Abstract
Background
The aim of the study was to characterize molecular diagnoses in patients with childhood-onset progressive neurological disorders of suspected genetic etiology.
Methods
We studied 48 probands (age range from newborn to 17 years old) with progressive neurological disorders of unknown etiology from the largest pediatric neurology clinic in Finland. Phenotypes included encephalopathy (54%), neuromuscular disorders (33%), movement disorders (11%), and one patient (2%) with hemiplegic migraine. All patients underwent whole-exome sequencing and disease-causing genes were analyzed.
Results
We found 20 (42%) of the patients to have variants in genes previously associated with disease. Of these, 12 were previously reported disease-causing variants, whereas eight patients had a novel variant on a disease-causing gene: ATP7A, CHD2, PURA, PYCR2, SLC1A4, SPAST, TRIT1, and UPF3B. Genetics also enabled us to define atypical clinical presentations of Rett syndrome (MECP2) and Menkes disease (ATP7A). Except for one deletion, all findings were single-nucleotide variants (missense 72%, truncating 22%, splice-site 6%). Nearly half of the variants were de novo.
Conclusions
The most common cause of childhood encephalopathies are de novo variants. Whole-exome sequencing, even singleton, proved to be an efficient tool to gain specific diagnoses and in finding de novo variants in a clinically heterogeneous group of childhood encephalopathies.
Impact
-
Whole-exome sequencing is useful in heterogeneous pediatric neurology cohorts.
-
Our article provides further evidence for and novel variants in several genes.
-
De novo variants are an important cause of childhood encephalopathies.
Similar content being viewed by others
Introduction
Pediatric neurological disorders are characterized with variable phenotypes, severity of symptoms, age at onset, and disease progression. Compared to adults, the developing nervous system and growth of the child sometimes mask classical neurological disease signs and symptoms are often unspecific, such as hypotonia.1 Furthermore, one gene can cause several different phenotypes, challenging the diagnostic process.2 Next-generation sequencing methods have greatly improved diagnostic accuracy in these heterogeneous disorders.3 Especially whole-exome sequencing (WES), which focuses on protein encoding regions of the genome, has been widely applied to pediatric patient cohorts4 and has become part of clinical diagnostics. Yet the success, often measured as diagnostic yield of WES, varies in different studies depending on the patient cohort phenotype and selection criteria. According to a recent meta-analysis, the average diagnostic yield of WES has been around 40% in all studies, with pediatric cohorts and studies with acute illness or neurological disorder as the test indication improving the yield.3 Achieving a molecular diagnosis has several benefits to the patient and their families: the possibility of a specific therapy, aid to family planning, and the conclusion to the diagnostic odyssey many patients must endure.
The aim of this study was to unravel the genetic etiologies of children with childhood-onset, unknown progressive neurological disorders in the largest pediatric neurology clinic in Finland. Our clinically mixed cohort of 48 children underwent WES. We found that WES, also for analyzing single probands and de novo variants, is a useful and effective diagnostic tool in this type of pediatric population, facilitating diagnosis of atypical presentations of classical syndromes.
Materials and methods
Ethics
The Ethics Review Board of the Hospital District of Helsinki and Uusimaa granted ethical approval for the study. Informed written consents were gathered from parents of child participants in accordance with the Declaration of Helsinki.
Study population
The procedure for recruiting patients and the study population have been previously described in Aaltio et al.5 To summarize, patients with a severe neurological disorder of infancy or childhood-onset progressive neurological disorder were recruited ongoingly, with special attempt to enroll them already early in their diagnostic investigations, by experienced pediatric neurologists in the Children’s Hospital of Helsinki University Hospital, a tertiary care center unit, during 2015–2018. Patients with a known genetic disorder, clinically recognizable disorder, non-progressive intellectual disability, or autistic spectrum disorder were not considered. A total of 48 probands were recruited from 47 families, including one sister-pair. Of the families, 90% were of Finnish, 8% African, and 2% of other European origin. In one family, the parents were first-degree cousins. Of the probands 62% were male with a median age at inclusion of 5.7 years (range 0–17 years). Patient phenotypes included encephalopathy (54%), neuromuscular disorders (33%), and movement disorders (11%), while one patient had a varying phenotype: severe hemiplegic migraine. Beyond the study subjects, four families had more than one affected individual. Table 1 compiles the study population demographics.
