Abstract
Background
The extracellular matrix modulates the hallmarks of cancer. Here we examined the role of agrin—a member of this matrix—in progression of oral squamous cell carcinoma (OSCC).
Methods
We evaluated the immunohistochemical expression of agrin in OSCC and dysplasias. Benign lesions were used as control. In subsequent experiments, we investigated whether the silencing of agrin interferes with tumour expansion both in vitro as well as in vivo. To gain insights into the role of agrin, we identified its protein network (interactome) using mass spectrometry-based proteomics and bioinformatics. Finally, we evaluated the clinical relevance of agrin interactome.
Results
Agrin was elevated in malignant and premalignant lesions. Further, we show that agrin silencing interferes with cancer cell motility, proliferation, invasion, colony and tumour spheroid formation, and it also reduces the phosphorylation of FAK, ERK and cyclin D1 proteins in OSCC cells. In orthotopic model, agrin silencing reduces tumour aggressiveness, like vascular and neural invasion. From a clinical perspective, agrin contextual hubs predict a poor clinical prognosis related with overall survival.
Conclusions
Altogether, our results demonstrate that agrin is a histological marker for the progression of oral cancer and is a strong therapeutic target candidate for both premalignant and OSCC lesions.
Similar content being viewed by others
Introduction
Worldwide, head and neck squamous cell carcinoma (HNSCC) affects over 500,000 patients per year.1 Oral squamous cell carcinoma (OSCC) represents 95% of all forms of HNSCC.2 It is the most common malignancy of the head and neck.3 Despite advancements in prevention and multimodality therapies, the prognosis of OSCC patients has remained unfavourable in the past few decades.4 A better understanding of cellular and molecular mechanisms that promote the progression of OSCC can help improve our approach to this disease.
The process of cancer progression (i.e. local invasion and metastasis) is characterised by rapid cellular growth accompanied by alterations of the microenvironment of the cancer cells.5 OSCC can be presented as a natural history, which originates from nontumourigenic keratinocytes which are chronically exposed to carcinogens, following a hyperplasia, oral epithelial dysplasia (OED; in different degrees) and an invasive carcinoma leading to the generation of metastases.6
The extracellular matrix (ECM) modulates the hallmarks of cancer, and changes in its dynamics contribute to tumour progression.7 Some components of the ECM, which include heparan sulphate proteoglycans, are frequently overproduced in cancer.8 Agrin is one of the main heparan sulphate proteoglycans present in the ECM.
Agrin is a multi-domain protein expressed as either a membrane protein or secreted in the ECM.9 Agrin has been shown to act as a sensor in developing oncogenic signals associated with the ECM in hepatic carcinomas.9 In the context of OSCC progression, a previous study of our group showed that agrin has high expression in OSCC and has a role on cell migration, adhesion and resistance to chemotherapy,10 suggesting that agrin also has an oncogenic role in oral cancer.
Agrin can be proteolytically cleaved which generates bioactive fragments that modulate cellular behaviour.11 One of the agrin cleavage products, the C-terminal fragment (hereafter called Ct-agrin), has been shown to be a promising new biomarker for pathological processes, including sarcopenia,12 renal dysfunction13 and colorectal cancer.14 The disintegration of the basement membrane upon local invasion processes can release agrin-processing products, such as Ct-agrin.14 Within OSCC are active proteases, such as matrix metalloproteinase (MMP)-3 and neurotrypsin,15, 16 which are capable of generating this soluble fragment. Focussing on the dynamics of tumour progression, the Ct-agrin could then help explain the role of agrin in oral cancer.
Despite the findings from the aforementioned studies, the contribution of agrin to cancer progression remains unknown. To better understand the role of this protein in OSCC, we studied the immunohistochemistry (IHC) staining of agrin in benign, premalignant (OEDs) and malignant lesions. In addition, we modulated the expression of agrin in aberrant keratinocytes and evaluated processes and characteristics associated with cancer progression in vitro and in vivo. Considering the potential of Ct-agrin, we identified its binding proteins (interactome) in an OSCC context using mass spectrometry-based proteomics and bioinformatics. Finally, we found that agrin interactome is related with clinical outcomes of head and neck cancers.
Materials and methods
General design
In the first experiment, we evaluated agrin immunoexpression on tissue biopsies. The effect of agrin silencing in cancer events was evaluated by in vitro and in vivo (orthotopic model) experiments. We induced the overexpression of secreted agrin fragment and ligands able to bind it were identified by mass spectrometry after protein immunoprecipitation. Once the agrin ligands were identified, we visualise the agrin network and evaluate its potential prognostic value.
Subjects
Patients
We retrospectively collected OSCCs (n = 58), OEDs (n = 40) and benign lesions (n = 35) from two oral pathology services (University of Talca and University of Campinas). The information of patients is provided in Supplementary Table S1.
Cells
The following cell lines were used: HMK and HaCaT (nontumourigenic human keratinocytes), SCC-9 and SCC-25 (OSCC), HSC-3 and SCC-9-LN1 (highly invasive and metastatic OSCC cells), and HEK-293 (variable tumourigenic potential). All cells were cultured in recommended media under standard conditions (Supplementary Table S2). HaCaT, SCC9 and HSC-3 were used to establish agrin-silenced cells. The HEK-293 cell line was used to establish cells that overexpress secreted agrin.
Mice
Age-matched NOD-SCID male mice (6 weeks old) were obtained under specific pathogen-free conditions (FMUSP, São Paulo, Brazil). Animals were maintained under controlled conditions with freely available food and water, in groups of four mice each.
