Abstract
Background:
The von Hippel–Lindau (VHL) gene encodes two mRNA variants. Variant 1 encodes two protein isoforms, pVHL213 and pVHL160, that have been extensively documented in the literature. Variant 2 is produced by alternative splicing of exon 2 and encodes a pVHL isoform of 172 amino acids with a theoretical molecular weight of 19 kDa (pVHL172), the expression of which has never been demonstrated so far due to the absence of suitable antibodies.
Methods:
We have generated an anti-pVHL monoclonal antibody (JD-1956) using pVHL172 recombinant protein. We tested the antibody against exogenous or endogenous expressed proteins in different cell lines. We identified the pVHL172 using a silencing RNA strategy. The epitope of the antibody was mapped using a peptide array.
Results:
We efficiently detected the three different isoforms of pVHL in cell lines and tumorigenic tissues by western blotting and immunohistochemistry and confirmed for the first time the endogenous expression of pVHL172.
Conclusions:
The endogenous expression of the three isoforms and particularly the pVHL172 has never been shown before due to a lack of a highly specific antibody since none of the available commercial antibodies distinguish the three isoforms of pVHL in cells or in both normal and cancerous human tissues. Evidence of pVHL172 expression emphasises the need to further study its implication in renal tumorigenesis and VHL disease.
Similar content being viewed by others
Main
The von Hippel–Lindau (VHL) gene was isolated in 1993 (Latif et al, 1993) and was then characterised as a tumour suppressor gene (Iliopoulos et al, 1995; Clark and Cookson, 2008). Germline mutations in the VHL gene cause the VHL disease, an autosomal dominant familial cancer syndrome that predisposes to the development of retinal angioma, cerebellar and spinal haemangioblastoma, clear-cell renal cell carcinoma (ccRCC) and pheochromocytoma, as well as pancreatic disease. Somatic VHL mutations have also been found in patients with sporadic renal cell carcinoma, particularly ccRCC, which is the most common type of kidney cancer (Gnarra et al, 1994). These mutations are often detected in malignant and frequently metastatic ccRCCs, suggesting that VHL has a key role in the development/progression of kidney cancer.
The VHL gene encodes pVHL, a substrate-binding component of an E3 ubiquitin ligase complex. This complex targets HIF-α, a hypoxia-inducible transcription factor, for proteasomal degradation in a pVHL-dependent manner. pVHL has also other functions that are considered to be independent from its role in the E3 ubiquitin ligase complex. These include, for instance, regulation of the extracellular matrix, microtubule stability and primary cilium maintenance, regulation of E-cadherin and stabilisation of p53 and Jade-1 (genes involved in apoptosis and epithelium differentiation, respectively) (Bader and Hsu, 2012; Hsu, 2012). Failure of pVHL to control these functions may contribute to tumour progression and metastasis formation.
The complexity of pVHL functions is not only due to its involvement in multiple cellular processes, but also to the fact that the VHL gene, which is located on the short arm of human chromosome 3 (3p25.3), produces two different VHL mRNAs (Gnarra et al, 1994; Richards et al, 1996): variant 1 (V1; NM_000551) includes exons 1, 2 and 3, while variant 2 (V2; NM_198156) lacks exon 2 (Richards et al, 1996).
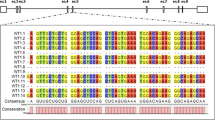
V1 encodes a protein of 213 amino acids with apparent molecular weight of ∼30 kDa (pVHL213) and a smaller 160 amino-acid long isoform of 19 kDa (pVHL160) (Schoenfeld et al, 1998) the translation of which initiates from an internal translation start site (codon 54) within the VHL open reading frame (Figure 1). Although they are both ubiquitously expressed, high expression levels are observed specifically in the urogenital system, brain, spinal cord, sensory ganglia, eyes and bronchial epithelium (Richards et al, 1996). Moreover, while pVHL213 is primarily found in the cytoplasm with minor pools in the nuclear and membrane compartments, pVHL160 is equally distributed in the nucleus and cytoplasm (Iliopoulos et al, 1998). When overexpressed in mammalian cells, the subcellular localisation of pVHL213 appears to vary in a cell density-dependent manner (Lee et al, 1996). Both pVHL213 and pVHL160 are functional tumour suppressors. Indeed, reintroduction of either isoform in VHL-defective ccRCC cells suppresses their ability to form tumours when xenografted in nude mice (Schoenfeld et al, 1998).
