Abstract
Autism Spectrum Disorder (ASD) is characterized mainly by social and sensory-motor abnormal and repetitive behavior patterns. Over hundreds of genes and thousands of genetic variants were reported to be highly penetrant and causative of ASD. Many of these mutations cause comorbidities such as epilepsy and intellectual disabilities (ID). In this study, we measured cortical neurons derived from induced pluripotent stem cells (iPSCs) of patients with four mutations in the genes GRIN2B, SHANK3, UBTF, as well as chromosomal duplication in the 7q11.23 region and compared them to neurons derived from a first-degree relative without the mutation. Using a whole-cell patch-clamp, we observed that the mutant cortical neurons demonstrated hyperexcitability and early maturation compared to control lines. These changes were characterized by increased sodium currents, increased amplitude and rate of excitatory postsynaptic currents (EPSCs), and more evoked action potentials in response to current stimulation in early-stage cell development (3–5 weeks post differentiation). These changes that appeared in all the different mutant lines, together with previously reported data, indicate that an early maturation and hyperexcitability may be a convergent phenotype of ASD cortical neurons.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD) was first defined by Leo Kanner in 1943 and named “early infantile autism” as an independent disorder from the psychotic disorder of schizophrenia, describing 11 children with social, biological, and emotional abnormalities such as the inability to relate to others or objects in a traditional way, an anxious and obsessive desire for consistency, early eating difficulties and hearing problems [1]. Recently, more symptoms entered the category of autistic-like behaviors such as attention deficit hyperactivity disorder (ADHD), poorly integrated verbal and non-verbal communication, abnormalities in eye contact, hyper or hypo-reactivity to sensory input, repetitive body movements, and more [2, 3].
On the genomic aspect, there has been more attention to many variants in which chromosomal subregions are deleted or duplicated in an inherited and de novo manner as well [2]. Various genes and mutations have been reported to be associated with ASD [4], and there seem to be a most definitive leaders: ADNP, CHD8, and SHANK3 [5].
On the neurobiological aspect, no specific brain area nor system has been confirmed to be entirely associated with the disorder, but an overall brain impairment has been shown starting from childhood [3]. The areas in the brain that are thought to be affected include cortical as well as non-cortical regions: the prefrontal cortex (PFC), Brodmann’s areas 9, 21, and 22, orbitofrontal cortex (OFC) fusiform gyrus, fronto-insular cortex, cingulate cortex, hippocampus, amygdala, cerebellum, and brainstem [6]. Our study focuses on the following four genetic mutations:
Dup7
7q11.23 Duplication Syndrome, briefly known as Dup7, is caused by a duplication of 1.5–1.8 Mb in section q11.23 of chromosome 7 [7], also known as the Williams-Beuren syndrome critical region (WBSCR) and is inherited in an autosomal dominant manner [8]. Extra copies of the genes are located in the critical region and include the Elastin gene (ELN)-encoding elastin, a structural protein and a critical component of elastic fibers [9], and the General Transcription Factor IIi gene (GTF2I)-encoding general transcription factor IIi, an inducible multifunctional transcription factor and a regulator of agonist-induced calcium entry to the cytoplasm [10]. These are thought to contribute significantly to the Syndrome’s symptoms [11, 12].
Common symptoms of Dup7 patients include anxiety disorders, selective mutism, ASD, ADHD, intellectual disability, seizures, severe speech delays, and hypotonia [13, 14] as well as Facial features including a broad forehead, a thin upper lip, and facial asymmetry [11]. Beyond that, magnetic resonance imaging (MRI) showed various abnormalities such as cerebral atrophy, increased cortical thickness, alterations in white matter’s volume and cerebellar hypoplasia, and an increased cortical extra-axial space [14].
SHANK3
The SHANK protein family, also known as ProSAP, including SHANK 1-3, is a family of scaffold proteins first identified using guanylate kinase-associated protein (GKAP)—a PSD-95-binding protein and a major component of the postsynaptic density (PSD), as bait [15, 16]. These proteins contain five key sites for protein-protein interaction: ankyrin repeats, Src homology 3 (SH3) domain, a vast proline-rich region, a C-terminal sterile α-motif (SAM) domain, and a PSD-95/discs large/zonula occludens-1 (PDZ) domain mediating its interaction with a variety of proteins as GKAP which moderates its binding to the NMDA and AMPA receptors [15, 17, 18].
The cognate gene, SHANK3 gene, is located at the terminal long arm of chromosome 22, coding for master scaffolding protein found in the body’s tissues and notably in the brain. It plays a critical role in the postsynaptic density of glutamatergic synapses and synaptic functions [15, 19]. Mutations in this gene have been linked to deficits in synaptic function and plasticity, accompanied by lower reciprocal social interactions [18, 20,21,22,23], ASD, and other neurodevelopmental symptoms [21, 24]. The individuals affected by the 22q13 deletion syndrome [24, 25], also known as Phelan-McDermid syndrome (PMS), suffer from developmental delays, a low muscular tone, a weakened perception of pain, delayed speech, seizures and ASD [26,27,28]. The patients also exhibit cortical alterations [29].
GRIN2B
Glutamate Ionotropic Receptor NMDA type Subunit 2B (GRIN2B) is a gene located on the ‘p’ arm of chromosome 12 (12p13.1 is its exact location) [30, 31]. This gene is one member of a family of genes encoding various proteins that together form the NMDA receptor [32]—glutamate-gated ion channel, which allows positively charged particles to flow through cells in the brain, activating neurons to send signals to each other [33, 34].
Mutations in this gene lead to a production of a nonfunctional GluN2B protein or completely prevent the production of GluN2B proteins from one copy of the gene in each cell; A shortage or dysfunction of this protein may cause an extreme reduction of the number of the functional NMDA receptors [33, 35, 36], causing neurodevelopmental disorders [37]. This disorder is associated with West syndrome, characterized by intellectual disability, delayed development of speech and motor skills, focal seizures, weak muscle tone (hypotonia), pediatric encephalopathy, movement disorders, schizophrenia, and behavioral delays common in ASD [30, 37,38,39,40,41,42]. In addition, MRI scans revealed a consistent malformation of cortical development (MCD) consistent with that of tubulinopathies, hypoplastic corpus callosum of varying degrees, enlarged and mildly dysplastic basal ganglia, hippocampal dysplasia as well as generalized cerebral volume loss [43]. GRIN2B-related disorders seem to have the highest prevalence among all GRIN-related disorders with 5.91 predicted incidence per 100,000 births [44]; This was previously reported to make up ~0.22% of the total neurodevelopmental disorders [43].
UBTF
The upstream binding transcription factor (UBTF) is a gene that codes for upstream binding factor (UBF)—a protein that acts as a transcription factor in RNA polymerase I (Pol I). UBF is critical for ribosomal RNA transcripts (rRNA) and synthesis from ribosomal DNA (rDNA) in the nucleolus [45]; A loss of UBF induces nuclear disruptions, including inhibition of cell proliferation, rapid and synchronous apoptosis as well as cell death [46]. Moreover, mutations in the UBTF gene lead to a production of an increased amount of ribosomal RNA transcript (rRNA), inducing depletion of RNA binding proteins, altered disposal machinery, and ribosome biogenesis [47]. These disruptions lead to pediatric neurodevelopmental regression starting at young onset (2.5 to 3 years). The degeneration starts with cognitive-motor deficits, reported in humans as well as knockout mice [48]. This regression is accompanied by dystonia, parkinsonism, severe intellectual disability, feeding difficulties, autistic-like behaviors, a slow loss of motor, cognitive, and speech capabilities, as well as severe epilepsy [47, 49,50,51,52]. Recent MRI studies showed gradual widespread brain atrophy in both supratentorial and infratentorial areas affecting both gray and white matter in patients with similar neuroregression patterns [47, 50]. Furthermore, increased pre-rRNA and 18S rRNA expression was reported in addition to nucleolar abnormalities quantified by quantitative real-time polymerase chain reaction (qRT-PCR), increased number of DNA breaks, defective cell cycle progression by comet assay, and apoptosis by TUNEL assay [47].
Materials and methods
After obtaining institutional committee approval and written informed consent from all participants or their respective legal guardians, iPSCs were generated from the peripheral blood mononuclear cells (PBMCs) of one male child carrying Dup7 (7q11.23 dup) (see supplementary Fig S1a), one female child carrying GRIN2B (c.2065 G- > T), one female child carrying SHANK3 [53] (C.3679insG) (see supplementary Fig S1b)and one female child carrying UBTF (E210K) mutations and a corresponding number of controls sharing the same genetic backgrounds (first-degree relatives from the same gender, except for GRIN2B), Table S1 presents a description of the patients’ cohort.
