Abstract
The persistent COVID-19 pandemic since 2020 has brought an enormous public health burden to the global society and is accompanied by various evolution of the virus genome. The consistently emerging SARS-CoV-2 variants harboring critical mutations impact the molecular characteristics of viral proteins and display heterogeneous behaviors in immune evasion, transmissibility, and the clinical manifestation during infection, which differ each strain and endow them with distinguished features during populational spread. Several SARS-CoV-2 variants, identified as Variants of Concern (VOC) by the World Health Organization, challenged global efforts on COVID-19 control due to the rapid worldwide spread and enhanced immune evasion from current antibodies and vaccines. Moreover, the recent Omicron variant even exacerbated the global anxiety in the continuous pandemic. Its significant evasion from current medical treatment and disease control even highlights the necessity of combinatory investigation of the mutational pattern and influence of the mutations on viral dynamics against populational immunity, which would greatly facilitate drug and vaccine development and benefit the global public health policymaking. Hence in this review, we summarized the molecular characteristics, immune evasion, and impacts of the SARS-CoV-2 variants and focused on the parallel comparison of different variants in mutational profile, transmissibility and tropism alteration, treatment effectiveness, and clinical manifestations, in order to provide a comprehensive landscape for SARS-CoV-2 variant research.
Similar content being viewed by others
Introduction
The COVID-19 pandemic has lasted for over 2 years and caused over 6 million death cases.1 A wide variety of SARS-CoV-2 variants emerged during its persistence and displayed evolving adaptation to global populational immunity,2,3,4,5 leading to rapid worldwide spread and heterogenous escape from available therapeutic drugs and vaccines.6,7,8,9 The mutations harbored in the genome of SARS-CoV-2 variants have a significant impact on viral protein structures, function, and immunogenicity, which was strongly associated with the immunological response and clinical outcome in humans.10,11,12,13
This review systematically describes the evolutionary and molecular characteristics of SARS-CoV-2 variants and summarizes the mutational impact on the critical viral proteins. Then it comprehensively describes the landscape of immune evasion of various critical variants from the currently approved antibody, small antiviral molecules, and vaccines. Lastly, it describes the epidemiological profile of SARS-CoV-2 variants and overview the different critical strains’ changes in infectivity, host tropism, and clinical manifestation and outcome. Detailed datasets for the parameterized depiction of the difference between SARS-CoV-2 variants in molecular characteristics, immune evasion, and clinical impact are also provided.
Molecular characteristics of sequence and the encoded proteins of SARS-CoV-2 variants
The genomic evolution of SARS-CoV-2
Since the emergency of SARS-CoV-2,14,15,16,17 its viral genome has been under constant and rapid mutation to adapt host system.18,19 Like other RNA virus,20,21,22,23,24,25 a high mutation rate benefits the emergence of novel variants with a significant change in viral phenotypes.20,26 Therefore, the global scientific community endeavors to construct systematic tracking systems of SARS-CoV-2 mutations and identified the clade with a genetically close relationship.27
The phylogenetic classification is widely used as a fundamental method for emergent SARS-CoV-2 strain classification in the clade-nomenclature system (terming the major strain as clade code such as GR) by Global Initiative of Sharing All Influenza Data (GISAID)28 or NextStrain29 or Pango lineage system (terming the major strain as letter and number with point interval such as B.1.1.7) by Pango Network30 (Fig. 1a). However, with the rapid increase in submitted sequence to the genomic database and wider observation of sequential distribution in the infected population, a more compact naming system for the critical variants was demanded to guide global anti-virus policy. Therefore World Health Organization (WHO) proposed using the Greek alphabet to name the critical SARS-CoV-2 clades or Pango lineages and raised the concept of Variant of Concern (VOCs) and Variants of Interest (VOIs) as a larger dynamic classification.17 Our review used the WHO naming system to indicate the strains in representing both sequence identity and their impact on disease control.
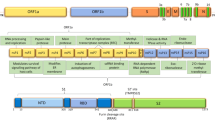
SARS-CoV-2 evolution, prevalence, and genome architecture. a Phylogenetic analysis of sequence divergence of SARS-CoV-2 circulating variants based on clade classification in February 2022. The WHO labeling of clades is marked besides. b Sequential frequency of major clades of SARS-CoV-2 variants from April 2021 to February 2022. c Linear genome architecture of encoded viral protein and structural overview of SARS-CoV-2. The phylogenetic analysis and sequential frequency data come from the Nextstrain GISAID database (https://nextstrain.org/ncov/gisaid/global), and figures in related (a, b) are generated under the CC-BY 4.0 permission. BioRender is used to generate the structure diagram of SARS-CoV-2 virus in Fig. 1c
Early 2020 has witnessed the emergence of the first widely reported spike mutation of SARS-CoV-2, D614G.31,32,33,34,35,36 In December 2020, the Alpha variant (B.1.1.7) harboring another critical mutation N501Y37,38 in spike protein, initially expanded in the southeast of England, soon became the first globally distributed VOC (Fig. 1b).39,40,41 Later the Beta variant (B.1.351) was found in South Africa and manifested a rapid domestic distribution to an over 80% prevalence.42,43 One month later, the Gamma variant (P.1) was reported in Brazil, and the travelers arriving in Japan from Brazil.44,45 Delta variant (B.1.617.2) was first detected in India in May 2021 and rapidly became the dominant variant worldwide by late 2021, while some sub-clade of Delta variant displayed a unique penchant in epidemic areas, such as Clade 20I (Delta) in some parts of Asia.41,46,47 Delta-dominant epidemic lasted quite long in the world until Omicron (B.1.1.529) in November 2021, which was first reported in South Africa,48,49 and soon in Chinese Hong Kong.50 Since its discovery, Omicron rapidly displaced Delta and became the major variant worldwide.48,51,52,53,54 By the time of 31 March 2022, only variants Alpha (B.1.1.7), Beta (B.1.351), Gamma (P.1), Delta (B.1.617.2), and Omicron (B.1.1.529) were labeled as Variant of Concern (VOCs), within which only Delta and Omicron were recognized as currently circulating VOCs.50,55,56,57,58,59,60,61
As the SARS-CoV-2 RNA genome encoded a set of structural and non-structural proteins (Fig. 1c),62,63,64,65 the mutations in these proteins lead to various molecular alterations in protein characteristics, shaping the difference between variant to variant.66
SARS-CoV-2 spike protein
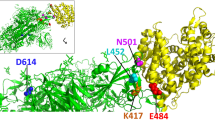
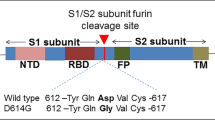
The SARS-CoV-2 spike protein, as the major structural protein, is embedded in the SARS-CoV-2 viral membrane in homo-trimeric form and recognizes human ACE2 as a receptor for viral entry.65,67,68,69,70 It consists of two subunits, S1 and S2, cleaved by host furin.65,71,72,73,74,75,76 The distal S1 subunit contains two important regions, RBD (receptor binding domain) and NTD (N-terminal domain),77 and the RBD acts as the binding region for ACE2,77,78,79 making it the most critical target affecting virus-host interaction and vulnerable site to antibody neutralization80,81,82,83,84(Fig. 2a). Currently, most neutralizing antibodies or vaccines are developed to target the RBD to block or inhibit viral infection.80,84,85,86,87,88,89,90,91,92,93 Furthermore, the binding with ACE2 of RBD requires conformational adaptation, and an easier transition from “closed” to “open” conformation of spike protein benefits the viral infection.32,94,95,96 Therefore, mutations in the spike protein of SARS-CoV-2 variants could significantly influence the structure of the spike protein conformation and further the interaction with ACE2 or neutralizing antibodies32,95,97,98,99,100 (Figs. 2b and 3).
Mutations and their effect on SARS-CoV-2 spike protein. a Structural overview of SARS-CoV-2 spike protein and its subdomain RBD and NTD. Mutations from variants were marked beside the colored surface. b Patterns of mutational impact on the spike protein. Mutations affect the spike affinity to ACE2 and neutralizing antibodies (nAbs) and influence the spike protein conformations. BioRender is used to generate the structural presentation and cartoon models. Pymol is used to generate the surface model of RBD or NTD region. 6VSB, 7CM4, and 6M0J structures are retrieved from PDB database
Heatmap of the mutation site and the mutational impact of SARS-CoV-2 VOCs on the spike protein characteristics. The mutations of VOCs vary from strain to strain and could exert multiple impacts on the protein characteristics of spike protein, including the ACE2 binding, antibody affinity, protein conformation, stability, and productivity. Quantitative results were recorded on the influence of ACE2 affinity and RBD production, and qualitative results were recorded on antibody affinity, conformational change, and protein stability. Details for each mutation can be found in Supplementary Table 1
Impact on ACE2 binding
In vitro binding experiments have shown that SARS-CoV-2 bound to human ACE2 with an affinity of about 10 nM, which was 10–20-folds higher than SARS-CoV,79,87,101,102 which is a potential reason for the higher infection rate of SARS-CoV-2.103 The residues of RBD directly participated in ACE2 binding79 are K417, Y449, Y453, N487, Y489, G496, T500, G502, Y505, L455, F456, F486, Q493, N501, Q498, and the mutations at or beside these sites may directly impact the interaction with ACE2.103 The N501Y is one of the most common mutations at spike protein and could be found in Alpha, Beta, Gamma, and Omicron. Various studies have demonstrated that this mutation enhances the binding with human ACE2 through the extra introduction of π-π packing between RBD Y501 and ACE2 Y41.103,104 Likewise, mutation E484K from Beta and Gamma variant or L452R and T478K from Delta variant was also reported to increase the affinity with ACE2 by mutational scanning or computational analysis.105,106,107,108 However, not all the mutations at RBD from variants benefited ACE2 binding. Studies showed that mutation at K417 from Beta and Gamma variants could impair the RBD binding with ACE2.103,104,105 More mutations from Omicron were yet to be studied on their impact on ACE2 binding affinity, despite some of them being quite far from the interface with ACE2.53,55,58,109,110 SARS-CoV-2 evolved to adapt to the host, leading to widespread circulation among animals while still retaining its ability to efficiently utilize human ACE2 for entry, thus allowing for transmission of the virus back into humans.99,111
Impact on antibody binding
The spike protein could elicit a protective antibody against viral infection through the key regions mediating viral entry and fusion. Therefore, there were early explorations of using anti-SARS-CoV spike antibodies to neutralize SARS-CoV-2 due to the high sequence similarity.81,87 Unfortunately, none of them (m396, S230, 80R, CR3014) manifested obviously neutralization activity against SARS-CoV-2 infection,67,87,102,112 and these results provided primary evidence of antibody escape caused by the amino acid substitution in antigen.79,101 As more antibodies targeting SARS-CoV-2 spike or RBD were isolated or developed, many of them with potent neutralizing capability and clinical perspective were reported and investigated in-depth with biological and structural experiments.79,106,113,114,115,116,117,118,119,120
For mutations located on RBD, a structural study revealed that K417 mutation from Beta, Gamma, and Omicron reduced antibody binding to spike protein of C682, C614, and C653,121,122 E484K from Beta and Gamma diminished binding of C602, C627, C628, C653, C643 and C6710,105,121,123,124 and N501Y from Alpha, Beta, Gamma and Omicron diminished binding of C613, C628, C663, and C670.121 It was also revealed that L452R from Delta variant reduced DH1041 binding to spike protein.124,125,126 Another mutational analysis evaluating the overall mutation sets from SARS-CoV-2 variants based on computational analysis and in vitro experiment also indicated that currently approved antibodies, including REGN-10933 (Casirivimab), REGN-10987 (Imdevimab), and CT-P59 (Regdanvimab) displayed decreased affinity with spike proteins from all VOCs.60,127
For mutations not located on RBD, L18F in Gamma, T19R in Delta and Omicron, D80A in Beta, G142-/D, and Y144- in Omicron reduced S2L28, S2X28, S2M28, S2X333 and 4A8 (PDB:7C2L) binding to spike protein.115,128 The deletion of amino acid residues 241~243 in NTD of the Beta variant nearly abolished the binding of 4A8, an antibody targeting the NTD domain.107,115
Impact on protein conformation, yield, and stability
The conformation of spike protein also determined the efficacy of ACE2 binding, as the spike protein had two conformations, “open” and “close,” in the representation of the sub-structural arrangement of RBD as “up” or “down”. Only spike protein with at least one RBD in “up” conformation could be bound by ACE2.94 Therefore, the mutational impact on the spike conformation would also influence the spike protein binding with ACE2. Some studies revealed that N501Y and D614G could facilitate the transition of spike protein from “closed” to “open“.31,32,34,95,100,129 Furthermore, mutations at H655 and P681 were reported to increase the cleavage efficiency of spike protein at furin site to promote viral-cell membrane fusion.104,130,131,132,133 As for the NTD domain, it was reported that H69del/V70del in the Omicron and Alpha variant resulted in the contraction of NTD and lead to a tighter NTD configuration.114,115,116
Moreover, the stability and yield of spike protein upon the viral membrane would also influence the overall infectivity, as associated with the availability of spike protein for viral entry.34,36 A series of mutations were associated with the instability of spike protein,55,57,134 while one study made a parallel analysis of the RBD productivity under various mutations and found that most prevalent mutations would facilitate the yield of RBD.103 These results provided another perspective to analyze the mutational impact on the spike protein.
SARS-CoV-2 structural proteins beyond spike
Except from the spike, there are other three structural proteins: Envelop (E), Membrane (M), and nucleocapsid (N), encoded by ORF4, ORF5, and ORF9.135,136,137,138 E and M majorly participate in virion assembly,136,139 while N forms the viral capsid structure associated with viral RNA and facilitates genome packaging. Despite not participating in the initiation of viral infection, these proteins had a significant role in viral replication, assembly, and release. Their mutations could also influence viral activity.137,140,141,142,143
Compared with E or M, much more mutations in N protein were observed,56,144,145 and Omicron had the most abundant mutations, including P13L, E31del, R32del, S33del, and S413R as unique mutations, and 203K (Alpha, Gamma, Delta, and Omicron), and G204R (Alpha, Gamma, and Omicron) as common mutations. And it’s also reported that R203K/G204R in N protein increased viral RNA binding ability.146,147,148,149
Hence, more evidence was required to reveal the mutational impact on the E, M, or N protein of the SARS-CoV-2 variants.149
SARS-CoV-2 non-structural proteins
Nearly two-thirds of the genome coded ORF1a/ORF1ab can be translated into polyprotein pp1a and pp1ab. The translation of ORF1ab is realized by −1 frameshift at C13468, allowing continued translation instead of termination. The translated product pp1a/pp1ab could be proteolytically cleaved into non-structural proteins (Nsps) with different functions by viral proteases.20,141,150,151
Among the Nsps, Nsp3, Nsp5, and Nsp12 received the most attention for both biological investigation and drug development, as Nsp3 and Nsp5 were viral protease PLpro (Papain-like protease)152,153 and Mpro/3CLpro (Main protease/3 chymotrypsin-like protease)154,155 mediating polyprotein cleavage and Nsp12 was the RNA-dependent RNA polymerase along with co-factor Nsp7/8 mediating genome replication and transcription.156,157 Therefore, mutations in these key viral proteins may further impact viral survival (Fig. 4). Nsp3 is now reported with the highest variation rate among the non-structural proteins and is closely related to the overall genome variation. Besides, great mutation diversity among the VOCs was observed since no shared mutation in Nsp3 was found among them. In comparison, relatively few mutations were found in the Nsp5 and Nsp12 (Nsp5:K90R-Beta, Nsp5: P132H-Omicron, Nsp12: P314L-All, Nsp12:G662S-Delta), and hence these two Nsps were recognized as relatively static in circulating variants. A mutational analysis using computational modeling is recently reported to reveal the influence of mutations in Nsps on protein stability and flexibility showed little mutation from VOCs with a significant impact on the Nsps, especially Nsp3, Nsp5, and Nsp12. Such as mutation P314L at Nsp12 was reported to decrease the protein stability.27,45,61,134,158,159,160,161,162,163
Heatmap of the mutation site and the mutational impact of SARS-CoV-2 variants on the non-structural proteins. The mutation site and its wild type and mutant residue in ORF sequence and detailed non-structural protein sequence among all variants are displayed. Each mutation is marked in a different color in the representation of the occurrence in VOCs and color-scale in reflection of the impact on protein stability and flexibility. Details for each mutation can be found in Supplementary Table 1
In summary, the evolution of SARS-CoV-2 endowed each clade with a distinct pattern of mutations. These mutations exerted various impacts on viral molecular characteristics. Current studies focused on investigating the mutational impact on spike protein due to its critical role in receptor binding and antibody evasion. It was broadly found that the mutations could alter the protein characteristics to benefit from ACE2 binding and diminish the binding with neutralization antibodies. In comparison, relatively less attention was paid to the mutational impact on other structural and non-structural proteins. Although sequential analysis has revealed mutations on these proteins from SARS-CoV-2 variants, little effort was given to the in-depth study of their influence on the protein structure and function and potential resistance to targeted drugs.141,153
Immune evasion of SARS-CoV-2 variants from current therapeutic agents
Neutralizing antibodies
The preliminary step for viral entry was the spike protein binding to the host cell receptor ACE2 (Fig. 5). Neutralizing antibodies (nAbs) targeting the spike protein, especially the RBD domain directly located at the interface with ACE2, can neutralize viral infection by blocking receptor recognition.164 Moreover, nAbs could also mediate antibody-dependent cellular cytotoxicity (ADCC) to eliminate infected host cells expressing the spike protein.165,166,167,168 Thus, many neutralizing antibodies against spike protein were developed as potential therapeutic agents against acute SARS-CoV-2 viral infection.169,170,171
Overview of SARS-CoV-2 infection and replication and the mechanism for anti-SARS-CoV-2 therapeutic agents. Step1: spike protein of SARS-CoV-2 binds to the host ACE2 to initiate viral attachment and endocytosis. The vaccine-induced antibody, neutralizing antibody, and ACE2-mimic protein could block this step. Step2: spike protein was cleaved by host TMPRSS2 to mediate spike protein conformation and viral fusion with the host membrane in an acidic endosomal environment. Small molecules such as Arbidol acted in this step. Step3: the released viral RNA genome is translated into polyproteins, and the translated product undergoes proteolysis by viral protein PLpro and 3CLpro to generate mature non-structural proteins and initiate viral replication. Protease inhibitors such as Nirmatrelvir target this step. Step4: the assembly of RNA-dependent RNA polymerase (RdRp) complex would start viral genome replication and transcription for structural proteins. RdRp inhibitors such as Remdesivir target this step. Step5: the transcripts for structural proteins would be translated further into the cytoplasm for N protein and in the endoplasmic reticulum for S, M, and E proteins. Step6: replicated RNA genome binds with N proteins to form nucleocapsid, and it would further assemble with other structural proteins in the membrane envelope. Immune modulators such as interferon act in this step. Step7: the assembled and enveloped virion is transported to the cell membrane and released. BioRender was used to generate this figure
According to the binding epitopes, SARS-CoV-2 nAbs can be classified as RBD-targeted nAbs, NTD-targeted nAbs and other protein-targeted nAbs172,173,174,175,176,177 and RBD-targeted nAbs accounted for the most widely-accepted antibody type entering the clinical trials for preventing SARS-CoV-2 infection.178,179,180,181 Currently, nine neutralizing monoclonal antibodies and their combinatorial cocktail, Bamlanivimab (LY-CoV555),182 Etesevimab (LY-CoV016),183 Casirivimab (REGN10933), Imdevimab (REGN10987)184,185 Cilgavimab (AZD1061/COV2-2130), Tixagevimab (AZD8895/COV2-2196),186 Sotrovimab (VIR-7831),187 Regdanvimab (CT-P59)188,189 and Bebtelovimab (LY-CoV1404)183 targeting SARS-COV-2 spike protein were issued with emergency use authorization (EUA) by FDA for treatment of mild to moderate SARS-CoV-2 infected individuals. These authorized antibodies all displayed potent neutralizing capability against the ancestral strain of SARS-CoV-2.5,164,190,191
With the emergence of SARS-CoV-2 variants, many studies investigated the altered efficacy against viral infection of these antibodies.4,192,193,194,195,196,197,198 The development of approaches in studying antibody’s molecular and structural characteristics has provided preliminary insight into the changes of binding dynamics of spike-antibody and the following potential impact on antibody neutralization efficacy as described above.197,198,199,200 With multiple critical mutations destabilizing the antibody interaction with spike or RBD, in vitro experiments also revealed the worrying performance of current antibodies against the SARS-CoV-2 variants.201,202
Considering that in vitro neutralization for the same therapeutic agent might differ under different circumstances, including virus types, cell culture types, and reference virus isolation, we listed the pseudovirus or authentic virus half-maximal inhibitory concentration (IC50) values of the mentioned authorized antibodies in range by reference to collective information from Stanford University Coronavirus Antiviral & Resistance Database and the contained corresponding studies on each antibody (Fig. 6).203,204,205
The summary of neutralization capability of approved antibodies against in vitro pseudovirus and authentic virus infection from SARS-CoV-2 variants. The ranges of reported neutralization capability against each VOC are listed. The lower-limit concentration IC50 are indicated in a red-to-blue color scale to represent the minimal neutralizing capability of each antibody reported. A bluer color indicates a reported stronger evasion of VOCs from antibody neutralization than ancestral strain. The detailed antibody neutralization concentration from each study could be referred in Supplementary Table 2
Most antibodies or cocktail pairs maintained the neutralizing efficacy against Alpha, Beta, Gamma, and Delta variants.5,117,206 Only Bamlanvimab showed great vulnerability toward all the VOCs, and the combination use of Etesevimab could barely improve the efficacy against specific strains.5 However, in terms of the Omicron strain, a significant reduction of neutralizing titer (>100 folds decrease) against both pseudovirus and authentic virus of all antibodies can be observed.5 In 2021, the use of Bamlanivimab alone was withdrawn from the authorization list due to its limited capability to block Beta and Gamma infection.207 Interestingly, the EUA of Bamlanivimab and Etesevimab in combination was later revised for post-exposure prophylaxis (prevention) because of their neutralizing potency against Delta, but the antibody cocktail received withdrawal again against Omicron for lack of effectiveness both in vitro and in vivo by November 2021.208
It should be noticed that an antibody pair, Cilgavimab+Tixagevimab (Evusheld) could maintain its efficacy against VOCs. Cilgavimab+ Tixagevimab showed a 5-folds to 12-folds decrease at a minimal neutralization against authentic Omicron, while the independent use in neutralizing assay displayed even significantly diminished efficacy (IC50 > 10,000/10,000 ng/mL in pseudovirus assay and >10,000/5000 ng/mL in authentic virus assay).193
In comparison, although individual use of Sotrovimab showed great vulnerability to the evasion of the Omicron variant,5 Sotrovimab even showed potency in controlling in vivo viral infection by ADCC and antibody-dependent cellular phagocytosis and cross-neutralizing capability against other sabecoronovirus,165 providing a possible mechanism for its use.
