Abstract
Although aberrant alveolar myofibroblasts (AMYFs) proliferation and differentiation are often associated with abnormal lung development and diseases, such as bronchopulmonary dysplasia (BPD), chronic obstructive pulmonary disease (COPD), and idiopathic pulmonary fibrosis (IPF), epigenetic mechanisms regulating proliferation and differentiation of AMYFs remain poorly understood. Protein arginine methyltransferase 7 (PRMT7) is the only reported type III enzyme responsible for monomethylation of arginine residue on both histone and nonhistone substrates. Here we provide evidence for PRMT7’s function in regulating AMYFs proliferation and differentiation during lung alveologenesis. In PRMT7-deficient mice, we found reduced AMYFs proliferation and differentiation, abnormal elastin deposition, and failure of alveolar septum formation. We further shown that oncogene forkhead box M1 (Foxm1) is a direct target of PRMT7 and that PRMT7-catalyzed monomethylation at histone H4 arginine 3 (H4R3me1) directly associate with chromatin of Foxm1 to activate its transcription, and thereby regulate of cell cycle-related genes to inhibit AMYFs proliferation and differentiation. Overexpression of Foxm1 in isolated myofibroblasts (MYFs) significantly rescued PRMT7-deficiency-induced cell proliferation and differentiation defects. Thus, our results reveal a novel epigenetic mechanism through which PRMT7-mediated histone arginine monomethylation activates Foxm1 transcriptional expression to regulate AMYFs proliferation and differentiation during lung alveologenesis and may represent a potential target for intervention in pulmonary diseases.
Similar content being viewed by others
Introduction
Alveologenesis is the terminal step of lung development in which mature pulmonary gas exchange units (alveoli) are formed through a dramatic remodeling of the large, thick-walled primitive saccules [1, 2]. Abnormal alveologenesis has devastating effects and is often associated with chronic lung diseases such as bronchopulmonary dysplasia (BPD) [2,3,4]. The mechanisms involved in alveologenesis include the expansion of alveolar epithelium and alveolar myofibroblasts (AMYFs) localizing to the tip of crests to deposit elastin, which produce the necessary force for lifting the alveolar crest from the primary septa wall [5, 6]. AMYFs is a special type of fibroblast, which has the characteristics of fibroblast and smooth muscle cells in both ultrastructure and physiological function [7]. In the pseudoglandular phase of lung development, a group of PDGFRα-expressing interstitial cells around the distal epithelium began to appear, which were the precursors of AMYFs [8]. Later on, in the saccular phase, PDGFRα positive cells migrated to the periphery of the terminal cyst wall to form the secondary crest, and differentiated into mature α-SMA+ AMYFs to produce elastin [9]. Disruption of AMYFs differentiation, proliferation, or migration leads to the failure formation of the secondary septum and arrested alveolarization during lung development [10], and are found in several pulmonary diseases including emphysema, idiopathic pulmonary fibrosis (IPF), BPD, and chronic obstructive pulmonary disease (COPD) [11]. Moreover, given that AMYFs are the primary cell types responsible for the accumulation of extracellular matrix (ECM) during fibrotic diseases, targeting AMYFs proliferation and differentiation is an important therapeutic strategy for the treatment of pulmonary diseases [12].
AMYFs specification, proliferation, and migration are tightly regulated during lung development. Platelet-derived growth factor receptor α (PDGFRα) signaling is crucial for AMYFs differentiation and maturation [8, 9, 13]. It is reported that disruption of PDGFA in mice results in failure of elastic fiber deposition and secondary septum formation by repressing myofibroblast (MYF) differentiation [9]. Insulin-like growth factor 1 (IGF1) and transforming growth factor β1 (TGF-β1) are known to be required for AMYFs specification in mice. IGF1 promotes α-SMA expression and elastin deposition in MYFs of developing lung tissue, and overexpression of IGF1 enhances MYFs migration and proliferation in vitro [14]. TGF-β1 was reported to promote the differentiation of lung fibroblasts into MYFs, and deletion of SMAD3, an important intermediate signal molecule of TGF-β1, and results in failure of septa formation and decrease elastin deposition phenotype in mice [15]. In contrast to the well-established roles of these signaling factors, relatively little is known about how epigenetic mechanisms regulating the proliferation and differentiation of AMYFs during lung development and pulmonary diseases.
Protein arginine methyltransferase 7 (PRMT7) was reported as a type III enzyme that catalyze monomethylation and symmetrically di-methylation of arginine residues on both histone and nonhistone substrates to modulate diverse biological processes, including cell proliferation, carcinogenesis, and stem cell biology [16]. Our previous studies have shown that PRMT7 catalyzes monomethylation at histone H4 arginine 3 (H4R3me1) to negatively regulate the expression of Bcl6 during B-cell maturation [17]. To ensure timely activation of antiviral defense, PRMT7 is tightly controlled to suppress MAVS activation [18]. PRMT7 is also involved in the regulation of germ cell proliferation and myoblast differentiation, PRMT7 deficiency causes impaired myogenic differentiation and defects in regenerative capacity upon muscle injury [19, 20]. PRMT7 is also reported to suppress cellular senescence through methylates GLI2 on arginine residues 225 and 227 in mouse embryonic fibroblasts (MEFs) [21]. Moreover, PRMT7 contributes to promoting metastasis in human breast cancer, non-small-cell lung cancer cells, and renal cell carcinoma [22,23,24], and it is increasingly being recognized as a potential drug target [24, 25].
In this study, we show that PRMT7-deficient mice exhibited enlarged distal airspaces secondary to the failure of alveolar septation, impaired AMYFs differentiation, proliferation, and elastin deposition. We also identified that PRMT7 and H4R3me1 directly associate with Foxm1 to activate its transcription, and thereby enhance AMYFs proliferation and differentiation. Collectively, these findings represent one mechanistic explanation for how PRMT7 regulating AMYFs proliferation and differentiation during lung alveologenesis, and may provide potential clues for the treatment of pulmonary diseases caused by AMYF proliferation and differentiation disorders.
Results
Inactivation of PRMT7 disrupts alveolar development in mice
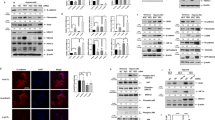
By analyzing the PRMT7 expression level in various tissues, we detected more abundant expressed PRMT7 in the lung, brain, stomach, and spleen tissues at postnatal day (P) 2 (Fig. 1A). We then examined the temporal expression level of PRMT7 at different lung developmental stages and found that the PRMT7 expression was notably increased from P2 to P14 (Fig. 1B), a time period when the parenchymal walls were undergoing the process of alveologenesis. Histological analysis revealed that whereas at embryonic day (E) 18.5, PRMT7−/− lungs were indistinguishable from controls (Fig. 1C), at P2 PRMT7−/− lungs failed to initiate alveolarization, secondary septa were less frequent, and distal airspaces were enlarged compared with controls, and the more apparent failure of septa formation was detected by P6 in PRMT7−/− lungs (Fig. 1C). Detailed analyses of inflated lung structures revealed that the average size of alveolar was significantly increased in PRMT7−/− lungs compared with control mice (Fig. 1D). The mean linear intercept, which describes the mean free distance in airspaces and a surrogate parameter for alveoli, was significantly larger in PRMT7−/− than in control lungs (Fig. 1E). Collectively, these findings suggest PRMT7 is required for septa formation during lung alveologenesis in mice.
