Abstract
Long intergenic non-coding RNA 152 (LINC00152) is a recently identified tumor-promoting long non-coding RNA. However, the biological functions of LINC00152 in colorectal cancer (CRC) remain unclear and require further research. The aim of the present study is to explore the roles of LINC00152 in cellular function and its possible molecular mechanism. In this study, we discovered that LINC00152 was overexpressed in CRC tissues and negatively related to the survival time of CRC patients. Functional analyses revealed that LINC00152 could promote cell proliferation. Furthermore, LINC00152 could increase the resistance of CRC cells to 5-fluorouracil (5-FU) by suppressing apoptosis. We also discovered that LINC00152 could enhance cell migration and invasion. Mechanistic studies demonstrated that LINC00152 could regulate the expression of NOTCH1 through sponging miR-139-5p and inhibiting its activity from promoting CRC progression and development. Altogether, our work points out a novel LINC00152/miR-139-5p/NOTCH1 regulatory axis in CRC progression and development.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the third most common cancer worldwide1. The occurrence and development of CRC involve a series of complex changes at the genetic and epigenetic levels2. Increasing number of studies have demonstrated that long non-coding RNAs (lncRNAs) are involved in the occurrence and development of CRC3.
LncRNAs are a kind of RNA molecules with more than 200 nucleotides and no protein translation ability. Recent advances have revealed the vital roles of lncRNAs in regulating tumorigenesis, and progression. Long intergenic non-coding RNA 152 (LINC00152) locates on chromosome 2p11.2 with 828 nt transcription length. LINC00152 was overexpressed in tumor tissues and plasma of gastric cancer (GC) patients, and could promote GC cell proliferation and cell cycle progression through regulating EGFR and EZH24,5,6,7. LINC00152 also plays an oncogenic role in liver8, gallbladder9, and lung cancer10. In addition, LINC00152 is likely to be an indicator of stress in a variety of cells11. These studies exhibit the key oncogenic role and complicated mechanisms of LINC00152 in cancers. However, the detailed functions and mechanisms of LINC00152 in CRC are mainly unclear.
In this study, we showed that LINC00152 was upregulated in CRC, and correlated with poor survival. Functional analyses showed that LINC00152 could enhance CRC growth, metastasis, and chemoresistance. Mechanistic studies demonstrated that LINC00152 promotes tumorigenesis and progression via working as a competitive endogenous RNA (ceRNA) of miR-139-5p, which is a key tumor suppressive microRNA (miRNA)12,13,14,15,16,17,18. The present work reveals a novel regulatory pathway of LINC00152/miR-139-5p/NOTCH1 in CRC, suggesting that LINC00152 is a new prognostic factor and potential therapeutic target in CRC.
Results
Overexpression of LINC00152 in CRC associates with poor prognosis
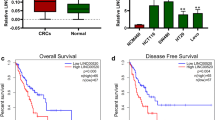
To study the role of LINC00152 in CRC, we first detected its expression in 108 paired CRC tissues and noncancerous tissues (NCTs). The results revealed that LINC00152 was obviously upregulated in CRC (P < 0.001, Fig. 1a), and 46.3% (50 of 108) of the CRC tissues showed > 2-fold upregulation of LINC00152 compared with their NCTs (Fig. 1b).
a Relative expression levels of LINC00152 in 108 paired CRC and NCTs were quantified by qRT-PCR. b LINC00152 was upregulated (> 2-fold) in 46.3% of the CRC tissues compared with the NCTs. c, d Kaplan–Meier survival analysis of the overall survival and disease-free survival in two groups defined by low and high expression of LINC00152 in patients with CRC
To assess the potential association of LINC00152 with clinicopathological features, we first divided the 108 patients into LINC00152-high and -low groups. We found that the LINC00152 levels in CRCs were significantly correlated with tumor stage (P = 0.013), whereas no obvious correlation between LINC00152 expression and other clinicopathological parameters was observed (Table 1).