The alarming evasion of VOCs from currently approved antibodies urged novel antibody discovery or development. Recently Bebtelovimab (LY-CoV1404), a novel antibody with the latest approval for clinical use, displayed outstanding performance (IC50 = 0.003 μg/ml) in neutralizing Omicron pseudovirus and authentic virus, shedding light on the development of effective antibody therapy against Omicron variant.122,205,209 Besides NTD domain targeting, RBD-NTD dual-targeting or multi-spike variants-targeting antibodies were under development, and early findings showed their potential advantage in neutralizing more mutated variants such as Omicron previously reported antibody-based on screening the binding affinity with wild-type RBD or spike.210,211,212,213,214 Moreover, cocktail usage such as Cilgavimab+Tixagevimab provided a potential strategy of rationally using currently available antibodies to avoid the significant deficiency in neutralizing specific VOCs during individual application.188,215 Nevertheless, antibody-based therapy still confronts a huge challenge from the current Omicron pandemic. A new strategy to optimize the discovery and development of novel antibodies in adaptation to the mutational impact on neutralizing efficacy of emerging SARS-CoV-2 variants is urgently demanded, and systematic surveillance of antibody efficacy.
Vaccines
Currently, prophylactic vaccines remain the mainstay preventing SARS-COV 2 infection.216 According to the WHO COVID-19 vaccine tracker and landscape, there are currently 153 candidate vaccines under clinical development and 196 candidate vaccines under preclinical development worldwide,217 among which 19 vaccines have been authorized or fully approved in various countries, and some have been incorporated into national vaccination programs and widely applied.218 At present, authorized vaccines using three major platforms, mRNA, adenovirus, and inactivated virus, account for over 95% of vaccination doses around the world, including the two mRNA vaccines BNT162b2 (Pfizer/BioNTech) and mRNA-1273 (Moderna), one adenovirus vaccine AZD-1222 (AstraZeneca) and two inactivated vaccines BBIBP-CorV and CoronaVac (Sinovac).219,220,221 Approved vaccines have shown substantial efficacy in both in vitro neutralization and populational scale against the ancestral strain of SARS-CoV-2,222 but emerging evidence indicated significant immune escape of VOCs from the vaccine-induced protection.59,117,223,224,225
Neutralizing antibodies induced by the antigen underlie the vaccine protection against SARS-COV 2 infection, and the neutralizing titer of serum from vaccinated people is a key indicator for evaluating vaccine effectiveness, making in vitro neutralization analysis a convenient but important method for monitoring vaccine-elicited humoral immunity against SARS-COV 2 variants.226,227 Nevertheless, due to the diversity in serum condition, experimental procedures, and statistical calculation for serum neutralization titer, current studies reported a wide range of 50% neutralization titer (NT50) of serum against each VOC strain. Therefore, we listed the reduction fold in NT50 of pseudovirus or authentic virus neutralization against each variant of the mentioned authorized vaccines in average and the range compared to the ancestral strain by reference to collective information from Stanford University Coronavirus Antiviral & Resistance Database (https://covdb.stanford.edu/) and the contained corresponding studies on each vaccine228 (Fig. 7).
The summary of reduction folds of neutralization titers of elicited antibody by approved vaccines against in vitro pseudovirus and authentic virus infection from SARS-CoV-2 variants. The average and ranges of reduction folds of neutralization titers against each VOC compared to ancestral strain are listed. The average folds are indicated in a red-to-blue color scale to represent the average neutralizing capability of each vaccine against variants. A bluer color indicates a stronger evasion of VOCs from neutralization of vaccine-elicited antibodies in comparison to the ancestral strain. The detailed neutralization titer reduction from each study could be referred in Supplementary Table 3
In summary, sera from people receiving programmed or even booster vaccines showed a generally declined neutralization titers against different SARS-COV 2 VOCs compared with neutralization titers against the ancestral strain. For programmed vaccination, Alpha,229,230,231 Gamma,232,233,234 and Delta235,236,237 variants manifested relatively mild escape from neutralization (mostly ≤ 5 folds reduction), followed by the Beta variant232,238,239 (mostly 5–10 folds reduction). The most prominent reduction in the Omicron variant (mostly 10 folds reduction) was observed.60,193,235,240 This phenomenon was consistent with the evasion of therapeutic antibodies by Beta and Omicron variants discussed above, and neutralization assay results obtained from authentic virus and pseudovirus were generally parallel. The booster dose for each vaccine appeared to increase the neutralizing capacity of vaccine sera against VOCs by large, but the Omicron variants still presented obvious evasion from vaccine sera, with 8, 4.7, and 9.3-folds decrease for BNT162b2,193,235,241,242,243 mRNA-1273,235,242,244 and BBIBP-CorV,118,245,246 in the pseudovirus-based assay, and 12.3, 9.1 and 16.3-folds decrease respectively for BNT162b2,4,60,242,247,248 mRNA-1273,242,247,248 and AZD1222 in the authentic virus-based assay.60
Further insight into the Omicron variant evasion from neutralization showed an even remarkable decrease in vaccine protection over time. In a study, sera were obtained from 17 people in the second week, third month, and the sixth-month post BNT162b2 programmed immunization, and their neutralization against SARS-COV 2 wild type or Omicron variant was measured by authentic virus neutralization assay.249 It was found that the proportion of samples with neutralizing titer below the limit of detection against Omicron variants increased from 23.5% (2nd week) to 41.2% (3rd month) and 64.7% (6th month), suggesting that programmed vaccination of BNT162b could not elicit durable protection against the Omicron variant, and strengthening the necessity of booster vaccination.
SARS-CoV-2 VOCs, especially the Omicron variant, seriously affect vaccine-induced immunity from sera neutralization. Booster seems to be an ideal strategy for handling the immune evasion of the Delta variant but still partially compromises the Omicron variant. More importantly, as we have witnessed the continuing decrease in neutralizing efficacy post programmed vaccination, it deserves further observation on how well the neutralization titer post-booster could be maintained, and the time durability of elicited neutralizing antibodies could be another decisive factor influencing the final protective efficacy of vaccines.
Viral inhibitor drugs
Compared to the antibody, the viral inhibitor for SARS-CoV-2 focused more on the host factor facilitating viral infection and SARS-CoV-2 non-structural proteins.205,250,251,252,253 These proteins played a critical role during viral replication and maintained relatively sequential stability compared to the spike.253,254,255,256,257,258 Two mainstay strategies are adopted in developing antiviral drugs against COVID-19, including novel design of virus-targeted drugs and repurposing of currently available drugs with potential antiviral activity.175,259,260 However, it often took decades to develop novel drugs due to drug management authorities’ comprehensive pharmacological and biosafety evaluation. Hence, broad-spectrum drugs with prior validation against other pathogens such as viruses (SARS-CoV, MERS-CoV, HIV, EboV) and parasites (Malaria Plasmodium) received more attention for evaluation of their potential use in the ongoing pandemic, and these repurposed drug candidates are prone to large-scale manufacturing and delivery once sufficient evidence was acquired examining their effectiveness against SARS-CoV-2.261,262,263,264,265 Currently, various viral inhibitors are under clinical trial, and some have entered clinical use.260,266,267,268,269
The emergence of SARS-CoV-2 variants and broad reports of their evasion from neutralization by nAbs and vaccines lift the expectation of an anti-SARS-CoV-2 viral inhibitor.270 More and more experimental and clinical studies revealed the unique advantage of viral inhibitors in controlling the viral infection of VOCs due to the conservative drug target.258,271,272,273
The antiviral effects of small molecule inhibitors are determined by 50% and 90% effective concentration (EC50 and EC90) values. Here, we listed EC50 of small molecules in ranges with the reported value from corresponding studies on each molecule (Fig. 8).274
The summary of neutralization capability of small molecule drugs against in vitro authentic virus infection from SARS-CoV-2 variants. The ranges of neutralization capability of small molecule drugs targeting both viral and non-viral targets against VOCs are listed. The lower-limit concentration EC50 are indicated in a red-to-blue color scale to represent the minimal neutralizing capability of each antibody reported. A bluer color indicates a stronger evasion of VOCs from neutralization of small molecule drug in comparison to ancestral strain. The detailed small molecule neutralization capability from each study could be referred in Supplementary Table 4
Broad-spectrum antiviral inhibitors
Various viruses, including influenza virus, SARS-CoV, and MERS-CoV, employ TMPRSS2 to cleavage the fusion protein to mediate viral-host membrane fusion. Thus, TMPRSS2 inhibitors have become a promising target for inhibiting virus infection.275,276,277 Camostat and Nafamostat mesylate are oral TMPRSS2 inhibitors, and both enter phase III clinical trials with previously reported applications on SARS-CoV and MERS-CoV infection.278 Furthermore, Nafamostat was reported superior to Camostat in specificity and effectiveness.261,279 Research reported that these two drugs could effectively block the virus infection of ancestral SARS-CoV-2.261,275,276 Moreover, another novel small molecule compound targeting TMPRSS2, N-0385, exerted equivalent potency against four VOCs, Alpha, Beta, Gamma, and Delta variants, with EC50 ranging from 2.1 to 13.9 nM from SARS-CoV-2 nucleocapsid staining assay, and 2.6–26.5 nM from dsRNA staining assay.251,280
Chloroquine is a widely used antimalaria and anti-autoimmune drug by modulating endosomal pH and disturbing the Clathrin-dependent endocytosis to inhibit pathogen entry into the host cell, for which it manifested broadly anti-pathogen activity.281 Chloroquine was first reported to be active against SARS-CoV-2 in vitro in early 2020, which promoted clinical trials and authorization.264,274,282,283 Later, it faded out of the list of antiviral agents against SARS-CoV-2 with disproved inhibiting infection at the cellular level and protection in clinical practice.284,285,286 Mpro/3CLpro and PLpro play essential roles in transforming viral polyprotein into an active form in SARS-COV-2 replication.287,288,289 Therefore, drugs targeting the two proteins may significantly reduce the viral replication in the host cell.290,291,292,293
Lopinavir was the firstly-reported 3CLpro inhibitor for SARS-CoV-2, repurposed from inhibiting human immunodeficiency virus 1 (HIV-1).294 It has entered Phase III/IV clinical trials (ClinicalTrials.gov number, NCT04738045, NCT04328285, NCT04364022) with preclinical support (estimated EC50 26.63 μM in vivo).271 However, Lopinavir-Ritonavir provided no benefit for severe Covid-19 patients and few studies investigate variants susceptibility to the drug.295 Later, Nirmatrelvir was raised by Pfizer as a novel 3CLpro inhibitor for SARS-CoV-2. The FDA approved the Paxlovid comprising Nirmatrelvir and another molecule Ritonavir, to postpone drug metabolism in vivo, with Emergency Use Authorization (EUA) in December 2021.296,297 Comprehensive studies were performed to investigate its efficacy against SARS-CoV-2 VOCs. Results indicated that Nirmatrelvir-maintained effective (EC50 0.08–0.18 μM—WT and 0.09–0.21 μM—Omicron) against various VOCs, including Omicron. This can be explained by the conserved bind pattern between P132H 3CLpro in complex with Nirmatrelvir under structural simulation.258,272,298,299 GRL-0617 was reported to be a novel drug targeting the PLpro of SARS-CoV-2. Despite the status as a preclinical study, the relatively high efficacy against ancestral strain infection in vitro (EC50 1.9–2.5 μM—WT) may suggest its bright future for further investigation.300,301,302 Studies also pointed out that combinatorial use of 3CLpro and Mpro inhibitors could significantly inhibit SARS-CoV-2 variants, broadening the use of drugs targeting these two proteins.303,304
RdRp inhibitor
RNA-dependent RNA polymerase (RdRp) is employed by RNA viruses, including SARS-CoV-2, to replicate the viral genome and translate the protein, and such critical functions make it a great target for drug design.253,305 Several RdRp inhibitors, repurposed from another pathogen usage, were approved or entered clinical trial as SARS-CoV-2 inhibitors, such as Remdesvir, GS-441524, Molnupiravir, EIDD-1931, and Favipiravir.306
Remdesivir, an adenosine analog created by Gilead as an Ebola virus inhibitor, was the first to show that it could bind SARS-CoV-2 RdRp and disrupt RNA replication, acting as a translocation barrier. Remdesivir was the first drug approved by FDA for COVID-19 in pediatric and adult hospitalized patients in May 2020.262 A parallel study manifested even stronger antiviral activity against all VOCs than Nirmatrelvir (EC50 0.05–0.06 μM—WT and 0.04–0.06 μM—Omicron).272 Although in vitro experiments have suggested that it displays great resistance to VOCs’ mutation,272 a recent meta-analysis revealed that the medication has no effect on COVID-19 protection, and WHO announced a conditional recommendation against remdesivir’s use in hospitalized patients.266,307 These controversial results raised questions about whether Remdesivir was clinically effective in inhibiting SARS-CoV-2 emergent variants and whether the RdRp-targeted drugs were an ideal strategy to overcome the evasion of variants from therapeutic agents.272 The active metabolite of Remdesivir, GS-441524, entered the Phase I clinical trial. It was once used to inhibit the Feline infectious peritonitis virus (FIPV), a coronavirus, targeting the viral RdRp.308,309 GS-441524 was also effective for inhibiting all VOCs, with almost unaffected efficacy for the Omicron variant (EC50 0.15-2.08 μM—WT and 0.42-0.57 μM—Omicron).272 Whereas EIDD-1931 and its active form Molnupiravir (EIDD-2801) is also a nucleoside analog developed by Drug Innovation Ventures from Emory University, also targeting the SARS-CoV-2 RdRp.310,311 Molnupiravir has been approved by FDA for clinical use as the first oral drug treating SARS-CoV-2, and it displayed broad efficacy against multiple RNA viruses.270,294,312 Studies found that it could effectively inhibit the infection of both SARS-CoV-2 ancestral strain and Omicron variant in vitro (EC50 3.57–4.15 μM—WT and 1.86–1.95 μM—Omicron).5,263
The viral inhibitor developed for SARS-CoV-2 mostly manifested maintained efficacy against the VOCs, significantly different from the performance of currently approved antibodies or vaccines.258,272,313 Although non-viral specific drugs such as Camostat could theoretically avoid the mutational impact of SARS-CoV-2 variants due to the independence of viral target,313,314,315,316 their general antiviral capability against authentic virus infection in vitro were relatively inferior to the viral-specific drugs such as Nirmatrelvir.258 However, more evidence has shown that non-structural proteins were also under the pressure of viral mutations.25,159 Despite the outstanding efficacy of currently approved 3CLpro inhibitors like Paxlovid and RdRp inhibitors like Molnupiravir against the Omicron variant, more attention should be paid to the future possible strain with significant mutations located on these viral proteins affecting the drug efficacy.
Impacts of SARS-CoV-2 variants on pandemic control
The epidemiological landscape of variants
The persistent pandemic has witnessed the epidemiological change in virus spread due to various factors, including the emergence of variants, application of vaccines, and disease control policy implemented by global society.317,318,319 Globally, over 450 million infection cases and 6 million death cases were reported by the WHO (February 2022). Europe and the Americas accounted for the most confirmed cases, followed by Asia, Oceania, and Africa. Each SARS-CoV-2 VOC strain superseded the previous one to become the regionally or globally dominant strain during the pandemic. All the VOCs manifested diverse transmission dynamics, responses to vaccines, and impacts on infection outcomes to the difference in molecular profiles and immune escape as we have described. Hence, the epidemiological characteristics of SARS-CoV-2 variants would provide the preliminary impression for the investigator to analyze their distinct behavior in the global spread and immune evasion.