A Western blot analysis showing the boundary expressed PRMT7 in lung tissues. B PRMT7 expression was significantly increased in lung tissues during P2 to P14. For (A) and (B), GAPDH serves as a loading control. The experiment was repeated three times with similar results. C Representative hematoxylin and eosin (H&E) staining showing severely enlarged alveolar airspace in PRMT7−/− lungs at P2 and day P6. Boxed regions are magnified in insets. Scale bars: 100 μm. D, E Quantitated bar graphs showing the average size of distal airways (D) and the mean linear intercept (E), which describes the mean free distance in airspaces, was significantly larger in PRMT7−/− lungs. Data were presented as mean ± SD. n = 8 mice per genotype. **P < 0.01 (Student’s t-test).
Decreased number of AMYFs and compromised elastin deposition in postnatal PRMT7 −/− lungs
To investigate the mechanism of the alveologenesis defects in PRMT7-knockout mice, we examined cell differentiation in PRMT7−/− lung tissues by immunofluorescence staining. All checked proximal airway epithelial cell types, including proximal progenitor cells (Sox2+), Clara cells (CC10+), ciliated cells (Ac-tubulin+), and neuroendocrine cells (CGRP+), were present in PRMT7−/− lungs at age of P6 (Fig. S1A). Alveolar type I (ATI) 5 lung epithelial cells (Aqp5+, RAGE+), and alveolar type II (ATII) lung epithelial cells (Sftpc+, Abca3+) in lung distal airway also shown corresponding expression and localization in PRMT7−/− lungs (Fig. S1A). qRT-PCR results showed that T1α (ATI cell marker), Sftpc (ATII cell marker), and Scgb1a1 (Clara cell marker) expression were comparable between control and PRMT7-deficient lungs (Fig. 2A). However, one of the mesenchyme cell markers, alpha-smooth muscle actin (α-SMA), which labels parabronchial smooth muscle cells and AMYFs in lung tissues [26], was significantly reduced expressed in PRMT7−/− mice at P6 (Fig. 2A, B). Immunohistology analysis revealed that the α-SMA+ AMYFs from control animals exhibited prominent α-SMA immune reactivity near tips of secondary septa, while the α-SMA immune reactivity was strongly reduced and α-SMA+ cells were rarely associated with recognizable secondary septa in PRMT7 mutant lungs at P6 (Fig. 2C). Moreover, no apparent differences of α-SMA expression were observed between control and PRMT7−/− lungs at E18.5 (Fig. 2C), and the α-SMA+ parabronchial smooth muscle cells and vessel associated α-SMA+ cells were also not significantly affected by PRMT7 deletion at P6 (Fig. S1B). Quantification of α-SMA+ cells in the alveolar region revealed a significantly decreased number of AMYFs in PRMT7−/− lungs at P6 (Fig. 2D). Given that the most important role of mature AMYFs during alveolar morphogenesis is product elastin during alveolar septa formation [5], we conducted Gomori’s aldehyde fuchsin staining for elastin. Enriched elastin was found at the tips of secondary septa in control lungs at P6, whereas the elastin was detected as a disorganized zone rather than foci in the lungs of PRMT7−/− mice (Fig. 2E). Collectively, these results revealed a crucial role of PRMT7 for the elastin fiber deposit of AMYFs during lung alveolarization.
A qRT-PCR validation of expression of indicated genes in control and PRMT7−/− lungs at P6. Data were normalized to β-actin and then to the control expression level. n = 6 biological replicates. Data were presented as mean ± SD. ns not significant, **P < 0.01 (Student’s t-test). B PRMT7 and α-SMA protein expression were detected by western blotting in the lung tissues of PRMT7−/− and control mice at P2 and P6. The relative level of α-SMA are listed above. C Representative α-SMA immunostaining images of PRMT7−/− and control lungs at E18.5 and P6. Scale bars: 100 μm. D Quantitative analysis of α-SMA positive cells in the alveolar region. Fifteen fields from at least three mice were calculated for each condition. ns not significant, **P < 0.01 (Student’s t-test). E Gomori’s aldehyde fuchsin staining of elastin from control and PRMT7−/− lungs at P6. Magnified pictures from the boxed area were shown on the right panel. Arrows point to positive elastin staining signals. Scale bars: 100 μm.
Inactivation of PRMT7 results in decreased AMYFs proliferation
During normal alveolarization, interstitial AMYFs markedly proliferates coinciding with intense alveolar formation [27]. The absence of AMYFs are associated with a lack of secondary septation and alveolarization failure as observed in PRMT7-knockout mice. Therefore, we investigate whether proliferation decrease or apoptosis increase in the PRMT7-deficient mice. Terminal -deoxynucleotidyl transferase-mediated nick end labeling (TUNEL) analysis indicated that only very few cell deaths occurred in PRMT7-deficient lungs as in control (Fig. S2A, B), implying the observed reduction in AMYFs number was not likely caused by apoptosis. Next, we analyzed the AMYFs proliferation status in PRMT7-deficient mice. Immunofluorescence co-staining of Ki67/pH3 and α-SMA was performed in lung tissues of control and PRMT7-knockout mice at P6. As shown in Fig. 3A–D, both Ki67 and pH3 positive AMYFs were decreased in PMRT7−/− lungs. Quantitative analysis showed a significant decrease in the number of proliferating α-SMA/Ki67 and α-SMA/pH3 double-labeled cells in the alveolar region of PRMT7−/− lungs (Fig. 3B, D), while the number of proliferating Ki67/Sftpc double-positive ATII epithelial cells remains comparable between control and PRMT7−/− lungs (Fig. 3E, F). The decreased AMYF proliferation in PRMT7-deficient lungs was further confirmed by using isolated MYFs from the distal lungs. Immunofluorescence staining results showing decreased α-SMA expression in isolated PRMT7−/− cells, recapitulated the in vivo phenotype of PRMT7−/− mice (Fig. S2C). Of note, we found expression of PRMT7 was higher in isolated MYFs than in lung tissues (Fig. S2D). Analysis of the proliferation rates by Ki67 and pH3 immunostaining shown a significantly decreased number of both Ki67+ and pH3+ cells in PRMT7−/− MYFs than in controls (Fig. S2E–H). Therefore, PRMT7 is required for intestinal MYFs proliferation during lung alveologenesis.
