Abstract
Background/objectives
There is a growing appreciation for individual responses to diet. In a previous study, mouse strain-specific responses to American and ketogenic diets were observed. In this study, we searched for genetic variants underlying differences in the responses to American and ketogenic diets between C57BL/6J (B6) and FVB/NJ (FVB) mouse strains.
Results
Genetic mapping of fat and lean mass gain revealed QTLs on Chromosome (Chr) 1 at 191.6 Mb (Fmgq1) (P < 0.001, CI = 180.2–194.4 Mb), Chr5 at 73.7 Mb (Fmgq2, Lmgq1) (P < 0.001, CI = 66.1–76.6 Mb), and Chr7 at 40.5 Mb (Fmgq3) (P < 0.01, CI = 36.6–44.5 Mb). Analysis of serum HDL cholesterol concentration identified a significant (P < 0.001, CI = 160.6–176.1 Mb) QTL on Chr1 at 168.6 Mb (Hdlq1). Causal network inference suggests that HDL cholesterol and fat mass gain are both linked to Fmgq1.
Conclusions
Strong sex effects were identified at both Fmgq2 and Lmgq1, which are also diet-dependent. Interestingly, Fmgq2 and Fmgq3 affect fat gain directly, while Fmgq1 influences fat gain directly and via an intermediate change in serum cholesterol. These results demonstrate how precision nutrition will be advanced through the integration of genetic variation and sex in physiological responses to diets varied in carbohydrate composition.
Similar content being viewed by others
Introduction
Efforts to provide individualized dietary recommendations based on genetic markers have been underway for several years. However, these efforts have largely detected alleles with small-effect sizes that are not clinically actionable [1,2,3]. As with studies investigating responses to other environmental factors, responses to diet are likely not due to the action of single alleles with a large effect, but rather the sum of multiple small-effect alleles, interactions among alleles, epigenetic modifications, and sex.
Recently, our group observed strong mouse strain-specific differences in response to feeding different human-relevant diets [4,5,6]. This initial study provided evidence for striking differences between C57BL/6J (B6) and FVB/NJ (FVB) mice in response to high-fat diets varying in carbohydrate content. Although studies increasingly consider the role of sex as a biological variable, the role of sex in the response to diets with varied macronutrient contents has been understudied [7].
To further investigate the genetic origin of differential response to carbohydrate restriction and sensitivity to high-fat diets, we generated an intercross population (F2) between B6 and FVB. All F2s were fed either an American (35% of energy from fat, 50% from carbohydrate) or a ketogenic (84% of energy from fat, 0% from carbohydrate) diet and changes to body composition and serum cholesterol concentrations were measured. In this population, we performed genome-wide linkage analysis to elucidate the genetic architecture contributing to differential responses to the specific diets. The data obtained provide evidence for genetic loci that not only directly affect body composition response but also loci that indirectly affect response through differences in serum cholesterol concentration. In addition, significant sex differences in the effects of the identified quantitative trait loci (QTL) were detected.
Methods
Animals and diets
Initially, we screened 6-week-old B6 and FVB mice for their response to American (35% of energy from fat, 50% from carbohydrates) and ketogenic (84% of energy from fat, 0% from carbohydrates) diets after a 6-month feeding trial. These strains have previously exhibited significantly different responses to these two diets [4]. Detailed diet compositions are provided in Supplementary Table 1. B6 females were crossed with FVB males to generate F1 mice and subsequently intercrossed to generate the F2 population. Four-week-old F1s and 3–5 week-old F2s were screened for their response to American and ketogenic diets during a 3-month feeding trial.
For the feeding trials, mice were randomly assigned to one of the two diet groups. Half of the B6, FVB, F1, and F2 mice were placed on American diet (B6: 11 males, 9 females, FVB: 10 males, 10 females, F1: 6 males, 6 females, F2: 102 males, 122 females) and a half on the ketogenic diet (B6: 9 males, 10 females, FVB: 10 males, 10 females, F1: 6 females, 9 males, F2: 126 males, 119 females). Researchers were not blinded to diet assignments. All animals were maintained in accordance with Texas A&M University Institution Animal Care and Use Committee guidelines at 22 °C under a 12-h light cycle. At the end of the feeding trial, mice were euthanized by carbon dioxide asphyxiation, blood was collected, and tissues were harvested and immediately flash-frozen in liquid nitrogen.
Phenotyping
Echo magnetic resonance spectroscopy (MRI) (EchoMRI, Houston, TX, USA) was used to measure the fat and lean mass of all individuals. During the initial screen of the B6 and FVB strains, body weight and body composition were measured at a 3-month time point of the 6-month feeding trial. In the F1 population, body weight and body composition were measured at the beginning and end of the 3-month feeding trial. The fat percentage of total body weight was calculated at the 3-month timepoint for B6, FVB, and F1 populations. Fat percentage is the percentage of total body fat mass measured by MRI relative to body weight at the time of the MRI measurement. In the F2 population, body weight and body composition from before and after the 3-month feeding trial allowed for changes in fat and lean mass to be calculated. Fat and lean mass gains were calculated as the difference in fat and lean mass prior to the feeding trial and after 3 months on the assigned diet.
In the F2 population, total cholesterol, HDL, and LDL fractions, as well as APOA2 were measured in serum obtained from blood at sacrifice at the end of the feeding trial. Total cholesterol, HDL, and LDL measurements were performed in duplicate using the EnzyChrom AF HDL and LDL/VLDL Assay kit (BioAssay Systems, Hayward, CA, USA). APOA2 measurements were performed in duplicate in a subset of the F2 population with the highest and lowest serum HDL cholesterol concentration (11 males, 27 females), using the Mouse Apolipoprotein A2 ELISA kit (ABclonal cat no. RK02605, Woburn, MA, USA).