PBMC isolation
Participants had undergone a regular blood test (except for the SHANK3 cohort and GRIN2B-mutant line, in which iPSCs were generated from skin biopsies); blood was collected into heparin-coated tubes, diluted with phosphate-buffered saline (PBS) and centrifuged for 30 min at 1800 × g at 23 °C. The mononuclear cells (MCs) were collected by pipetting the buffy coat—the cell layer between the gel barrier and the plasma, into a sterile 15 ml conical centrifuge tube, adding PBS to reach a volume of 10 ml. This was followed by centrifuging (300 × g at room temperature for 15 min), resuspended using fetal bovine serum (FBS, F7524 Merck) 10% DMSO to freeze a total of ~2 × 106 cells per cryovial in 1 ml volume were aliquoted, frozen and kept in liquid nitrogen.
Reprograming of PBMCs into iPSCs
PBMCs were grown in a 24-well plate and seeded in fresh StemProTM SFM medium (10639011, Thermo Fisher Scientific). Sendai viral particle factors from CytoTune-iPS Sendai Reprogramming Kit (A16518, Life Technologies, Carlsbad, CA, USA) were added based on the manufacturer’s recommendations [54], followed by centrifuging the cells (1160 × g, at 35 °C, for 30 min) and seeded on a 24-well plate and an overnight incubation in 5% CO2 at 37 °C. The next day, the cells were transferred into a low attachment coated 12-well plate. Later, they were plated in six-well Matrigel-coated plates for 3 days and fed with a complete PBMC medium: StemPro™-34 supplement and 2 mM of L-Glutamine, followed by 2 days of gradually transitioning into iPSC medium (DMEM/F-12, GlutaMAX, KnockOut™ Serum Replacement, 10 mM MEM Non-Essential Amino Acids Solution and 55 mM β-mercaptoethanol). This was followed by daily replacing spent medium with fresh mTeSR for 20 days. When iPSC colonies were large enough, manual picking of the colonies was done, and colonies were transferred onto six-well Matrigel-coated plates. Immunocytochemistry of TRA-1-60, SOX2, and OCT4 antibodies confirmed pluripotency.
Generating neuronal cultures
Cortical neurons were plated based on a previously described protocol [55]. In brief, iPSCs were grown to ~80% confluency, embryonic bodies (EBs) were formed by mechanical dissociation using dispase for 20 min and plated onto low-adherence plates in mTeSR medium with ROCK inhibitor for 1 day, followed by 1 day of mTeSR medium only. On the following 10 days, cells were fed with EB media: (DMEM/F12 with Glutamax (1:100), B27 with Retinoic Acid (RA) (1:100), N2 supplement (1:100) and 0.1uM LDN), followed by plating onto polyornithine/laminin (Sigma)-coated dishes in DMEM/F12 (Invitrogen) plus N2, B27 and laminin for the following 7 days. The rosettes were selected based on their morphology and were manually picked, dissociated with Accutase (Chemicon) and plated onto poly-l-ornithine/laminin-coated plates and fed with complete NPC medium (DMEM/F12 with Glutamax (1:100), B27 supplement with RA (1:100), N2 supplement (1:100), laminin (1 mg/ml) and 20 ng/ml FGF2) for the following 15–20 days (based on their confluency). NPCs were then differentiated into cortical neurons by feeding with the differentiation medium that contained: DMEM/F12, N2, B27, Glutamax, ascorbic acid (200 nM), cyclic AMP (cAMP; 500 mg/ml), laminin (1 mg/ml), BDNF (20 ng/ml), GDNF (20 ng/ml) for 10 days. Between days 11 and 14, the cells were dissociated again and then fed with Brainphys medium with B27 supplement (1:100), N2 supplement (1:100), ascorbic acid (200 nM), cyclic AMP (500 mg/ml), BDNF (20 ng/ml), GDNF (20 ng/ml) and laminin (1 mg/ml).
Electrophysiology
Whole-cell patch-clamp recordings have been performed on neurons derived from patients carrying the Dup7, GRIN2B, SHANK3, and UBTF mutations, as well as neurons derived from healthy controls based on previously described [56] with some modifications [57], 3 to 5 weeks post differentiation. Culture coverslips were placed inside a recording chamber filled with HEPES-based artificial cerebrospinal fluid (ACSF) containing (in mM): 10 HEPES,139 NaCl, 4 KCl, 2 CaCl2, 10 D-glucose, and 1MgCl2 (pH 7.5, osmolarity adjusted to 310 mOsm). The recording micropipettes (tip resistance of 9–12 MΩ) were filled with an internal solution containing (in mM): 130 K-gluconate, 6 KCl, 4 NaCl, 10 Na-HEPES, 0.2 K-EGTA, 0.3 GTP, 2 Mg-ATP, 0.2 cAMP, 10 D-glucose, 0.15% biocytin and 0.06% rhodamine (pH 7.5, osmolarity adjusted to 290–300 mOsm). All measurements were done at room temperature using Clampex v11.1 with a sampling rate of 20 kHz.
Analysis of electrophysiological recordings
Analysis was performed based on previously described analysis [57] using custom-written MATLAB scripts modified as follows:
Synaptic currents analysis
The mean and standard error (SE) of the excitatory postsynaptic currents (EPSCs) amplitudes for each active cell were calculated. The cumulative distribution of EPSCs amplitude was calculated for each group. For each cell, the rate of the events was calculated by dividing the number of events by the time period of the recording (non-active cells were included and had an event rate=0). The mean of all cells’ rates and the standard error of the frequencies were computed for the control and mutant groups. Non-parametric statistical tests (Wilcoxon signed-rank test) were performed for comparisons.
Sodium, fast, and slow potassium currents
Neurons were held in voltage clamp mode at −60 mV, and voltage steps of 400 ms were performed in the −100 to 90 mV range. Currents were typically normalized by the cells’ capacitance; the sodium current was computed by subtracting the sodium current after stabilization from the lowest value of the inward sodium current. The fast potassium currents were measured by the maximum outward currents that appeared within a few milliseconds after a depolarization step. The slow potassium currents were measured after the 400 ms depolarization phase. A one-way ANOVA test was performed for the statistical analysis.
Evoked action potentials (APs)
Neurons were held in current clamp mode at −60 mV with a constant holding current. Following this, current injections were given in 3-pA steps throughout 400 ms, starting 12 pA below the steady-hold current. A total of 38 depolarization steps were given. The total evoked action potential was the total number of action potentials that were counted in the 38 depolarization steps. Non-parametric statistical tests (Wilcoxon signed-rank test) were performed for comparisons.
Action potential shape analysis
The first evoked action potential generated with the lowest amount of injected current was used for spike shape analysis. The spike threshold was defined as the membrane potential at which the slope of the depolarizing membrane potential increased dramatically, resulting in an AP (the second derivative of the voltage versus time as the initial maximum). The spike height was calculated as the difference between the highest membrane potential during a spike and the threshold. The spike rise time is the time it takes the spike to reach the maximum. The spike width was calculated as the time it took the membrane potential to reach half of the spike amplitude in the rising part.
Analysis of brightfield images
Brightfield images of NPCs from both control and patient cultures were captured using a Nikon Eclipse Ts2 microscope with ×20 magnification. Neurite’s length was measured manually using ImageJ, statistically tested with Wilcoxon’s signed-rank test and visualized using MATLAB.
Immunocytochemistry (ICC)
Cells were fixed in 4% paraformaldehyde for 15 min, followed by three washes of DPBS, blocked and permeabilized in PBS containing 0.1–0.2% Triton X-100 and 10% horse serum. The coverslips were incubated with primary antibodies; for NPCs: rabbit anti-PAX6 (CST, mAb#60433, 1:250) and mouse anti-NESTIN (CST, mAb#33475,1:2000); for Neurons: chicken anti-MAP2 (Abcam, ab92434, 1:500), rabbit anti-TBR1 (Abcam, ab183032, 1:250), rabbit anti-VGLuT1 (Abcam, ab227805, 1:500) and Mouse anti-GABA (Abcam, ab86186, 1:400) in the blocking solution overnight at 4 °C. On the next day, they were washed in DPBS and incubated with DAPI (Abcam, ab228549, 1:1000) and corresponding secondary antibodies (Abcam, ab150084, ab150117, ab175711, 1:250) for 60 min at room temperature. Then the coverslips were washed three times, mounted on glass slides using Fluromount-G (mounting medium), and dried overnight while being protected from light. Fluorescence signals were detected using a Leica THUNDER imager and analyzed using ImageJ and MATLAB.