A, B Ki67 and α-SMA co-immunostaining showing the reduced myofibroblasts proliferation in PRMT7−/− lungs. Arrows indicate Ki67 and α-SMA double-positive cells. Scale bars: 20 μm. proliferating myofibroblasts were quantified as in (B). C Proliferating myofibroblasts were detected by α-SMA and pH3 co-immunostaining in control and PRMT7−/− lungs at P6. Arrows indicate pH3 and α-SMA double-positive cells. Scale bars: 20 μm. D Quantification of α-SMA and pH3 double-positive cells. E, F Ki67 and Sftpc co-immunofluorescence staining showing normal alveolar type II (ATII) cell proliferation in PRMT7−/− lungs. Arrows indicate Ki67 and Sftpc double-positive cells. Scale bars: 20 μm. Proliferating ATII cells were quantified as in (F). B, D, F Only the number of α-SMA+ cells located within the alveolar region were calculated (excluding parabronchial and vessel associated α-SMA+ cells). Data were shown as mean ± SD. Fifteen fields from at least six mice per genotype were quantified. ns not significant, ***P < 0.001 (Student’s t-test).
Impaired AMYFs differentiation in PRMT7-deficient lungs
Given that the α-SMA positive AMYFs are derived from a population of mesenchymal progenitors expressing PDGFRα [8], we then examined whether PDGFRα expression was affected by PRMT7 deletion. Western blotting analysis showed a significantly decreased α-SMA and elevated PDGFRα expression levels in PRMT7−/− lungs at P6 (Fig. 4A, B). Co-immunostaining of α-SMA and PDGFRα also showed a reduced α-SMA and increased PDGFRα expression in PRMT7−/− lungs at P6 (Fig. 4C). Quantitative analysis results revealed a significantly reduced number of α-SMA+ and increased number of PDGFRα+ cells at P6, whereas the number of α-SMA+ and PDGFRα+ cells were comparable between control and PRMT7-deficient mice at E18.5 and P2 (Fig. 4D, E). In isolated PRMT7−/− MYFs, we consistently detected a significantly reduced number of α-SMA+ and an increased number of PDGFRα+ cells (Fig. 4F, G). Taken together, these results implying that PRMT7 is required for AMYFs differentiation from PDGFRα+ precursors to α-SMA+ MYFs.
A Western blotting analysis showing the increased PDGFRα and decreased α-SMA expression levels in PRMT7−/− lungs at P6. B Quantitative analysis of relative PDGFRα and α-SMA expression levels. The protein expression levels were normalized to GAPDH and the value for control protein expression level being set as 1. n = 4 biological replicates from three independent experiments. ***P < 0.001 (Student’s t-test). C Representative immunofluorescence images showing fewer α-SMA+ cells and more PDGFRα+ cells in PRMT7−/− lungs. Scale bars: 50 μm. D, E Quantitative analysis of α-SMA+ (D) and PDGFRα+ (E) cells in control and PRMT7−/− lungs at E18.5, P2, and P6. Only the number of α-SMA+ or PDGFRα+ cells located within the alveolar region were calculated. Data were presented as mean ± SD. n = 8 mice for each phenotype and developmental stage. ns not significant,**P < 0.01, ***P < 0.001 (one-way ANOVA with Sidak’s test). F Representative double-immunofluorescence staining images of isolated myofibroblasts (MYFs) by using α-SMA and PDGFRα antibodies. Scale bars: 20 μm. G Quantitative analysis of α-SMA+ and PDGFRα+ cells in MYFs. n = 15 fields. ***P < 0.001 (one-way ANOVA with Sidak’s test).
PRMT7 directly binds to the promoter of Foxm1 to activate its expression
To investigate the molecular mechanism involving in AMYFs proliferation and differentiation defects in PRMT7−/− mice, we conducted RNA-sequencing analysis using total RNA from control and PRMT7−/− mice at P2. We found a total of 137 gene expression alterations (24 upregulated and 113 downregulated) with a fold change >2 (p value <0.05) in PRMT7−/− lung tissues (Table S1). To reveal whether particular gene classes were enriched among the differentially expressed genes (DEGs), we performed gene ontology (GO) terms related to cellular component using DAVID software, and found that genes associated with cell division, cell cycle, and DNA replication, were significant highly ranked among the top five categories (Fig. S3). Specifically, we found one set of cell cycle and cell proliferation regulating genes, including the proliferation-specific transcription factor Foxm1 (a typical proliferation factor that plays an important role in organ morphogenesis and development of cancer), and its target genes, such as Ccnb1, Ccnd1, Ki67, Cenpa, Cenpe, Cenpf, Aurka, Plk1, and Ccna2, were significantly decreased in PRMT7−/− lungs (Fig. 5A). The decreased expression of Foxm1 in PRMT7−/− lung tissues was further confirmed by qRT-PCR (Fig. 5B) and western blotting (Fig. 5C). Immunoflourance staining showed that Foxm1 expression was also decreased in isolated MYFs (Fig. 5D). Consistently, significantly decreased expression of Foxm1 target genes were detected in PRMT7−/− lung tissues by qRT-PCR at P6 (Fig. 5E). We further determined whether Foxm1 is a direct target of PRMT7 by chromatin immunoprecipitation (ChIP) assays using chromatin extracts from P2 lungs. With PRMT7 antibody and multiple primer sets within 2 kb region upstream of the mouse Foxm1 transcriptional start site (TSS), we detected appreciable signals at three sites (3, 4, 5) with a region between 416 and 958 bp upstream of TSS. In contrast, PRMT7 binding with Foxm1 was not observed in an unrelated intergenic region and the other two detected regions upstream of the Foxm1 locus (Fig. 5F, G). H4R3me1 binding with Foxm1 was also highly enriched at the site of 3, 4, and 5 in control lungs, which was significantly reduced in PRMT7−/− lungs (Fig. 5H), demonstrating specificity. Furthermore, in consistent with the downregulated expression of Foxm1 in PRMT7−/− lungs, we observed a significantly increased binding of H3K27me2 (a transcriptional inhibition histone modification), and significantly decreased binding of H3K9ac (a transcriptional activation histone modification) in the Foxm1 promoter regions of PRMT7 binding with (Fig. 5I, J). However, although the PRMT7 was enriched in chromatin fraction of lung tissues (Fig. S4A), the global level of H4R3me1, as well as other reported histone arginine methylations, such as H4R3mes2, H3R2me2s, H3R17me2s, H3R8me2s, and H3R8mm, were not significantly altered in PRMT7−/− lungs detected by western blotting (Fig. S4B–E). Thus, the results of these experiments suggest that PRMT7 and H4R3mm1 directly bind with Foxm1 and positively regulate its transcription, thereby modulating the downstream target genes to control AMYFs proliferation and differentiation.
