Abstract
Helicobacter pylori infection of the human stomach is associated with inflammation that leads to the release of reactive oxygen and nitrogen species (RONs), eliciting DNA damage in host cells. Unrepaired DNA damage leads to genomic instability that is associated with cancer. Base excision repair (BER) is critical to maintain genomic stability during RONs-induced DNA damage, but little is known about its role in processing DNA damage associated with H. pylori infection of normal gastric epithelial cells. Here, we show that upon H. pylori infection, abasic (AP) sites accumulate and lead to increased levels of double-stranded DNA breaks (DSBs). In contrast, downregulation of the OGG1 DNA glycosylase decreases the levels of both AP sites and DSBs during H. pylori infection. Processing of AP sites during different phases of the cell cycle leads to an elevation in the levels of DSBs. Therefore, the induction of oxidative DNA damage by H. pylori and subsequent processing by BER in normal gastric epithelial cells has the potential to lead to genomic instability that may have a role in the development of gastric cancer. Our results are consistent with the interpretation that precise coordination of BER processing of DNA damage is critical for the maintenance of genomic stability.
Similar content being viewed by others
Introduction
Helicobacter pylori colonizes the gastric mucosa of half of the world’s population1 and is a major etiopathogenic factor for chronic antral gastritis, duodenal ulcers and gastric cancer.1, 2, 3 Chronic inflammation associated with the long-term persistence of H. pylori infection leads to release of reactive oxygen and nitrogen species (RONs) from inflammatory cells. RONs can cause DNA base damage, strand breaks and damage to the tumor-suppressor genes and enhanced expression of proto-oncogenes.4, 5, 6
Nitric oxide has also been found to inhibit the function of 8-oxoguanine glycosylase (OGG1) to impair the removal of DNA lesions that likely contribute to carcinogenesis.7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17 In addition, bacterial products including cytotoxins, lipase, phospholipase or the urease-mediated release of toxic ammonia18, 19, 20 at the site of inflammation can damage DNA, which may represent an early step in gastric carcinogenesis via development of chromosomal aberrations and DNA mutations.21, 22 H. pylori also induces hypoxia-inducible factor-1α23 that in turn inhibits mismatch repair24 to promote infection-associated microsatellite instability and cancer.10, 25, 26
The cellular consequences of DNA oxidation by RONs can lead to a number of different types of damage, such as 7, 8-hydroxy-2′-deoxyguanosine (8oxodG), abasic sites (AP) and oxidized deoxyribose sugars, which in turn lead to single-stranded DNA (ssDNA) breaks and double-stranded DNA breaks (DSBs),27 crosslinking of DNA and mutation.10, 13, 24, 28, 29 The most common oxidative base modifications resulting from direct attacks by hydroxyl radicals are purine lesions (8oxodG and 8-oxoA) and pyrimidine lesions (thymine glycol and cytosine glycol) in the human body in association with human cancer.30, 31, 32, 33, 34, 35, 36 Thousands of these lesions may be formed in each cell daily and levels are increased upon exposure to a variety of environmental factors.37, 38 Oxidized bases, including 8oxodG, are removed predominantly by base excision repair (BER).39, 40, 41 BER is the major repair pathway of DNA damage induced by RONs and is critical for maintaining genome stability during chronic inflammation that occurs during bacterial infection.42 BER is initiated by DNA glycosylases that recognize and cleave the damaged bases. The OGG1 bifunctional DNA glycosylase is the major enzyme that catalyzes the removal of 8oxodG paired with C.43, 44, 45 OGG1 remains bound to its abasic site product and its turnover can be stimulated either by AP endonuclease 1 (APE1) or by NEIL1,46, 47 both of which can process the AP site. After AP site processing and end-remodeling, the single-nucleotide gap is filled by DNA polymerase beta and the nick is sealed to complete repair.48
In humans, defective removal of 8oxodG or other types of base lesions may increase susceptibility to H. pylori-induced cancer due to mutagenesis. For example, unrepaired 8oxodG mispairs with adenine and results in G:C to A:T transversion mutations.49 Importantly, every step of BER generates intermediates (AP sites, 5′-deoxyribose phosphate residues and ssDNA breaks), which have been shown to be both mutagenic and toxic to cells.11, 50, 51 An imbalance between the generation of excess AP sites and inefficient repair has the potential to lead to mutation or DNA replication fork collapse.52
Some studies showed that H. pylori infection in BER-deficient cells leads to increased levels of inflammation that consequently result in the production of more RONs and tumor-promoting cytokines.42 Furthermore, Toller et al.53 found that coculture of H. pylori with mouse and human gastric cancer cell lines led to increased levels of DSBs. Although H. pylori infection induces genomic instability,54, 55 the underlying mechanism is not clear. Because H. pylori induces oxidative base damage, we hypothesized that the processing of this small base damage will lead to the accumulation of BER intermediates.56, 57 Our results show that H. pylori infection significantly increases the number of AP sites in cells. These AP sites arise in replicating DNA and eventually lead to DSB formation. In contrast, downregulation of the OGG1 DNA glycosylase reduces the number of AP sites and DSBs during infection. These data suggest that OGG1 deficiency has a protective role against genomic instability induced by H. pylori infection. Our data support the conclusion that H. pylori infection induces the accumulation of AP sites in DNA that are further processed into DSBs, resulting in genomic instability and cellular transformation.
Results
H. pylori induces accumulation of AP sites
To determine whether H. pylori infection induces an increased number of AP sites compared with non-infected controls, we infected GES-1 immortal but non-transformed gastric epithelial cells with H. pylori at a multiplicity of infection (MOI) of 100 bacteria to 1 cells. We then measured the number of AP sites in DNA extracted from the cells following infection at different time intervals as previously described58, 59, 60 (Figures 1a and b). For these experiments, AP sites were measured using an AP site assay kit (Colorimetric; Abcam, Cambridge, MA, USA) that utilizes an aldehyde reactive probe (ARP) reagent that reacts specifically with an aldehyde group, which is the open ring form of an AP site. We found that H. pylori infection generates on average 16 AP sites/105 nucleotides within 24 h compared with ~4 AP sites/105 nucleotides in non-infected controls (P=0.0001), a fourfold difference. Strikingly, the numbers of AP sites remain high over several hours in cells infected with H. pylori, reaching a maximum level of ~22 AP sites/105 nucleotides within the first 3 h of infection, and this is not significantly reduced during the next consecutive 6–48 h of infection (Figure 1b). Remarkably, H. pylori infection induces on average a threefold increase in the level of AP sites compared with cells treated with 10 mM H2O2, a potent base-oxidizing agent (P=0.01).
(a–e) Global levels of AP site accumulation are enhanced in H. pylori-infected gastric epithelial cells. (a) A standard curve for estimating AP sites/105 nucleotides using the AP sites assay kit (see Materials and methods section). (b) Estimated numbers of AP sites/105 nucleotides before (unfilled columns) and after 3, 6,12, 24, 48 and 72 h of H. pylori infection of GES-1 cells (filled columns) and after 1 h treatment of GES-1 cells with 10mM H2O2 as a control. (c) There is no fold change difference observed in protein expression levels of APE1 and DNA polymerase beta in H. pylori-infected for 12 and 24 h versus non-infected control. We have applied the ImageJ program to measure the integrated density of each band and normalized with the loading control. We then calculated the ratio of adjusted integrated density of the sample from the infected versus non-infected group as the fold change. (d) Representative images of the lengths of DNA tracks before and after APE1 treatment of H. pylori-infected GES-1 cells. (e) The distribution of the lengths of the DNA tracks (μm) in cells treated with APE1 after H. pylori infection. ns, not significant.
