Abstract
Prenatal stress and poor maternal mental health are associated with adverse offspring outcomes; however, the biological mechanisms are unknown. Epigenetic modification has linked maternal health with offspring development. Epigenome-wide association studies (EWAS) have examined offspring DNA methylation profiles for association with prenatal maternal mental health to elucidate mechanisms of these complex relationships. The objective of this study is to provide a comprehensive, systematic review of EWASs of infant epigenetic profiles and prenatal maternal anxiety, depression, or depression treatment. We conducted a systematic literature search following PRISMA guidelines for EWAS studies between prenatal maternal mental health and infant epigenetics through May 22, 2023. Of 645 identified articles, 20 fulfilled inclusion criteria. We assessed replication of CpG sites among studies, conducted gene enrichment analysis, and evaluated the articles for quality and risk of bias. We found one repeated CpG site among the maternal depression studies; however, nine pairs of overlapping differentially methylatd regions were reported in at least two maternal depression studies. Gene enrichment analysis found significant pathways for maternal depression but not for any other maternal mental health category. We found evidence that these EWAS present a medium to high risk of bias. Exposure to prenatal maternal depression and anxiety or treatment for such was not consistently associated with epigenetic changes in infants in this systematic review and meta-analysis. Small sample size, potential bias due to exposure misclassification and statistical challenges are critical to address in future efforts to explore epigenetic modification as a potential mechanism by which prenatal exposure to maternal mental health disorders leads to adverse infant outcomes.
Similar content being viewed by others
Introduction
Maternal depression and anxiety, both during and after pregnancy, are common and a major public health problem in the United States, affecting over 1 in 10 mothers [1]. Epidemiologic studies have suggested an association between prenatal maternal depression and anxiety with adverse child outcomes such as low birthweight and preterm delivery [2, 3], as well as developmental delays and emotional and behavioral problems [4]. Low birthweight, preterm delivery, and developmental delays have been associated with changes in methylation profiles [5,6,7]. Epigenetic modifications have been proposed as possible mechanisms that may help explain the association between prenatal stress and adverse child developmental outcomes [8]. The motivation for this approach is that DNA methylation may alter gene expression in ways that influence early-infancy and later-life developmental outcomes.
Epigenetic markers, including DNA methylation, may be responsive to environmental factors throughout life, especially during in utero development [9]. With the exception of imprinted regions, the genome is demethylated prior to implantation with totipotency restored and the appropriate sex and tissue type specificity patterns reestablished throughout development [10,11,12,13,14,15,16]. Prenatal smoking [17], body mass index [18], and exposure to certain chemicals [19, 20] have been associated with changes to the infant’s epigenetic profile and the long-term health outcomes of offspring. It has thus been hypothesized that in utero exposure of the fetus to maternal depression may influence infant epigenetic profiles that alter fetal and child health and development. However, exposure to maternal depression is highly related and confounded with many potential variables such as smoking [21], maternal age [22], and maternal socioeconomic status [23]. Alterations in DNA methylation patterns in response to maternal depression may be adaptive changes that help an infant to anticipate a stressful or scarce environment; alternatively, they could be induced by pathological changes associated with medications or increased oxidative stress, or perhaps simply reflect different underlying sequence variation.
Epigenome-wide associations studies (EWAS) investigate associations between a phenotype (e.g. maternal mental health) and epigenetic variants in various tissues across the genome spanning 27,000 to a million or more CpG methylation sites [24]. While most studies of prenatal stress and infant epigenetic outcomes have focused on candidate gene methylation sites, epigenome-wide studies take an agnostic approach to identifying novel associations. Determining the influence of prenatal exposures on offspring epigenomic patterns requires complex study design and careful consideration of potential confounding and effect modification because the genetic, epigenetic, and environmental exposures over time of two linked but independent individuals must be considered. Although several reviews have attempted to synthesize related research [25,26,27], to our knowledge, ours is the first comprehensive, systematic review of EWAS of prenatal mental health and infant epigenetic profiles.
We performed a systematic review and critical assessment of EWAS studies on maternal mental health during pregnancy and the epigenetic profile of the offspring to assess whether depression, depression treatment, or anxiety in pregnancy women may influence offspring epigenetic profiles, compared with the epigenetic profiles in offspring born to mothers without depression or anxiety during pregnancy. We examined each study’s design, statistical analyses, and reporting, as well as its methodological quality and risk of bias. We assessed replication of CpG findings among studies and conducted gene enrichment analysis. Our findings underscore the importance of replication of research and study design in this area; more studies are needed to clarify the associations between maternal mental state and offspring epigenetic changes, which may influence offspring early-infancy and later-life health and developmental outcomes.
Methods
Literature search
We conducted our systematic review in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines [28] and registered it prospectively in PROSPERO (registration ID number: CRD42022335595). We conducted a review of EWAS to assess the association between mothers with vs mothers without either maternal depression, maternal anxiety, or depression treatment during pregnancy and their offspring’s epigenetic profile to explore the quality of these studies and compare their findings. The exposures were maternal depression, maternal anxiety, or depression treatment during pregnancy. The comparison groups were mothers who did not experience maternal depression, maternal anxiety, or depression treatment during pregnancy. The outcomes were epigenetic profiles of the offspring. Epigenome wide association studies were eligible for inclusion if they 1) specified the epigenetic profile of the offspring as the outcome, 2) measured exposures occurring during pregnancy, and 3) were published in the English language. Studies were excluded if they 1) analyzed only candidate genes, or 2) were published only as conference papers or abstracts. The search was conducted on May 22, 2023, on PubMed, Scopus, and Embase. The following search terms were applied: “epigenome wide association study”, “DNA methylation”, “pregnancy”, “prenatal”, “depression”, “anxiety”, and “psychiatric” (detailed search strategy is available in the Supplement).