A RNA-sequencing analysis showing representative genes that were differentially expressed between PRMT7−/− and control lung tissues. Red, upregulated genes; green, downregulated genes. B qRT-PCR analysis to validate the expression of Foxm1 in PRMT7−/− and control lung tissues at E18.5, P2, and P6. Gene expression levels were normalized to β-actin. Data were presented as mean ± SD. n = 5 biological replicates, **P < 0.01 (Student’s t-test). C Western blot analysis showing the decreased Foxm1 expression in PRMT7−/− lungs at P2 and P6; the relative Foxm1 levels are listed above. GAPDH was used as a loading control. The experiment was repeated three times with similar results. D Representative immunostaining images showing the decreased Foxm1 expression in isolated PRMT7−/− MYFs. Scale bars: 20 μm. E qRT-PCR validation of expression for levels of Foxm1 downstream genes. Gene expression levels were normalized to β-actin. Data were presented as mean ± SD. n = 6 biological replicates, *P < 0.05, **P < 0.01 (Student’s t-test). F Schematic diagrams of the mouse Foxm1 gene structure, with bars representing the regions examined by ChIP, white boxes represent exons; black lines represent introns. G–J ChIP assay performed with anti-PRMT7 (G), anti-H4R3me1 (H), anti-H3K27me2 (I), and anti-H3K9ac (J) antibodies, at indicated regions of the Foxm1 gene locus in control and PRMT7−/− lung tissues at P2, n = 5 animals for each group. All changes were normalized to inputs. The data shown are the mean ± SEM of three independent experiments. **P < 0.01 (Student’s t-test).
Overexpression of Foxm1 rescues the proliferation and differentiation defects in PRMT7-deficient MYFs
Given that the previously reported function of Foxm1 during MYFs proliferation and differentiation [28], we further determined whether the decreased Foxm1 expression in PRMT7 mutant contributes to the lung defects described above. We overexpressed Foxm1 in isolated MYFs, and MYFs isolated from PRMT7 mutant lungs exhibited significantly decreased proliferation (as evidenced by immunofluorescence staining of Ki67 and pH3) and differentiation (as evidenced by co-immunofluorescence staining of α-SMA and PDGFRα) phenotypes (Fig. 6A, B), recapitulating the in vivo PRMT7-deficient lung phenotypes, suggesting that this in vitro cell culture system would allow us to examine how Foxm1 expression levels contribute to the defects in PRMT7−/− lungs. Furthermore, expression of Foxm1 to the control level significantly rescues both the proliferation and differentiation defects in PRMT7-deficient MYFs (Fig. 6A–C), suggesting that the decreased Foxm1 expression significantly contributes to the lung AMYFs proliferation and differentiation defects in PRMT7−/− mice.
A Representative immunostaining images for Ki67, pH3, α-SMA, and PDGFRα in control and PRMT7−/− MYFs which were transfected with pCMV-Tag2B or pCMV-Tag2B-Foxm1. Note that both proliferation and differentiation were significantly rescued by Foxm1 overexpression in PRMT7-deficient MYFs. Scale bars: 20 μm. B Quantification of the number of Ki67+, pH3+, α-SMA+, and PDGFRα+ cells. Data were presented as mean ± SD. n = 15 fields from six biological replicates of three independent experiments. ns not significant, ***P < 0.001 (one-way ANOVA with Tukey’s test). C Western blots analysis showing elevated α-SMA and decreased PDGFRα expression after Foxm1 overexpression in cultured PRMT7−/− MYFs. The relative protein expression levels are listed above. GAPDH was used as a loading control. The experiment was repeated three times with similar results.
Mesenchymal-specific deletion of PRMT7 mimics the alveolarization defects in PRMT7 −/− mice
By examining the cellular localization of PRMT7 in lung tissues, we found that PRMT7 is ubiquitously expressed both in lung epithelium and mesenchyme (Fig. S5). To address whether epithelial- or mesenchymal-expressed PRMT7 contributes to the lung developmental defects in PRMT7-deficient mice, we conditionally inactivated the PRMT7 gene in either lung epithelium or mesenchyme using a previously reported inducible gene knockout approach [29, 30] (Fig. 7A). PRMT7fl/fl mice were bred into TetO-Cre and SPC-rtTA/Tbx4-rtTA mice to produce triple-transgenic mice, in which PRMT7 was selectively deleted in respiratory epithelial or mesenchymal cells upon administration of doxycycline (Dox). PRMT7fl/fl;SPC-rtTA;TetO-Cre (herein referred to as epPRMT7−/−) and PRMT7fl/fl;Tbx4-rtTA;TetO-Cre (herein referred to as mePRMT7−/−) triple-transgenic mice were generated by maintaining pregnant dams on Dox from E6.5 onwards. Specifically, Cre expression and the efficiency of Cre-mediated PRMT7 deletion was examined by PCR and western blotting respectively (Fig. 7B, C). Of note, PRMT7 protein expression was dramatically decreased in mePRMT7−/− lungs, whereas its expression was slightly decreased in epPRMT7−/− mice compared with controls (Fig. 7C), implying a higher expression of PRMT7 protein in lung mesenchyme than epithelium. By histology analysis of lung structures from epPRMT7−/− and mePRMT7−/− mice, we observed normal alveolar morphogenesis in the lung of epPRMT7−/− mice at P6 and P28 (Fig. 7D, E). However, mesenchymal-specific deletion of PRMT7 in lung tissues resulted in failure of septa formation, decreased α-SMA expression, and reduced elastin deposition (Fig. 7D–H), recapture the global PRMT7-knockout lung phenotypes. Therefore, these results demonstrate that the mesenchymal-expressed PRMT7 is predominantly required during lung alveologenesis.
A Schematic of the specifically knockout strategy used to delete the PRMT7 gene in lung mesenchyme or epithelium lineages. In triple-transgenic mice (epPRMT7−/− or mePRMT7−/−), rtTA is expressed in epithelium or mesenchyme under the control of SPC or Tbx4 promoter, respectively. In the presence of doxycycline (Dox), rtTA binds to the tetO-CMV promoter to activate Cre-recombinase expression, resulted in recombination and deletion of PRMT7 gene in PRMT7fl/fl mice. B PCR genotyping of the offspring from epithelium PRMT7-knockout (epPRMT7−/−) and mesenchyme PRMT7-knockout (mePRMT7−/−). C Western blotting showing downregulated PRMT7 expression levels in epPRMT7−/− and mePRMT7−/− lung tissues. The relative PRMT7 expression level was listed above. GAPDH was served as a loading control. D Representative H&E staining images from epPRMT7−/− and mePRMT7−/− lung tissues at P6 and P28. Scale bars: 100 μm. E Quantitated bar graphs of the mean linear intercept. The mean free distance in airspaces was significantly larger in mePRMT7−/− lungs at P6 and P28, but no difference between control and epPRMT7−/− mice. Data were shown as mean ± SD. n = 8 mice per genotype. ns not significant, ***P < 0.001 (one-way ANOVA with Tukey’s test). F Representative staining images showing decreased α-SMA expression in lung tissues of mePRMT7−/− at P6. Scale bars: 50 μm. G the quantification result of α-SMA positive cells as shown in (F). The α-SMA+ cells located in alveolar regions were quantified. Data were presented as mean ± SD. n = 15 fields from six mice per genotype. ***P < 0.001 (Student’s t-test). H Representative elastin staining images from lung tissue of control and mePRMT7−/− mice. Scale bars: 50 μm. The boxed area was magnified below. Arrows indicate a positive elastin signal.
