conditions using human breast tumour endothelial cells (hbTECs) [85]. The platform also introduced areas of high and low perfusion using two different chip designs to simulate the native TME, in combination with a polyethylene glycol (PEG)-fibrinogen (PF) hydrogel to serve as an ECM. PEG, a hydrophilic and biocompatible polymer, can provide a mechanically tunable hydrogel system by changing the ratio of PEG to fibrinogen prior to cross-linking. Fibrinogen is a protein secreted and deposited into the surrounding ECM by cancer cells, and has been demonstrated to promote the secretion of pro-angiogenic factors and transendothelial migration, as well as increase cancer cell metastatic potential. This platform also enabled the long-term co-culture of breast cancer cells and fibroblasts. Metastatic MDA-MB-231 or non-metastatic MCF-7 breast cancer cells were co-cultured with BJ-5ta human foreskin fibroblasts in a 5:1 ratio [85].
To induce tumour heterogeneity in the microfluidic system, microvascular pattern-dependent flow variations were introduced to generate zonal variations and a concentration gradient within the tumour culture chambers [85]. The efficacy of anti-cancer drugs by perfusing DOX or paclitaxel through the microvasculature for delivery to the tumour mass, and cell viability were assessed 48 h after treatment [85]. Both anti-cancer treatments had a greater cytotoxic effect in static 3D cultures of breast cancer cells co-cultured with fibroblasts in the PF hydrogels (in well-plates) compared to the microfluidic tumour-on-a-chip model, highlighting the importance of incorporating a vascular network in tumour models for drug screening [85]. One challenge associated with this microfluidic tissue model was the use of the PF matrix. These hydrogels tend to swell, exerting additional pressure in the tumour chamber and therefore on the cancer cells [85]. Overall, the tumour-on-a-chip platform allowed for the use of an intricate vascular network with “leaky” vessels, with variations in shear flow and the creation of tumour heterogeneity via concentration gradients [85]. It can provide important information on the distribution of drug molecules, the effects of treatments on encapsulated cancer cells within a tumour mass and therefore more predictive data on drug efficacy and mechanisms of actions [85].
Multicellular spheroids have also been incorporated in microfluidic tissue models to provide more spatial complexity. Nashimoto et al. reported a tumour-on-a-chip platform using tri-culture spheroids composed of MCF-7 breast cancer cells, HLFs and HUVECs with an engineered tumour vascular network to evaluate tumour behaviour under intraluminal flow, which can provide oxygen, nutrients or cancer therapies to the tumour spheroid [90]. Co-culture spheroids incorporating HLFs and MCF-7 s and tri-culture spheroids including HUVECs could induce abundant angiogenic sprouting from HUVECs cultured in surrounding channels [90]. Specifically, the inclusion of fibroblasts in breast cancer spheroids induced angiogenesis, leading to the construction of a perfusable vascular network in the microfluidic chip [90]. Interestingly, when treated with paclitaxel, spheroids in perfusion conditions did not decrease spheroid volume compared to spheroid in static conditions. The authors also observed that spheroid volume decreased in a dose-dependent manner in static conditions [90]. This complex vascular network demonstrated the importance of perfusion on tumour activities, as well as drug administration for preclinical screening tests and highlighted the careful considerations that must be made regarding the continuous supply of nutrients and oxygen for tumour-on-a-chip platforms during drug screening tests [90]. Kwak and Lee developed an optimization protocol for their previously developed tumour on a chip model by using tri-culture spheroids consisting of MDA-MB-231 breast cancer cells, MSCs and either HLFs or HUVECs [91]. To support vascular sprouting in the system, HLFs were embedded in the ECM to promote vascular sprouting and invasion, as HLFs secrete a number of pro-angiogenic factors including VEGF and FGF-2 [91]. The authors observed that multicellular tumour spheroids with HUVECs and MSCs embedded in an ECM containing HLFs significantly enhanced the sprouting and migration of tumour spheroids and promoted angiogenesis and vascular invasion, while preserving the structural integrity and functionality of HUVEC-lined microfluidic channels [91]. Such an optimization protocol enabled the generation and validation of microfluidic models of the TME for drug screening of anti-angiogenic therapies, or for testing patient samples for personalized medicine.
Pancreatic cancer
Nguyen et al. introduced a pancreatic-cancer-on-chip model to understand the interactions between pancreatic ductal adenocarcinoma (PDAC) and blood vessels [86]. PDAC enters circulation at the early stages of tumour development, and develops abnormal vasculature at later stages in the disease, thereby limiting the delivery of chemotherapies to the tumour [92, 93]. Therefore, a model of a biomimetic PDAC ductal channel co-cultured beside an endothelialized and perfused lumen channel to observe PDAC-endothelial interactions was created [86]. The hollow cylindrical PDAC and blood vessel channels were embedded in a collagen matrix [86]. Primary PD7591 cells were seeded into the biomimetic ductal channel, HUVECs were seeded in the blood vessel channel to form a perfusable lumen [86], with the co-culture configuration shown in the schematic in Fig. 4D (top left). After seeding the endothelium successfully deposited a collagen IV layer in the device [86].
The authors observed that part of the endothelium became occupied by the tumour cells, with increased number of apoptotic HUVECs in proximity to the PDAC cells. The collagen IV layer gradually disappeared as the tumour cells replaced parts of the endothelium, a phenomena termed as endothelial ablation as demonstrated on the device in Fig. 2D [86]. Endothelial ablation was also observed in in vivo experiments with mice. Using the pancreatic cancer-on-chip device, Nguyen et al. identified that the TGF-β receptor signalling pathway mediated endothelial ablation, that is, inhibition of TGF-β receptor signalling significantly reduced endothelial ablation of the microfluidic blood vessel [86]. Knockout of activin receptor-like kinase 7 (AlK7) receptors, required for endothelial ablation in HUVECs, resulted in significantly decreased endothelial ablation and PDAC invasion in 2D patterned co-cultures and in vivo studies using mice [86]. Using such a 3D tissue model allowed for the study of cancer-endothelial cell interactions and observed endothelial ablation as well as the signalling pathways involved. This model also permitted the use of spatiotemporal and genetic control to determine the signalling pathway for endothelial ablation [86].
Glioma
Truong et al. developed a 3D organotypic model to study glioma stem cell-vascular interactions [94]. This 3D microfluidic tissue model was integrated with hydrogels to create a biomimetic vascular niche and study the influence of endothelial cells on the behavior of glioma stem cells (GSCs) [94]. Co-culture with endothelial cells enhanced GSC migration, an invasive GSC morphology, and increased expression of neural epithelial stem cell protein Nestin (neural progenitor), SRY-box transcription factor 2 (SOX2) (pluripotency and self-renewal) and cluster of differentiation 44 (CD44) (hyaluronic acid in malignant glioblastoma) markers, suggesting that the platform allowed for GSCs to maintain their stemness [94]. Comparison with an in vivo mouse model revealed similar invasive GSC morphology, indicating that the microfluidic system is comparable with the native TME [94]. C-X-C motif ligand chemokine 12-C-X-C motif receptor chemokine 4 (CXCL12-CXCR4) signalling was confirmed to be involved in the migration of GSCs when co-cultured with endothelial cells in a 3D microenvironment [94]. Treatment with AMD3100, a CXCR4 inhibitor, significantly decreased invasion distance. Overall, the microfluidic model allowed for studying GSC interaction with the surrounding microenvironment and for the development of future cancer therapies that target key molecular signalling pathways key to GSC growth, invasion, or regulatory mechanisms [94].
Colorectal cancer
Hachey et al. presented a novel 3D vascularized micro-tumour (VMT) to recapitulate the colorectal cancer microenvironment [95]. Their tumour-on-a-chip model was transparent, allowing for real-time monitoring, and contained perfusable blood vessels and fibroblasts for drug screening studies. Tumour growth and response to chemotherapies using this new model more closely resembled the responses observed in preclinical in vivo xenograft models. RNA sequencing studies also confirmed that the VMT platform indeed captured tumour heterogeneity [95]. Tumours derived from the VMT model contained a distinct neural/glial antigen 2-positive (NG2 +) population that underwent the epithelial to mesenchymal transition (EMT), whereas cancer cells in 2D and 3D monocultures did not [95]. Additionally, by culturing tumours in a 3D stroma with fibroblasts, the authors observed a significant upregulation of TGF- β signalling. Treatment with TGF-β inhibitor galunisertib suppressed tumour growth whereas this effect was not observed in 2D or 3D cultures through impairing stromal fibroblast signalling [95]. The VMT platform was able to better capture the heterogeneity and drug response of in vivo tumours compared to standard clinical drug screening platforms, which included 2D monolayer culture and 3D tumour spheroids [95].
Cancer-on-a-chip models for studying tumour-stroma interactions
The tumour stroma is the non-neoplastic component of the TME, consisting of multiple types of support cells and an abundance of ECM and acellular factors (cytokines, chemokines, EVs, etc.) (Fig. 3A) [96]. The complex interactions that occur within the tumour stroma are a hallmark feature of solid tumours that promote tumour growth, progression, and metastasis as well as an adaptive response to chemotherapies that ultimately result in the resistance to anti-cancer drugs [96, 97].
A Tumour-stroma microenvironment including NFs, and CAFs. CAFs tend to exhibit elevated levels of ECM production and remodelling, and acellular factors. B Novel gene of interest GPNMB discovered by Truong et al. using their microfluidic tumour invasion model of breast cancer [98]. (i) Immunofluorescence analysis of GPNMB. Knockdown of GPNMB in SUM-159 breast cancer cell line shG #1 and #4 did not increase GPNMB expression when co-cultured with CAFs, compared to control [98]. GPNMB knockdown attenuated (ii.) migration speed and (iii.) persistence [98]. C Co-culture of breast cancer cells with varying fibroblast and matrix composition [99]. (i.) Co-culture with CAFs enhanced cancer cell migration distance compared with normal fibroblasts [99]. (ii.) Second harmonic generated images of collagen/fibronectin matrices [99]. (iii.) Measured matrix gap area in collagen and collagen-fibronectin matrices when cultured with NFs or CAFs [99]
Tumour-stroma structures in in vitro platforms can provide the necessary cellular and matrix complexity of the native tissue/cancer microenvironment. However, the accurate compartmentalization of tumour-stroma is critical and must allow for cellular interactions and migration between compartments [97]. When accurately modelled, cancer-on-a-chip platforms can aid in identifying important genes or markers of interest in cancer-stroma interactions that drive cancer invasion or metastasis. For example, Truong et al. identified glycoprotein non-metastatic B (GPNMB) as a key marker for the migration speed of breast cancer cells during co-culture with CAFs, as demonstrated in Fig. 3B [98]. Stromal components include CAFs, immune cells, endothelial cells, ECM and chemokines, growth factors, EVs and other soluble factors [96]. Modelling cancer microenvironments with the appropriate stromal components is also crucial for accurately measuring response during preclinical drug screenings, as in Fig. 2C which shows that incorporation of CAFs and fibronectin enhance cancer cell migration compared to normal fibroblasts. Some cancer types are stroma rich. In fact, the stroma can consist of the majority of the tumour volume, such as PDAC where the tumour consists of around 90% stroma by volume [97]. Here, recent 3D microfluidic tissue models highlighting tumour-stroma interactions are examined, with a particular focus on tumour-fibroblast (normal or cancer-associated) interactions.