The survival analysis showed that patients in the LINC00152-high group showed a shorter survival time than those in the LINC00152-low group (46.614 ± 3.366 vs. 69.338 ± 3.271 months; log rank = 9.456, P = 0.0021, Fig. 1c). In addition, high LINC00152 expression was also associated with poor disease-free survival (log rank = 4.383, P = 0.0363, Fig. 1d). Furthermore, multivariate analysis further identified that LINC00152 was an independent prognosis factor for CRC (hazard ratio (HR) = 2.514, 95% confidence interval (CI) = 1.125-5.621, P = 0.025, Table 2).
LINC00152 promotes CRC cell proliferation
The expression analyses of LINC00152 in six CRC cell lines showed that LoVo and SW480 have relatively high expressions of LINC00152, whereas HCT116 and HT29 have relatively low expressions of LINC00152 (Fig. 2a). To investigate the biological functions of LINC00152 in CRC, we overexpressed LINC00152 in HCT116 and HT29 cells, and inhibited LINC00152 expression in LoVo and SW480 cells (Fig. 2b). We observed that LINC00152 overexpression significantly promoted CRC cell proliferation and colony formation. In contrast, decreased cell growth, and colony formation abilities were showed in LINC00152-silenced cells (Fig. 2c–e). Furthermore, ectopic LINC00152 expression promoted CRC tumor growth in vivo (Fig. 2f). All these data reveal the growth-stimulating functions of LINC00152 in CRC.
a Relative expression of LINC00152 in CRC cell lines. b Validation of overexpression and knockdown efficacy of LINC00152 in CRC cell lines by qRT-PCR. c, d Effects of LINC00152 overexpression and downregulation on CRC cell proliferation were measured by a CCK-8 assay. e Effects of LINC00152 overexpression and knockdown on colony formation in CRC cells. f LINC00152 overexpression promoted CRC tumorigenesis in a xenograft mouse model. *P < 0.05; **P < 0.01
LINC00152 promotes cell cycle progression and confers resistance to 5-FU-induced apoptosis
To investigate the mechanism mediating the growth-promoting functions of LINC00152 in CRC, we measured the cell cycle distribution in the LINC00152-overexpressed and silenced CRC cells. As shown in Fig. 3a, ectopic LINC00152 expression resulted in an increased number of cells in S phase, whereas LINC00152 knockdown caused a decreased cell number in S phase, indicating the promotion of the cell cycle by LINC00152.
a Cell cycle analyses were performed in HCT116 cells transfected with pWPXL-LINC00152 and pWPXL, or LoVo cells transfected with si-LINC00152 and si-NC. b LINC00152 decreased the sensitivity of CRC cells to 5-FU. The IC50 of LINC00152-overexpressed HCT116 cells was significantly higher than that of the control (0.836 vs. 0.279 μg/ml), and the IC50 of LINC00152-silenced LoVo cells was lower than that the control (0.576 vs. 0.960 μg/ml). c Cell apoptosis analyses were performed in cell lines with LINC00152 overexpression or knockdown. *P < 0.05; **P < 0.01
5-fluorouracil (5-FU) is a basic drug for CRC treatment, and we evaluated the effect of LINC00152 on 5-FU sensitivity in CRC cells. After overexpression or knockdown of LINC00152, CRC cells were then assayed for their sensitivity to 5-FU by a CCK-8 assay. The results showed that ectopic LINC00152 expression decreased the sensitivity of HCT116 cells to 5-FU, whereas LINC00152 silencing increased the sensitivity to 5-FU in LoVo cells (Fig. 3b). Given the key role of apoptosis in cancer chemotherapy, we further measured the effect of LINC00152 on 5-FU-induced apoptosis. The results showed that the LINC00152 overexpression significantly antagonize 5-FU-induced apoptosis, whereas the LINC00152 knockdown could augment apoptosis caused by 5-FU (Fig. 3c).