Discussion
In this study, we provide direct evidence that protein arginine methyltransferase PRMT7 is required during lung alveologenesis through activation of Foxm1. We propose that decreased Foxm1 expression in PRMT7-deficient mice resulting in reduced AMYFs proliferation and differentiation, which contributes to impaired elastin deposition and failure of septa formation in PRMT7−/− lungs (Fig. 8). Our results thus revealed an intriguing epigenetic mechanism whereby PRMT7-mediated H4R3me1 transcriptionally activates Foxm1 expression to regulate proliferation and differentiation of AMYFs during lung alveologenesis.
During lung alveolar morphogenesis, PRMT7 and H4R3me1 directly bind to the promoter of Foxm1 to activate its expression, and thereby promote the proliferation and differentiation of AMYFs. In the absence of PRMT7 and H4R3me1, repressed Foxm1 results in reduced proliferation and differentiation of AMYFs, compromised elastin deposition, failure of septa formation, and eventually form a lung with enlarged distal airspaces structures. The red star indicates the secondary septa.
Our results reveal an important role for histone methylation during lung alveologenesis. Abnormal alveolarization has devastating effects and is present in human disease conditions such as emphysema or BPD that are characterized by destruction or failure of alveoli development [31]. Previous studies suggested that epigenetic mechanisms, such as histone acetylation, play an important role in regulating lung development. Hdac1/2 is required for proximal epithelium development, loss of which leads to defects in early lung branching morphogenesis [32]. Postnatally, Hdac1/2 was necessary for regeneration but not homeostasis of proximal airway epithelial [33]. The Hdac3-mediated epigenetic pathway is critical for the proper remodeling and expansion of the distal lung saccules into primitive alveoli, epithelial loss of Hdac3 in lung epithelium leads to a reduction of ATI cell spreading and a disruption of lung alveolarization [34]. However, the physiological effect of histone methylation during lung development remains largely unknown. The present study provided direct evidence that histone arginine methylation writer PRMT7 is essential for lung alveolar morphogenesis. Global or mesenchymal-specific deletion of PRMT7 gene in mice resulted in failure of septa formation and reduced elastin deposition. Consistently with the function of PRMT7 during lung alveolarization, a significantly highly expressed PRMT7 was detected in lung tissues during P2 to P14, a time period when lungs underwent alveolar morphogenesis and secondary septa formation [2].
Our results identify a typical transcription factor Foxm1, as the key target gene of PRMT7 to regulate AMYFs proliferation and differentiation during alveologenesis. Foxm1 is best recognized as a master regulator of cell cycle progression on the basis of studies in a variety of different cell types [28, 35,36,37], and it is also required for the differentiation and proliferation of many cell lineages during embryonic development [38,39,40,41]. Fibroblast-specific deletion of Foxm1 inhibits fibroblast proliferation, MYF differentiation and therefore protects mice from bleomycin-induced fibrosis [28]. Mice with global deletion of the Foxm1 gene (Foxm1−/−) exhibited an embryonic lethal phenotype between E13.5 and E16.5, due to structural abnormalities in various organs including the lung [38, 39, 42]. Foxm1−/− lungs exhibit a significant reduction in proliferation and differentiation of mesenchymal cells during the canalicular stage of lung development, yet proliferation levels were normal in the Foxm1-deficient lung epithelial cells [39]. In consistent with this, deletion of Foxm1 in developing respiratory epithelium using the surfactant-associated protein C (SPC) promoter also did not alter proliferation rates of the epithelium [43]. In support of the specific role of Foxm1 in lung mesenchymal cell proliferation, we observed a significant decreased proliferation of α-SMA+ AMYFs and normal proliferation of Sftpc+ epithelium in PRMT7−/− lungs. The observation of significant reduction of AMYFs proliferation, differentiation, and septa formation in the mePRMT7−/− but not in the epPRMT7−/− lungs, further support the idea of Foxm1 is specifically required for lung mesenchyme but not epithelium proliferation. Moreover, ectopic expression of Foxm1 was sufficient for rescuing the defective mitotic entry and differentiation in isolated PRMT7−/− MYFs, consistent with the previous finding of that Foxm1 was required not only for cell proliferation but also for MYFs differentiation [28].
Our findings presented here indicate that PRMT7 was recruited to the Foxm1 promoter regions to epigenetic change methylation status of H4R3me1, and thereby activates its transcription. It has been shown that PRMT7 regulates gene expression in a methyltransferase activity-dependent manner, we determine whether the methyltransferase activity of PRMT7 is required for the regulation of target gene expression. Firstly, we found PRMT7 deletion did not alter the global H4R3 methylation levels and other histone methylation directly or indirectly regulated by PRMT7, which is consistent with a previous study that PRMT7 knockdown had no significant effect on global H4R3 methylation [44, 45]. However, through ChIP-qPCR analysis, we found that PRMT7 deficiency markedly decreased H4K3me1 at the Foxm1 promoter in lung tissues. Thus, PRMT7 deletion may lead to the decrease of Foxm1 epigenetic regulation.
In conclusion, the results of this study demonstrate that arginine methylation writer PRMT7 is required for the postnatal lung alveolar morphogenesis by regulating AMYFs proliferation and differentiation through activating of Foxm1. These discoveries revealed a prominent role of PRMT7 and histone arginine methylation during lung alveologenesis, and provide a new approach for therapeutic intervention of pulmonary AMYFs dysfunction-related diseases.
Materials and methods
Animals
PRMT7fl/fl mice were generated as previously described [17]. PRMT7+/− mice were obtained by crossing with ZP3-Cre mice. PRMT7−/− mice were obtained by intercrossing of PRMT7+/−mice. PRMT7+/+, PRMT7+/−, and PRMT7−/− littermates, both male and female between the ages of E18.5, P2, and P6 were used in this study. PRMT7+/+littermates are denoted as control mice. The lung epithelial-specific PRMT7 knockout mice (epPRMT7) were generated by crossing PRMT7fl/fl mice with SPC-rtTA [30] and TetO-Cre mice [29]. Mesenchymal conditional knockout mice (mePRMT7) were generated by crossing PRMT7fl/fl mice with TBX4-rtTA [46] and TetO-Cre mice. SPC-rtTA and TetO-Cre transgenic mice were kindly gifted by Prof. Huajing Wan (Sichuan University). TBX4-rtTA transgenic mouse line was from Prof. Wen Ning (Nankai University). Administration of Dox started from E6.5 to the endpoint of the experiment by feeding the pregnant mice with 625 mg/kg of Dox-containing food. All mice used in this study were bred in the C57BL/6 strain background and housed in a specific pathogen-free condition with a 12 h light/12 h dark cycle in a temperature- and humidity-controlled environment. Mice were genotyped by genomic DNA PCR using primers listed in Table S2 and then were randomly assigned to their conditions.