The second mRNA variant (V2) is produced by alternative splicing of exon 2 and should generate a protein composed of 172 amino acids with a theoretical molecular weight of 19 kDa (pVHL172) (Figure 1). In healthy adult kidney or in cultured renal proximal tubule cells, the expression of V2 mRNA can be barely detected by quantitative RT–PCR analysis and is very low compared with V1 (Herman et al, 1994; Gnarra et al, 1994; Richards et al, 1996). As a consequence, functional studies have mainly focused on the pVHL213/160 isoforms. However, there are at least two contexts in which high V2 expression has been reported: (i) during human embryogenesis (8–10 gestational weeks) in kidney, brain, spinal cord, eyes, testis and lung; and (ii) in sporadic ccRCC with specific VHL point mutations in exon–intron boundaries or in the coding region (Martella et al, 2006; Taylor et al, 2012). In some cases, V2 is the only VHL transcript detected in ccRCC samples or cell lines (Herman et al, 1994; Gnarra et al, 1994), suggesting that the protein encoded by this alternatively spliced transcript is defective in tumour suppressor activity (Gnarra et al, 1994; Shuin et al, 1994; Whaley et al, 1994) and/or participates in tumour initiation or progression.
So far, pVHL172 expression has not been rigourously demonstrated due to the lack of adequate tools and thus it has been impossible to compare the expression profile of all three protein isoforms to study their respective roles in ccRCC development and/or progression. Indeed, based on their primary sequence similarity, pVHL213 and pVHL160 are expected to share many functions (Robinson and Ohh, 2014). Conversely, the absence of part of the β-domain (aa 114–154) in the pVHL172 isoform modifies the number of beta sheets in its structure, which is likely to be involved in altered protein folding with structural and functional consequences on its activity and protein interaction network.
We have therefore generated a monoclonal antibody that efficiently discriminates between closely related pVHL isoforms. Using highly purified recombinant human pVHL172 as the antigen, this new antibody recognises the full spectrum of endogenous pVHL isoforms, including the particular pVHL172 isoform, which has been a subject of debate within the scientific community. This new development enables us to perform detailed studies on the expression of the different pVHL isoforms with particular focus on pVHL172. We report for the first time the use of an unambiguous technology for the in vitro and in vivo study of isoform-specific pVHL expression in various cell lines and tumour tissues.
Materials and Methods
Real-time PCR analysis
Real-time PCR was performed on total RNA extracted from cells using the QIAamp total RNA kit (Qiagen, Courtaboeuf, France). Five micrograms of total RNA were reverse-transcribed using oligo-dT primers and M-MLV reverse transcriptase. The resulting cDNAs were then PCR amplified using the following primers designed from human cDNA sequences: 5′-CCCGTATGGCTCAACTTCG-3′ (forward) and 5′-TCAGGTCGCTCTACGAAGATCT-3′ (reverse) for VHL variant 1 (308 bp); 5′-CCCGTATGGCTCAACTTCG-3′ (forward) and 5′-TCAGGTCGCTCTACGAAGATCT-3′ (reverse) for VHL variant 2 (185 bp) (Martin et al, 2013). Assays were performed in triplicate, using the RotorGene 3000 instrument (Corbett Research, Biolabo, Archamps, France) with SYBR Green I master mix (Roche Diagnostics, Mannheim, Germany). For each sample, the relative amounts of VHL and GAPDH transcripts were calculated from these standard curves using the RotorGene software.
Expression and purification of pVHL isoforms
There is conflicting nomenclature used by different investigators to assign pVHL isoforms: pVHL213 is also known as pVHL30 or pVHL25, while pVHL160 is also referred as pVHL19 or pVHL21. In this study, to prevent this confusion, each isoform will be named according to the number of amino acids of the molecule, that is, pVHL213, pVHL160 and pVHL172.
All recombinant proteins were prepared using the E.coli strain BL21(DE3)pLysS transformed with pET21a-hVHL213, hVHL172 or hVHL160 and induced to express the proteins VHL213(His)6, VHL172(His)6 and VHL160(His)6 with IPTG. Proteins were prepared and purified by Talon affinity chromatography following the manufacturer’s instructions (Clontech, Mountain View, CA, USA) as described by Martin et al (2013). The purity of the eluted fractions was assessed by 12.5% SDS–PAGE and silver staining.
Antibodies
The rabbit polyclonal antibody against human VHL (#6030) (Martin et al, 2013) and the mouse monoclonal antibody JD-1956 (Patent No. 14305925.1-1402-2014) against human VHL were produced in our laboratory (CNRS-EFS). The commercial antibody against human VHL was from Santa Cruz Biotechnology (sc-5575; Heidelberg, Germany). The actin antibody was from Sigma-Aldrich (Saint-Quentin Fallavier, France). The horseradish peroxidase-conjugated secondary antibodies were from Jackson Immuno-Research Laboratories (Baltimore, MD, USA).
Production of anti-pVHL antibody
Anti-pVHL monoclonal antibodies were raised by immunising six mice with recombinant human pVHL172 (His)6 subcutaneously. After 3 months, blood was collected from each mouse and the immunoreactivity of the different blood samples was tested. Mouse n°28 was killed and splenocytes were collected. A first selection of fused cells was done using an ELISA assay with recombinant pVHL213(His)6, pVHL172(His)6 and Aurora(His)6 to identify cells producing antibodies against the (His)6 tag. Ten to 100 ng of each recombinant protein (pVHL213(His)6, pVHL172(His)6 and Aurora-A) were loaded on polyacrylamide gels and transferred onto nitrocellulose membranes to test the clones. We found 25 clones that were immunoreactive against the recombinant VHL proteins. The immunoreactivity of the positive clones was then tested using total protein extracts from cells expressing exogenous VHL proteins.
