Abstract
Chromatin regulation through histone modifications plays an essential role in coordinated expression of multiple genes. Alterations in chromatin induced by histone modifiers and readers regulate critical transcriptional programs involved in both normal development and tumor differentiation. Recently, we identified that histone deacetylases HDAC1 and HDAC7 are necessary to maintain cancer stem cells (CSCs) in both breast and ovarian tumors. Here, we sought to investigate the CSC-specific function of HDAC1 and HDAC7 mechanistically by using a stem-like breast cancer (BrCa) cell model BPLER and matched nonstem tumor cell (nsTC)-like HMLER, along with conventional BrCa cell lines with different CSC enrichment levels. We found that HDAC1 and HDAC3 inhibition or knockdown results in HDAC7 downregulation, which is associated with a decrease in histone 3 lysine 27 acetylation (H3K27ac) at transcription start sites (TSS) and super-enhancers (SEs) prominently in stem-like BrCa cells. Importantly, these changes in chromatin landscape also correlate with the repression of many SE-associated oncogenes, including c-MYC, CD44, CDKN1B, SLUG, VDR, SMAD3, VEGFA, and XBP1. In stem-like BrCa cells, HDAC7 binds near TSS and to SEs of these oncogenes where it appears to contribute to both H3K27ac and transcriptional regulation. These results suggest that HDAC7 inactivation, directly or through inhibition of HDAC1 and HDAC3, can result in the inhibition of the CSC phenotype by downregulating multiple SE-associated oncogenes. The CSC selective nature of this mechanism and the prospect of inhibiting multiple oncogenes simultaneously makes development of HDAC7 specific inhibitors a compelling objective.
Similar content being viewed by others
Introduction
We previously identified a cancer stem cell (CSC)-specific function for the histone deacetylases HDAC1 and HDAC7 in breast and ovarian cancer cells. Cumulatively, our results demonstrated that both HDAC1 and HDAC7 are necessary to maintain CSCs. Moreover, HDAC7 overexpression alone was sufficient to augment the CSC phenotype [1]. In brief, we showed that HDAC1 and/or HDAC7 inhibition downregulates stem cell markers CD44, CD49f, CD326, CD166, and BMI-1 in ten different breast, ovary, and colon cell lines and using six different HDAC inhibitors (HDACis). HDAC1 and/or HDAC7 siRNA knockdown and inhibition by HDACis, reduced 3D tumor sphere formation more significantly compared with 2D proliferation in vitro, and xenograft tumor growth in vivo [1].
HDAC1, a class I HDAC family member, is associated with transcriptional repression through its deacetylase activity [2, 3]. HDAC1 is recruited by DNA-binding transcription factors and is present in repressor multiprotein complexes with Sin3, nucleosome-remodeling deacetylase (NuRD) and REST corepressor 1 (RCOR1/CoREST) [3, 4]. For example, an HDAC1 and NuRD complex was found at transcription start sites (TSS) and active enhancers, where it contributes to their repression during embryonic stem cell (ESC) differentiation, and at super-enhancers (SEs) [5, 6]. Thus, in general HDAC1 inhibition has been associated with transcriptional activation of target genes.
HDAC7, a class II HDAC family member, is also associated with a number of transcription factors and corepressors including MEF2A/C/D [7, 8], STAT3 [9], HIF-1α [10], ERα, FOXP3 [11], KLF4 [12], RUNX2 [13], FOXA1 [14], nuclear receptor corepressor 1 (NCOR1), and nuclear receptor corepressor 2/silencing mediator for retinoid or thyroid hormone receptors (NCOR2/SMRT) [15]. However, the mechanism of transcriptional regulation by HDAC7 is more complex. While HDAC1 actions are dependent on its enzymatic activity, HDAC7 can mediate transcriptional repression by deacetylase-independent mechanisms [8, 13, 16]. In addition, while HDAC1 is predominantly associated with repression, HDAC7 can be associated with both transcriptional activation and repression. For instance, in hematopoietic malignancies, such as pro-B acute lymphoblastic leukemia and Burkitt lymphoma, ectopic expression of HDAC7 represses c-MYC transcription, induces apoptosis and inhibits in vivo oncogenic potential. In contrast, in cervix, breast, and colon epithelial cancer cell lines, such as HeLa, MCF7, and HCT-116, HDAC7 knockdown results in c-MYC repression and consequent increase in p21Cip1 and p27Kip1 protein levels [17, 18]. In human embryonic kidney HEK293 cell line, hypoxia-associated transcriptional activation is associated with cotranslocation of HDAC7 and HIF-1α into the nucleus and interaction with histone acetyltransferase p300 [10]. Thus, HDAC7 seems to activate or repress gene transcription depending on cell lineage (hematopoietic vs. epithelial), as well as microenvironmental context (normoxia vs. hypoxia).
While HDAC1 has been studied in many cancer types before [19,20,21], the role of HDAC7 has been emerging more recently. Since we first reported a specific role for HDAC7 in regulating CSCs, several papers showing that HDAC7 correlates with poor patient outcome, drug resistance, and metastasis in ovarian cancer and gastric cancer have been published [22, 23]. Recently, it was shown that HDAC7 also promotes lung, colorectal cancer, and salivary tumorigenesis [24,25,26].
Pan-HDACis that inhibit all 11 HDACs, such as SAHA (Vorinostat), have been widely used to target many types of cancer. More recently, several class-specific HDACis have been developed. While MS-275 (Entinostat) and MGCD0103 (Mocetinostat), members of the benzamide class of HDACis, were developed as specific inhibitors of HDAC1 and HDAC3 (HDAC1/3) enzymatic activity, we recently reported that these drugs unexpectedly also downregulate HDAC7 protein levels [1]. Interestingly, HDAC7 is also specifically downregulated by pan-HDACis (Vorinostat and Trichostatin A [TSA]); while these drugs inhibit the enzymatic activity of the entire HDAC family, they only seem to downregulate HDAC7 transcript and protein levels, with limited or no effects on the protein levels of other class I and II HDAC family members [27, 28]. These observations raise the possibility that HDAC7 might be one of the downstream convergence points for the actions of both class I and II HDACs.
Here we report that HDAC1, along with HDAC3, regulate HDAC7 expression, which in turn maintains histone 3 lysine 27 acetylation (H3K27ac) near TSS and to SEs and contributes to the expression of SE-associated oncogenes in stem-like BrCa cells. Enhancers are key cis-regulatory elements that can regulate gene transcription independent of distance and orientation through their juxtaposition to gene promoters as a result of folding and looping of the chromatin filament within topologically associating domains along the chromosome [29]. SEs are clusters of enhancers initially identified in mouse embryonic stem cells (mESCs) that are associated with H3K27ac enrichment. In addition, SEs are associated with Oct4, Sox2 and Nanog, and Mediator complex on genes that control mESC pluripotent state [30]. It was found that Hdac1 accumulates at SEs along with other components of the NuRD complex, the bromodomain protein family member Brd4, the histone acetyltransferase p300, and RNA polymerase II (RNA pol II) [6]. SEs are also found in other cell types, usually associated with key cell identity genes where they confer higher transcriptional activity than of genes associated with typical enhancers. Our findings suggest that inactivation of HDAC1/3-HDAC7 axis may inhibit SE-associated oncogene expression in CSCs, which can be a promising pathway to target for developing novel breast cancer (BrCa) therapies.
Results
MS-275 (Entinostat) represses HDAC7 in BrCa cells with stem-like phenotype
Previously, we reported that both HDAC1 and HDAC7 are necessary to maintain the CSC phenotype in breast and ovarian cancer cells and HDAC7 overexpression alone is sufficient to augment the CSC phenotype [1].
We now found that MS-275 (Entinostat), which is a HDAC1 and HDAC3 specific inhibitor, downregulates HDAC7 mRNA and protein in CSC-like BPLER cells. Importantly, HDAC7 mRNA and protein reduction in nonstem tumor cell (nsTC)-like HMLER cells by MS-275 was less pronounced (Fig. 1a, b). While HDAC7 mRNA downregulation (1.7- to 2.0-fold) is observed within 6 h of MS-275 treatment (Fig. 1a), HDAC7 protein reduction is detected after 24 h (Fig. 1b and S1). There was no change in HDAC1 levels (Fig. 1b). In addition, MS-275 increased histone acetyltransferase p300 and total H3K27ac within 6 h followed by total H3K27ac accumulation and p300 depletion at 24 h of treatment in both stem-like and nonstem BrCa cells, consistent with its HDACi activity (Fig. 1b and S1A). However, we observed an opposite effect around the TSS of many stem cell-associated transcription factor (SCTF) genes analyzed in CSC-like BPLER cells, in which MS-275 decreased H3K27ac (n = 50/84) and histone 3 lysine 9/14 acetylation (H3ac) (n = 59/84) (Fig. S2). Accordingly, MS-275 treatment decreased expression of SCTFs, such as c-MYC and VDR in CSC-like BPLER cells. No change in c-MYC and VDR mRNA and protein levels were detected in nsTC-like HMLER cells in response to MS-275 treatment (Fig. 1a, b).