Data extraction and quality assessment
The results of the search were exported to Excel and two reviewers (ED, KSC) conducted title abstract screening full text of articles that passed screening. Risk of bias was assessed by two reviewers (ED and DR) through the Risk of Bias in Non-randomized Studies of Interventions (ROBINS-I) tool [29] and the quality of reporting was assessed using the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines [30]. From each study, we extracted data pertaining to sample size, recruitment time period, time of enrollment, country, recruitment location, ethnicity/race, age, study design, analytic method, methylation chip, tissue sample type, cell type correction, covariates, covariate data collection methods, ancestral markers, genetic interactions, genome-wide associations, CpG sites excluded from analysis, P-values for significance, replication/validation analyses, exposures, exposure measurements, timing of exposure measurement, main CpG findings, main DMR findings, and relevant genes. The significant or top CpG sites and DMRs were compared within each exposure category to find any replicated CpG sites and DMRs or overlapping DMRs. Gene enrichment of the gene ontology (GO) categories of the significant and top ranked CpG sites and DMRs from the studies was conducted using the gometh function of the missMethyl R package [31] (version 1.31.0). Gometh tests GO enrichments for inputted CpG sites by empirically calculating the probability of differential methylation as a function of the number of CpGs, which accounts for biases in the number of probes per gene on the array and for CpGs that are annotated to multiple genes [31].
We conducted separate analyses for three specific associations: prenatal maternal depression and offspring DNA methylation profile (directed acyclic graph Fig. 1A), prenatal maternal depression treatment and offspring DNA methylation profile (directed acyclic graph Fig. 1B), and prenatal maternal anxiety and offspring DNA methylation profile (directed acyclic graph Fig. 1C). Due to the direct relationship between depression and depression treatment shown in Fig. 1B and the effect medications may have on DNA methylation, depression treatment was examined as its own category. Anxiety treatment was not examined due to the lack of articles on anxiety treatment in the search results.
A Directed acyclic graph for prenatal maternal depression and offspring DNA methylation profile. Many factors have been associated with prenatal maternal depression, offspring DNA methylation, or both resulting in a complex network between prenatal maternal depression and offspring DNA methylation. The arrows represent associations between two factors in which one factor influences the other. The hyphened arrows refer to a theoretical association. The purple arrows refer to associations where prenatal maternal depression influences another factor, and the black arrows refer to associations between the covariates and other factors. These covariates came from common covariates used in the studies for prenatal maternal depression and offspring DNA methylation in this review. Prenatal maternal depression can be connected to or influence offspring DNA methylation through multiple pathways. A main pathway through which prenatal maternal depression influences offspring methylation in this DAG is through preterm birth/gestational age [79, 80]. Multiple factors including maternal place of birth [21, 81], maternal age [22, 82], maternal smoking [21, 83, 84], maternal BMI [18, 85], maternal education [21, 86, 87], maternal SES [23, 88], parity [87, 89], and race/ethnicity [88, 90] influence both prenatal maternal depression and offspring DNA methylation. Though not “directly” connected in this DAG, offspring DNA methylation is influenced by maternal education through maternal smoking [87], by maternal socioeconomic status through preterm birth/gestational age [88], and by parity through maternal smoking [87] and maternal BMI [89]. Infant sex [91] and child age [91] also influence the offspring’s DNA methylation profile. B Directed acyclic graph for prenatal maternal depression treatment and offspring DNA methylation profile. Many factors have been associated with prenatal maternal depression treatment, offspring DNA methylation, or both resulting in a complex network between prenatal maternal depression and offspring DNA methylation. The arrows represent associations between two factors in which one factor influences the other. The hyphened arrows refer to a theoretical association. The arrows represent associations between two factors in which one factor influences the other. The green arrows refer to associations where prenatal maternal depression treatment influences another factor, the purple arrows refer to association where prenatal maternal depression influences another factor, and the black arrows refer to associations between the covariates and another factors. These covariates came from common covariates used in the studies for prenatal maternal depression treatment and offspring DNA methylation in this review. Like prenatal maternal depression, prenatal maternal depression treatment influences offspring DNA methylation through preterm birth/gestational age [92]. As with the previous DAG in A, maternal age [22, 82], maternal smoking [21, 83], maternal BMI [18, 85], and parity [87, 89, 93] maternal education [21, 81] influence both prenatal maternal depression and offspring DNA methylation. As a result, these factors also influence prenatal maternal depression treatment through prenatal maternal depression. Infant sex [91] and child age [91] also influence the offspring’s DNA methylation profile. C Directed acyclic graph for prenatal maternal anxiety and offspring DNA methylation profile. Many factors have been associated with prenatal maternal anxiety, offspring DNA methylation, or both. The arrows represent associations between two factors in which one factor influences the other. The blue arrows refer to associations where prenatal maternal anxiety influences another factors and the black arrows refer to associations between the covariates and another factors. The hyphened arrow refers to a theoretical association. These covariates came from common covariates used in the studies for prenatal maternal anxiety and offspring DNA methylation in this review. Prenatal maternal anxiety is connected to or influences offspring DNA methylation through multiple pathways. An important route in which prenatal maternal anxiety influences offspring DNA methylation is through preterm birth/gestational age [94]. Similarly, to A and B, except with prenatal maternal anxiety instead of prenatal maternal depression, maternal education [86, 95], maternal socioeconomic status [88, 95], maternal smoking [83, 96], maternal age [82, 87], maternal BMI [18, 85], and parity [87, 89] influence both prenatal maternal anxiety and offspring DNA methylation. Infant sex [91] also influences the offspring’s DNA methylation profile.
