Abstract
Mouse behavioral genetic mapping studies can identify genomic intervals modulating complex traits under well-controlled environmental conditions and have been used to study ethanol behaviors to aid in understanding genetic risk and the neurobiology of alcohol use disorder (AUD). However, historically such studies have produced large confidence intervals, thus complicating identification of potential causal candidate genes. Diversity Outbred (DO) mice offer the ability to perform high-resolution quantitative trait loci (QTL) mapping on a very genetically diverse background, thus facilitating identification of candidate genes. Here, we studied a population of 636 male DO mice with four weeks of intermittent ethanol access via a three-bottle choice procedure, producing a progressive ethanol consumption phenotype. QTL analysis identified 3 significant (Chrs 3, 4, and 12) and 13 suggestive loci for ethanol-drinking behaviors with narrow confidence intervals (1–4 Mbp for significant QTLs). Results suggested that genetic influences on initial versus progressive ethanol consumption were localized to different genomic intervals. A defined set of positional candidate genes were prioritized using haplotype analysis, identified coding polymorphisms, prefrontal cortex transcriptomics data, human GWAS data and prior rodent gene set data for ethanol or other misused substances. These candidates included Car8, the lone gene with a significant cis-eQTL within a Chr 4 QTL for week four ethanol consumption. These results represent the highest-resolution genetic mapping of ethanol consumption behaviors in mice to date, providing identification of novel loci and candidate genes for study in relation to the neurobiology of AUD.
Similar content being viewed by others
Introduction
Alcohol use disorder (AUD) poses a significant global healthcare burden, contributing yearly to around 3 million deaths worldwide [1]. In the United States, 140,000 people die yearly from causes related to alcohol use, making it the third-largest cause of preventable death [2]. Genetic factors are thought to play a significant role, with twin studies routinely estimating ~50% heritability in the risk for AUD [3].
Considerable progress in recent large human genome-wide association studies (GWAS) has identified a growing number of genetic loci associated with alcohol consumption [4, 5], dependence [6, 7], AUD [8], and problematic alcohol use [9]. Animal models complement human genetic studies by providing improved control over environmental variance and dependent variables, allowing for mechanistic and direct candidate gene identification studies infeasible in humans. Mouse models have identified broadly-applicable biological mechanisms underlying specific behaviors associated with AUD including ethanol consumption [10], preference [11, 12], and withdrawal [13]. Intermittent ethanol access (IEA) procedures have been shown to model escalation of alcohol consumption as seen in the early stages of AUD [14, 15]. IEA consumption varies across different mouse lines, suggesting that genetic variance impacting progressive consumption can be studied in such models [16, 17]. Additionally, rodent models have identified brain regions functioning in the transition to compulsive consumption, such as prefrontal cortex and nucleus accumbens [18, 19]. Medial prefrontal cortex (mPFC) has been implicated in molecular and neurocircuitry plasticity underlying dependence-like consumption and cue-induced reinstatement in rodent models [19, 20].
Behavioral quantitative trait loci (QTL) studies in rodents have been used to implicate genetic loci modulating ethanol behaviors, but generally with broad confidence intervals ( > 10–40 Mbp) confounding candidate gene identification [21,22,23]. Recent development of rodent genetic models with increased genetic diversity and recombination have greatly improved QTL analysis [24,25,26]. Diversity Outbred (DO) mice, derived from random outbreeding of 8 genetically and phenotypically diverse founder strains [27], provide high levels of allelic variation and recombination events for high-resolution genetic mapping [25]. The progenitor strains exhibit marked diversity in ethanol-drinking phenotypes [16, 17] and DO mice have recently been used to identify genetic loci for ethanol sensitivity and having narrow confidence intervals ( ~ 4 Mbp) [28].
Here we describe the first use of DO mice and an IEA procedure to map behavioral QTLs (bQTLs) associated with ethanol consumption. Our results show significant bQTLs on multiple chromosomes with narrow support intervals. Candidate genes were then prioritized by a multi-step strategy using haplotype analysis, integration of DO mouse transcriptome data from medial prefrontal cortex, and bioinformatics studies including merging of genetic data on AUD and other animal model data on ethanol. Our results identify novel candidate genes and biological mechanisms for modulation of ethanol consumption behaviors. The work may both validate and broaden existing human studies on the genetics of alcohol consumption and AUD, while also providing novel hypotheses for future studies on the neurobiology of ethanol in rodent models.
Methods
Full details on Methods and Materials are provided in Supplemental Materials. Additional data supporting these studies are provided on the Open Science Framework (https://osf.io/tfrzg/?view_only=232cfcc8d9a94d16846138f64e4e3e59)
Ethics statement
All animal care and euthanasia procedures were performed in accordance with the rules and regulations established by the United States Department of Agriculture Animal Welfare Act and Regulations, Public Health Services Policy on Humane Care and Use of Laboratory Animals, and American Association for Accreditation of Laboratory Animal Care. Humane endpoints were established by the same standards.
Animal studies
Male DO mice (n = 636) were acquired from Jackson Laboratories after weaning at 4–6 weeks of age in 7 cohorts (average n = 106) spanning DO generations 22–25. Mice were singly housed in temperature- and humidity-controlled vivariums on cedar shaving bedding with ad libitum access to water and standard chow (#7912, Harlan Teklad, Madison, WI, United States). Sample size considerations and exclusion of female mice were chosen to maximize power to detect significant bQTL and are described in in Supplemental Materials. Study of ethanol consumption in ~600 animals was predicted to have 80% power to detect alleles causing less than 5% variance in a trait at a p value of <0.05 [29].
Intermittent ethanol access
Mice (n = 587) were studied for ethanol consumption via a three-bottle choice (15% ethanol v/v, 30% ethanol v/v, and water) IEA procedure for 5 weeks with alternating 24-hour periods of ethanol access (Monday, Wednesday, Friday) (Fig. 1A). 49 additional animals were exposed to only water as controls for anxiety-related behavioral studies and RNAseq analyses. Only the first 4 uninterrupted weeks of IEA were used for the purposes of behavioral genetic studies in this report, excluding day 1 of IEA due to increased variance on initial exposure to ethanol bottles. As described in Supplemental Materials, a marble-burying assay was done after week 4 of IEA as a study on abstinence-induced anxiety. This data and that from a basal light-dark box assay done before IEA exposure are not discussed in this report. However, an additional week of IEA was conducted after the marble burying assay to re-establish consumption levels prior to tissue harvesting.