Western blot analysis
Frozen tissues or pelleted cells were lysed in RIPA buffer (50 mM Tris-HCl, pH 7.4, 1% NP-40, 0.5% sodium deoxycholate, 150 mM sodium chloride, 1 mM EDTA, 1 mM sodium fluoride, 1 mM AEBSF, 10 μg/ml aprotinin, 10 μg/ml leupeptin and 1 mM sodium orthovanadate) and centrifuged at 9000 g for 10 min. Fifty micrograms of total proteins from each extract were separated by SDS–PAGE on 12.5% polyacrylamide gels in denaturing conditions and transferred onto nitrocellulose membranes. Membranes washed with TBST were incubated with primary antibodies in 2.5% low-fat milk in TBST at 4 °C overnight, followed by incubation with horseradish peroxidase-conjugated anti-rabbit or anti-mouse IgG antibodies (1 : 15 000 in TBST/2.5% BSA at RT for 1 h). Sample loading was controlled with a polyclonal anti-tubulin antibody (1 : 500). Western blots were revealed by chemiluminescence using the Super Signal ECL reagent kit (Pierce, Rockford, IL, USA).
Tissue samples
Tumour and matched normal tissue samples were obtained from patients with ccRCC who underwent partial or total nephrectomy between 2002 and 2005. The Ethics Committee of Rennes University Medical School and Hospital approved this prospective study and all patients signed the informed consent. Tissue samples were immediately frozen in liquid nitrogen and stored at −80 °C in the Centre de Ressources Biologiques (CRB, Rennes, France) and formalin-fixed and paraffin-embedded. To determine the VHL status of the collected samples, denaturing high-performance liquid chromatography was carried out on a WAVE Nucleic Acid Fragment Analysis system (Transgenomic, Glasgow, UK) with a DNAsep column (Patard et al, 2009). Aberrant peaks were further analysed by direct sequencing using standard procedures. All mutations were confirmed by a second PCR and sequencing reaction. Multiplex ligation-dependent probe amplification was used for VHL deletion analysis.
Cell culture
Human tumour cell lines (HeLa, HEK-293T and RCC4) were maintained in DMEM (Life Technologies, St Aubin, France) supplemented with 10% foetal calf serum and antibiotics at 37 °C in a humidified 5% CO2 atmosphere. The two primary cell lines (R-180 and R-305) derived from two human ccRCCs and the 786-0 were cultured in RPMI-1640 (Life Technologies) supplemented with 10% foetal calf serum (Sigma-Aldrich), 1% penicillin/streptomycin (Sigma-Aldrich) and 10 mM HEPES buffer (Dugay et al, 2014). HeLa cells were transfected using JetPRIME as recommended by the manufacturer (PolyPlus, Ozyme, Montigny-le-Bretonneux, France). To overexpress pVHL in the various cell lines, we used the pCDNA-VHL213 and pCDNA-VHL160 plasmids (subcloned from pCMV-FlagHA-VHL213/160 that were a generous gift from Dr A Buchberger-Max Planck Institute of Biochemistry, Department of Molecular Cell Biology, Martinsried, Germany) and the pcDNA-FlagHApVHL172 plasmid.
Immunohistochemistry
Five-micron sections of formalin-fixed paraffin-embedded tissues were transferred to glass slides and incubated in TBST in the presence of 5% BSA. Reactivity to JD-1956 (1 : 100) was revealed with the biotin–streptavidin detection system (Dako, Glostrup, Denmark) using diaminobenzidine as chromogen (Sigma-Aldrich). Photographs were taken using a Leica DMRXA microscope equipped with a CoolSnapsHQ camera (Photometrics, Tucson, AZ, USA). The images were processed with ImageJ 1.4 software (the National Institute of Mental Health, Bethesda, MD, USA).
Epitope mapping
The 213 amino acids sequence of pVHL213 (Uniprot accession code P40337) was used to generate an overlapping peptide library. The library is composed of 20-mer peptides with 5 amino acid off-set (40 peptides). Peptides were synthesised on cellulose membrane by SPOT technology (Frank, 2002) using an automated multiple synthesiser (MultiPep RS, Intavis, Köln, Germany). The membrane was rinsed with a small volume of methanol for 5 min to avoid precipitation of hydrophobic peptides during the subsequent procedure. The membrane was processed as described in western blot analysis, then incubated with the antibody JD-1956 (dilution 1 : 200) in 2.5% low-fat milk in TBST at 4 °C overnight, followed by incubation with horseradish peroxidase-conjugated anti-mouse IgG antibodies (1 : 15 000 in TBST/2.5% BSA at room temperature for 1 h). The blot was revealed by chemiluminescence using the Super Signal ECL reagent kit (Pierce).