LINC00152 promotes CRC cell migration and invasion
Transwell assays were then performed to measure the impact of LINC00152 on CRC metastasis. We observed that ectopic LINC00152 expression significantly facilitated migration and invasion in HCT116 cells (Fig. 4a), whereas the LINC00152 knockdown suppressed migration and invasion in LoVo cells (Fig. 4b).
LINC00152 sponges miR-139-5p
To investigate underlying mechanisms of LINC00152 in CRC, we first measured the subcellular localization of LINC00152 in HCT116 cells, and revealed that LINC00152 was localized predominantly in the cell cytoplasm (Fig. 5a), suggesting that LINC00152 may regulate tumorigenesis at the post-transcriptional level. LncRNAs could act as molecular sponges to modulate mRNAs expression by competitively binding their common miRNA responsive elements (MREs). Previous studies have proved that LINC00152 could function as a ceRNA in human cancers19,20,21. We hypothesized that LINC00152 could promote CRC tumorigenesis and progression by suppressing the functions of certain miRNAs. Based on the bioinformatics analysis and Xia’s work22, we found that LINC00152 harbors a recognition sequence of miR-139-5p (Fig. 5b). In view of the opposite functions of miR-139-5p and LINC00152 in CRC12,13,14,15,16,17,18, we intended to explore the potential relationship between them in CRC.
a Subcellular localization of LINC00152 was determined by qRT-PCR in HCT116 cell line. b miR-139-5p-binding sequence in LINC00152 and NOTCH1 3'UTR. A mutation was generated in LINC00152 in the complementary site for miR-139-5p binding. c Luciferase activity of a luciferase reporter plasmid (pLuc) containing wild-type or mutant LINC00152 co-transfected with miR-139-5p was determined using the dual luciferase assay. d Cellular lysates from HCT116 cells were used for RIP with an anti-Ago2 antibody or IgG antibody. The levels of LINC00152 and miR-139-5p were detected by qRT-PCR. e MiR-139-5p and pLuc plasmid containing NOTCH1 3'UTRs were co-transfected with pWPXL-LINC00152 or empty vector into 293T cells to verify whether LINC00152 can function as a ceRNA of miR-139-5p. f The expression levels of NOTCH1 in HCT116 cells transfected with pWPXL-LINC00152 and LoVo cells transfected with si-LINC00152 were analyzed by qRT-PCR and western blot. g Correlation analysis between NOTCH1 and LINC00152 expression. *P < 0.05; **P < 0.01
We first constructed reporter vectors containing LINC00152 (pLuc-LINC00152-WT) or its mutant with mutations in the seed sequence of miR-139-5p (pLuc-LINC00152-Mut), and then evaluated this underlying correlation of miR-139-5p with LINC00152 using luciferase reporter assays. We observed that miR-139-5p overexpression led to a marked inhibition in the reporter activity of pLuc-LINC00152-WT compared with that of pLuc-LINC00152-Mut (Fig. 5c), suggesting sequence-specific binding and inhibition of LINC00152 by miR-139-5p. To further validate the potential binding of LINC00152 to miR-139-5p, an RNA Immunoprecipitation (RIP) assay using an anti-Ago2 antibody was performed. The data exhibited that both LINC00152 and miR-139-5p were obviously enriched in Ago2 complex, demonstrating that LINC00152 is included in miRNPs, probably through binding with miR-139-5p (Fig. 5d).
LINC00152 modulates NOTCH1 expression by competitively binding miR-139-5p
Previous studies have shown that miR-139-5p inhibit CRC tumorigenesis, development, and chemoresistance by regulating NOTCH112,13,14,15. To ascertain whether the above-observed effects depend on the regulation of LINC00152 on the miR-139-5p/NOTCH1 pathway, we first evaluated the relationship among LINC00152, miR-139-5p and NOTCH1 using luciferase assays. As a result, the overexpression of LINC00152, but not the vector control, blocked the inhibitory effect of miR-139-5p on the relative luciferase expression of pLuc-NOTCH1-3′UTR (Fig. 5e). These results confirmed that LINC00152 abolishes the miR-139-5p-mediated repressive activity on NOTCH1 by competitively binding miR-139-5p. In addition, LINC00152 knockdown significantly reduced the endogenous NOTCH1 expression in CRC cells (Fig. 5f). In contrast, NOTCH1 expression was obviously increased in LINC00152 overexpressing CRC cells (Fig. 5f). A positive relationship was also observed between the levels of NOTCH1 and LINC00152 in CRC tissues (Fig. 5g). These data demonstrate that LINC00152 can regulate NOTCH1 activity by sponging miR-139-5p both in CRC cell lines and clinical CRC tumors.