Results
Included studies
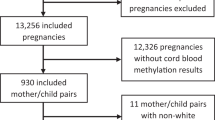
We identified 1321 articles from our PubMed, Scopus, Embase search. Of these, 676 were excluded as duplicates leaving 645 for abstract screening. After removing 598 during abstract screening, we performed full text screening on the remaining 47 articles and removed an additional 27 for one of the following reasons: non-human study population, candidate genes only, postnatal exposure, not being a full article, or exposure other than maternal depression, maternal depression treatment, or maternal anxiety, The remaining 20 articles were included our analysis (Fig. 2). Several articles described analyses of more than one exposure, resulting in 16 analyses [32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47] of maternal depression, 8 analyses [32, 35, 38, 39, 44, 45, 47, 48] of maternal depression treatment, and 9 analyses [27, 32, 33, 35, 38, 41, 47, 49, 50] of maternal anxiety (Table S1). Based on STROBE and ROBINS-I criteria, we found that 13 analyses had a moderate risk of bias and 7 had a severe risk of bias (Tables S2 and S3).
Depression during pregnancy
Of the 16 studies that analyzed associations between maternal depression during pregnancy and the infant’s epigenetic profile, we found the risk of bias to be moderate in 12 studies and severe in 4 studies. Main factors increasing the risk of bias were not accounting for all important confounders and not accounting for selection bias. Covariates and confounders adjusted for in more than one of the included studies are shown in Fig. 1A. No statistically significant CpG site was reported from more than one study. However, one CpG site, cg25157095, was reported as a top ranked CpG site in two studies. Nine pairs of overlapping DMRs were found in at least two maternal depression studies (Table 1). The gene enrichment analysis had 62 significant pathways for study significant DMRs and 65 signficant pathways for top ranked DMRs (Table 2). The gene enrichment analysis did not produce any significant results for the CpG sites. The top ten pathways for the CpG sites are presented in Table S4.
Depression treatment during pregnancy
Eight studies analyzed associations of maternal depression treatment during pregnancy with the infant’s epigenetic profile. We found the risk of bias to be moderate in 7 of these studies and severe in 1 study. Main factors increasing the risk of bias in these studies were not accounting for all important confounders, not accounting for selection bias, and missing data. Covariates and confounders adjusted for in more than one of the included studies are shown in Fig. 1B. No statistically significant or top-ranked CpG sites were reported from more than one study. Many studies either did not perform a DMR analysis or did not provide ranked results. Only one DMR, chr12: 56325797–56325867, was reported as a top ranked DMR. The gene enrichment analysis did not produce any significant results for the CpG sites. The top ten pathways are presented in Table S5.
Anxiety during pregnancy
Of the 9 studies that analyzed the association between maternal anxiety during pregnancy and the infant’s epigenetic profile, 7 had a moderate risk of bias while 2 had a severe risk of bias. Main factors increasing the risk of bias in these studies were not accounting for all important confounders, not accounting for selection bias, and missing data. Covariates and confounders adjusted for in more than one of the included studies are shown in Fig. 1C. No statistically significant or top-ranked CpG sites were reported from more than one study. The studies shared no overlapping DMRs. The gene enrichment analysis did not produce any significant results for the CpG sites or the DMRs. The top ten pathways are presented in Table S6.
Discussion
To our knowledge, the present work is the first systematic review of epigenome-wide association studies of prenatal maternal mental health and infant epigenetic profiles. We identified 20 EWAS studies which together reported 803 CpG sites and 19,440 DMRs in infants associated with maternal anxiety, depression, or depression treatment. Among the studies within any maternal mental health category, there was only one top ranked CpG site, cg25157095, reported in more than one study. Only nine overlapping DMRs were reported in at least two studies on maternal depression. We identified significant pathways only for maternal depression DMRs in the gene enrichment analysis. The main limitations of the studies we identified were small sample sizes, concerns about exposure misclassification and suboptimal statistical analyses that increased the risk of bias.