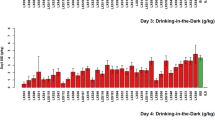
Mice were given voluntary access to ethanol via a three-bottle choice (H2O, 15% EtOH, 30% EtOH) procedure for 24 hours, starting at the beginning of each dark cycle (A). Hierarchical clustering of daily ethanol consumption identified two superclusters, one including drinking days 2–4 and one including drinking days 5–11 (B). Week four mean ethanol consumption was significantly higher than week one mean consumption in a within-sample one-tailed t-test (C). Drinking day was a significant predictor of ethanol consumption in linear regression, with consumption increasing by approximately 0.2 g EtOH/kg body weight each successive day (E). Similarly, week four ethanol preference was higher than week one ethanol preference (D) and drinking day was a significant predictor of ethanol preference, explaining a small proportion of overall variance (F).
Tissue sample collection and genotyping
Mice were euthanized 24 hours after the end of their last ethanol exposure period via cervical dislocation and decapitation and tissue samples were collected immediately afterwards, flash-frozen in liquid nitrogen and stored at −80 °C. Brains were microdissected into nine regions as previously described [30, 31]. Tail snips were collected for genotyping (NeoGen Inc; Lincoln, NE) using a GigaMUGA microarray (nSNP = 141,090; nCNV = 2169) designed to optimize genotyping of DO mice [32] Of the initial 636 DO mice, full datasets were only collected from 630 mice as 6 mice reached humane endpoints before the end of experimentation. Data cleaning removed an additional 29 mice (22 for poor genotyping quality and 7 for potential sample mix-ups), resulting in a final sample of 603 mice (554 ethanol-drinking mice and 49 ethanol-naïve controls) used for analyses presented here.
Behavioral QTL analysis
QTL mapping was carried out using the R/qtl2 software package. Dependent variables included whole study (drinking days 2–11), week one (days 2–4), and week four (days 9–11) averages for ethanol consumption (g ethanol/kg/24 h), ethanol preference (total ethanol consumed in ml/total fluid consumed in ml), and 30% choice (ml 30% ethanol/ml total ethanol) [33]. A week 4–week 1 difference in total ethanol consumption was used as an additional phenotype. Cohort was included as a fixed-effect covariate in all our analyses and kinship between mice was calculated using a linear mixed model and the leave-one-chromosome-out method. Phenotypes were either log- (for consumption and 30% choice) or square-root-transformed (for ethanol preference) to obtain normality before running analyses. Logarithm of the Odds (LOD) scores were calculated for each marker and permutation analysis (nperm = 1000) used to calculate genome-wide empirical p values. SNP variant LOD scores were similarly calculated for QTL intervals and significance level p values determined by permutation across the involved chromosome. To detect founder strain effects, haplotype analysis was conducted for chromosomes containing significant or suggestive bQTLs using best linear unbiased predictors (BLUPs) in R/qtl2.
Expression QTL analysis
To further implicate positional candidate genes from bQTL intervals, we utilized an existing expression QTL (eQTL) analysis of medial prefrontal cortex (mPFC) samples from a subset of 220 DO mice chosen for RNA-seq analysis based on average total ethanol consumption during the fourth week of IEA [34]. mPFC was chosen as a high-priority brain region due to its known role in decision-making, ethanol consumption and reinstatement and functional changes resulting from ethanol exposure as described in the Introduction. This included 100 mice from each extreme of the ethanol consumption distribution and 20 ethanol-naïve control mice. Detailed methodology for RNA-seq data, including alignment to individual genomes using Genotype-by-RNA-Seq (GBRS) [35], is described in Supplemental Methods and is reported elsewhere [34]. GBRS count data was used to generate eQTLs and haplotypes thereof within R/qtl2. Empirical significance of cis-eQTLs were determined by permutation analysis as described in Supplemental Methods and are defined as suggestive (p < 0.63, LOD ≥ 6.13) or significant (p < 0.05, LOD ≥ 8.45).
Results
DO mice exhibit variable patterns of initial versus progressive increase in ethanol consumption over time
Following exclusion of select mice as detailed in Methods, the remaining population of 554 ethanol-exposed DO mice exhibited wide variation in ethanol consumption, preference, and 30% choice over 4 weeks of IEA (Figs. 1 and S1) with ethanol consumption varying from <1 g/kg/24 h to >35 g/kg/24 h in week four of consumption. Consumption increased over time (Fig. 1E) and ethanol intake in week four was significantly higher than in week one (Fig. 1C, p = 1.006 × 10−05). Week one and week four consumption were significantly correlated (0.58, p = 1.08×10-52). Preference was significantly positively correlated with ethanol consumption across all time periods (p < 0.05) and increased significantly from week one to week four (Fig. 1D). 30% choice was not significantly correlated with ethanol consumption across all time periods, suggesting that mice preferring 30% ethanol did not necessarily increase ethanol intake overall (Fig. S2).
Hierarchical clustering of the ethanol consumption time course identified two major clusters, one containing drinking days 2–4 and one containing drinking days 5–11 (Fig. 1B). Principal component analysis of total daily ethanol consumption showed one principal component explained a large percentage (42.4%) of the variance in daily ethanol consumption (Fig. S3A) and was evenly positively loaded across all drinking days (Fig. S1B) and correlated with whole study mean total ethanol consumption on biplot analysis (Fig. S3E). In contrast, a second principal component loaded positively with week 1 drinking and negatively with drinking days 8–11 (Fig. S3B). This component also showed similar results in biplot analysis, being positively loaded with week one mean ethanol consumption (Fig. S3C) and negatively with week four mean ethanol consumption (Fig. S3D). While the first principal component may influence a major portion of overall ethanol consumption, the second component appears to explain variance between week one and week four consumption. This suggests possible genetic factors controlling initial consumption versus escalation in consumption over time, consistent with the hierarchical clustering data. We therefore chose to separately analyze week one and week four of ethanol access in addition to whole study ethanol drinking phenotypes for genetic mapping. The difference between week 4 and week 1 consumption was also studied to assess possible genetic influences on escalation.