LINC00152 exerts tumor-promoting function in CRC by regulating the miR-139-5p/NOTCH1 axis
Both miR-139-5p and NOTCH1 could regulate cell growth, apoptosis, and invasion in CRC12,13,14,15,16,17,18, 22. To investigate whether LINC00152 exerts tumor-promoting functions in CRC by modulating the miR-139-5p/NOTCH1 axis, we first checked the effects of miR-139-5p and NOTCH1 on LINC00152-induced cell proliferation, and observed that miR-139-5p overexpression or NOTCH1 knockdown blocked the LINC00152-induced CRC cell growth (Fig. 6a). We then evaluated the effects of miR-139-5p/NOTCH1 signaling on the LINC00152-induced 5-FU resistance in CRC cells. As shown in Fig. 6b, ectopic miR-139-5p expression or NOTCH1 knockdown significantly reversed the LINC00152-induced 5-FU resistance and counteracted the apoptosis-inhibiting effects of LINC00152 in CRC cells (Fig. 6c). In addition, the increased cell mobility in LINC00152 overexpressing CRC cells was also reversed by miR-139-5p overexpression or NOTCH1 knockdown (Fig. 6d). Altogether, these data demonstrate that LINC00152 exerts tumor-promoting functions in CRC, at least partly, through sponging miR-139-5p and then regulating NOTCH1.
a The increased cell viability in pWPXL-LINC00152 transfected CRC cells was abolished by ectopic miR-139-5p expression or NOTCH1 knockdown. The cell viability was measured by a CCK-8 assay. b Increased 5-FU resistance in pWPXL-LINC00152 transfected CRC cells was abolished by ectopic miR-139-5p expression or NOTCH1 knockdown. The IC50s for group (1) to (8) were 0.632, 1.180, 0.327, 0.512, 0.564, 1.014, 0.329, and 0.466 μg/ml, respectively. c Overexpression of LINC00152 decreased 5-FU-induced apoptosis, which was partly blocked by ectopic miR-139-5p expression or NOTCH1 knockdown. d Ectopic miR-139-5p expression or NOTCH1 knockdown could partly block LINC00152-induced cell migration. * or #P < 0.05; **or ##or &&P < 0.01 (* or **: (1) vs. (2) or (5) vs. (6); # or ##: (1) vs. (3) or (5) vs. (7); &&: (1) vs. (4))
Discussion
In this study, we observed that LINC00152 expression is obviously increased in clinical CRC tissues, and is correlated with tumor stage and poor patient survival. Functionally, we revealed that LINC00152 promotes CRC growth, metastasis, and induces 5-FU resistance. Moreover, we further demonstrated that LINC00152 executes tumor-promoting functions by sponging miR-139-5p and then modulating NOTCH1 in CRC.
