Abstract
Eukaryotic five-methylcytosine (m5C) is an important regulator of viral RNA splicing, stability, and translation. However, its role in HBV replication remains largely unknown. In this study, functional m5C sites are identified in hepatitis B virus (HBV) mRNA. The m5C modification at nt 1291 is not only indispensable for Aly/REF export factor (ALYREF) recognition to promote viral mRNA export and HBx translation but also for the inhibition of RIG-I binding to suppress interferon-β (IFN-β) production. Moreover, NOP2/Sun RNA methyltransferase 2 (NSUN2) catalyzes the addition of m5C to HBV mRNA and is transcriptionally downregulated by the viral protein HBx, which suppresses the binding of EGR1 to the NSUN2 promoter. Additionally, NSUN2 expression correlates with m5C modification of type I IFN mRNA in host cells, thus, positively regulating IFN expression. Hence, the delicate regulation of NSUN2 expression induces m5C modification of HBV mRNA while decreasing the levels of m5C in host IFN mRNA, making it a vital component of the HBV life cycle. These findings provide new molecular insights into the mechanism of HBV-mediated IFN inhibition and may inform the development of new IFN-α based therapies.
Similar content being viewed by others
Introduction
RNA epigenetic modifications exert important effects in myriad biological processes by regulating RNA export [1,2,3], stability [4,5,6], and translation [7,8,9]. The regulatory functions of certain eukaryotic RNA modifications, including N6-methyladenosine (m6A), 5-methylcytosine (m5C), and N4-acetylcytidine (ac4C), have been well characterized [10, 11]. In particular, m5C is formed via the addition of a methyl group from S-adenosylmethionine (SAM) to the carbon-5 position of cytosine in RNA; this process is catalyzed by NOL1/NOP2/Sun domain (NSUN) family members or DNA methyltransferase-2 (DNMT2) [12]. In addition to ribosomal RNAs (rRNAs) and transfer RNAs (tRNAs), m5C is present in messenger RNAs (mRNAs) [13]. NOP2/Sun RNA Methyltransferase 2 (NSUN2) functions as a major methyltransferase associated with m5C in mRNA. Meanwhile, Aly/REF export factor (ALYREF) and Y-box binding protein 1 (YBX1) serve as “reader” proteins that recognize m5C and promote mRNA export from the nucleus, or stabilize RNA, respectively [13,14,15,16]. However, the distribution of m5C in mRNA varies among cell types. That is, although m5C is distributed throughout the mRNA transcript, it may be enriched in specific regions, such as the coding sequence (CDS) [17, 18], 3′ untranslated region (UTR) [18, 19] or the region proximal to the start codon [20,21,22].
Hepatitis B virus (HBV) infection is a common cause of cirrhosis and cancer. Although the HBV vaccine offers 98%–100% protection against infection, HBV infection remains a global health issue, with significant morbidity and mortality (https://www.who.int/news-room/fact-sheets/detail/hepatitis-b). HBV contains a relaxed-circular DNA genome of approximately 3.2 kb that is transcribed into four HBV mRNA transcripts (3.5 kb, 2.4 kb, 2.1 kb, and 0.7 kb). Among them, the 3.5 kb mRNA encodes three viral proteins, core antigen (HBc), E antigen (HBe), and polymerase (Pol); the 2.4 and 2.1 kb mRNAs encode three S antigens, preS1, preS2, and HBs; whereas the 0.7 kb mRNA encodes X protein (HBx) [23]. HBx is a multi-functional protein that promotes HBV transcription by interacting with damage specific DNA binding protein 1 (DDB1) and recruiting E3 ligase to degrade the structural maintenance of chromosome protein (Smc)5/6 complex [24]. It also interferes with interferon (IFN) production and signaling by targeting mitochondrial antiviral signaling protein (MAVS) [25, 26], cytokine signaling 3 (SOCS3), and protein phosphatase 2 A (PP2A) [27].