We found that no replicated single significant CpG sites were reported from more than one study in any of the three categories (maternal depression, maternal depression treatment, and maternal anxiety). However, one CpG site, cg25157095 was found among the top non-significant CpG sites for maternal depression in two studies [40, 47]. This CpG site is in an intron for the RIPK4 gene which has been implicated in keratinocyte differentiation, modulation of the actin cytoskeleton, and restricting intercellular adhesion [51]. Three pairs of overlapping DMRs were reported in two of 15 studies of maternal depression during pregnancy. One pair of overlapping DMRs, chr8:70378380-70378994 and chr8:70378380-70378995, is in the SULF1 gene. The SULF1 gene is involved in the regulation of multiple cellular pathways for editing heparan sulfate chains [52]. This gene has been associated with nervous system development in studies of SULF1 deficient mice, providing evidence that a non-functioning SULF1 gene is associated with impaired neurite growth and impaired long-term potentiation [53, 54], a form of synaptic plasticity [55]. Single nucleotide polymorphisms (SNPs) in this gene have been associated with cancer risk [56]. Another pair of overlapping DMRs, chr8: 143859669-143859991 and chr8: 143859369-143860092, is closest to the LYNX1 gene. Evidence has been provided through mouse models for the possible role of this gene in synaptic plasticity [57].
The chr11: 85195094-85195206 and chr11: 85195119-85195288 pair of overlapping DMRs are in the DLG2 gene. This gene encodes for the postsynaptic density 93 protein which has been thought to have roles in synaptic stability and regulation [58, 59]. Genetic variations in DLG2 have been associated with schizophrenia [60], attention deficit hyperactivity disorder [61], and bipolar disorder [62]. Another pair of overlapping DMRs were chr7:27183643-27184853 and chr7:27183133-27184522, which were significantly associated in opposite directions in the two studies. This DMR is located in HOXA-5 in the HOXA gene cluster, which is important in human development [63]. Hypermethylation in this gene has also been associated with various cancers [64] and hypermethylation in this region of the gene has also been associated with Alzheimer’s disease [65]. Another pair of overlapping DMRs, chr1:62660188-62660861 and chr1:62660038-62661010, span the L1TD1 gene. The L1TD1 gene has been connected to pluripotency maintence [66]. Higher expression of L1TD1 has also been associated with longer disease free colon cancer survival [67] and L1TD1 has been found to have higher levels of methylation in non-small cell lung cancer tissue [68]. The chr6: 151125848-151125886 and chr6: 151125729-151125904 pair of DMRs are within the PLEKHG1 gene. A SNP within this gene has been associate with white matter hyperintensities and ischemic stroke [69].
Continued research in this area is needed to determine if changes in these DMR regions result in functional changes that may correspond to the adverse outcomes seen in the epidemiological literature [2,3,4]. The overall lack of replication across studies highlights the difficulty of interpreting these types of analyses.EWAS studies are limited by the types of CpG sites included and may systematically overlook critical regions [70, 71]. These types of studies also limited by available tissue type, typically blood or saliva. It is well established that DNA methylation patterns are tissue specific and the lack of data on critical regions such as the brain is problematic [11, 70, 72]. Poor correlation of DNA methylation results measured using different Illumina platforms (e.g., the EPIC, 450k, and 27k arrays) presents problems for replication [73]. Underlying sequence variation is also of concern in EWAS studies as methylation can be a direct result of the underlying genetic sequence. Correction for population stratification and genetic variation/interaction will be critical in future studies.
Methods for determining and classifying maternal depression and maternal anxiety during pregnancy varied among our included studies, as did measurements of medication intake and dosage. The studies we considered to have a severe risk of bias all lacked appropriate adjustment for confounding and consideration of selection bias.
Maternal depression is a critical public health problem that is both under-treated and under-diagnosed, especially in minority and underserved populations. Untreated prenatal depression has been associated with detrimental health outcomes for both the pregnant woman and the baby, including an elevated risk for postpartum depression in the mother and increased infant risk for preterm birth and low birth weight [74]. Antenatal maternal anxiety and stress can impact the psychological and intellectual development of the infant [75], with some studies suggesting increased risk for emotional and cognitive problems, attentional deficit, and language delay. Prenatal anxiety and depression are also associated with increased risk for suicidality in mothers [76], with the greatest increases seen among Non-Hispanic Black, low-income, and younger individuals. Maternal mental health is tied to racial and ethnic disparities, with a higher overall prevalence of maternal depression among non-Hispanic Blacks and Hispanics compared to non-Hispanic whites [77]. Emerging evidence indicates that the COVID-19 pandemic increased the prevalence of mental health issues during pregnancy, with a meta-analysis of 37 studies suggesting that more than one in four pregnant women experienced prenatal depression and one in three experienced clinically significant anxiety [78]. Early screening and treatment for women at risk for maternal depression and anxiety may help prevent long-term adverse outcomes on maternal and infant well-being.
The EWAS studies included in this systematic review explored associations of prenatal anxiety or depression with DNA methylation patterns in offspring. Among the included studies, there was a lack of replication for a majority of the studies’ findings. However, a limitation of this study includes the potential to miss relevant articles in the initial search. Further studies of larger sample sizes are needed to identify and replicate findings and further investigate the role of maternal mental health on infant epigenetic profiles as well as take greater steps to control for confounding and address selection bias.
References
Ko JY, Rockhill KM, Tong VT, Morrow B, Farr SL. Trends in Postpartum Depressive Symptoms - 27 States, 2004, 2008, and 2012. MMWR Morb Mortal Wkly Rep. 2017;66:153–8.
Ghimire U, Papabathini SS, Kawuki J, Obore N, Musa TH. Depression during pregnancy and the risk of low birth weight, preterm birth and intrauterine growth restriction- an updated meta-analysis. Early Hum Dev. 2021;152:105243.
Liou SR, Wang P, Cheng CY. Effects of prenatal maternal mental distress on birth outcomes. Women Birth. 2016;29:376–80.