Whole-exome sequencing and variant analysis
Exome sequencing was performed to 48 probands following the methods as described in Sainio et al.6 The variant calling pipeline of the Finnish Institute of Molecular Medicine (FIMM) was used for the reference genome alignment and variant calling.7 Variants were analyzed primarily by suspected mode of inheritance, cross-referencing against variant databases8,9 and by bioinformatical prediction tools: Combined Annotation Dependent Depletion (CADD) C-score,10 PolyPhen,11 and SIFT.12 Variants with allele frequencies lower than 1% were prioritized. Pathogenicity of novel sequence variants was assessed according to ACMG criteria.13 Every patient had a phenotype highly specific for a disease of genetic etiology, and all reported novel sequence variants had computational evidence of deleteriousness and resided on highly conserved amino acid residues. We reported only disease-causing variants, not variants of uncertain significance or variants that could not be confirmed as disease-causing.
Confirmation of findings
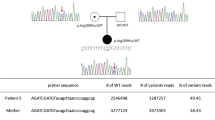
Most of the findings were confirmed by us with another independent method, Sanger sequencing, while all of them were confirmed by a clinical laboratory certified in genetic testing. In novel findings, Sanger sequencing was used to define variant segregation in the families. Paternity and maternity were confirmed by DNA fingerprinting to validate variant de novo status. For one family (P16 and P17), solid-phase minisequencing, a quantitative single-nucleotide detection assay,14 was used to confirm mosaicism in the family. Minisequencing is a method that permits quantifying the share of a single nucleotide variant in samples. The primers used for Sanger sequencing, minisequencing and parental testing, are reported in Supplementary Information.
Results
Genetic diagnoses in cohort
A total of 20 (42%) patients received a definitive molecular diagnosis by WES. Of the phenotypic groups, half of the patients with encephalopathy (n = 13) were diagnosed, while the proportions were smaller in the other groups: a third of patients with neuromuscular disorders (n = 5), and one out of five in the movement disorder group. Most findings indicated a dominant disorder (13/20) while autosomal recessive (AR) and X-linked recessive (XLR) disorders were only found in the encephalopathy group, five of the former and two the latter. Of the dominantly inherited disorders, the only sister-pair (P16 and P17) included in the study was diagnosed with a previously reported autosomal dominant (AD) missense variant in DNM2, causing Charcot–Marie–Tooth disease.15 The parents of the sisters were unaffected, initially eluding the inheritance pattern: Sanger sequencing was inconclusive, however pointing towards the mother being an unaffected carrier of the allele. Minisequencing performed on the mother’s blood, saliva, and urine revealed that the mother is a mosaic carrier, with a mutation load of 20–32% (see Supplementary Fig. 1 summarizing the findings of the family). A third patient had inherited a GDAP1 variant from his asymptomatic mother; however, there were five other maternal family members that were affected, carried the variant, and received a molecular diagnosis for their disease. The fourth inherited AD variant was in ATP1A2 causing childhood-onset hemiplegic migraine, affecting also the proband’s sister, father, and two other relatives. All the remaining inherited AD variants were de novo (7/11), including two unrelated patients with variants in the PURA gene (the patients have also been described in Reijnders et al.16). In addition, two X-linked dominant (XLD) variants were de novo, raising the total share of de novo mutations to 47%. One of the patients, P10, a girl of 2 years who had undergone extensive investigations for undetermined encephalopathy, with symptoms of vomiting, growth retardation, developmental delay, and lactic acidosis, had an XLD variant in MECP2 causing Rett syndrome. Of the maternally inherited XLR variants, patient P04 with UPF3B variant had an affected brother with the same genetic finding, while the hemizygote ATP7A variant of patient P03 was absent in other male family members. Of the five AR disorders, two of the children were of African descent, the other with consanguineous parents, and one child manifested with a Finnish disorder, the PEHO syndrome17 caused by ZNHIT3. Table 2 compiles all patients with genetic diagnoses and the Mendelian Inheritance of Man (MIM) names of disorders.