Similar to other RNA modifications, m5C in viral RNA influences viral replication by regulating RNA splicing, stability, and translation [28]. m5C in HIV-1 mRNA is modified by NSUN2, whereas the loss of NSUN2 perturbs HIV-1 alternative splicing and ribosomal recruitment [28]. Meanwhile, m5C in the EBV-encoded non-coding RNA, Epstein-Barr virus-encoded protein 1 (EBER1) is essential for viral lytic replication and negatively impacts RNA stability [29]. Despite these few viral studies emphasizing the importance of m5C modification during viral infection [28,29,30], it remains largely unclear how viruses hijack and manipulate the host m5C system to promote viral replication.
RNA modification also significantly suppresses RNA-induced innate immune responses in mammalian cells [31]. The most abundant internal RNA modification, m6A, serves as a molecular signature that assists viruses, including HBV, in evading the immune system [32,33,34]. Given that stimulator of IFN gene protein (STING)-mediated DNA sensing is deficient in hepatocytes [35], HBV infection triggers an innate immune response through retinoic acid-inducible protein I (RIG-I)-mediated RNA sensing [36]. RIG-I binds to the 5′ epsilon stem-loop of HBV pre-genomic RNA (pgRNA) to induce type III, not type I, interferon responses [36]. However, Adenosine1907 in the 5’ epsilon stem-loop is typically m6A-modified, which reduces the interaction between pgRNA and RIG-I and inhibits IRF-3–mediated IFN production [34]. Meanwhile, m5C-modified nucleotides do not trigger RIG-I-mediated immune signaling [37]. However, the function of HBV m5C in virus-induced immune evasion has not been elucidated.
In this study, we characterize the role of m5C and its methyltransferase NSUN2 in the HBV-induced innate immune response and demonstrate that the precise balance of NSUN2 expression is vital for the HBV life cycle. That is, reduced NSUN2 expression is sufficient for maintaining HBV m5C to facilitate efficient viral infection, however, does not increase IFN m5C abundance, preventing the induction of IFN expression. Our study reveals that HBV m5C and NSUN2 are important factors in HBV-induced innate immune responses.
Results
HBV m5Cs are essential for viral replication
To explore whether m5C is present in HBV mRNA, the overall level of m5C was determined using m5C-methylated RNA immunoprecipitation (m5C-RIP) in HepG2.2.15 cells stably transfected with the HBV ayw strain. Approximately 70-fold more HBV mRNA was immunoprecipitated with an m5C specific antibody compared with the negative control IgG antibody (Fig. 1A), indicating that HBV mRNA contained m5C modifications. The precise m5C positions were then determined by nanopore direct RNA sequencing (DRS-seq) using poly (A) + RNAs purified from HepG2.2.15 cells. Four sites, m5C-705, m5C-1204, m5C-1235, and m5C-1291, were identified by applying two stringent criteria: (1) the m5C rate of each candidate site was at least 90%; (2) the candidate sites could be re-detected by independent DRS (Fig. 1B and Table S1). Consistently, m5C-1204 and m5C-1291 with m5C rate more than 90% were also identified in DRS-seq of poly (A) + RNAs from AAV-HBV transduced mouse liver [38, 39] (Fig. 1B and Table S1). To further confirm the identified HBV m5Cs, bisulfite sequencing (BS-seq) was performed on HBV 1.1-mer-transfected cells. Three sites, m5C-1204, m5C-1235, and m5C-1291, were detected using BS-seq (Table S1), however, their detection rates were lower. Statistical m5C motif analysis using MEME [40] showed no obvious base preference around the candidate m5C sites in either DRS-seq or BS-seq (Fig. 1C).