Field T. Prenatal depression effects on early development: a review. Infant Behav Dev. 2011;34:1–14.
Hayashi I, Yamaguchi K, Sumitomo M, Takakura K, Nagai N, Sakane N. Full-term low birth weight infants have differentially hypermethylated DNA related to immune system and organ growth: a comparison with full-term normal birth weight infants. BMC Res Notes. 2020;13:199.
Sparrow S, Manning JR, Cartier J, Anblagan D, Bastin ME, Piyasena C, et al. Epigenomic profiling of preterm infants reveals DNA methylation differences at sites associated with neural function. Transl Psychiatry. 2016;6:e716.
Hüls A, Wedderburn CJ, Groenewold NA, Gladish N, Jones MJ, Koen N, et al. Newborn differential DNA methylation and subcortical brain volumes as early signs of severe neurodevelopmental delay in a South African Birth Cohort Study. The. World J Biol Psychiatry. 2022;23:601–12.
Wright ML, Starkweather AR, York TP. Mechanisms of the Maternal Exposome and Implications for Health Outcomes. ANS Adv Nurs Sci. 2016;39:E17–30.
Kundakovic M, Jaric I. The Epigenetic Link between Prenatal Adverse Environments and Neurodevelopmental Disorders. Genes. 2017;8:104.
Crider KS, Yang TP, Berry RJ, Bailey LB. Folate and DNA methylation: a review of molecular mechanisms and the evidence for folate’s role. Adv Nutr. 2012;3:21–38.
Kota SK, Feil R. Epigenetic transitions in germ cell development and meiosis. Dev Cell. 2011;19:675–86.
Weksberg R. Imprinted genes and human disease. Am J Med Genet C Semin Med Genet. 2010;154C:317–20.
Zhu P, Guo H, Ren Y, Hou Y, Dong J, Li R, et al. Single-cell DNA methylome sequencing of human preimplantation embryos. Nat Genet. 2018;50:12–9.
Clarke HJ. Nuclear and chromatin composition of mammalian gametes and early embryos. Biochem Cell Biol. 1992;70:856–66.
Ishida M, Moore GE. The role of imprinted genes in humans. Mol Asp Med. 2013;34:826–40.
Weaver JR, Susiarjo M, Bartolomei MS. Imprinting and epigenetic changes in the early embryo. Mamm Genome. 2009;20:532–43.
Wiklund P, Karhunen V, Richmond RC, Parmar P, Rodriguez A, De Silva M, et al. DNA methylation links prenatal smoking exposure to later life health outcomes in offspring. Clin Epigenetics. 2019;11:97.
Sharp GC, Salas LA, Monnereau C, Allard C, Yousefi P, Everson TM, et al. Maternal BMI at the start of pregnancy and offspring epigenome-wide DNA methylation: findings from the pregnancy and childhood epigenetics (PACE) consortium. Hum Mol Genet. 2017;26:4067–85.
Bozack AK, Rifas-Shiman SL, Coull BA, Baccarelli AA, Wright RO, Amarasiriwardena C, et al. Prenatal metal exposure, cord blood DNA methylation and persistence in childhood: an epigenome-wide association study of 12 metals. Clin Epigenetics. 2021;13:208.
McCabe CF, Padmanabhan V, Dolinoy DC, Domino SE, Jones TR, Bakulski KM, et al. Maternal environmental exposure to bisphenols and epigenome-wide DNA methylation in infant cord blood. Environ Epigenetics. 2020;6:dvaa021.
Field T. Prenatal Depression Risk Factors, Developmental Effects and Interventions: A Review. J Pregnancy Child Health. 2017;4:301.
Míguez MC, Vázquez MB. Risk factors for antenatal depression: A review. World J Psychiatry. 2021;11:325–36.
Hein A, Rauh C, Engel A, Häberle L, Dammer U, Voigt F, et al. Socioeconomic status and depression during and after pregnancy in the Franconian Maternal Health Evaluation Studies (FRAMES). Arch Gynecol Obstet. 2014;289:755–63.
Ryan J, Mansell T, Fransquet P, Saffery R. Does maternal mental well-being in pregnancy impact the early human epigenome? Epigenomics. 2017;9:313–32.
Edris A, den Dekker HT, Melén E, Lahousse L. Epigenome-wide association studies in asthma: A systematic review. Clin Exp Allergy. 2019;49:953–68.
Flanagan JM. Epigenome-wide association studies (EWAS): past, present, and future. Methods Mol Biol. 2015;1238:51–63.
Sammallahti S, Cortes Hidalgo AP, Tuominen S, Malmberg A, Mulder RH, Brunst KJ, et al. Maternal anxiety during pregnancy and newborn epigenome-wide DNA methylation. Mol Psychiatry. 2021;26:1832–45.
Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. BMJ. 2021;372:n71.
Schünemann HJ, Cuello C, Akl EA, Mustafa RA, Meerpohl JJ, Thayer K, et al. GRADE guidelines: 18. How ROBINS-I and other tools to assess risk of bias in nonrandomized studies should be used to rate the certainty of a body of evidence. J Clin Epidemiol. 2019;111:105–14.
Cuschieri S. The STROBE guidelines. Saudi J Anaesth. 2019;13:31.