We wished to determine whether AP sites could arise in newly synthesized DNA in cells infected with H. pylori. To determine whether this is the case, we infected GES-1 cells with H. pylori for 12 h and then added iodo-deoxyuridine (IdU) to label the DNA during the active infection. We then lysed the cells, spread the DNA fibers on glass slides, treated them with APE1 followed by fixation and measured the lengths of the DNA fibers (Figure 1d). Fixation after treatment with APE1 allows the measurement to work. The lengths of the DNA tracks are significantly decreased in APE1-treated DNA fibers derived from H. pylori-infected cells (P=0.001) versus infected cells without APE1 treatment (Figure 1e), confirming that DNA track length in these experiments is dependent upon lesion processing by addition of APE1 to the fibers on the slide. Importantly, we found that the average length of the DNA tracks in APE1-treated DNA fibers derived from non-infected cells is significantly higher than the length of APE1-treated DNA fibers from infected cells (11.8 μm versus 6.4 μm; P=0.0001; Figure 1e), suggesting that more AP sites arise in actively replicating DNA from infected cells. However, protein expression of APE1 and DNA polymerase beta were not altered after 12 and 24 h of H. pylori infection (Figure 1c).
OGG1 downregulation reduces the number of AP sites during H. pylori infection
The OGG1 DNA glycosylase removes 8oxodG from DNA to initiate BER, resulting in AP sites. The increase in the number of AP sites observed in GES-1 cells infected with H. pylori could be a result of deficient processing by APE1 after excision of the damaged base by OGG1. To determine whether AP site levels observed in infected cells resulted from the removal of damaged bases by OGG1, we used small interfering RNA (siRNA) to downregulate OGG1 in infected GES-1 cells and controls, and then quantified the numbers of AP sites in these cells (Figures 2a and b). Downregulation of OGG1 protects the cells from the accumulation of high levels of AP sites (Figure 2b) compared with controls that are downregulated for GADPH. We found that H. pylori infection generates on average 14 AP sites/105 nucleotides within 24 h compared with ~4 AP sites/105 nucleotides in infected but OGG1-downregulated cells (P=0.02; Figure 2b). Remarkably, H. pylori infection induces on average a threefold lower number of AP sites in OGG1-downregulated cells compared with control cells (P=0.01). Next, we wished to determine the levels of AP sites in newly synthesized DNA in cells that were downregulated for OGG1. We performed these experiments by treating cells with an siRNA against OGG1 and infected GES-1 cells with H. pylori for 12 h. We then labeled replicating DNA for 2 h in cells with an active H. pylori infection, lysed the cells, spread the DNA fibers on slides, treated with APE1 and fixed the cells. We then quantified the lengths of the DNA tracks (Figure 2c). For non-infected GES-1 cells, we compared control cells (GAPDH downregulated) versus OGG1-downregulated cells with or without APE1 treatment (Figure 2d). There are no statistical differences between the lengths of DNA tracks in OGG1-downregulated cells with or without APE1. In contrast, the average length of the DNA tracks is significantly higher in infected and OGG1-downregulated cells treated with APE1 (10 μm) than in infected GES-1 cells that are downregulated for GAPDH (3.9 μm; P=0.0001; Figure 2d). These data suggest that the downregulation of OGG1 results in fewer AP sites that serve as a substrate for APE1. Our results are consistent with the idea that removal of damaged bases by OGG1 leads to accumulation of AP sites in GES-1 cells infected with H. pylori.
OGG1 downregulation suppresses AP site accumulation. (a) Western blot analysis of siRNA mediated downregulation of OGG1, GAPDH (control) and immunoblot with anti-rabbit OGG1 antibody, anti-mouse GAPDH and the loading control, anti-alpha-tubulin. (b) The estimated numbers of AP sites in OGG1 and GAPDH downregulated cells before (unfilled columns) and after (filled columns) infection with for 3, 6, 12, 24, 48 and 72 h. Note that all the experiments were conducted after 72 h of downregulation of OGG1 and GAPDH genes. (c) Representative images of DNA fibers after OGG1 and GAPDH downregulation. Cells were infected with H. pylori for 12 h and labeled with Idu for 2 h after H. pylori infection. The cells were lysed and the DNA fibers were spread on slides and treated with APE1 for 1 h at 37 oC. (d) Average lengths of DNA tracks in OGG1 and GAPDH downregulated cells with or without APE1 treatment. ns, not significant.
OGG1 downregulation results in fewer DSBs in H. pylori-infected GES-1 cells
AP sites are replication-blocking lesions that could result in the accumulation of DSBs, leading to chromosomal fragmentation and genomic instability if not repaired in an accurate and timely manner.61 To determine whether DSBs accumulate in cells infected with H. pylori, we monitored γH2AX recruitment as a marker for DSBs in infected versus non-infected GES-1 cells. The number of cells with γH2AX foci is significantly increased in cells infected with live H. pylori (n=50) compared with non-infected control cells (n=70) (P=0.0001; Figures 3a and b). Moreover, we asked whether live bacteria are necessary for DSB generation by characterizing γH2AX foci in cells treated with an extract or bacterial supernatant for 24 h. We observe significantly fewer γH2AX foci in cells treated with a bacterial extract (total number of cells analyzed n=60 and 46, respectively) compared with cells infected with live bacteria (n=50, P=0.01). The protein level of γH2AX is also increased by seven- and eightfold during 12 and 24 h of infection, respectively, compared with non-infected cells (Figure 3c). Importantly, we find that the γH2AX-positive cells infected with H. pylori are distributed in all cell cycle stages after infection, with the most significant differences in γH2AX-positive cells between infected and non-infected GES-1 cells observed during the G1 phase (Figure 3d). Processing of clustered AP sites by BER enzymes is known to result in the accumulation of DSBs during the G1 phase of the cell cycle,62, 63 which could be occurring in the H. pylori-infected GES-1 cells. To determine whether AP sites are an intermediate in DSB formation, we downregulated OGG1 because we know that its regulation results in fewer AP sites in H. pylori-infected cells. Importantly, OGG1-downregulated cells exhibit significantly fewer cells with γH2AX foci (Figure 3e). This result was confirmed by Fluorescence-Activated Cell Sorting (FACS) analysis and showed that significantly fewer γH2AX-positive H. pylori-infected cells are observed when OGG1 is downregulated (Figure 3f). In combination, these results indicate that AP sites are a BER intermediate leading to DSB formation in H. pylori-infected cells.
H. pylori infection of normal human gastric epithelial cells induces DSBs. (a) Images of GES-1 cells that were infected with J99 strain of live H. pylori, dead bacterial extract or a supernatant of the bacterial cell culture for 24 h. Note that blue represents the nuclei of cells stained with 4′,6-diamidino-2-phenylindole and red represents γH2AX foci. (b) The percent of γH2AX-positive cells, meaning >10 foci/cell, was estimated from live bacteria-infected GES-1 cells (n=50), bacterial cell extract (n=46), supernatant (n=60) and non-infected (n=70). (c) Western blot analysis of γH2AX before and after 12 and 24 h of H. pylori infection. The level of γH2AX increased seven- and eightfold after infection with H. pylori for 12 and 24 h, respectively. Note that quantification of bands was carried out using the National Institutes of Health ImageJ program by measuring the integrative density of each γH2AX band that is normalized with alpha-tubulin band as internal control. (d) Estimated percentage of cells with γH2AX in different cell cycle stages based on FACS analysis after 24 h of infection. Cells were stained with γH2AX antibody and propidium iodide to assess the levels of DSBs and the cell cycle phase, respectively, and were analyzed by flow cytometry. (e) Percentage of cells with γH2AX foci in OGG1-deficient cells after 24 h of infection. (f) Percentage of cells with γH2AX in different stages of the cell cycle in OGG1-deficient versus OGG1-proficient cells after 24 h of infection based on FACS analysis. Note that all images were taken × 63 Zeiss microscope with constant exposure time and any cell with >10 foci of γH2AX/cell was categorized as a positive. The data were analyzed based on the paired t-test using GraphPad Prism program. ns, not significant.