We collected the data from Our World in Data320 and WHO COVID-19 dashboard321 from January 2020 to February 2022 and aligned the time scale to comparatively overview the influence of emerging SARS-CoV-2 VOCs (Fig. 9). The designating timepoint of each VOC was marked as a reference timeline for evaluation. It was shown that the emergence of all VOCs was closely correlated with an increased number of weekly reported infected cases and death cases, which also explained the meaning of “Variant of Concern”. In particular, the Omicron variant manifested astonishing capability in causing emergent infection, with a peak of over 20 million cases per week by the end of February 2022. It was yet to cause more death cases per week than Alpha, Beta Gamma, and Delta variants. Along with the emergence of VOCs, the worldwide application of vaccines also underwent steep expansion. It was shown that the early emergence of variant Alpha and Beta was associated with the initiation of global vaccination. Then, the emergence of the Gamma and Delta variants urged a more rapid application of vaccines around the world, and peak vaccination dose per day (around 40 million doses) could be observed right after the Delta variant was listed in the VOC. Despite that no emergent VOC was announced from June to October 2021, the global vaccination campaign maintained a relatively continuous trend until the emergence of the Omicron variant. With reports on the evasion of Delta, booster vaccination was started but still not widely applied. However, the significant evasion of novel SARS-CoV-2 variants from current antibodies and vaccines by prospective experiments revealed the deficiency of programmed vaccination, and it can be observed that the emergence of Omicron variants was highly associated with the rapid application of booster vaccination, indicating a global consensus on the necessity of improving vaccination efficacy against emergent SARS-CoV-2 variants.
Epidemiological profile of SARS-CoV-2 infection and populational vaccination. a The global distribution of accumulated confirmed COVID-19 cases was reported by the WHO COVID-19 dashboard in February 2022. b The aligned trend of global weekly cases per million people, weekly death per million people, daily vaccination doses per million people, the proportion of programmed vaccination per 100 people, and proportion of booster vaccination per 100 people from January 2020 to February 2022. The timeline of emergent VOC by WHO classification is marked, and the regional data curve is colored differently. c The regional proportion of emerging cases of COVID-19 in February 2022. d The regional data of applied doses for programmed vaccination and booster vaccination. All the data comes from the website Our World in Data (https://ourworldindata.org/coronavirus), WHO COVID-19 dashboard (https://covid19.who.int/), and the figures in related are generated under the CC-BY 4.0 permission
However, the documented infection and vaccination status displayed a prominent imbalance between regions. Europe and the Americas recorded the most infection and death cases but received more doses for both programmed and booster vaccination in the population. The proportions of people with completed programmed vaccination in South America, North America, and Europe were 69%, 61%, and 64%, but the proportion of confirmed cases in the world of these regions was around 50%, 8%, and 8% by the end of February 2022. In comparison, new cases proportion (0.6%) and applied doses for programmed vaccination (12%) in Africa were largely behind the world level. Reasonable speculation about the low number of infected cases and vaccination doses should include that the backward economy and relatively low administrative ability in Africa hinder a broad test for COVID-19 positive cases and wide use of vaccination, which could make this region a black box for actual viral transmission and affection to people’ health and a potential reservoir for viral spread and evolution in the human population. This could also be the case in other countries lacking the broad COVID-19 testing capacity, as the COVID-19 tests per one million people were reported to be low in Asian countries like Afghanistan.
Vaccine efficacy
The observed huge reduction in the neutralizing efficacy of vaccine sera against SARS-CoV-2 variants raises the concern of vaccine efficacy in preventing both infections and, more importantly, severe diseases,322,323 for which clinical studies were performed to evaluate the performance of vaccines in protection from viral infection and virus-induced diseases. The vaccine efficacy or effectiveness (VE) was used to describe the proportionate reduction in cases with endpoint signs among the vaccinated group compared to an unvaccinated group. It is calculated as follows:
Three major endpoints were broadly used to evaluate vaccine efficacy,324 documented infection, symptomatic infection, and more severe hospitalization cases. The four vaccines, BNT162b2, mRNA-1273, AZD1222, and CoronaVac, were all effective against the ancestral strain of SARS-CoV-2, displaying high efficacy in preventing both asymptomatic and symptomatic infection325,326 but are now challenged by the emergence of VOCs. Here, we summarized the VE of four widely applied vaccines against SARS-CoV-2 VOCs and listed their performance in preventing different outcomes after programmed vaccination and booster vaccination (Fig. 10). Unless specified as d/w/m for the time post certain vaccination program, the VE reported in the table was calculated from 14 days post-vaccination until the study reached the research endpoint.
The summary of vaccine efficacy (VE) against infection from SARS-CoV-2 variants after programmed or booster vaccination. Three major outcomes are recorded, including documented infection, symptomatic infection, and hospitalization or more severe cases. Each vaccine’s average VE and 95% confidential interval (CI) against variants are listed. The average VE value is indicated in a red-to-blue color scale to represent the average protective efficacy against each outcome. A bluer color indicates worse protection from the outcome of vaccines. Detailed efficacy data could be referred in Supplementary Table 5
Programmed vaccination
The two mRNA vaccines, BNT162b2 and mRNA-1273, maintained equivalent protection efficacy against Alpha327,328,329,330,331 Beta,221,330,332,333 Gamma,327,330,333,334 and Delta221,327,330,335,336,337 variants in all major endpoints, with only a moderate decrease in protection against documented Beta variant infection.338 For the adenovirus vaccine AZD1222, VE for Beta variant symptomatic infection was found to drop dramatically to 10.4% (95% CI: −76.8 to 54.8) in a prospective study performed in South Africa among 2026 participants,328 while the protection against Alpha,330,339,340 Gamma,330,334 and Delta339,341,342,343 variant was mostly retained. This result was consistent with the observed decline in neutralizing activity of vaccine sera against the Beta variant as described above. For CoronaVac, data was comparatively limited.344,345 One large national cohort study in Chile, including about 2 million 6–16 years old participants, suggests that programmed vaccination of CoronaVac remains effective against Delta variant, reducing 74.5% (95% CI, 73.8–75.2) of symptomatic infection and 91.0% (95% CI, 87.8–93.4) of hospitalization.346 However, CoronaVac VE against Gamma variant 14 days post the second dose was estimated to be 46.8% (95% CI, 38.7–53.8) regarding symptomatic infection and 55.5% (95% CI, 46.5–62.9) regarding hospitalization, suggesting that Gamma variant may dramatically compromise CoronaVac-elicited immune protection.341
Vaccine protection against Omicron variant conferred by complete vaccination of BNT162b2, mRNA-1273, and AZD1222, significantly reduced. Programmed vaccination of AZD1222 was found to offer no significant protection against Omicron documented infection 14 days post-immunization in 2 recent studies,347,348 and only very limited protection against Omicron documented infection within 3 months post-immunization and VE against Omicron symptomatic infection or severe cases after vaccination was around 50% (2–4 weeks post) or 85% (0–3 months).343 For the 2 mRNA vaccines, VE against Omicron documented or symptomatic infection also decreased dramatically within 2-4 weeks after the programmed vaccination, while VE against hospitalization for Omicron maintained over 70% in the same period. A study investigating the recent Omicron outbreak in Hong Kong reported that programmed vaccination of CoronaVac and BNT162b2 offered minimal protection against mild/moderate disease but relatively robust protection (VE all over 70% in different age groups) against severe outcomes.345 In general, current evidence suggests that programmed vaccine protection, especially against documented or symptomatic infection, is substantially evaded by Omicron.
Booster vaccination
Since the clear reduction in VE of programmed vaccination was observed, a booster dose is now widely accepted, especially in countries with a high programmed vaccination implementation rate. VEs of booster vaccination listed in the table are calculated in people receiving homologous boosters compared to unvaccinated people. As discussed above, vaccine protection declined over time. Reports showed that VEs of 2 doses of mRNA vaccines (BNT162b2 or mRNA-1273) against documented Delta variant infection declined to less than 60% over 6 months, and the booster dose reinforced the VE to over 70%.349 In a recent study, VE against symptomatic Delta or Omicron infection was investigated, with booster vaccination taken into consideration.343 It revealed that VEs with AZD1222, BNT162b2, or mRNA-1273 against Omicron symptomatic infection reached approximately 48.9%, 65.5%, or 75.1% within 2–4 weeks post programmed vaccination but then continuously declined to a half from the peak after the 15–19 weeks for AZD1222 and 10–14 weeks for the mRNA vaccines, while VE against symptomatic infection of all three vaccines barely existed 20 weeks post programmed vaccination.
Furthermore, VEs of primary AZD1222 vaccination followed by homologous or heterologous mRNA vaccine boost, and primary mRNA vaccination followed by a homologous or heterologous mRNA vaccine boost were investigated. Booster vaccination effectively elevates VEs against Omicron symptomatic infection to an equivalent or even higher level than VEs post programmed vaccination. The importance of booster vaccination, especially for senior citizens, has also been well addressed in the current Hong Kong outbreak of Omicron, in which the third dose of BNT162b2 or CoronaVac conferred a 71.9% (95% CI: 25.1–89.5) or 96.6% (95% CI: 85.7–99.2) extra protection against severe or fatal COVID-19 to people over 80 years as compared with programmed vaccination.345
In general, despite the reduction in neutralizing titers of vaccine sera against various VOCs, programmed vaccination displayed great performance in protecting people from symptomatic and severe infection of VOCs, except Omicron. All currently approved vaccines in programmed doses did not manifest clear protection from documented infection of the Omicron variant, and more importantly, the efficacy would decrease over time. Application of booster vaccination enhances vaccine efficacy regardless of the booster vaccine type, reinforces the declined protection, and protects against severe infection of the Omicron variant. Nevertheless, it remained a question of how durable the VE was after booster vaccination. More importantly, booster vaccination did not provide extra protection against documented infection of the Omicron variant, which alarmed the global society about the necessity of strict policy for controlling the virus transmission.
Virus transmission in human and animal populations
Multiple studies have identified the increased affinity of SARS-CoV-2 spike protein or RBD with receptor ACE2, leading to a possible higher viral infectivity or faster transmission.36,68,73,350,351,352 The SARS-CoV-2 fitness is a concept to depict the advantages of certain strains during virus spreading and transmission, including stronger binding to ACE2-expressing cells, higher fusion activity, more rapid transmission between hosts, faster replication, and increased viral loads in infected subjects.22,26,353,354,355,356,357 Moreover, as the ACE2 of other mammals share sequence similarity with human ACE2, the mutations located on RBD that affect the binding with human ACE2 could increase the affinity to ACE2 of other mammals and increase their susceptibility to SARS-CoV-2.357,358,359,360,361,362,363,364
Human transmission
As multiple in vitro and in vivo research have identified the viral fitness change under the influence of certain mutations in the spike protein from different VOCs, the transmission efficiency in the population could be altered. For in vitro study, single mutation N501Y, D614G, L452R, and P681R and set mutations from Alpha, Delta, and Omicron in spike protein were found to increase the viral fitness, enhancing both transmissibility and replication, indicating that except evasion from neutralization, mutations of SARS-CoV-2 can bring in advantage for viral transmission.56,206,365,366,367,368,369,370 Hence, populational studies were also performed to investigate the transmission dynamics of SARS-CoV-2 variants (Fig. 11).
Change in SARS-CoV-2 variants transmissibility and patterns for transmission among human and animal populations. The transmissibility of SARS-CoV-2 VOCs increased over strains during human-to-human transmission. And substantial evidence reveals the SARS-CoV-2 transmission from humans to various mammalian animals as reported by OIE (https://www.oie.int/en/what-we-offer/emergency-and-resilience/covid-19/). There have been cases reporting the transmission between animal populations and animals to humans, making wild animals or pets a potential reservoir for viral preservation and evolution. BioRender is used to generate this figure
In general, epidemiologists have studied the change in transmissibility of each of the VOCs compared to the co-circulating VOCs at their time. Alpha, Beta, Gamma, and Delta variants, were reported with higher transmissibility of 43–90%,371 around 50%,43 170–240%,372 or 130–170%373 than their co-circulating VOCs, respectively, and Omicron is estimated to be even more transmissible compared with Delta variant, especially in the vaccinated population.40,371,374,375,376,377,378,379 A recent study conducted in Denmark analyzed the transmission of Delta and Omicron variants among 11,937 households and found that Omicron was 2.61 (95% CI: 2.34–2.90) times or 3.66 (95% CI: 2.65–5.05) times more transmissible than Delta among fully-vaccinated households or booster-vaccinated households, but only merely 1.17 times (95% CI: 0.99–1.38) among unvaccinated people, suggesting that the real-world advantage of Omicron variant in transmission over Delta variant may be largely attributed to the evasion of vaccine-elicited protection of Omicron, especially in the context of broad vaccinated population.
Animal transmission
With increasing studies revealing the sequence similarity between human ACE2 and other mammalian ACE2, growing evidence indicated that SARS-CoV-2 can also infect animals.360,363,364,380,381,382,383 An international organization OIE (World Organization for Animal Health), recorded the investigation of animal infection of SARS-CoV-2. Over 600 outbreaks in animals have been reported worldwide, affecting 19 species in 35 countries (Fig. 7384,385). Currently, animals including the Feline family (Cat,386,387,388,389 Lion, Tiger,390 Snow leopard,391 Fishing, cat,392 Lynx), Dog,358,393,394,395 Mink,111,389 Otter,396 Ferret,394,397,398 Gorilla, Deer,399,400,401,402 Binturong, Coatimundi, Hippo,403 and Hamster404,405 were reported with positive cases of SARS-CoV-2 infection. These results suggested the existence of a human-to-animal transmission pathway.
Moreover, evidence shows animal-to-animal transmission between cats, minks, ferrets, and hamsters, which even shows that the animal population near human activity could be a repository for SARS-CoV-2.394,404,406,407,408 Studies on mink farms indicated that SARS-CoV-2 could transmit between human and mink and back to human,111,408 while non-synonymous mutations could be found in the mink sequence. These results further implied that SARS-CoV-2 evolution could occur during intra-animal populational transmission, and such mutant strain could be transmitted back to humans by animal-to-human transmission.
Besides the study on the general transmission among animals, SARS-CoV-2 variants manifested an inclination of broad host spectrum tropism. In vitro studies have shown that various VOCs displayed higher affinity to murine ACE2, and mice challenged with the authentic virus of Alpha variant developed pathological changes along the respiratory tract compared to the ancestral strain.409 The primary outcomes suggested that the host tropism of SARS-CoV-2 variants in animals tends to expand.
Generally, SARS-CoV-2 variants manifested an increase in transmissibility among the human population,410,411,412 and the Omicron variant, with its remarkable evasion from neutralization, displayed an even stronger populational prevalence.374,413 Furthermore, the findings of animal infection of SARS-CoV-2 further displayed that the virus enjoyed broader transmission due to the wider host tropism and huger reservoir for viral evolution, which could facilitate the emergence of SARS-CoV-2 variants and exacerbate the burden of global cost in containing the pandemic. More evidence was required to demonstrate an enhanced transmission of SARS-CoV-2 variants to animal populations in wild conditions.
Clinical presentation and complications
With the broader distribution, enhanced evasion, and improved transmissibility mentioned above, SARS-CoV-2 variants infection cause more heterogeneous outcomes in patients mainly in two ways: the stronger but comprehensive ability to cause severe diseases due to immune escape from host immunity and faster replication, or the strain-specific mutational impact on viral protein leading to diversity in pathogenesis.
Overall clinical outcome
Risks of hospitalization and severe cases of death related to Alpha,378,414 Beta,414 Gamma,414,415 or Delta416 variant infections increased. Conversely, the Omicron variant shows a decrease in disease severity. This is consistent with the laboratory finding that Omicron infected mice showed reduced replication in respiratory tracts and ameliorated lung pathology compared with ancestral strain or Delta variant infected mice, and weight loss and mortality rate of Omicron infected mice were also lowest.417 Epidemiologically, Omicron variant infection was associated with a lower risk of hospitalization, ICU admission, mechanical ventilation, and a shorter length of hospital stay than Delta variant infection by large.414,418
Acute clinical presentations in common
Clinically, the infection of SARS-CoV-2 is diagnosed with reverse-transcription PCR as the gold standard and could be classified into different clinical types according to clinical manifestations and radiological examinations.419 Severe COVID-19 in adults is defined as meeting any of the following conditions: dyspnea with the respiratory rate of 30/min, blood oxygen saturation of 93%, the ratio of the partial pressure of arterial oxygen to the fraction of inspired oxygen (Pao2:Fio2) 300 mm Hg, or lung imaging showing infiltrates in more than 50% of the field.420 In a large case series published by the Chinese Center for Disease Control and Prevention early in the pandemic, mild, severe, and critical cases accounted for 81%, 14%, and 5%, respectively, and the case-fatality rate was 2.3% in the 44,672 confirmed COVID-19 cases series.421 Nevertheless, severe cases and mortality rates vary as the vaccination campaign progresses. Pandemic lineage shifts variant and experience in treatment accumulates, which should be noticed in the following discussion about clinical manifestations of SARS-COV-2 infection, references of which would inevitably be limited by the representativeness of the study population and time.
In clinical practice, the average incubation period (interval between exposure to symptom onset) is approximately 5 days, and most people develop symptoms within 11.5 days after infection.422 Common symptoms include fever, dry cough, fatigue, and shortness of breath.423,424 Among hospitalized patients, common symptoms were fever (>90%), dry cough (60–86%), shortness of breath (53–80%), and fatigue (23–70%) (Fig. 8).425,426,427 In patients with mild symptomatic infection, fatigue, cough, and fever were reported with occurrence rates of 68%, 60%, and 56% as most frequent symptoms, and an altered sense of smell or taste was reported at a rate of around 3%.428 Besides these symptoms, sore throat, rhinorrhea, diarrhea, nausea, abdominal pain, myalgia, chest pain, dizziness, headache, anosmia, ageusia, testicle pain, and many other symptoms have been reported for SARS-COV-2 infection.429,430,431 As for laboratory findings, common laboratory abnormalities among hospitalized patients include lymphocytopenia (83.2%), thrombocytopenia (36.2%), and leukopenia (33.7%). Most patients had elevated C-reactive protein levels, and some had additional increased levels of aspartate aminotransferase, alanine aminotransferase, creatine kinase, or D-dimer.425,432 Chest radiographs or CT scans found that radiological abnormalities were common among hospitalized patients on admission, including consolidation (59%), ground-glass opacity (71%), and bilateral pulmonary infiltration (75%).427
Infected patients could develop more severe complications,433 especially with risk factors including older age, comorbidities, immunocompromise, obesity, and heavy smoke.433,434,435,436 Most hospitalized patients (91.1%) are diagnosed with pneumonia by physicians on hospital admission, with a mean incubation of 3 days after onset of symptoms, and 3.4–8% of hospitalized patients developed ARDS (acute respiratory distress syndrome426). Extrapulmonary complications were observed in different organs and systems, including myocarditis,437,438,439 arrhythmia,440,441,442 myocardial ischemia443,444,445 regarding the cardiovascular system, acute kidney injury446,447,448 and electrolyte abnormalities,449,450,451 hyperglycemia,452 and ketoacidosis453 in the urinary system, endocrine system, stroke,454,455,456 and encephalitis423,457,458 regarding the neurological system (Fig. 12).459
The overview of SARS-CoV-2 infection affecting the human system and summary of acute and long-term symptoms post. COVID-19 could lead to various symptoms occurring in various body systems except for the expiratory system. The summary table lists the clinical presentations of COVID-19, including severe and mild symptoms during acute infection and long-term symptoms with the occurrence time and rate. The reported occurrence rates of long-term symptoms are average with the 95% confidential interval (CI), and the occurrence time is presented as the time (m - month) post-infection. BioRender is used to generate the human body system presenetation
Long-term post-acute clinical presentations in common
Besides acute symptoms and complications, SARS-CoV-2 infection could leave a long-lasting or fluctuating impact on patients, termed long COVID.460,461 The World Health Organization has defined it as a condition that “occurs in individuals with a history of probable or confirmed SARS-CoV-2 infection, usually 3 months from the onset of COVID-19 with symptoms lasting at least 2 months and cannot be explained by an alternative diagnosis.” It should be noticed that, for practical reasons, many studies investigating post-acute COVID-19 syndrome were based on self-report data, and the claimed symptoms were not validated with a comprehensive inspection to exclude another possible diagnosis, which means that they should not be recognized equivalently as long COVID.462 One important finding is that the post-acute COVID-19 syndrome is not just limited to patients with severe COVID-19.463,464 In a study investigating post-acute syndromes in SARS-CoV-2 symptomatic infection patients over 18 years old, 35% of the 274 symptomatic respondents, with over half of them under 50 years old, reported “not having returned to their usual state of health 2 weeks or more after testing”, and the most common symptoms were fatigue (71%), cough (61%), and headache (61%). Another study included self-reported data of around 100 thousand people with diagnosed COVID-19 infection previously and found that 37.7% of them reported at least one persistent symptom lasting for at least 12 weeks, with fatigue, shortness of breath, myalgia, and insomnia being the most common symptoms.465,466 A meta-analysis analyzed the prevalence of post-acute COVID-19 syndrome symptoms and found that many COVID-19 patients experienced long-lasting post-acute COVID-19 syndrome after recovery from the acute phase of infection.467 The clinical manifestations involve a wide range of systems, and the most common symptoms were fatigue (32%), dyspnea (25%), sleeping disorder (24%), and difficulty in concentrating (22%) within 3–6 months following infection. Many other symptoms, including depression, anxiety, palpitations, effort intolerance, chest pain, diarrhea, joint pain, myalgia, cognitive disorder, headache, and cough, are also found in no less than 10% of convalescent patients in this period. Under the current pandemic of the Omicron variant, infections continuously occur for evasion of vaccine and infection-induced immunity, meaning that a large proportion of people may suffer the post-acute COVID-19 syndrome, which raised the importance of further studies exploring underlying mechanisms and treatment of the post-acute COVID-19 syndrome468 (Fig. 12).