Maksimovic J, Oshlack A, Phipson B. Gene set enrichment analysis for genome-wide DNA methylation data. Genome Biol. 2021;22:173.
Cardenas A, Faleschini S, Cortes Hidalgo A, Rifas-Shiman SL, Baccarelli AA, DeMeo DL, et al. Prenatal maternal antidepressants, anxiety, and depression and offspring DNA methylation: epigenome-wide associations at birth and persistence into early childhood. Clin Epigenetics. 2019;11:56.
Dean DC 3rd, Madrid A, Planalp EM, Moody JF, Papale LA, Knobel KM, et al. Cord blood DNA methylation modifications in infants are associated with white matter microstructure in the context of prenatal maternal depression and anxiety. Sci Rep. 2021;11:12181.
Drzymalla E, Gladish N, Koen N, Epstein MP, Kobor MS, Zar HJ, et al. Association between maternal depression during pregnancy and newborn DNA methylation. Transl Psychiatry. 2021;11:572.
Kallak TK, Bränn E, Fransson E, Johansson Å, Lager S, Comasco E, et al. DNA methylation in cord blood in association with prenatal depressive symptoms. Clin Epigenetics. 2021;13:78.
Kallak TK, Fransson E, Bränn E, Berglund H, Lager S, Comasco E, et al. Maternal prenatal depressive symptoms and toddler behavior: an umbilical cord blood epigenome-wide association study. Transl Psychiatry. 2022;12:186.
Nemoda Z, Massart R, Suderman M, Hallett M, Li T, Coote M, et al. Maternal depression is associated with DNA methylation changes in cord blood T lymphocytes and adult hippocampi. Transl Psychiatry. 2015;5:e545.
Non AL, Binder AM, Kubzansky LD, Michels KB, Genome-wide DNA. methylation in neonates exposed to maternal depression, anxiety, or SSRI medication during pregnancy. Epigenetics. 2014;9:964–72.
Schroeder JW, Smith AK, Brennan PA, Conneely KN, Kilaru V, Knight BT, et al. DNA methylation in neonates born to women receiving psychiatric care. Epigenetics. 2012;7:409–14.
Stonawski V, Roetner J, Goecke TW, Fasching PA, Beckmann MW, Kornhuber J, et al. Genome-Wide DNA Methylation Patterns in Children Exposed to Nonpharmacologically Treated Prenatal Depressive Symptoms: Results From 2 Independent Cohorts. Epigenet Insights. 2020;13:2516865720932146.
Tesfaye M, Chatterjee S, Zeng X, Joseph P, Tekola-Ayele F. Impact of depression and stress on placental DNA methylation in ethnically diverse pregnant women. Epigenomics. 2021;13:1485–96.
Viuff AC, Sharp GC, Rai D, Henriksen TB, Pedersen LH, Kyng KJ, et al. Maternal depression during pregnancy and cord blood DNA methylation: findings from the Avon Longitudinal Study of Parents and Children. Transl Psychiatry. 2018;8:244.
Wikenius E, Myhre AM, Page CM, Moe V, Smith L, Heiervang ER, et al. Prenatal maternal depressive symptoms and infant DNA methylation: a longitudinal epigenome-wide study. Nord J Psychiatry. 2019;73:257–63.
Inkster AM, Konwar C, Peñaherrera MS, Brain U, Khan A, Price EM, et al. Profiling placental DNA methylation associated with maternal SSRI treatment during pregnancy. Sci Rep. 2022;12:22576.
Olstad EW, Nordeng HME, Sandve GK, Lyle R, Gervin K. Effects of prenatal exposure to (es)citalopram and maternal depression during pregnancy on DNA methylation and child neurodevelopment. Transl Psychiatry. 2023;13:149.
Robakis TK, Roth MC, King LS, Humphreys KL, Ho M, Zhang X, et al. Maternal attachment insecurity, maltreatment history, and depressive symptoms are associated with broad DNA methylation signatures in infants. Mol Psychiatry. 2022;27:3306–15.
Bleker LS, Milgrom J, Sexton-Oates A, Roseboom TJ, Gemmill AW, Holt CJ, et al. Exploring the effect of antenatal depression treatment on children’s epigenetic profiles: findings from a pilot randomized controlled trial. Clin Epigenetics. 2019;11:18.
Gurnot C, Martin-Subero I, Mah SM, Weikum W, Goodman SJ, Brain U, et al. Prenatal antidepressant exposure associated with CYP2E1 DNA methylation change in neonates. Epigenetics 2015;10:361–72.
Kim HB, Kang MJ, Lee SY, Shin YJ, Hong SJ. Prenatal maternal anxiety promotes atopic dermatitis in offspring via placental DNA methylation changes. Asian Pac J Allergy Immunol. 2023;41:60–66.
Vangeel EB, Pishva E, Hompes T, van den Hove D, Lambrechts D, Allegaert K, et al. Newborn genome-wide DNA methylation in association with pregnancy anxiety reveals a potential role for GABBR1. Clin Epigenetics. 2017;9:107.
De Groote P, Tran HT, Fransen M, Tanghe G, Urwyler C, De Craene B, et al. A novel RIPK4-IRF6 connection is required to prevent epithelial fusions characteristic for popliteal pterygium syndromes. Cell Death Differ. 2015;22:1012–24.
