Abstract
The epidermal growth factor receptor (EGFR) is a clinically validated target in head and neck squamous cell carcinoma (HNSCC), where EGFR-blocking antibodies are approved for first-line treatment. However, as with other targeted therapies, intrinsic/acquired resistance mechanisms limit efficacy. In the FaDu HNSCC xenograft model, we show that combined blockade of EGFR and ERBB3 promotes rapid tumor regression, followed by the eventual outgrowth of resistant cells. RNA sequencing revealed that resistant cells express FGFR3-TACC3 fusion proteins, which were validated as drivers of the resistant phenotype by several approaches, including CRISPR-mediated inactivation of FGFR3-TACC3 fusion genes. Interestingly, analysis of signaling in resistant cell lines demonstrated that FGFR3-TACC3 fusion proteins promote resistance by preferentially substituting for EGFR/RAS/ERK signaling rather than ERBB3/PI3K/AKT signaling. Furthermore, although FGFR3-TACC3 fusion proteins promote resistance of additional EGFR-dependent HNSCC and lung cancer cell lines to EGFR blockade, they are unable to compensate for inhibition of PI3K signaling in PIK3CA-mutant HNSCC cell lines. Validation of FGFR3-TACC3 fusion proteins as endogenous drivers of resistance in our screen provides strong evidence that these fusions are capable of substituting for EGFR signaling. Thus, FGFR3-TACC3 fusion proteins may represent a novel mechanism of acquired resistance in EGFR-dependent cancers of multiple cell lineages.
Similar content being viewed by others
Introduction
Inhibitors of epidermal growth factor receptor (EGFR) signaling are approved for the treatment of multiple human cancers. For example, EGFR tyrosine kinase inhibitors (TKIs) are used to treat non-small cell lung cancer patients who have activating mutations in the EGFR kinase domain.1, 2 In addition, antibodies that block the binding of ligands to EGFR are used in KRAS wild-type (WT) colorectal cancer and in head and neck squamous cell carcinoma (HNSCC).3, 4, 5 However, the efficacy of EGFR inhibitors, as with other targeted therapies, is limited by multiple mechanisms of intrinsic and acquired resistance.6, 7
Signaling by the ERBB family member ERBB3 has been identified in recent years as a prominent mechanism of resistance to targeted therapies in several tumor types.8, 9 For example, preclinical studies from our group and others have demonstrated that ERBB3 antibodies can potentiate the effects of EGFR blockade in colorectal cancer and HNSCC models,10, 11, 12, 13, 14 most likely by inhibiting ERBB3-dependent activation of the PI3K/AKT survival pathway.15, 16, 17 On the basis of this preclinical rationale for blocking ERBB3 in human cancer, several ERBB3 antibodies, including our antibody REGN1400, have progressed into the clinic.10, 12, 13, 18, 19, 20
While combined blockade of EGFR/ERBB3 can have potent effects in tumor xenograft models, it is likely that there are resistance mechanisms that can limit the benefit of this combination. In this study, we perform an in vivo selection for FaDu HNSCC cells that are resistant to EGFR/ERBB3 blockade and demonstrate that FGFR3-TACC3 fusion proteins are major drivers of the resistant phenotype. We show that, although FGFR3-TACC3 fusion proteins can promote resistance to EGFR blockade in multiple cancer cell lines, apparently via strong activation of ERK signaling, they are not able to promote resistance of PIK3CA-mutant HNSCC cell lines to PI3K inhibition. This report is the first to identify FGFR3-TACC3 fusion proteins as a natural mechanism of resistance to blockade of ERBB family receptors, and it provides important insight into the functional capabilities of these fusion proteins. In addition, our findings suggest the possibility that combined EGFR/FGFR3 blockade might be an effective therapy in a subset of HNSCC patients.
Results
Head and neck cancer cells selected for resistance to EGFR/ERBB3 blockade express activated FGFR3-TACC3 fusion proteins
To discover mechanisms that mediate acquired resistance of FaDu tumors to combined blockade of EGFR/ERBB3, we generated variant cell lines that exhibit complete resistance to this treatment, as outlined in Figure 1. FaDu tumors treated with the combination of the EGFR antibody REGN955 plus the ERBB3 antibody REGN140013 shrink and become virtually undetectable, suggesting that the majority of the tumor cells have undergone apoptosis (Figure 1a, left panel). Eventually, however, individual tumors can regrow (Figure 1a, middle panel). These regrowing tumors were harvested, fragmented and re-passaged in vivo under drug treatment (Figure 1a, right panels). After being re-passaged twice in vivo, tumors that exhibited resistance to combined EGFR/ERBB3 blockade were used to generate cell lines. Tumors formed from two such variant cell lines, called FaDu V1 and FaDu V2, exhibited complete resistance to EGFR/ERBB3 blockade (Figures 1b and c). We also generated a control parental cell line, called FaDu P1, in which tumor fragments were similarly re-passaged in vivo, except that the mice were treated with control protein human Fc. The FaDu P1 cell line was the comparator cell line for the subsequent genetic and biochemical characterization of the resistant variants.
Generation of FaDu cell lines resistant to EGFR/ERBB3 blockade. (a) Severe combined immunodeficiency (SCID) mice bearing established FaDu tumors (~200 mm3 in volume) were randomized and treated continuously with control antibody (12.5 mg/kg), REGN1400 (ERBB3-blocking antibody; 2.5 mg/kg), REGN955 (EGFR-blocking antibody; 10 mg/kg) or the combination of REGN1400 plus REGN955. The line graph depicts the average tumor volumes over the course of treatment. Error bars show the s.d. A tumor in a combination-treated mouse that began to regrow at ~110 days after implantation (middle panel) was harvested and fragments of this tumor were re-implanted into mice. A tumor fragment that grew rapidly in the face of REGN1400 plus REGN1955 combination treatment was harvested (top right panel shows the growth of individual re-implanted fragments) and the re-implantation and treatment procedure was repeated. Finally, a tumor growing rapidly under combined EGFR/ERBB3 blockade was harvested (bottom right panel) and used to generate a resistant cell line. (b and c) Cultured FaDu V1 or V2 cells were implanted into SCID mice to generate tumors. Mice bearing established tumors were randomized and treated twice per week with control antibody or Fc protein (12.5 mg/kg), REGN1400 (2.5 mg/kg), REGN955 (10 mg/kg) or the combination of REGN1400 plus REGN955. The line graphs depict the average tumor volumes over the course of treatment. Error bars show the s.d.
To exclude a trivial explanation for the resistance of the FaDu V1 and V2 cell lines to EGFR/ERBB3 blockade, we demonstrated that REGN955 and REGN1400 still bind and block their respective targets in these cells (Supplementary Figure 1), confirming that the variants do not express mutated versions of EGFR or ERBB3 that can no longer be inhibited by these antibodies. To investigate the molecular basis for the resistance of the variant cell lines, the ability of REGN955 and REGN1400 to inhibit growth of these cells in vitro was assessed. Combined blockade of EGFR plus ERBB3 inhibited the growth of FaDu P1 parental cells by ~80% (as shown previously13) while only inhibiting growth of FaDu V1 and V2 cells by ~25% (Figures 2a–c), indicating that the mechanisms promoting in vivo resistance of these cell lines are largely operative in vitro as well. Interestingly, in both FaDu V1 and V2 cell lines, what was most different from the parental cells was the response to the EGFR-blocking antibody, which was able to significantly inhibit growth of parental cells (~40% inhibition) but had almost no effect (only 5–10% inhibition) in the variant cell lines (Figures 2a–c). In contrast, the effect of the ERBB3-blocking antibody was similar in the parental and variant cell lines (Figures 2a–c).