Novel findings in known disease-causing genes
Novel variant and possible Finnish founder allele in TRIT1
The proband, P02, was born at term uneventfully, with normal growth and no family history of hereditary disorders. At 4 months of age, she presented initially with vertical nystagmus, which later receded. At 9 months age, she suffered of seizures and was diagnosed with focal epilepsy, initially difficult to treat but later well in control with antiepileptic medication. Brain MRI at 4 and 9 months of age showed thin corpus callosum and white matter abnormality. She also had microcephaly, with her head circumference being −3 SD at 1 year of age, and significant spasticity in legs. She learned to crawl but lost that skill. At 7 years of age, she does not move independently or talk, but uses communication aids. WES revealed two variants on TRIT1 (NM_017646.6), c.70 C > T (p.Pro24Ser) and c.979 C > T (p.Arg327*). The gene, tRNA isopentenyltransferase 1, is associated with combined oxidative phosphorylation deficiency 35 (COXPD35, MIM#617873).18 We collected previously reported disease-causing variants of TRIT1 (Fig. 1a), and found that p.R327X has been previously reported as pathogenic by two independent studies,19,20 while p.P24S is novel, located on the mitochondrial transit peptide sequence, and conserved in species down to arthropods (Fig. 1b). The findings were confirmed by Sanger sequencing to have been inherited as compound heterozygous from the mother and father, respectively (Fig. 1c, d). The allele frequencies of disease associated TRIT1 variants (Fig. 1e) showed that p.R327X is the most common variant, most common in Finns with an allele frequency of 0.18% (globally 0.05%). The p.P24S variant is not found in genomic databases. Histological analysis of skeletal muscle sample showed normal morphology, but reduced activity of respiratory chain complex IV and low amount of complex I, fitting to a mitochondrial translation defect. The novel p.P24S replaces a hydrophobic proline with a polar serine in the mitochondrial targeting sequence, suggesting reduced entry of the mutant protein to mitochondria. Since the patient’s phenotype corresponds to previous descriptions of TRIT1 caused COXPD35 disease, the variant p.P24S was found in trans with a known pathogenic variant, and it affects a conserved amino acid altering the protein’s hydrophobicity, p.P24S was judged to be likely pathogenic (ACMG criteria and computational evidence of pathogenicity in Table 3).
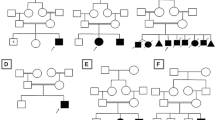
a TRIT1 protein with domains according to Uniprot, and previously reported disease-causing variants, with exons indicated below the linear graph. The vertical axis depicts the number of patients found in literature, including the variants found in this study. Our novel variant p.P24S resides on the mitochondrial transit peptide, where one previous variant has been described. References for the gene review can be found in supplementary information. b Protein conservation through species of amino acid residue Pro24 (yellow shade) and bordering regions. c Family pedigree, both parents were unaffected carriers of different variants, the affected proband’s genotype compound heterozygote. d Sanger sequencing results of family, the first row depicting the maternally inherited c.70 C > T variant, the second row the paternally inherited c.979 C > T. e Of the previously reported 16 pathogenic variants, 11 are found in gnomAD as rare variants. The bar graph depicts the allele frequency of those five variants with total allele frequency greater than 0.005%, in different genetic subpopulations (bars), in order of total frequency (dotted line).
SLC1A4 caused spastic tetraparesis, thin corpus callosum, and progressive microcephaly
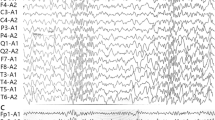
The proband, P01, was the first child of the family, born with microcephaly −2.45 SD, but otherwise healthy. At 7 months of age, he began having tonic seizures. EEG showed multifocal spikes, increasing during sleep. Brain MRI at 8 months showed hypomyelination, a thin corpus callosum, gray matter abnormality, and mildly enlarged ventricles. He had spastic tetraparesis, more prominently in the upper limbs and shoulder region. At the age of 1 year and 4 months, his microcephaly was remarkable, with his head circumference being −7.7 SD. He had cerebral visual impairment, did not appear to take any contact during clinical examination and could not move independently. WES revealed a novel homozygote SLC1A4 (NM_003038.5) missense variant c.1421 T > C (p.Leu474Pro). SLC1A4 is an L-serine transporter associated with spastic tetraparesis, thin corpus callosum, and progressive microcephaly (SPATCCM, MIM#616657). We collected previously reported disease-causing variants of SLC1A4 (Fig. 2a), and found that the variant resides on exon 8, associated with two previously reported variants. Sanger sequencing confirmed both the parents to be heterozygote carriers (Fig. 2b, c). The variant resides on a highly conserved amino acid position (Fig. 2d), it is very rare and does not appear as homozygote in genomic databases (allele frequency highest in Finns 0.0040%, total 0.0018%). The variant was classified as likely pathogenic (ACMG criteria and further computational evidence of pathogenicity in Table 3). Empirical experiences suggested usefulness of L-serine supplementation in SPATCCM,21 and the patient was begun L-serine substitution as a treatment trial. During the treatment, no improvement was seen clinically in the proband’s symptoms nor in an EEG control, and the medication was discontinued after 2 months.
a SLC1A4 protein with domains according to Uniprot, and previously reported disease-causing variants, with exons indicated below the linear graph. The vertical axis depicts the number of patients found in literature, including the variant found in this study. Our novel variant p.L474P resides on the 8th exon, where two previous variants have been described. References for the gene review can be found in supplementary information. b Family pedigree: the unaffected parents were both heterozygote carriers and the affected proband homozygote. c Sanger sequencing results, both parents were heterozygote for the c.1421 T > C variant, the proband homozygote. d Protein conservation through species of amino acid residue Leu474 (yellow shade) and previously reported variants (blue shade).