Infection induces replication fork collapse
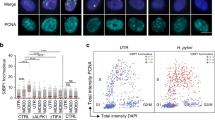
Chromosome breakage as a result of replication stress has been hypothesized to be a direct consequence of defective replication fork progression or collapsed replication forks. We asked whether the γH2AX staining that is observed during the S and G2 phases of the cell cycle is associated with arrested DNA replication forks. As PCNA is essential for numerous cellular processes including DNA replication and repair,64 we performed co-immunostaining of the cells with antisera against PCNA and γH2AX. This revealed that 45% of the γH2AX foci colocalized with PCNA in infected cells (P=0.001, n=78; Figures 4a and b) and colocalization was observed in very few uninfected cells (n=92; Figure 4a). Colocalization of PCNA with γH2AX is likely indicative of DNA damage within cells positioned in S phase.
Replication fork arrest is a source of genomic rearrangements, and the recombinogenic properties of blocked forks likely depend on the cause of blockage. We applied a DNA fiber approach to determine whether infection with H. pylori leads to slowed or stalled replication forks. The general schematic for these experiments is shown in Figure 5a. We labeled actively replicating DNA with IdU and then infected cells with H. pylori for 12 h, followed by incubation with CIdU, as shown in the general schematic in Figure 5a. The replication fork speed after infection was quantified by dividing the length of each fluorescent track by the time of incubation with the halogenated nucleotide, shown in Figures 5b and c. Analyses of replication fork speeds in H. pylori-infected versus non-infected cells showed a broad distribution of values ranging from 0.085 to 1.24 kb/min and 0.4 to 2.5 kb/min, respectively, (Figure 5c). Approximately 58% of the forks in infected cells exhibit fork speeds between 0.2 and 0.6 kb/min. In contrast, the distribution of fork speeds in non-infected cells is between 0.6 and 1.2 kb/min (Figure 5c). The average fork speed with infection (mean±s.e.m.) is 0.42±0.03 kb/min, which is significantly reduced compared with that of non-infected control groups (1.05±0.048 kb/min; P=0.0001; Figure 5d). Furthermore, we find that H. pylori infection induces stalled forks in 40% of cells (n=263) versus ~6% of non-infected control cells (n=201; P=0.0001; Figure 5e), as assessed by failure to incorporate CIdU. However, the percent of stalled forks is not significantly changed in cells infected with H. pylori for 6 versus 12 h (Figures 5d–f(i–iii)).
DNA combing analysis of DNA replication in GES-1 cells before and after infection. (a) Schematic representation of GES-1 cells cultured (blue), pulsed with IdU (30 min represented by red solid line), infected with H. pylori (black) for various times and followed by incubation with CIdU (30 min, represented by green solid line). The cells stained red in the cartoon represent the population of cells with stalled replication forks, whereas those stained yellow represent the cells with active replication forks. (b) Representative image of DNA fibers derived from H. pylori-infected (6 or 12 h) and non-infected control GES-1 cells. (c) The distribution of fork speed with (red columns) and without infection for 12 h (green columns) ranges between 0.2 and 2.6 kb/min. (d) Average DNA replication fork speed of H. pylori-infected versus non-infected control cells. Replication fork speed after infection was calculated by measuring the signal length of CIdU divided by pulse duration (30 min). (e) Estimated fraction of stalled forks of H. pylori-infected cells for 6 and 12 h (red columns) versus non-infected control cells (green columns). (f) Schematic representation of experimental design. (i) GES-1 cells cultured and pulsed with IdU (red) for 30 min and immediately followed by CIdu label (green) for 30 min. (ii) Idu label followed with H. pylori infection for 6 h then labeled with CIdU for 30 min. (iii) Idu label followed by 12 h with H. pylori infection then labeled with CIdU for 30 min. Note that fork stalling does occur within the time frame of a double-labeling experiment. To estimate the numbers of stalled forks, we counted the number of IdU-labeled tracks, which fail to incorporate the subsequent 30-min pulse of CldU. These events will therefore appear as an elevated number of red-only tracks (IdU). ns, not significant.
Replication protein A-coated ssDNA and appearance of Rad51 increases during infection
Replication protein A (RPA), the major ssDNA-binding protein in eukaryotic cells, accumulates along stretches of ssDNA generated by stalled replication forks and/or DNA damage.65, 66 To determine whether ssDNA levels increase during infection, we quantified the number of cells with RPA foci (Figures 6a and b). We find that the number of H. pylori-infected cells with RPA foci significantly increased (n=143, 63%) compared with non-infected controls (n=176, 18%; P=0.001; Figure 6b). Once DSBs are induced by H. pylori, 5′-DNA end resection of DSBs is a prerequisite for loading of Rad51 to promote strand exchange activity.67, 68 When the cells were analyzed for the presence of RAD51 foci (Figure 6c), 50% of the infected cells exhibited foci compared with 13% for non-infected controls (13%; n=67; P=0.0001; Figure 6d).
ssDNA is critical for RPA localization and the recruitment of Rad51 foci in H. pylori-infected cells. (a) Localization of RPA in infected H. pylori cells versus non-infected cells. (b) Percent of cells with RPA foci in infected versus non-infected cells (P=0.001). (c) Localization of Rad51 foci in H. pylori-infected cells. Rad51 foci are recruited to the DSB sites in H. pylori-infected GES-1 cells. (d) Estimated percent of cells with Rad51 foci in infected cells and non-infected cells.
H. pylori infection induces chromosomal aberrations
DSBs can lead to chromosomal fragmentation and genomic rearrangements if not repaired in an accurate and timely manner.61 Previous work has documented the existence of chromosomal aberrations but only in gastric cancer cells infected with H. pylori. However, it is not known whether H. pylori infection induces chromosomal aberrations in normal gastric epithelial GES-1 cells, which are the cells that are initially infected with this pathogen. Therefore, we asked whether H. pylori induces chromosomal aberrations in normal human gastric epithelial cells. Infection of GES-1 cells with H. pylori for 24 h results in significantly increased levels of chromosomal fusions (64%), chromosomal fragments (48%), chromatid breaks (36%) and chromosome breaks (n=21%) versus non-infected cells (P=0.0001; Figures 7a and b). Our observation is significant because the levels of chromosomal aberrations are more pronounced in normal gastric epithelial cells than in previously reported gastric cancer cell lines infected with H. pylori.53 We tested the hypothesis that the genomic instability resulting from H. pylori infection induces cellular transformation. We infected normal gastric epithelial cells with H. pylori for 35 days and monitored focus formation. In the focus formation assay, non-transformed cells will grow to confluence forming a monolayer (Figure 7c), whereas transformed cells will continue to grow after reaching confluence, thereby forming foci (Figure 7c). We find that the number of foci/field are significantly increased in infected cells versus non-infected control cells (Figure 7d; P=0.0001).