EGFR/ERBB3 blockade fails to inhibit ERK activation and cell growth in FaDu-resistant variant cell lines. (a–c) FaDu P1, V1 or V2 cells were grown for 72 h in the presence of control antibody (15 μg/ml), REGN1400 (5 μg/ml), REGN955 (10 μg/ml) or the combination of REGN1400 plus REGN955. The bar graphs show the relative cell growth in each treatment group, as determined by MTS assay. Error bars show the s.d., n=8. Cell growth was compared by one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test (***P<0.001; for comparisons with the control group, asterisks are shown only when the inhibition is >15%). (d) FaDu P1, V1 or V2 cells were treated for 2 h with control antibody (15 μg/ml), REGN1400 (5 μg/ml), REGN955 (10 μg/ml) or the combination of REGN1400 plus REGN955. Following treatment, cell lysates were subjected to western blot with antibodies against phospho-AKT, AKT, phospho-ERK and ERK. (e) FaDu V2 cells were treated for 2 h with control antibody (5 μg/ml) plus vehicle, REGN1400 (5 μg/ml), MEK inhibitor GSK1120212 (100 nM) or the combination of REGN1400 plus GSK1120212. Following treatment, cell lysates were subjected to western blot analysis with antibodies against phospho-AKT, AKT, phospho-ERK and ERK. (f) FaDu V2 cells were grown for 72 h in the presence of control antibody (5 μg/ml) plus vehicle, REGN1400 (5 μg/ml), MEK inhibitor GSK1120212 (100 nM) or the combination of REGN1400 plus GSK1120212. The bar graphs show the relative cell growth in each treatment group, as determined by MTS assay. Error bars show the s.d., n=8. Cell growth was compared by one-way ANOVA with Tukey’s multiple comparison test (***P<0.001).
To assess whether the relatively weak effect of EGFR/ERBB3 blockade on the growth of FaDu V1 and V2 cells reflected a failure to inhibit downstream signaling pathways, we tested the effects of EGFR/ERBB3 blockade on AKT and ERK activation. In FaDu cells, blockade of EGFR primarily inhibits activation of the ERK pathway, whereas ERBB3 blockade primarily inhibits activation of the AKT pathway,13 likely explaining the superior efficacy of the combination treatment. Interestingly, in both FaDu V1 and V2 cells, REGN1400 inhibited AKT activation as effectively as it did in FaDu P1 cells (Figure 2d). However, neither REGN955 nor the combination of REGN955 plus REGN1400 was able to effectively inhibit ERK activation in FaDu V1 or V2 cells, in contrast to the almost complete ERK inhibition observed in FaDu P1 cells (Figure 2d). Thus, despite the ability of REGN955 to effectively inhibit EGFR in the variant cell lines (Supplementary Figure 1), the antibody was unable to block downstream ERK activation. Consistent with the possibility that sustained activation of the MAP kinase pathway upon EGFR blockade is a key element of the resistant phenotype, combined treatment with REGN1400 plus the MEK inhibitor GSK1120212 (trametinib, GlaxoSmithKline (GSK)) effectively blocked both AKT and ERK phosphorylation in FaDu V2 cells (Figure 2e) and inhibited cell growth by ~70% (Figure 2f), similar to the effect of combined EGFR/ERBB3 blockade on the growth of parental FaDu cells.
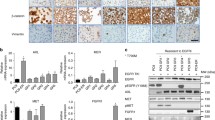
These findings suggested the possibility that another receptor tyrosine kinase (RTK), not active in FaDu P1 parental cells, maintains ERK signaling in the FaDu V1 and V2 cell lines when EGFR is blocked. Thus, we used a phospho-RTK array to assess the activation status of all RTKs in FaDu P1, V1 and V2 cells. As shown previously,13 parental FaDu cells exhibit activation of EGFR, HER2 and ERBB3 (Figure 3a). These RTKs remained active in FaDu V1 and V2 cells, but both of the resistant cell lines also exhibited FGFR3 phosphorylation, which was not detectable in parental cells (Figure 3a). In addition, FaDu V2 cells exhibited much stronger activation of MET than FaDu P1 or FaDu V1 cells (Figure 3a). Western blot analysis of whole-cell lysates confirmed the increased MET phosphorylation in FaDu V2 cells (Figure 3b). Immunoprecipitation with anti-phosphotyrosine antibody followed by western blot analysis for FGFR3 confirmed the presence of activated FGFR3 in both FaDu V1 and V2 cells, but not FaDu P1 cells (with a higher level of phospho-FGFR3 present in FaDu V2 cells; Figure 3c).
FGFR3 is activated in FaDu-resistant variant cell lines and maintains ERK signaling upon EGFR blockade. (a) Lysates were prepared from FaDu P1, V1 or V2 cells and used to assess tyrosine phosphorylation of 49 human RTKs with the Human Phospho-RTK Array Kit. Active RTKs of note are boxed and labeled. The unlabeled spots on the corners of the membranes are positive controls. (b) Lysates from FaDu P1, V1 or V2 cells were subjected to western blot analysis with antibodies against phospho-MET, MET and actin. (c) Lysates from FaDu P1, V1 or V2 cells were subjected to immunoprecipitation with anti-phosphotyrosine (p-Tyr) antibody 4G10 conjugated to agarose beads. The presence of FGFR3 and Src (a positive control) in the immunoprecipitates was assessed by western blot analysis. (d) FaDu V2 cells were treated for 30 min with control antibody (10 μg/ml) plus vehicle, REGN955 (10 μg/ml), 100 nM PHA665752 (a MET tyrosine kinase inhibitor) or the combination of REGN955 plus PHA665752. Following treatment, cell lysates were subjected to western blot analysis with antibodies against phospho-ERK, ERK, phospho-MET and MET. (e) FaDu V2 cells were treated for 1 h with vehicle or with 25 nM AZD4547, a pan-FGFR tyrosine kinase inhibitor. Following treatment, cell lysates were subjected to immunoprecipitation with p-Tyr antibody 4G10 conjugated to agarose beads. The presence of FGFR3 and Src in the immunoprecipitates was assessed by western blot analysis. (f) FaDu V1 or V2 cells were treated for 30 min with control antibody (10 μg/ml) plus vehicle, REGN955 (10 μg/ml), 25 nM AZD4547 or the combination of REGN955 plus AZD4547. Following treatment, cell lysates were subjected to western blot analysis with antibodies against phospho-ERK, ERK, phospho-AKT and AKT.
In FaDu V2 cells, the MET TKI PHA665752, as a single agent or in combination with REGN955, failed to inhibit ERK activation, despite completely blocking MET phosphorylation (Figure 3d). Thus, the failure of REGN955 to inhibit ERK is not a result of increased MET activation.
To assess the role of FGFR3 in maintaining ERK activation, we employed the selective pan-FGFR TKI (AZD4547), which completely blocked FGFR3 phosphorylation in FaDu V2 cells (Figure 3e). In both FaDu V1 and V2 cells, the combination of AZD4547 plus REGN955 completely inhibited ERK activation, whereas the single agents had either no effect or a partial effect (Figure 3f). Thus, FGFR3 signaling is necessary to maintain ERK activation in FaDu variant cells when EGFR is blocked. AZD4547 had no effect on AKT activation, either alone or in combination with REGN955 (Figure 3f), indicating that FGFR3 signaling is not required for activation of AKT in FaDu variant cells. This finding is consistent with the observation that inhibition of ERBB3 alone results in almost complete loss of activated AKT in these cells (Figure 2d).