Genotyping
The F2 population was genotyped on the Mouse Universal Genotyping Array (MUGA) that includes 7854 SNP markers [8]. Markers that were not polymorphic between B6 and FVB were removed from the dataset and uncertain genotype calls for individuals (GenCall score quality metric <0.7) were set to missing. The remaining markers were used to generate a genetic map to check for problematic markers and/or sample DNAs. After all corrections, 1667 markers were used for the association analyses. Updated MUGA marker annotation was obtained from Dr. Karl Broman (https://kbroman.org/MUGAarrays/new_annotations.html).
Heritability calculations
Broad-sense heritability for body fat percentage was calculated as the ratio of total genetic variance (variance in the F2 population) to environmental variance (variance in the F1 population).
Quantitative trait loci analysis
Outliers can have a strong influence on the results of QTL analyses. We observed biologically implausible errors in data that were greater than three standard deviations from the mean and suspect that these reflect the technical errors in the measurements. As such, outliers were defined as individuals with phenotypes that were more than three standard deviations away from the mean for each sex and phenotype in the F2 population. Outliers were set to missing. This procedure was repeated iteratively to discover all outliers in the data. Pearson’s correlation between phenotypes was determined after correcting for sex and diet effects (Supplementary Fig. 1). Code available by request.
The combined model includes all F2s, both sexes, and both diets (y~ sex * diet + marker). QTL peaks with a logarithm of the odds (LOD) greater than thresholds determined by 10,000 permutations were considered genome-wide significant (P < 0.05, LOD > 3.90) or highly significant (P < 0.01, LOD > 4.70). A LOD drop of 1.5 LOD from the top marker was used to determine the 95% confidence intervals, or support intervals, for each QTL. Linear models using ANOVA was used to check for any interactions between sex and/or diet with the top markers of each QTL. The variance explained by the top markers at each QTL was calculated by dividing the sum of squares of the model including the top marker by the total sum of squares of the model without QTL.
Candidate gene analysis using KEGG
KEGG pathways are a collection of pathway maps that reflect known genetic and metabolic relationships. All genes within each significant QTL confidence interval were annotated with KEGG pathway identifiers. We further characterized our candidate genes by KEGG pathways related to glucose, insulin, fatty acids, adipocytes, cholesterol, obesity, diabetes mellitus, metabolic syndrome, and digestion and absorption of carbohydrates, fats, and proteins. A comprehensive list of KEGG pathway queries is provided in Supplementary Table S4.
Conditioned and unconditioned linkage analysis
We performed a set of conditional QTL scans using traits as covariates in the analysis of other traits based on the individual QTL that were identified for each trait and suspected biological relationships. If conditioning the genome scan with a covariate resulted in a significant increase or decrease in the absolute value of the LOD score, it was interpreted that the traits are causally related to one another. When comparing conditioned and unconditioned QTL scans, an increase or decrease in the LOD of at least 2.0 corresponds to a 5% type I error rate [9, 10]. Only conditioned genome scans that resulted in a change of 2.0 LOD were considered pleiotropic, or shared, QTL between the two traits.
Causal network inferences
For traits with overlapping QTL we made causal inferences in networks based on methods described elsewhere [11]. Briefly, the first trait (T1) was regressed on the second trait (T2) and T2 was regressed on T1 in order to obtain the residual of each trait after adjusting for the other (R1 and R2). A bivariate t test between R1, R2, and the shared locus was used to infer the causal network among them. P values were Holm–Bonferroni corrected for the number of tests (i.e., number of residuals tested = 2) where P = α0.05/(number of tests – rank of ith hypothesis +1). A P value < 0.05 is considered significant. An initial pathway describing the relationship between T1 and T2 was defined based on the inferred causal networks, and the predicted causal models were compared with the Akaike’s information criterion (AIC) and Bayesian information criterion (BIC). A lower AIC and BIC indicates a better model.
Structural equation modeling
Structural equation modeling was used to illustrate observed and unobserved relationships between the QTL models. The models were refined until all path coefficients were significantly different from zero. Linear models using ANOVA was used to check for the amount of variation explained by each predictor in the structural model. The proportion of variation explained with the predictors modeled was calculated by dividing the sum of squares of the model including the predictor by the total sum of squares without the predictor.
Quantitative PCR (qPCR)
RNA from flash-frozen gonadal fat and liver were extracted using the simplyRNA Tissue kit on a Maxwell AS3000 (Promega). cDNA was generated using the Transcriptor First Strand cDNA Synthesis kit (Roche). qPCRs were performed on a LightCycler 480 (Roche) and CFX384 (BioRad) with LightCycler 480 SYBR Green I Master reagent (Roche). B2m was used as a housekeeping gene to correct for starting amounts of cDNA in both gonadal fat and liver. Primers were purchased from Integrated DNA Technologies as custom DNA oligos and sequences are provided in Supplementary Table S2.
Results
Genetic background and sex modulate parental strain fat differences
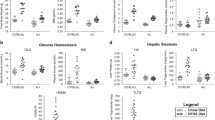
The fat percentage of B6 males consuming an American diet was 1.77-fold higher compared to B6 males consuming a ketogenic diet (B6 male American: 27.4 ± 5.2%, B6 male ketogenic: 15.5 ± 7.1%, P = 0.001, CI = 5.8–18.0%; Fig. 1 and Supplementary Table 3). Conversely, B6 females and FVB males and females fed the same two diets did not show diet-dependent differences of body fat percentage (B6 female American: 20.7 ± 8.8%, B6 female ketogenic: 17.2 ± 5.7%; FVB male American: 18.1 ± 6.9%, FVB male ketogenic: 20.4 ± 7.5%; FVB female American: 13.4 ± 6.7%, FVB female ketogenic: 16.2 ± 6.4%; Fig. 1 and Supplementary Table 3). The mean fat percentage in B6 males on the ketogenic diet is lower than the fat percentage in FVB males on the ketogenic diet, although the difference is not statistically significant (Fig. 1 and Supplementary Table 3).