Especially a recent study based on UK Biobank data found that participants infected with SARS-CoV-2 showed a greater reduction in grey matter thickness and global brain size than the controls, with the reduction still being significant after excluding hospitalized cases.469 This study raised concern about the impact of SARS-CoV-2 infection on the neurological system.
Diversity in clinical presentation among VOCs
Evidence regarding the impacts of SARS-CoV-2 variants on certain symptoms was relatively limited. A previous meta-analysis found that anosmia was much more prevalent among populations predominantly infected with the G614 virus (pooled prevalence of 31.8%) as compared with populations predominantly infected with the D614 virus (pooled anosmia prevalence of 5.3%), suggesting that the D614G mutation contributed to the prevalence of anosmia in COVID-19.470 The Omicron variant was linked to a decrease in disease severity, which has been discussed above and was also found to impact clinical manifestation.414 Compared to Delta variant, anosmia was reported less often in Omicron variant infection cases (13% of Omicron cases, 34% of Delta cases, OR: 0.22, 95% CI: 0.21–0.23), and sore throat was reported more often in Omicron variant infection cases (53% of Omicron cases, 34% of Delta cases, OR: 1.93, 95% CI: 1.88–1.98).471 A Laboratory study found that replication of the Omicron variant was similar to the Delta variant in human nasal cultures but lower in lung cells and gut cells.472
With a sharp increase in infected cases worldwide, even mild cases of infection could be extremely troublesome for regions with poor medical resources. A variety of COVID-19 acute infection symptoms could affect almost every critical body system in humans.459 More serious consideration should be given to a wider and more rapid application of booster vaccination globally. The long COVID further suggests that the infection of SARS-CoV-2, even if not deadly, may bring a long-term negative influence on the infected population and decrease the life quality post-infection.473
Discussion and perspective
As the COVID-19 pandemic persisted, various SARS-CoV-2 variants emerged and became a major threat to public health.110,474,475,476 These variants harbored critical mutations in structural and non-structural proteins, affecting protein stability, antigenicity, and function.317,477 The accumulated impact on viral proteins at the single-residue level led to significant changes in biological behaviors of the virus, including infection, transmission, replication, and response to host immunity, and finally influenced the clinical phenotypes and presentations post-viral infection. Therefore, systematic studies connecting the molecular, biological, epidemiological, and clinical evidence of SARS-CoV-2 variants would be greatly demanded to provide insightful and constructive ideas for fundamental research on the virus itself and pandemic disease control.
The sequencing data from SARS-CoV-2 isolated samples provided evolutionary trace and revealed a newly-emergent strain. The evolutionary and sequential abundance analysis witnessed the Omicron strain as the most-diverged strain from the ancestors and currently the most prevalent strain. As more mutations were found, the greater concern was given to the surveillance of mutational impact on the critical viral proteins.478 Among the structural proteins of SARS-CoV-2, spike protein was regarded as the most important target determining the fate of viral recognition and fusion due to its binding with host receptor ACE2, for which mutation located on the spike protein exerted versatile influence on the structural characteristics, including receptor affinity, antibody binding, structural stability, and protein yield.479 These factors had an enormous and direct impact on the viral activity and response to host immunity and highlighted the value of current effort in closely monitoring the critical mutations leading to a significant alteration in vitro structural and biological features of spike protein from variants.480 Whereas, less attention was given to the accumulated mutations in non-structural proteins with extremely important biological functions during the virus life cycle, as they were by the large recognized stable invariants. However, the discovered mutation locus from the variants of concern at non-structural proteins with reported evidence showing the impact on structural stability doubts their “uninfluenced” prospect as a target for drug development. More experimental evidence was required to reveal the mutational impact on these key viral proteins in modeling the structural characters and mediating viral replication.
Since the mutations brought new molecular characteristics to key proteins of SARS-CoV-2, therapeutic strategies against viral infection confronted more challenges. It has been widely reported that all the VOCs manifested varied immune escape, especially the Omicron.50,323,481,482 Extensive effort has been put into revealing both molecular and immunological basis of the resistance of these variants to an antibody or vaccine sera targeting spike protein, and increasing evidence has indicated the connection between the structural change in protein-antibody complex and the diminished neutralizing capability of antibodies.483,484,485,486 These results possibly suggested a positive selection of emergent strains harboring mutations in spike protein with a potent immune escape from humoral immunity in the global population. The breakthrough of currently available recombinant or vaccine-induced antibodies has led to growing worried about the future development of antibody-based therapy.190,322,487 In comparison, recently, good news came from the small molecule drug targeting non-structural proteins such as 3CL protease inhibitor Nirmatrelvir and RdRp inhibitor Molnupiravir, as they manifested well-maintained antiviral activity against SARS-CoV-2 variants including Omicron during in vitro experiment. Therefore, closer monitor of drug resistance due to mutational change in non-structural protein targets and more high-level clinical evidence of drug efficacy are in demand to provide clear guidance in the use of anti-SARS-CoV-2 drugs against variants,488,489,490 while a “game-changer” method for developing variant-effective antibody is under great expectation.
With the changes in response to current therapeutic agents, SARS-CoV-2 variants exhibited various epidemiological and clinical manifestations differed by strain.425 The global epidemiological profile has shown an unprecedentedly rapid spread of the Omicron variant. Interestingly, global vaccination against SARS-CoV-2 rapidly increased since the first identification of the Alpha variant as VOC. In particular, the distribution of emerging infection and vaccination quota displayed a huge geographical imbalance and a mismatched relationship. Moreover, the vaccination efficacy confronted great challenges from the SARS-CoV-2 variants. Clear breakthrough has been observed among all vaccines for their programmed vaccination, and booster vaccination cannot provide significant sterile immunity toward the infection but higher efficacy in preventing symptomatic and severe cases. However, the duration of protection from the booster vaccination could be a key point for its efficacy against SARS-CoV-2, and more studies are required to answer this question.491,492 Besides, the higher transmissibility among the human population and wider host tropism of SARS-CoV-2 urged more attention to the transmission dynamics of the virus between humans or between humans and animals.381,492 As the potential virus bank, animal infection of SARS-CoV-2 might become an “Achilles’ heel” for disease surveillance and containment.493 Although an overall reduction in the Omicron variant-related death rate was observed, the increasing number of infected cases raised to worry about the global medical resources for curing the symptomatic infection. As more evidence has demonstrated the acute and long-term impact of SARS-CoV-2 infection,420,443,494,495,496,497 greater effort should be given to reduce populational infection instead of merely focusing on the number of death cases. Governments should realize the importance of collaboration in COVID-19 disease control.
References
Koelle, K., Martin, M. A., Antia, R., Lopman, B. & Dean, N. E. The changing epidemiology of SARS-CoV-2. Science 375, 1116–1121 (2022).
Markov, P. V., Katzourakis, A. & Stilianakis, N. I. Antigenic evolution will lead to new SARS-CoV-2 variants with unpredictable severity. Nat. Rev. Micobiol. 20, 251–252 (2022).
Flemming, A. Omicron, the great escape artist. Nat. Rev. Immunol. 22, 75 (2022).
Planas, D. et al. Considerable escape of SARS-CoV-2 Omicron to antibody neutralization. Nature 602, 671–675 (2022).
Cao, Y. et al. Omicron escapes the majority of existing SARS-CoV-2 neutralizing antibodies. Nature 602, 657–663 (2022).
Grubaugh, N. D. & Cobey, S. Of variants and vaccines. Cell 184, 6222–6223 (2021).
Bushman, M., Kahn, R., Taylor, B. P., Lipsitch, M. & Hanage, W. P. Population impact of SARS-CoV-2 variants with enhanced transmissibility and/or partial immune escape. Cell 184, 6229–6242.e18 (2021).
Schmidt, F. et al. High genetic barrier to SARS-CoV-2 polyclonal neutralizing antibody escape. Nature 600, 512–516 (2021).
Oude, M. B. et al. The next phase of SARS-CoV-2 surveillance: real-time molecular epidemiology. Nat. Med. 27, 1518–1524 (2021).
Harvey, W. T. et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat. Rev. Microbiol. 19, 409–424 (2021).
Li, Q. et al. The impact of mutations in SARS-CoV-2 spike on viral infectivity and antigenicity. Cell 182, 1284–1294.e9 (2020).
Yurkovetskiy, L. et al. Structural and functional analysis of the D614G SARS-CoV-2 spike protein variant. Cell 183, 739–751.e8 (2020).
Zhang, J. et al. Structural impact on SARS-CoV-2 spike protein by D614G substitution. Science 372, 525–530 (2021).
Chan, J. F. et al. A familial cluster of pneumonia associated with the 2019 novel coronavirus indicating person-to-person transmission: a study of a family cluster. Lancet 395, 514–523 (2020).
Espenhain, L. et al. Epidemiological characterisation of the first 785 SARS-CoV-2 Omicron variant cases in Denmark, December 2021. Eur. Surveill. 26, 2101146 (2021).
Poon, L. & Peiris, M. Emergence of a novel human coronavirus threatening human health. Nat. Med. 26, 317–319 (2020).
Cucinotta, D. & Vanelli, M. WHO declares COVID-19 a pandemic. Acta Biomed. 91, 157–160 (2020).
Ghosh, N., Nandi, S. & Saha, I. A review on evolution of emerging SARS-CoV-2 variants based on spike glycoprotein. Int. Immunopharmocol. 105, 108565 (2022).
Kumar, S. U. et al. A review of novel coronavirus disease (COVID-19): based on genomic structure, phylogeny, current shreds of evidence, candidate vaccines, and drug repurposing. 3 Biotech. 11, 198 (2021).
Gupta, A. M., Chakrabarti, J. & Mandal, S. Non-synonymous mutations of SARS-CoV-2 leads epitope loss and segregates its variants. Microbes Infect. 22, 598–607 (2020).
Duffy, S. Why are RNA virus mutation rates so damn high? PLoS Biol. 16, e3000003 (2018).
Domingo, E. & Holland, J. J. RNA virus mutations and fitness for survival. Annu. Rev. Microbiol. 51, 151–178 (1997).
Karthic, A. et al. Computational analysis reveals monomethylated triazolopyrimidine as a novel inhibitor of SARS-CoV-2 RNA-dependent RNA polymerase (RdRp). Molecules 27, 801 (2022).
Jung, L. S., Gund, T. M. & Narayan, M. Comparison of binding site of remdesivir and its metabolites with NSP12-NSP7-NSP8, and NSP3 of SARS CoV-2 virus and alternative potential drugs for COVID-19 treatment. Protein J. 39, 619–630 (2020).
Anwar, M. Z., Lodhi, M. S., Khan, M. T., Khan, M. I. & Sharif, S. Coronavirus genomes and unique mutations in structural and non-structural proteins in Pakistani SARS-CoV-2 delta variants during the fourth wave of the pandemic. Genes 13, 552 (2022).
Banerjee, A., Mossman, K. & Grandvaux, N. Molecular determinants of SARS-CoV-2 variants. Trends Microbiol. 29, 871–873 (2021).
Mahmood, T. B. et al. A next generation sequencing (NGS) analysis to reveal genomic and proteomic mutation landscapes of SARS-CoV-2 in South Asia. Curr. Res. Microb. Sci. 2, 100065–100065 (2021).
Hamed, S. M., Elkhatib, W. F., Khairalla, A. S. & Noreddin, A. M. Global dynamics of SARS-CoV-2 clades and their relation to COVID-19 epidemiology. Sci. Rep. 11, 8435 (2021).
O’Toole, Á., Pybus, O. G., Abram, M. E., Kelly, E. J. & Rambaut, A. Pango lineage designation and assignment using SARS-CoV-2 spike gene nucleotide sequences. BMC Genom. 23, 121 (2022).
Rambaut, A. et al. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 5, 1403–1407 (2020).
Chakraborty, C. et al. D614G mutation eventuates in all VOI and VOC in SARS-CoV-2: is it part of the positive selection pioneered by Darwin? Mol. Ther. Nucleic Acids 26, 237–241 (2021).
Bhattacharya, M., Chatterjee, S., Sharma, A. R., Agoramoorthy, G. & Chakraborty, C. D614G mutation and SARS-CoV-2: impact on S-protein structure, function, infectivity, and immunity. Appl. Microbiol. Biotechnol. 105, 9035–9045 (2021).
Gobeil, S. M. et al. D614G mutation alters SARS-CoV-2 spike conformation and enhances protease cleavage at the S1/S2 junction. Cell Rep. 34, 108630 (2021).
Ozono, S. et al. SARS-CoV-2 D614G spike mutation increases entry efficiency with enhanced ACE2-binding affinity. Nat. Commun. 12, 848 (2021).
Hou, Y. J. et al. SARS-CoV-2 D614G variant exhibits enhanced replication ex vivo and earlier transmission in vivo. Science 370, 1446–1468 (2020).
Zhang, L. et al. SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nat. Commun. 11, 6013 (2020).
Cele, S. et al. Escape of SARS-CoV-2 501Y.V2 from neutralization by convalescent plasma. Nature 593, 142–146 (2021).
Luan, B., Wang, H. & Huynh, T. Enhanced binding of the N501Y-mutated SARS-CoV-2 spike protein to the human ACE2 receptor: insights from molecular dynamics simulations. FEBS Lett. 595, 1454–1461 (2021).
Galloway, S. E. et al. Emergence of SARS-CoV-2 B.1.1.7 lineage—United States, December 29, 2020–January 12, 2021. Morb. Mortal. Wkly Rep. 70, 95–99 (2021).
Volz, E. et al. Assessing transmissibility of SARS-CoV-2 lineage B.1.1.7 in England. Nature 593, 266–269 (2021).
Lauring, A. S. & Malani, P. N. Variants of SARS-CoV-2. JAMA 326, 880 (2021).
Chan, K. W., Wong, V. T. & Tang, S. COVID-19: an update on the epidemiological, clinical, preventive and therapeutic evidence and guidelines of Integrative Chinese-Western medicine for the management of 2019 novel coronavirus disease. Am. J. Chin. Med. 48, 737–762 (2020).
Tegally, H. et al. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 592, 438–443 (2021).
Naveca, F. G. et al. COVID-19 in Amazonas, Brazil, was driven by the persistence of endemic lineages and P.1 emergence. Nat. Med. 27, 1230–1238 (2021).
Zimerman, R. A. et al. Comparative genomics and characterization of SARS-CoV-2 P.1 (gamma) variant of concern from Amazonas, Brazil. Front. Med. 9, 806611 (2022).
Mlcochova, P. et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 599, 114–119 (2021).
Yadav, P. D. et al. SARS-CoV-2 Kappa Variant Shows Pathogenicity in a Syrian Hamster Model. Vector borne and zoonotic dis. 22, 289–296 (2022).
CDC COVID-19 Response Team. SARS-CoV-2 B.1.1.529 (Omicron) variant—United States, December 1–8, 2021. MMWR Morb. Mortal. Wkly Rep. 70, 1731–1734 (2021).
Jansen, L. et al. Investigation of a SARS-CoV-2 B.1.1.529 (Omicron) variant cluster— Nebraska, November–December 2021. MMWR Morb. Mortal. Wkly Rep. 70, 1782–1784 (2021).
Karim, S. & Karim, Q. A. Omicron SARS-CoV-2 variant: a new chapter in the COVID-19 pandemic. Lancet 398, 2126–2128 (2021).
Wang, L. & Cheng, G. Sequence analysis of the emerging SARS-CoV-2 variant omicron in South Africa. J. Med. Virol. 94, 1728–1733 (2022).
Thakur, V. & Ratho, R. K. OMICRON (B.1.1.529): a new SARS-CoV-2 variant of concern mounting worldwide fear. J. Med. Virol. 94, 1821–1824 (2022).
Del, R. C., Omer, S. B. & Malani, P. N. Winter of omicron—the evolving COVID-19 pandemic. JAMA 327, 319–320 (2022).
Brandal, L. T. et al. Outbreak caused by the SARS-CoV-2 omicron variant in Norway, November to December 2021. Eur. Surveill. 26, 2101147 (2021).
Kumar, S., Thambiraja, T. S., Karuppanan, K. & Subramaniam, G. Omicron and delta variant of SARS-CoV-2: a comparative computational study of spike protein. J. Med. Virol. 94, 1641–1649 (2022).
Suryawanshi, R. K. et al. Limited cross-variant immunity from SARS-CoV-2 Omicron without vaccination. Nature https://doi.org/10.1038/s41586-022-04865-0 (2022).
Bhattacharya, M., Sharma, A. R., Dhama, K., Agoramoorthy, G. & Chakraborty, C. Omicron variant (B.1.1.529) of SARS-CoV-2: understanding mutations in the genome, S-glycoprotein, and antibody-binding regions. Geroscience 44, 619–637 (2022).
Song, Y. & Masaki, F. Preparation for the challenge of heavily mutated Omicron variant. Clin. Transl. Med. 11, e679 (2021).
Cele, S. et al. Omicron extensively but incompletely escapes Pfizer BNT162b2 neutralization. Nature 602, 654–656 (2022).
Dejnirattisai, W. et al. SARS-CoV-2 Omicron-B.1.1.529 leads to widespread escape from neutralizing antibody responses. Cell 185, 467–484.e15 (2022).
Khan, A. et al. SARS-CoV-2 UK, South African and Brazilian Variants in Karachi-Pakistan. Front. Mol. Biosci. 8, 724208 (2021).