An atypical, less-severe Menkes disease
The proband, P03, was the second child of the family, born uneventfully at term, with normal initial growth. There was no family history of hereditary conditions. At 8 months of age he stopped reaching new motoric milestones and presented with severe hypotonia. At 2 years of age, brain MRI showed cerebellar and vermis atrophy, with signal intensities in the thalamus and dentate nuclei. He had severe ataxia and athetosis. At 6 years of age, he was diagnosed with epilepsy, has not reached any new milestones, does not talk nor move independently, and is extremely hypotonic with athetoid movement. At 8 years of age, he required a gastrostomy. WES revealed a novel hemizygote ATP7A (NM_000052.7) truncating variant c.412 C > T (p.Gln138*). Mutations on ATP7A, the ATPase copper transporting alpha protein, cause Menkes disease (MIM#309400), a copper metabolism disorder leading to infant-onset encephalopathy, known by the coarse, kinky or steely hair on patients.22 We collected previously reported disease-causing variants (Fig. 3a), and found that nonsense variants are a common cause of the disease. The variant resided on a highly conserved amino acid residue (Fig. 3b) and is absent from human genome databases.
a ATP7A protein with domains according to Uniprot, and previously reported disease-causing variants, with exons indicated below the linear graph. The vertical axis depicts the number of patients found in literature. Protein level consequences shown for truncating variants in exons 2–7. Our novel variant p.Q138X resides on the 3rd exon, on the second Heavy-Metal Associated domain. References for the gene review can be found in supplementary information. b Protein conservation through species of amino acid residue p.Gln138 (yellow shade) and previously reported variants (blue shade). c Family pedigree: the affected proband was hemizygote for the variant carried by the mother, while all other maternal relatives alive were not carries. d Sanger sequencing results of the proband, parents, maternal uncle, and grandfather. The mother is a heterozygote carrier of the X-chromosomal c.412 C > T variant, while the affected proband’s genotype is hemizygote.
The proband’s phenotype did not match the classical Menkes disease presentation. Due to the genetic finding, the proband’s copper metabolism was assessed by measuring serum copper and ceruloplasmin (S-Cu 11 µmol/l, ref. 12.6–23.6 µmol/l; S-Ceruloplasmin 143 mg/l, ref. 200–500 mg/l), which were slightly below normal, supporting a copper metabolism disorder. A brain MRI controlled at 3 years of age showed previous findings, but also elongation and tortuosity of intracranial arteria, common in Menkes disease.22 Family samples available were tested for the variant by Sanger sequencing: the mother was a carrier of the X-chromosomal variant, whereas neither the proband’s unaffected maternal uncle nor grandfather had the variant (Fig. 3c, d). The variant was classified as pathogenic (ACMG criteria and computational evidence of pathogenicity in Table 3).
Childhood-onset severe hereditary spastic paraplegia
The proband, P14, was the second child of the family, born uneventfully at term, with normal initial growth. His father had been clinically diagnosed with mild hereditary motor sensory neuropathy (HMSN1) that also affected his father and grandfather, but genetic testing, including WES, had been inconclusive. At 8 months of age, the proband could not sit up, and he learned to crawl first at 11 months of age. At clinical examination, the proband presented with severe lower limb spasticity. Brain and spine MRI, as well as electroneuromyography (ENMG), were all normal. The child has never been ambulatory. WES revealed a novel heterozygote SPAST (NM_014946.4) missense variant c.1166 C > T (p.Thr389Ile). We collected previously reported disease-causing variants (Fig. 4a), finding that the variant resides on the “ATPases Associated with diverse cellular Activities” (AAA) domain, showing clustering of SPAST-related pathogenic variants. Most variants cause adult-onset HSP, but there are some reports of childhood-onset disease. The variant p.T389I is not found in genomic databases and is highly conserved through species (Fig. 4b). Sanger sequencing as well as parental testing confirms it to be de novo (Fig. 4c, d). Another missense variant on the same amino acid site, p.Thr389Ala, has been reported in Clinvar (ID 188190) by multiple submitters as causing hereditary spastic paraplegia (HSP). The variant was classified as pathogenic (ACMG criteria and computational evidence of pathogenicity in Table 3), and the HSP-disease of the proband to be unrelated to HMSN1 in the family.
a SPAST protein with domains according to Uniprot, and previously reported disease-causing variants; the variants below the linear graph have been described in children. The vertical axis above depicts the number of patients found in literature. Our novel p.T389I variant resides on the AAA-cassette, considered a mutational hotspot. References for the gene review can be found in supplementary information. b The Thr389 amino acid residue, indicated with yellow shade, is highly conserved throughout species. c Family pedigree, the affected proband was found to have a de novo variant c.1166C>T. The father and his father were affected by a clinically diagnosed HSMN1. d Sanger sequencing confirmed the variant to be de novo. MIT microtubule interacting and trafficking, MTBD microtubule binding domain, AAA ATPase associated with diverse cellular activities.