Results
Dup7 cortical neurons display increased sodium and potassium currents, increased synaptic activity and hyperexcitability early in the differentiation
We performed whole-cell patch clamp experiments 5 weeks (day 34) after the start of the differentiation of 16 Dup7-mutant neurons and 13 control neurons derived from a first-degree relative of the same gender. In a voltage clamp mode, EPSC recordings were performed by holding the cell at −60 mV. We observed an increase in the rate of EPSCs of the mutant neurons compared to the controls (0.13 ± 0.08 Hz in Dup7-mutant neurons and 0.07 ± 0.07 Hz in the control neurons, p = 0.03), as shown in Fig. 1a–c. Figure 1a, b presents representative traces, while Fig. 1c represents the average over all the recordings. In addition, a significant increase in the mean amplitude of the EPSCs was observed. The Dup7-mutant neurons had a larger amplitude compared to the control neurons (12.72 ± 2.86 pA for the mutant neurons and 7.12 + 5.09 pA for the control neurons (p = 0.002, (Fig. 1d)). The cumulative distribution of the EPSC amplitudes for Dup7-mutant neurons is slightly right-shifted compared to control neurons indicating larger amplitudes of EPSCs (Fig. 1e).
a A representative trace of (EPSCs measured in control cortical neurons at 5 weeks post-differentiation. b A representative trace of EPSCs measured in a dup7-mutant neuron at 5 weeks post-differentiation. c The mean rate of synaptic events was higher in the dup7-mutant neurons compared to control neurons (p = 0.029). d The average amplitude of EPSCs was increased in the dup7-mutant neurons (p = 0.002). e The cumulative distribution of the amplitude of EPSCs is slightly right-shifted in the dup7-mutant neurons, indicating an increase in the amplitudes. f A Representative trace of sodium and potassium currents recorded in a voltage-clamp mode in control neurons. g A Representative trace of sodium and potassium currents recorded in a voltage-clamp mode in dup7-mutant neurons. h The average sodium currents in dup7-mutant neurons is increased compared to control neurons (p = 0.004). i The average slow potassium currents in dup7-mutant neurons is increased compared to control neurons (p = 0.03). j The average fast potassium currents is increased in dup7-mutant compared to control neurons (p = 0.01). k A representative recording of evoked action potentials in a current-clamp mode of a control neuron. l A representative recording of evoked action potentials in a current-clamp mode of a dup7-mutant neuron. m The total number of evoked action potentials is larger in dup7-mutant neurons compared to control neurons (p = 0.006). n, o A representative image of control n and mutant o NPCs that were immunostained for DAPI, PAX6, and Nestin. p, q A representative image of control p and mutant q neurons that were immunostained for DAPI, MAP2, and TBR1.
Next, we recorded in voltage clamp mode the sodium and potassium currents. We observed a significantly larger normalized sodium current in the Dup7-mutant neurons compared to the control neurons (F (1,38) = 9.43, p = 0.004). Representative traces of the recordings are shown in Fig. 1f (control) and 1g (mutant). The average sodium currents are presented in Fig. 1h. In addition, we observed increased slow and fast potassium currents (normalized by the capacitance) in the Dup7-mutant neurons compared to controls (F (1,14) = 5.61; p = 0.03 for the slow potassium currents and F (1,14) = 8.36; p = 0.01 for the fast potassium currents) over the 10–80 mV range (Fig. 1h–j). We next measured the number of evoked action potentials in a current clamp mode as a measure of the neuronal excitability. We observed a hyperexcitability pattern for the Dup7-mutant neurons compared to control neurons. The total number of evoked potentials (see “Materials and methods”) for the Dup7-mutant neurons was 47.63 ± 49.61; for the control neurons, it was 28.5 ± 69.48 (p = 0.006). A representative example is presented in Fig. 1k (control) and 1l (Dup7), and the average over all recordings is shown in Fig. 1m. Spike shape analysis (see “Materials and methods”) is presented in Table S2. Examples of ICC images for control and mutant lines are shown in Fig. 1n, o (typical NPC markers PAX6 and NESTIN) and 1p, q (neuronal markers MAP2 and the cortical marker TBR1).
GRIN2B cortical neurons display increased sodium and potassium currents and hyperexcitability early in the differentiation
We performed whole-cell patch clamp experiments 3 weeks (day 19) after the start of the differentiation of 12 GRIN2B-mutant neurons and 5 weeks (day 27) of 7 control neurons. In a voltage clamp mode, EPSC recordings were performed by holding the cell at −60 mV. We observed an increase in the rate of EPSCs of the mutant neurons compared to the controls (0.39 ± 0.27 Hz in GRIN2B-mutant neurons and 0.17 ± 0.2 Hz in the control neurons, p = 0.03) as shown in Fig. 2a–c. Figure 2a, b presents representative traces, while Fig. 2c represents the average over all the recordings. No significant difference in the mean amplitude of the EPSCs was observed (Fig. 2d). The cumulative distribution of the EPSC amplitudes was similar between the control and GRIN2B-mutant neurons (Fig. 2e).
a A representative trace of EPSCs measured in control cortical neurons at 5 weeks post-differentiation. b A representative trace of EPSCs measured in a GRIN2B-mutant neuron at 3 weeks post-differentiation. c The mean rate of synaptic events was higher in the GRIN2B-mutant neurons (p = 0.03). d The mean amplitude of EPSCs was increased but not significantly different in the GRIN2B-mutant neurons. e The cumulative distribution of the amplitude of the EPSCs of GRIN2B-mutant and control neurons looks similar. f A Representative trace of sodium and potassium currents recorded in a voltage-clamp mode in control neurons. g A Representative trace of sodium and potassium currents recorded in voltage-clamp in GRIN2B-mutant neurons. h The average sodium currents in GRIN2B-mutant neurons is severely increased compared to control neurons (p = 0.006). I The average slow potassium currents in GRIN2B-mutant neurons is increased compared to control neurons (p = 0.04). j The average fast potassium currents in GRIN2B-mutant neurons is increased compared to control neurons (p = 0.03). k A representative recording of evoked action potentials in a current-clamp mode of a control neuron. l A representative recording of evoked action potentials in a current-clamp mode of a GRIN2B-mutant neuron. m The total number of evoked action potentials is larger in the GRIN2B-mutant neurons compared to control neurons (p = 0.003). n, o A representative image of control (n) and mutant (o) NPCs that were immunostained for DAPI, PAX6, and Nestin. p, q A representative image of control (p) and mutant (q) neurons that were immunostained for DAPI, MAP2, and TBR1.
Next, we recorded in voltage clamp mode the sodium and potassium currents. We observed a significantly larger normalized sodium current in the GRIN2B-mutant neurons compared to the control neurons (F (1,38) = 14.1, p = 0.0006). Representative traces of the recordings are shown in Fig. 2f (control) and 2g (mutant). The average sodium currents are presented in Fig. 2h. In addition, we observed larger slow and fast potassium currents (normalized by the capacitance) in the GRIN2B-mutant neurons compared to controls (F (1,16) = 4.6, p = 0.04 for the slow potassium currents and F (1,16) = 5.39, p = 0.03 for the fast potassium currents) over the 0–80 mV range (Fig. 2h–j).
We next measured the number of evoked action potentials in current clamp mode as a measure of the neuronal excitability. We observed a hyperexcitability pattern for the GRIN2B-mutant neurons compared to control neurons. The total number of evoked potentials (see “Materials and methods”) in the GRIN2B-mutant neurons was 38.86 ± 10.17, and in the control neurons, it was 22.37 ± 19.5 (p = 0.003). A representative example is presented in Fig. 2k (control) and 2l (GRIN2B), and the average over all recordings is presented in Fig. 2m. Furthermore, we observed a significant increase in GRIN2B-mutant neurons’ spike amplitude compared to control neurons (41.2 ± 12.5 mV in GRIN2B-mutant neurons and 20.5 ± 14.01 mV in control neurons, p = 0.008); further spike shape analysis is presented in Table S2. Examples of ICC images for control and mutant lines are shown in Fig. 2n, o (typical NPC markers PAX6 and NESTIN) and 2p, q (neuronal markers MAP2, the cortical marker TBR1).