SNP-based heritability estimates for ethanol consumption ranged from 0.197 to 0.310, and values for ethanol preference ranged from 0.143 to 0.301. Generally, week one heritability estimates were lower than for either last week or whole study. Estimates for 30% choice were much lower, ranging from 0.00 to 0.073 (Table S1).
bQTL analysis identifies 3 significant loci and multiple suggestive loci for ethanol intake
Significant bQTLs (p < 0.05) were identified on Chromosome (Chr) 4 for week four mean ethanol consumption (W4_EC; LOD = 8.23), on Chr 3 for week four mean 30% choice (W4_30C; LOD = 8.63), and on Chr 12 for week one mean ethanol preference (W1_EP; LOD = 7.52) (Fig. 2). An additional 13 suggestive QTLs (p < 0.63) were identified, and 95% Bayesian credible intervals were estimated for all QTLs (Table 1). All significant and suggestive bQTLs explained between 4% and 6% of observed variance in their relevant phenotypes, with significant bQTLs identifying greater genetic variance (Table 1). No significant or suggestive QTLs were found for the drinking phenotype principal components described above (Figs. S3F and S3G). The week 4 – week 1 difference analysis identified a suggestive QTL on Chr 15 overlapping with a week 4 ethanol preference suggestive QTL in that location (Fig. S7).
Different genetic effects were identified for week one and week four ethanol consumption (A), preference (B), and 30% choice (C) behavioral QTLs. For ethanol consumption, no significant QTL was observed during week one of the study, but a significant QTL on Chromosome 4 was identified in week four (LOD = 8.23; p = 0.017). For ethanol preference, a significant QTL was observed on Chromosome 12 (LOD = 7.52, p = 0.043) during week one of the study. For 30% choice a significant QTL was identified on Chromosome 3 during week four (LOD = 8.63, p = 0.008). Empirical significance thresholds for each phenotype were calculated using permutation analysis (nperm = 1000); solid black lines represent p < 0.05 and dashed black lines represent p < 0.63 thresholds. D–F Haplotype analysis was carried out using best linear unbiased predictors for each significant bQTL. Support intervals for bQTLs are grayed. A/J alleles (yellow) contributed negatively to week four ethanol consumption (D) and positively to week one ethanol preference (E). PWK alleles (red) contributed negatively and C57BL/6 J (gray) contributed positively to week four 30% choice (F).
Haplotype analysis within R/qtl2 implicated strain-specific allelic contributions to variation in ethanol traits at the significant bQTLs. For the significant Chr 4 W4_EC locus, A/J alleles were associated with lower consumption, whereas NOD and PWK alleles were associated with higher ethanol intake in week 4 (Fig. 2D). The Chr 3 W4_30C locus had A/J, C57BL/6 J, and WSB/EiJ alleles correlated with increased in 30% choice whereas PWK/PhJ alleles associated with a decrease in this phenotype (Fig. 2F). NZO/HlLtJ and A/J alleles at the significant Chr 12 locus were associated with an increase in W1_EP, while CAST/EiJ, NOD/SHiLtJ, and WSB/EiJ alleles showed decreased ethanol preference (Fig. 2I).
Top variants identified within significant bQTL intervals
Top SNPs within significant QTL confidence intervals were identified using a 1.5 LOD-drop from the peak SNP and permutation analysis (nperm = 1000) to calculate empirical p values for variant LOD scores (Figs. 3A, S4 and S5). For the Chr 4 W4_EC QTL, the top three variants were intergenic SNPs located upstream of the Car8 gene and downstream of predicted gene Gm37386, with the highest LOD score at rs249655952 (LOD = 3.84) (Fig. 3A, Table S2). Among 7 variants within a 1.5 LOD drop, 5 were unique to the A/J progenitor strain and the remaining 2 were intergenic deletions unique to NOD/ShiLtJ, consistent with the pattern of these two strains on haplotype analysis (Fig. 2D).
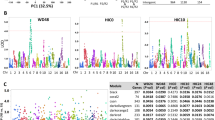
A SNP associations were estimated across the 95% Bayesian confidence interval identified for the significant bQTL, identified by dashed vertical lines (upper panel). Statistical thresholds identified by permutation analysis are indicated. Top variants were selected as those with a LOD score within 1.5 of the highest score as is conventional (Supplemental Table 2). Variants having suggestive p value significance for association with last week ethanol consumption are indicated (green). Known gene transcript annotations within this C.I. (middle panel) include Car8, Rab2a, Chd7, and Clvs1. As a highly ranked candidate gene (Table 2), Car8 is colored (green). Lower panel indicates ReMap regulatory element density across the interval. B eQTL analysis for Car8 indicates a highly significant cis-eQTL. C Haplotype analysis of Car8 eQTL indicates A/J alleles (yellow) contribute positively while NOD alleles contribute negatively to expression of this gene.
Similarly, variant LOD scores, transcripts, and regulatory elements were identified for the Chr 12 C.I. for W1_EP (Fig. S4) and the Chr 3 C.I. for W4_30C (Fig. S5). Within the Chr 12 W1_EP interval, 23 variants were identified within 1.5 LOD units of the top-scoring variant (rs46944281, LOD = 6.00), with 22 of these being intronic or intergenic and 1 (rs51613251, LOD = 4.52) a missense variant in Zfyve26. This variant alters the reference amino acid sequence from alanine in progenitor strains C57BL/6 J, 129, A/J, PWK/PhJ, WSB/EiJ and NZO/HlLtJ, to glutamine in CAST/EiJ and NOD/ShiLtJ mice (https://www.informatics.jax.org/mgv/). This is partially consistent with findings of the W1_EP haplotype analysis (Fig. 2E) discussed above. The Ala/Gln amino acid change in Zfyve26 is predicted to be structurally tolerated (https://useast.ensembl.org/info/docs/tools/vep/index.html).
Within the Chr 3 C.I. for W4_30C, 250 variants were identified within 1.5 LOD units of the top scoring SNP (rs49087152, LOD = 6.13). Again, most variants occurred in extragenic or intronic sequences; however, there was a missense variant (rs29605696, LOD = 5.10) for Celsr2, and 5’-UTR variants for both Prpf38b (rs29689050, LOD = 5.59) and Sypl2 (rs253441023, LOD = 4.69) (https://www.informatics.jax.org/mgv/). The missense variant in Celsr2 alters a leucine in the reference sequence (i.e., C57BL/6 J, A/J, 129S1/SvImJ, NOD/ShiLtJ, WSB/EiJ) to proline in the NZO/HILtJ, CAST/EiJ, and PWK/PhJ progenitor strains [36]. Haplotype analysis of W4_30C noted above was consistent with this variant strain distribution pattern, demonstrating a decrease in W4_30C for strains containing the proline substitution (Fig. 2I). The Leu/Pro amino acid change in Celsr2 is predicted to be structurally “tolerated” (https://useast.ensembl.org/info/docs/tools/vep/index.html). The 5’-UTR variant in Sypl2 is present in NZO/HILtJ, CAST/EiJ, PWK/PhJ, and WSB/EiJ founder strains and falls within a predicted promotor element (ENCODE EM10E0741064). The 5’-UTR variant in Prpf38b occurs only in PWK and NZO strains and falls within a predicted binding site for the Znf549 transcription factor. The strain distribution pattern of the 5’-UTR variants in both Sypl2 and Prpf38b do not fully match the haplotype pattern for the W4_30C phenotype (Fig. 2F), suggesting the rs29605696 missense variant in Celsr2 as a more likely causal variant for W4_30C.