Orange dots indicate a ketogenic diet. Green dots indicate American diet. Blue (male) and pink (female) boxes denote sex. B6 males on the ketogenic diet have a lower percentage of body fat than B6 males on the American diet (Welch’s two-sample t test when variances are unequal). This trend persists in the F1 population where F1 males on the ketogenic diet have a lower percentage of body fat than F1 males on the American diet. The F1 males on the ketogenic diet also have a significantly higher percentage of body fat than B6 males on the ketogenic diet.
Sex modulates hybrid population fat differences
The F1 males, but not females, also responded to the ketogenic diet. F1 males on the American diet had 1.25-fold higher body fat percentage than on the ketogenic diet (F1 male American: 32.5 ± 3.2%, F1 male ketogenic: 25.9 ± 4.7%, P = 0.020, CI = 1.3–11.9%; Fig. 1 and Supplementary Table 3). We also observed that the F1 males on the ketogenic diet had a 1.67-fold higher fat percentage than that observed in B6 males on the ketogenic diet (F1 male ketogenic: 25.9 ± 4.7%, B6 male ketogenic: 15.5 ± 7.1% P = 0.005, CI = 3.7–15.6%; Fig. 1 and Supplementary Table 3).
Sex and diet modulate phenotypes in the F2 population
As expected, sex had a profound effect on the phenotypes used for analysis (Supplementary Table 3). This effect was greater than the effect of diet for all phenotypes (Supplementary Table 3). Regardless of the carbohydrate composition of the diet, females had lower fat mass gain than males (F2 females, 4.5 ± 3.0 g, F2 males, 8.2 ± 4.5 g, P < 0.001, CI = 2.8–4.2 g), lower lean mass gain than males (F2 females, 5.5 ± 1.6 g, F2 males, 9.0 ± 3.2 g, P < 0.001, CI = 3.0–3.9 g), and lower serum HDL cholesterol concentration than males (F2 females, 157.6 ± 58.3 ng/mL, F2 males, 201.6 ± 32.5 ng/mL, P < 0.001, CI = 34.9–53.0 ng/mL).
Diet had a significant effect on lean mass gain and serum HDL cholesterol (Supplementary Table 3). Irrespective to sex, F2s on the ketogenic diet gained less lean mass than F2s on the American diet (F2 ketogenic, 6.9 ± 2.8 ng/mL, F2 American, 7.4 ± 3.2 ng/mL, P = 0.002, CI = 0.3–1.2 ng/mL) and had lower serum HDL cholesterol concentration on the ketogenic diet compared to F2s on the American diet (F2 ketogenic, 175.1 ± 52.2 ng/mL, F2 American, 183.3 ± 52.4 ng/mL, P = 0.027, CI = 1.2–19.2 ng/mL).
A significant interaction of sex and diet (Supplementary Table 3) was observed for lean mass gain where F2 males on the ketogenic diet gain significantly less lean mass than F2 males on the American diet (F2 male ketogenic, 8.2 ± 3.1 g, F2 male American, 10.0 ± 3.0 g, P < 0.001, CI = 0.9–2.7 g).
The amount of fat mass gain was highly heritable in the F2 population. Broad-sense heritability for the trait was 81% in males and 71% in females.
Linkage analysis reveals QTLs for a fat mass gain during the feeding trial
In the combined analysis that incorporates all F2s on both diets (y~ sex * diet + marker), we detected QTL for fat mass gain on Chr1 at 191.6 Mb (Fmgq1; P < 0.001, CI = 180.2–194.4 Mb), Chr5 at 73.7 Mb (Fmgq2; P < 0.001, CI = 66.1–76.6 Mb), and Chr7 at 40.5 Mb (Fmgq3; P < 0.01, CI = 36.6–44.5 Mb) (Fig. 2A and Table 1). At Fmgq1, the FVB allele contributes to higher fat mass gain, and the top marker (UNC010475128) accounts for 3.44% of the variance. Fmgq2 is a highly significant QTL where the FVB allele contributes to lower fat mass gain. The top marker (backupUNC050383757) accounts for 6.3% of the total variance in the F2 population (Table 1). The top marker explains 22.8% of the variation in males on the ketogenic diet, while it only explaining 5.9% of the variation in females on the ketogenic diet and 4.1% and 1.6% of the variation in males and females on the American diet, respectively. At Fmgq3, the FVB allele contributes to lower fat mass gain, and the top marker (JAX00150446) accounts for 3.4% of the variance in fat mass gain.
The QTL profiles show the logarithmic P values across the whole genome. Positive values indicate that the FVB allele increases the trait, while negative values indicate that the FVB allele decreases the trait. The horizontal lines present the genome-wide thresholds of high significance (P < 0.01, green) and significance (P < 0.05, orange) based on 10,000 permutations of the data. A QTL profile for fat mass gain during the feeding trial. In the combined model including both sexes and both diets (y~ sex * diet + marker), a highly significant QTL is located on Chr5 at 73.7 Mb (Fmgq2) (black confidence interval). There are additional significant QTLs on Chr1 at 191.6 Mb (Fmgq1) and on Chr7 at 40.5 Mb (Fmgq3). The top marker at Fmgq2 explains the most variation in fat mass gain in males on the ketogenic diet. B Interaction plots for Fmgq2 and Fmgq1 in ketogenic males and C ketogenic females represented as mean ± standard deviation. In ketogenic males, if Fmgq2 is homozygous for the B6 allele and Fmgq1 harbors at least one FVB allele, a functional interaction effect was evident. As long as Fmgq2 carries one B6 and one FVB allele, the effects of the two loci are additive, independent of the genotype at Fmgq1. D QTL profile on Chr5 for lean mass gain during the 3-month feeding trial. In the combined model including both sexes and both diets (y~ sex * diet + marker), a highly significant QTL is located on Chr5 at 73.7 Mb (Lmgq1) (black confidence interval). The top marker at Lmgq1 explains the most variation in a lean mass gain in males on the ketogenic. E QTL profile on Chr1 for serum HDL cholesterol concentration after the 3-month feeding trial. In the combined model that includes both sexes and diets (y~ sex * diet + marker), the peak position of the highly significant QTL is located at 168.6 Mb (Hdlq1) (black confidence interval).