Breast cancer
Co-culture with stromal cells is crucial for the study of in vivo-like cancer cell behaviours, secretion of cancer-derived factors, and expression of genes responsible for cancer cell invasion and migration in a controlled in vitro environment. Truong et al. developed a microfluidic tumour invasion model incorporated with either patient-derived CAFs or normal fibroblasts (NFs) to study tumour-stroma interactions [98]. Invasive SUM-159 breast cancer cells and patient-derived fibroblasts were co-cultured in their respective 3D tumour and stromal areas on the microfluidic chip to assess the molecular and cellular influence of patient-derived fibroblasts during tumour-stroma crosstalk [98]. The authors integrated transcriptome profiling as well as a conventional cancer cell migration analysis to examine how tumour-stroma interactions promote breast cancer invasion [98]. Transcriptional profiling during cancer invasion was used to reveal molecular changes that occurred during invasion. More specifically, co-culture with NFs exhibited a tumour-suppressive behaviour through a reduction in migration, proliferation of breast cancer cells and an enrichment in inflammatory pathways, whereas CAFs promoted a pro-tumorigenic environment through enhanced invasion, cell adhesion and ECM [98]. The authors also identified a gene of interest called GPNMB and performed a functional assay to determine the specific role of GPNMB. The authors established GPNMB knockdown lines (Fig. 3B (i.)) and determined that GPNMB knockdown was responsible for reduced breast cancer cell invasion through a 3D matrix, migration speed, and persistence (Fig. 3B (ii., iii.)). Overall, the platform was useful in uncovering key mechanistic pathways involving GPNMB expressed in CAFs, by studying tumour-stromal cell interactions on a biomimetic platform for the discovery of novel cancer therapies that target the TME [98].
Ayuso et al. developed a microfluidic tissue model to mimic the ductal carcinoma in situ (DCIS) microenvironment, which requires the generation of a luminal structures containing MCF10A-DCIS cancer cells with different metabolic phenotypes in addition to the presence of stromal cells [100]. Their platform enabled the co-culture of DCIS cancer cells in an epithelial mammary duct comprised of MCF-10A cells along with human mammary fibroblasts (HMFs) embedded in a collagen-based ECM [100]. The authors demonstrated that DCIS cells indeed rely on glycolysis to support cancer growth, as DCIS cells quickly consume glucose and glutamine compared to their normal epithelial counterparts (MCF-10A cells) [100]. Moreover, during cancer cell invasion over 3 days, optical metabolic imaging of the microfluidic platforms revealed a spatial dependence of DCIS cells in the device [100]. More specifically, cells invading the ECM had a decreased nicotinamide adenine dinucleotide phosphate (NADPH) fluorescent lifetime, suggesting that these cells underwent more glycolysis compared to the cancer cells remaining in the lumen, demonstrating spatial heterogeneity in cancer cell metabolic phenotype.
Several other microfluidic platforms were also designed for studying tumour-stroma interactions. They have been used to study cancer-secreted factors that promote invasion or upregulate mesenchymal markers such as MMPs or fibronectin. Fan et al. designed a 3D microfluidic system with microchamber arrays embedded in collagen with tunable biochemical gradients [101]. The microchamber provided a similar environment to that of the native TME, with the microfluidic channels offering precise control of complex concentration gradients of biomolecules or anti-cancer therapies. MCF-10A breast epithelial cells formed lumen-like structures similar to those found in epithelial layers in the microfluidic device [101]. However, when MCF-10A cells were co-cultured with invasive MDA-MB-231 breast cancer cells, the formation of lumen structures was inhibited, demonstrating how cancer cells can disrupt the formation of normal structures due to the secretion of MMPs [101].
Lugo-Cintrón et al. developed a microfluidic model to mimic the complex TME during breast cancer invasion [99]. MDA-MB-231 breast cancer cells were seeded into a lumen structure within a 3D collagen matrix to determine the effects of various fibronectin and fibroblast compositions on cancer cell migration distance and matrix remodelling [99]. Normal HMFs or CAFs were embedded into the 3D matrix to observe cancer-stroma crosstalk. The authors observed that a fibronectin-rich matrix containing CAFs increases cancer cell migration distance and promotes higher MMP secretion and ECM remodelling (Fig. 3C (i.)) [99]. Furthermore, a fibronectin-rich matrix dramatically increased ECM remodelling (matrix gap area) in NFs, whereas no change in remodelling occurred with the addition of fibronectin in CAF co-cultures (Fig. 3C(ii., iii.)). This may be attributed to the fact that CAFs already secrete enhanced quantities of fibronectin compared to NFs, and addition of fibronectin does not alter CAF behaviours [99]. Interestingly, MMP inhibitors for breast cancers were only effective in monoculture conditions and not in a fibronectin-rich matrix containing NFs, suggesting that the lack of fibroblast incorporation in in vitro studies could explain the limited success of MMP inhibitors in clinical trials [99, 102, 103]. Taken together, this study was the first to use a microfluidic tissue model to screen the effects of key TME components on cancer cell migration and on the efficacy of recently developed cancer treatments [99].
Incorporation of the stroma is also important for studying the expression of pro-inflammatory cytokines in the TME. Du et al. presented a microfluidic platform to control multiple factors of the in vitro TME and assess tumour invasion into the 3D stroma containing noncancerous MCF-10A breast epithelial cells and neonatal human dermal fibroblasts (HDF-n) [104]. The side-by-side arrangement of the tumour and stromal compartments allowed for paracrine and juxtracrine signalling between cell types, and a 3D endothelial layer between the collagen gel and nutrient reservoir allowed for the simulation of the trans-endothelial transport of nutrients [104]. The real-time visualization of MDA-MB-231 breast cancer cells also revealed that the ability of tumour invasion was directly correlated to cell density [104]. Additionally, migration distance of invading cancer cells was enhanced through co-culture with noncancerous cells. Analysis of interleukin-6 (IL-6), a proinflammatory cytokine in MCF-10A and MDA-MB-231 mono- and co-cultures revealed that IL-6 expression was significantly enhanced in both cell types during co-culture [104]. Inhibition of IL-6 decreased the invasion ability of breast cancer cells, demonstrating the importance of inflammatory pathways on the promotion of cancer invasion.
Pancreatic cancer
Lee et al. developed an in vitro 3D pancreatic tumour model to recapitulate the PDAC TME [105]. In PDAC, EMT in the pancreatic TME facilitates metastasis and has been thought to be a mechanism of drug resistance [106]. PANC-1 pancreatic cancer spheroids were co-cultured with pancreatic stellate cells (PSCs), known to be a source of CAFs, in collagen-supported microchannels [105].
Tumour spheroids co-cultured with PSCs resulted in a higher number and larger diameter of PANC-1 spheroids after five days of culture [105]. Moreover, the tumour spheroids acquired a migratory phenotype and enhanced cell migration in co-culture conditions [105]. Co-culture with the stroma also increased expression of EMT markers in tumour spheroids. More specifically, there was increased expression of EMT marker vimentin as well as EMT cytokines TGF-β and connective tissue growth factor (CTGF) [105]. Drug screening also revealed that co-culture conditions with PSCs promote resistance to chemotherapies, likely attributed to EMT of the tumour spheroids induced by stromal cells and activation of PSCs through co-culture induced upregulation of alpha smooth muscle actin (α-SMA). [105]. This platform allowed for the formation of stable pancreatic cancer aggregates, three dimensionality in collagen-gel stroma and heterotypic cell interactions with PSCs, an important component in the PDAC microenvironment.
Lung cancer
Kim et al. developed a microfluidic platform that recapitulated the unidirectional interstitial flow created by mesothelioma tumours, which is then delivered to fibroblasts located in the surrounding stroma [107]. Briefly, unidirectional interstitial flow containing tumour-secreted cytokines is generated by the tumour due to elevated hydrostatic and osmotic pressure and transported to the surrounding stroma, thereby activating neighboring fibroblasts and other host cells [107]. By receiving the interstitial flow generated by the bulk tumour containing proteins and EVs, surrounding host cells are transformed into cancer-associated phenotypes. Several co-culture methods typically utilize bi-directional signalling from cancer and host cells, as the intricate unidirectional signalling created by interstitial flow is difficult to simulate [107]. In this study, interstitial flow in the microfluidic device was generated by the evaporation of medium from reservoirs with different diameters and carried secreted factors from the tumour cells to the fibroblasts [107]. These factors stimulated the activation, migration, and stellate formation of the receiving fibroblasts. The activated fibroblasts expressed abundant MMPs 9 and 14 as well as CAF markers including vimentin, α-SMA, and fibroblast activation protein alpha (FAP-α) [107]. Overall, this platform captures and highlights the importance of recapitulating biomechanical aspects of the TME.
Melanoma
Ayuso et al. developed a microfluidic platform to study primary melanoma cell phenotype and metabolism during co-culture with fibroblasts and/or keratinocytes (stromal cells) by leveraging air-wall patterning to control the spatial distribution of each cell type [108]. The authors observed that co-culture with stromal cells induced an elongated melanoma cell morphology compared to monoculture conditions in the device, as well as elevated secretion of IL-6 and IL-8, both of which contribute to tumour proliferation and metastatic potential [108]. Interestingly, by utilizing optical metabolic imaging, the authors observed that triple culture conditions led to a marked decrease in melanoma redox ratio, thereby shifting the metabolic profile of cancer cells when compared to monoculture conditions [108]. Moreover, analysis of NADPH fluorescent lifetime revealed that co-culture with stromal cells provides a more supportive environment for melanoma cells [108]. A reduced NADPH fluorescent lifetime implies there is a higher degree of NADPH in the cytoplasm, thus reducing the degree of oxidative stress in melanoma cells. Taken together, this platform demonstrates that fibroblasts and keratinocytes play a supportive role in melanoma tumorigenesis [108].
Cancer-on-a-chip models for studying tumour-immune cell interactions
Immune cells play a key role in the TME. The TME forms an immunosuppressive microenvironment due to nutrient depletion and accumulation of waste and hypoxia, which can compromise the function of immune cells [109]. Tumours interact with a diverse number of adaptive or innate immune cells (Fig. 4A). Briefly, adaptive immune cells are cells activated upon the exposure to a specific antigen, and subsequently use immunological memory to evaluate the potential threat of the foreign particle [110]. Adaptive immune cells include T cells and B cells [110]. Innate immune cells utilize a non-specific defense mechanism upon the entry of a foreign antigen into the body [110]. These cells include macrophages, neutrophils, natural killer (NK) cells, and dendritic cells [110]. Immune cells can either function as a tumour suppressor or can promote tumorigenesis. The role that immune cells play in the TME depends on the tumour type and context [108]. For example, macrophages that infiltrate the TME can undergo polarization and become TAMs [111].