Activation of FGFR3 in the FaDu variant cell lines could result from either increased ligand-dependent stimulation of FGFR3 or from a genetic alteration of FGFR3. For example, activating point mutations of FGFR3 have been identified in multiple cancers, most prominently in bladder cancer.21 We therefore performed RNA sequencing (RNA-seq) to identify genetic alterations of FGFR3 and/or of other genes in the FaDu variant cell lines that might underlie the resistant phenotype. Consistent with the presence of activated FGFR3 in the resistant cell lines, we identified FGFR3-TACC3 fusion transcripts in both FaDu V1 and V2 cells (each cell line expressed a distinct fusion transcript) but not in parental FaDu cells. FGFR3-TACC3 fusions have recently been identified in multiple human cancers, and in all cases these fusion proteins contain most of the FGFR3 protein, including the tyrosine kinase domain and the TACC3 coiled coil domain, suggesting that constitutive dimerization of the fusion proteins mediated by the TACC3 coiled coil domain underlies FGFR3 kinase activation.22, 23, 24, 25 The fusion transcripts identified in FaDu V1 and V2 cells are similar to those previously reported (Figure 4a; see Supplementary Figures S2 and S3 for the RNA-seq reads supporting the fusion transcripts and for the chromosomal coordinates of the breakpoints). RT–PCR (with primers flanking the putative fusion junctions) followed by Sanger sequencing of the PCR products confirmed the presence of the respective fusion transcripts in FaDu V1 and V2 cells and confirmed the junction sequences (Figure 4b and Supplementary Figure S4). Consistent with this finding, quantitative real-time PCR revealed significant expression of the respective fusion transcripts in FaDu V1 and V2 cells, but not in parental FaDu cells, where these transcripts were undetectable (Figure 4c).
FaDu variant cell lines express constitutively active FGFR3-TACC3 fusion proteins. (a) Diagram of the structure of the FGFR3-TACC3 fusion proteins that were identified in FaDu V1 and V2 cells. (b) Overall, 100 ng of cDNA from FaDu P1, V1 or V2 cells was subjected to PCR with primers that flank the FGFR3-TACC3 fusion junctions identified by RNA-seq. As a control for the integrity of the cDNA, a fragment of the cyclophilin gene was amplified from all samples. Aliquots of the PCR reactions were run on a 2% agarose gel (M, molecular weight markers) and the fragments of the FGFR3-TACC3 fusion transcripts (expected PCR products are 122 bp (V1 cells) and 95 bp (V2 cells)) were gel-purified and subjected to Sanger sequencing. The nucleotide and amino-acid sequences immediately flanking the FaDu V2 fusion junction and the corresponding sequence trace are shown (see Supplementary Figure 4 for the sequence data for the FaDu V1 fusion junctions). The sequence GTG in black text represents shared sequence at the fusion junction. (c) RNA from FaDu P1, V1 or V2 cells was subjected to TaqMan real-time PCR analysis using primers/probe sets specific for the FGFR3-TACC3 fusion transcripts. For each sample, the threshold cycle (Ct) value for the control gene cyclophilin was subtracted from the Ct value for the FGFR3-TACC3 fusion transcript to give the delta Ct (ΔCt) value. The bars show the average 2-ΔCt for each sample. Error bars show the s.d., n=3. (d) Lysates from FaDu P1, V1 or V2 cells were subjected to immunoprecipitation with a TACC3 antibody (recognizes an epitope near the C terminus of TACC3 that is present in the FGFR3-TACC3 fusions), followed by western blot analysis for FGFR3 or TACC3. In addition, aliquots of lysate from FaDu P1, V1 or V2 cells were directly subjected to western blot analysis (last three lanes). The FGFR3 western blot antibody recognizes both native FGFR3 and the FGFR3-TACC3 fusion proteins. The fusions migrate slightly above native FGFR3 (in this experiment, proteins were resolved on a 4% SDS gel to maximize the separation). (e) Lysates from FaDu P1, V1 or V2 cells were subjected to immunoprecipitation with p-Tyr antibody 4G10 conjugated to agarose beads. The presence of TACC3 and Src (a positive control) in the immunoprecipitates was assessed by western blot analysis. (f) Lysate from FaDu V2 cells was subjected to immunoprecipitation with p-Tyr antibody 4G10 conjugated to agarose beads. Multiple aliquots of the immunoprecipitate were run on a single SDS gel. Lysate from FaDu P1 parental cells was also run to show the migration of native FGFR3 and TACC3. Following transfer, the PVDF membrane was cut in half and western blot analyses were performed for either FGFR3 or TACC3. The two halves of the membrane were put back together for signal development and exposure, illustrating the identical migration of the tyrosine-phosphorylated proteins detected by the FGFR3 and TACC3 antibodies.
Western blotting of whole-cell lysates from FaDu parental and variant cell lines with an FGFR3 antibody revealed the presence of higher molecular weight FGFR3-containing proteins in both V1 and V2 cells compared with parental cells (Figure 4d, last three lanes). The larger FGFR3-containing proteins in FaDu V1 and V2 cells migrate at a molecular weight consistent with the fusion transcripts we identified, that is, ~20 kDa larger than native FGFR3 (which migrates at ~110–130 kDa). To confirm that the larger FGFR3-containing proteins expressed in the FaDu variant cell lines are the FGFR3-TACC3 fusions, cell lysates were subjected to immunoprecipitaton with an antibody that recognizes a C-terminal epitope in TACC3 that is present in the putative FGFR3-TACC3 fusion proteins. As shown in Figure 4d, the TACC3 antibody selectively immunoprecipitated the larger FGFR3-containing proteins from both FaDu V1 and V2 cells, but did not immunoprecipitate native FGFR3 from any of the cell lines. Native TACC3 was immunoprecipitated from all three cell lines, controlling for the immunoprecipitation procedure (Figure 4d, bottom panel). The FGFR3-TACC3 fusion proteins are expressed at much lower levels than native TACC3, explaining why only native TACC3 is visible in the exposure shown in the bottom panel of Figure 4d. Thus, TACC3 antibody is able to immunoprecipitate FGFR3-containing proteins specifically from the FaDu variant cell lines, confirming the expression of the FGFR3-TACC3 fusions.
As shown earlier in Figure 3c, FGFR3 was detected in anti-phosphotyrosine immunoprecipitates from both FaDu V1 and V2 cells, but not from parental cells. If these tyrosine-phosphorylated FGFR3-containing proteins are the FGFR3-TACC3 fusion proteins, they should also be detectable by western blot analysis with TACC3 antibody. As shown in Figure 4e, TACC3 (like FGFR3) was detected in anti-phosphotyrosine immunoprecipitates from both FaDu V1 and V2 cells, but not from FaDu P1 cells. The tyrosine-phosphorylated proteins from FaDu V2 cells recognized by the FGFR3 and TACC3 antibodies migrate identically in an SDS gel (Figure 4f), confirming that they are the same proteins.
Thus, the FaDu V1 and V2 cell lines, but not parental FaDu cells, express tyrosine-phosphorylated FGFR3-TACC3 fusion proteins that appear to maintain ERK signaling upon EGFR blockade and may therefore have a role in the resistant phenotype of these two cell lines.