Procedures
Measuring agrin expression in oral lesions
Immunostaining was performed using the sodium citrate standard protocol for antigen retrieval. Primary antibody used was an agrin antibody (1:300 dilution; #374117, Santa Cruz). Staining was quantified using the IHC profiler plugin in ImageJ.17 Additionally, IHC slides were evaluated by two pathologists blinded to clinical data who provided a consensus opinion of staining patterns by light microscopy. According to the consensus, we described intensity (signal strength) and geographical spread. Intensity was classified as 0 = negative, 1 = low, 2 = positive and 3 = high. Geographical spread was classified as 0 = no epithelial staining, 1 = lower third, 2 = two thirds or more and 3 = full thickness. For relative risk (RR) analysis, grades 0–1 were merged into the ‘level 1’ group and grades 2–3 were labelled as ‘level 2’. Briefly, RR corresponded to the proportion of cancer or premalignant lesions in cases with higher expression of agrin divided by the proportion of cancer or premalignant lesions in cases with lower expression.
Generation of agrin-silenced cells
We performed agrin (isoform 1, also known as secreted agrin) silencing studies using short hairpin RNA (shRNA)-expressing vectors. We cloned agrin shRNA constructs #TRCN0000056390 and #TRCN0000056391 (RNAi Consortium) into pLKO.1-TRC plasmid (Addgene #10878). We chose the first target for functional experiments (target and template sequences shared a complete alignment). pLKO.1-shGFP was used as control (shControl).18 Target sequences are as follows: shAgrin 5′-CCTGCTCTACAACGGGCAGAA-3′ and shControl 5′-CAAGCTGACCCTGAAGTTCAT-3′. Procedures for packaging shRNA-encoding lentivirus were performed at the Viral Vector Laboratory (LNBio-CNPEM, Campinas, Brazil). We generated two cell groups: agrin-silenced cells (shAgrin) and control cells (shControl). Agrin silencing was verified by real-time quantitative PCR (RT-qPCR) and western blot.
Generation of C-terminal agrin-overexpressing cells
We simulated a secreted bioactive fragment of agrin using the C-agrin4,19-GFP construct19 (Ct-agrin; Supplementary Figure 2A). This fragment was kindly gifted by Dr Matthew P. Daniels (NIH, Bethesda, MD, USA). We used as control a FLAG-tagged green fluorescent protein (GFP) (named IP-control) cloned into a pcDNA3 vector (Invitrogen, Thermo Fisher Scientific Inc.). Ct-agrin construction produced cytotoxicity in SCC-9 and HSC-3 cells (Supplementary Figure 2B). This problem was solved by transfecting HEK-293 cells using polyethylenimine (Polysciences Inc.) (Supplementary Figure 2C).
Gene expression analysis
RNA isolation and RT-qPCR were performed according to previously published protocol.20 Primers used in this study can be found in Supplementary Table S3.
Immunoblotting detection
Cell lysate and secretome isolation for western blot analysis were performed as previously described.20, 21 Antibodies used are listed in Supplementary Table S4.
Agrin and cancer-associated events
Proliferation
HaCaT, SCC9 and HSC-3 cells either transduced with control shRNA or agrin shRNA (1 × 104 cells/well) were seeded in 96-well plates. Bromodeoxyuridine labelling assay was performed as described.22
Migration and tumour cell invasion activity
Motility assays were performed as described,23 with some modifications. For migration, we used 7.4 × 104 cells/well (HaCaT, SCC-9 and HSC-3) and 24-well chambers with uncoated 8-mm pore polycarbonate membranes. For invasion, we used 8 × 104 cells/well (SCC-9 and HSC-3) and 96-well chambers precoated with Matrigel Basement Membrane Matrix (BD Biosciences).
Cancer colony formation
Control or agrin shRNA-tranduced SCC-9 and HSC-3 cells (5 × 103 cells/well) were cultured in 6-well plates. The culture medium was changed every 2 days. After 9 days, cells were stained with 4% formaldehyde/0.005% gentian violet solution. Images were captured with an inverted microscope. We quantified colonies using ImageJ histogram tool (darkest pixels were analysed).
Multicellular tumour spheroid formation
To simulate cancer cells in the blood or lymphatic circulation, we performed a three-dimensional tumour sphere culture. Control or agrin shRNA-tranduced SCC9 and HSC-3 cells (6 × 105 cells/dish) were cultured in non-adhesive conditions, as described previously.24 The multicellular tumour sphere area was analysed using the ImageJ particle analysis tool.
In vivo tumourigenesis and aggressiveness of lesions
We utilised a murine orthotopic model for OSCC. Mice were randomly divided into the following 2 groups (n = 8 animals in each): HSC-3 shControl and shAgrin. Then 2.5 × 105 cells/tongue in 20 μL of Matrigel were intrabuccally implanted into the right lateral portion of the tongue. Animal health was monitored daily. After 21 days, tumour severity was established by the presence of ulcerations and conventional histopathological examination. We evaluated the growth pattern, keratinisation, cell morphology, angiogenesis and vascular (intravascular tumour thrombus) and neural invasion.
Rescue-like experiments
To evaluate whether the specificity of agrin rescues the phenotypes, we exchanged the conditioned media from HSC-3 shControl and shAgrin cells in proliferation, invasion and wound-healing assays. Conditioned media (10 μg) from each group (hereafter called rescue-like medium and shAgrin medium, respectively) was added in serum-free media as described previously.20 For wound-healing experiments, 2 × 104 cells/well were used as described previously.25
Determination of agrin-binding partners
Immunoprecipitation
We used the secretome extract of Ct-agrin and IP-control cells. Immunoprecipitation was performed at 4 °C with 2.5 μg of GFP antibody (#af4240, R&D Systems) in the presence of 30 µL of protein G-Sepharose 4 Fast Flow (GE Healthcare) for 2 h in a rocker. Sample proteins of 250 μg were added and incubated with the beads overnight at 4 °C. The sepharose-bound proteins were washed with cold TBST. Bound proteins were eluted with 4× Laemmli sample buffer at 95 °C for 10 min and resolved by 10% sodium dodecyl sulphate-polyacrylamide gel electrophoresis (SDS–PAGE) for subsequent western blot analysis.