A Tumour-immune microenvironment. B NK cell exhaustion in the TME. [112] (i) Spatial cell viability gradient from the proximal (outer proliferative zone) to the distal (necrotic core) zones of the solid tumour [112]. (ii) Upregulation of genes associated with exhaustion and downregulation of genes associated with proliferation and survival in NK cells isolated from the tumour chip [112]. C Microvascularized model for studying tumour extravasation [113] (i) Schematic of the microfluidic device for monocyte studies with the experimental timeline. Quantified increase in monocyte extravasation over 48 h after perfusion [113]. (ii.) Monocytes (white) undergoing extravasation through the endothelium (green), and subsequent interactions with fibroblasts (red) [113]
Several platforms have been recently developed to study the interactions between tumour and various types of immune cells and model the biological processes that occur, such as immune cell recruitment and migration [108]. Modelling these dynamic tumour-immune cell interactions can provide further understanding of the underlying mechanisms that lead to the formation of either an anti- or pro- tumorigenic microenvironment or to the cancer-induced formation of the immune-suppressive microenvironment [108, 114]. For example, 3D tumour chip models that mimic spatial aspects of tumour architecture show profound effects on immune cell exhaustion, indicative of an immunosuppressive microenvironment, as demonstrated in Fig. 4B i and ii. These models can also provide an opportunity for observing biological phenomena at high spatiotemporal resolution. The first high resolution imaging of monocyte transmigration performed by Boussomier-Calleja et al. is shown in Fig. 4C. They also investigated the effects of monocyte differentiation on tumour cell migration and extravasation. Such information can help in the development of novel cancer immunotherapies that target various immune cell types across multiple stages of immune cell differentiation [108, 114].
Breast cancer
The pro-inflammatory, immunosuppressive microenvironment is a hallmark of the TME. However, more insight on the effects of the TME on host immune cells is needed. Ayuso et al. developed a 3D in vitro tumour-on-a-chip platform to examine how NK cells respond to an immunosuppressive microenvironment [112]. The tumour-on-a-chip device consisted of a central microchamber where MCF-7 breast cancer cells were embedded in collagen with or without NK cells [112]. The asymmetric distribution of nutrients can generate multiple tumour scenarios and created gradients of cell viability (Fig. 4B(i.)). As a result of the heterogenous distribution of nutrient and oxygen, immune cells isolated from the chip expressed genes associated with immune exhaustion (indoleamine 2,3-dioxygenase (IDO1), programmed cell death protein 1 (PD-1) and cytotoxic T-lymphocyte-associated protein 4 (CTLA-4)), as well as nutrient starvation and hypoxia (HIF-1A, VEGF) were upregulated (Fig. 4B(i.)) [112]. Genes associated with cell survival, proliferation and activation (B-cell lymphoma 2 (BCL2), granzyme B (GZMB), Fas cell surface death ligand (FASL), IL-15 were significantly downregulated, suggesting an immunosuppressive tumour-on-a-chip microenvironment (Fig. 4B(ii.)) [112]. On the other hand, the NK cells isolated from the well-plate had a gene expression profile similar to naïve NK cells (NK cells that have never been exposed to cancer cells), compared to the profile NK cells from the microdevice [112]. More specifically, NK cells cultured in well plates exhibited a significant upregulation of activation/prosurvival markers, whereas NK cells from the microdevice exhibited an upregulation of genes associated with NK cell exhaustion [112]. Treatment with IDO1 and PD-1 inhibitors led to partial alleviation of immune exhaustion and an increase in MCF-7 cell death [112]. This platform allowed for the study of molecular alteration that drives immunosuppression and to identify new immunotherapeutic targets. Additionally, it highlights the importance of analyzing molecular changes of immune cells in a physiologically relevant platform, as well as the spatial dependence on cancer cell response to immunotherapies, as tumour cells in the necrotic region were observed to be highly resistant to treatment. Aung et al. developed a perfusable tumour-on-a-chip model to study the effects of breast cancer-immune interactions on T-cell recruitment [115]. Breast cancer cells, monocytes and endothelial cells were spatially controlled in a gelatin matrix using 3D micropatterning [115]. The effects of cancer-monocyte interactions on T-cell recruitment were examined by dispersing T-cells into the perfused culture medium and allowed to infiltrate through the endothelial barrier [115]. Microfluidic culture simulating the hypoxic environment commonly found in solid tumours/spheroids resulted in increased T-cell recruitment, compared to dispersed cancer cells [115]. Addition of monocytes into the system also resulted in higher T-cell recruitment, attributed to differences in chemokine secretion profiles that influence endothelial permeability [115]. Overall, this platform allowed for the study of immune cell recruitment using multiple cell types.
Host immune cells also have important effects on tumour development and progression. Mi et al. introduced a novel microfluidic platform to study the effects of macrophages on the invasion of breast cancer cells [116]. Breast cancer cells and either normal macrophages or TAMs were embedded in separate 3D matrices arranged in adjacent channels to observe paracrine and juxtracrine signalling [116]. A vascular endothelial layer was located at the interface of the 3D matrices to simulate the trans-endothelial barrier. The authors observed that invasive MDA-MB-231 breast cancer cells co-cultured with TAMs resulted in significantly higher cancer cell migration distance compared to co-culture with normal macrophages [116]. Moreover, the viability of breast cancer cells when treated with paclitaxel was the highest when co-cultured with TAMs, suggesting that transformed immune cells aid in imparting resistance to anti-cancer therapies [116]. This platform can be used for invasion studies in the context of tumour-macrophage crosstalk. Boussommier-Calleja et al. presented a 3D vascularized microfluidic model to study the effects of monocytes on tumour extravasation [113]. The central compartment of the device contains endothelial cells and fibroblasts suspended in a 3D fibrin gel (Fig. 4C (i.)). The studies on this platform provided evidence of direct interactions between tumour cells and monocytes while demonstrating the first high-resolution imaging of monocyte transmigration through human vasculature (Fig. 4C (ii.)) [113]. The platform also replicated monocyte extravasation patterns observed in vivo, where inflammatory monocytes (C–C chemokine receptor-2 (CCR2)-positive) have higher rates of extravasation compared to patrolling monocytes (CCR2-negative) [113]. When treated with blebbistabin, a myosin II inhibitor in monocytes, inflammatory monocytes significantly slowed migration. To determine monocyte effects on tumour extravasation, they used monocytes at various stages of differentiation [113]. Intravascular or undifferentiated monocytes reduced tumour cell extravasation by 42% within 10 h of co-culture [113]. Extravasated monocytes that took on a more macrophage-like phenotype did not have a significant effect on tumour extravasation [113]. Overall, this platform can be used as a tool to further study monocyte-tumour interactions and function as a drug screening model to study novel therapies that target monocyte differentiation.
Ovarian cancer
Ovarian cancer tends to metastasize early in disease progression and has an affinity to travel towards aggregated immune cells on the peritoneum [117]. Immunotherapeutic strategies for ovarian cancer are under development and are currently being tested in numerous clinical trials [118]. Surendran et al. developed a 3D microfluidic platform that served to recapitulate the tumour-immune microenvironment (TIME) [119]. This platform permitted the formation of ovarian cancer spheroids from OVCAR-3 cells embedded in a collagen matrix in hydrogel microwells [119]. These microwells were integrated with a microfluidic channel carrying neutrophils. The formation of neutrophil extracellular traps (NETs) generated a reciprocated response from the tumour spheroids, resulting in the collective invasion of tumour cells into the surrounding collagen matrix [119]. Inhibition of NET formation using Sivelestat reversed the effects induced by NETs. Moreover, NETs were only formed around spheroids in the collagen matrix, rather than NETs formed in contact with spheroids, mediated the collective invasion of tumours, providing further insight into neutrophil-tumour dynamics [119]. Carroll et al. developed a co-culture platform to determine the effects of alternatively activated macrophages (AAMs) on the adhesion of high grade serious ovarian cancer (HGSOC) to a mesothelial-lined surface [120]. The co-culture platform permitted the formation of the peritoneal microenvironment with concentrated paracrine signalling to facilitate the identification of AAM-secreted factors that could enhance cancer cell adhesion [120]. This study demonstrated that HGSOCs primarily adhere to transmembrane protein P-selectin through cluster of differentiation 24 (CD24), and that AAM-secreted macrophage inflammatory protein-1 beta (MIP-1β) activates the C–C chemokine receptor-5 (CCR5)/Pl3K pathway, enhancing P-selectin expression in mesothelial cells. This mechanism was indeed confirmed by analyzing samples from patients with HGSOC [120]. This study provided insight into peritoneal dissemination in HGSOC and new strategies for the development of therapies that target HGSOC metastasis.
Lung cancer
Guo et al. developed a microfluidic platform to simulate the lung cancer microenvironment and examined the role of M2 macrophages on the invasive ability of non-small cell lung cancers (NSCLCs) [121]. This microfluidic tissue model permitted the co-culture of lung cancer cells with macrophages. By combining the use of their platform with proteomic analysis, the authors found that M2 macrophages upregulate the expression of αB-Crystallin (CRYAB), a small heat shock protein that is known to induce EMT and therefore promote NSCLC migration and invasion [121].
Cancer-on-a-chip models for studying pre-metastatic niche interactions
The pre-metastatic niche is the local microenvironment that is primed or educated by the primary tumour to create conditions that permit the survival of tumour cells before metastatic colonization occurs [15,16,17]. Primary tumours secrete factors that can change the structure of the secondary site to enable the colonization of circulating tumour cells (Fig. 5A) [17]. Briefly, circulating tumour-secreted EVs reach the secondary site and prime the local microenvironments by inducing increased endothelial permeability to facilitate cancer cell extravasation, and angiogenesis to provide abundant oxygen and nutrients [15, 17]. Tumour-secreted EVs also activate fibroblasts, increasing their production of fibronectin and MMPs to remodel the surrounding ECM [15]. Microfluidic tissue models may allow for more reliable and physiological modelling of the in vivo pre-metastatic microenvironment and focus on specific interactions between cancer cells and other important cell types in the pre-metastatic niche, particularly immune and endothelial cells. For example, Kim et al. demonstrate that monocyte-endothelial cell interactions increase endothelial permeability upon the arrival of cancer cells to the secondary site, as shown in Fig. 5B [122]. Moreover, Crippa et al. demonstrated that inclusion of neutrophils and platelets on their pre-metastatic niche platform, with the schematic shown in Fig. 5C, enhanced cancer cell migration and EMT [123]. Microfluidic tissue models have recently been used to further reveal that the interactions that occur within the pre-metastatic niche indeed shape cancer cell behaviour during metastasis [122,123,124].