FGFR3-TACC3 fusion proteins promote resistance of parental FaDu cells to EGFR/ERBB3 blockade
To assess the ability of the FGFR3-TACC3 fusion proteins identified in FaDu V1 and V2 cells to drive resistance, the fusion proteins were stably expressed in FaDu P1 parental cells. As controls, we also expressed WT FGFR3 and a kinase-dead FGFR3-TACC3 fusion. The cell lines were generated by lentiviral infection at low multiplicity of infection (0.3) to minimize overexpression due to multiple integrations. As shown in Figure 5a, strong expression of WT FGFR3 and the FGFR3-TACC3 fusion proteins was detected in stably transduced FaDu parental cells. Whereas WT FGFR3 was very weakly phosphorylated as assessed by western blot analysis with a phospho-FGFR antibody (compare WT FGFR3 with empty vector control), both of the FGFR3-TACC3 fusion proteins were phosphorylated to a much greater extent (Figure 5a), indicating a higher level of constitutive activity. As expected, kinase-dead fusion protein was not tyrosine-phosphorylated (Figure 5a). Western blot analysis for TACC3 confirmed the presence of TACC3 sequence in the stably expressed FGFR3-TACC3 fusion proteins (Figure 5a). At the exposure shown in Figure 5a, expression of endogenous FGFR3 was undetectable in FaDu parental cells transduced with empty vector, indicating that the lentivirally encoded proteins are in fact overexpressed. However, analysis of changes in downstream signaling using a phospho-kinase array revealed that expression of the FGFR3-TACC3 fusions did not promote a general rewiring of the signaling pathways in parental FaDu cells; in fact, few changes were observed (Supplementary Figure S5).
FGFR3-TACC3 fusion proteins promote resistance to EGFR/ERBB3 blockade. (a) Parental FaDu cells were infected with an empty vector control lentivirus or with lentiviruses encoding wild-type FGFR3, the FGFR3-TACC3 fusion proteins (F3-T3) identified in the FaDu variants or a kinase-dead version of the V2 FGFR3-TACC3 fusion, and stable cell lines were generated. Cell lysates were prepared and subjected to western blot analysis with antibodies against FGFR3, phospho-FGFR, TACC3 or Actin. (b) Lysates were prepared from parental FaDu cells expressing wild-type FGFR3 or FGFR3-TACC3 fusion proteins (F3-T3) and were subjected to immunoprecipitation with anti-p-Tyr antibody 4G10 conjugated to agarose beads. The presence of FGFR3, TACC3 and Src in the immunoprecipitates was assessed by western blot analysis. (c) Parental FaDu cells expressing wild-type FGFR3 or FGFR3-TACC3 fusion protein (from V2 cells) were treated for 2 h with control antibody (15 μg/ml), REGN1400 (5 μg/ml) or REGN955 (10 μg/ml). Cell lysates were prepared and subjected to western blot analysis with antibodies against phospho-ERK, ERK, phospho-AKT or AKT. (d) Parental FaDu cells expressing wild-type FGFR3, FGFR3-TACC3 fusion proteins or kinase-dead FGFR3-TACC3 fusion were grown for 72 h in the presence of control antibody (15 μg/ml), REGN1400 (5 μg/ml), REGN955 (10 μg/ml) or the combination of REGN1400 plus REGN955. The bar graphs show the relative cell growth in each treatment group, as determined by MTS assay. Error bars show the s.d., n=8. Cell growth was compared by one-way ANOVA with Tukey’s multiple comparison test (***P<0.001; for comparisons with the control group, asterisks are shown only when the inhibition is >15%). (e) Parental FaDu cells expressing wild-type FGFR3 or FGFR3-TACC3 fusion protein (from FaDu V2 cells), or transduced with empty vector, were implanted into SCID mice. Mice bearing established tumors were randomized and treated twice per week with control antibody (12.5 mg/kg) or the combination of REGN1400 (2.5 mg/kg) plus REGN955 (10 mg/kg). The line graphs depict the average tumor volumes over the course of treatment. Error bars show the s.d.
Immunoprecipitation of cell lysates with a phosphotyrosine antibody confirmed the increased phosphorylation of the FGFR3-TACC3 fusion proteins compared with WT FGFR3 (Figure 5b). Consistent with the observation that EGFR blockade fails to inhibit ERK activation in FaDu variant cells (Figure 2d), expression of FGFR3-TACC3 fusion protein, but not WT FGFR3, prevented REGN955 from significantly inhibiting ERK activation in FaDu parental cells (Figure 5c), confirming that the fusion protein drives strong activation of the ERK pathway. In contrast, the FGFR3-TACC3 fusion protein did not prevent REGN1400 from inhibiting AKT activation (Figure 5c), consistent with the fact that REGN1400 effectively blocks this pathway in FaDu V1 and V2 cells (Figure 2d).
To determine whether expression of FGFR3-TACC3 fusion proteins is sufficient to drive resistance, parental FaDu cells expressing WT FGFR3 or the fusion proteins were treated with REGN1400 plus REGN955. Whereas parental cells expressing either of the FGFR3-TACC3 fusion proteins were resistant to growth inhibition, cells expressing WT FGFR3 or kinase-dead FGFR3-TACC3 fusion remained sensitive (Figure 5d). Thus, the ability of FGFR3-TACC3 fusion proteins to promote resistance of FaDu cells is dependent on FGFR3 kinase activity, consistent with a recent report showing that oncogenic function of FGFR3-TACC3 fusions requires an active FGFR3 kinase.26
Consistent with the ability of the fusion proteins to drive resistance in vitro, the FGFR3-TACC3 fusion protein from FaDu V2 cells, but not WT FGFR3, was sufficient to promote resistance of FaDu parental tumor xenografts to combined EGFR/ERBB3 blockade (Figure 5e). The faster growth of the tumors expressing the FGFR3-TACC3 fusion protein may reflect the fact that these cells exhibit increased baseline ERK activation compared with cells expressing WT FGFR3 (data not shown). Finally, we show that parental FaDu tumors engineered to overexpress FGF1 ligand exhibit complete resistance to EGFR/ERBB3 blockade in vivo (Supplementary Figure S6), indicating that ligand-activated FGFR3 can also drive resistance (although in this experiment we cannot rule out contributions from FGFR2 or FGFR4, which can be activated by FGF1 and which are expressed in FaDu cells).
FGFR3-TACC3 fusion proteins are required for resistance of FaDu variants
To assess whether the endogenous FGFR3-TACC3 fusion proteins expressed in the FaDu variants are responsible for the resistant phenotype, we employed CRISPR/Cas9 technology27 to inactivate the FGFR3-TACC3 fusion genes. We used lentivirus to deliver Cas9 nuclease and single guide RNAs (sgRNAs) to FaDu V1 and V2 cells. Four sgRNAs targeting early exons in FGFR3 (exon 2 or exon 3) were tested, and two of the four sgRNAs almost completely eliminated expression of the FGFR3-TACC3 fusion proteins (and native FGFR3) in FaDu V1 and V2 cells (Figure 6a). Consistent with our previous finding that combined treatment with FGFR TKI and EGFR antibody REGN955 effectively blocks ERK activation in resistant cells (Figure 3f), CRISPR-mediated inactivation of FGFR3-TACC3 fusion genes enabled REGN955 to inhibit ERK activation in both the V1 and V2 cell lines (Figure 6b), establishing that signaling by these fusion proteins maintains ERK activation when EGFR is blocked.
FGFR3-TACC3 fusion proteins are required for the resistant phenotype of FaDu variant cell lines. (a) FaDu V1 or V2 cells were infected with lentiviruses expressing the Cas9 nuclease alone (control) or expressing Cas9 plus a sgRNA specific for FGFR3 (sgRNAs 1 and 2 target distinct sequences within the FGFR3 gene). At 10 days after infection, the levels of FGFR3-TACC3 fusion proteins (F3-T3) were assessed by western blot analysis. (b) FaDu V1 or V2 cells stably expressing Cas9 nuclease (control) or Cas9 nuclease plus FGFR3 sgRNA 1 were treated with control antibody (10 μg/ml), REGN1400 (5 μg/ml) or REGN955 (10 μg/ml) for 2 h. Following treatment, cell lysates were subjected to western blot analysis with antibodies against phospho-ERK and ERK. (c) FaDu V1 or V2 cells stably expressing Cas9 nuclease (control) or Cas9 nuclease plus FGFR3 sgRNA 1 or FGFR3 sgRNA 2 were grown for 72 h in the presence of control antibody (15 μg/ml), REGN1400 (5 μg/ml), REGN955 (10 μg/ml) or the combination of REGN1400 plus REGN955. The bar graphs show the relative cell growth in each treatment group, as determined by MTS assay. Error bars show the s.d., n=8. Cell growth was compared by one-way ANOVA with Tukey’s multiple comparisons test (***P<0.001; for comparisons with the control group, asterisks are shown only when the inhibition is >15%). (d) Model depicting the role of FGFR3-TACC3 fusion proteins in resistance of FaDu variant cell lines. Constitutively active FGFR3-TACC3 fusions drive strong activation of the RAS/RAF/ERK pathway, functionally substituting for EGFR signaling. The model is discussed in detail in the text.