(a– e) Infection with H. pylori induces chromosomal aberrations and cellular transformation. (a) Representative images of metaphase spreads from non-infected and GES-1 cells infected with H. pylori for 24 h. Note that arrowheads show fusion, black arrows show chromatid breaks and gray arrows show chromosome breaks. (b) Estimated percentage of different types aberrations. (c) Representative image of transformed cells after H. pylori infection for 35 days coculture. (d) Estimated number of foci/field for transformed cells that were infected with H. pylori for 35 days. All statistical analysis was performed using the paired t-test on GraphPad prism software. (e) Proposed model of DNA damage response and repair in H. pylori-infected gastric epithelial cells. H. pylori induces RONS that could result in 8oxodG lesions (shown as orange circles) that are likely removed with OGG1, resulting in the formation of AP sites (represented by blue circles). AP sites are clustered in close proximity on the opposite strands of the DNA, and processed with AP lyase activity that cleaves the 3′-side of AP site40, 45, 88, 89 or with APE1 that cleaves the phosphate backbone and generates DSBs (left side of the model) in G1 phase of cell cycle. In contrast, OGG1 downregulation reduces the numbers of AP sites that accumulate during G1 phase of the cell cycle and reduces the levels of DSBs during infection. The second alternative is if AP sites are localized in the vicinity of active replication forks, then cleavage of the AP site at the replication fork could lead to replication fork collapse at DSBs (right side of the model) during the S/G2 phase of cell cycle. We propose that repair at G1 phase of DSBs may lead to chromosomal aberrations that are manifested at the metaphase stage of cell division and eventually repaired via non-homologous end-joining pathways that likely induce cellular transformation. In contrast repair at S/G2 phase is processed by homologous recombination-dependent repair.
Discussion
The response of gastric epithelial cells to H. pylori infection and their ability to maintain genomic stability via DNA repair mechanisms is essential in preventing gastric cancer initiation and progression. Here, we provide important mechanistic insight regarding the consequences of H. pylori infection of normal human gastric epithelial cells, a physiologically relevant model of gastric cancer. We show that infection of these cells with H. pylori induces the accumulation of AP sites. Our results show that AP sites are generated at a high frequency in genomic DNA during infection. In contrast, downregulation of OGG1 during H. pylori infection protects cells from accumulating AP sites. Our data suggest that H. pylori infection induces 8oxodG lesions in genomic DNA that are removed by OGG1, leading to the accumulation of AP sites, either as a result of continuous generation, deficient repair or an imbalance between generation of AP sites and their repair. Moreover, we show that many of these AP sites are further processed into DSBs, resulting in genomic instability in OGG1-proficient H. pylori-infected cells. We conclude that H. pylori infection induces BER intermediates that likely overwhelm the BER machinery. This alters the dynamics of AP sites in the DNA to drive genomic instability in gastric epithelial cells.
Our results show that processing of AP sites into DSBs occurs during all phases of the cell cycle in OGG1-proficient cells. The cell cycle profiles revealed an increase in the percentage of cells in G1 versus S and G2 after infection. (Supplementary Figure 1) Therefore, we propose that DNA DSBs accumulate as a result of AP site processing during infection by two different mechanisms. The first mechanism is associated with the G1 phase of the cell cycle in which we observe the most significant accumulation of DSBs (Figure 7e). We suggest that these DSBs arise as a result of the removal of clustered damaged bases by OGG1. OGG1 glycosylase activity could catalyze the removal of the damaged base, resulting in an AP site that is processed by the OGG1 lyase activity, leading to DSBs if the oxidative DNA damage is clustered.69, 70 However, OGG1 has weak lyase activity,71, 72, 73 so once the damaged base is removed by this enzyme, it is possible that other bifunctional DNA glycosylases such as NTH1, NEIL1 or NEIL2, or perhaps APE1 itself, could process the abasic sites, leading to the accumulation of DSBs. Our results show that OGG1 deficiency suppresses the accumulation of AP sites, resulting in a significant reduction of DSBs at the G1 cell cycle stage during H. pylori infection (Figure 7e). These data suggest that OGG1 functions in the removal of bases that are damaged during infection, and that an OGG1 deficiency protects cells from H. pylori-induced DSBs at the G1 cell cycle stage. The presence of other bifunctional glycosylases, such as NEIL1, do not appear to have a significant backup role in the removal of 8oxodG induced by H. pylori likely as a result of their weak glycosylase activity against 8oxodG. However, the lyase activity of glycosylases that recognize other types of base lesions may process AP sites owing to robust AP lyase activity73, 74, 75, 76, 77 and may be able to promote BER in OGG1-deficient cells infected with H. pylori. This could contribute to the lower numbers of AP sites in OGG1-deficient cells infected with H. pylori. Our observation of the critical role of OGG1 during H. pylori infection is supported by previously published data, showing that OGG1 deficiency promotes a protective role against inflammation and genotoxicity associated with H. pylori infection.78 Our results are consistent with the interpretation that OGG1 has a key role in the removal of RONs-mediated 8oxodG lesions during H. pylori infection. Initiation of BER of clustered oxidized bases on opposite strands of the DNA by OGG1 would lead directly to the formation of DSBs. These resulting DSBs are most likely repaired by the error-prone pathway of non-homologous end-joining during G1, leading to chromosomal aberrations.
The second mechanism of DSB generation occurs during S and G2 phases, as DSBs also accumulate during these phases of the cell cycle. Our results suggest that DSBs arise as a result of replication fork encounter of AP sites that may potentially block DNA polymerase.79, 80 It is also possible that the AP sites are processed by APE1 as they collide with the replication fork in cells infected with H. pylori. In fact, it has been proposed that homologous recombination is needed to repair replication forks that collapse owing to encountering oxidative damage81 and to assist in repairing DSBs that form as a consequence of repairing complex oxidative DNA lesions.81, 82, 83 In support of previous studies, our data on the presence of RPA and Rad51 in cells infected with H. pylori may likely indicate a replicative stress response including fork stabilization or replication fork restart.65, 84, 85 The homology-dependent repair pathway repairs DSBs arising during S and G2 phases, which is mostly error-free, but in some cases, can also lead to genomic instability.
Altogether, our data indicate that H. pylori infection induces the accumulation of unrepaired BER intermediates that can initiate a cascade of events to generate DSBs. We show that overwhelming the capacity of BER as a result of oxidative DNA damage induced by H. pylori results in the accumulation of AP sites and DSBs in normal gastric epithelial cells, leading to genomic instability, an underlying cause of gastric cancer. Our findings suggest that alteration of the activity of enzymes that function in BER as a result of genetic mutation could significantly impact gastric cancer risk.
Materials and Methods
H. pylori strains, culture and preparation of bacterial components
The H. pylori type strain, J99 strain ATCC 700824, was used in this study and was stored at −80 °C. The H. pylori strain was inoculated onto Columbia blood agar plates, which consisted of 39 g of Columbia Agar Base (CM0331, Becton Dickinson, Franklin Lakes, NJ, USA) per liter, 7% hemolyzed horse blood (SR0048) and supplemented with one vial of Dent (SR0147, Oxoid, Hampshire, UK). For growing visible colonies, plates were kept in a Gaspack (BD GasPak EZ products, Franklin Lakes, NJ, USA) at 37 °C from 3 to 10 days until colonies were observed. From the primary growth, single colonies were propagated in blood agar for an additional 48 h, harvested in phosphate-buffered saline (PBS) and inoculated into Hams F12 media supplemented with 5% horse serum and incubated in 5% CO2 incubator. Bacterial protein extraction was conducted using B-PER extraction reagents based on manufacturer’s protocols (Thermo Scientific, Rockford, IL, USA).