SHANK3 cortical neurons display increased sodium and slow potassium currents, a drastic increase in synaptic activity and hyperexcitability early in the differentiation
We performed whole-cell patch clamp experiments 5 weeks (days 29–32) after the start of the differentiation of 21 SHANK3-mutant and 17 control neurons. In a voltage clamp mode, EPSC recordings were performed by holding the cell at −60 mV. We observed a significant increase in the rate of EPSCs of the mutant neurons compared to the controls (0.28 ± 0.36 Hz in SHANK3-mutant neurons and 0.08 ± 0.06 Hz in the control neurons, p 0.003) as shown in Fig. 3a–c. Figure 3a, b presents representative traces, while Fig. 3c represents the average over all the recordings. In addition, a drastic increase in the mean amplitude of the EPSCs was observed. The SHANK3-mutant neurons had a larger amplitude compared to the control neurons (10.145 ± 3.26 pA for the mutant neurons and 5.29 + 1.65 pA for the control neurons, p = 1.15e−5 (Fig. 3d)). The cumulative distribution of the EPSC amplitudes for SHANK3-mutant neurons is right shifted compared to control neurons indicating larger amplitudes of EPSCs (Fig. 3e).
a A representative trace of excitatory postsynaptic currents (EPSCs) that were measured in control cortical neurons at 5 weeks post-differentiation. b A representative trace of EPSCs measured in a SHANK3-mutant neuron at 5 weeks post-differentiation. c The mean rate of synaptic events was higher in the SHANK3-mutant neurons (p = 0.003). d The average amplitude of EPSCs was increased in the SHANK3-mutant neurons (p = 1.15e−5). e The cumulative distribution of the amplitude of EPSCs of SHANK3-mutant is right-shifted, indicating an increase in the amplitudes. f A Representative trace of sodium and potassium currents recorded in voltage-clamp in control neurons. g A Representative trace of sodium and potassium currents recorded in a voltage-clamp mode in SHANK3-mutant neurons. h The average sodium currents in SHANK3-mutant neurons is increased compared to control neurons (p = 0.04). I The average slow potassium currents in SHANK3-mutant neurons is increased compared to control neurons (p = 0.03). j The average fast potassium currents look similar and not significantly different in SHANK3-mutant compared to control neurons. k A representative recording of evoked action potentials in a current-clamp mode of control neurons. l A representative recording of evoked action potentials in a current-clamp mode of SHANK3-mutant neurons. m The total number of evoked action potentials is larger in SHANK3-mutant neurons compared to control neurons (p = 0.03). n, o A representative image of control (n) and mutant (o) NPCs that were immunostained for DAPI, PAX6, and Nestin. p, q A representative image of control (p) and mutant (q) neurons that were immunostained for DAPI, MAP2, and TBR1.
Next, we recorded in voltage clamp mode the sodium and potassium currents. We observed a significantly larger normalized sodium current in the SHANK3-mutant neurons compared to the control neurons F (1,36) = 4.51, p = 0.04. Representative traces of the recordings are shown in Fig. 3f (control) and 3g (mutant). The average sodium currents are presented in Fig. 3h. In addition, we observed increased slow, but not fast, potassium currents (normalized by the capacitance) in the SHANK3-mutant neurons compared to controls, (F (1,10) = 5.68; p = 0.03) over the 40–90 mV range (Fig. 3h–j).
We next measured the number of evoked action potentials in current clamp mode as a measure of the neuronal excitability. We observed a hyperexcitability pattern for the SHANK3-mutant neurons compared to control neurons. The total number of evoked potentials (see “Materials and methods”) for the SHANK3-mutant neurons was 31.32 ± 39.8; for the control neurons, it was 18.4 ± 22.96 (p = 0.03). A representative example is presented in Fig. 3k (control) and 3l (SHANK3) and the average over all recordings are presented in Fig. 3m. Furthermore, we observed a more depolarized threshold (higher) in the SHANK3-mutant neurons compared to control neurons (30.4 ± 5.6 mV in the SHANK3-mutant neurons and 25.5 ± 4.2 mV in control neurons, p = 0.02); further spike shape analysis is presented in Table S2. Examples of ICC images of control and mutant lines are shown in Fig. 3n, o (typical NPC markers PAX6 and NESTIN) and 3p, q (neuronal markers MAP2, the cortical marker TBR1).
UBTF cortical neurons display increased sodium currents, an increase in synaptic amplitude, an increase in spontaneous activity and hyperexcitability early in the differentiation
We performed whole-cell patch clamp experiments 5 to 6 weeks (days 32–37) after the start of the differentiation of 22 UBTF-mutant neurons and 5 to 6 weeks (days 32–37) after the beginning of the differentiation of 18 control neurons. In a voltage clamp mode, EPSC recordings were performed by holding the cell at −60 mV. Figure 4a, b presents representative traces, while Fig. 4c represents the average EPSC amplitude over all the recordings. The EPSC rate of the UBTF-mutant neurons is slightly similar to the control neurons (Fig. 4c). We observed an increase in the mean amplitude of the EPSCs. The UBTF-mutant neurons had a larger amplitude compared to control neurons it was 7.81 ± 1.16 pA for the mutant neurons and 6.55 + 2.39 pA for control neurons, p = 0.02 (Fig. 4d). The cumulative distribution of the EPSC amplitudes for UBTF-mutant neurons is slightly right-shifted compared to control neurons indicating larger amplitudes of EPSCs (Fig. 4e).
a A representative trace of excitatory postsynaptic currents (EPSCs) that were measured in control cortical neurons at 6 weeks post-differentiation. b A representative trace of EPSCs measured in a UBTF-mutant neuron at five weeks post-differentiation. c The mean rate of synaptic events was not significantly different in UBTF-mutant neurons compared to control neurons. d The average amplitude of EPSCs was increased in the UBTF-mutant neurons (p = 0.02). e The cumulative distribution of the amplitude of EPSCs of UBTF-mutant neurons is slightly right-shifted, indicating an increase in the amplitudes. f A Representative trace of sodium and potassium currents recorded in a voltage-clamp mode in control neurons. g A Representative trace of sodium and potassium currents recorded in a voltage-clamp mode in UBTF-mutant neurons. h The average sodium currents in UBTF-mutant neurons is increased compared to control neurons (p < 0.05). I The average slow potassium currents look similar in the UBTF-mutant and control neurons. j The average fast potassium currents look identical in the UBTF-mutant and control neurons. k A representative recording of evoked action potentials in the current-clamp mode of control neurons. l A representative recording of evoked action potentials in a current-clamp mode of UBTF-mutant neurons. m The total number of evoked action potentials is higher in the UBTF-mutant neurons compared to control neurons (p = 5.56e−4). n The mean rate of spontaneous action potentials was significantly higher in UBTF-mutant neurons compared to control neurons (p = 0.013). o The mean amplitude of spontaneous action potentials in UBTF-mutant neurons was larger compared to control neurons (p = 0.012). p, q A representative image of control (p) and mutant (q) NPCs that were immunostained for DAPI, PAX6, and Nestin. r, s A representative image of control (r) and mutant (s) neurons that were immunostained for DAPI, MAP2, and TBR1.
Next, we recorded in voltage clamp mode the sodium and potassium currents. We observed a significantly larger normalized sodium current in the UBTF-mutant neurons compared to the control neurons F (1,28) = 5.58, p = 0.03 over the −50–90-mV range. Representative traces of the recordings are shown in Fig. 4f (control) and 4 g (mutant). The average sodium currents is presented in Fig. 4h. An ANOVA test indicated no significant differences in the slow and fast potassium currents (Fig. 4h–j).