Novel variants with typical phenotypes on previously disease-associated genes
We found novel variants on four more patients with typical geno-phenotypes (Table 3). Defects in Chromodomain Helicase DNA-Binding Protein 2 (CHD2) are known to cause developmental and epileptic encephalopathy 94 (MIM#615369), whereas Regulator of Nonsense-Mediated mRNA Decay (UPF3B) is associated with X-linked syndromic intellectual developmental disorder 14 (MIM#300676). For probands P05 and P04, both CHD2 (NM_001271.4) c.1312 C > T (p.Gln438Ter) and UPF3B (NM_080632.3) c.462_465delCGAT (p.Ile154Metfs*20) are loss of function (LOF) variants in genes where LOF is a known mechanism of disease,23,24 and are as such classified as pathogenic (ACMG criteria in Table 3, additional data on Supplementary Figs. 2 and 3). Purine-Rich Element-Binding Protein A (PURA) causes a neurodevelopmental disorder with neonatal respiratory insufficiency, hypotonia, and feeding difficulties (MIM#616158), and Pyrroline-5-Carboxylate Reductase 2 (PYCR2) causes hypomyelinating leukodystrophy 10 (MIM#616420). For probands P07 and P06, both PURA (NM_005859.5) c.299 T > G (p.Leu100Arg) and PYCR2 (NM_013328.4) c.596 G > A (p.Arg199Gln) resided on highly-conserved amino acid residues previously associated with other pathogenic missense variants, PURA (p.Leu100Pro)25 and PYCR2 (p.Arg199Trp),26 and the variants were assigned as likely pathogenic (ACMG criteria in Table 3, additional data on Supplementary Figs. 4 and 5).
Discussion
Here, we report genetic findings of progressive neurological disorders of children in a mixed cohort from the largest pediatric neurology clinic in Finland. In our patient cohort of 47 families, not including apparent known syndromes, 19 received a definitive genetic diagnosis. Most of the variants were sporadic and de novo born, further emphasizing the importance of next-generation sequencing techniques in the diagnostic process. We aimed also to further describe the genetic landscape of pediatric encephalopathies in Finland. While 90% of our patients were of Finnish heritage, all families carried their specific gene defects and only one patient had a Finnish disease heritage disorder, PEHO syndrome. This indicates that disorders of Finnish disease heritage are well identifiable and therefore not included in the cohort of diseases with unknown background.
Eight of the found variants were novel and have not been previously reported to the best of our knowledge. These include variants in TRIT1, SPAST, ATP7A, and SLC1A4. Only 17 patients have been reported with defects in TRIT1 gene (MIM#617873), which encodes a tRNA isopentenyltransferase modifying mitochondrial tRNAs.18,19,20 In addition to the novel p.P24S variant, we present the first Finnish patient with the p.R327X19,20 variant, previously described in two studies from France20 and USA/China.19 The p.R327X is the most common pathogenic TRIT1 variant in genomic databases, and three times more frequent in the Finnish population than in others, suggesting it might be a Finnish founder allele.
Only 20 published patients having spastic tetraparesis, thin corpus callosum and progressive microcephaly (SPATCCM) have been published with SLC1A4 variants. This protein transports L-serine and participates in brain serine metabolism, importing L-serine to neuronal cells. SPATCCM is mostly reported in Ashkenazi Jews,21 with only three occurrences in European populations.27,28,29 Here we report the fourth European case of SLC1A4 caused SPATCCM with the hallmark symptoms. The variant is very rare, and although its reported frequency is highest in Finns, the sample (n = 1) in gnomAD is insufficient for further conclusions. Based on empirical experience of potentially beneficial effects of L-serine supplementation in Ashkenazi patients, our patient underwent a treatment trial, but with no detectable effect.
ATP7A is an ATPase copper transporting alpha protein which causes Menkes syndrome. Our patient’s symptoms started late, at 8-months, and he is still alive at 8-years old. Typically, Menkes manifests at 2–3 months with life expectancy less than 4 years. Our patient’s variant p.Q138X lies in the third exon. Truncating variants in exons 3 and 4 have also previously been suggested to cause moderate phenotypes of Menkes disease due to translation reinitiation in exon 5, leading to a partially functional protein.30 Previous reports included frameshift variants p.Val41fsX and p.Asn137Lysfs*22; the latter manifesting at the same time as our patient, and both having less-severe phenotypes than typically.30,31 Our report supports the conclusion that truncating variants in exons 3 or 4 of ATP7A associate with a slightly delayed manifestation and progression of Menkes disease.