Consistent with this observation, CRISPR-mediated inactivation of FGFR3-TACC3 with either sgRNA 1 or sgRNA 2 enabled REGN955 to inhibit growth of FaDu V1 and V2 cells as a single agent and to significantly potentiate the modest growth inhibition mediated by REGN1400 (Figure 6c). In CRISPR-modified FaDu V1 and V2 cells, the magnitude of the growth inhibition mediated by the combination of REGN955 plus REGN1400 was similar to that observed in parental FaDu cells (Figure 2a). Thus, although we cannot exclude the involvement of additional resistance mechanisms in the FaDu variants, our data indicate that a substantial component of the resistant phenotype is attributable to signaling by FGFR3-TACC3 fusion proteins (see Figure 6d for a model).
FGFR3-TACC3 fusion proteins promote resistance to targeted therapy in cancer cell lines driven by EGFR, but not by mutated PI3K
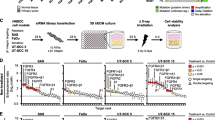
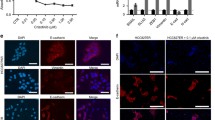
To further investigate the functional capabilities of FGFR3-TACC3 fusion proteins, we assessed their ability to promote resistance of additional cancer cell lines to targeted therapies. We employed cancer cell lines driven by EGFR/ERBB3 signaling (Cal27 HNSCC), mutated EGFR (NCI-H1975 non-small cell lung cancer) or mutated PIK3CA (SNU1076 and Detroit 562 HNSCC), as recent genomic data show that the PIK3CA gene is frequently mutated in HNSCC, suggesting that PI3K is an important driver in this indication.28 As in FaDu parental cells, FGFR3-TACC3 fusion proteins (but not WT FGFR3) were able to promote resistance of Cal27 cells to combined EGFR/ERBB3 blockade (Figure 7a) and of NCI-H1975 cells to the EGFR TKI AZD9291 (Figure 7b; see Supplementary Figure S7 for confirmation of the expression and phosphorylation of the fusion proteins in these cell lines).
FGFR3-TACC3 fusion proteins promote resistance in cancer cell lines driven by EGFR, but not by mutated PI3K. (a) Cal27 cells expressing wild-type FGFR3 or the FGFR3-TACC3 fusion proteins (F3-T3) identified in FaDu V1 or V2 cells were grown for 72 h in the presence of control antibody (15 μg/ml) or the combination of REGN1400 (5 μg/ml) plus REGN955 (10 μg/ml). The bar graphs show the relative cell growth in each treatment group, as determined by MTS assay. Error bars show the s.d., n=8 (applies to a–d). (b) NCI-H1975 cells expressing wild-type FGFR3 or FGFR3-TACC3 fusion proteins were grown for 72 h in the presence of vehicle or 50 nM EGFR TKI AZD9291, a third-generation irreversible TKI that inhibits the T790M EGFR mutant expressed in these cells. (c and d) SNU1076 or Detroit 562 cells expressing wild-type FGFR3 or FGFR3-TACC3 fusion proteins were grown for 72 h in the presence of vehicle or 5 μM PI3K inhibitor BYL719. (e) Cell lysates were prepared from Detroit 562 or SNU1076 cells stably expressing wild-type FGFR3 or FGFR3-TACC3 fusion proteins and subjected to western blot analysis with antibodies against FGFR3, phospho-FGFR or TACC3. (f) Control SNU1076 or Detroit 562 cells or cells expressing FGFR3-TACC3 fusion protein (from FaDu V1) were treated with vehicle or 5 μM BYL719 for 60 min. Cell lysates were prepared and subjected to western blot analysis with antibodies against phospho-ERK, ERK, phospho-AKT or AKT.
In contrast, neither of the FGFR3-TACC3 fusion proteins was able to confer substantial resistance of SNU1076 or Detroit 562 cells to the PI3K inhibitor BYL719 (alpelisib, Novartis, Basel, Switzerland) (Figures 7c and d), despite high expression and phosphorylation of the fusions (Figure 7e). Similar to our observations in FaDu cells, the FGFR3-TACC3 fusion protein strongly activated ERK signaling in both of these cell lines, either fully restoring ERK signaling in the presence of the PI3K inhibitor (SNU1076 cells) or substantially increasing the baseline level of ERK activation (Detroit 562 cells; Figure 7f). The FGFR3-TACC3 fusion protein did not restore AKT activation upon PI3K blockade (Figure 7f), although even if the FGFR3-TACC3 fusion was capable of activating AKT in these cells, the PI3K inhibitor would likely have prevented it. Thus, strong activation of ERK signaling by FGFR3-TACC3 fusion protein does not compensate for loss of PI3K/AKT signaling in HNSCC cells ‘addicted’ to the PI3K pathway, suggesting that FGFR3-TACC3 fusion proteins are unlikely to be relevant mediators of resistance to PI3K inhibitors in PIK3CA-mutant HNSCC.
Discussion
FGFR3-TACC3 fusion proteins have now been identified at low frequency in brain, bladder, lung and head and neck cancers.23, 24, 25, 29, 30, 31, 32, 33, 34 However, the functional capabilities of these fusions have not been fully elucidated. While glioma and bladder cancer cells expressing FGFR3-TACC3 fusion proteins are sensitive to FGFR TKIs,23, 25, 30, 35 data demonstrating that endogenous FGFR3-TACC3 fusions drive cancer cell growth/survival are still limited, as most studies have only assessed the function of ectopically expressed fusion proteins. Thus, our finding that the FGFR3-TACC3 fusion proteins endogenously expressed in the FaDu variants are mediators of the resistant phenotype adds significantly to the evidence that these fusion proteins are drivers in human cancer. In addition, our study provides important mechanistic insight into the capabilities of FGFR3-TACC3 fusions, clearly demonstrating that they can substitute for EGFR signaling and may therefore represent a novel mechanism of resistance to EGFR inhibitors.
Our finding that FGFR3-TACC3 fusion proteins preferentially substitute for EGFR/ERK signaling rather than for ERBB3/AKT signaling in FaDu cells is consistent with published studies that have shown that FGFR3 signaling can compensate for the blockade of EGFR and vice versa.36, 37, 38, 39 The fact that the FaDu variants are resistant to combined EGFR/ERBB3 blockade despite the inability of the fusion proteins to maintain AKT activation could be attributable to activation of other downstream pathways by the fusion proteins or to mechanisms unrelated to these fusions.
Interestingly, in bladder cancer, activating FGFR3 point mutations and RAS mutations appear to be mutually exclusive,40 consistent with the observation that mutated FGFR3 drives activation of RAS/ERK signaling.41 In contrast, FGFR3 mutations in bladder cancer do co-occur with activating mutations in PIK3CA,42, 43 suggesting that FGFR3 and PI3K provide complementary signals. These studies are consistent with our finding (and published data24) that in cancer cells FGFR3-TACC3 fusion proteins signal predominantly via the ERK pathway and with our observation that these fusions do not rescue PIK3CA-mutant HNSCC cells from a PI3K inhibitor.