HDAC1/3-specific inhibitor (MS-275) reduces HDAC7 and c-MYC mRNA and protein levels in stem-like BrCa cells. a Treatment with 1 μM MS-275 (Entinostat) reduced HDAC7 mRNA levels in CSC-like BPLER2 (black bars; 1.7-fold at 6 h and 15.0-fold at 24 h) and nsTC-like HMLER2 (gray bars; 2.0-fold at 6 h and 4.5-fold at 24 h) measured by real-time quantitative RT-PCR. In contrast, c-MYC and VDR transcript levels were reduced 2.2- and 1.9-fold at 24 h only in BPLER2 cells and HDAC1 transcription remained unaffected in both cell populations. Transcript levels were normalized to ACTB and B2M and expressed relative to values in vehicle control (DMSO). Results of real-time quantitative RT-PCR from three independent repeats were plotted in bar chart using logarithmic scale, error bars represent standard deviation. b Treatment with 1 μM MS-275 for 24 h reduced HDAC7, c-MYC, and VDR (upper isoform) protein levels in BPLER2 but not in HMLER2 cells. HDAC1 protein level remained largely unaffected while total H3K27ac levels increased considerably in both cell populations. Immunoblots of whole cell lysates were probed with the indicated antibodies, GAPDH and ACTB were used as loading control. c Treatment with 1 μM MS-275 for 24–48 h reduced HDAC7 (≥12-fold) and c-MYC (≥2-fold) mRNA levels in CSC-enriched SUM159 cells but not in ERα+ MCF7 cells. The mRNA for VDR increased up to 3.4-fold only in MCF7 cells. d Treatment with 1 μM MS-275 for 24 h reduced HDAC7 and c-MYC protein levels in SUM159 but not in MCF7, increased total H3K27ac in either BrCa cell line with no detectable change in HDAC1 protein levels
The above results were confirmed in CSC-enriched BrCa cell line (SUM159) vs. a BrCa cell line that is not as CSC-enriched (MCF7) [31]. Similar to BPLER/HMLER pairs, MS-275 treatment downregulated HDAC7 and c-MYC mRNA and protein in CSC-enriched SUM159 cell line without significant change in MCF7 (Fig. 1c, d) and increased total H3K27ac in both cell lines (Fig. 1c, d).
In summary, inhibition of HDAC1/3 by MS-275 represses HDAC7 more prominently in CSC-enriched cells (BPLER and SUM159), decreases H3K27ac and H3ac around the TSS of many SCTF genes and reduces their mRNA and protein levels, while increasing H3K27ac globally.
HDAC1/3/7 inhibition by MS-275 reduces H3K27ac at stem cell transcription factor genes
H3K27ac modifications are typically deposited on histones located next to transcriptionally active TSS, enhancers, and SEs by histone acetyltransferases p300 and CREB-binding protein [6, 32]. We found that while MS-275 treatment increased H3K27ac globally through HDAC1/3 inhibition (Fig. 1b and S1A), it also produced the opposite effect at the promoters of SCTF genes (n = 50/84), such as c-MYC, VDR, RB1, EZH2, c-JUN, HOXA2, and HOXA10 (Fig. S2).
These results prompted us to examine the genome-wide changes in H3K27ac produced by MS-275. Hence, we carried out ChIP-seq analysis in CSC-like BPLER cells treated with MS-275 and found 13,804 H3K27ac peaks at TSS and 24,095 peaks at putative enhancers that are decreased with MS-275 (Fig. 2a). In particular, there was a decrease in H3K27ac at the promoter-proximal regions of SCTF genes, including c-MYC, VDR, RB1, EZH2, RUNX1, c-JUN, HOXA2, and HOXA10 in CSC-like BPLER cells (Fig. 2b and S3A, B), which was not observed in nsTC-like HMLER cells (Fig. S3C). Interestingly, HDAC7 is among the genes with reduced H3K27ac at TSS after MS-275 treatment (Fig. 2b). The reduction in H3K27ac peaks extended several kb beyond the TSS of c-MYC, HDAC7, VDR, RUNX1, c-JUN, and HOXA genes both into intergenic and intragenic regions, similar to the pattern that was identified at SEs [6].
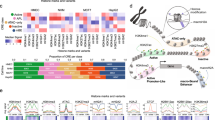
HDAC1/3/7 inhibition by MS-275 reduces H3K27ac on TSS, enhancers, and SEs in stem-like BrCa cells. a Heatmap of genome-wide H3K27ac ChIP-seq analysis revealed enrichment of H3K27ac around TSS (±1 kb) of 13,804 genes that was reduced following treatment with 1 μΜ MS-275 for 24 h in CSC-like BPLER cells. Coverage profiles for H3K27ac enrichment at non-TSS sites (>1 kb) also showed broad distribution of H3K27ac at 25,095 putative enhancers and SEs, which was reduced following MS-275 treatment. b ChIP-seq profiling in BPLER2 cells revealed reduction in H3K27ac peaks near TSS and to SEs of c-MYC, HDAC7, and VDR genes with MS-275 treatment. c Super-enhancers ranking plot based on H3K27ac signal using the ROSE algorithm [30, 33]. Under normal growth condition (DMSO treatment), we identified 1870 putative SE regions characterized by elevated levels of H3K27ac in CSC-like BPLER cells, which are reduced to 573 H3K27ac peak regions in cells under MS-275 treatment (see Supplementary Table 1). d Venn diagram showing the 1287 SE-associated genes (SEs) previously identified in MM, GM, and SCLC [33], among which 447 have been implicated in breast cancer (BrCa) according to the literature. Of these 447 BrCa/SE-associated genes, 361 demonstrated moderate to high H3K27ac peak levels at either proximal or overlapping SEs in BPLER2 cells (see Supplementary Table 2). With MS-275 treatment, nearly two thirds (61%) of BrCa/SE/H3K27ac associated genes (221/361) demonstrated maximum (117; ↓↓↓) to moderate (104; ↓↓) reduction in H3K27ac peaks, 136 (38%) demonstrated minimal reduction (= /↓), and four genes demonstrated an increase in H3K27ac peak levels (↑↑). The remaining 86 BrCa/SE-associated genes had no H3K27ac signal in BPLER2 cells
HDAC1/3/7 inhibition by MS-275 reduces H3K27ac at SE-associated genes specifically in stem-like BrCa cells
Next, we examined the changes in H3K27ac distribution specifically at SEs following MS-275 treatment in CSC-like BPLER cells with ChIP-seq. In general, SEs are identified as clusters of enhancers enriched in H3K27ac and associated with oncogenes, such as c-MYC in cancer cells [6, 33]. We found that H3K27ac is reduced in 1427 of the 1870 SEs after MS-275 treatment in BPLER cells (Fig. 2c; Supplementary Table 1) [30, 33].
In order to examine the MS-275 induced H3K27ac changes at SEs that are associated with BrCa, we first gathered a list of 1287 malignancy-associated SEs that were identified in multiple malignant neoplasms including small cell lung cancer, multiple myeloma, and glioblastoma after excluding noncoding RNAs, microRNAs, histone variants, and unidentified genes (Fig. 2d; Supplementary Table 2) [33]. Next, we determined the BrCa-associated SEs. We found that 447 of the 1287 malignancy-associated genes are implicated in breast cancer in previously published studies and 81% of these genes (n = 361/447) were associated with H3K27ac at TSS/SEs in CSC-like BPLER cells. Moreover, MS-275 treatment decreased H3K27ac in 61% of these BrCa/SE associated genes (n = 221/361) including c-MYC, CD44, CDKN1B, BCL-XL, SLUG, VDR, SMAD3, VEGFA, XBP1, and HDAC7 (Fig. 2b, d, S3A, B and Supplementary Table 2).
These results indicate that, 361 out of 447 malignancy-associated SEs identified in BrCa are potential targets of HDAC1/3/7 regulation.
HDAC1/3/7 inhibition by MS-275 activates transcription in BrCa cells
HDAC activity generally correlates with transcriptional repression due to removal of histone acetyl groups that leads to a more compact chromatin. Thus, inhibition of HDACs would be expected to correlate with transcriptional activation, as it was reported in BrCa cell lines in response to MS-275 [34]. We confirmed these results for several genes that are not typically associated with SEs such as DHRS2, CLU, and CDKN1A, which are upregulated with MS-275 treatment in both nsTC-like HMLER and CSC-like BPLER cells and in four other BrCa cell lines (Fig. 3a and S4A) [31, 33, 34]. Moreover, in BPLER cells no significant change was observed in H3K27ac peak levels at DHRS2 and CLU genes, with a decrease at CDKN1A locus with MS-275 treatment (Fig. S5A; Supplementary Table 1).