A The local premetastatic microenvironment, associated with increased endothelial permeability, angiogenesis, and enhanced secretion of fibronectin and MMPs to remodel the ECM. B (i.) Schematic of the microfluidic platform by Kim et al., showing the invasion of cancer cells once monocytes have pre-invaded the ECM [122]. (ii.) Downregulation of endothelial tight junction proteins in monocyte co-culture conditions [122]. (iii.) Immunostaining of endothelial layers for tight junction proteins ZO-1 and OCLN [122]. (iv.) Endothelial cell permeability with or without the presence of monocytes in contact and non-conditions [122]. C Early metastatic niche (EMN) chip by Crippa et al. [123] (i.) Schematic of the device channels [123] (ii.) Live cell imaging of the cellular components of the chip, including platelets (white, PAC-1), neutrophils (blue, CD-11b,), vessels (green), and cancer cells (red), trans-endothelial migration with (C + P + N) or without (C) the presence of platelets and neutrophils [123]
Kim et al. developed a 3D microfluidic platform to study the pre-metastatic niche (Fig. 5B(i)) [122]. Monocytes were seeded on the endothelial monolayer to mimic the early stages of pre-metastatic niche formation and subsequently allowed to differentiate in the presence of collagen [122]. Monocytes cultured with collagen had increased expression of macrophage differentiation markers [122]. Monocyte-secreted MMP-9 destroyed endothelial tight junctions to facilitate metastasis (Fig. 5(ii., iii., iv.)), and as a result, endothelial permeability was significantly increased through co-culture with monocytes in a non-contact dependent manner [122]. Pre-invasion of macrophages into the ECM also created a more favourable environment for cancer cell invasion. MDA-MB-231 cells invasion into the microfluidic ECM had significantly higher cell numbers and distance when pre-invaded macrophages were present (compared to an ECM with no macrophages) [122]. Macrophages invading the ECM were also demonstrated to remodel the ECM to facilitate cancer invasion through laminin degradation as well as the realignment and degradation of collagen fibers through the formation of macrophage invadopodia [122]. The studies from this microfluidic platform provided direct evidence on the role of monocytes and macrophages in the formation of the pre-metastatic niche to facilitate cancer metastasis and can be used for studying macrophage/monocyte-cancer crosstalk or for drug screening for macrophage-associated anti-cancer therapies.
Crippa et al. designed an early metastatic niche (EMN)-on-a-chip platform incorporating blood cells to better simulate the in vivo microenvironment (Fig. 5C (i.)) [123]. They showed that inclusion of neutrophils and platelets significantly increased trans-endothelial migration of MDA-MB-231 cells and that the presence of platelets in the EMN-on-a-chip promoted the EMT of breast cancer cells (Fig. 5C (ii.)) [123]. Additionally, their platform was applied to examine the specific action of β3 integrin inhibitor eptifibatide and demonstrated that this drug could have anti-metastatic potential. More specifically, treatment with eptifibatide reduced expression invasion marker MMP-9 in breast cancer cells and reduced permeability in endothelial cells through the proto-oncogene tyrosine-protein kinase/focal adhesion kinase/vascular endothelial cadherin (Src/FAK/VE cadherin) signalling axis [123]. The data acquired from the platform correlated well with previous in vivo studies which have demonstrated that platelets mediate early metastasis and transition to a mesenchymal phenotype by activating the TGF-β/Smad protein signalling pathway in cancer cells [125]. This platform also enabled the investigation of mechanisms within the pre-metastatic niche and could be used for drug screenings of already clinically approved drugs for anti-metastatic applications.
Kim et al. presented a 3D microfluidic human liver-chip to recapitulate the formation of the pre-metastatic niche and study the role of breast cancer-derived EVs before liver metastasis [124]. This model incorporated human liver sinusoidal endothelial cells (LSECs), primary liver fibroblasts and liver hepatocytes and can be used to model the normal liver microenvironment or as a pre-metastatic niche primed by tumour-secreted factors [124]. Importantly, the authors demonstrated that breast cancer-EVs activated liver sinusoidal endothelial cells, promoted the endothelial-to-mesenchymal transition through upregulation of vimentin and zonula occludens-1 (ZO-1), and destroyed endothelial cell barriers. Moreover, TGF-β1 present in the breast cancer-EVs upregulated the secretion of fibronectin in endothelial cells, ultimately facilitating the adhesion of breast cancer cells to the endothelium [124]. Furthermore, the microfluidic platform demonstrated that EVs derived from triple negative breast cancer patients with liver metastasis contained higher levels of TGF-β1 compared to non-metastatic triple negative breast cancer or healthy patients, and that the increased TGF-β1 levels correspond to increased adhesion of breast cancer cells to the endothelium [124]. The model and findings demonstrated that the platform can be used to investigate the mechanisms driving the formation of the pre-metastatic niche in the liver and for screenings anti-cancer treatments targeting the pre-metastatic niche in the liver.
Cancer-on-a-chip models for studying metastasis
Metastasis is primarily responsible for cancer deaths [126]. As the primary tumour develops, it undergoes several genetic mutations, co-opts the TME, and induces angiogenic sprouting once the tumour has a large enough necrotic core [127]. The metastatic cascade is complex but can generally be divided into five major steps [127], including primary cancer cell migration and invasion of basement membrane; intravasation into the surrounding blood or lymph vessels; circulation through the vasculature; extravasation into the secondary site; and colonization at the secondary tumour site (Fig. 6A) [127].
A Schematic of the metastatic progression of tumours demonstrating the five general steps that occur in the metastatic cascade (1) migration and invasion (2) intravasation (3) circulation through the blood stream (4) extravasation from the vasculature (5) colonization at the secondary site [127]. B (Left) Schematic of the microfluidic device by Saha et al., consisting of two PDMS compartments separated by a thin, porous membrane [128] (Right) Cross-sectional view of the device showing the organization of the cellular compartments and ovarian cancer cell migration triggered by platelet extravasation [128]. C (i.) EMT of cancer cells [129]. (ii.) The chip is designed to mimic metastatic cancer cells in lymph vessels that recruit and colonize blood vessels [129]. (iii.) Schematic of the lymph vessel-tissue-blood vessel (LTB) microfluidic device [129]. D Schematic of the microfluidic platform developed by Mei et al. showing the osteocyte and lumen channels [130]. MDA-MB-231 breast cancer cells are allowed to extravasate towards the osteocytes through the side channels [130]
Only 0.001–0.02% of tumour cells that enter circulation successfully metastasize [131]. However, metastasis significantly increases the mortality and morbidity of patients as the 5-year survival rate of metastatic cancer is < 30% [131]. Thus, in vitro platforms that can successfully mimic in vivo tumour metastasis are required for fundamental biological studies to better understand this process. Several microfluidic platforms to study tumour metastasis have been developed. Microfluidic metastasis models can use different stimuli to induce cancer cell invasion and migration through mechanical guidance, biochemical signalling, or through interaction with cells at the secondary site (extravasation) [131]. Microfluidic tissue models of metastasis can be complex in nature and typically modelled according to the secondary site of choice, as multiple cell types are involved in the metastatic cascade and require the relevant cell types, ECM, and spatial architecture for optimal recreation of the metastatic cascade at that location. Moreover, microfluidic metastatic models typically focus on a specific stage of the metastatic cascade. Figure 6B shows platelet-driven ovarian cancer metastasis on a multi-layered tumour chip model [128]. Figure 6C shows the schematic of lymphatic metastasis model developed by Cho et al. incorporating both lymph and blood vessels to demonstrate that circulating breast cancer cell EMT is driven by IL-6 [129]. Modelling in vivo biomechanical regimes are also crucial to developing relevant cancer metastasis models, particularly in environments with highly mechanosensitive cells. Figure 6D shows the platform developed by Mei et al. to model breast cancer bone metastasis [130]. This model could connect to an oscillatory flow pump to apply shear stress on osteocytes, which are sensitive to fluid shear stress and can modulate the functions of other cell types through loading-induced signalling [130]. In vitro microfluidic models to study metastasis can provide a complex but well-controlled microenvironment to study or discover the fundamental mechanisms that drive cancer metastasis and provide new information for the development of new cancer treatments that target key molecules involved in various steps of the metastatic cascade.
Ovarian cancer metastasis
Saha et al. developed a novel ovarian TME organ-on-a-chip model [128]. This platform incorporated a collagen-based ECM microenvironment with a tumour-interfacing platelet-perfused vascular endothelial channel (Fig. 6B) [128]. The authors studied the dynamics of ovarian cancer cell invasion following platelet extravasation through the endothelium into the TME using the novel microfluidic platform. The device allowed for time-lapse isolation of cancer cells based on degree of platelet perfusion to study effects on cancer cell growth and proliferation [128]. Gene editing and RNA sequencing of cancer cells from the tumour-on-a-chip platform determined that platelets and ovarian tumours interact through glycoprotein-VI (GPVI) and tumour Galectin-3 under shear stress [128]. Galectin-3 knockout reduced ovarian cancer invasion into the ECM when compared to wild type ovarian cancer cells [128]. Furthermore, treatment with GPVI inhibitor Revacept significantly reduced ovarian cancer cell invasion through the ECM and enhanced the effects of chemotherapy when combined with cisplatin [128]. This microfluidic model can be used for the identification of complex tumour-blood cells interactions and metastatic signalling as well as studies for antiplatelet therapeutics against tumour metastasis.
Lung cancer metastasis
Kim et al. presented a 3D microfluidic tri-culture platform to study crosstalk between cerebral metastatic lung cancer cells and the brain perivascular microenvironment [132]. The device consisted of four medium channels divided by three hydrogel channels [132]. Patient-derived lung carcinomas were isolated and subsequently cultured with human cerebral microvascular endothelial cells (hCMECs) and ACBRI 371 primary human brain astrocytes [132]. Tri-culture in the microfluidic device resulted in elevated secretions of tumour-promoting factors such as serpin E1, IL-8, and phosphoprotein 1 [132]. This platform allows for in vitro screening of potential molecular targets for anti-cancer therapies that target key components of the brain perivascular TME.
Liu et al. developed and established a multi-organ microfluidic platform that recapitulated the entire process of lung cancer brain metastasis [133]. This device consisted of an upstream “lung” and downstream “brain unit” [133] and was utilized to demonstrate that the recently discovered aldo–keto reductase family 1 member B10 (AKR1B10) protein is significantly elevated in brain metastatic lung cancer cells and is associated with lung cancer cell extravasation through the blood brain barrier. In the same research group, Xu et al. isolated brain metastatic cells from the microfluidic device and observed that these cells had developed significant resistance to several anti-cancer therapies compared to parent cells [134]. Proteomic analyses revealed that this acquired drug resistance in lung cancer derived brain metastasis was due to increased activity of the glutathione (GSH) metabolism pathway which contributes to intracellular antioxidant stress response [134]. Additionally, proteomics revealed an upregulation of aldehyde dehydrogenases and downregulation of epithelial growth factor receptor (EGFR), both of which contribute to resistance of EGFR-targeted therapies in the brain metastatic cells [134].
Breast cancer metastasis
Cho et al. developed a three-channel microfluidic device to understand the role of inflammatory cytokines on breast cancer lymphatic metastasis, with both blood and lymph vasculature (Fig. 6C(i., ii.)) [129]. A lumen in the lymph vessel channel was formed using human lymphatic endothelial cells (HLECs), and the blood vessel lumen was formed using HUVECs (Fig. 6C (iii.)) [129]. The inflammatory cytokine IL-6 mediated breast cancer cell and lymphatic vessel crosstalk [129]. Treatment with IL-6 resulted in higher expression of EMT markers in breast cancer cells and thereby higher invasion from the lymphatic vessel into the hydrogel ECM in the microfluidic platform [129]. The device can be used to better understand interactions between the tumour-lymph system interaction during lymphatic metastasis.