The accumulated genetic and functional data indicating that both the EGFR and FGFR pathways are important in HNSCC25, 33, 44, 45, 46 suggest that combined blockade of these pathways may be effective in HNSCC patients. Whereas one would not expect FGFR3-TACC3 fusions or gain-of-function mutations of FGFR3 to be homogeneously present in EGFR-driven tumors at diagnosis (because of the similar nature of the signals provided by these receptors), genetic alterations of FGFR3 present in rare subclones could drive acquired resistance, as suggested by our study (digital PCR data indicate that only very rare (<1 in 5000) parental FaDu cells harbor an FGFR3-TACC3 fusion (data not shown)). However, identifying patients in whom subclonal FGFR3 alterations might ultimately drive tumor regrowth is obviously challenging in the clinic. An alternate scenario, also supported by our data, is that ligand-dependent activation of FGFRs could drive innate resistance to EGFR inhibitors in a subset of HNSCCs. In this case, one might be able to identify tumors with active FGFR signaling at baseline by assessing expression of FGFRs and FGFs and/or by assessing FGFR phosphorylation status.
In summary, our study identifies signaling by FGFR3-TACC3 fusion proteins as a novel mechanism of resistance to EGFR/ERBB3 inhibition, providing further evidence for the importance of the FGFR3 pathway in HNSCC and suggesting the possibility that combined inhibition of EGFR and FGFR3 could be beneficial to a subset of HNSCC patients.
Materials and methods
ERBB3- and EGFR-blocking antibodies
Blocking antibodies against human ERBB3 (REGN1400) and human EGFR (REGN955) were generated using VelocImmune mice as described previously.13
Human tumor cell lines
FaDu, Cal27, NCI-H1975 and Detroit 562 cells were obtained from ATCC. SNU1076 cells were obtained from the Korean Cell Line Bank. Cell lines were authenticated by short tandem repeat profiling at ATCC (Promega, Madison, WI, USA). All experiments were conducted with low-passage cell cultures (< passage 10).
Generation of FaDu-resistant variant cell lines
Female C.B.-17 SCID mice (6–8 weeks old) bearing established FaDu tumors13 were randomized such that the average tumor size and variance for each treatment group were approximately equal (n=6 mice per treatment group, based on prior experience with this tumor model). Tumor regression was promoted by treatment (non-blinded) with EGFR/ERBB3-blocking antibodies. Mice were treated with a combination of REGN955 plus REGN1400 to isolate the V2 cell line and with a Regeneron EGFR/ERBB3-bispecific antibody to isolate the V1 cell line. The two treatment regimens provide an identical degree of tumor regression. Tumors that escaped treatment were harvested and re-passaged in vivo (generation of cell lines is described in detail in Figure 1a). All procedures were conducted according to the guidelines of the Regeneron Institutional Animal Care and Use Committee.
Analysis of tumor cell growth and signaling
The effect of various inhibitors on tumor cell growth and signaling was assessed as described previously.13 Cell growth between treatment groups was compared by one-way analysis of variance with Tukey’s multiple comparison test (groups were confirmed to have similar variances). Cells were treated with the following reagents (doses and treatment times are indicated in the Figure Legends): REGN1400 (ERBB3-blocking antibody), REGN955 (EGFR-blocking antibody), MEK inhibitor GSK1120212 (Selleckchem, Houston, TX, USA), FGFR TKI AZD4547 (Selleckchem), MET TKI PHA665752 (Sigma, St Louis, MO, USA), EGFR TKI AZD9291 (Selleckchem), PI3K inhibitor BYL719 (Selleckchem), human NRG1 (R&D Systems, Minneapolis, MN, USA) and human EGF (R&D Systems). The following antibodies were used for western blot analyses: ERBB3 (CST, Danvers, MA, USA, cat. #4754), phospho-ERBB3 (CST, cat. #4561), EGFR (CST, cat. #2646), phospho-EGFR (CST, cat. #2234), Akt (CST, cat. #9272), phospho-Akt (CST, cat. #4060), ERK (CST, cat. #4695), phospho-ERK (CST, cat. #4370), MET (CST, cat. #8198), phospho-MET (CST, cat. #3077) and phospho-FGFR (CST, cat. #3476).
The tyrosine phosphorylation status of 49 human RTKs was assessed using the Human Phospho-RTK Array Kit (R&D Systems) as described previously.
Identification of FGFR3-TACC3 fusion transcripts in FaDu variant cell lines
To identify genetic alterations unique to FaDu variants versus parental FaDu cells, mRNA was purified from 5 μg of total RNA using Dynabeads mRNA Purification Kit (Invitrogen, Carlsbad, CA, USA). Strand-specific RNA-seq libraries were prepared using ScriptSeq mRNA-Seq Library Preparation Kit (Epicentre, Madison, WI, USA). Twelve-cycle PCR was performed to amplify libraries. The amplified libraries were purified using 0.7X SPRIselect beads (Beckman Coulter, Brea, CA, USA) to enrich fragments larger than 300 bp. Sequencing was performed on Illumina HiSeq2500 by multiplexed paired-read runs with 2x100 cycles.
To confirm the sequences at the junctions of the fusion transcripts identified in FaDu V1 and V2 cells by RNA-seq, 100 ng of complementary DNA (cDNA) from FaDu P1, V1 or V2 cells was subjected to PCR. To amplify the region flanking the FGFR3 exon 18-TACC3 intron 9-TACC3 exon 11 fusion junctions (V1), nested PCR was performed using a forward primer in FGFR3 exon 17 (5′-AGAGGCCCACCTTCAAGC-3′) and a reverse primer in TACC3 exon 16 (5′-CAGATCCTGGTCAGCTCCTC-3′) for the first reaction. The second PCR reaction employed a forward primer in FGFR3 exon 18 (5′-AGCTCCTCAGGGGACGACTC-3′) and a reverse primer in TACC3 exon 11 (5′-TCACACCTGCTCCTCAGC-3′). To amplify the region flanking the FGFR3 exon 17-TACC3 exon 9 fusion junction (V2), a forward primer in FGFR3 exon 17 (5′-ATGCGGGAGTGCTGGCATG-3′) and a reverse primer in TACC3 exon 9 (5′-ACGTCCTGAGGGAGTCTCATTTG-3′) were used.
To quantitate the expression of the FGFR3-TACC3 fusion transcripts in FaDu V1 and V2 cells by Taqman real-time PCR, total RNA was extracted and cDNA was synthesized using High Capacity RNA-to-cDNA Master Mix (Applied Biosystems). To detect the FGFR3 exon 18-TACC3 intron 9-TACC3 exon 11 transcript (V1), the forward FGFR3 exon 18 primer and the reverse TACC3 exon 11 primer described above and the probe 5′-CGAAGGCGACACAGGAGGAGAACC-3′ were used. To detect the FGFR3 exon 17-TACC3 exon 9 transcript (V2), the forward FGFR3 exon 17 primer and the reverse TACC3 exon 9 primer described above and the probe 5′-CCTCCCAGAGGCCCACCTTCAAG-3′ were used. The assays were run under standard Taqman conditions on the ABI 7900HT instrument using the automatic setting for determining the threshold cycle. All probes were dual-labeled 5′ FAM/3′ BHQ-1 (Biosearch Technologies Inc, Novato, CA, USA).
Detection of FGFR3-TACC3 fusion proteins in FaDu variant cell lines
To assess the tyrosine phosphorylation status of FGFR3-TACC3 fusion proteins in FaDu V1 and V2 cell lines, cell lysates were subjected to immunoprecipitation with anti-phosphotyrosine (4G10) agarose conjugate (EMD Millipore, Darmstadt, Germany) followed by western blot analysis with antibodies against FGFR3 (Santa Cruz Biotechnology, Dallas, TX, USA, clone B-9), TACC3 (R&D Systems, cat. # AF5720) or Src (CST, cat. #2123).