Histologic analysis, immunohistochemical staining, and H&E staining
For histologic analysis, lungs from control or PRMT7 mutant mice were fixed in 4% (w/v) paraformaldehyde (PFA, Sigma-Aldrich, USA) overnight at 4 °C. After paraffin (Thermo, USA) embedding, tissues were sectioned at 5 µm. For immunohistochemical staining, sections were deparaffinized and rehydrated. Antigen retrieval was performed using the microwave method [17]. Sections were then pretreated with 0.1% Triton X-100 and blocked with 5% BSA for 2 h, followed by incubation with diluted primary antibodies in 3% BSA overnight at 4 °C, 1 h incubation with selected secondary antibodies, and counterstained with hematoxylin. Antibodies used for histologic analysis were listed in Table S3. H&E staining was performed as previously described [17]. Briefly, sections were deparaffinized and rehydrated, followed by staining in harris hematoxylin solution for 8 min. Then, differentiate in 1% acid alcohol for 30 s and counterstain in eosin-phloxine solution for 1 min. At last, dehydrate, clear, and mount.
Immunofluorescence staining, imaging, and quantification
Immunofluorescence staining was performed following a previously published protocol [47] with antibodies listed in Table S3. Briefly, fixed tissues were transferred into 30% sucrose (Sigma, S8501) overnight at 4 °C and embedded with Optimal Cutting Temperature Compound (TFM-5, USA). Six-μm-thick sections were pretreated with 0.5% Triton X-100 in PBS for 15 min at room temperature. Then, samples were incubated with primary antibodies in 3% BSA overnight at 4 °C, followed by secondary antibodies and counterstained with DAPI for 1 h at room temperature. Images were acquired with an Olympus microscope system (Olympus, Japan) and analyzed with Photoshop software (Adobe Systems Software Ireland Ltd, USA). For quantification of lung sections, images were obtained from a minimum of four mice per genotype, and at least three different areas were counted per image of each mouse. For quantification of in vitro cultured cells, multiple fields were calculated as indicated in the figure legends. All quantification experiments were repeated at least three times with similar results.
Gomori’s aldehyde fuchsin staining and TUNEL assay
Gomori’s aldehyde fuchsin staining was performed following g a previously described protocol [48]. Images were obtained from a minimum of three mice per genotype, and a representative image for each mouse was shown. TUNEL assay was performed as described previously [47]. For quantification, images were obtained from six mice per genotype. Cells were counted in at least three different areas of each image for each mouse.
Lung MYFs isolation, culture, and transfection
Lung MYFs were isolated from control and PRMT7−/− animals using a previously reported protocol [49]. In brief, the trachea and proximal airway were first removed, and the edge of the remaining distal lungs were cut and collected by manual dissection. Collected tissues were then finely minced using sterile scissors and digested with 2 mL trypsin (Gibco, USA) for 20 min at 37 °C with shaking at 5-min intervals. The resulting cell suspension was added to 2 mL of grown medium (Dulbecco’s modified Eagle’s medium (DMEM) (Invitrogen, 12100-046), supplemented with 10% fetal bovine serum (FBS). Cells were washed twice with grown medium and plated in six-well plates (NEST, China). Isolated MYFs were cultured in a growth medium plus 100 U/mL penicillin/streptomycin (Gbico, USA) at 37 °C with 5% carbon dioxide. Cells were tested for mycoplasma contamination using TaKaRa PCR Mycoplasma Detection Set (Clontech Laboratories, EUA).
For rescue experiments, MYFs at 60–70% confluence were transfected with the pCMV-Tag2B-Foxm1 or empty vector using LipofectamineTM 2000 (Invitrogen, 11668019) as described previously [50]. In brief, replace the growth medium before 1 h of the transfection, mixed the plasmids and transfection reagents at 1:3 ratios, and incubated for 20 min at room temperature before added to the cells. Six hours later, cells were maintained in a fresh medium for 48 h before being fixed or harvested.
Chromatin immunoprecipitation (ChIP) and western blotting assay
Approximately ten lungs from control and PRMT7−/− mice at age of P2 were isolated and pooled in a ChIP assay as previously described [47]. Primers for quantitative PCR detection of Foxm1 chromatin regions are listed in Table S2. For western blotting, proteins were extracted from lung tissues or cultured MYFs using RIPA buffer (50 mM Tris (pH7.4), 150 mM NaCl, 0.5% sodium deoxycholate, 0.1% SDS, and 1% NP-40) contains proteinase inhibitors (Sigma, P8340). Protein concentration was determined using the BCA assay as previously reported [51]. About 20 μg of total proteins were subjected to a standard western blot analysis using a protocol as previously reported [52]. The antibodies used were listed in Table S3.
RNA extraction, reverse transcription, and quantitative real-time PCR (qRT-PCR)
RNA was extracted using TRIZOL reagent (Ambion, USA) and reversed transcribed with a FastQuant RT Kit (TIANGEN, China) according to the manufacturer’s protocol. RNA was normalized and used for an Agilent Technologies StrataGene Mx3000P real-time PCR system (Agilent, USA). The transcription levels were normalized to internal β-actin expression. At least three biological replicates were performed per genotype and three technical replicates were performed for every independent experiment.
RNA-sequencing assays
For RNA-sequencing, total RNA was extracted from control and PRMT7−/− lungs at P2. For each sample, five lungs were collected and pooled. Sequencing libraries were generated using NEBNext® Ultra™ RNA Library Prep Kit (NEB, USA) and sequenced on an Illumina Hiseq platform (Illumina, USA). The 125 bp/150 bp paired-end reads were generated and reads were aligned to the mouse reference genome (mm10) using TopHat v2.0.12. HTSeqv0.6.1 was used to count the reads numbers, and then the FPKM of each gene was calculated based on the length of the gene and the reads count mapped to this gene. Differential expression analysis was performed using the DESeqR package (1.18.0).
Statistical analysis
All experiments were performed using 4–20 mice or at least three independent repeated experiments with cells. The exact sample size (n) for each experiment was specified in the figure legends. Animals were not excluded from the analysis. Data were expressed as means ± SD or ±SEM, as indicated in the figure legends. Statistical significance were analyzed by GraphPad Prism version 6 (GraphPad Software, San Diego, CA). A two-tailed unpaired Student’s t-test was used for the comparison between two experimental groups. When multiple comparisons were necessary, one-way ANOVA was performed. Prior to making comparisons across values, the normality of distributions was tested and the variance was similar between groups. All statistics are representative of biological replicates. ns, not significant, *P < 0.05, **P < 0.01, ***P < 0.001.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary files.