Cell lines and cultures
The GES-1 gastric epithelial cell line was obtained from the Chinese Academy of Sciences. GES-1 cells are derived from a human gastric mucosa epithelium and immortalized via SV40 and are non-tumorigenic.86, 87 GES-1 cells were maintained in RPMI supplemented with 10% fetal bovine serum, 1% glutamate and 1% penicillin–streptomycin.
Preparation of GES-1 cellular extracts
The cells were suspended in 20 mM HEPES, pH 7.9, 10 mM NaCl, 1.5 mM MgCl2, 10% glycerol 1 mM phenylmethylsulfonyl fluoride (PMSF) and a protease inhibitor cocktail and incubated for 1 h on a rocking platform at 4 °C. The whole-cell lysate was collected after centrifugation at 13 000 r.p.m.at 4 °C for 20 min and aliquoted and stored −80 °C.
Confocal microscopy of nuclear protein localization, antibodies used
For γH2AX foci staining, cells were grown on four chamber slides and infected with H. pylori for 24 h (100:1 bacteria to host cell ratio, MOI=100). After cell media was removed and rinsed with PBS, cells were fixed with methanol:acetic acid (3:1 ratio), incubated for 15 min at −20 °C, and permeabilized in PBS containing 0.5% Triton X-100 for 8 min at room temperature (RT). Cells were then incubated with 1:200 diluted rabbit polyclonal anti-γH2AX antibody (Bethyl Laboratory, Montgomery, TX, USA) for 1 h at RT and detected with a secondary fluorescein isothiocyanate (FITC)-conjugated goat anti-rabbit IgG (Jackson ImmunoResearch Laboratories, West Grove, PA, USA). Antibody dilutions and washes after incubations were performed in PBS containing 0.5% bovine serum albumin and 0.05% Tween-20. Finally, coverslips were mounted in Vectashield mounting medium with 4′,6-diamidino-2-phenylindole (H-1500; Vector Laboratories, Burlingame, CA, USA).
Flow cytometry
GES-1 cells were cultured in four chamber slides and infected with H. pylori (100: 1 bacteria to host cell ratio, MOI=100) for 24 h. The bacterial media was removed and the cells were rinsed with PBS then, cells were harvested by trypsinization, washed once with PBS and pelleted. The pellet was resuspended by adding 70% ice-cold ethanol dropwise while vortexing. Cells were fixed overnight at −20 °C. The cells were incubated with primary phospho-γH2AX antibody (Millipore, Billerica, MA, USA; 05-636) 1:500 overnight at 4 °C. Following the incubation, cells were washed twice with PBS and incubated with anti-mouse secondary antibody conjugated to FITC 1:500 for 1 h at RT. Cells were washed twice with PBS and resuspended in 500 μl propidium iodide (PI)/RNase staining buffer (BD Pharmingen, Franklin Lakes, NJ, USA). Fluorescence was analyzed by flow cytometry using the BD FACSCalibur (Franklin Lakes, NJ, USA) and also by using FlowJo 8.8.6 software (Ashland, OR, USA).
Metaphase spread preparation
For preparation of metaphase chromosome spreads in the absence of infection, GES-1 cells were treated with colcemid (final concentration of 0.1 μg/ml) for 6 h before harvesting. In contrast, GES-1 cells were cultured and infected with H. pylori (with 100:1 bacteria to host cell ratio, MOI =100) for 24 h including colcemid treatment for the last 6 h. The cells were trypsinized and were washed once with PBS. Mitotic cells were collected and centrifuged at 200 g for 5 min at RT, and the harvested cells were treated with 75 mM KCl at 30 min at 37 °C. After centrifugation, the cells were fixed three times in a freshly prepared mixture of 3:1 methanol:acetic acid. Ten microliters of cell suspension were dropped onto slides and allowed to dry, followed by rinsing the slides in PBS and staining with 5% Giemsa stain for 8 min. The slides were rinsed with water and were air-dried. Images were acquired with a Ziess microscope (Peabody, MA, USA).
DNA fiber analysis
For DNA replication analysis, sequential labeling of DNA with IdU and 5-chloro-2′-deoxyuridine (CldU) were performed. A subconfluent, asynchronous population of GES-1 cells was first labeled for 30 min with 25 μM IdU, washed with medium three times and infected for 12 h with H. pylori at an MOI of 100:1 (bacteria to host cell ratio; MOI=100). The cells were then labeled for another 30 min with 250 μM CldU. After incubation, cells were washed and resuspended at a concentration of 7.5 × 105 cells/ml. The number of cells lysed per slide ranged between 1500 and 5000 cells using fiber lysis buffer (50 mM EDTA, 0.5% SDS and 200 mM Tris–HCl, pH=7.5) for 2 min, and the slides were tilted at 20 ° for gravity flow. The control non-infected cells used were pulsed for 30 min with IdU, followed by 12 h with Ham’s F12 media and then pulsed with CIdU label for 30 min, and the cells were harvested for the fiber assay. For immunoflourescence staining, the slides were fixed for 10 min with methanol:acetic acid (3:1) and air-dried. The slides were treated with 2.5 M HCl for 30 min, washed with 1x PBS three times and then blocked with 3% bovine serum albumin/PBS for 1 h. CldU was detected by incubating acid-treated fiber spreads with rat anti-BrdU monoclonal antibody (Abcam) and IdU was detected using mouse anti-BrdU monoclonal antibody (1:1000; Becton Dickinson) for 1 h at RT, followed by washing three times with 1x PBS and stained with secondary antibody conjugated with sheep anti-mouse Cy3 and goat anti-rat Alexa flour 488 for 1 h at RT. The slides were mounted with Vectashield mounting media and covered with coverslips. Images were acquired with × 63 magnification using a Zeiss microscope and processed and analyzed using the ImageJ program (NIH, Bethesda, MD, USA). The lengths of red (Cy3) or green (AF 488) labeled patches were measured using the ImageJ software (National Institutes of Health; http://rsbweb.nih.gov/ij/) and arbitrary length values were converted into micrometers using the scale bars created by the microscope. Fluorescence images were captured using a Zeiss LSM 510 inverted confocal microscope using × 63/numerical aperture 1.4 oil immersion objective, and data analysis was carried on using the ImageJ software. We applied a conversion factor used is 1 μm=2.59 kb.85
AP site distribution on DNA tracks
A subconfluent, asynchronous population of GES-1 cells was infected for 12 h with H. pylori (MOI=100) then labeled for 2 h with 25 μM IdU. Cells were harvested and placed onto a glass slide and allowed to gently adhere to the glass surface. Next the cells were lysed causing their genomic DNA to be released, and then the DNA is straightened and aligned on the slide using gravitational flow. After air-drying, the slides were treated with APE1 (50 and 100 U) with reaction buffer containing 1 × NEB4 buffer for 1 h at 37 oC. The slides were washed with 1 × PBS three times, and fixed for 10 min with methanol:acetic acid (3:1) and processed for immunofluorescence based on the above described methods. To determine the relative distribution of AP site formation on the length of DNA track, we treated the slides with APE1 (100 U) for 1 h at 37 °C before fixation, then fixed the cells and labeled with secondary antibody conjugated with sheep anti-mouse Cy3. The slides were mounted with Vectashield mounting media and covered with coverslips. Images were acquired with × 63 magnification using a Ziess microscope, and were processed and analyzed using the ImageJ program.