Mei et al. presented the first microfluidic tissue model capable of applying in vivo mechanical loading conditions to osteocytes to study breast cancer bone metastasis [130]. The authors aimed to determine the effects of osteocyte mechanical loading on breast cancer cell extravasation [130]. This platform consisted of a 3D lumen structure lined with HUVECs in one channel, and MLO-Y4 osteocytes seeded in the adjacent channel (Fig. 6D) [130]. Mechanical stimulation of osteocytes using oscillatory fluid flow (1 Pa peak shear stress, 1 Hz frequency for 2 h) significantly decreased breast cancer cell extravasation distance and the percentage of side channels containing extravasated cancer cells at the end of the experiment [130]. This platform can be utilized to study other cancers that tend to metastasize to bone such as prostate cancer, or for drug studies to study the effects of novel bone metastasis treatments in the bone-cancer microenvironment. Other 3D tumour models of bone metastasis have also been developed. Marturano-Kruik et al. developed a perfused human bone perivascular niche-on-a-chip to study the breast cancer colonization of the bone [135]. Human endothelial cells and bone marrow-derived MSCs were embedded in a native bone matrix within the microfluidic chip [135]. Cells were exposed to physiological flow velocities and oxygen gradient to mimic the in vivo microenvironment, resulting in a self-assembled vascular network for long-term cell culture [135]. Once established capillaries structures were formed, cancer cells were infused into the system. Cancer cells that were exposed to the physiological fluid flow exhibited a slower proliferative state compared to static cancer cells when colonizing the bone matrix, resulting in an increased resistance to chemotherapy [135].
Many anti-cancer therapies are being developed to target circulating tumour cell intravasation and extravasation. Microfluidic tissue models have been developed to study these phenomena to identify potential target markers for novel therapies during adhesion and trans-endothelial migration. Offeddu et al. utilized self-assembled perfusable microvascular networks (MVNs), a well-established platform that was previously developed by their research group [136]. In this study, the authors examined the role of the glycocalyx in tumour cell arrest, adhesion and trans-endothelial migration. They observed that removal of hyaluronic acid from the endothelial glycocalyx increases extravasation efficiency of MDA-MB-231 breast cancer cells, whereas removal of hyaluronic acid from the tumour cell glycocalyx significantly hindered cancer extravasation [136]. Using their microfluidic platform, they demonstrated that hyaluronic acid in the tumour glycocalyx promotes cancer cell adhesion through CD44 binding [136]. Although important components of the TME such as immune cells were missing, the use of the in vitro MVNs can provide a controlled environment to study glycocalyx-mediated mechanisms during metastasis [136]. Gilardi et al. utilized three microfluidic in vitro models to determine the role of the cyclin-dependent kinase 5 (Cdk5)/talin-1 (Tln1)/FAK axis in breast cancer cell adhesion, trans-endothelial migration, and early invasion during extravasation to the secondary site [137]. They observed that Cdk5 silencing affects vascular adhesion, and that the structural function, instead of activation through phosphorylation of Tln1 and FAK is required for the formation of invadopodia, which are required for extravasation through the endothelium [137]. In addition, Gilardi et al. found that silencing or inhibiting Tln1 and FAK significantly reduces breast cancer colonization of the lung [137]. Shirure et al. presented a tumour-on-a-chip platform to examine and compare tumour progression and drug sensitivity in common breast cancer cell lines and patient-derived tumour organoids [138]. The platform allowed for the observation of tumour growth and proliferation, angiogenesis, cell migration, and finally tumour intravasation at high spatiotemporal resolution [138].
Summary and future perspectives
This review article provided an overview on organ-on-a-chip models and the TME. The potential utility of cancer-on-a-chip models as an in vitro drug screening platform for preclinical studies or personalized medicine was discussed. Table 2 shows the main findings from microfluidic tissue models that included drug screening trials.
Recent advances of tumour-on-a-chip models from the last five years were presented. Studies that focused on tumour-blood vessel, tumour-stroma, or tumour-immune cell interactions and metastasis were examined. A summary of the major works discussed in this review can be found in Table 3, with main findings listed.
Although many tumour-on-a-chip models have been developed, several challenges remain.
-
I
Industrial manufacturing of microfluidic devices: The production process of microfluidic devices for 3D tumour-on-a-chip studies at a commercial level must be considered. The complexity in the fabrication of such devices must be reduced, and quality control must be ensured. Current approaches to manufacturing of devices at the industrial (large volume) scale include hot embossing and micro-injection moulding [139], however, hot embossing requires a vacuum environment for optimal device production, and injection moulding can results in several replication imperfections at the macro- and micro- scale level [139].
-
II
Optimization of device materials: The commonly used material for microfluidic devices is polydimethyl siloxane (PDMS). PDMS is hydrophobic and thereby prone to adsorbing drug molecules [140], which can lead to inaccurate drug screening results. Therefore, new materials for microfluidic platforms involving cancer therapies must be developed and optimized to be biocompatible, low-cost and inert.
-
III
Technical robustness and difficulty of use: Microfluidic devices for biological studies using multiple cell types can be generally more difficult to use compared to conventional methods. The use of these platforms may require rigorous training and/or specialized personnel. The scale and complexity of the platforms also require many factors to interplay perfectly to achieve full functionality of the device. Simple factors such as bubble formation can ruin a microfluidic experiment, reducing the technical robustness of these platforms [141]. Thus, platforms must be designed to be robust and easy to use [141].
-
IV
Standardization and reproducibility: Experimental data acquired from different devices need to be standardized, and reproducibility of the results obtained from these platforms must be ensured.
-
V
Long-term microfluidic studies: Long-term studies require the maintenance of cell viability of multiple cell types and of the structural integrity of the tissue/ECM using a common media and/or consistent perfusion. Other challenges associated with long-term microfluidic cell culture include effects from biofouling, which may cause issues with optical monitoring of samples, and unwanted nonspecific interactions that may occur in the system.
-
VI
Simulation and quantification of complex cell signalling: The simulation of complex cell signalling and the regulated response of other organs in the body to cancer such as signalling from the endocrine or immune system have not yet been fully replicated [15, 17, 26]. While tumour-immune cell interactions have been studied, the full signalling response of the immune system has not been replicated on a microfluidic platform. Only interactions between cancer and one or two immune cell types have been examined. Additionally, the complete simulation of primary tumours with distant organs such as bone to form the pre-metastatic niche has yet to be further developed and investigated. Moreover, developing advanced microfluidic tissue models with compatible and functional biosensors to measure real-time cell-secreted products such as metabolites provides a great challenge.
The development of new techniques and methods to further increase the feasibility of microfluidic platforms for TME studies will continue. Advanced microfluidic tissue models integrated with biosensors for the real-time monitoring of metabolites (glucose, lactate, oxygen), pH, temperature, or the measurement of electrical and mechanical properties of the cancer microenvironment are anticipated and could potentially help with the development of cancer-on-chip model that aim to simulate complex signalling as well as cancer metabolism studies. The use of biosensors in microfluidic systems can provide more information on the TME, increase the functionalities of these systems, and avoid the conventional endpoint monitoring of biological microfluidic models. A few microfluidic models have probed into cancer metabolism and have performed screening for metabolic anti-cancer treatments, and these studies are expected to continue. Future directions using microfluidic tissue models include examining how cancer metabolism changes from the onset of cancer, growth and development of the primary tumour, all the way to invasion or metastasis, which would aid in developing novel metabolic anti-cancer therapies to target tumour at various stages of development. Primary tumours may undergo metabolic reprogramming due to oncogenic alterations or signalling from host cells to induce a malignant phenotype and resistance to anti-cancer therapies [71, 73]. Observation of metabolic reprogramming of cancers in a controlled in vitro environment would provide opportunities for the development of effective metabolic therapies. The incorporation of biosensors on microfluidic platforms to monitor cancer metabolic products or waste (oxygen uptake, glucose, lactate, etc.) would be of great use by providing real-time measurements during tumour progression and for studying patient-derived samples in terms of the selection of cancer therapies targeting cancer cell/tumour metabolism for personalized medicine. Moreover, the design of cancer-on-a-chip devices that accounts for the three dimensionality of tissues and permits the spatiotemporal mapping and analysis of morphophysical features of the TME will continue to be improved [26, 142, 143]. Some of the works presented in this review demonstrated the first steps towards high-resolution spatiotemporal mapping [113]. Patient-derived samples or biopsies to generate the TME-on-a-chip for personalized medicine will also continue to be developed in the future. In this review, both patient derived CAFs, NFs, and cancer cells have been used in microfluidic studies. The increased incorporation of autologous cell sources demonstrates a shift away from the use of immortalized cell lines. More patient-derived cell types are expected to be used in microfluidic platforms to recapitulate the native TME. Artificial intelligence (AI) for analysis of the complex TME has the potential to identify novel features that may be missed from manual data analysis [144]. Automated analysis using AI allows for the interpretation of multinomic data in an objective and efficient manner [144].
Furthermore, the use of advanced microfluidic tissue models to study complex cellular interactions such as the formation of the pre-metastatic niche and cancer cell dormancy will be intensively conducted. Cancer cell dormancy is a phenomenon by which single cancer cells acquire a reversible, quiescent state in an attempt to adapt to environmental stressors [145]. These dormant cells are resistant to chemotherapies and radiation, which typically target cell division and are undetectable in current diagnostic tests [146]. Dormant cancer cells can be reactivated years after remission, resulting in recurrence [147]. However, the mechanisms by which cancer cell dormancy is induced or spontaneously reactivated remain largely unknown. Microfluidic tissue models to study dormancy mechanisms in a controlled, in vitro environment may provide answers on the induction or reactivation of cancer cell dormancy, as well as potential methods to target and sensitize dormant cancer cells to chemotherapies. While the concept of the pre-metastatic niche is relatively recent and has grown much interest, there are many aspects of this niche that are still unknown [77]. More specifically, the mechanisms behind which primary tumour are driven to release soluble factors to establish the pre-metastatic niche at the secondary site is not well known. The discovery of more components of the pre-metastatic niche, such as other host/resident or recruited cells, remains to be elucidated [78]. As microfluidic tissue models offer a controlled but physiologically relevant microenvironment, this may present an advantage in studying the entire process of the pre-metastatic niche, as well as a platform for screening novel treatments that aim to target or prevent pre-metastatic niche formation.
Finally, microfluidic tissue models will lead to more biological discoveries. While many microfluidic models were shown to demonstrate proof-of-concept through a drug screening or observation of specific in vivo behaviours, this review also introduced a few works that utilized microfluidic platforms for the identification of novel genes, markers or cell behaviours that play a key role in tumour growth or invasion [86, 94, 98, 112, 113, 120, 121, 123, 128, 133, 134, 136, 137]. Many of the tissue models reported in this review also demonstrate good correlation with in vivo studies either in terms of drug response [95] or of biological phenomena [86, 94, 95, 113, 123], such as endothelial ablation [86]. Ultimately, the advancement of microfluidic cnacer-on-a-chip models for practical applications will require a multidisciplinary approach through the knowledge and collaboration of microfluidics, tissue engineering, biomaterials, and cancer biology.