eQTL analysis
To identify genetic variance in gene expression within significant bQTL S.I., cis-eQTLs were mapped using RNA-seq data from mPFC. These eQTL and transcriptomic studies are published in detail elsewhere [34] and are described in Supplemental Methods. This analysis identified suggestive (p < 0.63, LOD > 6.13) and significant cis-eQTLs (p < 0.05, LOD > 8.45). When filtered to identify cis-eQTLs overlapping the significant W4_EC, W1_EP and W4_30C bQTL confidence intervals, this produced 53 cis-eQTL for 48 genes in these 3 bQTL intervals based on suggestive or significant cis-eQTL, with 45/48 genes having significant (p < 0.05) cis-eQTLs (Table 2).
Only one gene within the significant bQTL confidence interval for W4_EC on Chr 4 had a cis-eQTL which met filtering criteria: Carbonic anhydrase 8 (Car8) (Fig. 3B). Haplotype analysis of the Car8 cis-eQTL indicated that A/J alleles at the locus correlated with increased expression of Car8 in prefrontal cortex (Fig. 3C). Car8 expression had a significant negative Spearman correlation with both W4_EC (r = −0.22; p = 0.008) and week four ethanol preference (r = −0.23; p = 0.006), consistent with the relationship suggested by the effect of A/J alleles. Taken together, these results suggest a role for variants unique to A/J mice in both decreased Car8 expression and increased last week ethanol consumption in DO mice.
The significant Chr 3 bQTL for W4_30C contained 40 genes having cis-eQTLs with LOD scores > 6.13 (Table 2). Of these, Sypl2 and Prpf38b also contained 5’-UTR sequence variation as noted above. No cis-eQTL was observed for Celsr2 at this locus. Eight genes within the Chr 12 W1_EP bQTL C.I had significant cis-eQTLs with LOD scores > 8.45 and four of these having multiple significant eQTLs. Zfyve26 had two significant eQTLs and a coding sequence variant in the top variants as noted above.
Bioinformatics analysis identifies genes with alcohol trait human GWAS associations
Bioinformatics analysis identified 7 genes from significant or suggestive bQTLs support intervals as having significant or suggestive associations (p values < 6.0E-6) with alcohol-related traits in the GWAS Catalog database (https://www.ebi.ac.uk/gwas/home) and multiple genes with supporting literature associations with ethanol or substance misuse (Table 2). Of the significant bQTLs, 4 human GWAS hits occurred within the LW30C Chr 3 S.I. Gstm5 is related to glutathione metabolism, Eps8l3 and Celsr2 function in epidermal growth factor pathways, and Ubl4b codes for a ubiquitin-like protein functioning in protein targeting.
Discussion
This report is the first to analyze genetic variance in ethanol consumption using DO mice. The design of this work allowed study of a progressive ethanol consumption phenotype at high genetic resolution in a model assessing a large proportion of genetic variance across mouse populations. Our results show remarkable diversity in ethanol consumption across the study population and a suggestion of differing genetic influences on initial ethanol intake (week one) versus chronic consumption (week four). We identified 3 significant and 13 suggestive bQTLs across 7 chromosomes. In most cases, the support intervals for these bQTL in DO mice were much narrower (Table 2) than seen in prior mouse genetic studies on ethanol behaviors. The integration of cis-eQTL and bioinformatic analyses allowed prioritization of an experimentally tractable number of novel high priority candidate genes, with some candidates overlapping results from prior rodent or human GWAS studies on alcohol.
bQTL analysis identifies specific, novel loci for ethanol consumption behaviors
Our high-resolution genetic mapping for ethanol consumption phenotypes identified significant or suggestive bQTLs across assays in week one or week four of ethanol access (Table 1), but importantly, we did not observe any bQTLs in common between those time periods, suggesting differing genetic influences on initial ethanol consumption and more chronic, progressive consumption. This is also consistent with our multivariate analysis of the longitudinal consumption data, showing a clear separation in initial vs. late ethanol access. Only one suggestive bQTL was identified for week 4–week 1 consumption, on Chr 15 in a region overlapping with a suggestive QTL for W4_EP, despite the strong evidence for significant escalation of ethanol consumption in the overall study. This likely resulted from decreased statistical power due to the mathematical manipulation in the difference score.
Several of the mapped bQTLs overlapped regions implicated in prior rodent genetic model analysis of ethanol or substance use behaviors (Table 1). For example, all 3 bQTLs on Chr 3 were located within the confidence interval for alcohol preference identified by Belknap and Atkins in 2001 [11, 37]. However, the QTL confidence intervals identified here were generally an order of magnitude smaller than prior reports, with the largest being the significant interval for W4_30C at approximately 3.77 Mbp. These comparisons with prior studies demonstrate both the power of our DO mouse analysis for high resolution identification of novel loci modulating ethanol consumption, and the likely validation and improved mapping of prior rodent QTL analyses, consistent with other behavioral genetic studies using DO mice [28, 38].
eQTL and bioinformatics analysis identifies Car8 as a top candidate gene for Chr4 LWEC QTL
We identified cis-eQTL within bQTL support intervals to annotate candidate gene lists since such colocalization provides strong evidence for genetic modulation of gene expression influencing ethanol-related behaviors. A prime example of this approach was Car8, located within the significant Chr 4 bQTL for W4_EC (Fig. 3). Car8 was the only gene in the interval having a significant cis-eQTL in our mPFC data. Furthermore, haplotype analysis across the W4_EC bQTL and Car8 eQTL revealed that A/J alleles at this locus correlated with both decreased week 4 ethanol consumption and increased Car8 expression in prefrontal cortex. Car8 expression in mPFC notably had a significant negative correlation with ethanol consumption in our male Diversity Outbred mice. We consider these data strong support for Car8 as a high priority candidate gene in the Chr4 W4_EC bQTL.