We next measured the number of evoked action potentials (see “Materials and methods”). For the UBTF-mutant neurons, it was 62.95 ± 31.56 and for the control neurons, it was 31.46 ± 22.3 (p = 5.56e−4). A representative example is presented in Fig. 4k (control) and 4 l. (UBTF) and the average over all recordings are presented in Fig. 4m. Furthermore, we observed a significant increase in UBTF-mutant neurons’ spike amplitude compared to control neurons (50 ± 17.05 mV in the UBTF-mutant neurons and 36.04 ± 18.01 mV in control neurons, p = 0.03). Besides, we observed a narrower spike in the UBTF-mutant neurons compared to the control neurons (3.3 ± 1.5 ms in the UBTF-mutant neurons and 12.5 ± 17.8 ms in the control neurons, p = 0.003); further spike shape analysis is presented in Table S2. The spontaneous neuronal activity (spontaneous action potentials) was measured in a holding potential of −45 mV. We observed a significant increase in the spontaneous activity rate (p = 0.013) and amplitudes (p = 0.012) in the UBTF-mutant neurons compared to control neurons (Fig. 4n, o). Examples of ICC images of control and mutant lines are shown in Fig. 4p, q (typical NPC markers PAX6 and NESTIN) and 4r, s (neuronal markers MAP2, the cortical marker TBR1).
Mutant neural progenitor cells exhibit longer neurite lengths compared to controls, alongside with decreased GABA-positive neurons in ASD-related mutant neuronal cultures
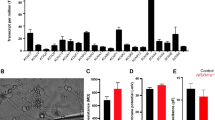
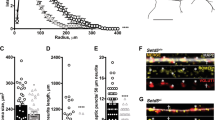
We observed that the lengths of neurites of neural progenitor cells (NPCs) in mutant groups were significantly increased compared to control groups (Fig. 5a). This difference was determined by analyzing brightfield images captured at ×20 magnification and measuring the length of neurites using a standardized methodology (see “Materials and methods”). Specifically, the average length of NPC neurites in the Dup7-mutant NPCs (left) was 0.12 ± 0.02 cm compared to control NPCs was 0.07 ± 0.02 cm (p = 4.36e−04), in the SHANK3-mutant NPCs (middle) the average length was 0.09 ± 0.03 cm compared to control NPCs 0.07 ± 0.02 cm (p = 6.75e−08) and in the UBTF-mutant NPCs it was 0.09 ± 0.03 cm compared to control NPCs 0.05 ± 0.01 cm (p = 0.01), as determined by Wilcoxon signed-rank test. Example images of control (b) and patient (c) are shown in Fig. 5b, c.
a A longer neurites in Dup7-NPCs (left) p = 4.36e−04, SHANK3-NPCs (middle) p = 6.75e−08 and UBTF-NPCs (right) p = 0.01 mutants compared to control groups. b The averages of the percentage of GABA-expressing neurons (left) out of the total neurons in the ASD-mutant neuronal cultures are significantly lower than the percentage of GABA-expressing neurons in the control neuronal cultures (p = 0.04), The averages of the percentage of VGluT1-expressing neurons (middle) out of the total neurons in the ASD-mutant neuronal cultures are similar to the percentage of VGluT1-expressing neurons in the control neuronal cultures (p = 0.56) and a higher E-I ratio (right) in ASD-mutant neuronal cultures compared to control cultures (p = 0.03). c, d A representative image of mutant (c) and control (d) neurons that were immunostained for DAPI, GABA, and VGluT1.
Next, we used GABA and VGluT1 antibodies to confirm the existence of GABA and VGluT1-positive cells in our cultures. Indeed, the control cultures contained 15.08 ± 1.13% GABA-expressing and 77.87 ± 2% VGluT1-expressing neurons. However, the ASD-related mutant cultures contained only 11.73 ± 1.14% GABA-expressing and 79.99 ± 2.93% VGluT1-expressing neurons (PGABA = 0.04, PVGluT1 = 0.56). In addition, we observed a higher ratio of excitation–inhibition (E-I ratio) in the Patient group compared to Control (8.183 ± 1.05 and 5.51 ± 0.38, respectively; p = 0.03) as shown in Fig. 5d. Figure 5e, f shows an example image of control (e) and mutant (f) neuronal cultures that were immunostained for DAPI (blue), VGluT1 (red), and GABA (green).
Discussion
In this study, iPSC technology was used to investigate the physiological features of cortical neurons derived from human patients with different ASD-related mutations: Dup7, GRIN2B, SHANK3, and UBTF. For that purpose, we differentiated patient-derived iPSCs into cortical neurons [55] since cortical alterations and malformations within the brain were identified in these mentioned mutations [14, 29, 43, 50].
A broad range of mutations has been associated with ASD, sharing autistic-like behaviors such as difficulties in social communication and interaction and restricted or repetitive behaviors or interests. The broad spectrum of mutations is characterized by affecting different genes and pathways. Various genes affecting the cytoskeletal [58,59,60,61,62] and microtubule [63] dynamics are altered in ASD-associated genes, disturbing a critical stage in axons and dendrites development [64]; For Dup7 mutation, for example, a rare genetic syndrome caused by a micro-duplication in section q11.23 of chromosome 7, alterations in the ELN gene such as inherited and de novo deletions were reported, coding for the extracellular matrix protein, elastin, associated with connective-tissue malformations as reported in human patients [12]. SHANK3 is a gene located at the terminal long arm of chromosome 22, coding for a master scaffolding protein found in the body’s tissues, and importantly in the brain, and has a critical role in the postsynaptic density of glutamatergic synapses and synaptic functions in rat and human brains [15, 19]. The GRIN2B mutation results in a production of a nonfunctional GluN2B protein. A shortage or dysfunction of this protein may cause an extreme reduction in the number of the functional NMDA receptors [35] causing neuronal impairments in both mice and human models [65, 66]. UBTF is a gene coding for UBF, a transcription factor in RNA Pol I, critical for rRNA transcripts synthesis from rDNA in the nucleolus [45]. Loss of UBF induces nuclear disruptions, including inhibition of cell proliferation, rapid and synchronous apoptosis, and cell death in mice models [46].
Although these mutations are functionally very different from one another, they cause similar symptoms in the patients. Interestingly, we observed a hyperexcitability pattern in all these mentioned ASD-related mutations in an early-stage cell development (3–5 weeks post differentiation) that we measured by electrophysiological recordings. This hyperexcitability involved different aspects; In terms of sodium-potassium currents and activity, a consistent increase in sodium currents was observed within the four patient-derived neurons, which could, in turn, increase the excitability of the neurons by decreasing the action potential threshold [67]. These alterations of sodium currents can lead to abnormal neuronal activity, a phenomenon that also occurs in epilepsy [68]. It is interesting to note that all these four mutations have a strong association with epilepsy, and many of the patients also suffer from epilepsy [14, 27, 44, 53]. We also observed an increase in the EPSC rate and amplitude, indicating pre and postsynaptic changes that occur in the mutant neurons compared to the controls neurons. Similar findings were reported in mice models of autism [69,70,71,72,73]. Furthermore, we observed more evoked action potentials in response to current stimulation in the mutant-derived neurons, alongside with a decrease in GABA-positive cells and an increase in the E-I ratio, which may contribute to the occurrence of infantile epileptic seizures in affected human subjects. We previously reported such changes also in another ASD model of an IQSEC2 mutation [74]. Furthermore, the NPCs derived from individuals with ASD-related mutations display longer neurite-like branches compared to control groups supporting our conclusions of their early maturation.
All these changes can indicate that the ASD mutant neurons develop faster, and at this early stage, when the control neurons are still very immature, they are already spiking and connecting with other neurons.
Several previous genetic studies showed a rise in cortical activity by documenting excitation-to-inhibition ratio system alterations [74,75,76], suggesting that periodic seizures and sensory hyperreactivity in ASD are caused by cortical hyperexcitability [77, 78], which was also previously observed in fragile-X syndrome [79]. Previously, we reported a similar early time point hyperexcitability pattern in another ASD-related mutation—the A350V IQSEC2mutation [74]. In that study, we followed the IQSEC2-mutant neurons that started more active and more connected (5 weeks post differentiation) as they became hypoexcitable with reduced synaptic connections later on in the differentiation process. An additional hyperexcitability pattern in 15q11-q13 Duplication Syndrome, a different ASD-associated mutation, was reported [80]. A reduction in synaptic connections was reported in a long line of ASD-related and ID-related studies using mice and human models [6, 81,82,83], suggesting a potential shared accelerated aging mechanisms in ASD-associated mutations [62, 63, 84,85,86].
We speculate that there may be a connection between this early hyperexcitability and the later synaptic deficits, as perhaps this early hyperexcitability is neurotoxic to the cell at such an early stage of development.