References
Basil MC, Katzen J, Engler AE, Guo M, Herriges MJ, Kathiriya JJ, et al. The cellular and physiological basis for lung repair and regeneration: past, present, and future. Cell Stem Cell. 2020;26:482–502.
Massaro D, Massaro GD. Developmental alveologenesis: longer, differential regulation and perhaps more danger. Am J Physiol Lung Cell Mol Physiol. 2007;293:L568–569.
Joss-Moore LA, Albertine KH, Lane RH. Epigenetics and the developmental origins of lung disease. Mol Genet Metab. 2011;104:61–66.
Zepp JA, Morrisey EE. Cellular crosstalk in the development and regeneration of the respiratory system. Nat Rev Mol Cell Biol. 2019;20:551–66.
Wendel DP, Taylor DG, Albertine KH, Keating MT, Li DY. Impaired distal airway development in mice lacking elastin. Am J Respir Cell Mol Biol. 2000;23:320–6.
Frank DB, Peng T, Zepp JA, Snitow M, Vincent TL, Penkala IJ, et al. Emergence of a wave of Wnt signaling that regulates lung alveologenesis by controlling epithelial self-renewal and differentiation. Cell Rep. 2016;17:2312–25.
Thannickal VJ, Toews GB, White ES, Lynch JP 3rd, Martinez FJ. Mechanisms of pulmonary fibrosis. Annu Rev Med. 2004;55:395–417.
Boström H, Willetts K, Pekny M, Levéen P, Lindahl P, Hedstrand H, et al. PDGF-A signaling is a critical event in lung alveolar myofibroblast development and alveogenesis. Cell. 1996;85:863–73.
Gouveia L, Betsholtz C, Andrae J. PDGF-A signaling is required for secondary alveolar septation and controls epithelial proliferation in the developing lung. Development. 2018;145:dev161976.
Roelofs AJ, Kania K, Rafipay AJ, Sambale M, Kuwahara ST, Collins FL, et al. Identification of the skeletal progenitor cells forming osteophytes in osteoarthritis. Ann Rheum Dis. 2020;79:1625–34.
Froidure A, Marchal-Duval E, Homps-Legrand M, Ghanem M, Justet A, Crestani B, et al. Chaotic activation of developmental signalling pathways drives idiopathic pulmonary fibrosis. Eur Respiratory Rev. 2020;29:190140.
Glass DS, Grossfeld D, Renna HA, Agarwala P, Spiegler P, Kasselman LJ, et al. Idiopathic pulmonary fibrosis: molecular mechanisms and potential treatment approaches. Respir Investig. 2020;58:320–35.
McGowan SE, Holmes AJ, Mecham RP, Ritty TM. Arg-Gly-Asp-containing domains of fibrillins-1 and −2 distinctly regulate lung fibroblast migration. Am J Respir Cell Mol Biol. 2008;38:435–45.
Nawabi J, Vohlen C, Dinger K, Thangaratnarajah C, Klaudt C, Lopez Garcia E, et al. Novel functional role of GH/IGF-I in neonatal lung myofibroblasts and in rat lung growth after intrauterine growth restriction. Am J Physiol Lung Cell Mol Physiol. 2018;315:L623–37.
Lopez-Rodriguez E, Boden C, Echaide M, Perez-Gil J, Kolb M, Gauldie J, et al. Surfactant dysfunction during overexpression of TGF-beta1 precedes profibrotic lung remodeling in vivo. Am J Physiol Lung Cell Mol Physiol. 2016;310:L1260–71.
Agolini E, Dentici ML, Bellacchio E, Alesi V, Radio FC, Torella A, et al. Expanding the clinical and molecular spectrum of PRMT7 mutations: 3 additional patients and review. Clin Genet. 2018;93:675–81.
Ying Z, Mei M, Zhang P, Liu C, He H, Gao F, et al. Histone arginine methylation by PRMT7 controls germinal center formation via regulating Bcl6 transcription. J Immunol. 2015;195:1538–47.
Zhu J, et al. Arginine monomethylation by PRMT7 controls MAVS-mediated antiviral innate immunity. Molecular Cell. 2021;81:3171–86.
Chen M, Wang Y, Lin L, Dong F, Wu H, Bao S, et al. PRMT7 is involved in regulation of germ cell proliferation during embryonic stage. Biochem Biophys Res Commun. 2020;533:938–44.
Jeong HJ, Lee SJ, Lee HJ, Kim HB, Anh Vuong T, Cho H, et al. Prmt7 promotes myoblast differentiation via methylation of p38MAPK on arginine residue 70. Cell Death Differ. 2020;27:573–86.
Vuong TA, Jeong HJ, Lee HJ, Kim BG, Leem YE, Cho H, et al. PRMT7 methylates and suppresses GLI2 binding to SUFU thereby promoting its activation. Cell Death Differ. 2020;27:15–28.
Geng P, Zhang Y, Liu X, Zhang N, Liu Y, Liu X, et al. Automethylation of protein arginine methyltransferase 7 and its impact on breast cancer progression. Faseb J. 2017;31:2287–300.
Cheng D, He Z, Zheng L, Xie D, Dong S, Zhang P. PRMT7 contributes to the metastasis phenotype in human non-small-cell lung cancer cells possibly through the interaction with HSPA5 and EEF2. Onco Targets Ther. 2018;11:4869–76.
Liu F, Wan L, Zou H, Pan Z, Zhou W, Lu X. PRMT7 promotes the growth of renal cell carcinoma through modulating the β-catenin/C-MYC axis. Int J Biochem Cell Biol. 2020;120:105686.
Szewczyk MM, Ishikawa Y, Organ S, Sakai N, Li F, Halabelian L, et al. Author correction: pharmacological inhibition of PRMT7 links arginine monomethylation to the cellular stress response. Nat Commun. 2020;11:2683.
McGowan SE, Grossmann RE, Kimani PW, Holmes AJ. Platelet-derived growth factor receptor-alpha-expressing cells localize to the alveolar entry ring and have characteristics of myofibroblasts during pulmonary alveolar septal formation. Anat Rec. 2008;291:1649–61.
Ruiz-Camp J, Morty RE. Divergent fibroblast growth factor signaling pathways in lung fibroblast subsets: where do we go from here? Am J Physiol Lung Cell Mol Physiol. 2015;309:L751–55.
Penke LR, Speth JM, Dommeti VL, White ES, Bergin IL, Peters-Golden M. FOXM1 is a critical driver of lung fibroblast activation and fibrogenesis. J Clin Invest. 2018;128:2389–405.
Sauer B. Inducible gene targeting in mice using the Cre/lox system. Methods. 1998;14:381–92.