We expected that FVB alleles would drive higher fat mass gain across the genome based on the initial screen of B6 and FVB showing that B6 males have a differential response to American and ketogenic diets, while FVB mice do not have a differential response to these two high-fat diets. Surprisingly, the FVB allele contributes to lower fat mass gain in response to the ketogenic diet at the most highly significant QTL at Fmgq2 as well as Fmgq3, but higher fat mass gain at Fmgq1.
In the initial analysis of fat percentage in B6, FVB, and F1 populations, we also observed that the male response to the ketogenic diet was greater in the F1 population compared to the B6 parent strain (Fig. 1). This prompted us to test if the combination of parent alleles at Fmgq1 with either Fmgq2 or Fmgq3 resulted in a higher fat mass gain in the F2 population. We found an interaction between Fmgq2 and Fmgq1 (P = 0.007, CI = 1.58–9.07 g) affecting fat mass gain. The interaction effect was specific to males on the ketogenic diet. Males that are homozygous for the B6 allele at Fmgq2 and that carry at least one FVB allele at Fmgq1 gained about 5 g more fat on a ketogenic diet than males homozygous for the B6 allele at both loci (Fig. 2B, C).
Linkage analysis reveals QTLs for a lean mass gain during the feeding trial
In the combined model that includes male and female F2s on both diets (y~ sex * diet + marker), we detected a significant QTL for lean body mass gain during the 3-month feeding trial (Lmgq1; P < 0.001, CI = 66.1–76.6 Mb) (Table 1 and Fig. 2D). Lmgq1 overlapped with Fmgq2. We observe again that males on the ketogenic diet drive this QTL. In the combined model, the top marker (backupUNC050383757) explains 3.3% of the variance in lean mass gain. In males on the ketogenic diet, the top marker (JAX00131700) explains 19.0% of the variation in lean mass gain while it only explaining 1.2% of the variation in females on the ketogenic diet and 8.2% and 4.1% of the variation in males and females on the American diet, respectively.
Linkage analysis reveals QTLs for serum HDL concentration at the end of the feeding trial
In the combined model that includes male and female F2 animals on both diets (y~ sex * diet + marker), we detected a highly significant QTL on Chr1 at 168.6 Mb (Hdlq1) for serum HDL concentration after the 3-month feeding trial (P < 0.001, CI = 160.6–176.1 Mb; Fig. 2E). All genotype classes (B6/B6 155.4 ± 48.8 ng/mL; B6/FVB 179.2 ± 51.5 ng/mL; FVB/FVB 200.6 ± 48.5 ng/mL) are significantly different from each other at this locus (B6/B6: B6/FVB P < 0.001, CI = 9.7–38.1 ng/mL; B6J/B6J: FVB/FVB P < 0.001, CI = 28.7–61.8 ng/mL; FVB/FVB: B6/FVB P < 0.001, CI = 7.4–35.4 ng/mL; Supplementary Fig. S5). This locus harbors the Apoa2 gene which has previously been associated with serum HDL cholesterol concentration [12,13,14]. Therefore, we measured serum APOA2 concentrations in F2s with the highest and lowest serum HDL cholesterol concentrations. These measurements confirm significant differences between homozygous genotype classes at Hdlq1 (B6/B6 2.2 ± 0.8 mg/dL; FVB/FVB 3.0 ± 0.9 mg/dL, P = 0.004, CI = 0.3–2.2 mg/dL; Fig. 3A).
A Serum APOA2 concentration for F2 animals which are homozygous for the alternative alleles at Hdlq1. The FVB/FVB genotype at Hdlq1 drives higher serum APOA2 concentration in addition to higher serum HDL cholesterol concentration (B6/B6 n = 14, FVB/FVB n = 24, Wilcoxon two-sample t test). B Fold change in Srd5a3 expression among F2 animals of genotype FVB/FVB (n = 15) relative to B6/B6 (n = 16) at Fmgq2. A decrease in Srd5a3 expression is associated with the FVB/FVB genotype at Fmgq2 relative to the B6/B6 genotype at Fmgq2 (Welch’s two-sample t test when variances are unequal). C Fold change in Srd5a3 expression among F2 animals of different genotypes at Fgmq1 and Fmgq2. A 1.59-fold increase in Srd5a3 expression is associated with F2 animals that are homozygous for the FVB allele at Fmgq1 and B6/B6 at Fmgq2 (n = 9) relative to those that are homozygous for the B6 allele at both loci (n = 7) (Welch’s two-sample t test when variances are unequal). D Fold change in Ppp2r5a expression among F2 animals of different genotypes at Fgmq1 and Fmgq2 (Welch’s two-sample t test when variances are unequal). A 1.61-fold increase in Ppp2r5a expression is associated with F2 animals that are homozygous for the FVB allele at Fmgq1 and B6/B6 at Fmgq2 (n = 9) relative to those that are homozygous for the B6 allele at both loci (n = 7) (Wilcoxon two-sample t test). E Graphical representation of SEM for fat mass gain during the feeding trial and serum HDL cholesterol concentrations after the feeding trial in the combined model that contains both sexes and both diets. Solid arrows indicate the direction of paths and the weight of each arrow is proportional to the path coefficient from the predictor to the variable and the percentage of variation in the variable that is explained by each predictor. Positive effects (green arrows) indicate that the FVB allele increases the trait; negative effects (red arrows) indicate that the FVB allele decreases the trait. The single-headed dashed arrow represents the inferred pleiotropic causal pathway at Fmgq1 that was identified in the causal network inference for fat mass gain and serum HDL cholesterol concentration. The double-headed dashed arrow represents a covariate pathway detected by the structural model between fat mass gain and serum HDL cholesterol concentration.