HDAC1/3/7 inhibition by MS-275 represses SE-associated genes in stem-like BrCa cells. a Real-time quantitative RT-PCR analysis of MS-275 activated DHRS2, CLU, and CDKN1A genes [34], at 24 h (HMLER2 and BPLER2) and 48 h (MCF7 and SUM159) of treatment with 1 μM MS-275 in BrCa cell lines. All genes demonstrated >2.0-fold increase in transcript levels in all cell lines. b SCTF genes demonstrated 1.8- to 6.2-fold decrease in transcript levels after MS-275 treatment in CSC-like BPLER2 cells. There was no change in nsTC-like HMLER2 cells except for a 2.0-fold decrease in HOXA mRNA levels. c Real-time quantitative RT-PCR analysis of 21 BrCa/SE-associated genes, 20 of which revealed significantly reduced H3K27ac in BPLER2 cells in response to MS-275 treatment (see Supplementary Table 2), demonstrated 1.6- to 15.0-fold decrease in mRNA levels in most of the genes (15/21) and 1.6- to 2.1-fold increase in transcript levels in only ZFP36, c-JUN, and ID1. In contrast, only 4 of 21 genes demonstrated a 1.6- to 4.5-fold decrease in mRNA in HMLER2 cells. d Nine out of 18 BrCa/SE-associated genes demonstrated 1.7- to 14.3-fold decrease in transcript levels, whereas c-JUN, ZFP36, and ID1 confirmed a >2.0-fold activation in MS-275 treated, CSC-enriched SUM159 cells (black bars). Only 2 out of 18 were decreased in ERα+ MCF7 cells (gray bars). Transcript levels were normalized to ACTB and B2M and expressed relative to values in vehicle control (DMSO). Real-time quantitative RT-PCR with each specific set of primers were obtained from 2–5 independent biological at three technical repeats each and plotted in bar chart using logarithmic scale, error bars represent standard deviation
HDAC1/3/7 inhibition by MS-275 represses stem cell transcription factor genes
While HDAC1/3/7 inhibition has been generally correlated with transcriptional activation as described in the previous section, we observed the opposite result in SCTF genes such as c-MYC and VDR in stem-like BrCa cells (Fig. 1). We extended this observation to additional SCTF genes including EZH2, RB1, EP300, HOXA2, and HOXA10, which responded to MS-275 with decreased mRNA (1.8- to 6.2-fold) and reduced H3K27ac and H3ac peaks around their TSS in CSC-like BPLER cells (Fig. 3b and S3A, B). These results indicate that a subset of SCTF genes may be differentially regulated by HDAC1/3/7 in stem-like BrCa cells, compared with global transcriptional regulation by these HDACs.
HDAC1/3/7 inhibition by MS-275 represses SE-associated gene transcription in stem-like BrCa cells
Next, we examined whether BrCa/SE-associated genes are also differentially regulated by HDAC1/3/7 in CSCs. We found that MS-275 reduced H3K27ac peaks in BrCa/SE -associated genes in stem like BPLER cells (Fig. S3; Supplementary Table 2). Consistent with this, we also found that the majority (15 out of 21) of BrCa/SE-associated genes responded to MS-275 with 1.6- to 15.0-fold decrease in mRNA levels in CSC-like BPLER cells (Fig. 3c). Among these, c-MYC, CD44, CDKN1B, SLUG, VDR, SMAD3, and VEGFA are among the most cited in BrCa literature. In SE-associated genes without H3K27ac, such as BCL2, there was no response to MS-275. On the other hand, in agreement with previous reports on the repressive functions exerted by HDAC1 on c-JUN and ID1 gene expression [35, 36], HDAC1/3/7 inhibition demonstrated a 1.6- to 2.1-fold increase in ZFP36, c-JUN, and ID1 transcript levels despite a reduction in H3K27ac peaks at SEs associated with c-JUN and ID1 (Fig. 3c and S5B). In nsTC-like HMLER cells, only VEGFA, CDKN1B, and XBP1 demonstrated a 1.7- to 2.2-fold decrease in transcript levels in agreement with a less pronounced MS-275 repression of HDAC7 (4.5-fold) on these cells.
In order to confirm these results in additional models, we examined the CSC-enriched SUM159 and SUM149 cell lines [31], in which many of the BrCa/SE-associated genes (7/18 and 13/18 respectively) including c-MYC, CDKN1B, BCL2, SMAD3, VEGFA, and XBP1 responded to MS-275 and HDAC7 downregulation, with 1.7- to 23.8-fold reduction in mRNA levels. Similar to BPLER, the control genes c-JUN, ZFP36, and ID1 were activated by MS-275 treatment in SUM159 cells (Fig. 3d and S4B). In contrast, nearly equal number of BrCa/SE-associated genes were upregulated (3–4/18) and downregulated (2–3/18) in ERα+ MCF7 and MDA-MB-157 in response to MS-275 treatment, consistent with the lack of HDAC7 repression in these BrCa cell lines (Fig. 3d and S4C).
Cumulatively these data indicate that M-275 inhibition of HDAC1/3/7 axis differentially alters H3K27ac landscape and gene transcription in breast CSCs vs. nsTCs, particularly in SE regulated oncogenes and SCTFs (Fig. S6).
Short-term HDAC1 knockdown alone is sufficient to downregulate HDAC7
In the above experiments we used a HDAC1/3 inhibitor MS-275, which specifically downregulates HDAC7 in stem-like BrCa cells, suggesting that HDAC7 expression might be regulated by HDAC1 and/or HDAC3. Since drugs that inhibit HDAC1 or HDAC3 individually are not available, we next used HDAC-specific siRNAs in the remainder of the experiments to examine the role of HDAC1 and HDAC3 separately and in combination.
While HDAC1 knockdown for 72 h by siH1 did not change HDAC7 mRNA expression (Fig. 4a), it resulted in reduced HDAC7 protein (Fig. 4b). However, in contrast with MS-275, there was no significant change in c-MYC or VDR expression (mRNA and protein), total H3K27ac or H3ac levels (Fig. 4a, b).
Short-term HDAC1 knockdown downregulates HDAC7. a Real-time quantitative RT-PCR analysis confirmed a significant siRNA-mediated knockdown of HDAC7 (4.7- and 8.0-fold) and HDAC1 (37.5- and 29.8-fold) mRNAs respectively in nsTC-like HMLER2 (gray bars) and CSC-like BPLER2 (black bars) cells at 72 h post transfection. HDAC1, MYC, and VDR mRNA levels decreased 1.7- to 2.3-fold with HDAC7 specific siRNAs (siH7) in BPLER2 but not HMLER2 cells. As for VDR and c-MYC, HDAC7, mRNA levels did not change significantly with HDAC1-specific siRNAs (siH1) transient transfection in both cell populations. Transcript levels were normalized to ACTB and B2M and expressed relative to values in nontargeting (NT) siRNAs control. Results of real-time quantitative RT-PCR from three independent repeats were plotted in a bar chart using logarithmic scale. Error bars represent standard deviations. b HDAC7 protein expression was reduced with transient transfection of siH1 for 72 h in HMLER2 and BPLER2 cells, but no change was observed in HDAC1 protein expression with transient transfection of siH7. Nontargeting siRNAs (NT) were used as control. No change in c-MYC and VDR expression or total H3K27ac and H3K9/14ac (H3ac) levels was found at 72 h after transient transfection with either siH1 or siH7. Immunoblot of whole cell lysates were probed with the indicated antibodies
HDAC7 knockdown for 72 h by siH7 resulted in 1.7- to 2.3-fold reduction in c-MYC, HDAC1, and VDR mRNA only in BPLER cells (Fig. 4a). However, there were no associated changes in protein expression at this short-time point (Fig. 4b).
Therefore, the decrease in HDAC7 protein can be induced by short-term HDAC1 knockdown alone, without inhibition of HDAC3. Furthermore, the reduction in HDAC7 protein precedes changes in HDAC7 and c-MYC mRNAs, or total H3K27ac/H3ac, suggesting that this activity is not through HDAC7 transcriptional regulation by c-MYC or H3K27ac/H3ac.