Results
Immunodetection of separate pVHL isoforms
As most of the available antibodies against pVHL are unspecific and cannot efficiently distinguish or detect the different pVHL isoforms (Supplementary Figure S1 and Supplementary Table S1), we produced a new monoclonal antibody against VHL (JD-1956) using recombinant human pVHL172 as antigen. This antibody has been developed with the intention of specifically recognising the pVHL172 isoform. However, given the high sequence similarity of the three isoforms (Figure 1), it was expected to target also the isoforms encoded by V1 (pVHL213 and pVHL160). The antibody specificity was first assessed by immunoblot analysis using T7- and His6-tagged recombinant pVHL213, pVHL172 and pVHL160. The amount of loaded proteins was controlled by silver staining (Figure 2A, left panel) and immunodetection with the anti-T7 antibody (Figure 2A, right panel). The membrane was then immunoblotted with different antibodies against pVHL: (i) the polyclonal rabbit antibody VHL-6030 that preferentially recognises the NH2-terminal part of full-length pVHL (Martin et al, 2013, ii) the polyclonal antibody sc-5575 from Santa Cruz Biotechnology raised against full-length pVHL and that was one of the most efficient among the tested commercial antibodies (Supplementary Table S1) and (iii) our new monoclonal antibody JD-1956. All three antibodies could detect recombinant pVHL213 (Figure 2B). The pVHL172 isoform was clearly detected by the antibodies VHL-6030 and JD-1956, but not by the commercial antibody (only a very faint signal). Finally, pVHL160 was mainly revealed by the JD-1956 antibody. Thus, the JD-1956 antibody recognised efficiently all three pVHL isoforms. It should be noted that the apparent molecular weight of pVHL172 (23 kDa) differed significantly from the expected theoretical molecular weight (19 kDa), whereas in the case of pVHL160 the difference was minor (19 kDa instead of 18 kDa).
Immunodetection of recombinant pVHL isoforms with different anti-VHL antibodies. (A) Loading of the different recombinant pVHL proteins was evaluated by silver staining and by western blotting with an anti-T7 antibody. (B) Immune detection of the three recombinant pVHL proteins was performed using the polyclonal antibody VHL-6030 (1 : 1500), a commercial polyclonal anti-VHL antibody from Santa Cruz Biotechnology (1 : 500) and the JD-1956 monoclonal antibody (1 : 100). (C) Total protein extracts from HeLa cells transiently transfected with pCDNA-FlagHA plasmids encoding pVHL213, pVHL172 and pVHL160 were analysed by western blotting using an anti-Flag antibody, the commercial polyclonal anti-VHL (1 : 500), VHL-6030 (1 : 1500) and JD-1956 (1 : 100).
We then assessed how efficiently these antibodies could detect Flag-HA-tagged pVHL213, pVHL172 and pVHL160 in cell lysates. HeLa cells were transfected with the corresponding plasmids (Figure 2C) and cell lysates were then immunoblotted with the three antibodies against pVHL. All three antibodies detected pVHL213 and pVHL172. In contrast, the pVHL160 isoform was detected only with the commercial antibody (Santa Cruz Biotechnology) and the monoclonal JD-1956 antibody. However, non-specific bands were not observed when using the JD-1956 antibody, differently from the commercial antibody (*, panel commercial VHL-Ab of Figure 2C). These results indicate that only the mouse monoclonal antibody JD-1956 is appropriate to detect all known pVHL isoforms with high specificity in complex protein extracts.
Characterisation of the antibody specificity
Western blot analysis of HeLa cell extracts (Figure 3A, left panel) showed that the JD-1956 antibody detects three bands that could correspond to the three pVHL isoforms based on their respective electrophoretic mobility (*, pVHL213; **, pVHL172 and ***, pVHL160). Pre-incubation of the antibody with an excess of (His)6-tagged recombinant pVHL213, fully abolished immunodetection of the three bands by western blotting (compare the left and right panels of Figure 3A). To further investigate the specificity of the JD-1956 antibody, HeLa cells were transfected with siRNAs to downregulate all variants (siRNA VHL) or only pVHL172 (siRNA VHL-172). Western blot analysis of HeLa cell extracts 24 h after transfection showed that, in cells transfected with the siRNA VHL, the intensity of the three bands corresponding to the three pVHL isoforms was strongly decreased compared with non-transfected cells (Figure 3B, lanes 3 and 1, respectively). Conversely, in cells transfected with the siRNA VHL-172, only the intensity of the band expected to correspond to pVHL172 was reduced compared with non-transfected cells (Figure 3B, asterisk in lane 2). This effect was dose-dependent as indicated by the progressive reduction in the intensity of the band corresponding to pVHL172 in cells transfected with increasing amounts (25–75 nM) of siRNA VHL-172 (Figure 3C). Transfection with scrambled siRNA (50 nM) had no effect on the detection of the three bands (Figure 3C). Again, these pVHL isoform-specific knockdowns indicate that antibody JD-1956 is directed against all three pVHL isoforms (recombinant or endogenously expressed) with high specificity.