Car8 shares sequence similarity to the carbonic anhydrase family of genes, but lacks carbonic anhydrase activity; instead, it is known to inhibit the IP3R1 calcium signaling channel [39]. IP3R1 has been suggested to play a role in ethanol-enhanced GABA release in cerebellum, a potential mechanism by which ethanol-induced cerebellar ataxia occurs [40]. Car8 is expressed across multiple brain regions and implicated in locomotor function and analgesia [41,42,43]. Car8 has been shown to be regulated by ethanol within cerebellum, ventral midbrain and anterior cingulate in mice (Table 2). Additionally, recent reports have suggested that IP3R1 signaling in prefrontal cortex astrocytes can regulate ethanol consumption and that Car8 is regulated in astrocytes within that brain region by chronic ethanol consumption [20, 44]. Given this ethanol regulation in brain, the significant negative correlations between Car8 expression and W4_EC (Fig. S6), and the strong Car8 eQTL in linkage disequilibrium with the Chr 4 bQTL for W4_EC, we hypothesize that altered expression of Car8 may modulate chronic ethanol consumption.
Other bQTL candidates with structural variants or links to human GWAS
Several candidate genes within the significant bQTL for W4_30C on Chr 3 (Celsr2, Prpf38b and Sypl2) and W1_EP on Chr 12 (Zfyve26) contained exonic sequence variants (Table 2) that might modulate gene expression or function. Of note, Prpf38b is poorly characterized functionally but is regulated by ethanol in prefrontal cortex astrocytes, as noted above for Car8 [45]. Celsr2 encodes a transmembrane protein and has been implicated in axon development in the forebrain [46] and regulation of motor neuron regeneration following injury [47]. Notably, Celsr2 had a missense variant in the DO population and GWAS data show that Celsr2 is strongly associated with liver fibrosis in individuals with high alcohol intake [48] and with interaction terms between alcohol consumption and both LDL and HDL cholesterol levels [49]. Additionally, Celsr2 has previously been shown to be regulated by ethanol in chronic ethanol consuming rhesus macaques [50] and in multiple rodent studies [30, 45, 51]. These genes with exonic variants clearly warrant further study to characterize any direct involvement in ethanol-related behaviors. Overall, the consilience between studies here, prior model organism behavioral genetics and human GWAS studies serve to reinforce our findings and justify pursuit of further validation or mechanistic studies in mice.
Limitations
Despite the novel findings of this analysis, there are several limitations which should be considered. Firstly, the Diversity Outbred mouse population used for this study only included male animals due to power considerations; while future studies are planned to validate identified loci and candidate genes in female mice, these results may not be generalizable to both sexes. Additionally, RNA-seq data used for eQTL analysis came from bulk sequencing of only prefrontal cortex tissue, which may not reflect transcriptomic differences in individual cell types or other brain regions. Larger sample sizes, incorporation of expression data from additional brain regions, and a single cell RNA-seq approach would provide more detailed insight into the genetic and transcriptomics mechanisms underlying observed differences in ethanol consumption. Finally, the addition of human GWAS data as an additional criterion for identifying top candidate genes is limited by the methods in which GWAS associations are linked to individual genes and the size of existing studies.
Conclusions
These findings identified novel genes and potential mechanisms modulating ethanol consumption phenotypes in DO mice, thus adding to our understanding of the complex genetic and molecular architecture of ethanol consumption and AUD in humans. Further transcriptome, bioinformatic and behavioral genetic analyses of this powerful dataset promise to identify additional gene networks or individual targets that may aid in development of future therapeutic approaches for AUD [34].
Data availability
All data supporting the conclusions of this manuscript are contained in figures and tables or supplemental materials. Cleaned behavioral and genotype data and expression QTL data are provided on the Open Science Framework (https://osf.io/tfrzg/?view_only=232cfcc8d9a94d16846138f64e4e3e59). Additional raw data will be provided upon request.
References
World Health Organization. 2018. Global Status Report on Alcohol and Health 2018; Geneva: Switzerland.
Esser M, Sherk A, Liu Y. Deaths and years of potential life lost from excessive alcohol use - United States, 2011-2015. MMWR Morb Mortal Wkly Rep. 2020;69:1428–33.
Verhulst B, Neale MC, Kendler KS. The heritability of alcohol use disorders: a meta-analysis of twin and adoption studies. Psychol Med. 2015;45:1061–72.
Backman JD, Li AH, Marcketta A, Sun D, Mbatchou J, Kessler MD, et al. Exome sequencing and analysis of 454,787 UK Biobank participants. Nature. 2021;599:628–34.
Zhou H, Sealock JM, Sanchez-Roige S, Clarke TK, Levey DF, Cheng Z, et al. Genome-wide meta-analysis of problematic alcohol use in 435,563 individuals yields insights into biology and relationships with other traits. Nat Neurosci. 2020;23:809–18.
Kranzler HR, Zhou H, Kember RL, Vickers Smith R, Justice AC, Damrauer S, et al. Genome-wide association study of alcohol consumption and use disorder in 274,424 individuals from multiple populations. Nat Commun. 2019;10:1499.
Sanchez-Roige S, Palmer AA, Fontanillas P, Elson SL, Me Research Team tSUDWGotPGC, Adams MJ, et al. Genome-wide association study meta-analysis of the alcohol use disorders identification test (AUDIT) in two population-based cohorts. Am J Psychiatry. 2019;176:107–18.
Walters RK, Polimanti R, Johnson EC, McClintick JN, Adams MJ, Adkins AE, et al. Transancestral GWAS of alcohol dependence reveals common genetic underpinnings with psychiatric disorders. Nat Neurosci. 2018;21:1656–69.
Zhou H, Kember RL, Deak JD, Xu H, Toikumo S, Yuan K, et al. Multi-ancestry study of the genetics of problematic alcohol use in over 1 million individuals. Nat Med. 2023;29:3184–92.
Ehlers CL, Walter NA, Dick DM, Buck KJ, Crabbe JC. A comparison of selected quantitative trait loci associated with alcohol use phenotypes in humans and mouse models. Addict Biol. 2010;15:185–99.
Belknap JK, Atkins AL. The replicability of QTLs for murine alcohol preference drinking behavior across eight independent studies. Mamm Genome. 2001;12:893–9.
Bice PJ, Foroud T, Carr LG, Zhang L, Liu L, Grahame NJ, et al. Identification of QTLs influencing alcohol preference in the High Alcohol Preferring (HAP) and Low Alcohol Preferring (LAP) mouse lines. Behav Genet. 2006;36:248–60.
Buck KJ, Finn DA. Genetic factors in addiction: QTL mapping and candidate gene studies implicate GABAergic genes in alcohol and barbiturate withdrawal in mice. Addiction. 2001;96:139–49.