Numerous studies have revealed varied regulatory roles of lncRNAs in human diseases, especially in tumorigenesis and development23. For example, our previous work revealed that UCA1 could promote cell proliferation and 5-FU chemoresistance in CRC via competitively inhibiting miR-204-5p24. LINC00152 is recently identified cancer-related lncRNA that play oncogenic roles in several kinds of human cancers, especially in digestive tract tumors4,5,6,7,8,9, 11. Yue et al.19 reported that LINC00152 expression is increased in CRC. Interestingly, in contradictory to their conclusions, a recently published work demonstrated that LINC00152 is downregulated in CRC, inhibits viability and promotes apoptosis of CRC cells25. Here, we demonstrated that LINC00152 expression was obviously increased in CRC and correlated with patient’s survival, which was also observed by Yue et al.19. Our detailed functional studies revealed the promoting effects of LINC00152 on CRC growth and metastasis, which is coincident with the oncogenic role of LINC00152 in GC4,5,6,7, liver cancer8, gallbladder cancer9, 26, and clear cell renal cell carcinoma27. In addition, we also showed that LINC00152 confers resistance to 5-FU-induced apoptosis, which was similar to that reported by Yue et al.19. In their study, Yue et al. demonstrated that LINC00152 works as a ceRNA of miR-193a-3p to induce oxaliplatin resistance. These data demonstrate that LINC00152 is a key lncRNA with extensive tumor-promoting functions in human cancers.
Several studies have reported that LINC00152 promotes tumor development and progression by regulating several key tumor-related pathways, including EGFR, mTOR, and PI3K/AKT signaling4, 8, 9. Recent studies revealed a new mechanism of lncRNA by acting as ceRNA20. In this situation, lncRNAs can block the repression of miRNA on its target genes by competitively binding their common MREs28. LINC00152 could bind several miRNAs in cancer cells, including miR-138, miR-376c-3p, and miR-193a-3p19, 25, 26, suggesting that ceRNA is a key mechanism by which LINC00152 regulates tumorigenesis and development.
Due to the upregulation and tumor-promoting role of LINC00152 in CRC, it is reasonably concluded that LINC00152 promotes CRC development and progression by inhibiting tumor suppressive miRNAs. Based on previous works by us and other researchers, miR-139-5p levels are markedly reduced in CRC, and has exact opposite functions to those of LINC0015212,13,14,15,16,17,18. MiR-139-5p can repress CRC growth, metastasis, and chemoresistance by regulating several genes, such as NOTCH1, BCL2, and AMFR12,13,14,15,16,17,18. MiR-139-5p was reported to play a suppressive role in other cancers, including gastric, breast, and hepatocellular carcinoma29. As a key member of the NOTCH family, NOTCH1 is frequently upregulated in human cancers, including CRC30. Previous researches have proved that miR-139-5p can regulate CRC growth, metastasis, stemness, and chemoresistance via targeting NOTCH112,13,14,15, 31. We speculated that LINC00152 exerts its functions by regulating the miR-139-5p/NOTCH1 pathway. As expected, both the luciferase and RIP assays confirmed the binding of LINC00152 to miR-139-5p. Subsequent functional and mechanistic assays proved that LINC00152 regulates CRC development, progression, and drug resistance by competitively sponging miR-139-5p and then restoring NOTCH1 activity.
In summary, our work shows that LINC00152 is upregulated in CRC, correlated with patients’ survival and appears to be a potential biomarker for predicting chemoresistance. LINC00152 contributes to the tumorigenesis, progression, and chemoresistance of CRC by inhibiting miR-139-5p, uncovering a novel ceRNA network of LINC00152/miR-139-5p/NOTCH1 in CRC cells. These data suggest that targeting LINC00152 may be a promising therapeutic strategy for CRC.
Materials and methods
Clinical samples
A total of 108 paired human CRC tissues and NCTs were collected with informed consent at Affiliated Hospital of Jiangnan University, and the detailed patient information are shown in Table 1. This study was carried out under the permission of the Clinical Research Ethics Committees of Affiliated Hospital of Jiangnan University.
Cell lines
HEK-293T and six CRC cell lines (HCT8, HT29, LoVo, HCT116, SW480, and SW620) were obtained from the American Type Culture Collection. These cells were maintained in Dulbecco's modified Eagle's medium supplemented with 10% fetal bovine serum (Gibco, USA) and have been recently authenticated.