Availability of data and materials
Not applicable.
References
“Cancer.” World Health Organization, World Health Organization, 2021. https://www.who.int/health-topics/cancer#tab=tab_1. Accessed 21 Feb 2023.
“What Is Cancer?” National Cancer Institute, National Institutes of Health, 2021. https://www.cancer.gov/about-cancer/understanding/what-is-cancer#:~:text=Cancer%20is%20a%20disease%20in,up%20of%20trillions%20of%20cells. Accessed 21 Feb 2023.
Dagogo-Jack I, Shaw AT. Tumour heterogeneity and resistance to cancer therapies. Nat Rev Clin Oncol. 2018;15:81–94. https://doi.org/10.1038/nrclinonc.2017.166.
Zhang A, et al. Tumor heterogeneity reshapes the tumor microenvironment to influence drug resistance. Int J Biol Sci. 2022;18(7):3019–33. https://doi.org/10.7150/ijbs.72534.
Ramos A, et al. Battling Chemoresistance in Cancer: Root Causes and Strategies to Uproot Them. Int J Mol Sci. 2021;22(17):9451. https://doi.org/10.3390/ijms22179451.
Ades F, et al. The past and future of breast cancer treatment-from the papyrus to individualised treatment approaches. Ecancermedicalscience. 2017;11:746. https://doi.org/10.3332/ecancer.2017.746.
Arneth B. Tumor Microenvironment. Medicina. 2019;56(1):15. https://doi.org/10.3390/medicina56010015.
Anderson NM, Simon MC. The Tumor Microenvironment. Curr Biol. 2020;30(16). https://doi.org/10.1016/j.cub.2020.06.081.
Ungefroren H, et al. Interaction of tumor cells with the microenvironment. Cell Commun Signal. 2011;9(18). https://doi.org/10.1186/1478-811X-9-18.
Sahai E, et al. A framework for advancing our understanding of cancer-associated fibroblasts. Nat Rev Cancer. 2020;20:174–86. https://doi.org/10.1038/s41568-019-0238-1.
Zhou J, et al. Tumor-associated macrophages: recent insights and therapies. Front Oncol. 2020;10:188. https://doi.org/10.3389/fonc.2020.00188.
Choi H, Moon A. Crosstalk between cancer cells and endothelial cells: implications for tumor progression and intervention. Arch Pharmacal Res. 2018;41(7):711–24. https://doi.org/10.1007/s12272-018-1051-1.
Hida K, et al. Contribution of tumor endothelial cells in cancer progression. Int J Mol Sci. 2018;19(5):1272. https://doi.org/10.3390/ijms19051272.
Furesi G, et al. Emerging Players in Prostate Cancer-Bone Niche Communication. Trends Cancer. 2020;7(2):112–21. https://doi.org/10.1016/j.trecan.2020.09.006.
Dong Q, et al. Pre-metastatic Niche Formation in Different Organs Induced by Tumor Extracellular Vesicles. Front Cell Dev Biol. 2021;9. https://doi.org/10.3389/fcell.2021.733627.
Sun IO, et al. Circulating miRNAs in extracellular vesicles related to treatment response in patients with idiopathic membranous nephropathy. J Transl Med. 2022;20:224. https://doi.org/10.1186/s12967-022-03430-7.
Peinado H, et al. Pre-metastatic niches: organ-specific homes for metastases. Nat Rev Cancer. 2017;17:302–17. https://doi.org/10.1038/nrc.2017.6.
Alfarouk KO, et al. Resistance to cancer chemotherapy: failure in drug response from ADME to P-gp. Cancer Cell Int. 2015;15:71. https://doi.org/10.1186/s12935-015-0221-1.
Rosa R, et al. In vitro and in vivo models for analysis of resistance to molecular therapies. Curr Med Chem. 2014;21(14):1595–606. https://doi.org/10.2174/09298673113209990226.
Liu X, et al. Tumor-on-a-Chip: From Bioinspired Design to Biomedical Application. Microsyst Nanoeng;7(1) 2021. https://doi.org/10.1038/s41378-021-00277-8
Sontheimer-Phelps A, et al. Modelling Cancer in Microfluidic Human Organs-on-Chips. Nat Rev Cancer. 2019;19(2):65–81. https://doi.org/10.1038/s41568-018-0104-6.
Nam H, et al. Cancer cell migration and cancer drug screening in oxygen tension gradient chip. Biomicrofluidics. 2020;14(4):044107. https://doi.org/10.1063/5.0011216.
Rosser J, et al. Microfluidic nutrient gradient–based three-dimensional chondrocyte culture-on-a-chip as an in vitro equine arthritis model. Materials Today Bio. 2019;4:100023. https://doi.org/10.1016/j.mtbio.2019.100023.
Ahmed MAM, Nagelkerke A. Current developments in modelling the tumour microenvironment in vitro: Incorporation of biochemical and physical gradients. Organs-on-a-Chip. 2021;3:100012. https://doi.org/10.1016/j.ooc.2021.100012.
Trujillo-de Santiago G, et al. The Tumor-on-Chip: Recent Advances in the Development of Microfluidic Systems to Recapitulate the Physiology of Solid Tumors. Materials. 2019;12(18):2945. https://doi.org/10.3390/ma12182945.
Imparato G, et al. Organ on Chip Technology to Model Cancer Growth and Metastasis. Bioengineering. 2022;9(1):28. https://doi.org/10.3390/bioengineering9010028.
Obino D, et al. An Overview on Microfluidic Systems for Nucleic Acids Extraction from Human Raw Samples. Sensors (Basel, Switzerland). 2021;21(9):3058. https://doi.org/10.3390/s21093058.
Arter WE, et al. Microfluidic approaches for the analysis of protein-protein interactions in solution. Biophys Rev. 2020;12(2):575–85. https://doi.org/10.1007/s12551-020-00679-4.
Gaa R, et al. Versatile and rapid microfluidics-assisted antibody discovery. mAbs. 2021;13(1):1978130. https://doi.org/10.1080/19420862.2021.1978130.
Gómez FA. Using Microfluidics to Understand and Control the Cellular Microenvironment. Biological Applications of Microfluidics: Wiley, Hoboken, NJ; 2008. p. 10–28.
Preetam S, et al. Emergence of Microfluidics for next Generation Biomedical Devices. Biosens Bioelectron X. 2022;10:100106. https://doi.org/10.1016/j.biosx.2022.100106.
Cooksey GA, et al. A multi-purpose microfluidic perfusion system with combinatorial choice of inputs, mixtures, gradient patterns, and flow rates. Lab Chip. 2009;9(3):417–26. https://doi.org/10.1016/10.1039/b806803h.
Lin B, Levchenko A. Spatial Manipulation with Microfluidics. Front Bioeng Biotechnol. 2015;3. https://doi.org/10.3389/fbioe.2015.00039.
Smith Q, Gerecht S. Going with the flow: microfluidic platforms in vascular tissue engineering. Curr Opin Chem Eng. 2014;3:42–50. https://doi.org/10.1016/j.coche.2013.11.001.
Wu Q, et al. Organ-on-a-Chip: Recent Breakthroughs and Future Prospects. Biomed Eng OnLine. 2020;19(1). https://doi.org/10.1186/s12938-020-0752-0.
Abulaiti M, et al. Establishment of a heart-on-a-chip microdevice based on human iPS cells for the evaluation of human heart tissue function. Sci Rep. 2020;10:19201. https://doi.org/10.1038/s41598-020-76062-w.
Francis I, et al. Recent advances in lung-on-a-chip models. Drug Discov Today. 2022;27(9):2593–602. https://doi.org/10.1016/j.drudis.2022.06.004.
Liu J, et al. Design and Fabrication of a Liver-on-a-chip Reconstructing Tissue-tissue Interfaces. Front Oncol. 2022;12. https://doi.org/10.3389/fonc.2022.959299
Wang D, et al. Kidney-on-a-Chip: Mechanical Stimulation and Sensor Integration. Sensors. 2022;22(18):6889. https://doi.org/10.3390/s22186889.
Maoz BM. Brain-on-a-Chip: Characterizing the next generation of advanced in vitro platforms for modeling the central nervous system. APL Bioeng. 2021;5(3):030902. https://doi.org/10.1063/5.0055812.
Ahn J, et al. Tumor Microenvironment on a Chip: The Progress and Future Perspective. Bioengineering. 2017;4(4):64. https://doi.org/10.3390/bioengineering4030064.
Nagaraju S, et al. Microfluidic Tumor-Vascular Model to Study Breast Cancer Cell Invasion and Intravasation. Adv Healthc Mater. 2018;7(9):e1701257. https://doi.org/10.1002/adhm.201701257.
Azadi S, et al. Characterizing the effect of substrate stiffness on the extravasation potential of breast cancer cells using a 3D microfluidic model. Biotechnol Bioeng. 2021;118(2):823–35. https://doi.org/10.1002/bit.27612.
Liu Y, et al. Angiogenesis and Functional Vessel Formation Induced by Interstitial Flow and Vascular Endothelial Growth Factor Using a Microfluidic Chip. Micromachines. 2022;13(2):225. https://doi.org/10.3390/mi13020225.
Mondadori C, et al. Advanced microfluidic models of cancer and immune cell extravasation: a systematic review of the literature. Front Bioeng Biotechnol. 2020;8. https://doi.org/10.3389/fbioe.2020.00907.
Mehta P, et al. Microfluidics meets 3D cancer cell migration. Trends Cancer. 2022;8(8):683–97. https://doi.org/10.1016/j.trecan.2022.03.006.
Xie H, et al. Going with the flow: modeling the tumor microenvironment using microfluidic technology. Cancers. 2021;13(23):6052. https://doi.org/10.3390/cancers13236052.
Shang M, et al. Microfluidic modelling of the tumor microenvironment for anti-cancer drug development. Lab Chip. 2019;19(3):369–86. https://doi.org/10.1002/bit27612.
Rothbauer M, et al. Recent advances in microfluidic technologies for cell-to-cell interaction studies. Lab Chip. 2018;18(2):249–70. https://doi.org/10.1039/C7LC00815E.
Baka Z, et al. Cancer-on-chip technology: current applications in major cancer types, challenges and future prospects. Prog Biomed Eng. 2022;4(3):032001. https://doi.org/10.1088/2516-1091/ac8259.
Baghban R, et al. Tumor microenvironment complexity and therapeutic implications at a glance. Cell Commun Signal. 2020;18(1). https://doi.org/10.1186/s12964-020-0530-4.
Wan L, et al. Tumor-on-a-Chip for Integrating a 3D Tumor Microenvironment: Chemical and Mechanical Factors. Lab Chip. 2020;20(5):873–88. https://doi.org/10.1039/c9lc00550a.
Boedtkjer E, Pedersen SF. The Acidic Tumor Microenvironment as a Driver of Cancer. Annu Rev Physiol. 2020;82(1):103–26. https://doi.org/10.1146/annurev-physiol-021119-034627.