Since, especially with mice studies, it is much harder to measure this early developmental stage, perhaps this is a stage that precedes the synaptic degradation in many other ASD mutations. More evidence of this early maturation was presented in a study with ASD patients with macrocephaly where the neurons derived from the patients were more arborized early in the development, and gene expression profiles also suggested an earlier maturation [87]. More evidence for functional hyperactivity in epilepsy and ASD-related mutations in human models were reported; briefly, an engineered iPSC-derived neuron with the homozygous P924L mutation (one of many epilepsy-associated Slack mutations) displayed increased KNa currents and more evoked action potentials in both single neurons and a connected neuronal network [88].
Our Findings present a shared phenotype of early maturation and hyperexcitability in four ASD-related mutations using patient-derived cortical neurons, indicating that there may be a common neurophysiological phenotype in ASD-related variants, sharing similar behavioral phenotypes but a different genotype. iPSC-derived neurons were previously used as a research tool for investigating physiological and cellular alterations characterizing various disorders including autism [74, 87, 89, 90], epilepsy [88, 91, 92] and other neurodegenerative diseases such as Parkinson’s [93,94,95,96] and schizophrenia [97,98,99]. Here we concentrated on the early developmental physiological alterations in 4 different ASD and epilepsy-related genes. The enhanced maturation and excitability in such young neurons may be deleterious to the cells and may later result in synaptic degeneration as was previously described in neurons derived from ASD and epilepsy patients [74, 83, 87,88,89, 91, 92].
Code availability
The custom-written scripts used for the analysis can be shared upon reasonable request.
References
Kanner L. Autistic disturbances of affective contact. Nerv Child. 1943;2:217–50.
Lord C, Elsabbagh M, Baird G, Veenstra-Vanderweele J. Autism spectrum disorder. Lancet. 2018;392:508–20.
Chaste P, Leboyer M. Autism risk factors genes environment and gene-environment interactions. Dialogues Clin Neurosci. 2012;14:281–92.
Devlin B, Boone BE, Levy SE, Lihm J, Buxbaum JD, Wu Y, et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature. 2012;485:242–6. p
Larsen E, Menashe I, Ziats MN, Pereanu W, Packer A, Banerjee-Basu S. A systematic variant annotation approach for ranking genes associated with autism spectrum disorders. Mol Autism. 2016;7:44.
Varghese M, Keshav N, Jacot-Descombes S, Warda T, Wicinski B, Dickstein DL, et al. Autism spectrum disorder: neuropathology and animal models. Acta Neuropathol. 2017;134:537–66.
Francke U. Williams-Beuren syndrome: genes and mechanisms. Hum Mol Genet. 1999;8:1947–54.
Mervis CB, Morris CA, Klein-Tasman BP, Velleman SL, Osborne LR. 7q11.23 duplication syndrome. GeneReviews. Seattle; 2015.
Debelle L, Tamburro AM. Elastin: molecular description and function. Int J Biochem Cell Biol. 1999;31:261–72.
Caraveo G, Van Rossum DB, Patterson RL, Snyder SH, Desiderio S. Action of TFII-I outside the nucleus as an inhibitor of agonist-induced calcium entry. Science 2006;314:122–5.
Merla G, Brunetti-Pierri N, Micale L, Fusco C. Copy number variants at Williams-Beuren syndrome 7q11.23 region. Hum Genet. 2010;128:3–26.
Ewart’ AK, Morris CA, Atkinson D, Jin’ W, Sternes K, Spallone P, et al. Hemizygosity at the elastin locus in a developmental disorder, Williams syndrome. Nat Genet. 1993;5:11–6.
Sanders SJ, Ercan-Sencicek AG, Hus V, Luo R, Murtha MT, Moreno-De-Luca D, et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 2011;70:863–85.
Morris CA, Mervis CB, Paciorkowski AP, Abdul-Rahman O, Dugan SL, Rope AF, et al. 7q11.23 Duplication syndrome: physical characteristics and natural history. Am J Med Genet A 2015;167:2916–35.
Naisbitt S, Kim E, Cheng TU J, Xiao B, Sala C, Valtschanoff J, et al. Shank, a novel family of postsynaptic density proteins that binds to the NMDA receptor/PSD-95/GKAP complex and cortactin. Neuron. 1999;23:569–82.
Sheng M. Glutamte receptors put in their place. Nature 1997;386:221–3.
Sheng M, Kim E. The shank family of scaffold proteins. J Cell Sci. 2000;113:1851–6.
Bozdagi O, Sakurai T, Papapetrou D, Wang X, Dickstein DL, Takahashi N, et al. Haploinsufficiency of the autism-associated Shank3 gene leads to deficits in synaptic function, social interaction, and social communication. Mol Autism. 2010;1:15.
Boeckers TM. The postsynaptic density. Cell Tissue Res. 2006;326:409–22.
Monteiro P, Feng G. SHANK proteins: roles at the synapse and in autism spectrum disorder. Nat Rev Neurosc. 2017;18:147–57.
Durand CM, Betancur C, Boeckers TM, Bockmann J, Chaste P, Fauchereau F, et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet. 2007;39:25–7.
Peça J, Feliciano C, Ting JT, Wang W, Wells MF, Venkatraman TN, et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 2011;472:437–42.
Amal H, Barak B, Bhat V, Gong G, Joughin BA, Wang X, et al. Shank3 mutation in a mouse model of autism leads to changes in the S-nitroso-proteome and affects key proteins involved in vesicle release and synaptic function. Mol Psychiatry. 2020;25:1835–48.
Wilson HL, Wong ACC, Shaw SR, Tse WY, Stapleton GA, Phelan MC, et al. Molecular characterisation of the 22q13 deletion syndrome supports the role of haploinsufficiency of SHANK3/PROSAP2 in the major neurological symptoms. J Med Genet. 2003;40:575–84.
Bonaglia MC, Giorda R, Mani E, Aceti G, Anderlid BM, Baroncini A, et al. Identification of a recurrent breakpoint within the SHANK3 gene in the 22q13.3 deletion syndrome. J Med Genet. 2006;43:822–8.
Phelan K, Curtis Rogers R, Boccuto L. Phelan-McDermid syndrome. GeneReviewsTM. Seattle, WA; 2005.
Soorya L, Kolevzon A, Zweifach J, Lim T, Dobry Y, Schwartz L, et al. Prospective investigation of autism and genotype-phenotype correlations in 22q13 deletion syndrome and SHANK3 deficiency. Mol Autism. 2013;4:18.
Holder JL, Quach MM. The spectrum of epilepsy and electroencephalographic abnormalities due to SHANK3 loss-of-function mutations. Epilepsia 2016;57:1651–9.
Li D, Liu C, Huang Z, Li H, Xu Q, Zhou B, et al. Common and distinct disruptions of cortical surface morphology between autism spectrum disorder children with and without SHANK3 deficiency. Front Neurosci. 2021;15:751364.
Dimassi S, Andrieux J, Labalme A, Lesca G, Cordier MP, Boute O, et al. Interstitial 12p13.1 deletion involving GRIN2B in three patients with intellectual disability. Am J Med Genet A 2013;161:2564–9.
Mandich P, Schito AM, Bellone E, Antonacci R, Finelli P, Rocchi M, et al. Mapping of the Human NMDAR2B Receptor subunit Gene (GRIN2B) to chromosome 12p12. Genomics 1994;22:216–8.
Monyer H, Burnashev N, Laurie DJ, Sakmann B, Seeburg PH. Developmental and regional expression in the rat brain and functional properties of four NMDA receptors. Neuron. 1994;12:529–40.
Cull-Candy S, Brickley S, Farrant M. NMDA receptor subunits: diversity, development and disease. Curr Opin Neurobiol. 2001;11:327–35.
Traynelis SF, Wollmuth LP, McBain CJ, Menniti FS, Vance KM, Ogden KK, et al. Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev. 2010;62:405–96. p
Akashi K, Kakizaki T, Kamiya H, Fukaya M, Yamasaki M, Abe M, et al. NMDA receptor GluN2B (GluRε2/NR2B) subunit is crucial for channel function, postsynaptic macromolecular organization, and actin cytoskeleton at hippocampal CA3 synapses. J Neurosci. 2009;29:10869–82.