Khailany, R. A., Safdar, M. & Ozaslan, M. Genomic characterization of a novel SARS-CoV-2. Gene Rep. 19, 100682 (2020).
Voloch, C. M. et al. Genomic characterization of a novel SARS-CoV-2 lineage from Rio de Janeiro, Brazil. J. Virol. 95, e00119–e00121 (2021).
Lu, R. et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: implications for virus origins and receptor binding. Lancet 395, 565–574 (2020).
Arya, R. et al. Structural insights into SARS-CoV-2 proteins. J. Mol. Biol. 433, 166725–166725 (2021).
Gralinski, L. E. & Menachery, V. D. Return of the Coronavirus: 2019-nCoV. Viruses 12, 135 (2020).
Walls, A. C. et al. Unexpected receptor functional mimicry elucidates activation of coronavirus fusion. Cell 183, 1732 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280.e8 (2020).
Shang, J. et al. Structural basis of receptor recognition by SARS-CoV-2. Nature 581, 221–224 (2020).
Yan, R. et al. Structural basis for the different states of the spike protein of SARS-CoV-2 in complex with ACE2. Cell Res. 31, 717–719 (2021).
Mittal, A., Khattri, A. & Verma, V. Structural and antigenic variations in the spike protein of emerging SARS-CoV-2 variants. PLoS Pathog. 18, e1010260 (2022).
Coutard, B. et al. The spike glycoprotein of the new coronavirus 2019-nCoV contains a furin-like cleavage site absent in CoV of the same clade. Antivir. Res. 176, 104742 (2020).
Walls, A. C. et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell 181, 281–292.e6 (2020).
Hoffmann, M., Kleine-Weber, H. & Pöhlmann, S. A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol. Cell 78, 779–784.e5 (2020).
Jaimes, J. A., Millet, J. K. & Whittaker, G. R. Proteolytic cleavage of the SARS-CoV-2 spike protein and the role of the novel S1/S2 site. iScience 23, 101212 (2020).
Xia, S. et al. The role of furin cleavage site in SARS-CoV-2 spike protein-mediated membrane fusion in the presence or absence of trypsin. Signal Transduct. Target Ther. 5, 92 (2020).
Adhikari, P. et al. Intra- and intermolecular atomic-scale interactions in the receptor binding domain of SARS-CoV-2 spike protein: implication for ACE2 receptor binding. Phys. Chem. Chem. Phys. 22, 18272–18283 (2020).
Ortega, J. T., Serrano, M. L., Pujol, F. H. & Rangel, H. R. Role of changes in SARS-CoV-2 spike protein in the interaction with the human ACE2 receptor: an in silico analysis. EXCLI J. 19, 410–417 (2020).
Lan, J. et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 581, 215–220 (2020).
Paul, D., Pyne, N. & Paul, S. Mutation profile of SARS-CoV-2 spike protein and identification of potential multiple epitopes within spike protein for vaccine development against SARS-CoV-2. Virusdisease 32, 703–726 (2021).
Yuan, M. et al. A highly conserved cryptic epitope in the receptor binding domains of SARS-CoV-2 and SARS-CoV. Science 368, 630–633 (2020).
Tai, W. et al. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell. Mol. Immunol. 17, 613–620 (2020).
Premkumar, L. et al. The receptor binding domain of the viral spike protein is an immunodominant and highly specific target of antibodies in SARS-CoV-2 patients. Sci. Immunol. 5, 48 (2020).
Panda, P. K. et al. Structure-based drug designing and immunoinformatics approach for SARS-CoV-2. Sci. Adv. 6, b8097 (2020).
Vishwakarma, P. et al. Severe acute respiratory syndrome coronavirus 2 spike protein based novel epitopes induce potent immune responses in vivo and inhibit viral replication in vitro. Front. Immunol. 12, 613045 (2021).
Coghi, P. et al. Exploring SARS-CoV-2 delta variant spike protein receptor-binding domain (RBD) as a target for tanshinones and antimalarials. Nat. Prod. Res. 25, 1–6 (2022).
Tian, X. et al. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg. Microbes Infect. 9, 382–385 (2020).
Zost, S. J. et al. Rapid isolation and profiling of a diverse panel of human monoclonal antibodies targeting the SARS-CoV-2 spike protein. Nat. Med. 26, 1422–1427 (2020).
Prajapat, M. et al. Drug targets for corona virus: a systematic review. Indian J. Pharm. 52, 56–65 (2020).
Mukherjee, R. Global efforts on vaccines for COVID-19: since, sooner or later, we all will catch the coronavirus. J. Biosci. 45, 1 (2020).
Ralph, R. et al. 2019-nCoV (Wuhan virus), a novel Coronavirus: human-to-human transmission, travel-related cases, and vaccine readiness. J. Infect. Dev. Ctries 14, 3–17 (2020).
Yadav, P. D. & Kumar, S. Global emergence of SARS-CoV-2 variants: new foresight needed for improved vaccine efficacy. Lancet Infect. Dis. 22, 298–299 (2022).
Chen, Z. et al. Global landscape of SARS-CoV-2 genomic surveillance and data sharing. Nat. Genet. 54, 499–507 (2022).
Peters, M. H., Bastidas, O., Kokron, D. S. & Henze, C. E. Static all-atom energetic mappings of the SARS-Cov-2 spike protein and dynamic stability analysis of “Up” versus “Down” protomer states. PLoS ONE 15, e241168 (2020).
Teruel, N., Mailhot, O. & Najmanovich, R. J. Modelling conformational state dynamics and its role on infection for SARS-CoV-2 Spike protein variants. PLoS Comput. Biol. 17, e1009286 (2021).
Cai, Y. et al. Distinct conformational states of SARS-CoV-2 spike protein. Science 369, 1586–1592 (2020).
Saito, A. et al. Enhanced fusogenicity and pathogenicity of SARS-CoV-2 Delta P681R mutation. Nature 602, 300–306 (2022).
Barton, M. I. et al. Effects of common mutations in the SARS-CoV-2 Spike RBD and its ligand, the human ACE2 receptor on binding affinity and kinetics. eLife 10, e70658 (2021).
Kim, Y. et al. Effects of Spike Mutations in SARS-CoV-2 Variants of Concern on Human or Animal ACE2-Mediated Virus Entry and Neutralization. Microbiol. Spectr. e0178921 (2022).
Peters, M. H., Bastidas, O., Kokron, D. S. & Henze, C. E. Transformations, lineage comparisons, and analysis of down-to-up protomer states of variants of the SARS-CoV-2 prefusion spike protein, including the UK variant B.1.1.7. Microbiol. Spectr. 9, e3021 (2021).
Zahradník, J. et al. SARS-CoV-2 variant prediction and antiviral drug design are enabled by RBD in vitro evolution. Nat. Microbiol. 6, 1188–1198 (2021).
Ye, G., Liu, B. & Li, F. Cryo-EM structure of a SARS-CoV-2 omicron spike protein ectodomain. Nat. Commun. 13, 1214 (2022).
Starr, T. N. et al. Deep mutational scanning of SARS-CoV-2 receptor binding domain reveals constraints on folding and ACE2 binding. Cell 182, 1295–1310.e20 (2020).
Gupta, D. et al. Structural and functional insights into the spike protein mutations of emerging SARS-CoV-2 variants. Cell. Mol. Life Sci. 78, 7967–7989 (2021).
Tanaka, S. et al. An ACE2 Triple Decoy that neutralizes SARS-CoV-2 shows enhanced affinity for virus variants. Sci Rep. 11, 12740 https://doi.org/10.1038/s41598-021-91809-9 (2021).
Thomson, E. C. et al. Circulating SARS-CoV-2 spike N439K variants maintain fitness while evading antibody-mediated immunity. Cell 184, 1171–1187.e20 (2021).
Mishra, T. et al. SARS-CoV-2 spike E156G/Δ157-158 mutations contribute to increased infectivity and immune escape. Life Sci. Alliance 5, e202201415 (2022).
Cheng, M. H. et al. Impact of new variants on SARS-CoV-2 infectivity and neutralization: A molecular assessment of the alterations in the spike–host protein interactions. iScience 25, 103939 (2022).
Fontanet, A. et al. SARS-CoV-2 variants and ending the COVID-19 pandemic. Lancet 397, 952–954 (2021).
Callaway, E. Heavily mutated Omicron variant puts scientists on alert. Nature 600, 21 (2021).
Ren, W. et al. Mutation Y453F in the spike protein of SARS-CoV-2 enhances interaction with the mink ACE2 receptor for host adaption. PLoS Pathog. 17, e1010053 (2021).
Prabakaran, P. et al. Structure of severe acute respiratory syndrome coronavirus receptor-binding domain complexed with neutralizing antibody. J. Biol. Chem. 281, 15829–15836 (2006).
Andreano, E. et al. SARS-CoV-2 escape from a highly neutralizing COVID-19 convalescent plasma. Proc. Natl Acad. Sci. USA 118, e2103154118 (2021).
Kemp, S. A. et al. SARS-CoV-2 evolution during treatment of chronic infection. Nature 592, 277–282 (2021).
McCarthy, K. R. et al. Recurrent deletions in the SARS-CoV-2 spike glycoprotein drive antibody escape. Science 371, 1139–1142 (2021).
Meng, B. et al. Recurrent emergence of SARS-CoV-2 spike deletion H69/V70 and its role in the Alpha variant B.1.1.7. Cell Rep. 35, 109292 (2021).
Planas, D. et al. Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature 596, 276–280 (2021).
Yu, X. et al. Reduced sensitivity of SARS-CoV-2 Omicron variant to antibody neutralization elicited by booster vaccination. Cell. Discov. 8, 4 (2022).
Wang, M. et al. Reduced sensitivity of the SARS-CoV-2 Lambda variant to monoclonal antibodies and neutralizing antibodies induced by infection and vaccination. Emerg. Microbes Infect. 11, 18–29 (2022).
Starr, T. N., Greaney, A. J., Dingens, A. S. & Bloom, J. D. Complete map of SARS-CoV-2 RBD mutations that escape the monoclonal antibody LY-CoV555 and its cocktail with LY-CoV016. Cell Rep. Med. 2, 100255 (2021).
Wang, Z. et al. mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants. Nature 592, 616–622 (2021).
Iketani, S. et al. Antibody evasion properties of SARS-CoV-2 Omicron sublineages. Nature 604, 553–556 (2022).
Rahmani, A., Baee, M., Saleki, K., Moradi, S. & Nouri, H. R., Applying high throughput and comprehensive immunoinformatics approaches to design a trivalent subunit vaccine for induction of immune response against emerging human coronaviruses SARS-CoV, MERS-CoV and SARS-CoV-2. J. Biomol. Struct. Dyn. 1–17 (2021).
Cherian, S. et al. SARS-CoV-2 Spike Mutations, L452R, T478K, E484Q and P681R, in the Second Wave of COVID-19 in Maharashtra, India. Microorganisms 9, 1542 (2021).
Li, D. et al. In vitro and in vivo functions of SARS-CoV-2 infection-enhancing and neutralizing antibodies. Cell 184, 4203–4219.e32 (2021).
Bates, T. A. et al. Antibody response and variant cross-neutralization after SARS-CoV-2 breakthrough infection. JAMA 327, 179–181 (2022).
Mader, A. L. et al. Omicron’s binding to sotrovimab, casirivimab, imdevimab, CR3022, and sera from previously infected or vaccinated individuals. iScience 25, 104076 (2022).
McCallum, M. et al. N-terminal domain antigenic mapping reveals a site of vulnerability for SARS-CoV-2. Cell 184, 2332–2347.e16 (2021).
Quarleri, J., Galvan, V. & Delpino, M. V. Omicron variant of the SARS-CoV-2: a quest to define the consequences of its high mutational load. Geroscience 44, 53–56 (2022).
Gong, S. Y. et al. Contribution of single mutations to selected SARS-CoV-2 emerging variants spike antigenicity. Virology 563, 134–145 (2021).
Laha, S. et al. Characterizations of SARS-CoV-2 mutational profile, spike protein stability and viral transmission. Infect. Genet. Evol. 85, 104445 (2020).
Johnson, B. A. et al. Loss of furin cleavage site attenuates SARS-CoV-2 pathogenesis. Nature 591, 293–299 (2021).
Haynes, W. A. et al. High-resolution epitope mapping and characterization of SARS-CoV-2 antibodies in large cohorts of subjects with COVID-19. Commun. Biol. 4, 1317 (2021).
Shah, A. et al. Comparative mutational analysis of SARS-CoV-2 isolates from Pakistan and structural–functional implications using computational modelling and simulation approaches. Comput. Biol. Med. 141, 105170 (2022).
Schoeman, D. & Fielding, B. C. Coronavirus envelope protein: current knowledge. Virol. J. 16, 69 (2019).
Cao, Y. et al. Computational study of the ion and water permeation and transport mechanisms of the SARS-CoV-2 pentameric E protein channel. Front. Mol. Biosci. 7, 565797 (2020).
Marques-Pereira, C. et al. SARS-CoV-2 membrane protein: from genomic data to structural new insights. Int. J. Mol. Sci. 23, 2986 (2022).
Zeng, W. et al. Biochemical characterization of SARS-CoV-2 nucleocapsid protein. Biochem. Biophys. Res. Commun. 527, 618–623 (2020).
Fu, Y. Z. et al. SARS-CoV-2 membrane glycoprotein M antagonizes the MAVS-mediated innate antiviral response. Cell. Mol. Immunol. 18, 613–620 (2021).
Kumar, B. K. et al. Deletion in the C-terminal region of the envelope glycoprotein in some of the Indian SARS-CoV-2 genome. Virus Res. 291, 198222 (2021).
Sarkar, M. & Saha, S. Structural insight into the role of novel SARS-CoV-2 E protein: a potential target for vaccine development and other therapeutic strategies. PLoS ONE 15, e237300 (2020).
Collins, L. T. et al. Elucidation of SARS-Cov-2 budding mechanisms through molecular dynamics simulations of M and E protein complexes. J. Phys. Chem. Lett. 12, 12249–12255 (2021).
Mou, K. et al. Emerging mutations in envelope protein of SARS-CoV-2 and their effect on thermodynamic properties. Inf. Med. Unlocked 25, 100675 (2021).
Jian, M. J. et al. SARS-CoV-2 variants with T135I nucleocapsid mutations may affect antigen test performance. Int. J. Infect. Dis. 114, 112–114 (2022).
Quaglia, F. et al. SARS-CoV-2 variants preferentially emerge at intrinsically disordered protein sites helping immune evasion. FEBS J. https://doi.org/10.1111/febs.16379 (2022).
Mourier, T. et al. SARS-CoV-2 genomes from Saudi Arabia implicate nucleocapsid mutations in host response and increased viral load. Nat. Commun. 13, 601 (2022).
Carlson, C. R. et al. Phosphoregulation of phase separation by the SARS-CoV-2 N protein suggests a biophysical basis for its dual functions. Mol. Cell 80, 1092–1103.e4 (2020).
Zhao, H. et al. Energetic and structural features of SARS-CoV-2 N-protein co-assemblies with nucleic acids. iScience 24, 102523 (2021).
Khan, M. T. et al. Structures of SARS-CoV-2 RNA-binding proteins and therapeutic targets. Intervirology 64, 55–68 (2021).
Ziebuhr, J. The coronavirus replicase. Curr. Phase Separ. Top. Microbiol. Immunol. 287, 57–94 (2005).
Finkel, Y. et al. The coding capacity of SARS-CoV-2. Nature 589, 125–130 (2021).
Shin, D. et al. Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 587, 657–662 (2020).
Wu, C. R., Yin, W. C., Jiang, Y. & Xu, H. E. Structure genomics of SARS-CoV-2 and its Omicron variant: drug design templates for COVID-19. Acta Pharmacol. Sin. https://doi.org/10.1038/s41401-021-00851-w (2022).
Dai, W. et al. Structure-based design of antiviral drug candidates targeting the SARS-CoV-2 main protease. Science 368, 1331–1335 (2020).
Naidu, S., Tripathi, Y. B., Shree, P., Clemens, R. A. & Naidu, A. S., Phytonutrient iInhibitors of SARS-CoV-2/NSP5-encoded main protease (M(pro)) autocleavage enzyme critical for COVID-19 pathogenesis. J. Diet Suppl. https://doi.org/10.1080/19390211.2021.2006388 (2021).
Yan, L. et al. Cryo-EM structure of an extended SARS-CoV-2 replication and transcription complex reveals an intermediate state in Cap synthesis. Cell 184, 184–193.e10 (2021).
Subissi, L. et al. One severe acute respiratory syndrome coronavirus protein complex integrates processive RNA polymerase and exonuclease activities. Proc. Natl Acad. Sci. USA 111, E3900–E3909 (2014).
Rahnavard, A. et al. Epidemiological associations with genomic variation in SARS-CoV-2. Sci. Rep. 11, 23023–23023 (2021).
Mou, K. et al. Emerging mutations in Nsp1 of SARS-CoV-2 and their effect on the structural stability. Pathogens 10, 1285 (2021).
Hartley, P. D. et al. Genomic surveillance of Nevada patients revealed prevalence of unique SARS-CoV-2 variants bearing mutations in the RdRp gene. J. Genet. Genom. 48, 40–51 (2021).
Vogels, C. et al. Multiplex qPCR discriminates variants of concern to enhance global surveillance of SARS-CoV-2. PLoS Biol. 19, e3001236 (2021).
Shalayel, M. H., Al-Mazaideh, G. M., Aladaileh, S. H., Al-Swailmi, F. K. & Al-Thiabat, M. G. Vitamin D is a potential inhibitor of COVID-19: in silico molecular docking to the binding site of SARS-CoV-2 endoribonuclease Nsp15. Pak. J. Pharm. Sci. 33, 2179–2186 (2020).
Biswal, M. et al. Two conserved oligomer interfaces of NSP7 and NSP8 underpin the dynamic assembly of SARS-CoV-2 RdRP. Nucleic Acids Res. 49, 5956–5966 (2021).
Corti, D., Purcell, L. A., Snell, G. & Veesler, D. Tackling COVID-19 with neutralizing monoclonal antibodies. Cell 184, 4593–4595 (2021).
Pinto, D. et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature 583, 290–295 (2020).
Tortorici, M. A. et al. Ultrapotent human antibodies protect against SARS-CoV-2 challenge via multiple mechanisms. Science 370, 950–957 (2020).
Tso, F. Y. et al. Presence of antibody-dependent cellular cytotoxicity (ADCC) against SARS-CoV-2 in COVID-19 plasma. PLoS ONE 16, e247640 (2021).
Chen, X. et al. The development and kinetics of functional antibody-dependent cell-mediated cytotoxicity (ADCC) to SARS-CoV-2 spike protein. Virology 559, 1–9 (2021).
Reis, G. et al. Effect of early treatment with hydroxychloroquine or lopinavir and ritonavir on risk of hospitalization among patients with COVID-19: the TOGETHER Randomized Clinical Trial. JAMA Netw. Open 4, e216468 (2021).
Cohen, M. S. et al. Effect of bamlanivimab vs. placebo on incidence of COVID-19 among residents and staff of skilled nursing and assisted living facilities: a Randomized Clinical Trial. JAMA 326, 46–55 (2021).
Dougan, M. et al. Bamlanivimab plus Etesevimab in mild or moderate Covid-19. N. Engl. J. Med. 385, 1382–1392 (2021).
Lok, S. M. An NTD supersite of attack. Cell Host Microbe 29, 744–746 (2021).