Dominiak A, et al. Communication in the Cancer Microenvironment as a Target for Therapeutic Interventions. Cancers. 2020;12(5):1232. https://doi.org/10.3390/cancers12051232.
Wei R, et al. Cellular and extracellular components in tumor microenvironment and their application in early diagnosis of cancers. Anal Cell Pathol. 2020;2020:1–13. https://doi.org/10.1155/2020/6283796.
Salvador E, et al. Tight junctions and the tumor microenvironment. Curr Pathobiol Rep. 2016;4:135–45. https://doi.org/10.1007/s40139-016-0106-6.
Zhou M, et al. The roles of connexins and gap junctions in the progression of cancer. Cell Commun Signal 2023;21(8). https://doi.org/10.1186/s12964-022-01009-9.
Thomas SK, et al. Paracrine and cell autonomous signalling in pancreatic cancer progression and metastasis. EBioMedicine. 2020;53:102662. https://doi.org/10.1016/j.ebiom.2020.102662.
Brassart-Pasco S, et al. Tumor microenvironment: extracellular matrix alterations influence tumor progression. Front Oncol. 2020;10:397. https://doi.org/10.3389/fonc.2020.00397.
Oudin MJ, Weaver VM. Physical and chemical gradients in the tumor microenvironment regulate tumor cell invasion, migration, and metastasis. Cold Spring Harb Symp Quant Biol. 2016;81:189–205. https://doi.org/10.1101/sqb.2016.81.030817.
Baumann Z, et al. Feed-forward loops between metastatic cancer cells and their microenvironment—the stage of escalation. EMBO Mol Med. 2022;14(6):e14283. https://doi.org/10.15252/emmm.202114283.
Emon B, et al. Biophysics of tumor microenvironment and cancer metastasis - a mini review. Comput Struct Biotechnol J. 2018;16:279–87. https://doi.org/10.1016/j.csbj.2018.07.003.
Winkler J, et al. Concepts of extracellular matrix remodeling in tumour progression and metastasis. Nat Commun. 2020;11:5120. https://doi.org/10.1038/s41467-020-18794-x.
Levental KR, et al. Matrix crosslinking forces tumor progression by enhancing integrin signaling. Cell. 2009;139(5):891–906. https://doi.org/10.1016/j.cell.2009.10.027.
Spill F, et al. Impact of the physical microenvironment on tumor progression and metastasis. Curr Opin Biotechnol. 2016;40:41–8. https://doi.org/10.1016/j.copbio.2016.02.007.
Bauer J, et al. Increased stiffness of the tumor microenvironment in colon cancer stimulates cancer associated fibroblast-mediated prometastatic activin A signaling. Sci Rep. 2020;10:50. https://doi.org/10.1038/s41598-019-55687-6.
Deng B, et al. Biological role of matrix stiffness in tumor growth and treatment. J Transl Med. 2022;20:540. https://doi.org/10.1186/s12967-022-03768-y.
Efthymiou G, et al. Shaping up the tumor microenvironment with cellular fibronectin. Front Oncol. 2020;10. https://doi.org/10.3389/fonc.2020.00641.
Barkovskaya A, et al. Proteoglycans as mediators of cancer tissue mechanics. Front Cell Dev Biol. 2020;8:569377. https://doi.org/10.3389/fcell.2020.569377.
Cox TR, et al. LOX-mediated collagen crosslinking is responsible for fibrosis-enhanced metastasis. Cancer Res. 2013;73(6):1721–32. https://doi.org/10.1158/0008-5472.CAN-12-2233.
Faubert B, et al. Metabolic Reprogramming and cancer progression. Science. 2020;368(6487):eeaw5473. https://doi.org/10.1126/science.aaw5473.
Korkaya H, Orsulic S. Editorial: the tumor microenvironment: recent advances and novel therapeutic approaches. Front Cell Dev Biol. 2020;8:586176. https://doi.org/10.3389/fcell.2020.586176.
Elia I, Haigis MC. Metabolites and the tumour microenvironment: from cellular mechanisms to systemic metabolism. Nat Metab. 2021;3:21–32. https://doi.org/10.1038/s42255-020-00317-z.
Shi R, et al. Metabolism in tumor microenvironment: Implications for cancer immunotherapy. MedComm. 2020;1(1):47–68. https://doi.org/10.1002/mco2.6.
DeBerardinis RJ, Chandel NS. Fundamentals of cancer metabolism. Sci Adv. 2016;2(5):e1600200. https://doi.org/10.1126/sciadv.1600200.
Martin TA, et al. “Cancer Invasion and Metastasis: Molecular and Cellular Perspective.” In: Madame Curie Bioscience Database. Austin (TX): Landes Bioscience; 2000–2013. https://www.ncbi.nlm.nih.gov/books/NBK164700/. Accessed 20 Feb 2023.
Doglioni G, et al. Interactions in the (Pre)metastatic Niche Support Metastasis Formation. Front Oncol. 2019;9. https://doi.org/10.3389/fonc.2019.00219.
Peinado H, et al. The secreted factors responsible for pre-metastatic niche formation: old sayings and new thoughts. Semin Cancer Biol. 2011;21(2):139–46. https://doi.org/10.1016/j.semcancer.2011.01.002.
Zeng H, et al. Cancer-associated fibroblasts facilitate premetastatic niche formation through lncRNA SNHG5-mediated angiogenesis and vascular permeability in breast cancer. Theranostics. 2022;12(17):7351–70. https://doi.org/10.7150/thno.74753.
Paolillo M, Schinelli S. Extracellular matrix alterations in metastatic processes. Int J Mol Sci. 2019;20(19):4947. https://doi.org/10.3390/ijms20194947.
Fong MY, et al. Breast-cancer-secreted miR-122 reprograms glucose metabolism in premetastatic niche to promote metastasis. Nat Cell Biol. 2015;17(2):183–94. https://doi.org/10.1038/ncb3094.
Kitamura T, et al. Immune cell promotion of metastasis. Nat Rev Immunol. 2015;15(2):73–86. https://doi.org/10.1038/nri3789.
Brassard-Jollive N, et al. In Vitro 3D Systems to Model Tumor Angiogenesis and Interactions with Stromal Cells. Front Cell Dev Biol 2020;8. https://doi.org/10.3389/fcell.2020.594903.
Park S, et al. Three-dimensional vascularized lung cancer-on-a-chip with lung extracellular matrix hydrogels for in vitro screening. Cancers. 2021;13(16):3930. https://doi.org/10.3390/cancers13163930.
Pradhan S, et al. A microvascularized tumor-mimetic platform for assessing anti-cancer drug efficacy. Sci Rep. 2018;8(1). https://doi.org/10.1038/s41598-018-21075-9.
Nguyen DT, et al. A biomimetic pancreatic cancer on-chip reveals endothelial ablation via alk7 signaling. Sci Adv. 2019;5(8). https://doi.org/10.1126/sciadv.aav6789.
Lim J, et al. Microvascularized tumor organoids-on-chips: advancing preclinical drug screening with pathophysiological relevance. Nano Converg. 2021;8(1). https://doi.org/10.1186/s40580-021-00261-y.
Michna R, et al. Vascularized microfluidic platforms to mimic the tumor microenvironment. Biotechnol Bioeng. 2018;115(11):2793–806. https://doi.org/10.1002/bit.26778.
Aazmi A, et al. Engineered vasculature for organ-on-a-chip systems. Engineering. 2022;9:131–47. https://doi.org/10.1016/j.eng.2021.06.020.
Nashimoto Y, et al. Vascularized cancer on a chip: the effect of perfusion on growth and drug delivery of tumor spheroid. Biomaterials. 2020;229:119547. https://doi.org/10.1016/j.biomaterials.2019.119547.
Kwak TJ, Esak L. In vitro modeling of solid tumor interactions with perfused blood vessels. Sci Rep. 2020;10(1). https://doi.org/10.1038/s41598-020-77180-1
Ayres Pereira M, Chio IIC. Metastasis in pancreatic ductal adenocarcinoma: current standing and methodologies. Genes. 2019;11(1):6. https://doi.org/10.3390/genes11010006.
Hosein AN, et al. Pancreatic cancer stroma: an update on therapeutic targeting strategies. Nat Rev Gastroenterol Hepatol. 2020;17(8):487–505. https://doi.org/10.1038/s41575-020-0300-1.
Truong D, et al. A Three-Dimensional (3D) Organotypic Microfluidic Model for Glioma Stem Cells – Vascular Interactions. Biomaterials. 2019;198:63–77. https://doi.org/10.1016/j.biomaterials.2018.07.048.
Hachey SJ, et al. An in vitro vascularized micro-tumor model of human colorectal cancer recapitulates in vivo responses to standard-of-care therapy. Lab Chip. 2021;21(7):1333–51. https://doi.org/10.1039/d0lc01216e.
Rodrigues J, et al. 3D In Vitro Model (r)Evolution: Unveiling Tumor-Stroma Interactions. Trends Cancer. 2021;7(3):249–64. https://doi.org/10.1016/j.trecan.2020.10.009.
Pape J, et al. 3D Cancer Models: The Need for a Complex Stroma, Compartmentalization and Stiffness. Front Bioeng Biotechnol. 2021;9. https://doi.org/10.3389/fbioe.2021.660502.
Truong DD, et al. A human organotypic microfluidic tumor model permits investigation of the interplay between patient-derived fibroblasts and breast cancer Cells. Cancer Res. 2019;79(12):3139–51. https://doi.org/10.1158/0008-5472.CAN-18-2293.
Lugo-Cintrón KM, et al. Breast Fibroblasts and ECM Components Modulate Breast Cancer Cell Migration Through the Secretion of MMPs in a 3D Microfluidic Co-Culture Model. Cancers. 2020;12(5):1173. https://doi.org/10.3390/cancers12051173.
Ayuso JM, et al. Organotypic microfluidic breast cancer model reveals starvation-induced spatial-temporal metabolic adaptations. eBioMedicine. 2018;37:144–57. https://doi.org/10.1016/j.ebiom.2018.10.046.
Fan Q, et al. A Novel 3-D Bio-Microfluidic System Mimicking in Vivo Heterogeneous Tumour Microstructures Reveals Complex Tumour-Stroma Interactions. Lab Chip. 2018;17(16):2852–60. https://doi.org/10.1039/C7LC00191F.
Fields GB. The Rebirth of Matrix Metalloproteinase Inhibitors: Moving Beyond the Dogma. Cells. 2019;8(9):984. https://doi.org/10.3390/cells8090984.
Winer A, et al. Matrix Metalloproteinase Inhibitors in Cancer Therapy: Turning Past Failures in Future Successes. Mol Cancer Ther. 2018;17(6):1147–55. https://doi.org/10.1158/1535-7163.MCT-17-0646.
Du Z, et al. Microfluidic System for Modelling 3D Tumour Invasion into Surrounding Stroma and Drug Screening. Biofabrication. 2018;10(3):034102. https://doi.org/10.1088/1758-5090/aac70c.