Long-term HDAC7 knockdown represses SCTFs in stem-like BrCa cells
HDAC7 knockdown was previously shown to repress c-MYC expression in HeLa, HCT-116, and MCF7 cell lines [18]. Since short-term HDAC1 or HDAC7 knockdown did not produce the reduction in SCTFs such as c-MYC, EZH2, and VDR that was previously observed by MS-275 treatment of stem-like BrCa cells (Fig. 1) [1], we next evaluated long-term HDAC1 and HDAC7 depletion with two consecutive rounds of knockdown up to 6 days.
In CSC-like BPLER cells, two rounds of HDAC7 knockdown resulted in reduction of c-MYC and CD44 proteins and changes in VDR isoforms expression without significantly affecting HDAC1 or EZH2 mRNA or protein levels (Fig. 5a, b). As previously shown in HeLa cells [18], cell-cycle inhibitor p27Kip1 is upregulated by long-term HDAC7 knockdown in BPLER cells, likely due to c-MYC repression (Fig. 5b) [37].
Long-term HDAC1/3 double knockdown maintains HDAC7 downregulation while HDAC7 knockdown represses SCTFs in stem-like BrCa cells. a Long-term HDAC7 knockdown (5.8- to 11.2-fold) with siH7 resulted in decreased HDAC1 (1.7-fold), VDR (1.9-fold), and c-MYC (2.7-fold) mRNAs in CSC-like BPLER2 (black bars) but not in nsTC-like HMLER2 (gray bars) cells. In contrast, HDAC1 knockdown with siH1 in either cell line (55.6- and 34.5-fold) did not significantly change c-MYC or HDAC7 mRNA levels while increasing EZH2 and VDR transcripts (3.3- and 1.8-fold) in BPLER2 cells after 6 days and two rounds of transient siRNA transfection. Transcript levels were normalized to ACTB and B2M and expressed relative to values in nontargeting (NT) siRNAs control. Results of real-time quantitative RT-PCR from three independent repeats were plotted in a bar chart using logarithmic scale. Error bars represent standard deviation. b Long-term transient transfections with siH1, siH7, and NT siRNAs as control resulted in stable HDAC1 and HDAC7 knockdown in BPLER2 and HMLER2 cell populations. In HDAC7-depleted BPLER2 cells, there was significant reduction of c-MYC and CD44, alterations in VDR isoforms and p27Kip1 upregulation. HMLER2 cells revealed only minimal or no change in c-MYC, CD44, VDR, EZH2, or p27Kip1 protein levels after second run of transfection with siH1 and siH7, which is consistent with a lack of MS-275 effect on these cells. c Long-term HDAC1 and HDAC3 double knockdown by combined suboptimal siH1 and siH3 concentrations resulted in reduced HDAC7 protein levels in CSC-like BPLER cells, which is also accompanied by a decrease in c-MYC and CD44 protein levels and change in VDR isoforms expression when compared with siH1/NT and siH3/NT siRNA combinations. Immunoblots of whole cell lysates were probed with the indicated antibodies, ACTB was used as loading control
It appears that while HDAC7 expression is downregulated with short-term HDAC1 knockdown (Fig. 4b), it recovers during long-term knockdown in BPLER2 cells (Fig. 5c and S7A). Consistent with this observation, we found that long-term HDAC1 knockdown alone resulted in no major changes in c-MYC, CD44, VDR, or EZH2 expression in BPLER cells, with partial to no reduction in HDAC7 protein levels (Fig. 5c and S7A).
In nsTC-like HMLER cells there was no change in c-MYC, CD44, EZH2, VDR, or p27Kip1 expression after extended HDAC1 or HDAC7 knockdown and no change in HDAC7 protein levels by long-term HDAC1 depletion (Fig. 5a, b and S7A), which is consistent with the absence of any MS-275 downregulating effect on SCTFs in these cells.
Long-term HDAC1/3 double knockdown maintains HDAC7 downregulation in stem-like BrCa cells
Inhibition of HDAC1/3 with MS-275 downregulates both HDAC7 and SCTFs in stem-like BrCa cells (Fig. 1). In contrast, HDAC1 knockdown alone produces only partial HDAC7 loss and no SCTFs downregulation. Therefore, we examined the role of HDAC3. Interestingly, long-term HDAC3 knockdown with HDAC3-specific siRNAs (siH3) in CSC-like BPLER cells did not significantly affect HDAC7 expression. In contrast, the long-term HDAC1/3 double knockdown did reduce HDAC7, as well as c-MYC and CD44 protein levels (Fig. 5c).
These results indicate that the long-term HDAC1/3 double knockdown mimics MS-275 (Entinostat) in maintaining HDAC7 downregulation, which is necessary and sufficient for repression of c-MYC and CD44 expression in stem-like BrCa cells.
HDAC7 knockdown decreases total H3K27ac in stem-like BrCa cells
We next used H3K27ac that is associated with transcriptionally active chromatin as a marker to examine the changes produced by HDAC1/3 and HDAC7 depletion.
Consistent with the general repressive activity of HDACs, we found that inhibition of HDAC1/3 by MS-275 results in a global increase in H3K27ac levels in both stem-like and nonstem BrCa cells (Fig. 1b, d and S1A). In agreement with this, long-term knockdown of HDAC1 and HDAC3 with combined siH1 and siH3 also increased total H3K27ac levels in CSC-like BPLER cells, mimicking the MS-275 effect. Interestingly, selective long-term knockdown of HDAC3 with siH3 was sufficient to increase total H3K27ac levels indicating that this activity of MS-275 is likely to be through HDAC3 inhibition (Fig. 6a).
Long-term HDAC7 knockdown decreases total H3K27ac in stem-like BrCa cells. a In CSC-like BPLER2 cells, selective long-term knockdown of HDAC3 with siH3 resulted in significant increase in total H3K27ac levels (3.3-fold) indicating a major role exerted by HDAC3 in maintaining total H3K27ac homeostasis. Extensive HDAC1/3 co-depletion also increased total H3K27ac levels (2.5-fold), all effects that recapitulate the MS-275 effect on stem-like BrCa cells. b In CSC-like BPLER2 cells, serial transfection with HDAC7-specific siRNAs caused a substantial decrease in total H3K27ac levels (10.0-fold). HMLER2 cells did not show substantial reduction in total H3K27ac levels following long-term HDAC1 or HDAC7 depletion. The H3K27ac levels were normalized towards total H3 levels
In contrast, selective long-term downregulation of HDAC7 directly with siH7 (Fig. 6b) or indirectly through siH1 (Fig. S7) resulted in a significant decrease in total H3K27ac levels in CSC-like BPLER cells. Furthermore, in nsTC-like HMLER cells neither HDAC1 nor HDAC7 knockdown had any effect on total H3K27ac levels (Fig. 6b and S7).
These results demonstrate a significant difference between HDAC7 and HDAC1/3 function. The total H3K27ac levels are increased by HDAC1/3 inhibition with MS-275 or HDAC3 knockdown in all cells we tested. In contrast, the total H3K27ac levels are decreased by HDAC7 knockdown, and this is observed only in CSC-like BPLER cells and not in nsTC-like HMLER cells.
HDAC7 regulates global gene expression differentially in stem-like vs. nonstem BrCa cells
In order to evaluate the role played by HDAC7 in transcriptional changes induced by MS-275, we performed RNA-seq analysis after 72 h HDAC7 knockdown.
HDAC7 knockdown by siH7 altered the expression of threefold as many genes in BPLER vs. HMLER cells (2545 vs. 763), with the two cell lines sharing altered expression of 328 genes. In contrast, HDAC1 knockdown by siH1 resulted in nearly equal number of altered transcripts in BPLER vs. HMLER cells (3187 vs. 2627), with the two cell lines sharing altered expression of 637 genes (Fig. 7a; Supplementary Table 3). In addition, HDAC7 knockdown resulted in alterations of significantly fewer gene transcripts compared with HDAC1 knockdown (508 vs. 2681) in HMLER cells (Fig. S8A). Our results cumulatively indicate that HDAC7 knockdown causes more significant changes in overall transcription in CSC-like cells compared with nonstem like tumor cells.