Protein identification
From 3 independent experiments, we excised, reduced, alkylated, trypsin-digested and desalted 60 SDS-PAGE gel bands containing proteins of agrin complexes according to previous protocols26 (Supplementary Figure 2D). Tryptic digested peptides were identified in a LTQ Orbitrap Velos mass spectrometer (Thermo Fisher Scientific Inc.) according to previous protocols.26 Identification of proteins was performed using the MaxQuant27 and Perseus28 software, as described previously.29 For bioinformatics analysis, we focussed only on proteins identified exclusively in the Ct-agrin group. See Supplementary file for further details and explanation.
Agrin interactome characterisation
Bioinformatics
We used Integrated Pathway Analysis Database for Systematic Enrichment Analysis (IPAD)30 to evaluate the inter-association between our identified protein lists and diseases. In IPAD, Fisher Exact test is adopted to measure the gene enrichment in annotation terms and the enrichment between components. IPAD adjust the P-value by Benjamini–Hochberg method.31 We considered a strong model if oral cancer appeared within the top five ranked epithelial diseases. Agrin-binding partners (Ct-agrin exclusive proteins) were described using FunRich.32 Then networks and hubs were visualised using Contextual Hub Analysis Tool (CHAT app) in the Cytoscape software.33, 34 To prioritise proteins, we used the cBio cancer genomics portal35 and HNSCC-TCGA data set36 selecting the genomic profiles by default (mutations, putative copy-number alterations from GISTIC, mRNA Expression z-Scores [RNA Seq V2 RSEM] and protein expression Z-scores [RPPA]). To explore whether prioritised candidates represent a community, we used the STRING database.37 In addition, we evaluated the gene-expression levels of agrin contextual hubs in shControl and shAgrin cells using RT-qPCR. The agrin group also was analysed using SMART38 and PAZAR database.39
Overall survival data
We evaluated the prognostic potential of agrin contextual hubs. Gene expression data in head and neck cancers was obtained from the following publicly available databases: PROGgeneV240 (GSE65858 data set) and SurvExpress41 (HNSCC-TCGA provisional data set).
Statistics
All independent experiments were performed in triplicate. The results are presented as the mean ± standard deviation (SD). We analysed differences between groups using Chi-Square, Student’s t-test, Fisher’s exact test and one-way analysis of variance (ANOVA with post hoc Tukey) tests. In all the procedures, we used a 95% confidence level (P-value ≤ 0.05).
Results
Agrin is elevated in malignant and premalignant lesions
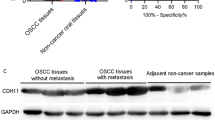
Oral cancer is a multi-stage disease.2 We believe that its progression can be studied by comparing benign tissues, OEDs and malignant lesions. By IHC, we detected higher expression of agrin in OSCCs (n = 58) and OEDs (n = 40) than in benign tissues (fibrous hyperplasia and hyperkeratosis, n = 35) (Fig. 1a, b). Epithelial staining intensity and reactivity were dichotomised, as shown in Fig. 1c. To further investigate the clinical significance of agrin expression in OSCC progression, we examined the association between agrin staining status and patient diagnosis. The results indicated that high expression of agrin was associated with the presence of OEDs and OSCC (see RRs in Fig. 1d). These results may indicate that, as cells increase agrin expression, the premalignant or malignant changes may become enhanced, shifting the balance from reversible status to tumour progression.
Agrin is overexpressed in oral cancer and epithelial precursor lesions. a Representative microphotographs of agrin immunohistochemical staining in oral tissues. Positive staining of agrin is shown in brown. Scale bars, 200 μm. b Stronger agrin score staining is observed in oral squamous cell carcinoma (OSCC) and oral epithelial dysplasia (OED) (ANOVA followed by Tukey’s test). Data are present as the mean ± SD (*P-value ≤ 0.05, **≤0.001). c Intensity and geographical spread for agrin staining. Premalignant and malignant diagnosis can be correlated with the increase in agrin staining (visualised by black coloured bars). d Expression level of agrin was closely associated with patient diagnosis (relative risk; CI confidence interval, Fisher’s exact test). Agrin intensity and geographical spread were dichotomised as levels 1 and 2
Agrin silencing suppresses cancer progression events, but conditioned media enriched with Ct-agrin rescue these effects
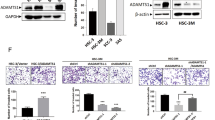
Most oral cancer cell lines secreted high levels of agrin (Fig. 2a, right panel, band having a molecular weight of ~ 72 kDa). We used shRNA technology to knockdown agrin expression in three cell lines. The knockdown efficiency of agrin shRNA was confirmed by RT-qPCR and dot blot (Fig. 2b). Compared to non-target shRNA, treatment with agrin shRNA resulted in a significant decrease in cell proliferation, migration, invasion (Fig. 2c) and EGFR mRNA levels (Supplementary Figure 3). Recent studies indicate that agrin provides stimulatory signals to augment focal adhesion kinase (FAK) activity during cancer growth and invasion.9, 42 FAK induces cell cycle progression through cyclin D1, involving extracellular-signal-regulated kinase (ERK), among others.43 In OSCC cells, treatment with agrin shRNA reduced the levels of FAK and ERK phosphorylated forms, as well as reduced the expression of cyclin D1 (Fig. 2d). In addition, agrin silencing suppresses the colony and tumour spheroid formation (Fig. 2e, f). Taken together, the data suggest that silencing agrin in oral cancer cells results in an impairment of in vitro proliferative and invasive growth programmes. Next, we examined the effect of shControl secretome (conditioned medium) on cell migration, invasion and wound-healing experiments. The results showed that exogenous shControl secretome rescues cell motility in agrin-silenced cells (Fig. 3a, b). Rescue-like medium had a higher capability to stimulate proliferation, invasion and wound healing compared to the secretome originating from shAgrin cells. These results demonstrate that progression of oral cancer may depend, at least in part, on the availability of agrin in the tumour microenvironment.