Kellner S, Berlin S. Two extreme Loss-of-Function GRIN2B-mutations are detrimental to tri-heteromeric NMDAR-function, but rescued by pregnanolone-sulfate. bioRxiv:520218v12022 [Preprint]. 2022. Available from: https://doi.org/10.1101/2022.12.13.520218
Hu C, Chen W, Myers SJ, Yuan H, Traynelis SF. Human GRIN2B variants in neurodevelopmental disorders. J Pharmacol Sci Jpn Pharmacol Soc. 2016;132:115–21. p
Platzer K, Lemke JR. GRIN2B-related neurodevelopmental disorder. GeneReviews. 2018 May [cited 2022 Aug 11]; Available from: http://www.ncbi.nlm.nih.gov/books/NBK501979/
Ohtsuki T, Sakurai K, Dou H, Toru M, Yamakawa-Kobayashi K, Arinami T. Mutation analysis of the NMDAR2B (GRIN2B) gene in schizophrenia. Mol Psychiatry. 2001;6:211–6.
Freunscht I, Popp B, Blank R, Endele S, Moog U, Petri H, et al. Behavioral phenotype in five individuals with de novo mutations within the GRIN2B gene. Behav Brain Funct. 2013;9:20.
Lemke JR, Hendrickx R, Geider K, Laube B, Schwake M, Harvey RJ, et al. GRIN2B mutations in west syndrome and intellectual disability with focal epilepsy. Ann Neurol. 2014;75:147–54.
Kellner S, Abbasi A, Carmi I, Heinrich R, Garin-Shkolnik T, Hershkovitz T, et al. Two de novo GluN2B mutations affect multiple NMDAR-functions and instigate severe pediatric encephalopathy. Elife. 2021;10:e67555.
Platzer K, Yuan H, Schütz H, Winschel A, Chen W, Hu C, et al. GRIN2B encephalopathy: novel findings on phenotype, variant clustering, functional consequences and treatment aspects. J Med Genet. 2017;54:460–70.
López-Rivera JA, Pérez-Palma E, Symonds J, Lindy AS, McKnight DA, Leu C, et al. Predicting incidences of neurodevelopmental disorders. Brain 2020;143:1099–105.
McStay B, Grummt I. The epigenetics of rRNA genes: from molecular to chromosome biology. Annu Rev Cell Dev. Biol. 2008;24:131–57.
Hamdane N, Herdman C, Mars JC, Stefanovsky V, Tremblay MG, Moss T. Depletion of the cisplatin targeted HMGB-box factor UBF selectively induces p53-independent apoptotic death in transformed cells. Oncotarget. 2015;6:27519–36.
Toro C, Hori RT, Malicdan MCV, Tifft CJ, Goldstein A, Gahl WA, et al. A recurrent de novo missense mutation in UBTF causes developmental neuroregression. Hum Mol Genet. 2018;27:691–705.
Hori RT, Moshahid Khan M, Xiao J, Hargrove PW, Moss T, LeDoux MS. Behavioral and molecular effects of Ubtf knockout and knockdown in mice. Brain Res. 2022;1793:148053.
Bastos F, Quinodoz M, Addor MC, Royer-Bertrand B, Fodstad H, Rivolta C, et al. Childhood neurodegeneration associated with a specific UBTF variant: A new case report and review of the literature. BMC Neurol. 2020;20:17.
Edvardson S, Nicolae CM, Agrawal PB, Mignot C, Payne K, Prasad AN, et al. Heterozygous de novo UBTF gain-of-function variant is associated with neurodegeneration in childhood. Am J Hum Genet. 2017;101:267–73.
Ikeda C, Kawarai T, Setoyama C, Orlacchio A, Imamura H. Recurrent de novo missense variant E210K in UBTF causes juvenile dystonia-parkinsonism. Neurol Sci. 2021;42:1217–9.
Sedláčková L, Laššuthová P, Štěrbová K, Haberlová J, Vyhnálková E, Neupauerová J, et al. UBTF mutation causes complex phenotype of neurodegeneration and severe epilepsy in childhood. Neuropediatrics 2019;50:57–60.
Nayak R, Rosh I, Rabinski T, Falik D, Mendel Percia M, Stern S. Generation and characterization of iPSC lines (UOHi003-A, UOHi002-A) from a patient with SHANK3 mutation and her healthy mother. Stem Cell Res. 2022;64:102899.
Thermo Fisher Scientific. CytoTuneTM-iPS 2.0 Sendai Reprogramming Kit USER GUIDE. Waltham, Massachusetts, United States. 2016 Nov [cited 2022 Jun 4]. Available from: https://tools.thermofisher.com/content/sfs/manuals/cytotune_ips_2_0_sendai_reprog_kit_man.pdf
Brennand KJ, Simone A, Jou J, Gelboin-Burkhart C, Tran N, Sangar S, et al. Modelling schizophrenia using human induced pluripotent stem cells. Nature 2011;473:221–5.
Stern S, Sarkar A, Galor D, Stern T, Mei A, Stern Y, et al. A Physiological instability displayed in hippocampal neurons derived from lithium-nonresponsive bipolar disorder patients. Biol Psychiatry. 2020;88:150–8.
Stern S, Santos R, Marchetto MC, Mendes APD, Rouleau GA, Biesmans S, et al. Neurons derived from patients with bipolar disorder divide into intrinsically different sub-populations of neurons, predicting the patients’ responsiveness to lithium. Mol Psychiatry. 2018;23:1453–65.
Ivashko-Pachima Y, Ganaiem M, Ben-Horin-Hazak I, Lobyntseva A, Bellaiche N, Fischer I, et al. SH3- and actin-binding domains connect ADNP and SHANK3, revealing a fundamental shared mechanism underlying autism. Mol Psychiatry. 2022;27:3316–27.
Griesi-Oliveira K, Suzuki AM, Alves AY, Mafra ACCN, Yamamoto GL, Ezquina S, et al. Actin cytoskeleton dynamics in stem cells from autistic individuals. Sci Rep. 2018;8:11138.
Karnik SK, Brooke BS, Bayes-Genis A, Sorensen L, Wythe JD, Schwartz RS, et al. A critical role for elastin signaling in vascular morphogenesis and disease. Development 2003;130:411–23. p
Li C, Su M, Wang H, Hu Y. Global view of transcriptome in the brains of aged NR2B transgenic mice. Neural Regen Res. 2013;8:2734–43.
Ivashko-Pachima Y, Hadar A, Grigg I, Korenková V, Kapitansky O, Karmon G, et al. Discovery of autism/intellectual disability somatic mutations in Alzheimer’s brains: mutated ADNP cytoskeletal impairments and repair as a case study. Mol Psychiatry. 2021;26:1619–33.
Hadar A, Kapitansky O, Ganaiem M, Sragovich S, Lobyntseva A, Giladi E, et al. Introducing ADNP and SIRT1 as new partners regulating microtubules and histone methylation. Mol Psychiatry. 2021;26:6550–61.
Conde C, Cáceres A. Microtubule assembly, organization and dynamics in axons and dendrites. Nat Rev Neurosci. 2009;10:319–32. p
Bell S, Maussion G, Jefri M, Peng H, Theroux JF, Silveira H, et al. Disruption of GRIN2B Impairs Differentiation in Human Neurons. Stem Cell Rep. 2018;11:183–96.
Kutsuwada T, Sakimura K, Manabe T, Takayama C, Katakura N, Kushiya E, et al. Impairment of suckling response, trigeminal neuronal pattern formation, and hippocampal LTD in NMDA receptor e2 subunit mutant mice. Neuron 1996;16:333–44.
Carrasquillo Y, Gereau RW. Pain sensitization. Learning and memory: a comprehensive reference. 2008;65–90.
Stafstrom CE. Persistent sodium current and its role in epilepsy. Epilepsy Curr. 2007;7:15–22.
Hosie S, Malone DT, Liu S, Glass M, Adlard PA, Hannan AJ, et al. Altered amygdala excitation and CB1 receptor modulation of aggressive behavior in the neuroligin-3R451C mouse model of autism. Front Cell Neurosci. 2018;12:234.
Lee E, Lee J, Kim E. Excitation/inhibition imbalance in animal models of autism spectrum disorders. Biol Psychiatry. 2017;81:838–47.
Han S, Tai C, Westenbroek RE, Yu FH, Cheah CS, Potter GB, et al. Autistic-like behaviour in Scn1a+- mice and rescue by enhanced GABA-mediated neurotransmission. Nature 2012;489:385–90.
Han S, Tai C, Jones CJ, Scheuer T, Catterall WA. Enhancement of inhibitory neurotransmission by GABAA receptors having α2,3-subunits ameliorates behavioral deficits in a mouse model of autism. Neuron 2014;81:1282–9.