HDAC7 regulates gene expression predominantly in stem-like BrCa cells. a RNA-seq analysis after 72 h of HDAC7 knockdown with siH7 revealed predominant alteration of transcriptional activity in CSC-like BPLER2 (black circles) compared with nsTC-like HMLER2 (white circles) cells (2545 vs. 763; 328 shared), with 1068 vs. 448 repressed and 1477 vs. 315 activated genes. Otherwise, short-term HDAC1 knockdown with siH1 demonstrated comparable deregulated gene numbers in BPLER vs. HMLER cells (3187 vs. 2627; 637 shared), with 1570 vs. 1307 repressed and 1617 vs. 1320 activated transcripts in the two cell lines. b Differential expression analysis after short-term HDAC7 knockdown revealed SE-associated transcriptional alteration predominantly in CSC-rich BPLER cells (391 vs. 161; 85 shared), with >1.5-fold repression of 210 vs. 84 and activation of 181 vs. 77 SE-associated genes when compared with ns-TC-like HMLER cells. HDAC1-depleted BPLER and HMLER cells showed comparable SE-associated gene deregulation (458 vs. 404; 107 shared), although with higher number of repressed (334 vs. 142) and lower number of activated (124 vs. 262) mRNA levels in CSC-like BPLER cell line. Differential expression based on DESeq2 analysis with adjusted cutoff of p < 0.05
HDAC7 regulates SE-associated gene expression differentially in stem-like vs. nonstem BrCa cells
Next, we examined the regulation of 1870 SE regions identified in CSC-like BPLER cells (Fig. 2c). Similar to the global changes, we found that HDAC7 knockdown caused more significant alterations in BrCa/SE-associated gene transcripts in CSC-like BPLER (210 down and 181 up) compared with nsTC-like HMLER cells (84 down and 77 up), with 58 repressed and 27 activated genes shared between the two HDAC7-depleted cell lines (Fig. 7b; Supplementary Table 3). In contrast, HDAC1 knockdown resulted in nearly equal number of changes in BPLER vs. HMLER (458 vs. 404) with the two cell lines sharing altered expression of 107 BrCa/SE-associated genes.
Furthermore, in HMLER cells, HDAC7 knockdown resulted in alterations of significantly fewer BrCa/SE-associated gene transcripts compared with HDAC1 knockdown (112 vs. 377); especially in activated (38 vs. 220) transcripts (Fig. S8B), in agreement with a CSC-specific regulation of BrCa/SE-associated gene expression by HDAC7.
We confirmed the above RNA-seq results with real-time quantitative RT-PCR analysis in a subset of BrCa/SE-associated genes. The short-term knockdown of HDAC7 resulted in repression (1.7- to 4.6-fold) in 50% (9/18) of BrCa/SE-associated genes tested in CSC-like BPLER cells vs. 22% (4/18) in nsTC-like HMLER cells (Fig. S9A). In contrast, knockdown of HDAC1 resulted in repression of only 1 out of 18 BrCa/SE-associated genes in BPLER cells and increased expression (1.7- to 3.6-fold) of 4 and 6 genes in BPLER and HMLER cells, respectively (Fig. S9B).
We found that 127 BrCa/SE-associated genes are repressed by both HDAC1 and HDAC7 knockdown in CSC-like BPLER cells. It is beyond the scope of this initial study to examine the relative contribution of each of these genes to the CSC phenotype. Nevertheless, as a proof-of-concept we selected two cytokines IL6 and IL34 for functional testing among these 127 genes (Fig. S8B; Supplementary Table 3). Interestingly, while IL6 is a well-known factor that stimulates CSC phenotype, IL34 was an unexpected finding that has not been examined in epithelial tumors. We found that IL34 alone and in combination with IL6 stimulated mammosphere formation, which is a functional assay for CSCs (Fig. S10).
Cumulatively, these results highlight that HDAC7 regulates nearly three times as many genes in stem-like compared with nonstem BrCa cells. In contrast, HDAC1 regulates equal number of genes in the same cell context, suggesting a predominant association of HDAC7 with the CSC phenotype.
HDAC7 directly regulates transcriptional activity at SE-associated genes
In order to confirm direct binding of HDAC7 to the BrCa/SE-associated genes, we carried out Quantitative chromatin immunoprecipitation (qChIP) analysis and found that HDAC7 knockdown in CSC-like BPLER cells resulted in a 3.2- to 11.1-fold reduction in HDAC7 binding to BrCa/SE-associated c-MYC, c-JUN, SMAD3, CDKN1B, BHLHE40, VDR, VEGFA, and CD44 gene loci (Fig. S11A). Importantly, the decrease in HDAC7 binding to BrCa/SE-associated SMAD3, c-MYC, VEGFA, and VDR gene loci were coupled with a 1.7- to 2.6-fold decrease in mRNA (Fig. S9A), suggesting that these genes are under a positive transcriptional control by HDAC7. In these experiments, HDAC7 binding was also reduced at SE regions overlapping the HDAC7 locus, suggesting an autoregulation mechanism.
As shown above, short-term HDAC1 knockdown results in partial reduction in HDAC7 protein levels without decrease in BrCa/SE-associated gene transcripts (see Fig. 4 and S9B). Consistent with these findings, there was no significant change in HDAC7 binding in the majority of the BrCa/SE-associated genes (seven out of ten) following HDAC1 knockdown (Fig. S11B). In addition, SE regions in the VDR locus revealed a significant increase in HDAC7 binding directly correlated with an increase in VDR mRNA levels following HDAC1 depletion (Figs. S11B and S9), once again in support of a positive VDR transcriptional control by HDAC7.
In summary, HDAC7 knockdown results in reduced HDAC7 binding to TSS and SEs, coupled with decreased H3K27ac and mRNA expression of SCTF and BrCa/SE-associated genes, specifically in stem-like BrCa cells.
Discussion
The mechanistic model we elaborate in this study (Fig. 8) is emerging from multiple comparisons that contrast the function of different HDAC isoforms (HDAC7 vs. HDAC1/3), the role of the cellular context (stem vs. nonstem cells), chromatin markers (H3 vs. H3K27ac) and SE vs. genome-wide patterns.
Model of SE-associated gene regulation by HDAC1/3-HDAC7 axis in CSCs. The cumulative results of our experiments with HDAC1/3/7 inhibitor MS-275 (Entinostat) as well as HDAC1-, HDAC3-, and HDAC7-specific siRNAs are depicted in a mechanistic model. The direct repressive activity of HDAC1 and HDAC3 is shown by “MS-275 activated genes”. In addition to this well-known activity, our results indicate a novel HDAC1/3 mechanism through HDAC7, which in turn maintains H3K27ac and expression of BrCa relevant, SE-associated oncogenes such as c-MYC, CD44, CDKN1B, BCL-XL, SLUG, VDR, VEGFA, and SMAD3. [18, 42] Importantly, this HDAC1/3-HDAC7 axis is only active in stem-like cells and not in other cells
A major question in cancer biology is how CSC differentiation phenotype is maintained over multiple generations of cell divisions. HDACs play a critical role in normal stem cell and tissue differentiation and our results suggest that CSC may appropriate a similar mechanism of regulation. At least in the case of breast CSCs HDAC7 seems to play a critical role in regulating a differentiation program involving many oncogenes and stem cell transcription factors. It is possible that other HDACs may play a similar role in other tissue types, also HDAC7 may be involved in other tumors, as suggested by recent studies in ovarian, gastric, lung, colon, and salivary cancers [19,20,21,22,23,24].
Regulation of HDAC7 through HDAC1/3 in breast CSCs
We previously reported that HDAC1/3/7 inactivation by MS-275 (Entinostat) and MGCD0103 (Mocetinostat) inhibits CSCs proliferation both in vitro and in vivo [1]. In addition, MS-275, which inhibits enzymatic activity of HDAC1/3, downregulates HDAC7, c-MYC, and CD44 only in CSCs and increases total H3K27ac levels. However, while knockdown of HDAC1 alone by siRNA downregulated HDAC7, it did not downregulate c-Myc and CD44 or upregulate H3K27ac, suggesting that HDAC3 may also play a role in MS-275 activity.
In this study, we found that double HDAC1 and HDAC3 knockdown produces the reduction in HDAC7, MYC, and CD44 protein levels and upregulation of H3K27ac as seen with MS-275 (Figs. 5c and 6a). In contrast, long-term HDAC3 knockdown was not sufficient to downregulate HDAC7, c-MYC, or CD44 protein levels (Fig. 5c), but upregulated H3K27ac.
Cumulatively these results suggest that MS-275 downregulates HDAC7 through HDAC1, upregulates H3K27ac through HDAC3, and downregulates c-MYC and CD44 through combined inhibition of HDAC1 and HDAC3.
Breast CSC specific regulation of H3K27ac and SEs by HDAC7
Increased H3K27ac is associated with transcriptionally active chromatin. Our results indicate that HDAC1/3 and HDAC7 differentially alter H3K27ac landscape and gene transcription in breast CSCs vs. nsTCs.
We discovered a significant difference between HDAC7 vs. HDAC1/3 function in the regulation of H3K27ac in CSCs. While HDAC3 knockdown, or HDAC1/3 inhibition by MS-275, increased total H3K27ac levels in both stem and nonstem BrCa cells, HDAC7 knockdown reduced total H3K27ac levels, but only in CSCs and not in other cells.