AP site analysis
GES-1 cells were cultured until 70% confluent and infected with H. pylori (with 100:1 bacteria to host cell ratio, MOI=100), for different time intervals. The bacterial media was removed and the cells were washed three times with PBS, then genomic DNA was isolated using DNAzol genomic DNA isolation reagents that contains guanidine/detergent lysis buffer that we purchased from Invitrogen (Grand Island, NY, USA). The AP assay was conducted after the DNA was labeled with ARP. The AP sites assay kit (Colorimetric; Abcam) utilizes the ARP reagent that reacts specifically with an aldehyde group, which is the open ring form of the AP sites. After treating DNA containing AP sites with ARP reagents, AP sites are tagged with biotin residues, which can be quantified using an avidin–biotin assay followed by a colorimetric detection. The kit provides the necessary reagents for convenient determination of abasic sites in purified DNA in 96-well plate format. The number of AP sites was measured and calculated based upon a standard curve generated using ARP standard DNA solutions as described previously (DNA Damage AP sites assay kit, Colorimetric, Abcam).
siRNA for OGG1 downregulation
For the siRNA studies, accell SMARTpool siRNA against OGG1 with target sequence 5′-GCCUUUUCUCUUUUAUCUG-3′ (A-005147-20), 5′-GCCUUCUGGACAAUCUUUC-3′ (A-005147-19), 5′-GGAUCAAGUAUGGACACUG-3′ (A-005147-18) and 5′-CUCAGAAAUUCCAAGGUGU-3′ (A-005147-17) were used to downregulate OGG1. In addition, ON-TARGETplus GAPDH control siRNA with target sequence 5′-GUCAACGGAUUUGGUCGUA-3′ were synthesized by Dharmacon Research Inc. (Lafayette, CO, USA). All DharmaFECT transfection reagent and protocols for downregulation of OGG1 and GAPDH were carried out based on modified Thermo Scientific DharmaFECT Transfection Reagents-siRNA Transfection Protocol.
Statistical analysis
All the reported data were evaluated in a pairwise manner, comparing infected versus non-infected control cells using GraphPad Prism (La Jolla, CA, USA).
Cellular transformation assay
Viable cells (200 cells/well) were seeded into wells of a six-well plate (in triplicate) and cultured in an incubator with 5% CO2. Then, bacteria were added in the ratio (GES-1 cells:bacterial cells) of 1:10 for coculture. GES-1 cells cultured under the same conditions but without coculture with H. pylori were used as controls. The culture continued for 35 days, and the cells were used for experiments and analysis during that period of time. Cells in the plate were washed with PBS after removal of the medium, and then fixed followed by staining with crystal violet for 15 min. The stain was washed off under running tap water, and the plate was allowed to dry. The number of distinctly stained cell growth pattern was counted under microscope.
References
Peek RM Jr, Blaser MJ . Helicobacter pylori and gastrointestinal tract adenocarcinomas. Nat Rev Cancer 2002; 2: 28–37.
Correa P . Human gastric carcinogenesis: a multistep and multifactorial process—First American Cancer Society Award Lecture on Cancer Epidemiology and Prevention. Cancer Res 1992; 52: 6735–6740.
Figueiredo C, Machado JC, Pharoah P, Seruca R, Sousa S, Carvalho R et al. Helicobacter pylori and interleukin 1 genotyping: an opportunity to identify high-risk individuals for gastric carcinoma. J Natl Cancer Inst 2002; 94: 1680–1687.
Cerutti PA . Oxy-radicals and cancer. Lancet 1994; 344: 862–863.
Feig DI, Reid TM, Loeb LA . Reactive oxygen species in tumorigenesis. Cancer Res 1994; 54: 1890s–1894s.
Schreck RR . Tumor suppressor gene (Rb and p53) mutations in osteosarcoma. Pediatric Hematol Oncol 1992; 9: ix–i.
Bagchi D, Bhattacharya G, Stohs SJ . Production of reactive oxygen species by gastric cells in association with Helicobacter pylori. Free Radic Res 1996; 24: 439–450.
Farinati F, Cardin R, Degan P, Rugge M, Mario FD, Bonvicini P et al. Oxidative DNA damage accumulation in gastric carcinogenesis. Gut 1998; 42: 351–356.
Izzotti A, De Flora S, Cartiglia C, Are BM, Longobardi M, Camoirano A et al. Interplay between Helicobacter pylori and host gene polymorphisms in inducing oxidative DNA damage in the gastric mucosa. Carcinogenesis 2007; 28: 892–898.
Kim JJ, Tao H, Carloni E, Leung WK, Graham DY, Sepulveda AR . Helicobacter pylori impairs DNA mismatch repair in gastric epithelial cells. Gastroenterology 2002; 123: 542–553.
Loeb LA, Preston BD . Mutagenesis by apurinic/apyrimidinic sites. Annu Rev Genet 1986; 20: 201–230.
Obst B, Wagner S, Sewing KF, Beil W . Helicobacter pylori causes DNA damage in gastric epithelial cells. Carcinogenesis 2000; 21: 1111–1115.
Park DI, Park SH, Kim SH, Kim JW, Cho YK, Kim HJ et al. Effect of Helicobacter pylori infection on the expression of DNA mismatch repair protein. Helicobacter 2005; 10: 179–184.
Yao Y, Tao H, Park DI, Sepulveda JL, Sepulveda AR . Demonstration and characterization of mutations induced by Helicobacter pylori organisms in gastric epithelial cells. Helicobacter 2006; 11: 272–286.
Jaiswal M, LaRusso NF, Nishioka N, Nakabeppu Y, Gores GJ . Human Ogg1, a protein involved in the repair of 8-oxoguanine, is inhibited by nitric oxide. Cancer Res 2001; 61: 6388–6393.
Jaiswal M, LaRusso NF, Shapiro RA, Billiar TR, Gores GJ . Nitric oxide-mediated inhibition of DNA repair potentiates oxidative DNA damage in cholangiocytes. Gastroenterology 2001; 120: 190–199.
Wink DA, Vodovotz Y, Laval J, Laval F, Dewhirst MW, Mitchell JB . The multifaceted roles of nitric oxide in cancer. Carcinogenesis 1998; 19: 711–721.
Berstad K, Sjodahl R, Berstad A . Phospholipase A2 activity in gastric juice from patients with active and H. pylori-eradicated healed duodenal ulcer. Aliment Pharmacol Ther 1994; 8: 175–180.
Langton SR, Cesareo SD . Helicobacter pylori associated phospholipase A2 activity: a factor in peptic ulcer production? J Clin Pathol 1992; 45: 221–224.
Slomiany BL, Slomiany A . Mechanism of Helicobacter pylori pathogenesis: focus on mucus. J Clin Gastroenterol 1992; 14: S114–S121.
Du MQ, Carmichael PL, Phillips DH . Induction of activating mutations in the human c-Ha-ras-1 proto-oncogene by oxygen free radicals. Mol Carcinog 1994; 11: 170–175.
Floyd RA . Role of oxygen free radicals in carcinogenesis and brain ischemia. FASEB J 1990; 4: 2587–2597.
Park JH, Kim TY, Jong HS, Chun YS, Park JW, Lee CT et al. Gastric epithelial reactive oxygen species prevent normoxic degradation of hypoxia-inducible factor-1alpha in gastric cancer cells. Clin Cancer Res 2003; 9: 433–440.
Koshiji M, To KK, Hammer S, Kumamoto K, Harris AL, Modrich P et al. HIF-1alphainduces genetic instability by transcriptionally downregulating MutSalpha expression. Mol Cell 2005; 17: 793–803.
Mirzaee V, Molaei M, Shalmani HM, Zali MR . Helicobacter pylori infection and expression of DNA mismatch repair proteins. World J Gastroenterol 2008; 14: 6717–6721.