Gkogkas CG, Khoutorsky A, Ran I, Rampakakis E, Nevarko T, Weatherill DB, et al. Autism-related deficits via dysregulated eIF4E-dependent translational control. Nature 2013;493:371–7.
Brant B, Stern T, Shekhidem HA, Mizrahi L, Rosh I, Stern Y, et al. IQSEC2 mutation associated with epilepsy, intellectual disability, and autism results in hyperexcitability of patient-derived neurons and deficient synaptic transmission. Mol Psychiatry. 2021;26:7498–508.
Rojas DC. The role of glutamate and its receptors in autism and the use of glutamate receptor antagonists in treatment. J Neural Transm. 2014;121:891–905.
Coghlan S, Horder J, Inkster B, Mendez MA, Murphy DG, Nutt DJ. GABA system dysfunction in autism and related disorders: From synapse to symptoms. Neurosci Biobehav Rev. 2012;36:2044–55. p
Rubenstein JLR, Merzenich MM. Model of autism: increased ratio of excitation/ inhibition in key neural systems. Genes Brain Behav. 2005;2:255–67.
Takarae Y, Sweeney J. Neural hyperexcitability in autism spectrum disorders. Brain Sci. 2017;7:129.
Gildin L, Rauti R, Vardi O, Kuznitsov-Yanovsky L, Maoz BM, Segal M, et al. Impaired functional connectivity underlies fragile X syndrome. Int J Mol Sci. 2022;23:2048.
Fink JJ, Schreiner JD, Bloom JE, James J, Baker DS, Robinson TM, et al. Hyperexcitable phenotypes in induced pluripotent stem cell–derived neurons from patients with 15q11-q13 duplication syndrome, a genetic form of autism. Biol Psychiatry. 2021;90:756–65.
Bourgeron T. From the genetic architecture to synaptic plasticity in autism spectrum disorder. Nat Rev Neurosci. 2015;16:551–63.
Sacai H, Sakoori K, Konno K, Nagahama K, Suzuki H, Watanabe T, et al. Autism spectrum disorder-like behavior caused by reduced excitatory synaptic transmission in pyramidal neurons of mouse prefrontal cortex. Nat Commun. 2020;11:5140.
Zoghbi HY, Bear MF. Synaptic dysfunction in neurodevelopmental disorders associated with autism and intellectual disabilities. Cold Spring Harb Perspect Biol. 2012;4:a009886.
Grigg I, Ivashko-Pachima Y, Hait TA, Korenková V, Touloumi O, Lagoudaki R, et al. Tauopathy in the young autistic brain: novel biomarker and therapeutic target. Transl Psychiatry. 2020;10:228.
Mencer S, Kartawy M, Lendenfeld F, Soluh H, Tripathi MK, Khaliulin I, et al. Proteomics of autism and Alzheimer’s mouse models reveal common alterations in mTOR signaling pathway. Transl Psychiatry. 2021;11:480.
Mason D, Ronald A, Ambler A, Caspi A, Houts R, Poulton R, et al. Autistic traits are associated with faster pace of aging: evidence from the Dunedin study at age 45. Autism Res. 2021;14:1684–94.
Schafer ST, Paquola ACM, Stern S, Gosselin D, Ku M, Pena M, et al. Pathological priming causes developmental gene network heterochronicity in autistic subject-derived neurons. Nat Neurosci. 2019;22:243–55.
Quraishi IH, Stern S, Mangan KP, Zhang Y, Ali SR, Mercier MR, et al. An epilepsy-associated KCNT1 mutation enhances excitability of human iPSC-derived neurons by increasing slack KNa currents. J Neurosci. 2019;39:7438–49.
Avazzadeh S, Quinlan LR, Reilly J, McDonagh K, Jalali A, Wang Y, et al. NRXN1α+/- is associated with increased excitability in ASD iPSC-derived neurons. BMC Neurosci. 2021;22:56.
Avazzadeh S, McDonagh K, Reilly J, Wang Y, Boomkamp SD, McInerney V, et al. Increased Ca2+ signaling in NRXN1α +/- neurons derived from ASD induced pluripotent stem cells. Mol Autism. 2019;10:52.
Que Z, Olivero-Acosta MI, Zhang J, Eaton M, Tukker AM, Chen X, et al. Hyperexcitability and pharmacological responsiveness of cortical neurons derived from human iPSCs carrying epilepsy-associated sodium channel Nav1.2-L1342P genetic variant. J Neurosci. 2021;41:10194–208.
Bell S, Rousseau J, Peng H, Aouabed Z, Priam P, Theroux JF, et al. Mutations in ACTL6B cause neurodevelopmental deficits and epilepsy and lead to loss of dendrites in human neurons. Am J Hum Genet. 2019;104:815–34.
Stern S, Lau S, Manole A, Rosh I, Percia MM, Ben Ezer R, et al. Reduced synaptic activity and dysregulated extracellular matrix pathways in midbrain neurons from Parkinson’s disease patients. NPJ Parkinsons Dis. 2022;8:103.
Tripathi U, Rosh I, Ezer B, Nayak R, Choudhary A, Djamus J, et al. Upregulated extracellular matrix-related genes and impaired synaptic activity in dopaminergic and hippocampal neurons derived from Parkinson’s disease patients with PINK1 and PARK2 mutations. bioRxiv:519781v2 [Preprint]. 2023. Available from: https://doi.org/10.1101/2022.12.09.519781
Rike WA, Stern S. Proteins and transcriptional dysregulation of the brain extracellular matrix in Parkinson’s disease: a systematic review. Int J Mol Sci. 2023;24:7435.
Rosh I, Tripathi U, Hussein Y, Rike WA, Manole A, Cordeiro D, et al. Synaptic dysfunction and dysregulation of extracellular matrix-related genes in dopaminergic neurons derived from Parkinson’s disease sporadic patients and with GBA1 mutations. bioRxiv:536143v1 [Preprint]. 2023. Available from: https://doi.org/10.1101/2023.04.10.536143
Stern S, Zhang L, Wang M, Wright R, Cordeiro D, Peles D, et al. Monozygotic twins discordant for schizophrenia differ in maturation and synaptic transmission. 2023; Available from: https://doi.org/10.1101/2022.05.13.491776
Choudhary A, Peles D, Nayak R, Mizrahi L, Stern S. Current progress in understanding schizophrenia using genomics and pluripotent stem cells: a meta-analytical overview. Schizophr Res. 2022. https://doi.org/10.1016/j.schres.2022.11.001.
Sarkar A, Mei A, Paquola ACM, Stern S, Bardy C, Klug JR, et al. Efficient generation of CA3 neurons from human pluripotent stem cells enables modeling of hippocampal connectivity in vitro. Cell Stem Cell. 2018;22:684–697.
Acknowledgements
This work was supported by Zuckerman STEM Leadership Program, Israel science foundation (ISF) grants 1994/21 and 352/21, new childhood – The organization for UBTF, Almy Foundation, and the GRIN2B Disorder Research Foundation (GDRF). We would like to thank the Technion Genomics Center for their contribution to STR testing services.
Author information
Authors and Affiliations
Contributions
SS designed and directed the study, wrote the MATLAB scripts for the analysis, and edited the manuscript, YH, UT, AC, RN, DP, JD, and IR performed the experiments; YH analyzed the data and wrote the manuscript; DP, RS, and TGS provided the patient material, TR and GDV provided some of the iPSC lines.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hussein, Y., Tripathi, U., Choudhary, A. et al. Early maturation and hyperexcitability is a shared phenotype of cortical neurons derived from different ASD-associated mutations. Transl Psychiatry 13, 246 (2023). https://doi.org/10.1038/s41398-023-02535-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-023-02535-x
- Springer Nature Limited
This article is cited by
-
Pharmacological modulation of developmental and synaptic phenotypes in human SHANK3 deficient stem cell-derived neuronal models
Translational Psychiatry (2024)
-
mGluR7 allosteric modulator AMN082 corrects protein synthesis and pathological phenotypes in FXS
EMBO Molecular Medicine (2024)
-
Uncovering convergence and divergence between autism and schizophrenia using genomic tools and patients’ neurons
Molecular Psychiatry (2024)
-
Upregulated ECM genes and increased synaptic activity in Parkinson’s human DA neurons with PINK1/ PRKN mutations
npj Parkinson's Disease (2024)
-
Monozygotic twins discordant for schizophrenia differ in maturation and synaptic transmission
Molecular Psychiatry (2024)