Furthermore, we found that HDAC7 knockdown altered the expression of more than threefold as many genes in breast CSCs vs. nonstem BrCa cells, including SE and malignancy associated genes. In contrast, HDAC1 knockdown altered similar number of SE and malignancy associated genes in breast CSC vs. nonstem BrCa.
Cumulatively, these results suggest that HDAC7 is more closely associated with the CSC phenotype compared with HDAC1 and HDAC3. Furthermore, results from many studies suggest that HDAC7 may be a downstream target for the entire HDAC family. While all tested HDACis such as SAHA (Vorinostat), TSA, sodium butyrate, Entinostat, Mocetinostat, and valproic acid inhibit selective or the entire members of the HDAC family, these agents specifically downregulate HDAC7, with minimal effects on other HDACs [1, 27, 28, 38].
Pan-HDAC inhibitors have been used in the clinic to treat several cancer types. These drugs inhibit all 11 HDAC family members and their mechanism of action has been difficult to discern due to their nonspecificity. It stands to reason that such a broad spectrum may produce side effects to limit their use in the clinic. Our results suggest that selective inhibition of HDAC7 may suffice to achieve the anti-CSC activity of pan-HDACis.
Activation of transcription by HDACs
Despite the seemingly global control of gene expression by HDAC proteins, it is reported that less than 5–10% of gene transcription is influenced by HDAC inhibitors. Moreover, approximately equal numbers of genes are upregulated or downregulated by HDACis [34, 39].
While activation of gene expression by HDAC inhibitors is easily explained by release of repressive HDAC enzymatic activity, the mechanism of gene repression by HDAC inhibitors has been less clear. HDACs are generally thought to repress transcription through creating a compact chromatin state by enzymatically removing acetyl groups from histones. Consequently, HDACis are expected to cause transcriptional activation by inhibiting the removal of histone acetyl groups [34, 40].
Several studies previously reported repression of gene expression by HDAC inhibitors and decrease in H3K27ac, H3ac, and H4ac near the promoters of many genes, despite an increase in total histone acetylation in breast and colon cancer cell lines [41,42,43]. In other studies it was shown that HDAC7 knockdown reduced RNA pol II association with c-MYC in HeLa cells [18], and HDAC1 overexpression relieved repression of SE-associated ERBB2 and c-MYC oncogenes by HDAC inhibitors in BrCa cells [6, 42]. Transcriptional repression by HDAC inhibitors could be due to indirect secondary effects. For example, some of the genes that are directly activated by HDAC inhibitors may in turn repress secondary targets. Our results provide another potential mechanism for these seemingly unanticipated results. It appears that the simultaneous HDAC1 and HDAC3 inhibition may lead to increase in total histone acetylation and transcriptional activation. In contrast, HDAC1/3-HDAC7 axis inhibition is associated with reduced H3K27ac and repression at SE-associated genes, particularly in stem-like BrCa cells (Fig. 8).
In summary, we previously showed that HDAC7 is overexpressed in breast, ovary, and colon CSCs. Moreover, our results indicated that HDAC7 is necessary and sufficient to maintain the CSC phenotype in these cells [1]. Independently, studies from several laboratories showed that HDAC7 expression correlates with poor patient outcome, drug resistance, and metastasis in ovarian, gastric, lung, colon, and salivary cancers [19,20,21,22,23,24]. Interestingly, HDAC7 is the only HDAC family member that is degraded by all HDAC inhibitors tested, suggesting that it may be a point of convergence of HDAC inhibition [1, 27, 28, 38]. Here we show that HDAC7 regulates H3K27ac levels and transcriptional activity of SE-associated oncogenes and SCTFs differentially in breast CSCs vs. nsTCs, by directly binding to SEs. To our knowledge, there are no small molecules that inhibit HDAC7 selectively. Such a HDAC7-specific inhibitor can be a powerful agent to inhibit multiple oncogenes simultaneously with less side effects compared with existing pan- or class I-specific HDAC inhibitors.
Materials and methods
Cell lines and culture conditions
BPLER and HMER cells were described before. [44] Both cells and BPLER culture medium (BMI-T) are available from Live Tissue Culture Service Center (contact LTCC@med.miami.edu). HMLER cells were cultured in MEGM medium (Lonza). SUM159PT and SUM149PT TNBC cell lines were cultured in F12 medium containing 5% FBS. MCF7 was cultured in DMEM medium containing 10% FBS. MDA-MB-157 was cultured in L-15 medium with 10% FBS in the absence of CO2. There were not significant changes in morphology after treatment with MS-275 or with HDAC1-, HDAC3-, and HDAC7-specific siRNAs.
siRNA knockdown of HDAC1, HDAC3, and HDAC7
BPLER and HMER cells were plated at 7 × 105 cells per plate 24 h before transfection and transfected with a pool of 50 nM each of double-stranded RNAs using Lipofectamine RNAiMAX (Invitrogen) in OptiMEM (GIBCO, Life Technologies) added directly to the medium. The cells were harvested 72 h later, stained with trypan blue, counted, and plated for long-term knockdown analysis. The next day, a second round of transfection was carried out under identical conditions. For double knockdown analysis, a final concentration (f.c.) of 30 nM each of the combined siRNAs was used. The siRNAs used were siGENOME Non-Targeting siRNA Pool #2, ON-TARGETplus Non-Targeting Pool, and the following SMARTpools: siGENOME and ON-TARGETplus Human HDAC7 (51564), ON-TARGETplus Human HDAC3 (8841) and siGENOME, and ON-TARGETplus Human HDAC1 (3065) siRNAs (Dharmacon, GE Healthcare).
Immunoblotting and antibodies
Whole-cell lysates of 1 × 106–1.5 × 106 cells in RIPA buffer containing protease inhibitors were sonicated and 30–50 μg of each protein extract was resolved on 4–12% NuPAGE gels (Invitrogen). Blocking and incubation with primary and secondary antibodies were performed with 5% skim milk in phosphate-buffered saline. Rabbit polyclonal/monoclonal antibodies: anti-HDAC7 (ab12174), anti-HDAC3 (ab1791), anti-HDAC1 (ab7028), anti-H3K27ac (ab4729), and anti-c-MYC (ab32072) were obtained from Abcam, Inc. Rabbit polyclonal anti-H3ac (06–599) antibody was obtained from Millipore and anti-CD44 Pan Specific (BBA10) antibody from R&D Systems. Rabbit polyclonal/monoclonal antibodies: anti-EZH2 (#5246), anti-p27Kip1 (#3686), and anti-GAPDH (#5174) were obtained from Cell Signaling Technology. Rabbit polyclonal anti-VDR (sc-1008), anti-p300 (sc-584), and mouse monoclonal anti-ACTB (sc-69879) antibodies were obtained from Santa Cruz Biotech. Horseradish peroxidase-conjugated secondary antibodies were obtained from Amersham Biosciences. The protein bands were visualized with SuperSignal West Dura or Femto ECL substrate (ThermoScientific). All experiments were repeated a minimum of three times. Autoradiographic signals were quantified by densitometric analysis using ImageJ from National Institute of Health.
Real-time RT-PCR analysis
Total RNA preparations were cleaned with Turbo DNA-Free Kit (Ambion), quantified, and reverse-transcribed in equal amounts with random primers using SuperScript III reverse transcriptase (Invitrogen) at 55 °C or qScript XLT cDNA SuperMix (Quanta Biosciences) at 42 °C. For SYBR green incorporation-based, real-time PCR analysis of HDAC1, HDAC7, SCTF, and SE-associated genes, primers (Supplementary Table 4) and PCR conditions described previously were used along with SsoFast EvaGreen Supermix (Bio-Rad) or PerfeCta SYBR Green FastMix (Quanta, Biosciences). The results were normalized toward real-time PCR analysis of human β-actin (ACTB), and β-2-microglobulin (B2M) expression levels. Relative quantification of gene transcripts was performed using the comparative threshold cycle method on the CFX96 Real-Time Detection system and analysis software (BioRad).
RNA-seq analysis
DNase I-treated total RNA preparations were qualitatively validated by Eukaryote Total RNA 6000 Nano analysis on 2100 Bioanalyzer (Agilent) and libraries were prepared using the KAPA RNA HyperPrep kit according to manufacturer instructions (KAPA). Libraries were sequenced on a 75 bp paired-end run using the NextSeq 500 High Output Kit (150-cycle; 400 M flow cell). RNA-seq data were uploaded to the Galaxy web platform and we used the public server at galaxy.org to analyze the data [45]. RNA-seq data sets were aligned using HISAT2 (version 2.1.0) to reference version NCBIv37/hs19 of the human genome; transcript assembly, quantification, and merging with StringTie (version 1.3.4). Number of reads per transcript was counted using feature counts (version 1.6.3) and differential expression analysis was performed using DESeq2 (version 2.11.40.2). A cutoff of 0.6 < log2 < −0.6, p < 0.05 was applied for the identification of genes with significant differential expression.