Agrin silencing decreased oral cancer progression in vitro in less (SCC9) and highly (HSC-3) invasive OSCC cells. a Western blotting confirmed the presence of agrin in different cell lines both in the cell lysate and secretome. Multiple bands may represent proteolytic cleavage products, besides the alternative splicing variants and posttranslational modifications. Vinculin, actin and ponceau red were used as loading control. b Verification of agrin silencing was performed by RT-qPCR and dot blotting. Actin expression was used as an internal control. (c) Agrin silenced cells (shAgrin) proliferate, migrate and invade less compared to the control (shControl; absorbance at 450, 600 and 620 nm, respectively). d Agrin silencing decreased pFAK, pERK and cyclin D1 protein levels. e Focus-formation assay demonstrates less number of colonies in agrin silencing compared to the control. Darkest intensities are represented in millions of pixels. Scale bars, 2 cm. f Agrin silencing interferes with tumour sphere formation. Scale bars, 100 µm. Area in µm2 (x100,000 to SCC9 and x10,000 to HSC-3). For all RT-qPCR experiments, data were normalised with GAPDH gene. Data are represented by the mean ± SD of three independent experiments performed in triplicates (Student’s t-test, *P-value ≤ 0.05, **≤0.001)
Secretome containing agrin plays a role in cancer progression events. In rescue-like experiments, HSC-3 shControl secretome (rescue-like medium) enhanced proliferation, invasion (a) and wound healing (b) in shAgrin cells. Data are represented by the mean ± SD of three independent experiments performed in triplicates (*P-value ≤ 0.05 ANOVA followed by Tukey’s test). Scale bars, 0.1 cm
Agrin silencing reduces tumour aggressiveness
As shown in Fig. 4a, mice that received agrin-silenced cells (shAgrin) developed less aggressive tumours. These tumours did not have ulcers, instead they showed a well-defined mass tumour formation. According to the histopathological evaluation, shAgrin group produced smaller tumours with few instances of vascular and nervous invasion (Fig. 4b). Tumours generated from control cells show significantly greater aggressiveness in comparison to tumours originating from agrin-silenced cells. An additional panel of histological images can be seen in Supplementary Figure 1.
Agrin regulates the aggressiveness of oral cancer. a An orthotopic model of OSCC was established, inoculating HSC-3 cells into the lateral border of the tongue of NOD-SCID mice. Animals received control (shControl) or silenced (shAgrin) cells. Representative images are shown (day 21). Scale bars, 0.2 cm and 400 µm (microphotographs). b Main histopathological characteristics of oral cancers are demonstrated above (*P-value ≤ 0.05 Student’s t-test and Pearson’s Chi-square test)
Agrin interactome reveals an enrichment in cellular growth process
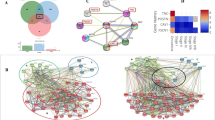
To better understand how agrin participates in malignant progression, we aimed to identify proteins interacting with Ct-agrin in HEK-293 cells. After the immunoprecipitation experiments, proteins bound to secreted agrin complex were identified by mass spectrometry. 197 proteins were analysed exclusively interacting with agrin complex (Supplementary dataset 1). Since that the interactome of a protein can be highly cell-type dependent, we examined all interacting partners using the IPAD30 with focus in malignant diseases. As shown in Fig. 5a, tongue neoplasms are in the top five predicted protein diseases related to agrin partners. We also submitted these candidate proteins to FunRich32 tool for functional analysis. We found that cell growth is the major biological category among the agrin-interacting candidates (Fig. 5b). Interestingly, FunRich identifies the interaction between the growth receptor-bound protein 2 factor (GRB2, not identified in our liquid chromatography–tandem mass spectrometric experiments) with many candidates potentially associated with agrin (Fig. 5c). The interaction of GRB2 with FAK and other proteins leads to activation of the Ras and ERK pathways, which induce tumour cell spreading through cytoskeleton rearrangement.44
High expression of agrin contextual hubs predict poor prognosis. a IPAD predicted diseases with exclusive Ct-agrin ligands as input. Cellular model was strong due to the presence of tongue cancers within the top 5 ranking (top neoplasms from 2,630 diseases). b Classification of biological processes of candidate proteins was performed by FunRich. A total of 197 proteins were analysed. Asterisk (*) represents regulation of nucleobase, nucleoside, nucleotide and nucleic acid metabolism. c FunRich identifies the interaction between GRB2 (predicted) with many candidates potentially associated with agrin. d Prioritised candidates using CHAT app and cBio cancer genomics portal. STRING clustering coefficient and protein–protein interaction (PPI) P-value are shown. e Agrin silencing in OSCC cells alters gene expression of agrin contextual hubs (data for independent replicates are presented in Supplementary dataset 2). f–g Analysis of HNSCC patient survival. Tumour samples that exhibit higher expression of agrin interactome show poor patient prognosis. f, g represent different cohorts. h A hypothetical model suggesting the role of agrin in oral cancer: (i) overexpression of secreted or cleaved agrin triggers elevated binding to its receptors promotes FAK activation and (ii) pro-tumourigenic activation of agrin contextual hubs could potentiate this pathway
Prioritisation of agrin-interacting partners reveals the “agrin contextual hubs”
We assigned numeric values to all identified agrin interacting candidates as follows: −3 (Ip-control-exclusive proteins), +3 (Ct-agrin-exclusive proteins), and 1 (common proteins). Networks were visualised using CHAT app. Using a hypergeometric test, CHAT identifies hub nodes that interact with more “contextual” nodes (i.e. Ct-agrin-exclusive proteins) than statistically expected in networks integrated with user-supplied contextual data (e.g. gene-expression data). CHAT term these nodes as contextual hubs. Contextual hubs are considerably more relevant than degree-based hubs to the specific experimental context under investigation.33 Neighbour interactors were sourced from four databases (InnateDB-all, Mentha, IntAct and UniProt). P-values calculated by CHAT are automatically corrected using the Benjamini–Hochberg procedure. Then we identified the most important centres of activity (the top 20 smallest P-values). To prioritise candidates, we used the cBio cancer genomics portal (2016 version).35 From that top 20, we chose those proteins that presented any alteration (amplification, deep deletion, mRNA upregulation or downregulation, truncating or missense mutations) in a percentage equal to or higher than 20% of The Cancer Genome Atlas HNSCC sample (TCGA Nature 2015, n = 279)36 (Supplementary dataset 1). We selected 9 proteins: cullin-1 (CUL1), cullin-5 (CUL5), eukaryotic initiation factor 4A-II (EIF4A2), protein NDRG1 (NDRG1), polyadenylate-binding protein 1 (PABPC1), dolichyl-diphosphooligosaccharide-protein glycosyltransferase subunit 1 (RPN1), double-stranded RNA-binding protein Staufen homologue 1 (STAU1), titin (TTN), and 14-3-3 protein zeta/delta (YWHAZ). Henceforth, we refer agrin and its nine partners as the “agrin contextual hubs” (the function of each member is described in Table 1). Additionally, Supplementary Table S5 shows OncoPrints (distinct genomic alterations, including somatic mutations, copy number alterations and mRNA expression changes) across a subset of OSCC cases.