Tichelaar JW, Lu W, Whitsett JA. Conditional expression of fibroblast growth factor-7 in the developing and mature lung. J Biol Chem. 2000;275:11858–64.
Lignelli E, Palumbo F, Myti D, Morty RE. Recent advances in our understanding of the mechanisms of lung alveolarization and bronchopulmonary dysplasia. Am J Physiol Lung Cell Mol Physiol. 2019;317:L832–87.
Wang Y, Tian Y, Morley MP, Lu MM, Demayo FJ, Olson EN, et al. Development and regeneration of Sox2+ endoderm progenitors are regulated by a Hdac1/2-Bmp4/Rb1 regulatory pathway. Dev Cell. 2013;24:345–58.
Chokas AL, Trivedi CM, Lu MM, Tucker PW, Li S, Epstein JA, et al. Foxp1/2/4-NuRD interactions regulate gene expression and epithelial injury response in the lung via regulation of interleukin-6. J Biol Chem. 2010;285:13304–13.
Wang Y, Frank DB, Morley MP, Zhou S, Wang X, Lu MM, et al. HDAC3-dependent epigenetic pathway controls lung alveolar epithelial cell remodeling and spreading via miR-17-92 and TGF-β signaling regulation. Dev Cell. 2016;36:303–15.
Xue L, Chiang L, He B, Zhao YY. Winoto A. FoxM1, a forkhead transcription factor is a master cell cycle regulator for mouse mature T cells but not double positive thymocytes. PLoS ONE. 2010;5:e9229.
Zhang H, Ackermann AM, Gusarova GA, Lowe D, Feng X, Kopsombut UG, et al. The FoxM1 transcription factor is required to maintain pancreatic beta-cell mass. Mol Endocrinol. 2006;20:1853–66.
Tan Q, Link PA, Meridew JA, Pham TX, Caporarello N, Ligresti G, et al. Spontaneous lung fibrosis resolution reveals novel antifibrotic regulators. Am J Respir Cell Mol Biol. 2021;64:453–64.
Krupczak-Hollis K, Wang X, Kalinichenko VV, Gusarova GA, Wang IC, Dennewitz MB, et al. The mouse Forkhead Box m1 transcription factor is essential for hepatoblast mitosis and development of intrahepatic bile ducts and vessels during liver morphogenesis. Dev Biol. 2004;276:74–88.
Kim IM, Ramakrishna S, Gusarova GA, Yoder HM, Costa RH, Kalinichenko VV. The forkhead box m1 transcription factor is essential for embryonic development of pulmonary vasculature. J Biol Chem. 2005;280:22278–86.
Korver W, Schilham MW, Moerer P, van den Hoff MJ, Dam K, Lamers WH, et al. Uncoupling of S phase and mitosis in cardiomyocytes and hepatocytes lacking the winged-helix transcription factor Trident. Curr Biol. 1998;8:1327–30.
Laoukili J, Kooistra MR, Brás A, Kauw J, Kerkhoven RM, Morrison A, et al. FoxM1 is required for execution of the mitotic programme and chromosome stability. Nat Cell Biol. 2005;7:126–36.
Ramakrishna S, Kim IM, Petrovic V, Malin D, Wang IC, Kalin TV, et al. Myocardium defects and ventricular hypoplasia in mice homozygous null for the Forkhead Box M1 transcription factor. Dev Dyn. 2007;236:1000–13.
Kalin TV, Wang IC, Meliton L, Zhang Y, Wert SE, Ren X, et al. Forkhead Box m1 transcription factor is required for perinatal lung function. Proc Natl Acad Sci USA. 2008;105:19330–5.
Karkhanis V, Wang L, Tae S, Hu YJ, Imbalzano AN, Sif S. Protein arginine methyltransferase 7 regulates cellular response to DNA damage by methylating promoter histones H2A and H4 of the polymerase delta catalytic subunit gene, POLD1. J Biol Chem. 2012;287:29801–14.
Dhar SS, Lee SH, Kan PY, Voigt P, Ma L, Shi X, et al. Trans-tail regulation of MLL4-catalyzed H3K4 methylation by H4R3 symmetric dimethylation is mediated by a tandem PHD of MLL4. Genes Dev. 2012;26:2749–62.
Zhang W, Menke DB, Jiang M, Chen H, Warburton D, Turcatel G, et al. Spatial-temporal targeting of lung-specific mesenchyme by a Tbx4 enhancer. BMC Biol. 2013;11:111.
Li Q, et al. Histone arginine methylation by Prmt5 is required for lung branching morphogenesis through repression of BMP signaling. J Cell Sci. 2018;131:jcs217406.
Gomori G. Aldehyde-fuchsin: a new stain for elastic tissue. Am J Clin Pathol. 1950;20:665–6.
Edelman BL, Redente EF. Isolation and characterization of mouse fibroblasts. In: Alper S, Janssen WJ, editors. Lung innate immunity and inflammation: methods and protocols. New York: Springer; 2018. p. 59–67.
Pang Q, et al. GM130 regulates pulmonary surfactant protein secretion in alveolar type II cells. Sci China Life Sci. 2021 Mar 16. Pubmed PMID: 33740186.
Li Q, Zhao Y, Yue M, Xue Y, Bao S. The protein arginine methylase 5 (PRMT5/SKB1) gene is required for the maintenance of root stem cells in response to DNA damage. J Genet Genomics. 2016;43:187–97.
Li Q, Li Y, Gu B, Fang L, Zhou P, Bao S, et al. Akt phosphorylates Wnt coactivator and chromatin effector Pygo2 at serine 48 to antagonize its ubiquitin/proteasome-mediated degradation. J Biol Chem. 2015;290:21553–67.
Funding
This work was supported by the National Natural Sciences Foundation of China (31730051), the National Key R&D Program of China (2018YFA0800900), the Educational Commission of Anhui Province of China (Grants KJ2020A0044 and KJ2019A0041), and the Open Project of State Key Laboratory of Respiratory Disease (SKLRD-OP-202109).
Author information
Authors and Affiliations
Contributions
H.H., S.B., and Q.L. conceived, designed, and coordinated the study and wrote the manuscript. J.C., J.Z., P.Z., and Y.Q. performed the experiments and analyzed the data. H.H., J.Z., S.B., and Q.L. contributed to experimental design and data analysis. H.W., J.W., and M.M. edited the manuscript. All authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
This study was performed in accordance with the principles of the Helsinki Declaration of the World Medical Association. All animal experiments were carried out in accordance with the protocols approved by the animal welfare committee of the Institute of Genetics and Developmental Biology, Chinese Academy of Sciences (IGDB-CAS).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Anastasis Stephanou
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
He, H., Chen, J., Zhao, J. et al. PRMT7 targets of Foxm1 controls alveolar myofibroblast proliferation and differentiation during alveologenesis. Cell Death Dis 12, 841 (2021). https://doi.org/10.1038/s41419-021-04129-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-021-04129-1
- Springer Nature Limited