KEGG pathway annotation of candidate genes at Fmgq1 and Fmgq2 highlights steroid hormone biosynthesis
We searched for candidate genes that might elucidate the functional interaction between Fmgq1 and Fmgq2. Out of 98 positional candidate genes, 4 genes at Fmgq1 are found on metabolically relevant KEGG pathways: Hsd11b1 (steroid hormone biosynthesis; mmu00140), Ephx1 (bile secretion; mmu004976), Camk1g (aldosterone synthesis and secretion; mmu004925), and Ppp2r5a (AMPK signaling pathway; mmu004152) (Table 2). Out of 482 positional candidate genes, 2 genes at Fmgq2 are found on metabolically relevant KEGG pathways: Srd5a3 (steroid hormone biosynthesis; mmu00140) and Cox7b2 (nonalcoholic fatty liver disease; mmu004932) (Table 2).
On the steroid hormone biosynthesis pathway, Srd5a3 converts inactive testosterone to active dihydrotestosterone. This makes Srd5a3 a particularly interesting candidate at Fmgq2 given the sex specificity we see at this QTL and the functional interaction we observed between Fmgq2 and Fmgq1. Consistent with Srd5a3 underlying Fmgq2, we observed lower expression of Srd5a3 in gonadal fat of F2s that are homozygous for the FVB allele at Fmgq2 relative to F2s that are homozygous for the B6 allele at Fmgq2 (Fig. 3B). Interestingly, we also observe a 1.59-fold increase in Srd5a3 expression in F2s that are B6/B6 at Fmgq2 and carry FVB/FVB alleles at Fmgq1 relative to F2s that are B6/B6 at both loci (Fig. 3C). This makes Srd5a3 a strong candidate gene at Fmgq2 and for the functional interaction between Fmgq1 and Fmgq2.
At Fmgq1, we did not observe any genotype-dependent differences in expression of Hsd11b1 (liver), Ephx1 (liver), or Ppp2r5a (gonadal fat) (Supplementary Fig. S6ABC). We did, however, observe a 1.61-fold increase in Ppp2r5a expression in F2s that are B6/B6 at Fmgq2 and carry FVB/FVB alleles at Fmgq1 relative to F2s that are B6/B6 at both loci (Fig. 3D). This makes Ppp2r5a a strong candidate gene at Fmgq1 for the functional interaction between Fmgq1 and Fmgq2.
The remaining candidate at Fmgq1, Camk1g, is exclusively expressed in brain tissue, which was unavailable to measure expression. Given the involvement of Camk1g in aldosterone synthesis and secretion, we instead measured serum aldosterone concentration (after the feeding trial) and observed no genotype-dependent differences (Supplementary Fig. S6D). The remaining candidate at Fmgq2, Cox7b2 is expressed exclusively in the testis, which was unavailable to measure its expression.
Conditioned linkage analysis of fat and lean mass gain
Fmgq2 and Lmgq1 overlap in the combined model of F2s that includes both sexes and diets. For both traits, this QTL appears to be driven largely by males on the ketogenic diet. This shared QTL prompted us to model the relationship between fat and lean mass gain. There is no significant change to the LOD score on Chr5 at 73.7 Mb when the fat mass gain is conditioned on the lean mass gain, nor when the lean mass gain is conditioned on fat mass gain. Thus, the relationship between fat and lean mass gain at this locus could not be elucidated in this model. This would suggest that the overlapping QTL region harbors one or two tightly linked genes that affect both lean and fat mass in the same direction.
Conditioned linkage analysis of fat mass gain and serum HDL cholesterol concentration
The close proximity of Hdlq1 and Fmgq1 in the combined models prompted us to model the relationship between fat mass gain and serum HDL cholesterol concentration at these loci. We observed that when the fat mass gain is conditioned on serum HDL cholesterol concentration, there is no significant change to the LOD score on Chr1 at 191.6 Mb (Table 3). Thus, the relationship between fat mass gain and serum HDL cholesterol concentration could not be elucidated in this model. Similarly, when serum HDL cholesterol concentration is conditioned on the fat mass gain, the LOD score on Chr1 at 168.6 Mb drops by only 1.08 LOD suggesting that Hdlq1 is not shared between fat mass gain and serum HDL cholesterol concentration (Table 3).
Causal network between Fmgq1, fat mass gain, and serum HDL cholesterol concentration
We also explored causal networks between fat mass gain and serum HDL cholesterol concentration at Fmgq1. After adjusting for multiple testing, the inferred network shows that serum HDL cholesterol concentration and fat mass gain are independently related to Fmgq1 (Table 3). The AIC and BIC model selection scores suggest that the model in which serum HDL cholesterol concentration occurs upstream of fat mass gain at Fmgq1 is most consistent with the data (Table 3).
Structural equation modeling of fat mass gain and serum HDL cholesterol concentration
We built a structural model from this inferred network to illustrate the magnitude of the effects of each predictor in the models of fat mass gain and serum HDL cholesterol concentration (Fig. 3E). The path coefficients are all significantly different from zero (Table 3). The proportion of variation explained with the predictors modeled for fat mass gain and serum HDL cholesterol concentration is 29.95% and 27.00%, respectively.