Amanat, F. et al. SARS-CoV-2 mRNA vaccination induces functionally diverse antibodies to NTD, RBD, and S2. Cell 184, 3936–3948.e10 (2021).
Du, S. et al. Structurally resolved SARS-CoV-2 antibody shows high efficacy in severely infected hamsters and provides a potent cocktail pairing strategy. Cell 183, 1013–1023.e13 (2020).
Du, Y. et al. A broadly neutralizing humanized ACE2-targeting antibody against SARS-CoV-2 variants. Nat. Commun. 12, 5000 (2021).
Ge, J. et al. Antibody neutralization of SARS-CoV-2 through ACE2 receptor mimicry. Nat. Commun. 12, 250 (2021).
Graham, C. et al. Neutralization potency of monoclonal antibodies recognizing dominant and subdominant epitopes on SARS-CoV-2 spike is impacted by the B.1.1.7 variant. Immunity 54, 1276–1289.e6 (2021).
Tuccori, M. et al. Anti-SARS-CoV-2 neutralizing monoclonal antibodies: clinical pipeline. MAbs 12, 1854149 (2020).
Cameroni, E. et al. Broadly neutralizing antibodies overcome SARS-CoV-2 Omicron antigenic shift. Nature 602, 664–670 (2022).
Hastie, K. M. et al. Defining variant-resistant epitopes targeted by SARS-CoV-2 antibodies: a global consortium study. Science 374, 472–478 (2021).
Starr, T. N. et al. Prospective mapping of viral mutations that escape antibodies used to treat COVID-19. Science 371, 850–854 (2021).
An EUA for bamlanivimab—a monoclonal antibody for COVID-19. JAMA 325, 880–881 (2021).
An EUA for bamlanivimab and etesevimab for COVID-19. Med. Lett. Drugs Ther. 63, 49–50 (2021).
Deeks, E. D. Casirivimab/Imdevimab: First Approval. Drugs 81, 2047–2055 (2021).
Casirivimab and imdevimab (REGEN-COV) for post-exposure prophylaxis of COVID-19. Med. Lett. Drugs Ther. 63, 130–131 (2021).
Tixagevimab and cilgavimab (Evusheld) for pre-exposure prophylaxis of COVID-19. JAMA 327, 384–385 (2022).
An EUA for sotrovimab for treatment of COVID-19. Med. Lett. Drugs Ther. 63, 97−xx98 (2021).
Syed, Y. Y. Correction to: regdanvimab: first approval. Drugs 81, 2139 (2021).
Streinu-Cercel, A. et al. Efficacy and safety of regdanvimab (CT-P59): a phase 2/3 randomized, double-blind, placebo-controlled trial in outpatients with mild-to-moderate coronavirus disease 2019. Open Forum Infect. Dis. 9, ofac053 (2022).
Taylor, P. C. et al. Neutralizing monoclonal antibodies for treatment of COVID-19. Nat. Rev. Immunol. 21, 382–393 (2021).
Kaplon, H. & Reichert, J. M. Antibodies to watch in 2021. MAbs 13, 1860476 (2021).
Copin, R. et al. The monoclonal antibody combination REGEN-COV protects against SARS-CoV-2 mutational escape in preclinical and human studies. Cell 184, 3949–3961.e11 (2021).
Hoffmann, M. et al. The Omicron variant is highly resistant against antibody-mediated neutralization: Implications for control of the COVID-19 pandemic. Cell 185, 447–456.e11 (2022).
Tong, P. et al. Memory B cell repertoire for recognition of evolving SARS-CoV-2 spike. Cell 184, 4969–4980.e15 (2021).
Chen, R. E. et al. Resistance of SARS-CoV-2 variants to neutralization by monoclonal and serum-derived polyclonal antibodies. Nat. Med. 27, 717–726 (2021).
Chen, R. E. et al. In vivo monoclonal antibody efficacy against SARS-CoV-2 variant strains. Nature 596, 103–108 (2021).
Gobeil, S. M. et al. Effect of natural mutations of SARS-CoV-2 on spike structure, conformation, and antigenicity. Science 373, eabi6226 (2021).
Mannar, D. et al. SARS-CoV-2 Omicron variant: antibody evasion and cryo-EM structure of spike protein-ACE2 complex. Science 375, 760–764 (2022).
McCallum, M. et al. Structural basis of SARS-CoV-2 Omicron immune evasion and receptor engagement. Science 375, 864–868 (2022).
Cui, Z. et al. Structural and functional characterizations of infectivity and immune evasion of SARS-CoV-2 Omicron. Cell 185, 860–871.e13 (2022).
Liu, Z. et al. Identification of SARS-CoV-2 spike mutations that attenuate monoclonal and serum antibody neutralization. Cell Host Microbe 29, 477–488.e4 (2021).
Rees-Spear, C. et al. The effect of spike mutations on SARS-CoV-2 neutralization. Cell Rep. 34, 108890 (2021).
McCallum, M. et al. SARS-CoV-2 immune evasion by the B.1.427/B.1.429 variant of concern. Science 373, 648–654 (2021).
Li, C. et al. Broad neutralization of SARS-CoV-2 variants by an inhalable bispecific single-domain antibody. Cell 185, 1389–1401.e18 (2022).
Westendorf, K. et al. LY-CoV1404 (bebtelovimab) potently neutralizes SARS-CoV-2 variants. Cell Rep. 39, 110812 (2022).
Li, Q. et al. SARS-CoV-2 501Y.V2 variants lack higher infectivity but do have immune escape. Cell 184, 2362–2371.e9 (2021).
Bamlanivimab (Drugs and Lactation Database (LactMed), National Library of Medicine (US), 2022).
Etesevimab and Bamlanivimab (Drugs and Lactation Database (LactMed), National Library of Medicine (US), 2022).
Takashita, E. et al. Efficacy of antibodies and antiviral drugs against Covid-19 Omicron variant. N. Engl. J. Med. 386, 995–998 (2022).
Beaudoin-Bussières, G. et al. A Fc-enhanced NTD-binding non-neutralizing antibody delays virus spread and synergizes with a nAb to protect mice from lethal SARS-CoV-2 infection. Cell Rep. 38, 110368 (2022).
Du, S. et al. Structures of SARS-CoV-2 B.1.351 neutralizing antibodies provide insights into cocktail design against concerning variants. Cell Res. 31, 1130–1133 (2021).
Guo, Y. et al. A SARS-CoV-2 neutralizing antibody with extensive Spike binding coverage and modified for optimal therapeutic outcomes. Nat. Commun. 12, 2623 (2021).
Cho, H. et al. Bispecific antibodies targeting distinct regions of the spike protein potently neutralize SARS-CoV-2 variants of concern. Sci. Transl. Med. 13, eabj5413 (2021).
Wratil, P. R. et al. Three exposures to the spike protein of SARS-CoV-2 by either infection or vaccination elicit superior neutralizing immunity to all variants of concern. Nat. Med. 28, 496–503 (2022).
Boschi, C., Colson, P., Bancod, A., Moal, V. & La Scola, B. Omicron variant escapes therapeutic mAbs including recently released Evusheld ®, contrary to eight prior main VOC. Clin. Infect. Dis. ciac143 (2022).
Shrotri, M., Swinnen, T., Kampmann, B. & Parker, E. An interactive website tracking COVID-19 vaccine development. Lancet Glob. Health 9, e590–e592 (2021).
Kashte, S., Gulbake, A., El-Amin, I. S. & Gupta, A. COVID-19 vaccines: rapid development, implications, challenges and future prospects. Hum. Cell 34, 711–733 (2021).
Shilo, S., Rossman, H. & Segal, E. Signals of hope: gauging the impact of a rapid national vaccination campaign. Nat. Rev. Immunol. 21, 198–199 (2021).
Wagner, C. E., Saad-Roy, C. M. & Grenfell, B. T. Modelling vaccination strategies for COVID-19. Nat. Rev. Immunol. 22, 139–141 (2022).
Pattni, K. et al. Effectiveness of the BNT162b2 (Pfizer-BioNTech) and the ChAdOx1 nCoV-19 (Oxford-AstraZeneca) vaccines for reducing susceptibility to infection with the Delta variant (B.1.617.2) of SARS-CoV-2. BMC Infect. Dis. 22, 270 (2022).
Tang, P. et al. BNT162b2 and mRNA-1273 COVID-19 vaccine effectiveness against the SARS-CoV-2 Delta variant in Qatar. Nat. Med. 27, 2136–2143 (2021).
Tregoning, J. S., Flight, K. E., Higham, S. L., Wang, Z. & Pierce, B. F. Progress of the COVID-19 vaccine effort: viruses, vaccines and variants versus efficacy, effectiveness and escape. Nat. Rev. Immunol. 21, 626–636 (2021).
Garcia-Beltran, W. F. et al. Multiple SARS-CoV-2 variants escape neutralization by vaccine-induced humoral immunity. Cell 184, 2372–2383.e9 (2021).
Zhou, D. et al. Evidence of escape of SARS-CoV-2 variant B.1.351 from natural and vaccine-induced sera. Cell 184, 2348–2361.e6 (2021).
Lu, L. et al. Neutralization of SARS-CoV-2 Omicron variant by sera from BNT162b2 or Coronavac vaccine recipients. Clin. Infect. Dis. ciab1041 (2021).
Addetia, A. et al. Neutralizing antibodies correlate with protection from SARS-CoV-2 in humans during a fishery vessel outbreak with a high attack rate. J. Clin. Microbiol. 58, e02107–20 (2020).
Khoury, D. S. et al. Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection. Nat. Med. 27, 1205–1211 (2021).
Tzou, P. L. et al. Coronavirus antiviral research database (cov-rdb): an online database designed to facilitate comparisons between candidate anti-coronavirus compounds. Viruses 12, 1006 (2020).
Liu, Y. et al. Neutralizing activity of BNT162b2-elicited serum. N. Engl. J. Med. 384, 1466–1468 (2021).
Supasa, P. et al. Reduced neutralization of SARS-CoV-2 B.1.1.7 variant by convalescent and vaccine sera. Cell 184, 2201–2211.e7 (2021).
Geers, D. et al. SARS-CoV-2 variants of concern partially escape humoral but not T-cell responses in COVID-19 convalescent donors and vaccinees. Sci. Immunol. 6, eabj1750 (2021).
Tada, T. et al. Convalescent-phase sera and vaccine-elicited antibodies largely maintain neutralizing titer against global SARS-CoV-2 variant spikes. MBio 12, e0069621 (2021).
Alenquer, M. et al. Signatures in SARS-CoV-2 spike protein conferring escape to neutralizing antibodies. PLoS Pathog. 17, e1009772 (2021).
Wang, P. et al. Increased resistance of SARS-CoV-2 variant P.1 to antibody neutralization. Cell Host Microbe 29, 747–751.e4 (2021).
Garcia-Beltran, W. F. et al. mRNA-based COVID-19 vaccine boosters induce neutralizing immunity against SARS-CoV-2 Omicron variant. Cell 185, 457–466.e4 (2022).
Uriu, K. et al. Neutralization of the SARS-CoV-2 Mu variant by convalescent and vaccine serum. N. Engl. J. Med. 385, 2397–2399 (2021).
Wang, B. et al. Resistance of SARS-CoV-2 Delta variant to neutralization by BNT162b2-elicited antibodies in Asians. Lancet Reg. Health West Pac. 15, 100276 (2021).
Stamatatos, L. et al. mRNA vaccination boosts cross-variant neutralizing antibodies elicited by SARS-CoV-2 infection. Science eabg9175 (2021).
Goel, R. R. et al. Distinct antibody and memory B cell responses in SARS-CoV-2 naïve and recovered individuals following mRNA vaccination. Sci. Immunol. 6, eabi6950 (2021).
Gruell, H. et al. mRNA booster immunization elicits potent neutralizing serum activity against the SARS-CoV-2 Omicron variant. Nat. Med. 28, 477–480 (2022).
Schmidt, F. et al. Plasma Neutralization of the SARS-CoV-2 Omicron Variant. N. Engl. J. Med. 386, 599–601 (2022).
Liu, L. et al. Striking antibody evasion manifested by the Omicron variant of SARS-CoV-2. Nature 602, 676–681 (2022).
Muik, A. et al. Neutralization of SARS-CoV-2 Omicron by BNT162b2 mRNA vaccine-elicited human sera. Science 375, 678–680 (2022).
Pajon, R. et al. SARS-CoV-2 Omicron Variant Neutralization after mRNA-1273 Booster Vaccination. N. Engl. J. Med. 386, 1088–1091 (2022).
Wang, X. et al. Homologous or heterologous booster of inactivated vaccine reduces SARS-CoV-2 Omicron variant escape from neutralizing antibodies. Emerg. Microbes Infect. 11, 477–481 (2022).
Ai, J. et al. Omicron variant showed lower neutralizing sensitivity than other SARS-CoV-2 variants to immune sera elicited by vaccines after boost. Emerg. Microbes Infect. 11, 337–343 (2022).
Edara, V. V. et al. mRNA-1273 and BNT162b2 mRNA vaccines have reduced neutralizing activity against the SARS-CoV-2 omicron variant. Cell Rep. Med. 3, 100529 (2022).
Carreño, J. M. et al. Activity of convalescent and vaccine serum against SARS-CoV-2 Omicron. Nature 602, 682–688 (2022).
Lapa, D. et al. Retention of Neutralizing Response against SARS-CoV-2 Omicron Variant in Sputnik V-Vaccinated Individuals. Vaccines 10, 817 (2022).
Yin, W. et al. Structural basis for inhibition of the RNA-dependent RNA polymerase from SARS-CoV-2 by remdesivir. Science 368, 1499–1504 (2020).
Shapira, T. et al. A TMPRSS2 inhibitor acts as a pan-SARS-CoV-2 prophylactic and therapeutic. Nature 605, 340–348 (2022).
Ibrahim, M. et al. Rutin and flavone analogs as prospective SARS-CoV-2 main protease inhibitors: In silico drug discovery study. J. Mol. Graph. Model. 105, 107904 (2021).
Hillen, H. S. et al. Structure of replicating SARS-CoV-2 polymerase. Nature 584, 154–156 (2020).
Liu, T., Luo, S., Libby, P. & Shi, G. P. Cathepsin L-selective inhibitors: a potentially promising treatment for COVID-19 patients. Pharm. Ther. 213, 107587 (2020).
Osipiuk, J. et al. Structure of papain-like protease from SARS-CoV-2 and its complexes with non-covalent inhibitors. Nat. Commun. 12, 743 (2021).
Lv, Z. et al. Targeting SARS-CoV-2 proteases for COVID-19 antiviral development. Front. Chem. 9, 819165 (2021).
Sacco, M. D. et al. The P132H mutation in the main protease of Omicron SARS-CoV-2 decreases thermal stability without compromising catalysis or small-molecule drug inhibition. Cell Res. 32, 498–500 (2022).
Ullrich, S., Ekanayake, K. B., Otting, G. & Nitsche, C. Main protease mutants of SARS-CoV-2 variants remain susceptible to nirmatrelvir. Bioorg. Med. Chem. Lett. 62, 128629 (2022).
Martinez, D. R. et al. A broadly cross-reactive antibody neutralizes and protects against sarbecovirus challenge in mice. Sci. Transl. Med. 14, eabj7125 (2022).
Apaydın, Ç. B., Çınar, G. & Cihan-Üstündağ, G. Small-molecule antiviral agents in ongoing clinical trials for COVID-19. Curr. Drug Targets 22, 1986–2005 (2021).
Yamamoto, M. et al. The anticoagulant nafamostat potently inhibits SARS-CoV-2 S protein-mediated fusion in a cell fusion assay system and viral infection in vitro in a cell-type-dependent manner. Viruses 12, 629 (2020).
Rubin, D., Chan-Tack, K., Farley, J. & Sherwat, A. FDA approval of remdesivir—a step in the right direction. N. Engl. J. Med. 383, 2598–2600 (2020).
Rosenke, K. et al. Molnupiravir (MK-4482) is efficacious against Omicron and other SARS-CoV-2 variants in the Syrian hamster COVID-19 model. Preprint at bioRxiv https://doi.org/10.1101/2022.02.22.481491 (2022).
Ledford, H. Chloroquine hype is derailing the search for coronavirus treatments. Nature 580, 573 (2020).
Mahoney, M. et al. A novel class of TMPRSS2 inhibitors potently block SARS-CoV-2 and MERS-CoV viral entry and protect human epithelial lung cells. Proc. Natl Acad. Sci. USA 118, e2108728118 (2021).
Aschenbrenner, D. S. Remdesivir approved to treat COVID-19 amid controversy. Am. J. Nurs. 121, 22–24 (2021).
Owen, D. R. et al. An oral SARS-CoV-2 M(pro) inhibitor clinical candidate for the treatment of COVID-19. Science 374, 1586–1593 (2021).
Molnupiravir for treatment of COVID-19. Med. Lett. Drugs Ther. 64, 10–11 (2022).
Paxlovid for treatment of COVID-19. Med. Lett. Drugs Ther. 64, 9 (2022).
Abdelnabi, R. et al. Molnupiravir inhibits replication of the emerging SARS-CoV-2 variants of concern in a hamster infection model. J. Infect. Dis. 224, 749–753 (2021).
Choy, K. T. et al. Remdesivir, lopinavir, emetine, and homoharringtonine inhibit SARS-CoV-2 replication in vitro. Antivir. Res. 178, 104786 (2020).
Vangeel, L. et al. Remdesivir, Molnupiravir and Nirmatrelvir remain active against SARS-CoV-2 Omicron and other variants of concern. Antivir. Res. 198, 105252 (2022).
Li, P. et al. SARS-CoV-2 Omicron variant is highly sensitive to molnupiravir, nirmatrelvir, and the combination. Cell Res. 32, 322–324 (2022).
Wang, M. et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 30, 269–271 (2020).
Hoffmann, M. et al. Camostat mesylate inhibits SARS-CoV-2 activation by TMPRSS2-related proteases and its metabolite GBPA exerts antiviral activity. EBiomedicine 65, 103255 (2021).
Hoffmann, M. et al. Nafamostat mesylate blocks activation of SARS-CoV-2: new treatment option for COVID-19. Antimicrob. Agents Chemother. 64, e00754–20 (2020).
Iwata-Yoshikawa, N. et al. TMPRSS2 contributes to virus spread and immunopathology in the airways of murine models after coronavirus infection. J. Virol. 93, e01815–e01818 (2019).
Mouffak, S., Shubbar, Q., Saleh, E. & El-Awady, R. Recent advances in management of COVID-19: a review. Biomed. Pharmacother. 143, 112107 (2021).
Zhu, H. et al. Spontaneous binding of potential COVID-19 drugs (Camostat and Nafamostat) to human serine protease TMPRSS2. Comput. Struct. Biotechnol. J. 19, 467–476 (2021).
Lee, J. et al. TMPRSS2 and RNA-dependent RNA polymerase are effective targets of therapeutic intervention for treatment of COVID-19 caused by SARS-CoV-2 variants (B.1.1.7 and B.1.351). Microbiol. Spectr. 9, e47221 (2021).
Gies, V. et al. Beyond anti-viral effects of chloroquine/hydroxychloroquine. Front. Immunol. 11, 1409 (2020).
Manivannan, E., Karthikeyan, C., Moorthy, N. & Chaturvedi, S. C. The rise and fall of chloroquine/hydroxychloroquine as compassionate therapy of COVID-19. Front. Pharm. 12, 584940 (2021).
Ferner, R. E. & Aronson, J. K. Chloroquine and hydroxychloroquine in covid-19. BMJ 369, m1432 (2020).