Agrin contextual hubs can potentially modulate transcription factors
To explore whether prioritised proteins represent a community (group of nodes with common processes, purposes or functions), we used the STRING database.37 STRING analysis resulted in a network with a high clustering coefficient (proteins have more interactions among themselves than what would be expected for a random set of proteins of similar size, drawn from the genome) (Fig. 5d). To explore this relationship, we evaluated the gene-expression levels of the agrin contextual hubs in shControl and shAgrin cells. Surprisingly, agrin silencing affected gene expression of different members of the agrin group (Fig. 5e). To explore whether agrin serves as a transcriptional regulator, we used SMART.38 According to prediction domain tools, agrin does not contain a DNA-binding domain; however, it does not exclude the possibility that agrin modulates RNA expression, since it can bind to transcription or enhancer factors. Therefore, we have also evaluated whether the transcription factors for the 10 genes overlap, which partially explains the gene-expression regulation that was observed in agrin knockdown condition. Analysing the transcription factors for agrin contextual hubs in the PAZAR database for gene-regulatory information,39 we observed that EGR-1 is the transcription factor for nine out of the ten genes in the agrin group (Supplementary dataset 1).
Agrin contextual hubs represent a community with prognostic potential
Since OSCC is the most common type of malignancy arising from the epithelial cells of the head and neck region,45 we evaluated a clinical relevance of agrin contextual hubs in head and neck cancers. We used ProgGene40 and SurvExpress41 online tools in HNSCC publically available gene-expression databases. High agrin contextual hubs expression was associated with lower overall survival (hazard ratio 7.6, confidence interval 1.7–33.3, P-value ≤ 0.05) in a German cohort followed up for >5 years (GSE65858, n = 269)46 (Fig. 5f). Similarly, in the HNSCC–TCGA provisional (SurvExpress, June 2016) data set (by splitting 502 patients into two maximised risk groups according to their prognostic index), we found that high-risk patients had decreased overall survival (hazard ratio 1.6, confidence interval 1.3–2.7, P-value ≤ 0.05) (Fig. 5g). These results suggest that higher expression of agrin contextual hubs is associated with a poor clinical prognosis.
Discussion
A class of molecules with relevant clinical potential, particularly for HNSCC, is heparan sulphate proteoglycans.47 They can be found on the cell surface and soluble in the ECM. Investigation of these molecules as participants in cancer progression is of great importance and reveals complex relationships occurring at the microenvironment, cellular and subcellular levels.48 In this study, we show that agrin promotes the progression of oral cancer and that its contextual hubs can predict a poor clinical prognosis.
We found that invasive oral carcinomas and premalignant lesions show a strong expression of agrin compared with benign lesions. We calculated RR to use as strength of association49 and biopsies with high agrin staining have 2–7 times the rate of malignant or dysplasia diagnoses compared to samples with low staining. Previous research reported that IHC evaluation of agrin is useful to differentiate benign lesions, dysplasias and hepatocellular carcinomas.50 In fact, intriguingly, we observed the presence of cleaved or secreted agrin in oral cancer cell secretomes but not in normal or immortalised cells. In addition, agrin can help distinguish between primary lesions of liver and metastasis with a high sensitivity and specificity.51 Oral cancer is a multi-stage disease, generated by sequentially malignant events, from epithelial precursor lesions to invasive carcinoma.2 Considering the higher expression of agrin in dysplastic and malignant keratinocytes, and its secretion mainly by cancer cells, it could play a role and/or reflect the process of malignant progression. It was a surprise to identify secreted agrin as a band of ~72 kDa, since the proteolytic results of C-terminal fragment of agrin have molecular weights of ~135, ~ 110, ~90 and ~22 kDa. In a model that evaluated agrin expression in synaptogenesis induced by a traumatic brain injury, the authors found agrin expressed only as species of a molecular weight between 75–55 kDa.52 They suggest that proteolysis may not be the major regulator of agrin expression since fragments generated by MMP-3 and neurotrypsin were not observed. Recent evidence shows that agrin is a target of several metalloproteinases, generating protein subfragments that can have diverse regulatory activities.53 The presence of ~72 kDa agrin band in OSCC secretome may be due to an intermediate processing of known proteases, a cleavage performed by some other protease, or that some alteration in secreted agrin by aberrant squamous cells causes an atypical processing. The nature of this band should be clarified in future work.
To further explore these findings, here we present evidence demonstrating that agrin silencing interferes with oncogenic cell functions. This is consistent with our previous findings where agrin siRNA knockdown promoted a decrease on OSCC cell migration and adhesion.10 Conversely, agrin rescue-like experiments restored proliferative and invasive behaviour in agrin-silenced cells. The secretome is a relevant component for cell–cell communication and the crosstalk between tumour and stroma has a key influence on cancer progression.20 Our results suggest that the observed cell phenotype may be due to the presence of agrin in the tumour microenvironment.