Discussion
This study provides evidence that individual responses to a high-fat diet significantly depends upon three factors: (1) the presence or absence of carbohydrates in the high-fat diet; (2) the combination of alleles occurring in the study population; and (3) sex. In our experiment, B6 males have a lower percentage of body fat in response to the high fat, no carbohydrate, ketogenic diet, while in contrast, B6 females and FVB males and females do not have a differential response to American and ketogenic diets. This sex-specific response to carbohydrate restriction on ketogenic diet observed in B6 males persists in F1s males, and we observed that the combination of B6 and FVB alleles in the hybrids results in a higher percentage of body fat in response to the feeding trial. The difference in ages at the beginning of the parent and F1 feeding trials might also contribute to the higher percentage of body fat we observed in the F1s. In the F2 population, we observed a very high heritability for fat mass gain in both sexes and were able to identify significant QTLs for fat mass gain at Fmgq1, Fmgq2, and Fmgq3. This raises the question of whether these loci could be used to make predictions about the individual responses to carbohydrate restriction. Fmgq2 explains the most variation in the fat mass gain in males on the ketogenic diet. We observed that the FVB allele drives lower fat mass gain at Fmgq2 and Fmgq3, while at Fmgq1 the FVB allele drives higher fat mass gain. In addition, we provided evidence that male F2s exposed to the ketogenic diet that is homozygous for B6 alleles at Fmgq2 and carry at least one FVB allele at Fmgq1 gain the most fat mass during the feeding trial. We observed that Fmgq2 and Lmgq1 represent the same QTL on Chr5 for fat and lean mass gain. Likely, a single gene or two tightly linked genes are responsible for the change in lean and fat mass.
The QTL for serum HDL cholesterol concentration at Hdlq1 is significant in both males and females and contains Apoa2. Apoa2 has been described repeatedly for its relationship to serum HDL cholesterol concentration, especially in these two strains [12,13,14]. The initial observation of the Apoa2 locus affecting serum concentrations of HDL cholesterol was made in an association study of females using an advanced intercross line between B6 and NZB/BINJ [12]. Later, in a panel of inbred strains that included B6 and FVB mice, it was determined that the amino acid substitution Ala61Val in APOA2 led to increased serum HDL concentrations [13]. B6 carry Ala61 while FVB mice carry amino acid substitution 61Val. Both males and females of all strains carrying the 61Val had increased serum HDL cholesterol concentrations. The results of our linkage analysis for serum cholesterol concentrations are consistent with the amino acid differences. They show that Hdlq1 affects HDL in males and females. We confirmed that FVB alleles at Hdlq1 drive higher concentrations of serum HDL cholesterol and serum APOA2.
In our causal network of the relationship between fat mass gain and serum HDL cholesterol concentration, we found that serum HDL cholesterol concentration and fat mass gain are independently linked to Fmgq1. The close proximity of the Fmgq1 and Hdlq1 QTLs puts them in linkage disequilibrium and this makes their direct and indirect effects difficult to detangle. In general, we would have expected that serum HDL cholesterol concentration would have a negative relationship with fat mass gain given the well-established relationship between serum HDL cholesterol and obesity in humans. It is likely that the two traits are independently linked to Fmgq1 because of the linkage disequilibrium. However, the AIC and BIC scores for the models suggest that differences in serum HDL cholesterol concentration occur upstream of the differences we observe in fat mass gain at Fmgq1.
Overlaying genes within Fmgq1 and Fmgq2 onto metabolically relevant KEGG pathways revealed candidate genes involved in the biogenesis of cortisol (Hsd11b1), aldosterone (Camk1g), and testosterone (Srd5a3). Cholesterol falls upstream of the production of each of these steroid hormones. Ephx1 resides inside of Fmgq1 and plays a role in the transfer of bile acids to the liver, a process that is critical for the endogenous synthesis of cholesterol. Each of these candidate genes has in common, a relationship with cholesterol. This offers insight into the role these particular genes could play in the causal network we inferred that showed serum HDL cholesterol concentration occurring upstream of fat mass gain. The inferred network might be reflective of the relationship between fat mass gain and one of these candidate genes more so than a direct relationship between serum HDL cholesterol concentration and fat mass gain.
Further, our candidate genes offer insight into the sex-specific interaction in males on the ketogenic diet that we observed between Fmgq1 and Fmgq2. Our most prominent QTL at Fmgq2 harbors Srd5a3 that converts inactive testosterone to active dihydrotestosterone. We confirmed that F2s that are homozygous for FVB alleles at Fmgq2 express Srd5a3 less than F2s that are homozygous for B6 alleles. At this QTL, FVB alleles also drive lower fat mass gain. This suggests that more active dihydrotestosterone promotes more fat mass gain at Fmgq2. The functional interaction we observed between Fmgq1 and Fmgq2 showed us that male F2s that are homozygous B6 at Fmgq2 and carry at least one FVB allele at Fmgq1 gained the most fat mass. Interestingly, we observed that F2s that carry this interaction have a 1.59-fold increase in expression of Srd5a3 relative to animals that are homozygous B6 at both Fmgq1 and Fmgq2. This provides further evidence that more active dihydrotestosterone promotes more fat mass gain in this population. Ppp2r5a at Fmgq1 has been associated with polycystic ovary syndrome (PCOS) through pathway analysis of genes enriched in genome-wide association studies (GWAS). Ppp2r5a is a member of the oocyte meiosis KEGG pathway that was significantly associated with PCOS, a pathway affected by the hyperandrogenemia that frequently occurs with PCOS [15]. The association of Ppp2r5a with PCOS makes Srd5a3, and Ppp2r5a primary candidate genes of interest for further investigation.
Each of the remaining candidate genes has a potential relationship with Srd5a3 and active dihydrotestosterone levels. Hsd11b1 at Fmgq1 reversibly converts active cortisol to inactive cortisone. Epidemiological data in humans show that the ratio of androgens to glucocorticoids is critical to metabolic homeostasis and alterations in this ratio can lead to increased incidence of obesity and metabolic syndrome [16]. In addition, excessive levels of active dihydrotestosterone have been shown to stimulate aldosterone secretion by a mechanism that is dependent upon the calmodulin/calmodulin-dependent protein kinase and protein kinase C intracellular signaling pathways [17]. Camk1g at Fmgq1 is a member of this pathway and people with obesity often experience hyperaldosteronism [18]. Ephx1 at Fmgq1 has previously been associated with other Srd5a isoforms in GWAS of PCOS [19], and similarly, Cox7b2 at Fmgq2, has been identified in another GWAS for PCOS [20]. The association with PCOS establishes a probable link between both Ephx1 and Cox7b2 and abnormal androgen secretion.