Characterisation of the JD-1956 antibody. (A) HeLa cell total protein extracts were analysed by western blotting with the JD-1956 antibody (−, left lane) or with JD-1956 after incubation with an excess of recombinant pVHL213 protein at 4 °C for 30 min (+, right lane). (B) HeLa cells were transfected with siRNAs (60 pmol) to downregulate all the three pVHL isoforms (siRNA VHL) or pVHL172 only (siRNA VHL-172). Cells were harvested 24 h after transfection and the expression level of the different pVHL isoforms was investigated by western blotting using the JD-1956 antibody (1 : 100). Non-transfected HeLa cells (−) were used to assess the expression of the endogenous pVHL isoforms. (C) HeLa cells were transfected with increasing amounts of siRNA VHL-172 (50–150 pmol) for 30 h and then the expression of the pVHL isoforms was investigated by immunoblotting using JD-1956 (1 : 100). Non-transfected HeLa cells (−) or transfected with scrambled siRNA (Scr, 100 pmol) were used as controls. In B and C, signals corresponding to VHL isoforms (as well as tubulin) were quantified using ImageQuant software (GE Healthcare Europe, Velizy-Villacoublay, France), VHL signals were normalised against tubulin and results presented as mean percentages±s.e.m. (n=3) of the signal (for each pVHL isoform) observed in control HeLa cells.
Finally, epitope mapping study showed that JD-1956 recognises an amino acid sequence located between amino acids 46 and 72 (peptides #10–12), which is an epitope region shared by all pVHL isoforms (Supplementary Figure S2).
pVHL expression profiling in tumour cell lines and tumour tissues
We then sought to determine whether the JD-1956 antibody could also recognise endogenous pVHL expressed in different cell lines derived from primary ccRCC tumours (786-O, R-180, R-305, Caki-1 and RCC4+ cells), HEK-293T cells and non-tumoral endothelial cells (Huvec). Figure 4A shows the VHL status of all tested cell lines. First, we checked by RT–PCR analysis whether the two VHL variants (V1 and V2) were expressed in selected wild-type VHL (HeLa and Huvec) and mutated/deleted VHL (R-305) cell lines. Two bands of the expected molecular size (308 bp for V1 and 185 bp for V2) were detected (Figure 4B). Western blot analysis using the commercial antibody revealed a single major band with a molecular weight of about 23 kDa in all tested cell lines (upper panel, Figure 4C), although 786-0 cells do not express full-length pVHL (Gnarra et al, 1994). Thus, this band likely corresponds to a non-specific signal. Two additional faster migrating bands were detected in R-305 and RCC4+ cells, but their molecular weight did not correspond to the theoretical molecular weight of the smaller pVHL isoforms (Figure 4C, upper panel). The VHL-6030 antibody detected bands that theoretically corresponded to pVHL213 and pVHL160, but not to the putative pVHL172 isoform, in most of the tested cell extracts (Figure 4C, middle panel). On the other hand, the JD-1956 antibody revealed three strong bands (*, pVHL213; **, pVHL172 and ***, pVHL160) in HeLa, RCC4+ (VHL-deficient RCC cells transfected with a plasmid coding for full-length pVHL213) and HEK-293T cell extracts. As expected, no band was observed in lysates from 786-0 cells in which a VHL mutation has introduced a stop codon in the middle of the mRNA coding sequence (Figure 4B, lower panel). These results show that the JD-1956 antibody can efficiently recognise the three different pVHL isoforms in various contexts (recombinant proteins, overexpressed or endogenous proteins). The expression profile of non-tumoral Huvec cells was similar to that of HeLa cells. The JD-1956 antibody was then used to examine the relative expression of the different pVHL isoforms in ccRCC samples with different VHL status (Figure 5A). HeLa cell extract was used as a positive control (Figure 5B right lane). Western blot analysis with the JD-1956 antibody revealed three bands that likely corresponded to *, pVHL213; **, pVHL172 and ***, pVHL160, based on their electrophoretic mobility. The relative intensity of each of the three bands could vary within and between samples, particularly if their VHL status was different (Table 1). The intensity of the lowest band (theoretically corresponding to pVHL160) (‘***’ in Figure 5B) was higher compared with the other bands particularly in tumour samples in which VHL was deleted in one allele and mutated in the other (del/mut) (Figure 5B, lanes 2, 5 and 8). These results suggest that JD-1956 is a suitable tool for tracking the expression of specific pVHL isoforms and to establish relative expression ratios in a variety of cell lines and tumour tissues.
Detection of the VHL isoforms (RNA and proteins) in cell lines. (A) VHL status of the different cell lines. (B) Total RNA was reverse-transcribed and PCR amplified using VHL primers for V1 (V1, 308 bp) and V2 (V2, 185 bp). (C) 40 μg of total protein extracts from HeLa cells, kidney cell lines (786-O, R-180, R-305, Caki-1, RCC4+ and HEK-293T) and HUVEC cells were analysed by western blotting with the commercial anti-VHL antibody (1 : 500), VHL-6030 (1 : 1500) and JD-1956 (1 : 10). β-tubulin was used as a loading control.