Moreover, in the context of liver cancer, agrin promotes proliferation, invasion and oncogenic cellular signalling.9 The invasive and proliferative phenotypes constitute fundamental biological activities for the progression of malignant diseases,54 and agrin contributes in maintaining these phenotypes. Recent data have shown that FAK pathways are crucial downstream signalling axes of agrin function in liver tumourigenesis.9 In our experiments, we found that inhibition of agrin expression decreased pFAK, pERK and cyclin D1 protein levels. This clearly demonstrates the relationship between agrin and cell cycle progression promoted by FAK also in oral cancer cells. Notably, orthotopic tumours produced by agrin-silenced cells exhibited reduced aggressiveness, showing less vascular and neural invasion, which are associated with a better clinical prognosis.3, 55
We used an immunoprecipitation-based proteomic approach to identify partners of secreted agrin. We discovered 197 proteins potentially interacting with agrin complex. These proteins are mainly related with cell growth, RNA and ubiquitin ligase-binding processes. A close examination using FunRich tool indicated GRB2 (a member of FAK signalling pathway) as an interactor with many proteins potentially presented in agrin complex. It is evident that, in OSCC cells, agrin can be a secreted or cleaved molecule, and as a soluble fragment, agrin can bind to integrins or dystroglycan complex and then could activate FAK,56 which induces cell growth, invasion and metastasis through GRB2, SRC, ERK and cyclin D, among others.43 Our contextual analysis of potential agrin-binding partners, immunopurified from the extracellular space, identified highly connected nodes located in the intracellular compartment. Within these nodes, we prioritised nine hubs to construct the agrin contextual hubs: CUL1, CUL5, EIF4A2, NDRG1, PABPC1, RPN1, STAU1, TTN, and YWHAZ. These proteins may interact directly or indirectly with secreted agrin. Agrin contextual hubs could interfere and enhance the FAK signalling pathway (a proposed model is presented in Fig. 5h). For example, CUL1 is necessary for the expression of SRC family kinases and FAK.57 Conversely, a low expression of CUL558 and NDRG159 allows SRC and FAK activation.
According to our bioinformatics analysis, agrin contextual hubs represent a biologically connected community. This was confirmed by changes in the expression profile of network members when agrin was silenced. For example, an increased expression of NDRG1 was observed in shAgrin cells compared to control cells. Interestingly, NDRG1 lower expression has been previously correlated with tumour progression and poor prognosis in patients with solid tumours.60 Then we further investigated how agrin expression affected gene expression of its contextual hubs. Surprisely, we observed using PAZAR database analysis that EGR-1 is a common transcription factor among all the members of the agrin group. In addition to influencing agrin expression, EGR-1 is involved, either directly or indirectly, in the process of agrin cleavage.61 Agrin is known to activate FAK and it is a critical regulator of YAP/TAZ function.62 A recent work indicates that YAP/TAZ are major oncogenes associated with OSCC.63 Since EGR-1 is a factor that also regulates the transcription of YAP and TAZ (according to PAZAR database), it may be interesting to explore whether EGR-1 silencing affects tumour progression or if agrin depletion affects YAP/TAZ activity in oral cancer cells. In this way, further investigations are needed to elucidate the underlying mechanisms of agrin contextual hubs.
In this research, the histopathological data cumulatively support the pathophysiological role of agrin in oral cancer progression. On the clinical perspective, we demonstrated that patients with HNSCC who show a high gene expression of agrin contextual hubs have a lower survival rate. Since agrin is a gateway for a set of proteins with clinical relevance, it is plausible to think that agrin could be a potential therapeutic alternative for future research. In conclusion, our results underscore agrin expression as a novel marker for malignant and premalignant oral lesions and indicates agrin contextual hubs as a prognostic signature for head and neck cancers.
References
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J. Clin. 65, 87–108 (2015).
Rivera, C. Essentials of oral cancer. Int. J. Clin. Exp. Pathol. 8, 11884–11894 (2015).
Chi, A. C., Day, T. A. & Neville, B. W. Oral cavity and oropharyngeal squamous cell carcinoma–an update. CA Cancer J. Clin. 65, 401–421 (2015).
Rivera, C., Oliveira, A. K., Costa, R. A. P., De Rossi, T. & Leme, A. F. P. Prognostic biomarkers in oral squamous cell carcinoma: a systematic review. Oral Oncol. 72, 38–47 (2017).
Vaupel, P. The role of hypoxia-induced factors in tumor progression. Oncologist 9, 10–17 (2004).
Tanaka, T. & Ishigamori, R. Understanding carcinogenesis for fighting oral cancer. J. Oncol. 2011, 603740 (2011).
Pickup, M. W., Mouw, J. K. & Weaver, V. M. The extracellular matrix modulates the hallmarks of cancer. EMBO Rep. 15, 1243–1253 (2014).
Lu, P., Weaver, V. M. & Werb, Z. The extracellular matrix: a dynamic niche in cancer progression. J. Cell. Biol. 196, 395–406 (2012).
Chakraborty, S. et al. An oncogenic role of Agrin in regulating focal adhesion integrity in hepatocellular carcinoma. Nat. Commun. 6, 6184 (2015).
Kawahara, R. et al. Agrin and perlecan mediate tumorigenic processes in oral squamous cell carcinoma. PLoS ONE 9, e115004 (2014).
Neill, T., Schaefer, L. & Iozzo, R. V. Decoding the matrix: instructive roles of proteoglycan receptors. Biochemistry 54, 4583–4598 (2015).
Scherbakov, N. et al. Evaluation of C-terminal Agrin fragment as a marker of muscle wasting in patients after acute stroke during early rehabilitation. J. Cachex. Sarcopenia Muscle 7, 60–67 (2016).
Yu, D. et al. The reference intervals for serum C-terminal agrin fragment in healthy individuals and as a biomarker for renal function in kidney transplant recipients. J. Clin. Lab. Anal. https://doi.org/10.1002/jcla.22059 (2017).
Klein-Scory, S. et al. Immunoscreening of the extracellular proteome of colorectal cancer cells. BMC Cancer 10, 70 (2010).
Hardt, M., Lam, D. K., Dolan, J. C. & Schmidt, B. L. Surveying proteolytic processes in human cancer microenvironments by microdialysis and activity-based mass spectrometry. Proteomics Clin. Appl. 5, 636–643 (2011).