Fleisher AS, Esteller M, Harpaz N, Leytin A, Rashid A, Xu Y et al. Microsatellite instability in inflammatory bowel disease-associated neoplastic lesions is associated with hypermethylation and diminished expression of the DNA mismatch repair gene, hMLH1. Cancer Res 2000; 60: 4864–4868.
Cooke MS, Evans MD, Dizdaroglu M, Lunec J . Oxidative DNA damage: mechanisms, mutation, and disease. FASEB J 2003; 17: 1195–1214.
Halliwell B, Aruoma OI . DNA damage by oxygen-derived species. Its mechanism and measurement in mammalian systems. FEBS Lett 1991; 281: 9–19.
Touati E, Michel V, Thiberge JM, Wuscher N, Huerre M, Labigne A . Chronic Helicobacter pylori infections induce gastric mutations in mice. Gastroenterology 2003; 124: 1408–1419.
Wang D, Kreutzer DA, Essigmann JM . Mutagenicity and repair of oxidative DNA damage: insights from studies using defined lesions. Mutat Res 1998; 400: 99–115.
Altieri F, Grillo C, Maceroni M, Chichiarelli S . DNA damage and repair: from molecular mechanisms to health implications. Antioxidants Redox Signal 2008; 10: 891–937.
Teoule R, Bert C, Bonicel A . Thymine fragment damage retained in the DNA polynucleotide chain after gamma irradiation in aerated solutions. II. Radiat Res 1977; 72: 190–200.
Tremblay S, Douki T, Cadet J, Wagner JR . 2'-Deoxycytidine glycols, a missing link in the free radical-mediated oxidation of DNA. J Biol Chem 1999; 274: 20833–20838.
Gajewski E, Rao G, Nackerdien Z, Dizdaroglu M . Modification of DNA bases in mammalian chromatin by radiation-generated free radicals. Biochemistry 1990; 29: 7876–7882.
Grollman AP, Moriya M . Mutagenesis by 8-oxoguanine: an enemy within. Trends Genet 1993; 9: 246–249.
Melvin T, Cunniffe SM, O'Neill P, Parker AW, Roldan-Arjona T . Guanine is the target for direct ionisation damage in DNA, as detected using excision enzymes. Nucleic Acids Res 1998; 26: 4935–4942.
Dizdaroglu M . Chemical determination of free radical-induced damage to DNA. Free Radic Biol Med 1991; 10: 225–242.
Lindahl T . Instability and decay of the primary structure of DNA. Nature 1993; 362: 709–715.
Radicella JP, Dherin C, Desmaze C, Fox MS, Boiteux S . Cloning and characterization of hOGG1, a human homolog of the OGG1 gene of Saccharomyces cerevisiae. Proc Natl Acad Sci USA 1997; 94: 8010–8015.
Aburatani H, Hippo Y, Ishida T, Takashima R, Matsuba C, Kodama T et al. Cloning and characterization of mammalian 8-hydroxyguanine-specific DNA glycosylase/apurinic, apyrimidinic lyase, a functional mutM homologue. Cancer Res 1997; 57: 2151–2156.
Demple B, Harrison L . Repair of oxidative damage to DNA: enzymology and biology. Annu Rev Biochem 1994; 63: 915–948.
Meira LB, Bugni JM, Green SL, Lee CW, Pang B, Borenshtein D et al. DNA damage induced by chronic inflammation contributes to colon carcinogenesis in mice. J Clin Invest 2008; 118: 2516–2525.
Fortini P, Parlanti E, Sidorkina OM, Laval J, Dogliotti E . The type of DNA glycosylase determines the base excision repair pathway in mammalian cells. J Biol Chem 1999; 274: 15230–15236.
Nishimura S . Involvement of mammalian OGG1(MMH) in excision of the 8-hydroxyguanine residue in DNA. Free Radic Biol Med 2002; 32: 813–821.
Bjoras M, Luna L, Johnsen B, Hoff E, Haug T, Rognes T et al. Opposite base-dependent reactions of a human base excision repair enzyme on DNA containing 7,8-dihydro-8-oxoguanine and abasic sites. EMBO J 1997; 16: 6314–6322.
Mokkapati SK, Wiederhold L, Hazra TK, Mitra S . Stimulation of DNA glycosylase activity of OGG1 by NEIL1: functional collaboration between two human DNA glycosylases. Biochemistry 2004; 43: 11596–11604.
Sidorenko VS, Nevinsky GA, Zharkov DO . Mechanism of interaction between human 8-oxoguanine-DNA glycosylase and AP endonuclease. DNA Repair 2007; 6: 317–328.
Fortini P, Pascucci B, Parlanti E, D'Errico M, Simonelli V, Dogliotti E . The base excision repair: mechanisms and its relevance for cancer susceptibility. Biochimie 2003; 85: 1053–1071.
Klaunig JE, Kamendulis LM . The role of oxidative stress in carcinogenesis. Annu Rev Pharmacol Toxicol 2004; 44: 239–267.
Gentil A, Cabral-Neto JB, Mariage-Samson R, Margot A, Imbach JL, Rayner B et al. Mutagenicity of a unique apurinic/apyrimidinic site in mammalian cells. J Mol Biol 1992; 227: 981–984.
Guillet M, Boiteux S . Origin of endogenous DNA abasic sites in Saccharomyces cerevisiae. Mol Cell Biol 2003; 23: 8386–8394.
Glassner BJ, Rasmussen LJ, Najarian MT, Posnick LM, Samson LD . Generation of a strong mutator phenotype in yeast by imbalanced base excision repair. Proc Natl Acad Sci USA 1998; 95: 9997–10002.
Toller IM, Neelsen KJ, Steger M, Hartung ML, Hottiger MO, Stucki M et al. Carcinogenic bacterial pathogen Helicobacter pylori triggers DNA double-strand breaks and a DNA damage response in its host cells. Proc Natl Acad Sci USA 2011; 108: 14944–14949.
Ding SZ, O'Hara AM, Denning TL, Dirden-Kramer B, Mifflin RC, Reyes VE et al. Helicobacter pylori and H2O2 increase AP endonuclease-1/redox factor-1 expression in human gastric epithelial cells. Gastroenterology 2004; 127: 845–858.
Machado AM, Figueiredo C, Touati E, Maximo V, Sousa S, Michel V et al. Helicobacter pylori infection induces genetic instability of nuclear and mitochondrial DNA in gastric cells. Clin Cancer Res 2009; 15: 2995–3002.
Ensminger M, Iloff L, Ebel C, Nikolova T, Kaina B, Lbrich M . DNA breaks and chromosomal aberrations arise when replication meets base excision repair. J Cell Biol 2014.
Yamtich J, Nemec AA, Keh A, Sweasy JB . A germline polymorphism of DNA polymerase beta induces genomic instability and cellular transformation. PLoS Genet 2012; 8: e1003052.
Kubo K, Ide H, Wallace SS, Kow YW . A novel, sensitive, and specific assay for abasic sites, the most commonly produced DNA lesion. Biochemistry 1992; 31: 3703–3708.
Kow YW, Dare A . Detection of abasic sites and oxidative DNA base damage using an ELISA-like assay. Methods 2000; 22: 164–169.
Kurisu S, Miya T, Terato H, Masaoka A, Ohyama Y, Kubo K et al. Quantitation of DNA damage by an aldehyde reactive probe (ARP). Nucleic Acids Res Suppl 2001; 1: 45–46.
Rogakou EP, Boon C, Redon C, Bonner WM . Megabase chromatin domains involved in DNA double-strand breaks in vivo. J Cell Biol 1999; 146: 905–916.