RNA isolation and quantitative reverse transcription (RT)-PCR assays
Total RNA was isolated with RNAiso Plus (Takara, Japan). Cytoplasmic and nuclear RNA was purified using PARISTM Kit (Ambion, USA). Complimentary DNA was synthesized using the HiFiScript 1st Strand cDNA Synthesis Kit (CWBIO, China). Real time RT-PCR was performed using an UltraSYBR Mixture (CWBIO).
Vector construction and siRNA
LINC00152 was synthesized at GENEray Biotechnology (China) and was inserted into the lentivirus vector pWPXL. The fragment of LINC00152 with miR-139-5p-binding site and the NOTCH1 3'UTR were cloned into pLuc. The LINC00152 with the mutated seed sequence of miR-139-5p was constructed by an overlap extension PCR32. The primers used in vector construction are shown in Supplementary Table 1. The siRNAs of LINC00152 and NOTCH1 were purchased from GenePharma (China).
Generation of cell lines with stable overexpression of LINC00152
HEK-293T cells were transfected with pWPXL-LINC00152 (or pWPXL), pMD2G, and ps-PAX2 plasmids using Lipofectamine 2000 (Invitrogen, USA). These virus particles were centrifuged and filtered to infect HCT116 and HT29 cells to generate corresponding stable cells.
Cell proliferation and colony formation assays
Cell Counting Kit 8 (CCK-8, Beyotime, China) was used to measure cell viability. A colony formation assay was performed as we previously described33.
Cell cycle and apoptosis analyses
The cell cycle and apoptosis analyses of LINC00152-overexpressed and silenced CRC cells were applied using the Cell Cycle and Apoptosis Detection Kit purchased from CWBIO.
Cell migration and invasion assay
Transwell assays were performed to measure cell migration and invasion using Boyden chambers (8-mm pore size, BD Biosciences) as we previously described33.
Xenograft tumor assay
Twenty-four male athymic nude BALB/c mice at 5 weeks of age were randomly divided into four groups, and the number of mice is determined according to prior experience of our laboratory. HCT116 cells stably expressing LINC00152 or the bank vector were subcutaneously injected into flank of nude mouse. Four (HCT116) or six weeks (HT29) after injection, these mice were sacrificed to measure the growth of subcutaneous tumors. The investigator was blinded to group allocation during the experiments. All animal experiments were approved by the Clinical Research Ethics Committees of our Hospital.
Luciferase reporter assay
HEK-293T cells were co-transfected with pLuc, pRL-CMV, miR-139-5p mimics (negative control, NC), and pWPXL-LINC00152 (pWPXL). These cells were then assayed for luciferase activity using the Dual-Luciferase® Reporter Assay System (Beyotime, China).
RNA Immunoprecipitation (RIP) assay
A RIP assay was performed using the EZ-Magna RIP Kit (Millipore, USA) as we previously described24.
Western blotting
Total protein was separated by 8% (or 10%) sodium dodecyl sulfate polyacrylamide gel electrophoresis and transferred to a PVDF membrane. After blocking with non-fat milk, the polyvinylidene difluoride membrane was incubated with a rabbit anti-human NOTCH1 antibody (1:1000, 20687-1-AP, Proteintech, USA) or a mouse anti-β-actin antibody (1:1000, AA128, Beyotime, China).
Statistical analyses
Data were presented as the mean ± s.d. Student’s t-test, the Mann–Whitney U-test and the χ2 test were performed to analyze the differences among different groups. The differences in survival rates were determined by the Kaplan–Meier method and compared by the log-rank test. HRs and 95% CIs were calculated by a Cox proportional hazards model. P-values < 0.05 were considered statistically significant.
Change history
16 August 2018
This article was originally published under Nature Research's License to Publish, but now has been made available under a CC BY 4.0 license. The PDF and HTML versions of the Article have been modified accordingly.
References
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J. Clin. 65, 87–108 (2015).
Migliore, L., Migheli, F., Spisni, R. & Coppede, F. Genetics, cytogenetics, and epigenetics of colorectal cancer. J. Biomed. Biotechnol. 2011, 792362 (2011).