Hwa LS, Chu A, Levinson SA, Kayyali TM, DeBold JF, Miczek KA. Persistent escalation of alcohol drinking in C57BL/6J mice with intermittent access to 20% ethanol. Alcohol Clin Exp Res. 2011;35:1938–47.
Rosenwasser AM, Fixaris MC, Crabbe JC, Brooks PC, Ascheid S. Escalation of intake under intermittent ethanol access in diverse mouse genotypes. Addict Biol. 2013;18:496–507.
Bagley JR, Chesler EJ, Philip VM, Center for the Systems Genetics of A, Jentsch JD. Heritability of ethanol consumption and pharmacokinetics in a genetically diverse panel of collaborative cross mouse strains and their inbred founders. Alcohol Clin Exp Res. 2021;45:697–708.
MacLeod LC, Miles MF. Analysis of intermittent versus daily ethanol consumption in genetically diverse mouse models. Alcohol-Clin Exp Res. 2016;40:75a–75a.
Avchalumov Y, Oliver RJ, Trenet W, Heyer Osorno RE, Sibley BD, Purohit DC, et al. Chronic ethanol exposure differentially alters neuronal function in the medial prefrontal cortex and dentate gyrus. Neuropharmacology. 2021;185:108438.
Keistler CR, Hammarlund E, Barker JM, Bond CW, DiLeone RJ, Pittenger C, et al. Regulation of alcohol extinction and cue-induced reinstatement by specific projections among medial prefrontal cortex, nucleus accumbens, and basolateral amygdala. J Neurosci. 2017;37:4462–71.
Salem NA, Manzano L, Keist MW, Ponomareva O, Roberts AJ, Roberto M, et al. Cell-type brain-region specific changes in prefrontal cortex of a mouse model of alcohol dependence. Neurobiol Dis. 2024;190:106361.
Mulligan MK, Zhao W, Dickerson M, Arends D, Prins P, Cavigelli SA, et al. Genetic contribution to initial and progressive alcohol intake among recombinant inbred strains of mice. Front Genet. 2018;9:370.
Bennett B, Downing C, Carosone-Link P, Ponicsan H, Ruf C, Johnson TE. Quantitative trait locus mapping for acute functional tolerance to ethanol in the L x S recombinant inbred panel. Alcohol Clin Exp Res. 2007;31:200–8.
Solberg Woods LC. QTL mapping in outbred populations: successes and challenges. Physiol Genomics. 2014;46:81–90.
Attie AD, Churchill GA, Nadeau JH. How mice are indispensable for understanding obesity and diabetes genetics. Curr Opin Endocrinol Diabetes Obes. 2017;24:83–91.
Svenson KL, Gatti DM, Valdar W, Welsh CE, Cheng R, Chesler EJ, et al. High-resolution genetic mapping using the Mouse Diversity outbred population. Genetics. 2012;190:437–47.
Threadgill DW, Churchill GA. Ten years of the collaborative cross. G3 (Bethesda). 2012;2:153–6.
Churchill GA, Gatti DM, Munger SC, Svenson KL. The Diversity Outbred mouse population. Mamm Genome. 2012;23:713–8.
Parker CC, Philip VM, Gatti DM, Kasparek S, Kreuzman AM, Kuffler L, et al. Genome-wide association mapping of ethanol sensitivity in the Diversity Outbred mouse population. Alcohol Clin Exp Res. 2022;46:941–60.
Gatti DM, Svenson KL, Shabalin A, Wu LY, Valdar W, Simecek P, et al. Quantitative trait locus mapping methods for diversity outbred mice. G3 (Bethesda). 2014;4:1623–33.
Wolen AR, Phillips CA, Langston MA, Putman AH, Vorster PJ, Bruce NA, et al. Genetic dissection of acute ethanol responsive gene networks in prefrontal cortex: functional and mechanistic implications. PLoS One. 2012;7:e33575.
Kerns RT, Ravindranathan A, Hassan S, Cage MP, York T, Sikela JM, et al. Ethanol-responsive brain region expression networks: implications for behavioral responses to acute ethanol in DBA/2J versus C57BL/6J mice. J Neurosci. 2005;25:2255–66.
Morgan AP, Fu CP, Kao CY, Welsh CE, Didion JP, Yadgary L, et al. The mouse universal genotyping array: from substrains to subspecies. G3 (Bethesda). 2015;6:263–79.
Broman KW, Gatti DM, Simecek P, Furlotte NA, Prins P, Sen S, et al. R/qtl2: software for mapping quantitative trait loci with high-dimensional data and multiparent populations. Genetics. 2019;211:495–502.
Smith ML, Sergi Z, Mignogna KM, Rodriguez NE, Tatom Z, MacLeod L, et al. Identification of Genetic and Genomic Influences on Progressive Ethanol Consumption in Diversity Outbred Mice. 2023;bioRxiv https://doi.org/10.1101/2023.09.15.554349.
Choi K, He H, Gatti DM, Philip VM, Raghupathy N, Gyuricza IG, et al. Genotype-free individual genome reconstruction of multiparental population models by RNA sequencing data. 2020;bioRxiv https://doi.org/10.1101/2020.10.11.335323.
Bult CJ, Blake JA, Smith CL, Kadin JA, Richardson JE, Mouse Genome Database G. Mouse Genome Database (MGD) 2019. Nucleic Acids Res. 2019;47:D801–D06.
Saba L, Bhave SV, Grahame N, Bice P, Lapadat R, Belknap J, et al. Candidate genes and their regulatory elements: alcohol preference and tolerance. Mamm Genome. 2006;17:669–88.
Recla JM, Bubier JA, Gatti DM, Ryan JL, Long KH, Robledo RF, et al. Genetic mapping in Diversity Outbred mice identifies a Trpa1 variant influencing late-phase formalin response. Pain. 2019;160:1740–53.
Hirota J, Ando H, Hamada K, Mikoshiba K. Carbonic anhydrase-related protein is a novel binding protein for inositol 1,4,5-trisphosphate receptor type 1. Biochem J. 2003;372:435–41.
Kelm MK, Weinberg RJ, Criswell HE, Breese GR. The PLC/IP 3 R/PKC pathway is required for ethanol-enhanced GABA release. Neuropharmacology 2010;58:1179–86.
Miterko LN, White JJ, Lin T, Brown AM, O’Donovan KJ, Sillitoe RV. Persistent motor dysfunction despite homeostatic rescue of cerebellar morphogenesis in the Car8 waddles mutant mouse. Neural Dev. 2019;14:6.