Ai X, Kusche-Gullberg M, Lindahl U, Emerson CP. Chapter 8 - Remodeling of Heparan Sulfate Sulfation by Extracellular Endosulfatases. In: Garg HG, Linhardt RJ, Hales CA, editors. Chemistry and Biology of Heparin and Heparan Sulfate. Amsterdam: Elsevier Science; 2005. p. 245–58.
Joy MT, Vrbova G, Dhoot GK, Anderson PN. Sulf1 and Sulf2 expression in the nervous system and its role in limiting neurite outgrowth in vitro. Exp Neurol. 2015;263:150–60.
Kalus I, Salmen B, Viebahn C, von Figura K, Schmitz D, D’Hooge R, et al. Differential involvement of the extracellular 6-O-endosulfatases Sulf1 and Sulf2 in brain development and neuronal and behavioural plasticity. J Cell Mol Med. 2009;13:4505–21.
Leal G, Bramham CR, Duarte CB Chapter Eight - BDNF and Hippocampal Synaptic Plasticity. In: Litwack G, editor. Vitamins and Hormones. 104: Academic Press; 2017. p. 153–95.
Han CH, Huang YJ, Lu KH, Liu Z, Mills GB, Wei Q, et al. Polymorphisms in the SULF1 gene are associated with early age of onset and survival of ovarian cancer. J Exp Clin Cancer Res. 2011;30:5.
Shenkarev ZO, Shulepko MA, Bychkov ML, Kulbatskii DS, Shlepova OV, Vasilyeva NA, et al. Water-soluble variant of human Lynx1 positively modulates synaptic plasticity and ameliorates cognitive impairment associated with α7-nAChR dysfunction. J Neurochem. 2020;155:45–61.
Qin XY, Shan QH, Fang H, Wang Y, Chen P, Xiong ZQ, et al. PSD-93 up-regulates the synaptic activity of corticotropin-releasing hormone neurons in the paraventricular nucleus in depression. Acta Neuropathol. 2021;142:1045–64.
Parker MJ, Zhao S, Bredt DS, Sanes JR, Feng G. PSD93 regulates synaptic stability at neuronal cholinergic synapses. J Neurosci. 2004;24:378–88.
Marshall CR, Howrigan DP, Merico D, Thiruvahindrapuram B, Wu W, Greer DS, et al. Contribution of copy number variants to schizophrenia from a genome-wide study of 41,321 subjects. Nat Genet. 2017;49:27–35.
Alemany S, Ribasés M, Vilor-Tejedor N, Bustamante M, Sánchez-Mora C, Bosch R, et al. New suggestive genetic loci and biological pathways for attention function in adult attention-deficit/hyperactivity disorder. Am J Med Genet B Neuropsychiatr Genet. 2015;168:459–70.
Noor A, Lionel AC, Cohen-Woods S, Moghimi N, Rucker J, Fennell A, et al. Copy number variant study of bipolar disorder in Canadian and UK populations implicates synaptic genes. Am J Med Genet B Neuropsychiatr Genet. 2014;165b:303–13.
Mark M, Rijli FM, Chambon P. Homeobox genes in embryogenesis and pathogenesis. Pediatr Res. 1997;42:421–9.
Paço A, de Bessa Garcia SA, Freitas R. Methylation in HOX Clusters and Its Applications in Cancer Therapy. Cells. 2020;9:1613.
Smith RG, Hannon E, De Jager PL, Chibnik L, Lott SJ, Condliffe D, et al. Elevated DNA methylation across a 48-kb region spanning the HOXA gene cluster is associated with Alzheimer’s disease neuropathology. Alzheimers Dement. 2018;14:1580–8.
Närvä E, Rahkonen N, Emani MR, Lund R, Pursiheimo JP, Nästi J, et al. RNA-binding protein L1TD1 interacts with LIN28 via RNA and is required for human embryonic stem cell self-renewal and cancer cell proliferation. Stem Cells. 2012;30:452–60.
Chakroborty D, Emani MR, Klén R, Böckelman C, Hagström J, Haglund C, et al. L1TD1 - a prognostic marker for colon cancer. BMC Cancer. 2019;19:727.
Altenberger C, Heller G, Ziegler B, Tomasich E, Marhold M, Topakian T, et al. SPAG6 and L1TD1 are transcriptionally regulated by DNA methylation in non-small cell lung cancers. Mol Cancer. 2017;16:1.
Traylor M, Tozer DJ, Croall ID, Lisiecka-Ford DM, Olorunda AO, Boncoraglio G, et al. Genetic variation in PLEKHG1 is associated with white matter hyperintensities (n = 11,226). Neurology. 2019;92:e749–e57.
Gunasekara CJ, Scott CA, Laritsky E, Baker MS, MacKay H, Duryea JD, et al. A genomic atlas of systemic interindividual epigenetic variation in humans. Genome Biol. 2019;20:105.
Gunasekara CJ, Waterland RA. A new era for epigenetic epidemiology. Epigenomics. 2019;11:1647–9.
Waterland RA. Early environmental effects on epigenetic regulation in humans. Epigenetics. 2009;4:523–5.
Olstad EW, Nordeng HME, Sandve GK, Lyle R, Gervin K. Low reliability of DNA methylation across Illumina Infinium platforms in cord blood: implications for replication studies and meta-analyses of prenatal exposures. Clinical. Epigenetics. 2022;14:80.