Lee JH, et al. Microfluidic Co-Culture of Pancreatic Tumor Spheroids with Stellate Cells as a Novel 3D Model for Investigation of Stroma-Mediated Cell Motility and Drug Resistance. J Exp Clin Cancer Res. 2018;37(1). https://doi.org/10.1186/s13046-017-0654-6.
Palamaris K, et al. Epithelial to Mesenchymal Transition: Key Regulator of Pancreatic Ductal Adenocarcinoma Progression and Chemoresistance. Cancers. 2021;13(21):5532. https://doi.org/10.3390/cancers13215532.
Kim J, et al. Microfluidic One-Directional Interstitial Flow Generation from Cancer to Cancer Associated Fibroblast. Acta Biomater. 2022;144:258–65. https://doi.org/10.1016/j.actbio.2022.03.044.
Ayuso JM, et al. Microfluidic model with air-walls reveals fibroblasts and keratinocytes modulate melanoma cell phenotype, migration, and metabolism. Lab Chip. 2021;21(6):113–1149. https://doi.org/10.1039/D0LC00988A.
Paterson K, et al. Microfluidic Technologies for Immunotherapy Studies on Solid Tumours. Lab Chip. 2021;21(12):2306–29. https://doi.org/10.1039/D0LC01305F.
Marshall JS, et al. An introduction to immunology and immunopathology. Allergy Asthma Clin Immunol. 2018;14(Suppl 2):49. https://doi.org/10.1186/s13223-018-0278-1.
He Z. and Zhang, Shuixing, “Tumor-Associated Macrophages and Their Functional Transformation in the Hypoxic Tumor Microenvironment.” Front Immunol. 2021;12:741305. https://doi.org/10.3389/fimmu.2021.741305.
Ayuso JM, et al. Microfluidic Tumor-on-a-Chip Model to Evaluate the Role of Tumor Environmental Stress on NK Cell Exhaustion. Sci Adv 2021;7(8). https://doi.org/10.1126/sciadv.abc2331.
Boussommier-Calleja A, et al. The Effects of Monocytes on Tumor Cell Extravasation in a 3D Vascularized Microfluidic Model. Biomaterials. 2019;198:180–93. https://doi.org/10.1016/j.biomaterials.2018.03.005.
Shelton SE, et al. Engineering approaches for studying immune-tumor cell interactions and immunotherapy. iScience. 2020;24(1):101985. https://doi.org/10.1016/j.isci.2020.101985.
Aung A, et al. An Engineered Tumor-on-a-Chip Device with Breast Cancer-Immune Cell Interactions for Assessing T-Cell Recruitment. Cancer Res. 2019;80(2):263–75. https://doi.org/10.1158/0008-5472.CAN-19-0342.
Mi S, et al. Three-dimensional microfluidic tumor-macrophage system for breast cancer cell invasion. Biotechnol Bioeng. 2019;116(7):1731–41. https://doi.org/10.1002/bit.26961.
Macpherson AM, et al. Epithelial ovarian cancer and the immune system: biology, interactions, challenges and potential advances for immunotherapy. J Clin Med. 2020;9(9):2967. https://doi.org/10.3390/jcm9092967.
Odunsi K. Immunotherapy in ovarian cancer. Ann Oncol. 2017;28(Suppl 8):viii1–7. https://doi.org/10.1093/annonc/mdx444.
Surendran V, et al. A Novel Tumor-Immune Microenvironment (Time)-on-Chip Mimics Three Dimensional Neutrophil-Tumor Dynamics and Neutrophil Extracellular Traps (Nets)-Mediated Collective Tumor Invasion. Biofabrication. 2021;13(3):035029. https://doi.org/10.1088/1758-5090/abe1cf.
Carroll MJ, et al. AlternativelyActivated Macrophages Upregulate Mesothelial Expression of P-Selectin to Enhance Adhesion of Ovarian Cancer Cells. Cancer Res. 2018;78(13):3560–73. https://doi.org/10.1158/0008-5472.CAN-17-3341.
Guo Z, et al. M2 Macrophages Promote NSCLC Metastasis by Upregulating CYRAB. Cell Death Dis. 2019;10:377. https://doi.org/10.1038/s41419-019-1618-x.
Kim H, et al. Macrophages-Triggered Sequential Remodeling of Endothelium-Interstitial Matrix to Form Pre-Metastatic Niche in Microfluidic Tumor Microenvironment. Adv Sci. 2019;6(11):1900195. https://doi.org/10.1002/advs.201900195.
Crippa M, et al. A microphysiological early metastatic niche on a chip reveals how heterotypic cell interactions and inhibition of integrin subunit β3 impact breast cancer cell extravasation. Lab Chip. 2021;21(6):1061–72. https://doi.org/10.1039/D0LC01011A.
Kim J, et al. Three-Dimensional Human Liver-Chip Emulating Premetastatic Niche Formation by Breast Cancer-Derived Extracellular Vesicles. ACS Nano. 2020;14(11):14971–88. https://doi.org/10.1021/acsnano.0c04778.
Labelle M, et al. Direct signalling between platelets and cancer cells induces an epithelial-mesenchymal-like transition and promotes metastasis. Cancer Cell. 2011;20(5):576–90. https://doi.org/10.1016/j.ccr.2011.09.009.
“Metastatic Cancer: When Cancer Spreads.” National Cancer Institute, National Institutes of Health, 2020. https://www.cancer.gov/types/metastatic-cancer. Accessed 21 Feb 2023.
Hapach LA, et al. Engineered Models to Parse Apart the Metastatic Cascade. NPJ Precis Oncol. 2019;3(1). https://doi.org/10.1038/s41698-019-0092-3.
Saha B, et al. Human Tumor Microenvironment Chip Evaluates the Consequences of Platelet Extravasation and Combinatorial Antitumor-Antiplatelet Therapy in Ovarian Cancer. Sci Adv. 2021;7(30). https://doi.org/10.1126/sciadv.abg5283.
Cho HY, et al. Microfluidic System to Analyze the Effects of Interleukin 6 on Lymphatic Breast Cancer Metastasis. Front Bioeng Biotechnol. 2021;8. https://doi.org/10.3389/fbioe.2020.611802.
Mei X, et al. Microfluidic Platform for Studying Osteocyte Mechanoregulation of Breast Cancer Bone Metastasis. Integr Biol. 2019;11(4):119–29. https://doi.org/10.1093/intbio/zyz008.
Ma YHV, et al. A review of microfluidic approaches for investigating cancer extravasation during metastasis. Microsyst Nanoeng;4(1). 2018. https://doi.org/10.1038/micronano.2017.104.
Kim H, et al. Recapitulated Crosstalk between Cerebral Metastatic Lung Cancer Cells and Brain Perivascular Tumor Microenvironment in a Microfluidic Co-Culture Chip. Adv Sci (Weinheim, Baden-Wurttemberg, Germany). 2022;9(22):e2201785. https://doi.org/10.1002/advs.202201785.
Liu W, et al. AKR1B10 (Aldo-keto reductase family 1 B10) promotes brain metastasis of lung cancer cells in a multi-organ microfluidic chip model. Acta Biomater. 2019;91:195–208. https://doi.org/10.1016/j.actbio.2019.04.053.
Xu M, et al. Proteomic Reveals Reasons for Acquired Drug Resistance in Lung Cancer Derived Brain Metastasis Based on a Newly established Multi-Organ Microfluidic Chip Model. Front Bioeng Biotechnol. 2020;8. https://doi.org/10.3389/fbioe.2020.612091.
Marturano-Kruik A, et al. Human Bone Perivascular Niche-on-a-Chip for Studying Metastatic Colonization. Proc Natl Acad Sci. 2018;115(6):1256–61. https://doi.org/10.1073/pnas.1714282115.
Offeddu G, et al. The Cancer Glycocalyx Mediates Intravascular Adhesion and Extravasation during Metastatic Dissemination. Commun Biol. 2021;4:255. https://doi.org/10.1038/s42003-021-01774-2.
Gilardi M, et al. The Driving Role of the Cdk5/Tln1/FAKS732 Axis in Cancer Cell Extravasation Dissected by Human Vascularized Microfluidic Models. Biomaterials. 2021;276:12097. https://doi.org/10.1016/j.biomaterials.2021.120975.
Shirure VS, et al. Tumor-on-a-Chip Platform to Investigate Progression and Drug Sensitivity in Cell Lines and Patient-Derived Organoids. Lab Chip. 2018;18(23):3687–702. https://doi.org/10.1039/c8lc00596f.
Scott SM, Ali Z. Fabrication methods for microfluidic devices: an overview. Micromachines (Basel). 2021;12(3):319. https://doi.org/10.3390/mi12030319.
van Meer BJ, et al. Small molecule absorption by PDMS in the context of drug response bioassays. Biochem Biophys Res Commun. 2017;482(2):323–8. https://doi.org/10.1016/j.bbrc.2016.11.062.
Liu Y, et al. Human in vitro vascularized micro-organ and micro-tumor models are reproducible organ-on-a-chip platforms for studies of anticancer drugs. Toxicology. 2020;445:152601. https://doi.org/10.1016/j.tox.2020.152601.
Yu J, et al. Reconfigurable open microfluidics for studying the spatiotemporal dynamics of paracrine signalling. Nat Biomed Eng. 2019;3:830–41. https://doi.org/10.1038/s41551-019-0421-4.
Ronteix G. High resolution microfluidic assay and probabilistic modeling reveal cooperation between T cells in tumor killing. Nat Commun. 2022;13:3111. https://doi.org/10.1038/s41467-022-30575-2.
Fetah KL, et al. Cancer Modeling-on-a-Chip with Future Artificial Intelligence Integration. Small. 2019;15(50):1901985. https://doi.org/10.1002/smll.201901985.
Santos-de-Frutos K, Djouder N. When dormancy fuels tumour relapse. Commun Biol. 2021;4:747. https://doi.org/10.1038/s42003-021-02257-0.
Gomis RR, Gawrzak S. Tumor cell dormancy. Mol Oncol. 2017;11(1):62–78. https://doi.org/10.1016/j.molonc.2016.09.009.
Aguirre-Ghiso JA. Models, mechanisms and clinical evidence for cancer dormancy. Nat Rev Cancer. 2007;7(11):834–46. https://doi.org/10.1038/nrc2256.
Acknowledgements
L.Y. acknowledges the Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grant number 06465-14. K.S. thanks the Ontario Graduate Scholarship (OGS) program and the Barbara and Frank Milligan Graduate Fellowship for the student scholarships.
Funding
Open access funding provided by Shanghai Jiao Tong University. This research was supported by Natural Sciences and Engineering Research Council of Canada (NSERC) Discovery Grant, grant number 06465–14.
Author information
Authors and Affiliations
Contributions
Conceptualization, K.S.; writing – original draft preparation, K.S.; writing – reviewing and editing, K.S., Y.S., and L.Y.; Y.S. and L.Y. revised and provided critical feedback of the manuscript. All authors have read and agreed to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests that are relevant to the context of this article.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Seaman, K., Sun, Y. & You, L. Recent advances in cancer-on-a-chip tissue models to dissect the tumour microenvironment. Med-X 1, 11 (2023). https://doi.org/10.1007/s44258-023-00011-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44258-023-00011-1