Xie, X. et al. Long non-coding RNAs in colorectal cancer. Oncotarget 7, 5226–5239 (2016).
Zhou, J. et al. Linc00152 promotes proliferation in gastric cancer through the EGFR-dependent pathway. J. Exp. Clin. Cancer Res. 34, 135 (2015).
Chen W. M. et al. Long intergenic non-coding RNA 00152 promotes tumor cell cycle progression by binding to EZH2 and repressing p15 and p21 in gastric cancer. Oncotarget 7, 9773–9787 (2016).
Pang, Q. et al. Increased expression of long intergenic non-coding RNA LINC00152 in gastric cancer and its clinical significance. Tumour Biol. 35, 5441–5447 (2014).
Li, Q. et al. Plasma long noncoding RNA protected by exosomes as a potential stable biomarker for gastric cancer. Tumour Biol. 36, 2007–2012 (2015).
Ji, J. et al. LINC00152 promotes proliferation in hepatocellular carcinoma by targeting EpCAM via the mTOR signaling pathway. Oncotarget 6, 42813–42824 (2015).
Cai, Q. et al. Upregulation of long non-coding RNA LINC00152 by SP1 contributes to gallbladder cancer cell growth and tumor metastasis via PI3K/AKT pathway. Am. J. Transl. Res. 8, 4068–4081 (2016).
Chen, Q. N. et al. Long intergenic non-coding RNA 00152 promotes lung adenocarcinoma proliferation via interacting with EZH2 and repressing IL24 expression. Mol. Cancer 16, 17 (2017).
Tani, H. & Torimura, M. Identification of short-lived long non-coding RNAs as surrogate indicators for chemical stress response. Biochem. Biophys. Res. Commun. 439, 547–551 (2013).
Song, M. et al. MiR-139-5p inhibits migration and invasion of colorectal cancer by downregulating AMFR and NOTCH1. Protein Cell 5, 851–861 (2014).
Zhang, L. et al. microRNA-139-5p exerts tumor suppressor function by targeting NOTCH1 in colorectal cancer. Mol. Cancer 13, 124 (2014).
Liu, H. et al. miR-139-5p sensitizes colorectal cancer cells to 5-fluorouracil by targeting NOTCH-1. Pathol. Res. Pract. 212, 643–649 (2016).
Xu, K. et al. MiR-139-5p reverses CD44+/CD133+-associated multidrug resistance by downregulating NOTCH1 in colorectal carcinoma cells. Oncotarget 7, 75118–75129 (2016).
Li, Q. et al. miR-139-5p inhibits the epithelial-mesenchymal transition and enhances the chemotherapeutic sensitivity of colorectal cancer cells by downregulating BCL2. Sci. Rep. 6, 27157 (2016).
Shen, K. et al. MiR-139 inhibits invasion and metastasis of colorectal cancer by targeting the type I insulin-like growth factor receptor. Biochem. Pharmacol. 84, 320–330 (2012).
Zou, F. et al. Targeted deletion of miR-139-5p activates MAPK, NF-kappaB and STAT3 signaling and promotes intestinal inflammation and colorectal cancer. FEBS J. 283, 1438–1452 (2016).
Yue, B., Cai, D., Liu, C., Fang, C. & Yan, D. Linc00152 functions as a competing endogenous rna to confer oxaliplatin resistance and holds prognostic values in colon cancer. Mol. Ther. 24, 2064–2077 (2016).
Yang, C. et al. Competing endogenous RNA networks in human cancer: hypothesis, validation, and perspectives. Oncotarget 7, 13479–13490 (2016).
Xia, T. et al. Long noncoding RNA associated-competing endogenous RNAs in gastric cancer. Sci. Rep. 4, 6088 (2014).
Fender, A. W., Nutter, J. M., Fitzgerald, T. L., Bertrand, F. E. & Sigounas, G. Notch-1 promotes stemness and epithelial to mesenchymal transition in colorectal cancer. J. Cell Biochem. 116, 2517–2527 (2015).