Wiegand, S. et al. Metaanalysis of the significance of matrix metalloproteinases for lymph node disease in patients with head and neck squamous cell carcinoma. Cancer 104, 94–100 (2005).
Varghese, F., Bukhari, A. B., Malhotra, R. & De, A. IHC Profiler: an open source plugin for the quantitative evaluation and automated scoring of immunohistochemistry images of human tissue samples. PLoS ONE 9, e96801 (2014).
Redis, R. S. et al. Allele-specific reprogramming of cancer metabolism by the long non-coding RNA CCAT2. Mol. Cell 61, 520–534 (2016).
Neuhuber, B. & Daniels, M. P. Targeting of recombinant agrin to axonal growth cones. Mol. Cell. Neurosci. 24, 1180–1196 (2003).
Kawahara, R. et al. Deciphering the role of the ADAM17-dependent secretome in cell signaling. J. Proteome Res. 13, 2080–2093 (2014).
Rodrigues, P. C. et al. Fascin promotes migration and invasion and is a prognostic marker for oral squamous cell carcinoma. Oncotarget 8, 74736–74754 (2017).
Simabuco, F. M. et al. ADAM17 mediates OSCC development in an orthotopic murine model. Mol. Cancer 13, 24 (2014).
Granato, D. C. et al. Integrated proteomics identified up-regulated focal adhesion-mediated proteins in human squamous cell carcinoma in an orthotopic murine model. PLoS ONE 9, e98208 (2014).
Rivera, C. Agarose floor technique: a simple scaffold-free method for 3D cell culture and multicellular tumor spheroid formation. Anal. Quant. Cytopathol. Histpathol. 39, 114–115 (2017).
Rivera, C. Gentian violet used as an epithelial cell monolayer stain in the scratch wound healing assay. Anal. Quant. Cytopathol. Histpathol. 38, 137–138 (2016).
Aragao, A. Z. et al. Novel processed form of syndecan-1 shed from SCC-9 cells plays a role in cell migration. PLoS ONE 7, e43521 (2012).
Tyanova, S., Temu, T. & Cox, J. The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 11, 2301–2319 (2016).
Tyanova, S., Temu, T. & Sinitcyn, P. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Winck, F. V. et al. Insights into immune responses in oral cancer through proteomic analysis of saliva and salivary extracellular vesicles. Sci. Rep. 5, 16305 (2015).
Zhang, F. & Drabier, R. IPAD: the integrated pathway analysis database for systematic enrichment analysis. BMC Bioinformatics 13, S7 (2012).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Stat. Methodol. 57, 289–300 (1995).
Pathan, M. et al. FunRich: an open access standalone functional enrichment and interaction network analysis tool. Proteomics 15, 2597–2601 (2015).
Muetze, T. et al. Contextual Hub Analysis Tool (CHAT): a Cytoscape app for identifying contextually relevant hubs in biological networks. F1000Res. 5, 1745 (2016).
Rivera, C. Immune system and zinc are associated with recurrent aphthous stomatitis. An assessment using a network-based approach. J. Oral. Res 6, 245–251 (2017).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
The Cancer Genome Atlas Network. Comprehensive genomic characterization of head and neck squamous cell carcinomas. Nature 517, 576–582 (2015).
Szklarczyk, D. et al. STRINGv10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–D452 (2015).
Letunic, I. & Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. https://doi.org/10.1093/nar/gkx922 (2017).
Portales-Casamar, E. et al. The PAZAR database of gene regulatory information coupled to the ORCA toolkit for the study of regulatory sequences. Nucleic Acids Res. 37, D54–D60 (2009).
Goswami, C. P. & Nakshatri, H. PROGgeneV2: enhancements on the existing database. BMC Cancer 14, 970 (2014).
Aguirre-Gamboa, R. et al. SurvExpress: an online biomarker validation tool and database for cancer gene expression data using survival analysis. PLoS ONE 8, e74250 (2013).
Chakraborty, S. et al. Agrin as a mechanotransduction signal regulating YAP through the Hippo pathway. Cell Rep. 18, 2464–2479 (2017).
Sulzmaier, F. J., Jean, C. & Schlaepfer, D. D. FAK in cancer: mechanistic findings and clinical applications. Nat. Rev. Cancer 14, 598–610 (2014).
Giubellino, A., Burke, T. R. Jr & Bottaro, D. P. Grb2 signaling in cell motility and cancer. Expert Opin. Ther. Targets 12, 1021–1033 (2008).
Dahiya, K. & Dhankhar, R. Updated overview of current biomarkers in head and neck carcinoma. World J. Methodol. 6, 77–86 (2016).
Wichmann, G. et al. The role of HPV RNA transcription, immune response-related gene expression and disruptive TP53 mutations in diagnostic and prognostic profiling of head and neck cancer. Int. J. Cancer 137, 2846–2857 (2015).
Farnedi, A. et al. Proteoglycan-based diversification of disease outcome in head and neck cancer patients identifies NG2/CSPG4 and syndecan-2 as unique relapse and overall survival predicting factors. BMC Cancer 15, 352 (2015).
Soares, M. A. et al. Heparan sulfate proteoglycans may promote or inhibit cancer progression by interacting with integrins and affecting cell migration. Biomed. Res. Int. 2015, 453801 (2015).
LaMorte, W. W. Measures of association: risk ratios and rate ratios (relative risk). Boston University School of Public Health. http://sphweb.bumc.bu.edu/otlt/mph-modules/ep/ep713_association/ep713_association3. html (2016).
Tatrai, P. et al. Agrin and CD34 immunohistochemistry for the discrimination of benign versus malignant hepatocellular lesions. Am. J. Surg. Pathol. 33, 874–885 (2009).
Somoracz, A. et al. Agrin immunohistochemistry facilitates the determination of primary versus metastatic origin of liver carcinomas. Hum. Pathol. 41, 1310–1319 (2010).