Shimobayashi E, Wagner W, Kapfhammer JP. Carbonic anhydrase 8 expression in purkinje cells is controlled by PKCgamma activity and regulates purkinje cell dendritic growth. Mol Neurobiol. 2016;53:5149–60.
Fu ES, Erasso DM, Zhuang GZ, Upadhyay U, Ozdemir M, Wiltshire T, et al. Impact of human CA8 on thermal antinociception in relation to morphine equivalence in mice. Neuroreport. 2017;28:1215–20.
Erickson EK, DaCosta AJ, Mason SC, Blednov YA, Mayfield RD, Harris RA. Cortical astrocytes regulate ethanol consumption and intoxication in mice. Neuropsychopharmacology. 2021;46:500–08.
Erickson EK, Farris SP, Blednov YA, Mayfield RD, Harris RA. Astrocyte-specific transcriptome responses to chronic ethanol consumption. Pharmacogenomics J. 2018;18:578–89.
Qu Y, Huang Y, Feng J, Alvarez-Bolado G, Grove EA, Yang Y, et al. Genetic evidence that Celsr3 and Celsr2, together with Fzd3, regulate forebrain wiring in a Vangl-independent manner. Proc Natl Acad Sci USA. 2014;111:E2996–3004.
Wen Q, Weng H, Liu T, Yu L, Zhao T, Qin J, et al. Inactivating Celsr2 promotes motor axon fasciculation and regeneration in mouse and human. Brain. 2022;145:670–83.
Innes H, Buch S, Hutchinson S, Guha IN, Morling JR, Barnes E, et al. Genome-wide association study for alcohol-related cirrhosis identifies risk loci in MARC1 and HNRNPUL1. Gastroenterology. 2020;159:1276–89 e7.
de Vries PS, Brown MR, Bentley AR, Sung YJ, Winkler TW, Ntalla I, et al. Multiancestry genome-wide association study of lipid levels incorporating gene-alcohol interactions. Am J Epidemiol. 2019;188:1033–54.
Bogenpohl JW, Smith ML, Farris SP, Dumur CI, Lopez MF, Becker HC, et al. Cross-species co-analysis of prefrontal cortex chronic ethanol transcriptome responses in mice and monkeys. Front Mol Neurosci. 2019;12:197.
McCarthy GM, Farris SP, Blednov YA, Harris RA, Mayfield RD. Microglial-specific transcriptome changes following chronic alcohol consumption. Neuropharmacology. 2018;128:416–24.
Radcliffe RA, Bludeau P, Asperi W, Fay T, Deng XS, Erwin VG, et al. Confirmation of quantitative trait loci for ethanol sensitivity and neurotensin receptor density in crosses derived from the inbred high and low alcohol sensitive selectively bred rat lines. Psychopharmacol (Berl). 2006;188:343–54.
Terenina-Rigaldie E, Moisan MP, Colas A, Beauge F, Shah KV, Jones BC, et al. Genetics of behaviour: phenotypic and molecular study of rats derived from high- and low-alcohol consuming lines. Pharmacogenetics. 2003;13:543–54.
Gill KJ, Boyle AE. Genetic basis for the psychostimulant effects of nicotine: a quantitative trait locus analysis in AcB/BcA recombinant congenic mice. Genes Brain Behav. 2005;4:401–11.
Risinger FO, Cunningham CL. Ethanol-induced conditioned taste aversion in BXD recombinant inbred mice. Alcohol Clin Exp Res. 1998;22:1234–44.
McClearn GE, Tarantino LM, Hofer SM, Jones B, Plomin R. Developmental loss of effect of a Chromosome 15 QTL on alcohol acceptance. Mamm Genome. 1998;9:991–4.
Bell RL, Kimpel MW, McClintick JN, Strother WN, Carr LG, Liang T, et al. Gene expression changes in the nucleus accumbens of alcohol-preferring rats following chronic ethanol consumption. Pharm Biochem Behav. 2009;94:131–47.
Osterndorff-Kahanek E, Ponomarev I, Blednov YA, Harris RA. Gene expression in brain and liver produced by three different regimens of alcohol consumption in mice: comparison with immune activation. PLoS One. 2013;8:e59870.
Philip VM, Duvvuru S, Gomero B, Ansah TA, Blaha CD, Cook MN, et al. High-throughput behavioral phenotyping in the expanded panel of BXD recombinant inbred strains. Genes Brain Behav. 2010;9:129–59.
Baumert J, Huang J, McKnight B, Sabater-Lleal M, Steri M, Chu AY, et al. No evidence for genome-wide interactions on plasma fibrinogen by smoking, alcohol consumption and body mass index: results from meta-analyses of 80,607 subjects. PLoS One. 2014;9:e111156.
Li W, Thygesen JH, O’Brien NL, Heydtmann M, Smith I, Degenhardt F, et al. The influence of regression models on genome-wide association studies of alcohol dependence: a comparison of binary and quantitative analyses. Psychiatr Genet. 2021;31:13–20.
Iwamoto K, Bundo M, Yamamoto M, Ozawa H, Saito T, Kato T. Decreased expression of NEFH and PCP4/PEP19 in the prefrontal cortex of alcoholics. Neurosci Res. 2004;49:379–85.
Lewohl JM, Wang L, Miles MF, Zhang L, Dodd PR, Harris RA. Gene expression in human alcoholism: microarray analysis of frontal cortex. Alcohol Clin Exp Res. 2000;24:1873–82.
Ponomarev I, Wang S, Zhang L, Harris RA, Mayfield RD. Gene coexpression networks in human brain identify epigenetic modifications in alcohol dependence. J Neurosci. 2012;32:1884–97.
Pochareddy S, Edenberg HJ. Chronic alcohol exposure alters gene expression in HepG2 cells. Alcohol Clin Exp Res. 2012;36:1021–33.
Farris SP, Harris RA, Ponomarev I. Epigenetic modulation of brain gene networks for cocaine and alcohol abuse. Front Neurosci. 2015;9:176.
Yin HQ, Kim M, Kim JH, Kong G, Kang KS, Kim HL, et al. Differential gene expression and lipid metabolism in fatty liver induced by acute ethanol treatment in mice. Toxicol Appl Pharm. 2007;223:225–33.
Konu O, Kane JK, Barrett T, Vawter MP, Chang R, Ma JZ, et al. Region-specific transcriptional response to chronic nicotine in rat brain. Brain Res. 2001;909:194–203.