Immunodetection of endogenous pVHL isoforms in tumour tissues. (A) Histo-pathological characteristics of the ccRCC samples used for VHL immunodetection. (B) Western blot analysis of protein extracts from ccRCC tissue samples using the JD-1956 monoclonal antibody (1 : 100). β-tubulin was used as loading control. Expression of endogenous pVHL isoforms in HeLa cells was added (right panel) as a migration control of the three pVHL isoforms which are indicated by asterisks. The asterisks (*, **, ***) indicate pVHL213, pVHL172 and pVHL160, respectively. (C) Microtome sections of formalin-fixed and paraffin-embedded 786-0 cells (a), 786-0 cells overexpressing pVHL172 (b and c), normal kidney tissue (d) and ccRCC samples (e and f) were stained with (a, b, d and e) or without (c and f) the monoclonal antibody JD-1956 (1 : 100). Images were acquired with an Olympus microscope and processed with ImageJ 1.4 software. Scale bars, 20 μm (cells) and 50 μm (tumour sections). The arrows indicate the localization of VHL in the cytoplasm (d) and the nucleus (e).
Immunocytochemical and immunohistochemical detection of pVHL isoforms with the JD-1956 antibody
The JD-1956 antibody was further tested using IHC in cells that overexpress or not the pVHL172 isoform and in paraffin-embedded normal kidney and ccRCC tissue samples. The antibody JD-1956 revealed a strong signal in cells that overexpress pVHL172 (Figure 5Cb). Likewise, the antibody detected pVHL expression in epithelial cells of renal tubules in healthy kidney samples, mostly in the cytoplasm (Figure 5Cd). In ccRCC tumour samples, expression seemed to localise mainly to the cell membranes, more strongly in samples with del/mut VHL (panel e) than in normal tissue expressing wild-type VHL (Figure 5Cd). In few cells, VHL was detected both in the cytoplasm and the nucleus (arrow in Figure 5Ce). Control IHC performed on transfected cells as well as tumour tissue in the same experimental conditions but without incubation with primary antibody revealed no signal (Figure 5Cc and f, respectively).
Discussion
The function of pVHL172 isoform is still very elusive owing to the lack of analytic tools to detect its expression experimentally. Although detectable levels of mRNA can be measured in tissues, the relative abundance of this isoform was poorly documented in the literature as the use of current anti-VHL antibodies have resulted in conflicting results. Consequently, the role of pVHL172 in tumorigenesis and cancer progression is not clearly defined. Since commercially available and lab-made antibodies against pVHL do not allow to discriminate the isoforms generated by V1 and V2 mRNA variants (Supplementary Data S1), we decided to produce a new monoclonal antibody using human recombinant pVHL172 as antigen. However, due to the high similarity of the three isoforms (see Figure 1), raising an antibody that would be specific to the pVHL172 isoform is very challenging. Indeed, the results of this study show that the selected monoclonal antibody JD-1956 can detect all three isoforms (pVHL213, pVHL172 and pVHL160) by recognising a peptide sequence right after the internal initiation codon. When tested using recombinant proteins, the JD-1956 antibody detected equally well the three isoforms, whereas the others antibodies (a lab-made polyclonal antibody VHL-6030 and a commercial antibody from Santa Cruz Biotechnology) weakly recognised recombinant pVHL160. The antibody VHL-6030 was produced using a combination of two antigenic peptides and recognises preferentially the NH2-terminal part of the full-length protein, thus leading to the weak detection of the pVHL160 (Martin et al, 2013). As for the commercial antibody, it revealed some non-specific bands of higher molecular weight that would hamper the unambiguous detection of the pVHL isoforms, for instance, in whole cell extracts or tissue sections. Although not uniquely directed to pVHL172 isoform, JD-1956 is the first antibody that specifically and unequivocally recognises all pVHL isoforms. JD-1956 represents a valuable tool to accurately and reliable detect changes in VHL expression within tumours.
The detection of pVHL172 isoform, now amenable to a variety of complex biological samples, gave rise to some interesting observations. The significant difference between the apparent and the theoretical molecular weight of pVHL172 observed when using recombinant proteins produced in bacteria or protein overexpressed in mammalian cells could be explained by the presence of acidic repeats (eight copies of GXEEX) in the NH2-terminal part (codons 1 to 54) of the pVHL213 and pVHL172, but not in pVHL160. Furthermore, analysis of the amino acid composition of the different isoforms revealed that pVHL160 has an isoelectric point of 8.7 and pVHL172 of 4.7. Electrophysiological alterations and structural changes within pVHL isoforms might explain why they behave anomalously in SDS–PAGE.
The weaker detection of pVHL160 compared with the other isoforms by all antibodies in HeLa cells overexpressing the three pVHL proteins could be explained by pVHL160 amino acid composition that confers to the protein a strong hydrophobic behaviour. As a consequence, the detection of this isoform requires the use of strong dissociating agents, such as deoxycholate, that were not present in the buffer used for the preparation of the cell extracts, thus leading to an underestimation of the amount of pVHL160 in cells (Schoenfeld et al, 1998). This confirmed that the weaker immunodetection of pVHL160 in cells was not due to protein instability, but rather due to epitope accessibility (Schoenfeld et al, 1998).