Mutations in SPAST, a member of the AAA protein family, are the most frequent cause of both sporadic and familial HSP (MIM#182601), but childhood-onset, such as that of our patient, is rare.32 Schieving et al.33 concluded recently that all pediatric SPAST-related phenotypes have been caused by de novo variants on the AAA-cassette of SPAST. This conclusion is supported by our finding. The pathogenicity of our patient’s variant is supported by its conservation and also the fact that a previous patient with childhood-onset HSP, manifesting at 1 year of age, had a de novo SPAST variant changing an amino acid residue adjacent to that of our patient (p.Met390Val34 vs our patient’s p.Thr389Ile). It has been proposed that early childhood-onset HSP caused by SPAST could be accompanied by other modifier variants in SPAST such as the p.Ser44Leu, and one of such variants is typically found on one of the parents.33 Our patient, however, does not have other variants, and the variant is de novo. With the age-at-onset of 8 months, our patient is one of the earliest-onset patients with the disease so far reported.
Our approach used singleton WES. Previously we have ascertained that such analysis is a cost-effective and efficient tool in the diagnostic workup.5 Here, families with more than one affected child were more likely to gain a diagnosis (75% success rate) than families with a single affected child (40% success; de novo mutation in 53% of the cases). Several studies have investigated the difference in performance between singletons and trios. A recent study with 700 patients investigated exomes with a mix of singletons, duos and trios, and found no considerable difference in the diagnostic yield.4 Another study examined the performance between singletons and trios in a double-blinded randomized exome cohort of comparable size to this study. Their trios provided one additional diagnosis over singletons.35 Trios are better at distinguishing de novo and biallelic variants in probands, but with higher costs. Our study suggests, however, that disease causing variants in children can usually be found with singleton WES with the help of in silico predictions, absence of the variant in genomic databases, and segregation analyses with Sanger sequencing. This applies particularly when the finding is a previously reported pathogenic variant with matching phenotype (60% of the diagnoses in our study). However, Sanger sequencing is required to investigate segregation patterns for completely novel variants. We analyzed seven additional candidate variants (for five patients) by Sanger sequencing, showing that these were neutral variants, inherited from parents or present in unaffected siblings. An eighth variant, in a phenotypically well-matched gene, previously reported pathogenic but with a dissimilar phenotype, could not be confirmed de novo due to a missing parental sample and was not reported.
WES reached a genetic diagnosis in 42% of the cases. The number is comparable to other studies using WES.3 The majority still remained undiagnosed, and the limitations of WES have been widely described in literature.36,37 For instance, WES does not cover intronic areas, and cannot reliably identify e.g. chromosomal rearrangements. In our study, most of the patients had chromosomal microarray done without actionable findings before study inclusion. CNVs were not analyzed in this study. Some disorders can be of polygenic origin, there could still be unrecognized disease-causing genes overlooked in the analysis, and sometimes, disorders are of other than genetic origin. In the case of children, progressive neurological disorders are however often of genetic origin compared to some adulthood onset disorders where lifestyle choices can affect disease presentation. Noteworthy, some of our patients had had years of extensive diagnostic workups including karyotyping, different gene tests and gene panels without success, and received their diagnosis only after WES. An interesting avenue of research is finding the best strategy in implementing WES to benefit most patients and decrease the number of undiagnosed. A recent consensus statement reported that WES should precede chromosomal microarrays in some neurological disorders in children due to higher diagnostic yields.38 Considering cost-effectiveness, Tan et al.35 find that due to the high chance of a positive singleton WES, it would be cost-effective to always start with singleton and only later escalate to a trio if needed. Wortmann et al.37 describe widely the pitfalls of WES and suggest a genomic checklist to help focusing and combining different lines of investigations after a negative WES. Finding the optimal strategy requires still research efforts, but as prices of sequencing are expected to decrease, there will be increased availability for WES.
To conclude, we provide novel variants and further evidence for several rare pathogenic genes. In line with previous exome cohorts with childhood-onset neurological disorders, de novo variants are an important cause of these disorders.4 Also, due to the clinical variability of pediatric disorders, even classical syndromes can be challenging to diagnose in early stages or when presenting atypically. In these cases WES provides an opportunity for early diagnosis by interrogating all genes at once. Singleton WES led to diagnosis of several family members in three families. WES also provided actionable, treatment-modifying effect for at least three patients (15% of the diagnosed, 6% of all): ketogenic diet was recommended for the patients with GLUT1 deficiency syndrome (MIM#612126; SLC2A1) and pyruvate dehydrogenase deficiency (#312170; PDHA1), and L-serine supplementation was initiated for the patient suffering of SPATCCM (MIM#616657; SLC4A1). According to our study, singleton WES is useful in diagnosing sporadic disorders.