Sequence variation between B6 and FVB result in amino acid changes in EPHX1, CAMK1G, and PPP2R5A, sometimes at multiple locations within the proteins. In HSD11B1 and SRD5A3, no amino acid changes have been documented, but sequence variation in Hsd11b1 and Srd5a3 does result in noncoding transcript variants that might be of interest in further investigation. These noncoding transcript variants would suggest differences in expression of relevant candidate genes like we observed for Srd5a3. However, we did not observe genotype-dependent differences in expression of Hsd11b1, but did find in the F2 population that mice that are B6/B6 at Fmgq2 and carry FVB/FVB alleles at Fmgq1 express Ppp2r5a 1.61-fold more than F2s that are B6/B6 at both loci.
Overall, our data parallel a clinical trial showing a greater fat mass loss among men than women on very low energy, ketogenic diet [21,22,23]. The differences could not be attributed to the attenuation of appetite-stimulating hormones, and concentrations of serum steroid hormones were not considered in the study. Unfortunately, studies in humans that utilize very low carbohydrate, ketogenic diets typically lack the power to detect sex differences or fail to incorporate realistic control diets [24,25,26,27,28]. Nonetheless, meta-analyses of human, genome-wide association studies of body mass index and waist-to-hip circumference have identified several sex-specific loci [29,30,31,32,33]. Future sex-specific genome-wide association studies that are powered to detect differential responses to macronutrients between men and women are needed.
Conclusions
It is possible that the genetic architecture for fat gain in response to high-fat diets is more complex in females than in males, and as such, the genetic component we are able to detect in our model contributes more to the overall response in males than in females. The observation of these sex-specific QTLs for fat and lean mass gain at Fmgq2 and Lmgq1 and serum HDL cholesterol concentration at Hdlq2 and Hdlq3 highlights the importance of sex-specific analyses during the development of individualized dietary guidelines. We identified a known QTL at Hdlq1 for serum HDL cholesterol concentration that harbors Apoa2. Interestingly, the QTLs at Fmgq2 and Fmgq3 affect fat gain directly while Fmgq1 seems to influence fat gain directly as well as via an intermediate physiological change in serum cholesterol related to the strain-specific Apoa2 phenotypes. We have shown that genotypes at Fmgq2 alter the expression of Srd5a3 in a sex- and genotype-specific manner. It remains unclear if these loci explain the differential response to carbohydrate restriction we observed in the initial parent strain screen or if instead, they characterize the increased fat mass gain that we observe after the parental genomes are combined in hybrid and F2 populations. Further investigation of the candidate genes presented here will elucidate their roles in response to carbohydrate restriction in the parent, hybrid, and F2 populations.
References
Yengo L, Sidorenko J, Kemper KE, Zheng Z, Wood AR, Weedon MN, et al. Meta-analysis of genome-wide association studies for height and body mass index in ~700 000 individuals of European ancestry. Hum Mol Genet. 2018;27:3641–9.
Hoffmann TJ, Choquet H, Yin J, Banda Y, Kvale MN, Glymour M, et al. A large multiethnic genome-wide association study of adult body mass index identifies novel loci. Genetics. 2018;210:499–515.
Heianza Y, Qi L. Gene-diet interaction and precision nutrition in obesity. Int J Mol Sci. 2017;18:787.
Barrington WT, Wulfridge P, Wells AE, Rojas CM, Howe SYF, Perry A, et al. Improving metabolic health through precision dietetics in mice. Genetics. 2018;208:399–417.
Wells A, Barrington WT, Dearth S, May A, Threadgill DW, Campagna SR, et al. Tissue level diet and sex-by-diet interactions reveal unique metabolite and clustering profiles using untargeted liquid chromatography-mass spectrometry on adipose, skeletal muscle, and liver tissue in C57BL6/J mice. J Proteome Res. 2018;17:1077–90.
Cuomo D, Porreca I, Ceccarelli M, Threadgill DW, Barrington WT, Petriella A, et al. Transcriptional landscape of mouse-aged ovaries reveals a unique set of non-coding RNAs associated with physiological and environmental ovarian dysfunctions. Cell Death Discov. 2018;4:1–14.
Comitato R, Saba A, Turrini A, Arganini C, Virgili F. Sex hormones and macronutrient metabolism. Crit Rev Food Sci. Nutr. 2015;55:227–41.
Morgan AP, Fu CP, Kao CY, Welsh CE, Didion JP, Yadgary L, et al. The mouse universal genotyping array: from substrains to subspecies. G3 Genes, Genomes, Genet. 2016;6:263–79.
Brockmann GA, Tsaih SW, Neuschl C, Churchill GA, Li R. Genetic factors contributing to obesity and body weight can act through mechanisms affecting muscle weight, fat weight, or both. Physiol Genomics. 2009;36:114–26.
Li R, Tsaih S-W, Shockley K, Stylianou IM, Wergedal J, Paigen B, et al. Structural model analysis of multiple quantitative traits. PLoS Genet. 2006;2:e114.
Li Y, Tesson BM, Churchill GA, Jansen RC. Critical reasoning on causal inference in genome-wide linkage and association studies. Trends Genet. 2010;26:493–8.
Wang X, Le Roy I, Nicodeme E, Li R, Wagner R, Petros C, et al. Using advanced intercross lines for high-resolution mapping HDL cholesterol quantitative trait loci. Genome Res. 2003;13:1654–64.