Lomax ME, Cunniffe S, O'Neill P . Efficiency of repair of an abasic site within DNA clustered damage sites by mammalian cell nuclear extracts. Biochemistry 2004; 43: 11017–11026.
Shikazono N, Noguchi M, Fujii K, Urushibara A, Yokoya A . The yield, processing, and biological consequences of clustered DNA damage induced by ionizing radiation. J Radiat Res 2009; 50: 27–36.
Moldovan GL, Pfander B, Jentsch S . PCNA the maestro of the replication fork. Cell 2007; 129: 665–679.
Wold MS . Replication protein A: a heterotrimeric, single-stranded DNA-binding protein required for eukaryotic DNA metabolism. Annu Rev Biochem 1997; 66: 61–92.
Raderschall E, Golub EI, Haaf T . Nuclear foci of mammalian recombination proteins are located at single-stranded DNA regions formed after DNA damage. Proc Natl Acad Sci USA 1999; 96: 1921–1926.
Rassool FV, North PS, Mufti GJ, Hickson ID . Constitutive DNA damage is linked to DNA replication abnormalities in Bloom's syndrome cells. Oncogene 2003; 22: 8749–8757.
Davies SL, North PS, Dart A, Lakin ND, Hickson ID . Phosphorylation of the Bloom's syndrome helicase and its role in recovery from S-phase arrest. Mol Cell Biol 2004; 24: 1279–1291.
Blaisdell JO, Harrison L, Wallace SS . Base excision repair processing of radiation-induced clustered DNA lesions. Radiat Prot Dosimetry 2001; 97: 25–31.
Weinfeld M, Rasouli-Nia A, Chaudhry MA, Britten RA . Response of base excision repair enzymes to complex DNA lesions. Radiat Res 2001; 156: 584–589.
Hill JW, Hazra TK, Izumi T, Mitra S . Stimulation of human 8-oxoguanine-DNA glycosylase by AP-endonuclease: potential coordination of the initial steps in base excision repair. Nucleic Acids Res 2001; 29: 430–438.
Vidal AE, Hickson ID, Boiteux S, Radicella JP . Mechanism of stimulation of the DNA glycosylase activity of hOGG1 by the major human AP endonuclease: bypass of the AP lyase activity step. Nucleic Acids Res 2001; 29: 1285–1292.
Dou H, Mitra S, Hazra TK . Repair of oxidized bases in DNA bubble structures by human DNA glycosylases NEIL1 and NEIL2. J Biol Chem 2003; 278: 49679–49684.
Wiederhold L, Leppard JB, Kedar P, Karimi-Busheri F, Rasouli-Nia A, Weinfeld M et al. AP endonuclease-independent DNA base excision repair in human cells. Mol Cell 2004; 15: 209–220.
Dizdaroglu M, Karahalil B, Senturker S, Buckley TJ, Roldan-Arjona T . Excision of products of oxidative DNA base damage by human NTH1 protein. Biochemistry 1999; 38: 243–246.
Ide H . DNA substrates containing defined oxidative base lesions and their application to study substrate specificities of base excision repair enzymes. Prog Nucleic Acid Res Mol Biol 2001; 68: 207–221.
Luna L, Bjoras M, Hoff E, Rognes T, Seeberg E . Cell-cycle regulation, intracellular sorting and induced overexpression of the human NTH1 DNA glycosylase involved in removal of formamidopyrimidine residues from DNA. Mutat Res 2000; 460: 95–104.
Touati E, Michel V, Thiberge JM, Ave P, Huerre M, Bourgade F et al. Deficiency in OGG1 protects against inflammation and mutagenic effects associated with H. pylori infection in mouse. Helicobacter 2006; 11: 494–505.
Higuchi K, Katayama T, Iwai S, Hidaka M, Horiuchi T, Maki H . Fate of DNA replication fork encountering a single DNA lesion during oriC plasmid DNA replication in vitro. Genes Cells 2003; 8: 437–449.
Shibutani S, Takeshita M, Grollman AP . Translesional synthesis on DNA templates containing a single abasic site. A mechanistic study of the ‘A rule’. J Biol Chem 1997; 272: 13916–13922.
Swanson RL, Morey NJ, Doetsch PW, Jinks-Robertson S . Overlapping specificities of base excision repair, nucleotide excision repair, recombination, and translesion synthesis pathways for DNA base damage in Saccharomyces cerevisiae. Mol Cell Biol 1999; 19: 2929–2935.
Jeggo P, O'Neill P . The Greek Goddess, Artemis, reveals the secrets of her cleavage. DNA Repair 2002; 1: 771–777.
Letavayova L, Markova E, Hermanska K, Vlckova V, Vlasakova D, Chovanec M et al. Relative contribution of homologous recombination and non-homologous end-joining to DNA double-strand break repair after oxidative stress in Saccharomyces cerevisiae. DNA Repair 2006; 5: 602–610.
Budzowska M, Kanaar R . Mechanisms of dealing with DNA damage-induced replication problems. Cell Biochem Biophys 2009; 53: 17–31.
Henry-Mowatt J, Jackson D, Masson JY, Johnson PA, Clements PM, Benson FE et al. XRCC3 and Rad51 modulate replication fork progression on damaged vertebrate chromosomes. Mol Cell 2003; 11: 1109–1117.
Wang BJ, Zhang ZQ, Ke Y . Conversion of cadherin isoforms in cultured human gastric carcinoma cells. World J Gastroenterol 2006; 12: 966–970.
Lian H, Cui J, Wang Y, Liu J, Wang J, Shen H et al. Downregulation of Rad51 participates in OTA-induced DNA double-strand breaks in GES-1 cells in vitro. Toxicol Lett 2014; 226: 214–221.
Shinmura K, Kasai H, Sasaki A, Sugimura H, Yokota J . 8-hydroxyguanine (7,8-dihydro-8-oxoguanine) DNA glycosylase and AP lyase activities of hOGG1 protein and their substrate specificity. Mutat Res 1997; 385: 75–82.
Arai K, Morishita K, Shinmura K, Kohno T, Kim SR, Nohmi T et al. Cloning of a human homolog of the yeast OGG1 gene that is involved in the repair of oxidative DNA damage. Oncogene 1997; 14: 2857–2861.
Acknowledgements
We thank Shanghai Jiao Tong University School of Medicine for generous gift of GES-1 cells. We also thank Professor Raymond Monnat and Dr Julia Sidrova for scientific advice. DK was supported by United States National Institutes of Health (NIH/National Cancer Institute (NCI)) K01 CA15485401 and the work was also supported by CA 116753 (to JBS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogenesis website
Supplementary information
Rights and permissions
Oncogenesis is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Kidane, D., Murphy, D. & Sweasy, J. Accumulation of abasic sites induces genomic instability in normal human gastric epithelial cells during Helicobacter pylori infection. Oncogenesis 3, e128 (2014). https://doi.org/10.1038/oncsis.2014.42
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/oncsis.2014.42
- Springer Nature Limited
This article is cited by
-
Probing photoprotection properties of lipophilic chain conjugated thiourea-aryl group molecules to attenuate ultraviolet-A induced cellular and DNA damages
Scientific Reports (2022)
-
Nucleosomal embedding reshapes the dynamics of abasic sites
Scientific Reports (2020)
-
Sequencing abasic sites in DNA at single-nucleotide resolution
Nature Chemistry (2019)
-
Effective gene editing by high-fidelity base editor 2 in mouse zygotes
Protein & Cell (2017)
-
Subversion of host genome integrity by bacterial pathogens
Nature Reviews Molecular Cell Biology (2016)