Data availability
The data that support the findings of this study are available upon reasonable request.
References
Neville, B. G. R. Pediatric neurology: the diagnostic process. Handb. Clin. Neurol. 111, 27–33 (2013).
De Vrieze, J. et al. Expanding phenotype of ATP1A3-related disorders: a case series. Child Neurol. Open 8, 2329048X211048068 (2021).
Shickh, S., Mighton, C., Uleryk, E., Pechlivanoglou, P. & Bombard, Y. The clinical utility of exome and genome sequencing across clinical indications: a systematic review. Hum. Genet. 140, 1403–1416 (2021).
Murrell, J. R. et al. Molecular diagnostic outcomes from 700 cases: what can we learn from a retrospective analysis of clinical exome sequencing? J. Mol. Diagn. 24, 274–286. (2022).
Aaltio, J. et al. Cost-effectiveness of whole-exome sequencing in progressive neurological disorders of children. Eur. J. Paediatr. Neurol. 36, 30–36 (2022).
Sainio, M. T. et al. Effectiveness of clinical exome sequencing in adult patients with difficult‐to‐diagnose neurological disorders. Acta Neurol. Scand. 145, 63–72 (2021).
Sulonen, A. M. et al. Comparison of solution-based exome capture methods for next generation sequencing. Genome Biol. 12, R94 (2011).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Lim, E. T. et al. Distribution and medical impact of loss-of-function variants in the Finnish founder population. PLoS Genet. 10, e1004494 (2014).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Sim, N. L. et al. SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40, W452–W457 (2012).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Suomalainen, A. & Syvänen, A.-C. Quantitative analysis of human DNA sequences by PCR and solid-phase minisequencing. Mol. Biotechnol. 15, 123–131 (2000).
Gallardo, E. et al. Magnetic resonance imaging findings of leg musculature in Charcot-Marie-Tooth disease type 2 due to dynamin 2 mutation. J. Neurol. 255, 986–992 (2008).
Reijnders, M. R. F. et al. PURA syndrome: clinical delineation and genotype-phenotype study in 32 individuals with review of published literature. J. Med. Genet. 55, 104–113 (2018).
Anttonen, A. K. et al. ZNHIT3 is defective in PEHO syndrome, a severe encephalopathy with cerebellar granule neuron loss. Brain 140, 1267–1279 (2017).
Yarham, J. W. et al. Defective i6A37 modification of mitochondrial and cytosolic tRNAs results from pathogenic mutations in TRIT1 and its substrate tRNA. PLoS Genet. 10, e1004424 (2014).
Muylle, E. et al. TRIT1 defect leads to a recognizable phenotype of myoclonic epilepsy, speech delay, strabismus, progressive spasticity, and normal lactate levels. J. Inherit. Metab. Dis. 45, 1039–1047 (2022).
Smol, T. et al. TRIT1 deficiency: two novel patients with four novel variants. Eur. J. Med. Genet. 65, 104603 (2022).
Damseh, N. et al. Mutations in SLC1A4, encoding the brain serine transporter, are associated with developmental delay, microcephaly and hypomyelination. J. Med. Genet. 52, 541–547 (2015).
Møller, L. B., Mogensen, M. & Horn, N. Molecular diagnosis of Menkes disease: genotype-phenotype correlation. Biochimie 91, 1273–1277 (2009).
Carvill, G. L. et al. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat. Genet. 45, 825–830 (2013).
Tarpey, P. S. et al. Mutations in UPF3B, a member of the nonsense-mediated mRNA decay complex, cause syndromic and nonsyndromic mental retardation. Nat. Genet. 39, 1127–1133 (2007).
Lalani, S. R. et al. Mutations in PURA cause profound neonatal hypotonia, seizures, and encephalopathy in 5q31.3 microdeletion syndrome. Am. J. Hum. Genet. 95, 579–583 (2014).
Zaki, M. S. et al. PYCR2 mutations cause a lethal syndrome of microcephaly and failure to thrive. Ann. Neurol. 80, 59–70 (2016).
Conroy, J. et al. Novel European SLC1A4 variant: infantile spasms and population ancestry analysis. J. Hum. Genet. 61, 761–764 (2016).
Sarigecili, E., Bulut, F. D. & Anlas, O. A rare cause of microcephaly, thin corpus callosum and refractory epilepsy due to a novel SLC1A4 gene mutation. Clin. Neurol. Neurosurg. 218, 107283 (2022).
Sedlackova, L. et al. Severe neurodevelopmental disorder with intractable seizures due to a novel SLC1A4 homozygous variant. Eur. J. Med. Genet. 64, 104263 (2021).