Kozell LB, Lockwood D, Darakjian P, Edmunds S, Shepherdson K, Buck KJ, et al. RNA-Seq Analysis of Genetic and Transcriptome Network Effects of Dual-Trait Selection for Ethanol Preference and Withdrawal Using SOT and NOT Genetic Models. Alcohol Clin Exp Res. 2020;44:820–30.
Mulligan MK, Ponomarev I, Hitzemann RJ, Belknap JK, Tabakoff B, Harris RA, et al. Toward understanding the genetics of alcohol drinking through transcriptome meta-analysis. Proc Natl Acad Sci USA. 2006;103:6368–73.
Bae JS, Jung MH, Lee BC, Cheong HS, Park BL, Kim LH, et al. The genetic effect of copy number variations on the risk of alcoholism in a Korean population. Alcohol Clin Exp Res. 2012;36:35–42.
Mulligan MK, Rhodes JS, Crabbe JC, Mayfield RD, Harris RA, Ponomarev I. Molecular profiles of drinking alcohol to intoxication in C57BL/6J mice. Alcohol Clin Exp Res. 2011;35:659–70.
Choi MR, Han JS, Chai YG, Jin YB, Lee SR, Kim DJ. Gene expression profiling in the hippocampus of adolescent rats after chronic alcohol administration. Basic Clin Pharm Toxicol. 2020;126:389–98.
Renthal W, Kumar A, Xiao G, Wilkinson M, Covington HE 3rd, Maze I, et al. Genome-wide analysis of chromatin regulation by cocaine reveals a role for sirtuins. Neuron 2009;62:335–48.
McClintick JN, McBride WJ, Bell RL, Ding ZM, Liu Y, Xuei X, et al. Gene Expression Changes in Glutamate and GABA-A Receptors, Neuropeptides, Ion Channels, and Cholesterol Synthesis in the Periaqueductal Gray Following Binge-Like Alcohol Drinking by Adolescent Alcohol-Preferring (P) Rats. Alcohol Clin Exp Res. 2016;40:955–68.
Stankiewicz AM, Goscik J, Dyr W, Juszczak GR, Ryglewicz D, Swiergiel AH, et al. Novel candidate genes for alcoholism-transcriptomic analysis of prefrontal medial cortex, hippocampus and nucleus accumbens of Warsaw alcohol-preferring and non-preferring rats. Pharm Biochem Behav. 2015;139:27–38.
Kapoor M, Wang JC, Wetherill L, Le N, Bertelsen S, Hinrichs AL, et al. Genome-wide survival analysis of age at onset of alcohol dependence in extended high-risk COGA families. Drug Alcohol Depend. 2014;142:56–62.
Mulligan MK, Mozhui K, Pandey AK, Smith ML, Gong S, Ingels J, et al. Genetic divergence in the transcriptional engram of chronic alcohol abuse: A laser-capture RNA-seq study of the mouse mesocorticolimbic system. Alcohol. 2017;58:61–72.
Repunte-Canonigo V, van der Stap LD, Chen J, Sabino V, Wagner U, Zorrilla EP, et al. Genome-wide gene expression analysis identifies K-ras as a regulator of alcohol intake. Brain Res. 2010;1339:1–10.
Marballi K, Genabai NK, Blednov YA, Harris RA, Ponomarev I. Alcohol consumption induces global gene expression changes in VTA dopaminergic neurons. Genes Brain Behav. 2016;15:318–26.
Melendez RI, McGinty JF, Kalivas PW, Becker HC. Brain region-specific gene expression changes after chronic intermittent ethanol exposure and early withdrawal in C57BL/6J mice. Addict Biol. 2012;17:351–64.
Sokolov BP, Jiang L, Trivedi NS, Aston C. Transcription profiling reveals mitochondrial, ubiquitin and signaling systems abnormalities in postmortem brains from subjects with a history of alcohol abuse or dependence. J Neurosci Res. 2003;72:756–67.
Tabakoff B, Bhave SV, Hoffman PL. Selective breeding, quantitative trait locus analysis, and gene arrays identify candidate genes for complex drug-related behaviors. J Neurosci. 2003;23:4491–8.
Kimpel MW, Strother WN, McClintick JN, Carr LG, Liang T, Edenberg HJ, et al. Functional gene expression differences between inbred alcohol-preferring and -non-preferring rats in five brain regions. Alcohol. 2007;41:95–132.
Grice DE, Reenila I, Mannisto PT, Brooks AI, Smith GG, Golden GT, et al. Transcriptional profiling of C57 and DBA strains of mice in the absence and presence of morphine. BMC Genomics. 2007;8:76.
Albertson DN, Schmidt CJ, Kapatos G, Bannon MJ. Distinctive profiles of gene expression in the human nucleus accumbens associated with cocaine and heroin abuse. Neuropsychopharmacology. 2006;31:2304–12.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet. 2000;25:25–9.
Gene Ontology C. The Gene Ontology resource: enriching a GOld mine. Nucleic Acids Res. 2021;49:D325–D34.
Buck KJ, Metten P, Belknap JK, Crabbe JC. Quantitative trait loci involved in genetic predisposition to acute alcohol withdrawal in mice. J Neurosci. 1997;17:3946–55.
Ghezzi A, Krishnan HR, Lew L, Prado FJ 3rd, Ong DS, Atkinson NS. Alcohol-induced histone acetylation reveals a gene network involved in alcohol tolerance. PLoS Genet. 2013;9:e1003986.
Acknowledgements
The authors thank members of the Miles Laboratory for their support during the performance of these analyses. Nicholas Rodriguez was instrumental in performing GBRS analyses. We also thank Drs. Vivek Philip and K.B. Choi at The Jackson Laboratory for assistance in utilizing GBRS for RNAseq analysis.
Funding
This research was supported by NIAAA grants P50AA022537 to MFM and 1F31AA031189 to ZT and by intramural funding to the VCU Alcohol Research Center provided by the VCU School of Medicine and the VCU Office of the Vice-President for Research and Innovation
Author information
Authors and Affiliations
Contributions
Conceived and designed the study: KM, LM, MFM. Acquired the data: KM, LM, ZT. Analyzed the data: ZT, KM, ZS, AN, MM. Wrote the paper: ZT, KM, ZS, MS, MFM.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mignogna, K.M., Tatom, Z., Macleod, L. et al. Identification of novel genetic loci and candidate genes for progressive ethanol consumption in diversity outbred mice. Neuropsychopharmacol. (2024). https://doi.org/10.1038/s41386-024-01902-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41386-024-01902-6
- Springer Nature Switzerland AG