We then tested the specificity of the JD-1956 antibody in embryonic and cancer kidney cell lines. Indeed, although the expression of the second mRNA variant has been reported in cells and tissues by different groups (including the present work), the expression of the encoded pVHL isoform (pVHL172) has never been substantiated due to the absence of a specific antibody. The JD-1956 antibody allowed highlighting the presence of a band that should correspond to pVHL172 both in embryonic kidney cells and in different cancer cell lines. The specificity of this detection was confirmed by VHL silencing using siRNAs specific for V2. The intensity of the bands corresponding to the different pVHL isoforms varied in the different cell lines. No signal was detected in 786-0 cells in which the VHL gene mutation exon 1 generates a very short and unstable protein (Iliopoulos et al, 1995). Similarly, other ccRCC cell lines (R-180 and R-305 cells) did not show any signal, confirming that mutations in the VHL gene can affect the expression and/or stability of the protein (Dugay et al, 2014). Conversely, in HeLa, RCC4 and HEK cells, different bands corresponding to pVHL213, pVHL172 and pVHL160 were observed. In MCF7 and HUVEC cells, only a smear signal corresponding to pVHL172 was detected, despite the confirmation of V2 mRNA presence. This indicates that the translation of V2 and/or the stability of the corresponding protein could be regulated differently in various cell lines. Finally, analysis of tumour tissue extracts led to the heterogeneous detection of the three bands corresponding to the pVHL isoforms. The signal variability could be explained by the heterogeneity of the tumour tissues.
In conclusion, we describe herein a new anti-pVHL antibody that allowed confirming, for the first time, the existence of the pVHL172 isoform translated from VHL V2 in different cell lines and tissues. In the last two decades, researchers have made significant contributions to the understanding of the functions of pVHL213/pVHL160; however, the interplay of the different isoforms, including pVHL172, in tumour development remains unsolved. Indeed, the VHL gene encodes a multifunctional protein with a crucial role in the ubiquitin-mediated degradation of HIF alpha. However, alternative functions, independent of HIF, have been identified and other interacting proteins need to be characterised. While it is clear that loss of pVHL can result in the activation of cellular pathways that are strongly associated with tumour initiation and progression, the concomitant presence of the three isoforms in some cells and tissues raises the question about their respective cellular functions and the reason for their concomitant presence. This new antibody represents an important tool for better understanding the tumour-suppressive function of pVHL, the specific role of pVHL172 and its critical targets in cancer.
Change history
14 July 2015
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Bader HL, Hsu T (2012) Systemic VHL gene functions and the VHL disease. FEBS Lett 586: 1562–1569.
Clark PE, Cookson MS (2008) The von Hippel-Lindau gene: turning discovery into therapy. Cancer 113: 1768–1778.
Dugay F, Le Goff X, Rioux-Leclerq N, Chesnel F, Jouan F, Henry C, Cabillic F, Verhoest G, Vigneau C, Arlot-Bonnemains Y, Belaud-Rotureau MA (2014) Overexpression of the polarity protein PAR-3 in clear cell renal cell carcinoma is associated with poor prognosis. Int J Cancer 134: 2051–2060.
Frank R (2002) The SPOT-synthesis technique. Synthetic peptide arrays on membrane supports—principles and applications. J Immunol Methods 267: 13–26.
Gnarra JR, Tory K, Weng Y, Schmidt L, Wei MH, Li H, Latif F, Liu S, Chen F, Duh FM, Lubensky I, Duan DR, Florence C, Pozzatti R, Walther MM, Bander NH, Grossman HB, Brauch H, Pomer S, Brooks JD, Isaacs WB, Lerman MI, Zbar B, Linehan WM (1994) Mutations of the VHL tumour suppressor gene in renal carcinoma. Nat Genet 7: 85–90.
Herman JG, Latif F, Weng Y, Lerman MI, Zbar B, Liu S, Samid D, Duan DS, Gnarra JR, Linehan WM (1994) Silencing of the VHL tumor-suppressor gene by DNA methylation in renal carcinoma. Proc Natl Acad Sci USA 91: 9700–9704.
Hsu T (2012) Complex cellular functions of the von Hippel-Lindau tumor suppressor gene: insights from model organisms. Oncogene 31: 2247–2257.
Iliopoulos O, Kibel A, Gray S, Kaelin WG Jr (1995) Tumour suppression by the human von Hippel-Lindau gene product. Nat Med 1: 822–826.
Iliopoulos O, Ohh M, Kaelin WG Jr (1998) pVHL19 is a biologically active product of the von Hippel-Lindau gene arising from internal translation initiation. Proc Natl Acad Sci USA 95: 11661–11666.