Hoffmann, M. et al. Chloroquine does not inhibit infection of human lung cells with SARS-CoV-2. Nature 585, 588–590 (2020).
Touret, F. & de Lamballerie, X. Of chloroquine and COVID-19. Antivir. Res. 177, 104762 (2020).
Das, S. et al. The controversial therapeutic journey of chloroquine and hydroxychloroquine in the battle against SARS-CoV-2: a comprehensive review. Med. Drug Discov. 10, 100085 (2021).
Riva, L. et al. Discovery of SARS-CoV-2 antiviral drugs through large-scale compound repurposing. Nature 586, 113–119 (2020).
Vuong, W. et al. Feline coronavirus drug inhibits the main protease of SARS-CoV-2 and blocks virus replication. Nat. Commun. 11, 4282 (2020).
Rajpoot, S., Alagumuthu, M. & Baig, M. S. Dual targeting of 3CL(pro) and PL(pro) of SARS-CoV-2: a novel structure-based design approach to treat COVID-19. Curr. Res. Struct. Biol. 3, 9–18 (2021).
Abdelnabi, R. et al. The oral protease inhibitor (PF-07321332) protects Syrian hamsters against infection with SARS-CoV-2 variants of concern. Nat. Commun. 13, 719 (2022).
Drayman, N. et al. Masitinib is a broad coronavirus 3CL inhibitor that blocks replication of SARS-CoV-2. Science 373, 931–936 (2021).
Rudrapal, M. et al. Repurposing of phytomedicine-derived bioactive compounds with promising anti-SARS-CoV-2 potential: Molecular docking, MD simulation and drug-likeness/ADMET studies. Saudi J. Biol. Sci. 29, 2432–2446 (2022).
Verma, D. et al. Potential inhibitors of SARS-CoV-2 (COVID 19) proteases PL(pro) and M(pro)/3CL(pro): molecular docking and simulation studies of three pertinent medicinal plant natural components. Curr. Res. Pharm. Drug Discov. 2, 100038 (2021).
Saravolatz, L. D., Depcinski, S. & Sharma, M. Molnupiravir and nirmatrelvir-ritonavir: oral COVID antiviral drugs. Clin. Infect. Dis. ciac180 (2022).
Arabi, Y. M. et al. Lopinavir-ritonavir and hydroxychloroquine for critically ill patients with COVID-19: REMAP-CAP randomized controlled trial. Intensive Care Med 47, 867–886 (2021).
Lamb, Y. N. Nirmatrelvir plus ritonavir: first approval. Drugs 82, 585–591 (2022).
McDonald, E. G. & Lee, T. C. Nirmatrelvir-ritonavir for COVID-19. CMAJ 194, E218 (2022).
Uraki, R. et al. Characterization and antiviral susceptibility of SARS-CoV-2 Omicron/BA.2. Nature https://doi.org/10.1038/s41586-022-04856-1 (2022)
Saeed, A. et al. Targeting omicron and other reported SARS-CoV-2 lineages by potent inhibitors of main protease 3CL Mpro: molecular simulation analysis. J. Infect. 84, e133–e136 (2022).
Liang, J. J. et al. Investigation of small molecule inhibitors of the SARS-CoV-2 papain-like protease by all-atom microsecond modelling, PELE Monte Carlo simulations, and in vitro activity inhibition. Chem. Phys. Lett. 139294 (2021).
Pitsillou, E. et al. Identification of small molecule inhibitors of the deubiquitinating activity of the SARS-CoV-2 papain-like protease: in silico molecular docking studies and in vitro enzymatic activity assay. Front. Chem. 8, 623971 (2020).
Hajbabaie, R., Harper, M. T. & Rahman, T. Establishing an analogue based in silico pipeline in the pursuit of novel inhibitory scaffolds against the SARS coronavirus 2 papain-like protease. Molecules 26, 1134 (2021).
Narayanan, A. et al. Identification of SARS-CoV-2 inhibitors targeting Mpro and PLpro using in-cell-protease assay. Commun. Biol. 5, 169 (2022).
Zhang, L. C. et al. Design of SARS-CoV-2 Mpro, PLpro dual-target inhibitors based on deep reinforcement learning and virtual screening. Future Med Chem. 14, 393–405 (2022).
Abbott, T. R. et al. Development of CRISPR as an Antiviral Strategy to Combat SARS-CoV-2 and Influenza. Cell 181, 865–876.e12 (2020).
Tian, D. et al. An update review of emerging small-molecule therapeutic options for COVID-19. Biomed. Pharmacotherp. 137, 111313 (2021).
WHO Solidarity Trial Consortium. Remdesivir and three other drugs for hospitalised patients with COVID-19: final results of the WHO Solidarity randomised trial and updated meta-analyses. Lancet 399, 1941–1953 (2022).
Cox, R. M. et al. Oral prodrug of remdesivir parent GS-441524 is efficacious against SARS-CoV-2 in ferrets. Nat. Commun. 12, 6415 (2021).
Murphy, B. G. et al. The nucleoside analog GS-441524 strongly inhibits feline infectious peritonitis (FIP) virus in tissue culture and experimental cat infection studies. Vet. Micobiol. 219, 226–233 (2018).
Zhang, Y. & Tang, L. V. Overview of targets and potential drugs of SARS-CoV-2 according to the viral replication. J. Proteome Res 20, 49–59 (2021).
Wahl, A. et al. SARS-CoV-2 infection is effectively treated and prevented by EIDD-2801. Nature 591, 451–457 (2021).
Syed, Y. Y. Molnupiravir: first approval. Drugs 82, 455–460 (2022).
Zhao, H. et al. SARS-CoV-2 Omicron variant shows less efficient replication and fusion activity when compared with Delta variant in TMPRSS2-expressed cells. Emerg. Microbes Infect. 11, 277–283 (2022).
Hashimoto, R. et al. Dual inhibition of TMPRSS2 and Cathepsin Bprevents SARS-CoV-2 infection in iPS cells. Mol. Ther. Nucleic Acids 26, 1107–1114 (2021).
Hoffmann, M. et al. SARS-CoV-2 variants B.1.351 and P.1 escape from neutralizing antibodies. Cell 184, 2384–2393.e12 (2021).
Hoffmann, M. et al. SARS-CoV-2 variant B.1.617 is resistant to bamlanivimab and evades antibodies induced by infection and vaccination. Cell Rep. 36, 109415 (2021).
Hu, B., Guo, H., Zhou, P. & Shi, Z. L. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 19, 141–154 (2021).
Lu, J. et al. Genomic epidemiology of SARS-CoV-2 in Guangdong Province, China. Cell 181, 997–1003.e9 (2020).
Stower, H. Clinical and epidemiological characteristics of children with COVID-19. Nat. Med. 26, 465 (2020).
Coronavirus Pandemic (COVID-19). https://ourworldindata.org/coronavirus (2020).
WHO Coronavirus (COVID-19) Dashboard. https://covid19.who.int/ (2021).
Gupta, R. K. Will SARS-CoV-2 variants of concern affect the promise of vaccines? Nat. Rev. Immunol. 21, 340–341 (2021).
Callaway, E. Omicron likely to weaken COVID vaccine protection. Nature 600, 367–368 (2021).
Polack, F. P. et al. Safety and efficacy of the BNT162b2 mRNA Covid-19 vaccine. N. Engl. J. Med. 383, 2603–2615 (2020).
Tanriover, M. D. et al. Efficacy and safety of an inactivated whole-virion SARS-CoV-2 vaccine (CoronaVac): interim results of a double-blind, randomised, placebo-controlled, phase 3 trial in Turkey. Lancet 398, 213–222 (2021).
Al, K. N. et al. Effect of 2 inactivated SARS-CoV-2 vaccines on symptomatic Covid-19 infection in adults: a randomized clinical trial. JAMA 326, 35–45 (2021).
Bruxvoort, K. J. et al. Effectiveness of mRNA-1273 against delta, mu, and other emerging variants of SARS-CoV-2: test negative case-control study. BMJ 375, e068848 (2021).
Abu-Raddad, L. J., Chemaitelly, H. & Butt, A. A. Effectiveness of the BNT162b2 Covid-19 Vaccine against the B.1.1.7 and B.1.351 Variants. N. Engl. J. Med. 385, 187–189 (2021).
Chemaitelly, H. et al. mRNA-1273 COVID-19 vaccine effectiveness against the B.1.1.7 and B.1.351 variants and severe COVID-19 disease in Qatar. Nat. Med 27, 1614–1621 (2021).
Nasreen, S. et al. Effectiveness of COVID-19 vaccines against symptomatic SARS-CoV-2 infection and severe outcomes with variants of concern in Ontario. Nat. Microbiol. 7, 379–385 (2022).
Munitz, A., Yechezkel, M., Dickstein, Y., Yamin, D. & Gerlic, M. BNT162b2 vaccination effectively prevents the rapid rise of SARS-CoV-2 variant B.1.1.7 in high-risk populations in Israel. Cell Rep. Med. 2, 100264 (2021).
Mor, O. et al. BNT162b2 vaccine effectiveness was marginally affected by the SARS-CoV-2 beta variant in fully vaccinated individuals. J. Clin. Epidemiol. 142, 38–44 (2022).
Vicenti, I. et al. BNT162b2 SARS-CoV-2 vaccination elicits high titers of neutralizing antibodies to both B.1 and P.1 variants in previously infected and uninfected subjects. Life (Basel) 11, 896 (2021).
Skowronski, D. M. et al. Two-dose SARS-CoV-2 vaccine effectiveness with mixed schedules and extended dosing intervals: test-negative design studies from British Columbia and Quebec, Canada. Clin. Infect. Dis. ciac290 (2022).
Chin, E. T. et al. Effectiveness of the mRNA-1273 vaccine during a SARS-CoV-2 delta outbreak in a prison. N. Engl. J. Med. 385, 2300–2301 (2021).
Wu, K. et al., mRNA-1273 vaccine induces neutralizing antibodies against spike mutants from global SARS-CoV-2 variants. Preprint at bioRxiv https://doi.org/10.1101/2021.01.25.427948 (2021).
Kaplonek, P. et al. mRNA-1273 vaccine-induced antibodies maintain Fc effector functions across SARS-CoV-2 variants of concern. Immunity 55, 355–365.e4 (2022).
de Gier, B. et al. Vaccine effectiveness against SARS-CoV-2 transmission and infections among household and other close contacts of confirmed cases, the Netherlands, February to May 2021. Eur. Surveill. 26, 2100640 (2021).
Eyre, D. W. et al. Effect of Covid-19 vaccination on transmission of alpha and delta variants. N. Engl. J. Med. 386, 744–756 (2022).
Lopez, B. J. et al. Effectiveness of Covid-19 vaccines against the B.1.617.2 (Delta) variant. N. Engl. J. Med. 385, 585–594 (2021).
Ranzani, O. T. et al. Effectiveness of the CoronaVac vaccine in older adults during a gamma variant associated epidemic of covid-19 in Brazil: test negative case-control study. BMJ 374, n2015 (2021).
Pouwels, K. B. et al. Effect of Delta variant on viral burden and vaccine effectiveness against new SARS-CoV-2 infections in the UK. Nat. Med. 27, 2127–2135 (2021).
Andrews, N. et al. Covid-19 vaccine effectiveness against the Omicron (B.1.1.529) Variant. N. Engl. J. Med. 386, 1532–1546 (2022).
Sritipsukho, P. et al. Comparing real-life effectiveness of various COVID-19 vaccine regimens during the delta variant-dominant pandemic: a test-negative case-control study. Emerg. Microbes Infect. 11, 585–592 (2022).
Cheng, S. M. S. et al. Neutralizing antibodies against the SARS-CoV-2 Omicron variant BA.1 following homologous and heterologous CoronaVac or BNT162b2 vaccination. Nat. Med. 28, 486–489 (2022).
Wu, D. et al. Effectiveness of inactivated COVID-19 vaccines against symptomatic, pneumonia, and severe disease caused by the delta variant: Real World Study and evidence—China, 2021. China CDC Wkly 4, 57–65 (2022).
Spensley, K. J. et al. Comparison of Vaccine Effectiveness Against the Omicron (B.1.1.529) Variant in Hemodialysis Patients. Kidney Int. Rep. 7, 1406–1409 (2022).
Willett, B. J. et al. The hyper-transmissible SARS-CoV-2 Omicron variant exhibits significant antigenic change, vaccine escape and a switch in cell entry mechanism. Preprint at medRxiv https://doi.org/10.1101/2022.01.03.21268111 (2022).
COVID-19 vaccine surveillance report. https://www.gov.uk/government/publications/covid-19-vaccine-weekly-surveillance-reports (2022).
Beyerstedt, S., Casaro, E. B. & Rangel, É. B. COVID-19: angiotensin-converting enzyme 2 (ACE2) expression and tissue susceptibility to SARS-CoV-2 infection. Eur. J. Clin. Microbiol Infect. Dis. 40, 905–919 (2021).
Ou, J. et al. V367F mutation in SARS-CoV-2 spike RBD emerging during the early transmission phase enhances viral infectivity through increased human ACE2 receptor binding affinity. J. Virol. 95, e0061721 (2021).
Daniloski, Z. et al. Identification of required host factors for SARS-CoV-2 infection in human. Cells Cell 184, 92–105.e16 (2021).
Syed, A. M. et al. Rapid assessment of SARS-CoV-2-evolved variants using virus-like particles. Science 374, 1626–1632 (2021).
Cai, Y. et al. Structural basis for enhanced infectivity and immune evasion of SARS-CoV-2 variants. Science 373, 642–648 (2021).
Sanjuán, R. The social life of viruses. Annu. Rev. Virol. 8, 183–199 (2021).
Park, G. & Hwang, B. H. SARS-CoV-2 variants: mutations and effective changes. Biotechnol. Bioprocess Eng. 26, 859–870 (2021).
Rothenburg, S. & Brennan, G. Species-specific host–virus interactions: implications for viral host range and virulence. Trends Micobiol. 28, 46–56 (2020).
Sit, T. H. C. et al. Infection of dogs with SARS-CoV-2. Nature 586, 776–778 (2020).
Liu, K. et al. Cross-species recognition of SARS-CoV-2 to bat ACE2. Proc. Natl Acad. Sci. USA 118, e2020216118 (2021).
Hossain, M. G., Javed, A., Akter, S. & Saha, S. SARS-CoV-2 host diversity: An update of natural infections and experimental evidence. J. Microbiol Immunol. Infect. 54, 175–181 (2021).
Conceicao, C. et al. The SARS-CoV-2 spike protein has a broad tropism for mammalian ACE2 proteins. Plos Biol. 18, e3001016 (2020).
Zhao, X. et al. Broad and differential animal angiotensin-converting enzyme 2 receptor usage by SARS-CoV-2. J. Virol. 94, e00940–20 (2020).
Liu, Y. et al. Functional and genetic analysis of viral receptor ACE2 orthologs reveals a broad potential host range of SARS-CoV-2. Proc. Natl Acad. Sci. USA 118, e2025373118 (2021).
Bonilla-Aldana, D. K. & Rodriguez-Morales, A. J. The threat of the spread of SARS-CoV-2 variants in animals. Vet. Q 41, 321–322 (2021).
Ulrich, L. et al. Enhanced fitness of SARS-CoV-2 variant of concern Alpha but not Beta. Nature 602, 307–313 (2022).
Plante, J. A. et al. Spike mutation D614G alters SARS-CoV-2 fitness. Nature 592, 116–121 (2021).
Zhou, B. et al. SARS-CoV-2 spike D614G change enhances replication and transmission. Nature 592, 122–127 (2021).
Kuzmina, A. et al. SARS CoV-2 Delta variant exhibits enhanced infectivity and a minor decrease in neutralization sensitivity to convalescent or post-vaccination sera. iScience 24, 103467 (2021).
Deng, X. et al. Transmission, infectivity, and neutralization of a spike L452R SARS-CoV-2 variant. Cell 184, 3426–3437.e8 (2021).
Liu, Y. et al. Delta spike P681R mutation enhances SARS-CoV-2 fitness over Alpha variant. bioRxiv (2021).
Davies, N. G. et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 372, eabg3055 (2021).
Faria, N. R. et al. Genomics and epidemiology of the P.1 SARS-CoV-2 lineage in Manaus, Brazil. Science 372, 815–821 (2021).
Dhar, M. S. et al. Genomic characterization and epidemiology of an emerging SARS-CoV-2 variant in Delhi, India. Science 374, 995–999 (2021).
Dong, R. et al. Assessing the transmissibility of the new SARS-CoV-2 variants: from Delta to Omicron. Vaccines 10, 496 (2022).
Davies, N. G. et al. Increased mortality in community-tested cases of SARS-CoV-2 lineage B.1.1.7. Nature 593, 270–274 (2021).
Buchan, S. A. et al. Increased household secondary attacks rates with variant of concern severe acute respiratory syndrome coronavirus 2 index cases. Clin. Infect. Dis. 74, 703–706 (2022).
Altarawneh, H. N. et al. Protection against the Omicron variant from previous SARS-CoV-2 infection. N. Engl. J. Med. 386, 1288–1290 (2022).
Holgersen, E. M. et al. Transcriptome-wide off-target effects of steric-blocking oligonucleotides. Nucleic Acid Ther. 31, 392–403 (2021).
Lyngse, F. P. et al. SARS-CoV-2 Omicron VOC transmission in Danish households. Preprint at medRxiv https://doi.org/10.1101/2021.12.27.21268278 (2021).
Pandey, K. et al. Animal models for SARS-CoV-2 research: a comprehensive literature review. Transbound. Emerg. Dis. 68, 1868–1885 (2021).
Muñoz-Fontela, C. et al. Animal models for COVID-19. Nature 586, 509–515 (2020).
Parolin, C. et al. Animal hosts and experimental models of SARS-CoV-2 infection. Chemotherapy 66, 8–16 (2021).
Farag, E. A. et al. SARS-CoV-2 at the human-animal interphase: a review. Heliyon 7, e08496 (2021).
Jo, W. K. et al. Potential zoonotic sources of SARS-CoV-2 infections. Transbound. Emerg. Dis. 68, 1824–1834 (2021).
Mahdy, M. A. A., Younis, W. & Ewaida, Z. An overview of SARS-CoV-2 and animal infection. Front. Vet. Sci. 7, 596391 (2020).
Gaudreault, N. N. et al. SARS-CoV-2 infection, disease and transmission in domestic cats. Emerg. Microbes Infect. 9, 2322–2332 (2020).
Zhang, Q. et al. A serological survey of SARS-CoV-2 in cat in Wuhan. Emerg. Microbes Infect. 9, 2013–2019 (2020).
Fritz, M. et al. Detection of SARS-CoV-2 in two cats during the second wave of the COVID-19 pandemic in France. Vet. Med. Sci. 8, 14–20 (2022).
Sharun, K., Tiwari, R., Natesan, S. & Dhama, K. SARS-CoV-2 infection in farmed minks, associated zoonotic concerns, and importance of the One Health approach during the ongoing COVID-19 pandemic. Vet. Quart. 41, 50–60 (2021).
McAloose, D. et al. From People to Panthera: natural SARS-CoV-2 infection in tigers and lions at the Bronx Zoo. MBio 11, e02220–e02220 (2020).
Pramod, R. K. et al. Reverse zoonosis of coronavirus disease-19: present status and the control by one health approach. Vet. World 14, 2817–2826 (2021).