Wang, K. C. & Chang, H. Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 43, 904–914 (2011).
Bian, Z. et al. LncRNA-UCA1 enhances cell proliferation and 5-fluorouracil resistance in colorectal cancer by inhibiting miR-204-5p. Sci. Rep. 6, 23892 (2016).
Zhang, Y. H., Fu, J., Zhang, Z. J., Ge, C. C. & Yi, Y. LncRNA-LINC00152 down-regulated by miR-376c-3p restricts viability and promotes apoptosis of colorectal cancer cells. Am. J. Transl. Res. 8, 5286–5297 (2016).
Cai, Q. et al. Long non-coding RNA LINC00152 promotes gallbladder cancer metastasis and epithelial-mesenchymal transition by regulating HIF-1alpha via miR-138. Open Biol. 7, 160247 (2017).
Wu, Y. et al. Long non-coding RNA Linc00152 is a positive prognostic factor for and demonstrates malignant biological behavior in clear cell renal cell carcinoma. Am. J. Cancer Res. 6, 285–299 (2016).
Rashid, F., Shah, A. & Shan, G. Long non-coding rnas in the cytoplasm. Genomics Proteomics Bioinformatics 14, 73–80 (2016).
Zhang, H. D., Jiang, L. H., Sun, D. W., Li, J. & Tang, J. H. MiR-139-5p: promising biomarker for cancer. Tumour Biol. 36, 1355–1365 (2015).
Chu, D. et al. High level of Notch1 protein is associated with poor overall survival in colorectal cancer. Ann. Surg. Oncol. 17, 1337–1342 (2010).
Vinson, K. E., George, D. C., Fender, A. W., Bertrand, F. E. & Sigounas, G. The Notch pathway in colorectal cancer. Int. J. Cancer 138, 1835–1842 (2016).
Huang, Z. et al. MicroRNA-95 promotes cell proliferation and targets sorting Nexin 1 in human colorectal carcinoma. Cancer Res. 71, 2582–2589 (2011).
Yin, Y. et al. miR-204-5p inhibits proliferation and invasion and enhances chemotherapeutic sensitivity of colorectal cancer cells by downregulating RAB22A. Clin. Cancer Res. 20, 6187–6199 (2014).
Acknowledgements
The study was supported by grants from National Natural Science Foundation of China (81672328, 81602033, 81272299, and 81301784), Natural Science Foundation of Jiangsu Province (BK20150004 and BK20151108), Medical Key Professionals Program of Jiangsu Province, Fundamental Research Funds for the Central Universities (JUSRP51710A and NOJUSRP51619B), Medical Innovation Team Program of Wuxi, and Hospital Management Center of Wuxi (YGZXZ1401 and YGZXM1524).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bian, Z., Zhang, J., Li, M. et al. Long non-coding RNA LINC00152 promotes cell proliferation, metastasis, and confers 5-FU resistance in colorectal cancer by inhibiting miR-139-5p. Oncogenesis 6, 395 (2017). https://doi.org/10.1038/s41389-017-0008-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-017-0008-4
- Springer Nature Limited
This article is cited by
-
LINC01852 inhibits the tumorigenesis and chemoresistance in colorectal cancer by suppressing SRSF5-mediated alternative splicing of PKM
Molecular Cancer (2024)
-
Targeting LINC00152 activates cAMP/Ca2+/ferroptosis axis and overcomes tamoxifen resistance in ER+ breast cancer
Cell Death & Disease (2024)
-
Interaction of lncRNAs with mTOR in colorectal cancer: a systematic review
BMC Cancer (2023)
-
Integrative Analysis Revealed LINC00847 as a Potential Target of Tumor Immunotherapy
Applied Biochemistry and Biotechnology (2023)
-
SLCO4A1-AS1 promotes colorectal tumourigenesis by regulating Cdk2/c-Myc signalling
Journal of Biomedical Science (2022)