Falo, M. C., Reeves, T. M. & Phillips, L. L. Agrin expression during synaptogenesis induced by traumatic brain injury. J. Neurotrauma 25, 769–783 (2008).
Patel, T. R. et al. Site specific cleavage mediated by MMPs regulates function of agrin. PLoS ONE 7, e43669 (2012).
Gao, C. F. et al. Proliferation and invasion: plasticity in tumor cells. Proc. Natl. Acad. Sci. USA 102, 10528–10533 (2005).
Tai, S. K. et al. Risks and clinical implications of perineural invasion in T1-2 oral tongue squamous cell carcinoma. Head Neck 34, 994–1001 (2012).
Bezakova, G. & Ruegg, M. A. New insights into the roles of agrin. Nat. Rev. Mol. Cell Biol. 4, 295–308 (2003).
Bai, J. et al. Overexpression of Cullin1 is associated with poor prognosis of patients with gastric cancer. Hum. Pathol. 42, 375–383 (2011).
Teckchandani, A. et al. Cullin 5 destabilizes Cas to inhibit Src-dependent cell transformation. J. Cell. Sci. 127(Pt 3), 509–520 (2014).
Wangpu, X. et al. Targeting the metastasis suppressor, N-Myc downstream regulated gene-1, with novel di-2-pyridylketone thiosemicarbazones: suppression of tumor cell migration and cell-collagen adhesion by inhibiting focal adhesion kinase/paxillin signaling. Mol. Pharmacol. 89, 521–540 (2016).
Ando, T. et al. Decreased expression of NDRG1 is correlated with tumor progression and poor prognosis in patients with esophageal squamous cell carcinoma. Dis. Esophagus 19, 454–458 (2006).
MacDonald, R. et al. A novel Egr-1-Agrin pathway and potential implications for regulation of synaptic physiology and homeostasis at the neuromuscular junction. Front. Aging Neurosci. 9, 258 (2017).
Chakraborty, S. & Hong, W. Linking extracellular matrix agrin to the hippo pathway in liver cancer and beyond. Cancers (Basel) 10, pii: E45 (2018).
Hiemer, S. E. et al. A YAP/TAZ-regulated molecular signature is associated with oral squamous cell carcinoma. Mol. Cancer Res. 13, 957–968 (2015).
World Medical Association. World Medical Association Declaration of Helsinki: ethical principles for medical research involving human subjects. JAMA 310, 2191–2194 (2013).
Watson, I. R., Irwin, M. S. & Ohh, M. NEDD8 pathways in cancer, sine quibus non. Cancer Cell 19, 168–176 (2011).
Xie, C. M., Wei, W. & Sun, Y. Role of SKP1-CUL1-F-box-protein (SCF) E3 ubiquitin ligases in skin cancer. J. Genet. Genom. 40, 97–106 (2013).
Okumura, F., Joo-Okumura, A., Nakatsukasa, K. & Kamura, T. The role of cullin 5-containing ubiquitin ligases. Cell Div. 11, 1 (2016).
Modelska, A. et al. The malignant phenotype in breast cancer is driven by eIF4A1-mediated changes in the translational landscape. Cell Death Dis. 6, e1603 (2015).
Dos Santos, M. et al. Prognostic significance of NDRG1 expression in oral and oropharyngeal squamous cell carcinoma. Mol. Biol. Rep. 39, 10157–10165 (2012).
Bekhouche, I. et al. High-resolution comparative genomic hybridization of inflammatory breast cancer and identification of candidate genes. PLoS ONE 6, e16950 (2011).
D’Arcy, P., Wang, X. & Linder, S. Deubiquitinase inhibition as a cancer therapeutic strategy. Pharmacol. Ther. 147, 32–54 (2015).
Oliveira-Ferrer, L. et al. 71P - Relevance of glycosylation-associated genes for tumor progression and metastasis localization in breast cancer. Ann. Oncol. 26, iii23–iii24 (2015).
Kretz, M. TINCR, staufen1, and cellular differentiation. RNA Biol. 10, 1597–1601 (2013).
Boulay, K. et al. Cell cycle-dependent regulation of the RNA-binding protein Staufen1. Nucleic Acids Res. 42, 7867–7883 (2014).
Tadeu, A. M. et al. Transcriptional profiling of ectoderm specification to keratinocyte fate in human embryonic stem cells. PLoS ONE 10, e0122493 (2015).
Watanabe, N. et al. Overexpression of YWHAZ as an independent prognostic factor in adenocarcinoma of the esophago-gastric junction. Am. J. Cancer Res. 6, 2729–2736 (2016).
Funding
CONICYT Becas-Chile Scholarship 8540/2014 (to C.R.), FONDECYT 11140507 (to W.A. G.-A.) and FAPESP Grants 2009/54067-3, 2010/19278-0 and 2016/07846-0 (to A.F.P.L.) supported this work.
Author information
Authors and Affiliations
Contributions
A.F.P.L. and C.R. conceived this research. C.R. and W.A.G.-A. performed the clinical sample collection and classification. C.R., F.S.Z., C.S.-R., C.D.S. and D.C.G. carried out experiments. C.R. analysed the data. C.R. and A.F.P.L. drafted the manuscript. All authors have read and agreed with the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval
All procedures were in accordance with the Helsinki Declaration and the guidelines for the welfare and use of animals in cancer research.64 This research was approved by the Ethics Committee of FOP-UNICAMP (CAAE protocol #78069317.0.0000.5418) and the Animal Ethics Committee CEUA-CNPEM (protocol #13-2015).
Competing interests
The authors declare no competing interests.
Additional information
Note: This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution 4.0 International (CC BY 4.0).
Electronic supplementary material
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Rivera, C., Zandonadi, F.S., Sánchez-Romero, C. et al. Agrin has a pathological role in the progression of oral cancer. Br J Cancer 118, 1628–1638 (2018). https://doi.org/10.1038/s41416-018-0135-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-018-0135-5
- Springer Nature Limited
This article is cited by
-
Oral lichen planus interactome reveals CXCR4 and CXCL12 as candidate therapeutic targets
Scientific Reports (2020)