Braun, K. M. et al. Transmission of SARS-CoV-2 in domestic cats imposes a narrow bottleneck. PLoS Pathog. 17, e1009373 (2021).
Bosco-Lauth, A. M. et al. Experimental infection of domestic dogs and cats with SARS-CoV-2: Pathogenesis, transmission, and response to reexposure in cats. Proc. Natl Acad. Sci. USA 117, 26382–26388 (2020).
Shi, J. et al. Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS–coronavirus 2. Science 368, 1016–1020 (2020).
Lee, D. H. et al. Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in a dog in Connecticut in February 2021. Viruses 13, 2141 (2021).
Padilla-Blanco, M. et al. The finding of the severe acute respiratory syndrome coronavirus (SARS-CoV-2) in a Wild Eurasian River Otter (Lutra lutra) highlights the need for viral surveillance in wild mustelids. Front. Vet. Sci. 9, 826991 (2022).
Kim, Y. et al. Infection and rapid transmission of SARS-CoV-2 in Ferrets. Cell Host Microbe 27, 704–709.e2 (2020).
Cox, R. M., Wolf, J. D. & Plemper, R. K. Therapeutically administered ribonucleoside analogue MK-4482/EIDD-2801 blocks SARS-CoV-2 transmission in ferrets. Nat. Microbiol. 6, 11–18 (2021).
Palmer, M. V. et al. Susceptibility of white-tailed deer (Odocoileus virginianus) to SARS-CoV-2. J. Virol. 95, e00083–21 (2021).
Griffin, B. D. et al. SARS-CoV-2 infection and transmission in the North American deer mouse. Nat. Commun. 12, 3612 (2021).
Hale, V. L. et al. SARS-CoV-2 infection in free-ranging white-tailed deer. Nature 602, 481–486 (2022).
Chandler, J. C. et al. SARS-CoV-2 exposure in wild white-tailed deer (Odocoileus virginianus). Proc. Natl Acad. Sci. USA 118, e2114828118 (2021).
Hayashi, T., Abiko, K., Mandai, M., Yaegashi, N. & Konishi, I. Highly conserved binding region of ACE2 as a receptor for SARS-CoV-2 between humans and mammals. Vet. Q 40, 243–249 (2020).
Sia, S. F. et al. Pathogenesis and transmission of SARS-CoV-2 in golden hamsters. Nature 583, 834–838 (2020).
Rosenke, K. et al. Defining the Syrian hamster as a highly susceptible preclinical model for SARS-CoV-2 infection. Emerg. Microbes Infect. 9, 2673–2684 (2020).
Sailleau, C. et al. First detection and genome sequencing of SARS‐CoV‐2 in an infected cat in France. Transbound. Emerg. Dis. 67, 2324–2328 (2020).
Freuling, C. M. et al. Susceptibility of Raccoon dogs for experimental SARS-CoV-2 infection. Emerg. Infect. Dis. 26, 2982–2985 (2020).
Oude Munnink, B. B. et al. Transmission of SARS-CoV-2 on mink farms between humans and mink and back to humans. Science 371, 172–177 (2021).
Shuai, H. et al. Emerging SARS-CoV-2 variants expand species tropism to murines. EBiomedicine 73, 103643 (2021).
Campbell, F. et al. Increased transmissibility and global spread of SARS-CoV-2 variants of concern as at June 2021. Eur. Surveill. 26, 2100509 (2021).
Lyngse, F. P. et al. Increased transmissibility of SARS-CoV-2 lineage B.1.1.7 by age and viral load. Nat. Commun. 12, 7251 (2021).
Lindstrøm, J. C. et al. Increased transmissibility of the alpha SARS-CoV-2 variant: evidence from contact tracing data in Oslo, January to February 2021. Infect. Dis. 54, 72–77 (2022).
Ledford, H. How severe are Omicron infections? Nature 600, 577–578 (2021).
Paredes, M. I. et al. Associations between SARS-CoV-2 variants and risk of COVID-19 hospitalization among confirmed cases in Washington State: a retrospective cohort study. Preprint at medRxiv https://doi.org/10.1101/2021.09.29.21264272 (2022).
Freitas, A. et al. The emergence of novel SARS-CoV-2 variant P.1 in Amazonas (Brazil) was temporally associated with a change in the age and sex profile of COVID-19 mortality: a population based ecological study. Lancet Reg. Health Am. 1, 100021 (2021).
Fisman, D. N. & Tuite, A. R. Evaluation of the relative virulence of novel SARS-CoV-2 variants: a retrospective cohort study in Ontario, Canada. CMAJ 193, E1619–E1625 (2021).
Shuai, H. et al. Attenuated replication and pathogenicity of SARS-CoV-2 B.1.1.529 Omicron. Nature 603, 693–699 (2022).
Wang, L. et al. Comparison of outcomes from COVID infection in pediatric and adult patients before and after the emergence of Omicron. Preprint at medRxiv https://doi.org/10.1101/2021.12.30.21268495 (2022).
Watson, J., Whiting, P. F. & Brush, J. E. Interpreting a covid-19 test result. BMJ 369, m1808 (2020).
Berlin, D. A., Gulick, R. M. & Martinez, F. J. Severe Covid-19. N. Engl. J. Med. 383, 2451–2460 (2020).
Wu, Z. & McGoogan, J. M. Characteristics of and important lessons from the coronavirus disease 2019 (COVID-19) Outbreak in China: summary of a report of 72314 cases from the Chinese Center for Disease Control and Prevention. JAMA 323, 1239–1242 (2020).
Lauer, S. A. et al. The incubation period of coronavirus disease 2019 (COVID-19) from publicly reported confirmed cases: estimation and application. Ann. Intern. Med. 172, 577–582 (2020).
McAbee, G. N., Brosgol, Y., Pavlakis, S., Agha, R. & Gaffoor, M. Encephalitis associated with COVID-19 Infection in an 11-year-old child. Pediatr. Neurol. 109, 94 (2020).
Docherty, A. B. et al. Features of 20133 UK patients in hospital with covid-19 using the ISARIC WHO Clinical Characterisation Protocol: prospective observational cohort study. BMJ 369, m1985 (2020).
Wiersinga, W. J., Rhodes, A., Cheng, A. C., Peacock, S. J. & Prescott, H. C. Pathophysiology, transmission, diagnosis, and treatment of coronavirus disease 2019 (COVID-19): a review. JAMA 324, 782–793 (2020).
Wang, D. et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA 323, 1061–1069 (2020).
Godeau, D., Petit, A., Richard, I., Roquelaure, Y. & Descatha, A. Return-to-work, disabilities and occupational health in the age of COVID-19. Scand. J. Work Environ. Health 47, 408–409 (2021).
Spinato, G. et al. Alterations in smell or taste in mildly symptomatic outpatients with SARS-CoV-2 infection. JAMA 323, 2089–2090 (2020).
Shi, S. et al. Association of Cardiac Injury With Mortality in Hospitalized Patients With COVID-19 in Wuhan, China. JAMA Cardiol. 5, 802–810 (2020).
Ediz, C. et al. Is there any association of COVID-19 with testicular pain and epididymo-orchitis? Int J. Clin. Pract. 75, e13753 (2021).
Garg, S. et al. Hospitalization rates and characteristics of patients hospitalized with laboratory-confirmed coronavirus disease 2019—COVID-NET, 14 States, March 1–30, 2020. Morb. Mortal. Wkly Rep. 69, 458–464 (2020).
Levi, M., Thachil, J., Iba, T. & Levy, J. H. Coagulation abnormalities and thrombosis in patients with COVID-19. Lancet Haematol. 7, e438–e444 (2020).
Drake, T. M. et al. Characterisation of in-hospital complications associated with COVID-19 using the ISARIC WHO Clinical Characterisation Protocol UK: a prospective, multicentre cohort study. Lancet 398, 223–237 (2021).
Zhang, Y. J., Sun, X. F., Xie, B., Feng, W. J. & Han, S. L. Exploration of severe Covid-19 associated risk factor in China: meta-analysis of current evidence. Int. J. Clin. Pract. 75, e14900 (2021).
Wu, C. et al. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern. Med. 180, 934–943 (2020).
Jordan, R. E., Adab, P. & Cheng, K. K. Covid-19: risk factors for severe disease and death. BMJ 368, m1198 (2020).
Paul, J. F., Charles, P., Richaud, C., Caussin, C. & Diakov, C. Myocarditis revealing COVID-19 infection in a young patient. Eur. Heart J. Cardiovasc. Imaging 21, 776 (2020).
Halushka, M. K. & Vander, H. R. Myocarditis is rare in COVID-19 autopsies: cardiovascular findings across 277 postmortem examinations. Cardiovasc. Pathol. 50, 107300 (2021).
Ma, K. et al. COVID-19 myocarditis and severity factors: an adult cohort study. Preprint at medRxiv https://doi.org/10.1101/2020.03.19.20034124 (2020).
Coromilas, E. J. et al. Worldwide survey of COVID-19-associated arrhythmias. Circ. Arrhythm. Electrophysiol. 14, e009458 (2021).
O’Shea, C. J. et al. Ventricular arrhythmia burden during the coronavirus disease 2019 (COVID-19) pandemic. Eur. Heart J. 42, 520–528 (2021).
Yarmohammadi, H. et al. Frequency of atrial arrhythmia in hospitalized patients with COVID-19. Am. J. Cardiol. 147, 52–57 (2021).
Naz, A. & Billah, M. COVID-19 and coronary heart disease. Encyclopedia 1, 340–349 (2021).
Bularga, A., Chapman, A. R. & Mills, N. L. Mechanisms of myocardial injury in COVID-19. Clin. Chem. 67, 1044–1046 (2021).
Solomon, M. D. et al. Changes in patterns of hospital visits for acute myocardial infarction or ischemic stroke during COVID-19 surges. JAMA 326, 82–84 (2021).
Legrand, M. et al. Pathophysiology of COVID-19-associated acute kidney injury. Nat. Rev. Nephrol. 17, 751–764 (2021).
Batlle, D. et al. Acute kidney injury in covid-19: emerging evidence of a distinct pathophysiology. J. Am. Soc. Nephrol. 31, 1380–1383 (2020).
Cheng, Y. et al. The incidence, risk factors, and prognosis of acute kidney injury in adult patients with coronavirus disease 2019. Clin. J. Am. Soc. Nephrol. 15, 1394–1402 (2020).
Malieckal, D. A., Uppal, N. N., Ng, J. H., Jhaveri, K. D. & Hirsch, J. S. Electrolyte abnormalities in patients hospitalized with COVID-19. Clin. Kidney J. 14, 1704–1707 (2021).
Long, B. et al. Electrocardiographic manifestations of COVID-19. Am. J. Emerg. Med. 41, 96–103 (2021).
Lippi, G., South, A. M. & Henry, B. M. Electrolyte imbalances in patients with severe coronavirus disease 2019 (COVID-19). Ann. Clin. Biochem. 57, 262–265 (2020).
Rubino, F. et al. New-onset diabetes in Covid-19. N. Engl. J. Med. 383, 789–790 (2020).
Li, J. et al. COVID-19 infection may cause ketosis and ketoacidosis. Diabetes Obes. Metab. 22, 1935–1941 (2020).
Ntaios, G. et al. Characteristics and outcomes in patients with COVID-19 and acute ischemic stroke: the global COVID-19 stroke registry. Stroke 51, e254–e258 (2020).
Hoyer, C. et al. Acute stroke in times of the COVID-19 pandemic: a Multicenter Study. Stroke 51, 2224–2227 (2020).
Nannoni, S., de Groot, R., Bell, S. & Markus, H. S. Stroke in COVID-19: a systematic review and meta-analysis. Int. J. Stroke 16, 137–149 (2021).
Ye, M., Ren, Y. & Lv, T. Encephalitis as a clinical manifestation of COVID-19. Brain Behav. Immun. 88, 945–946 (2020).
Zambreanu, L. et al. A case of limbic encephalitis associated with asymptomatic COVID-19 infection. J. Neurol. Neurourg. Psychiatry 91, 1229–1230 (2020).
Gupta, A. et al. Extrapulmonary manifestations of COVID-19. Nat. Med. 26, 1017–1032 (2020).
Nalbandian, A. et al. Post-acute COVID-19 syndrome. Nat. Med. 27, 601–615 (2021).
Raveendran, A. V., Jayadevan, R. & Sashidharan, S. Long COVID: an overview. Diabetes Metab. Syndr. 15, 869–875 (2021).
Burke, M. J. & Del, R. C. Long COVID has exposed medicine’s blind-spot. Lancet Infect. Dis. 21, 1062–1064 (2021).
Tenforde, M. W. et al. Symptom duration and risk factors for delayed return to usual health among outpatients with COVID-19 in a Multistate Health Care Systems Network—United States, March–June 2020. Morb. Mortal. Wkly Rep. 69, 993–998 (2020).
Ayoubkhani, D. et al. Post-covid syndrome in individuals admitted to hospital with covid-19: retrospective cohort study. BMJ 372, n693 (2021).
Whitaker, M. et al. Persistent symptoms following SARS-CoV-2 infection in a random community sample of 508,707 people. Preprint at medRxiv https://doi.org/10.1101/2021.06.28.21259452 (2021).
O’Dowd, A. Covid-19: third of people infected have long term symptoms. BMJ 373, n1626 (2021).
Alkodaymi, M. S. et al. Prevalence of post-acute COVID-19 syndrome symptoms at different follow-up periods: a systematic review and meta-analysis. Clin. Microbiol. Infect. 28, 657–666 (2022).
Ambrosino, P., Lanzillo, A. & Maniscalco, M. COVID-19 and post-acute COVID-19 Syndrome: From Pathophysiology To Novel Translational Applications. Biomedicines 10, 47 (2022).
Douaud, G. et al. SARS-CoV-2 is associated with changes in brain structure in UK Biobank. Nature 604, 697–707 (2022).
von Bartheld, C. S., Hagen, M. M. & Butowt, R. The D614G virus mutation enhances anosmia in COVID-19 patients: evidence from a systematic review and meta-analysis of studies from South Asia. ACS Chem. Neurosci. 12, 3535–3549 (2021).
Investigation of SARS-CoV-2 variants: technical briefings. https://www.gov.uk/government/publications/investigation-of-sars-cov-2-variants-technical-briefings (2022).
Meng, B. et al. Altered TMPRSS2 usage by SARS-CoV-2 Omicron impacts infectivity and fusogenicity. Nature 603, 706–714 (2022).
Spiers, N. Recognising and bearing the burden of long COVID-related disability. Br. J. Gen. Pract. 72, 70 (2022).
Dhama, K. et al. Coronavirus disease 2019-COVID-19. Clin. Micobiol Rev. 33, e00028–20 (2020).
Kupferschmidt, K. & Vogel, G. How bad is Omicron? Some clues are emerging. Science 374, 1304–1305 (2021).
Callaway, E. & Ledford, H. How bad is Omicron? What scientists know so far. Nature 600, 197–199 (2021).
Plante, J. A. et al. The variant gambit: COVID-19’s next move. Cell Host Microbe 29, 508–515 (2021).
V’Kovski, P., Kratzel, A., Steiner, S., Stalder, H. & Thiel, V. Coronavirus biology and replication: implications for SARS-CoV-2. Nat. Rev. Microbiol. 19, 155–170 (2021).
Dai, L. & Gao, G. F. Viral targets for vaccines against COVID-19. Nat. Rev. Immunol. 21, 73–82 (2021).
Telenti, A. et al. After the pandemic: perspectives on the future trajectory of COVID-19. Nature 596, 495–504 (2021).
Burki, T. K. Omicron variant and booster COVID-19 vaccines. Lancet Respir. Med. 10, e17 (2022).
Mahase, E. Covid-19: Omicron and the need for boosters. BMJ 375, n3079 (2021).
Jeyanathan, M. et al. Immunological considerations for COVID-19 vaccine strategies. Nat. Rev. Immunol. 20, 615–632 (2020).
Sette, A. & Crotty, S. Adaptive immunity to SARS-CoV-2 and COVID-19. Cell 184, 861–880 (2021).
Vabret, N. et al. Immunology of COVID-19: current state of the science. Immunity 52, 910–941 (2020).
Randolph, H. E. & Barreiro, L. B. Herd immunity: understanding COVID-19. Immunity 52, 737–741 (2020).
Sadarangani, M., Marchant, A. & Kollmann, T. R. Immunological mechanisms of vaccine-induced protection against COVID-19 in humans. Nat. Rev. Immunol. 21, 475–484 (2021).
Tao, K. et al. SARS-CoV-2 antiviral therapy. Clin. Mircobiol. Rev. 34, e0010921 (2021).
Asselah, T., Durantel, D., Pasmant, E., Lau, G. & Schinazi, R. F. COVID-19: discovery, diagnostics and drug development. J. Hepatol. 74, 168–184 (2021).
Siemieniuk, R. A. et al. Drug treatments for covid-19: living systematic review and network meta-analysis. BMJ 370, m2980 (2020).
Teijaro, J. R. & Farber, D. L. COVID-19 vaccines: modes of immune activation and future challenges. Nat. Rev. Immunol. 21, 195–197 (2021).
Leung, N. Transmissibility and transmission of respiratory viruses. Nat. Rev. Microbiol. 19, 528–545 (2021).
Meyerowitz, E. A., Richterman, A., Gandhi, R. T. & Sax, P. E. Transmission of SARS-CoV-2: a review of viral, host, and environmental factors. Ann. Intern. Med. 174, 69–79 (2021).
Ellul, M. A. et al. Neurological associations of COVID-19. Lancet Neurol. 19, 767–783 (2020).
Solomon, T. Neurological infection with SARS-CoV-2—the story so far. Nat. Rev. Neurol. 17, 65–66 (2021).
Clerkin, K. J. et al. COVID-19 and cardiovascular disease. Circulation 141, 1648–1655 (2020).
Crook, H., Raza, S., Nowell, J., Young, M. & Edison, P. Long covid-mechanisms, risk factors, and management. BMJ 374, n1648 (2021).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82030046, 81621004), Guangdong Science and Technology Department (2020B1212030004), and the Program for Guangdong Introducing Innovative and Entrepreneurial Teams (2019BT02Y198). BioRender (https://biorender.com/) is used to generate figures in Figs. 1c, 2, 5, 7, and Fig. 8.
Author information
Authors and Affiliations
Contributions
C.S. and M.S.Z. organized the manuscript. C.S. constructed the manuscript outline and wrote the manuscript. X.C., G.L.B., and L.Y.Z. participated in manuscript drafting and data collection. All authors have read and approved the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sun, C., Xie, C., Bu, GL. et al. Molecular characteristics, immune evasion, and impact of SARS-CoV-2 variants. Sig Transduct Target Ther 7, 202 (2022). https://doi.org/10.1038/s41392-022-01039-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-022-01039-2
- Springer Nature Limited
This article is cited by
-
Liver injury in COVID-19: an insight into pathobiology and roles of risk factors
Virology Journal (2024)
-
The symptoms and interval of Omicron SARS-CoV-2 reinfection among healthcare workers in a hospital of Southern China: a cross-sectional study
BMC Infectious Diseases (2024)
-
mRNA vaccine boosters and impaired immune system response in immune compromised individuals: a narrative review
Clinical and Experimental Medicine (2024)
-
Multiplexed discrimination of SARS-CoV-2 variants via plasmonic-enhanced fluorescence in a portable and automated device
Nature Biomedical Engineering (2023)
-
Elucidation of the neutralizing antibody evasion of emergent SARS-CoV-2 Omicron sub-lineages using structural analysis
Science China Life Sciences (2023)