Jarde A, Morais M, Kingston D, Giallo R, MacQueen GM, Giglia L, et al. Neonatal Outcomes in Women With Untreated Antenatal Depression Compared With Women Without Depression: A Systematic Review and Meta-analysis. JAMA Psychiatry. 2016;73:826–37.
Van den Bergh BR, Mulder EJ, Mennes M, Glover V. Antenatal maternal anxiety and stress and the neurobehavioural development of the fetus and child: links and possible mechanisms. A review. Neurosci Biobehav Rev. 2005;29:237–58.
Admon LK, Dalton VK, Kolenic GE, Ettner SL, Tilea A, Haffajee RL, et al. Trends in Suicidality 1 Year Before and After Birth Among Commercially Insured Childbearing Individuals in the United States, 2006-2017. JAMA Psychiatry. 2021;78:171–6.
Mukherjee S, Trepka MJ, Pierre-Victor D, Bahelah R, Avent T. Racial/Ethnic Disparities in Antenatal Depression in the United States: A Systematic Review. Matern Child Health J. 2016;20:1780–97.
Tomfohr-Madsen LM, Racine N, Giesbrecht GF, Lebel C, Madigan S. Depression and anxiety in pregnancy during COVID-19: A rapid review and meta-analysis. Psychiatry Res. 2021;300:113912.
Liu C, Cnattingius S, Bergström M, Östberg V, Hjern A. Prenatal parental depression and preterm birth: a national cohort study. Bjog. 2016;123:1973–82.
Merid SK, Novoloaca A, Sharp GC, Küpers LK, Kho AT, Roy R, et al. Epigenome-wide meta-analysis of blood DNA methylation in newborns and children identifies numerous loci related to gestational age. Genome Med. 2020;12:25.
Barchetta I, Arvastsson J, Sarmiento L, Cilio CM. Epigenetic Changes Induced by Maternal Factors during Fetal Life: Implication for Type 1 Diabetes. Genes. 2021;12:887.
Adkins RM, Thomas F, Tylavsky FA, Krushkal J. Parental ages and levels of DNA methylation in the newborn are correlated. BMC Med Genet. 2011;12:47.
Nakamura A, François O, Lepeule J. Epigenetic Alterations of Maternal Tobacco Smoking during Pregnancy: A Narrative Review. Int J Environ Res Public Health. 2021;18:5083.
Soneji S, Beltrán-Sánchez H. Association of Maternal Cigarette Smoking and Smoking Cessation With Preterm Birth. JAMA Netw Open. 2019;2:e192514.
Insan N, Slack E, Heslehurst N, Rankin J. Antenatal depression and anxiety and early pregnancy BMI among White British and South Asian women: retrospective analysis of data from the Born in Bradford cohort. BMC Pregnancy Childbirth. 2020;20:502.
Jackson M, Kiernan K, McLanahan S. Maternal Education, Changing Family Circumstances, and Children’s Skill Development in the United States and UK. Ann Am Acad Pol Soc Sci. 2017;674:59–84.
Sequí-Canet JM, Sequí-Sabater JM, Marco-Sabater A, Corpas-Burgos F, Collar Del Castillo JI, Orta-Sibú N. Maternal factors associated with smoking during gestation and consequences in newborns: Results of an 18-year study. J Clin Transl Res. 2022;8:6–19.
Goldenberg RL, Culhane JF, Iams JD, Romero R. Epidemiology and causes of preterm birth. Lancet. 2008;371:75–84.
Iversen DS, Kesmodel US, Ovesen PG. Associations between parity and maternal BMI in a population-based cohort study. Acta Obstet Gynecol Scand. 2018;97:694–700.
Xia YY, Ding YB, Liu XQ, Chen XM, Cheng SQ, Li LB, et al. Racial/ethnic disparities in human DNA methylation. Biochim Biophys Acta. 2014;1846:258–62.
Boks MP, Derks EM, Weisenberger DJ, Strengman E, Janson E, Sommer IE, et al. The relationship of DNA methylation with age, gender and genotype in twins and healthy controls. PLoS One. 2009;4:e6767.
Olivier JD, Åkerud H, Skalkidou A, Kaihola H, Sundström-Poromaa I. The effects of antenatal depression and antidepressant treatment on placental gene expression. Front Cell Neurosci. 2014;8:465.
Opaneye A. The influence of parity on the mode of delivery. Cent Afr J Med. 1988;34:26–8.
Rose MS, Pana G, Premji S. Prenatal Maternal Anxiety as a Risk Factor for Preterm Birth and the Effects of Heterogeneity on This Relationship: A Systematic Review and Meta-Analysis. Biomed Res Int. 2016;2016:8312158.
Wallace K, Araji S. An Overview of Maternal Anxiety During Pregnancy and the Post-Partum Period. J Ment Health Clin Psychol. 2020;4:47–56.
Silva MMJ, Nogueira DA, Clapis MJ, Leite E. Anxiety in pregnancy: prevalence and associated factors. Rev Esc Enferm Usp. 2017;51:e03253.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Drzymalla, E., Crider, K.S., Wang, A. et al. Epigenome-wide association studies of prenatal maternal mental health and infant epigenetic profiles: a systematic review. Transl Psychiatry 13, 377 (2023). https://doi.org/10.1038/s41398-023-02620-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-023-02620-1
- Springer Nature Limited