Latif F, Tory K, Gnarra J, Yao M, Duh FM, Orcutt ML, Stackhouse T, Kuzmin I, Modi W, Geil L, Schmidt L, Zhou F, Li H, Wei MH, Chen F, Glenn G, Choyke P, Walther MM, Weng Y, Duan DSR, Dean M, Glavac D, Richards FM, Crossey PA, Ferguson-Smith MA, Le Paslier D, Chumakov I, Cohen D, Chinault AC, Maher ER, Linehan WM, Zbar B, Lerman MI (1993) Identification of the von Hippel-Lindau disease tumor suppressor gene. Science 260: 1317–1320.
Lee S, Chen DY, Humphrey JS, Gnarra JR, Linehan WM, Klausner RD (1996) Nuclear/cytoplasmic localization of the von Hippel-Lindau tumor suppressor gene product is determined by cell density. Proc Natl Acad Sci USA 93: 1770–1775.
Martella M, Salviati L, Casarin A, Trevisson E, Opocher G, Polli R, Gross D, Murgia A (2006) Molecular analysis of two uncharacterized sequence variants of the VHL gene. J Hum Genet 51: 964–968.
Martin B, Chesnel F, Delcros JG, Jouan F, Couturier A, Dugay F, Le Goff X, Patard JJ, Fergelot P, Vigneau C, Rioux-Leclerq N, Arlot-Bonnemains Y (2013) Identification of pVHL as a novel substrate for Aurora-A in clear cell renal cell carcinoma (ccRCC). PloS One 8: e67071.
Patard JJ, Rioux-Leclercq N, Masson D, Zerrouki S, Jouan F, Collet N, Dubourg C, Lobel B, Denis M, Fergelot P (2009) Absence of VHL gene alteration and high VEGF expression are associated with tumour aggressiveness and poor survival of renal-cell carcinoma. Br J Cancer 101: 1417–1424.
Richards FM, Schofield PN, Fleming S, Maher ER (1996) Expression of the von Hippel-Lindau disease tumour suppressor gene during human embryogenesis. Hum Mol Genet 5: 639–644.
Robinson CM, Ohh M (2014) The multifaceted von Hippel-Lindau tumour suppressor protein. FEBS Lett 588: 2704–2711.
Schoenfeld A, Davidowitz EJ, Burk RD (1998) A second major native von Hippel-Lindau gene product, initiated from an internal translation start site, functions as a tumor suppressor. Proc Natl Acad Sci USA 95: 8817–8822.
Shuin T, Kondo K, Torigoe S, Kishida T, Kubota Y, Hosaka M, Nagashima Y, Kitamura H, Latif F, Zbar B, Lerman MI, Yao M (1994) Frequent somatic mutations and loss of heterozygosity of the von Hippel-Lindau tumor suppressor gene in primary human renal cell carcinomas. Cancer Res 54: 2852–2855.
Taylor C, Craven RA, Harnden P, Selby PJ, Banks RE (2012) Determination of the consequences of VHL mutations on VHL transcripts in renal cell carcinoma. Int J Oncol 41: 1229–1240.
Whaley JM, Naglich J, Gelbert L, Hsia YE, Lamiell JM, Green JS, Collins D, Neumann HP, Laidlaw J, Li FP, Klein-Szanto AJP, Seizinger BR, Kley N (1994) Germ-line mutations in the von Hippel-Lindau tumor-suppressor gene are similar to somatic von Hippel-Lindau aberrations in sporadic renal cell carcinoma. Am J Hum Genet 55: 1092–1102.
Acknowledgements
We would like to acknowledge the Biosit Biogenouest-Inserm histopathology platform H2P2 for immunohistochemistry analysis and the SFR Biosit CNRS UMS3480 MRic microscopy platform for fluorescence microscopy. The Bretagne Region, the INCA and the LCC supported this study. We acknowledge C Tascon (SIM Team-IGDR) for his participation in protein purification and S Dreano for his contribution in plasmids sequencing.
Authors contributions
All authors contributed to the conception and design, acquisition of data or analysis and interpretation of data.
Author information
Authors and Affiliations
Corresponding author
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Chesnel, F., Hascoet, P., Gagné, J. et al. The von Hippel–Lindau tumour suppressor gene: uncovering the expression of the pVHL172 isoform. Br J Cancer 113, 336–344 (2015). https://doi.org/10.1038/bjc.2015.189
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2015.189
- Springer Nature Limited
Keywords
This article is cited by
-
Mutation of the proline P81 into a serine modifies the tumour suppressor function of the von Hippel–Lindau gene in the ccRCC
British Journal of Cancer (2022)
-
Epigenetic crosstalk between hypoxia and tumor driven by HIF regulation
Journal of Experimental & Clinical Cancer Research (2020)
-
Insights into the molecular features of the von Hippel–Lindau-like protein
Amino Acids (2019)
-
Novel interactions of the von Hippel-Lindau (pVHL) tumor suppressor with the CDKN1 family of cell cycle inhibitors
Scientific Reports (2017)
-
VHLdb: A database of von Hippel-Lindau protein interactors and mutations
Scientific Reports (2016)