Wang X, Korstanje R, Higgins D, Paigen B. Haplotype analysis in multiple crosses to identify a QTL gene. Genome Res. 2004;14:1767–72.
Wang X, Paigen B. Genetics of variation in HDL cholesterol in humans and mice. Circ Res. 2005;96:27–42.
Shim U, Kim HN, Lee H, Oh JY, Sung YA, Kim HL. Pathway analysis based on a genome-wide association study of polycystic ovary syndrome. PLoS ONE. 2015;10:e0136609.
Alemany M. Do the interactions between glucocorticoids and sex hormones regulate the development of the metabolic syndrome? Front Endocrinol. 2012;3:27.
Yanes LL, Romero DG. Dihydrotestosterone stimulates aldosterone secretion by H295R human adrenocortical cells. Mol Cell Endocrinol. 2009;303:50–6.
Kawarazaki W, Fujita T. The role of aldosterone in obesity-related hypertension. Am J Hyperten. 2016;29:415–23.
Sang Q, Li X, Wang H, Wang H, Zhang S, Feng R, et al. Quantitative methylation level of the EPHX1 promoter in peripheral blood DNA is associated with polycystic ovary syndrome. PLoS ONE. 2014;9:e88013.
Lee H, Oh JY, Sung YA, Chung H, Kim HL, Kim GS, et al. Genome-wide association study identified new susceptibility loci for polycystic ovary syndrome. Hum Reprod. 2015;30:723–31.
Nymo S, Coutinho SR, Jørgensen J, Rehfeld JF, Truby H, Kulseng B, et al. Timeline of changes in appetite during weight loss with a ketogenic diet. Int J Obes. 2017;41:1224–31.
Nymo S, Coutinho SR, Rehfeld JF, Truby H, Kulseng B, Martins C. Physiological predictors of weight regain at 1-year follow-up in weight-reduced adults with obesity. Obesity. 2019;27:925–31.
Lyngstad A, Nymo S, Coutinho SR, Rehfeld JF, Truby H, Kulseng B, et al. Investigating the effect of sex and ketosis on weight-loss-induced changes in appetite. Am J Clin Nutr. 2019;109:1511–8.
Foster GD, Wyatt HR, Hill JO, McGuckin BG, Brill C, Mohammed BS, et al. A randomized trial of a low-carbohydrate diet for obesity. N Engl J Med. 2003;348:2082–90.
Samaha FF, Iqbal N, Seshadri P, Chicano KL, Daily DA, McGrory J, et al. A low-carbohydrate as compared with a low-fat diet in severe obesity. N Engl J Med. 2003;348:2074–81.
Gardner CD, Kiazand A, Alhassan S, Kim S, Stafford RS, Balise RR. et al. Comparison of the Atkins, Zone, Ornish, and LEARN diets for change in weight and related risk factors among overweight premenopausal women: the A to Z weight loss study: a randomized trial. J Am Med Assoc. 2007;297:969–77.
Hall KD, Chen KY, Guo J, Lam YY, Leibel RL, Mayer LES, et al. Energy expenditure and body composition changes after an isocaloric ketogenic diet in overweight and obese men. Am J Clin Nutr. 2016;104:324–33.
Rosenbaum M, Hall KD, Guo J, Ravussin E, Mayer LS, Reitman ML, et al. Glucose and lipid homeostasis and inflammation in humans following an isocaloric ketogenic diet. Obesity. 2019;27:971–81.
Winkler TW, Justice AE, Graff M, Barata L, Feitosa MF, Chu S. et al. The influence of age and sex on genetic associations with adult body size and shape: a large-scale genome-wide interaction study. PLoS Genet.2015;11:e1005378.
Randall JC, Winkler TW, Kutalik Z, Berndt SI, Jackson AU, Monda KL, et al. Sex-stratified genome-wide association studies including 270,000 individuals show sexual dimorphism in genetic loci for anthropometric traits. PLoS Genet. 2013;9:e1003500.
McGrice M, Porter J. The effect of low carbohydrate diets on fertility hormones and outcomes in overweight and obese women: a systematic review. Nutrients. 2017;9:204.
Gupta L, Khandelwal D, Kalra S, Gupta P, Dutta D, Aggarwal S. Ketogenic diet in endocrine disorders: current perspectives. J Postgraduate Med. 2017;63:242.
Mavropoulos JC, Yancy WS, Hepburn J, Westman EC. The effects of a low-carbohydrate, ketogenic diet on the polycystic ovary syndrome: a pilot study. Nutr Metab. 2005;2:1–5.
Acknowledgements
We thank Dr. Andrew Hillhouse and the Texas A&M Institute for Genome Sciences and Society (TIGSS) for assistance in the TIGSS Molecular Genomics Workspace and TIGSS Rodent Preclinical Phenotyping Core; Dr. Karl Broman for his discussion and guidance during the QTL analysis as well as for organizing and managing annotation files for the MUGA arrays; Daniel Genung and Aaron Van Wettering for assistance with mouse husbandry and phenotyping; Anthony Gacasan and Ryan McGovern for assistance with preparing samples for genotyping; and all members of the laboratory for their helpful insights. This work was supported by National Institutes of Health (NIH) grants RM1HG008529 and P30ES029067.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salvador, A.C., Arends, D., Barrington, W.T. et al. Sex-specific genetic architecture in response to American and ketogenic diets. Int J Obes 45, 1284–1297 (2021). https://doi.org/10.1038/s41366-021-00785-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41366-021-00785-7
- Springer Nature Limited
This article is cited by
-
A mathematically rigorous algorithm to define, compute and assess relevance of the probable dissociation constants in characterizing a biochemical network
Scientific Reports (2024)
-
Analysis of strain, sex, and diet-dependent modulation of gut microbiota reveals candidate keystone organisms driving microbial diversity in response to American and ketogenic diets
Microbiome (2023)