To detect FGFR3-TACC3 fusion proteins by immunoprecipitation, cell lysates were subjected to immunoprecipitation with TACC3 antibody followed by western blot analysis with antibodies against FGFR3 or TACC3. In experiments aimed at separating FGFR3-TACC3 fusion proteins from native FGFR3, 4% SDS gels were employed as they enabled better resolution than the 4–20% gradient gels that were used for other western blot analyses. FGFR3-TACC3 fusion proteins were also detected in FaDu V1 and V2 cells by direct western blotting of cell lysates with FGFR3 antibody.
Generation of cell lines expressing FGFR3-TACC3 fusion proteins
To enable the expression of WT FGFR3 and the FaDu V1 and V2 FGFR3-TACC3 fusion proteins in cancer cells, DNA fragments encoding these proteins were cloned into the lentiviral expression vector pLVX-IRES-Neo (Clontech, Mountain View, CA, USA) in which the cytomegalovirus (CMV) promoter was replaced by the EF1a promoter. In addition, a kinase-dead version (K510M mutation) of the V2 FGFR3-TACC3 fusion was generated using the Q5 Site-Directed Mutagenesis Kit (NEB, Ipswich, MA, USA). To generate lentiviruses, 293T cells were co-transfected with the various pLVX-IRES-Neo plasmids plus the packaging vector psPAX2 and the envelope vector pMD2.G using FuGENE 6 transfection reagent (Promega). At 72 h after transfection, the virus-containing supernatant was collected and filtered. To generate pooled stable cell lines, cells (FaDu parental, Cal27, NCI-H1975, SNU1076, Detroit 562) were infected at a multiplicity of infection of 0.3 with the various lentiviruses and selected in 400–800 μg/ml G418 for ~2 weeks.
Disruption of FGFR3-TACC3 fusion genes using CRISPR/Cas9
Double-stranded oligonucleotides encoding sgRNAs specific to FGFR3 were cloned into the lentiCRISPR plasmid, a lentiviral expression vector that encodes Cas9 endonuclease, a sgRNA and the puromycin selection marker.47 To generate lentiviruses, 293T cells were co-transfected with lentiCRISPR plasmids plus the packaging vector psPAX2 and the envelope vector pMD2.G using FuGENE 6 transfection reagent (Promega). At 72 h after transfection, the virus-containing supernatant was collected, filtered and concentrated by ultracentrifugation. FaDu V1 and V2 cells were infected at a multiplicity of infection of 0.3 with lentiviruses encoding Cas9 endonuclease plus FGFR3 sgRNA 1 or FGFR3 sgRNA 2 or with a control lentivirus encoding only the Cas9 endonuclease. The sequence of the DNA encoding the CRISPR RNA portion of FGFR3 sgRNA 1 is 5′-GGGGACGGAGCAGCGCGTCG-3′ (binds in FGFR3 exon 2) and of FGFR3 sgRNA 2 is 5′-CGCGCTGCGTGAGCCGCTGC-3′ (binds in FGFR3 exon 3). At ~24 h after infection, cells were treated with 1 μg/ml puromycin to kill uninfected cells. Stably transduced cells were used for experiments (cell growth or cell signaling) between 10 and 14 days post infection.
References
Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science 2004; 304: 1497–1500.
Sharma SV, Bell DW, Settleman J, Haber DA . Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer 2007; 7: 169–181.
Bonner JA, Harari PM, Giralt J, Azarnia N, Shin DM, Cohen RB et al. Radiotherapy plus cetuximab for squamous-cell carcinoma of the head and neck. N Engl J Med 2006; 354: 567–578.
Cunningham D, Humblet Y, Siena S, Khayat D, Bleiberg H, Santoro A et al. Cetuximab monotherapy and cetuximab plus irinotecan in irinotecan-refractory metastatic colorectal cancer. N Engl J Med 2004; 351: 337–345.
Jonker DJ, O'Callaghan CJ, Karapetis CS, Zalcberg JR, Tu D, Au HJ et al. Cetuximab for the treatment of colorectal cancer. N Engl J Med 2007; 357: 2040–2048.
Chong CR, Janne PA . The quest to overcome resistance to EGFR-targeted therapies in cancer. Nat Med 2013; 19: 1389–1400.
Misale S, Di Nicolantonio F, Sartore-Bianchi A, Siena S, Bardelli A . Resistance to anti-EGFR therapy in colorectal cancer: from heterogeneity to convergent evolution. Cancer Discov 2014; 4: 1269–1280.
Arteaga CL, Engelman JA . ERBB receptors: from oncogene discovery to basic science to mechanism-based cancer therapeutics. Cancer Cell 2014; 25: 282–303.
Gala K, Chandarlapaty S . Molecular pathways: HER3 targeted therapy. Clin Cancer Res 2014; 20: 1410–1416.
Garner AP, Bialucha CU, Sprague ER, Garrett JT, Sheng Q, Li S et al. An antibody that locks HER3 in the inactive conformation inhibits tumor growth driven by HER2 or neuregulin. Cancer Res 2013; 73: 6024–6035.
Huang S, Li C, Armstrong EA, Peet CR, Saker J, Amler LC et al. Dual targeting of EGFR and HER3 with MEHD7945A overcomes acquired resistance to EGFR inhibitors and radiation. Cancer Res 2013; 73: 824–833.
Schaefer G, Haber L, Crocker LM, Shia S, Shao L, Dowbenko D et al. A two-in-one antibody against HER3 and EGFR has superior inhibitory activity compared with monospecific antibodies. Cancer Cell 2011; 20: 472–486.
Zhang L, Castanaro C, Luan B, Yang K, Fan L, Fairhurst JL et al. ERBB3/HER2 signaling promotes resistance to EGFR blockade in head and neck and colorectal cancer models. Mol Cancer Ther 2014; 13: 1345–1355.
Jiang N, Wang D, Hu Z, Shin HJ, Qian G, Rahman MA et al. Combination of anti-HER3 antibody MM-121/SAR256212 and cetuximab inhibits tumor growth in preclinical models of head and neck squamous cell carcinoma. Mol Cancer Ther 2014; 13: 1826–1836.
Engelman JA, Janne PA, Mermel C, Pearlberg J, Mukohara T, Fleet C et al. ErbB-3 mediates phosphoinositide 3-kinase activity in gefitinib-sensitive non-small cell lung cancer cell lines. Proc Natl Acad Sci USA 2005; 102: 3788–3793.
Holbro T, Beerli RR, Maurer F, Koziczak M, Barbas CF 3rd, Hynes NE . The ErbB2/ErbB3 heterodimer functions as an oncogenic unit: ErbB2 requires ErbB3 to drive breast tumor cell proliferation. Proc Natl Acad Sci USA 2003; 100: 8933–8938.
Soltoff SP, Carraway KL 3rd, Prigent SA, Gullick WG, Cantley LC . ErbB3 is involved in activation of phosphatidylinositol 3-kinase by epidermal growth factor. Mol Cell Biol 1994; 14: 3550–3558.
LoRusso P, Janne PA, Oliveira M, Rizvi N, Malburg L, Keedy V et al. Phase I study of U3-1287, a fully human anti-HER3 monoclonal antibody, in patients with advanced solid tumors. Clin Cancer Res 2013; 19: 3078–3087.
Mirschberger C, Schiller CB, Schraml M, Dimoudis N, Friess T, Gerdes CA et al. RG7116, a therapeutic antibody that binds the inactive HER3 receptor and is optimized for immune effector activation. Cancer Res 2013; 73: 5183–5194.