Chromatin immunoprecipitation and real-time PCR analysis
qChIP analysis was carried out as previously described [40]. After MS-275 or siRNA treatment, BPLER2, and HMLER2 cells were crosslinked for 10 min at RT by adding formaldehyde to an f.c. of 1%. The reaction was stopped by adding glycine to 0.125 M f.c. and the cells were resuspended in SDS lysis buffer (50 μM Tris-Cl, pH 8.0; 10 μM EDTA; 1% SDS; 1 μM PMSF) plus complete protease inhibitor cocktail tablets (Roche). The chromatin was sonicated 10–25 cycles of 30 s ON and 30 s OFF each at High setting on a Bioruptor sonication system (Diagenode) to generate fragments averaging 0.5–1.5 kb. Protein A Sepharose beads (Amersham Biosciences) were incubated with the chromatin for 1 h at 4 °C for preclearing. The precleared chromatin was incubated overnight at 4 °C with 4 μg of rabbit polyclonal anti-H3K27Ac (ab4729) or anti-HDAC7 (ab12174) antibody. Immune complexes were collected on Protein A Sepharose beads for 1–2 h at 4 °C. Following washes, the beads were resuspended in TE buffer (10 μM Tris-Cl, pH 8.0; 1 μM EDTA) and RNase-treated with 0.5 μg/μl RNase A for 30 min at 37 °C. The immune complex was eluted from the beads in 1% SDS and 0.1 M NaHCO3 and crosslinks were reversed by incubation in 0.2 M NaCl for 6 h at 65 °C. The resulting DNA was purified by proteinase K treatment, phenol/chloroform/isoamyl alcohol extraction, ethanol precipitation, and suspended in water. Purified DNA was quantified by the Qubit dsDNA HS assay (Life Technologies) and 20 ng or more of DNA was mixed in 1× SSoFast EvaGreen Supermix (Bio-Rad) containing sets of primers for H3K27ac-rich regions of SCTF and SE-associated genes. Real-time PCR and dissociation curve analyses were performed on the CFX96 Real-Time Detection system and analysis software (BioRad). The results were calculated according to the ∆∆Ct method. To account for differences in chromatin preparation between samples, input normalized Ct values for every IP DNA (∆Ct[IP] = Ct[IP] − Ct[input]) were used for calculation of genomic site occupancy: % input = 2(−[Ct[IP]−Ct[input]) × 5%, where 5% is the percentage of total chromatin used as input. The percent input enrichment, or input normalized value, was calculated by subtracting the % input of the control IP (% input [noAb]) from the % input of the antibody IP (% input [Ab]): percent input enrichment = % input [Ab] − % input [noAb]. The fold enrichment of each IP towards control IP (or signal over background) was calculated by the linear conversion of differences between ∆Ct[IP] and ∆Ct[noAb], or ∆∆Ct[IP]: fold enrichment = 2(−∆∆Ct[IP]). For each of the loci analyzed, the difference in fold enrichment between MS-275 and control (DMSO) treated HMLER2 and BPLER2 cells or between siHDAC1/7 (siH1/7) and nontargeting (NT) siRNAs transfected BPLER2 cells were determined by dividing the treatment (MS-275 or siH1/7) fold enrichment by the control (DMSO or NT siRNAs) fold enrichment: fold change = treatment fold enrichment/control fold enrichment. Each ChIP experiment was repeated three times and the results on a representative experiment are shown.
ChIP-seq analysis
Chromatin for ChIP-seq analysis was prepared as described above with extended sonication conditions to generate fragments averaging 0.2–0.6 kb. Libraries for paired-end sequencing and alignment were prepared according to the TruSeq ChIP Library Preparation Kit according to manufacturer instructions (Illumina). After AMPure XP® Beads (Beckman Coulter, Inc.) purification and PCR amplification, the DNA library was quantified using the KAPA Library Quantification Kit (KAPA Biosystems), quality controlled on Bioanalyzer® (Agilent Technologies, Inc.), and sequenced on a 75 bp paired-end run using the NextSeq 500 Mid Output Kit. ChIP-seq data sets were aligned using Bowtie (version 0.7.12) to reference version NCBIv37/hs19 of the human genome. We used SICER version 0.1.1 broad-domain peak finding algorithm at the false discovery rate (FDR) of <0.01, window size of 200 and gap size of 600 to identify regions of H3K27ac enrichment over background. Peak regions were further filtered after SICER peak calling with adjusted p-value (FDR) <0.01 and fold change >2.5 to identify peaks overlapping with promoter regions (±1 kb from TSS) and with intergenic and intragenic loci by excluding the promoter-associated regions (>1 kb from TSS), also known as putative enhancers. SEs were mapped using the ROSE software package described in Lovèn at al. 2013 and Whyte et al. 2013 (https://bitbucket.org/young_computation/rose) [30, 33]. Genome peak profiles where visualized using Integrated Genome Browser (IGB, BioViz.com).
Mammosphere formation assay
Mammospheres assays were carried out in BPE excluded MEGM medium (Lonza) supplemented with 2% B27 (Invitrogen, Carlsbad, CA, USA), 20 ng/ml EGF, 20 ng/ml basic FGF (BD Biosciences), 4 μg/ml heparin (Sigma, St Louis, MO, USA), and 1% methylcellulose. Single cells were plated in ultralow attachment plates (Corning) at a density of 5000 viable cells in 2 ml per well. After 10 days, the colonies were stained with 2-(4-iodophenyl)-3-(4-nitrophenyl)-5-phenyl-2H-tetrazolium chloride (INT), and scanned using automated colony counting system GelCount™ (Oxford Optronix, Oxford, UK). All experiments were carried out with triplicates. Error bars represent the standard error of the mean of three replicates.
Public data deposition
The ChIP-seq and RNA-seq data were deposited to Gene Expression Omnibus (GSE131632), accession codes for deposited data includes ChIP-seq (GSE131436), RNA-seq (GSE131631).
References
Witt AE, Lee C-W, Lee TI, Azzam DJ, Wang B, Caslini C, et al. Identification of a cancer stem cell-specific function for the histone deacetylases, HDAC1 and HDAC7, in breast and ovarian cancer. Oncogene. 2017;36:1707–20.
Minucci S, Pelicci PG. Histone deacetylase inhibitors and the promise of epigenetic (and more) treatments for cancer. Nat Rev Cancer. 2006;6:38–51.
Seto E, Yoshida M. Erasers of histone acetylation: the histone deacetylase enzymes. Cold Spring Harb Perspect Biol. 2014;6:a018713–a018713.
Laugesen A, Helin K. Chromatin repressive complexes in stem cells, development, and cancer. Cell Stem Cell. 2014;14:735–51.
Whyte WA, Bilodeau S, Orlando DA, Hoke HA, Frampton GM, Foster CT, et al. Enhancer decommissioning by LSD1 during embryonic stem cell differentiation. Nature. 2012;482:221–5.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-André V, Sigova AA, et al. Super-enhancers in the control of cell identity and disease. Cell. 2013;155:934–47.
Kao H-Y, Verdel A, Tsai C-C, Simon C, Juguilon H, Khochbin S. Mechanism for nucleocytoplasmic shuttling of histone deacetylase 7. J Biol Chem. 2001;276:47496–507.
Dressel U, Bailey PJ, Wang S-CM, Downes M, Evans RM, GEO Muscat. A dynamic role for HDAC7 in MEF2-mediated muscle differentiation. J Biol Chem. 2001;276:17007–13.
Xiao H, Chung J, Kao H-Y, Yang Y-C. Tip60 is a co-repressor for STAT3. J Biol Chem. 2003;278:11197–204.
Kato H, Tamamizu-Kato S, Shibasaki F. Histone deacetylase 7 associates with hypoxia-inducible factor 1?? and increases transcriptional activity. J Biol Chem. 2004;279:41966–74.
Li B, Samanta A, Song X, Iacono KT, Bembas K, Tao R, et al. FOXP3 interactions with histone acetyltransferase and class II histone deacetylases are required for repression. Proc Natl Acad Sci. 2007;104:4571–6.
Ai W, Zheng H, Yang X, Liu Y, Wang TC. Tip60 functions as a potential corepressor of KLF4 in regulation of HDC promoter activity. Nucleic Acids Res. 2007;35:6137–49.
Jensen ED, Schroeder TM, Bailey J, Gopalakrishnan R, Westendorf JJ. Histone deacetylase 7 associates with Runx2 and represses its activity during osteoblast maturation in a deacetylation-independent manner. J Bone Miner Res. 2008;23:361–72.