Paulsen, M. et al. Evidence that translation reinitiation leads to a partially functional menkes protein containing two copper-binding sites. Am. J. Hum. Genet. 79, 214–229 (2006).
Gérard-Blanluet, M. et al. Early development of occipital horns in a classical Menkes patient. Am. J. Med. Genet. A 130A, 211–213 (2004).
Lo Giudice, T., Lombardi, F., Santorelli, F. M., Kawarai, T. & Orlacchio, A. Hereditary spastic paraplegia: clinical-genetic characteristics and evolving molecular mechanisms. Exp. Neurol. 261, 518–539 (2014).
Schieving, J. H. et al. De novo SPAST mutations may cause a complex SPG4 phenotype. Brain 142, e31 (2019).
Álvarez, V. et al. Mutational spectrum of the SPG4 (SPAST) and SPG3A (ATL1) genes in Spanish patients with hereditary spastic paraplegia. BMC Neurol. 10, 89 (2010).
Tan, T. Y. et al. A head-to-head evaluation of the diagnostic efficacy and costs of trio versus singleton exome sequencing analysis. Eur. J. Hum. Genet. 27, 1791–1799 (2019).
Srivastava, S. et al. Clinical whole exome sequencing in child neurology practice. Ann. Neurol. 76, 473–483 (2014).
Wortmann, S. B. et al. How to proceed after "negative" exome: A review on genetic diagnostics, limitations, challenges, and emerging new multiomics techniques. J. Inherit. Metab. Dis. 45, 663–681 (2022).
Srivastava, S. et al. Meta-analysis and multidisciplinary consensus statement: exome sequencing is a first-tier clinical diagnostic test for individuals with neurodevelopmental disorders. Genet. Med. 21, 2413–2421 (2019).
Amir, E. et al. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nat. Genet. 23, 185–188 (1999).
Calame, D. G. et al. Atypical phenotypes caused by the ATP1A3 varaint p.P775L. Neurology 94, 1946 (2020).
Kleppa, L., Kanavin, O. J., Klungland, A. & Stromme, P. A novel splice site mutation in the Cockayne syndrome group A gene in two siblings with Cockayne syndrome. Neuroscience 145, 1397–1406 (2007).
Joao Silva, M. et al. Pyruvate dehydrogenase deficiency: identification of a novel mutation in the PDHA1 gene which responds to amino acid supplementation. Eur. J. Pediatr. 168, 17–22 (2009).
Zimon, M. et al. Dominant GDAP1 mutations cause predominantly mild CMT phenotypes. Neurology 77, 540–548 (2011).
Weber, Y. G. et al. GLUT1 mutations are a cause of paroxysmal exertion-induced dyskinesias and induce hemolytic anemia by a cation leak. J. Clin. Investig. 118, 2157–2168 (2008).
Chen, Y. Z. et al. Autosomal dominant familial dyskinesia and facial myokymia: single exome sequencing identifies a mutation in adenylyl cyclase 5. Arch. Neurol. 69, 630–635 (2012).
Riant, F. et al. ATP1A2 mutations in 11 families with familial hemiplegic migraine. Hum. Mutat. 26, 281 (2005).
Acknowledgements
The authors thank all families and patients for contributing to this study. In addition, we thank Dr. Henna Tyynismaa and Markus Sainio for collaboration in the Neurogenomics project. Dr. Lisbeth Birk Møller for consultation in the case of the ATP7A variant, and Markus Innilä and Tuula Manninen for their lab expertise. The authors wish to acknowledge CSC – IT Center for Science, Finland for computational resources.
Funding
The pHealth academy program of the Academy of Finland (subgrant number 292584) funded this study, as part of the Neurogenomics Finland consortium project. Arvo and Lea Ylppö Foundation funded J.A. for research in the field of pediatric neurology. A.S. has been funded by Academy of Finland, University of Helsinki, Sigrid Juselius Foundation, and Jane and Aatos Erkko Foundation during this study. The funding sources had no role in conducting this study. Open Access funding provided by University of Helsinki including Helsinki University Central Hospital.
Author information
Authors and Affiliations
Contributions
J.A. conceptualized and designed the study, drafted the initial manuscript, and revised the manuscript. A.E., V.B., and S.O. helped in the conceptualization of the study, reviewed, and revised the manuscript. T.L. and P.I. supervised patient recruitment and, together with A.S., designed, provided medical expertise, revised the manuscript, and supervised the study. All authors approved the final manuscript as submitted and agree to be accountable for all aspects of the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
Patient consent was required for participation in the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Aaltio, J., Etula, A., Ojanen, S. et al. Genetic etiology of progressive pediatric neurological disorders. Pediatr Res 95, 102–111 (2024). https://doi.org/10.1038/s41390-023-02767-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41390-023-02767-z
- Springer Nature America, Inc.