Schoeberl B, Pace EA, Fitzgerald JB, Harms BD, Xu L, Nie L et al. Therapeutically targeting ErbB3: a key node in ligand-induced activation of the ErbB receptor-PI3K axis. Sci Signal 2009; 2: ra31.
Knowles MA . Novel therapeutic targets in bladder cancer: mutation and expression of FGF receptors. Future Oncol 2008; 4: 71–83.
Parker BC, Annala MJ, Cogdell DE, Granberg KJ, Sun Y, Ji P et al. The tumorigenic FGFR3-TACC3 gene fusion escapes miR-99a regulation in glioblastoma. J Clin Invest 2013; 123: 855–865.
Singh D, Chan JM, Zoppoli P, Niola F, Sullivan R, Castano A et al. Transforming fusions of FGFR and TACC genes in human glioblastoma. Science 2012; 337: 1231–1235.
Williams SV, Hurst CD, Knowles MA . Oncogenic FGFR3 gene fusions in bladder cancer. Hum Mol Genet 2013; 22: 795–803.
Wu YM, Su F, Kalyana-Sundaram S, Khazanov N, Ateeq B, Cao X et al. Identification of targetable FGFR gene fusions in diverse cancers. Cancer Discov 2013; 3: 636–647.
Nelson KN, Meyer AN, Siari A, Campos AR, Motamedchaboki K, Donoghue DJ . Oncogenic gene fusion FGFR3-TACC3 is regulated by tyrosine phosphorylation. Mol Cancer Res 2016; 14: 458–469.
Sander JD, Joung JK . CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol 2014; 32: 347–355.
Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 2015; 517: 576–582.
Capelletti M, Dodge ME, Ercan D, Hammerman PS, Park SI, Kim J et al. Identification of recurrent FGFR3-TACC3 fusion oncogenes from lung adenocarcinoma. Clin Cancer Res 2014; 20: 6551–6558.
Di Stefano AL, Fucci A, Frattini V, Labussiere M, Mokhtari K, Zoppoli P et al. Detection, characterization and inhibition of FGFR-TACC fusions in IDH wild type glioma. Clin Cancer Res 2015; 21: 3307–3317.
Kim Y, Hammerman PS, Kim J, Yoon JA, Lee Y, Sun JM et al. Integrative and comparative genomic analysis of lung squamous cell carcinomas in East Asian patients. J Clin Oncol 2014; 32: 121–128.
Yuan L, Liu ZH, Lin ZR, Xu LH, Zhong Q, Zeng MS . Recurrent FGFR3-TACC3 fusion gene in nasopharyngeal carcinoma. Cancer Biol Ther 2014; 15: 1613–1621.
Stransky N, Cerami E, Schalm S, Kim JL, Lengauer C . The landscape of kinase fusions in cancer. Nat Commun 2014; 5: 4846.
Wang R, Wang L, Li Y, Hu H, Shen L, Shen X et al. FGFR1/3 tyrosine kinase fusions define a unique molecular subtype of non-small cell lung cancer. Clin Cancer Res 2014; 20: 4107–4114.
Lamont FR, Tomlinson DC, Cooper PA, Shnyder SD, Chester JD, Knowles MA . Small molecule FGF receptor inhibitors block FGFR-dependent urothelial carcinoma growth in vitro and in vivo. Br J Cancer 2011; 104: 75–82.
Crystal AS, Shaw AT, Sequist LV, Friboulet L, Niederst MJ, Lockerman EL et al. Patient-derived models of acquired resistance can identify effective drug combinations for cancer. Science 2014; 346: 1480–1486.
Herrera-Abreu MT, Pearson A, Campbell J, Shnyder SD, Knowles MA, Ashworth A et al. Parallel RNA interference screens identify EGFR activation as an escape mechanism in FGFR3-mutant cancer. Cancer Discov 2013; 3: 1058–1071.
Oliveras-Ferraros C, Cufi S, Queralt B, Vazquez-Martin A, Martin-Castillo B, de Llorens R et al. Cross-suppression of EGFR ligands amphiregulin and epiregulin and de-repression of FGFR3 signalling contribute to cetuximab resistance in wild-type KRAS tumour cells. Br J Cancer 2012; 106: 1406–1414.
Ware KE, Marshall ME, Heasley LR, Marek L, Hinz TK, Hercule P et al. Rapidly acquired resistance to EGFR tyrosine kinase inhibitors in NSCLC cell lines through de-repression of FGFR2 and FGFR3 expression. PLoS One 2010; 5: e14117.
Jebar AH, Hurst CD, Tomlinson DC, Johnston C, Taylor CF, Knowles MA . FGFR3 and Ras gene mutations are mutually exclusive genetic events in urothelial cell carcinoma. Oncogene 2005; 24: 5218–5225.
di Martino E, L'Hote CG, Kennedy W, Tomlinson DC, Knowles MA . Mutant fibroblast growth factor receptor 3 induces intracellular signaling and cellular transformation in a cell type- and mutation-specific manner. Oncogene 2009; 28: 4306–4316.
Juanpere N, Agell L, Lorenzo M, de Muga S, Lopez-Vilaro L, Murillo R et al. Mutations in FGFR3 and PIK3CA, singly or combined with RAS and AKT1, are associated with AKT but not with MAPK pathway activation in urothelial bladder cancer. Hum Pathol 2012; 43: 1573–1582.
Helsten T, Elkin S, Arthur E, Tomson BN, Carter J, Kurzrock R . The Fgfr landscape in cancer: analysis of 4853 tumors by next generation sequencing. Clin Cancer Res 2016; 22: 259–267.
Gaykalova DA, Mambo E, Choudhary A, Houghton J, Buddavarapu K, Sanford T et al. Novel insight into mutational landscape of head and neck squamous cell carcinoma. PLoS One 2014; 9: e93102.
Marshall ME, Hinz TK, Kono SA, Singleton KR, Bichon B, Ware KE et al. Fibroblast growth factor receptors are components of autocrine signaling networks in head and neck squamous cell carcinoma cells. Clin Cancer Res 2011; 17: 5016–5025.
Seiwert TY, Zuo Z, Keck MK, Khattri A, Pedamallu CS, Stricker T et al. Integrative and comparative genomic analysis of HPV-positive and HPV-negative head and neck squamous cell carcinomas. Clin Cancer Res 2015; 21: 632–641.
Shalem O, Sanjana NE, Hartenian E, Shi X, Scott DA, Mikkelsen TS et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 2014; 343: 84–87.
Acknowledgements
We thank Elizabeth Pasnikowski, Chandrika Taduriyasas, Jennifer Espert and David D’Ambrosio for technical assistance and Neil Stahl and George Yancopoulos for helpful comments and suggestions. All research was fully supported by Regeneron Pharmaceuticals Inc.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Daly, C., Castanaro, C., Zhang, W. et al. FGFR3-TACC3 fusion proteins act as naturally occurring drivers of tumor resistance by functionally substituting for EGFR/ERK signaling. Oncogene 36, 471–481 (2017). https://doi.org/10.1038/onc.2016.216
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2016.216
- Springer Nature Limited
This article is cited by
-
Recurrent UBE3C-LRP5 translocations in head and neck cancer with therapeutic implications
npj Precision Oncology (2024)
-
Conundrum of 3N: nasopharyngeal nonkeratinizing non-viral carcinoma—reappraisal of fusions and report of two consecutive cases with unusual molecular findings
Virchows Archiv (2024)
-
Precision drugging of the MAPK pathway in head and neck cancer
npj Genomic Medicine (2022)
-
Fusion genes in gynecologic tumors: the occurrence, molecular mechanism and prospect for therapy
Cell Death & Disease (2021)
-
FGFR3 signaling and function in triple negative breast cancer
Cell Communication and Signaling (2020)