Malik S, Jiang S, Garee JP, Verdin E, Lee AV, O’Malley BW, et al. Histone deacetylase 7 and FoxA1 in estrogen-mediated repression of RPRM. Mol Cell Biol. 2010;30:399–412.
Fischle W, Dequiedt F, Fillion M, Hendzel MJ, Voelter W, Verdin E. Human HDAC7 histone deacetylase activity is associated with HDAC3 in Vivo. J Biol Chem. 2001;276:35826–35.
Ma C, D’Mello SR. Neuroprotection by histone deacetylase-7 (HDAC7) occurs by inhibition of c- jun expression through a deacetylase-independent mechanism. J Biol Chem. 2011;286:4819–28.
Barneda-Zahonero B, Collazo O, Azagra A, Fernández-Duran I, Serra-Musach J, Islam ABMMK, et al. The transcriptional repressor HDAC7 promotes apoptosis and c-Myc downregulation in particular types of leukemia and lymphoma. Cell Death Dis. 2015;6:e1635.
Zhu C, Chen Q, Xie Z, Ai J, Tong L, Ding J, et al. The role of histone deacetylase 7 (HDAC7) in cancer cell proliferation: regulation on c-Myc. J Mol Med. 2011;89:279–89.
Wilson AJ, Byun DS, Popova N, Murray LB, L’Italien K, Sowa Y, et al. Histone deacetylase 3 (HDAC3) and other class I HDACs regulate colon cell maturation and p21 expression and are deregulated in human colon cancer. J Biol Chem. 2006;281:13548–58.
Halkidou K, Gaughan L, Cook S, Leung HY, Neal DE, Robson CN. Upregulation and Nuclear Recruitment of HDACl in Hormone Refractory Prostate Cancer. Prostate. 2004;59:177–89.
Toh Y, Yamamoto M, Endo K, Ikeda Y, Baba H, Kohnoe S, et al. Histone H4 acetylation and histone deacetylase 1 expression in esophageal squamous cell carcinoma. Oncol Rep. 2003;10:333–8.
Yu Y, Cao F, Yu X, Zhou P, Di Q, Lei J, et al. The expression of HDAC7 in cancerous gastric tissues is positively associated with distant metastasis and poor patient prognosis. Clin Transl Oncol. 2017;19:1045–54.
Yano M, Yasuda M, Sakaki M, Nagata K, Fujino T, Arai E, et al. Association of histone deacetylase expression with histology and prognosis of ovarian cancer. Oncol Lett. 2018;15:3524–31.
Lei Y, Liu L, Zhang S, Guo S, Li X, Wang J, et al. Hdac7 promotes lung tumorigenesis by inhibiting Stat3 activation. Mol Cancer. 2017;16:170.
Gao S, Liu H, Hou S, Wu L, Yang Z, Shen J et al. MiR-489 suppresses tumor growth and invasion by targeting HDAC7 in colorectal cancer. Clin Transl Oncol. 2017. https://doi.org/10.1007/s12094-017-1770-7.
Ahn M-Y, Yoon J-H. Histone deacetylase 7 silencing induces apoptosis and autophagy in salivary mucoepidermoid carcinoma cells. J Oral Pathol Med. 2017;46:276–83.
Dokmanovic M, Perez G, Xu W, Ngo L, Clarke C, Parmigiani RB, et al. Histone deacetylase inhibitors selectively suppress expression of HDAC7. Mol Cancer Ther. 2007;6:2525–34.
Morrison BE, Majdzadeh N, Zhang X, Lyles A, Bassel-Duby R, Olson EN, et al. Neuroprotection by histone deacetylase-related protein. Mol Cell Biol. 2006;26:3550–64.
Rickels R, Shilatifard A Enhancer Logic and Mechanics in Development and Disease. Trends Cell Biol. 2018. https://doi.org/10.1016/j.tcb.2018.04.003.
Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell. 2013;153:307–19.
Fillmore CM, Kuperwasser C. Human breast cancer cell lines contain stem-like cells that self-renew, give rise to phenotypically diverse progeny and survive chemotherapy. Breast Cancer Res. 2008;10:R25.
Pasini D, Malatesta M, Jung HR, Walfridsson J, Willer A, Olsson L, et al. Characterization of an antagonistic switch between histone H3 lysine 27 methylation and acetylation in the transcriptional regulation of Polycomb group target genes. Nucleic Acids Res. 2010;38:4958–69.
Lovén J, Hoke HA, Lin CY, Lau A, Orlando DA, Vakoc CR, et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell. 2013;153:320–34.
Glaser KB, Staver MJ, Waring JF, Stender J, Ulrich RG, Davidsen SK. Gene expression profiling of multiple histone deacetylase (HDAC) inhibitors: defining a common gene set produced by HDAC inhibition in T24 and MDA carcinoma cell lines. Mol Cancer Ther. 2003;2:151–63.
Singh J, Murata K, Itahana Y, Desprez P-Y. Constitutive expression of the Id-1 promoter in human metastatic breast cancer cells is linked with the loss of NF-1/Rb/HDAC-1 transcription repressor complex. Oncogene. 2002;21:1812–22.
Aude-Garcia C, Collin-Faure V, Bausinger H, Hanau D, Rabilloud T, Lemercier C. Dual roles for MEF2A and MEF2D during human macrophage terminal differentiation and c-Jun expression. Biochem J. 2010;430:237–44.
Yang W, Shen J, Wu M, Arsura M, FitzGerald M, Suldan Z, et al. Repression of transcription of the p27 Kip1 cyclin-dependent kinase inhibitor gene by c-Myc. Oncogene. 2001;20:1688–1702.
Kaletsch A, Pinkerneil M, Hoffmann MJ, Jaguva Vasudevan AA, Wang C, Hansen FK, et al. Effects of novel HDAC inhibitors on urothelial carcinoma cells. Clin Epigenetics. 2018;10:100.
Hull EE, Montgomery MR, Leyva KJ. HDAC inhibitors as epigenetic regulators of the immune system: impacts on cancer therapy and inflammatory diseases. Biomed Res Int. 2016;2016:8797206.
Caslini C, Capo-chichi CD, Roland IH, Nicolas E, Yeung AT, Xu X-X. Histone modifications silence the GATA transcription factor genes in ovarian cancer. Oncogene. 2006;25:5446–61.
Kim YJ, Greer CB, Cecchini KR, Harris LN, Tuck DP, Kim TH. HDAC inhibitors induce transcriptional repression of high copy number genes in breast cancer through elongation blockade. Oncogene. 2013;32:2828–35.
Greer CB, Tanaka Y, Kim YJ, Xie P, Zhang MQ, Park I-H, et al. Histone deacetylases positively regulate transcription through the elongation machinery. Cell Rep. 2015;13:1444–55.
Sanchez GJ, Richmond PA, Bunker EN, Karman SS, Azofeifa J, Garnett AT, et al. Genome-wide dose-dependent inhibition of histone deacetylases studies reveal their roles in enhancer remodeling and suppression of oncogenic super-enhancers. Nucleic Acids Res. 2018;46:1756–76.
Ince TA, Richardson AL, Bell GW, Saitoh M, Godar S, Karnoub AE, et al. Transformation of different human breast epithelial cell types leads to distinct tumor phenotypes. Cancer Cell. 2007;12:160–70.
Afgan E, Baker D, Batut B, van den Beek M, Bouvier D, Čech M, et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 2018;46:W537–44.
Acknowledgements
We gratefully acknowledge the Sylvester Comprehensive Cancer Center, Onco-Genomics Shared Resource for ChIP, and RNA sequencing services. This work was funded by grants to TAI from Breast Cancer Research Foundation (NY), Florida Breast Cancer Foundation (FL), and Sylvester Comprehensive Cancer Center.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Caslini, C., Hong, S., Ban, Y.J. et al. HDAC7 regulates histone 3 lysine 27 acetylation and transcriptional activity at super-enhancer-associated genes in breast cancer stem cells. Oncogene 38, 6599–6614 (2019). https://doi.org/10.1038/s41388-019-0897-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-019-0897-0
- Springer Nature Limited
This article is cited by
-
Transcriptional regulation of cancer stem cell: regulatory factors elucidation and cancer treatment strategies
Journal of Experimental & Clinical Cancer Research (2024)
-
KLF7 regulates super-enhancer-driven IGF2BP2 overexpression to promote the progression of head and neck squamous cell carcinoma
Journal of Experimental & Clinical Cancer Research (2024)
-
Effects of super-enhancers in cancer metastasis: mechanisms and therapeutic targets
Molecular Cancer (2024)
-
Super-enhancer omics in stem cell
Molecular Cancer (2024)
-
HDAC inhibitors as a potential therapy for chemotherapy-induced neuropathic pain
Inflammopharmacology (2024)