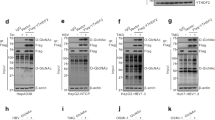
A Total level of HBV m5C quantified by m5C-RIP. HBV RNA transcripts were immunoprecipitated by m5C specific antibody in HepG2.2.15 cells stably transfected with HBV genomic DNA. The enriched HBV RNA was then measured by qPCR with primers targeting the 3′-UTR. B Top panel: rate of each m5C site, named according to HBV genomic position, quantified by Direct-RNA sequencing with purified polyA+ RNAs from HepG2.2.15 cells or AAV-HBV transduced mouse liver. Identified m5C sites with a rate > 90% in two independent experiments with HepG2.2.15 cells are labeled in red, while those with a rate > 90% in one experiment were labeled in green or blue. The m5C sites with a rate > 90% identified in AAV-HBV transduced mouse liver were labeled in yellow. Bottom panel: distribution of four confident m5C locations (color dots) in HBV RNA transcripts. Four major transcripts from HBV genome with coding potentials are shown. C m5C motif predicted by MEME [40]. D–G HBV m5Cs are essential for viral replication. The table shows point mutations introduced to each m5C and the altered amino acids in proteins. D M-ALL mutant containing all m5C mutations. Huh7 cells were transfected with HBV 1.1-mer wild-type or mutants. The core-associated DNA was detected by Southern blot using a probe spanning from nt 1 to nt 3182 at 72 h post-transfection. E RC relaxed circular DNA, DL duplex-linear DNA, SS single-stranded DNA. Secretion of HBeAg (F) and HBsAg (G) in cell culture supernatant quantified by ELISA at 72 h post-transfection.
To investigate the function of HBV m5C in the HBV life cycle, point mutations were introduced at four identified m5C positions in the HBV genome (Fig. 1D). Three non-synonymous mutations were generated due to limited mutagenesis options (Fig. 1D). The viral DNA replication of mutants was then examined by Southern blot. As shown in Fig. 1E, the synthesis of core-associated DNA including relaxed circular DNA, duplex-linear DNA, and single-stranded DNA was relatively eliminated in the C705T, C1291T, C1235G, and M-ALL mutants, and decreased in the C1204T mutant. In addition, compared to wild-type HBV, all mutants resulted in a significant reduction in HBeAg and HBsAg secretion which was quantified by ELISA in Huh7 cells, while the M-ALL mutant exhibited a synergistic inhibitory effect (Fig. 1F, G). These data indicate that the m5Cs are functional and facilitate HBV replication.
HBV m5C-1291 promotes viral RNA export and translation and suppresses RIG-I recognition
m5C has important roles in RNA export, stability, and translation [13, 15]. The HBV genome primarily encodes at least four mRNAs (Fig. 1B); among the four identified HBV m5Cs, only m5C-1291 was located in all four mRNAs and was in the 5’-UTR of the 0.7 kb mRNA. Therefore, m5C-1291 was selected for further functional analysis. ALYREF is an m5C reader that promotes mRNA export [30]. HBV mRNA was co-immunoprecipitated with ALYREF in a formaldehyde-crosslinked RNA immunoprecipitation (RIP) assay, indicating ALYREF bound HBV mRNA (Fig. 2A). Fractionation experiments revealed that ALYREF overexpression significantly increased the export of nuclear HBV mRNA into the cytoplasm (Fig. 2B). The fractionation efficiency was verified by qPCR analysis of glyceraldehyde 3-phosphate dehydrogenase (GAPDH) and U6 (Fig. S1A). To further confirm this result, we constructed a cDNA reporter of 0.7 kb mRNA driven by the CMV promoter according to the reported HBV mRNA isoforms (Fig. 2C) [41]. Notably, the m5C-1291 mutation did not affect the stability of the 0.7 kb mRNA after transcription inhibition by Actinomycin D (Fig. S1B). Meanwhile, ALYREF overexpression significantly promoted the export of the wild-type 0.7 kb mRNA, but not that of the mutant 0.7 kb mRNA (Fig. 2C). The binding efficiency of ALYREF to the mutant 0.7 kb mRNA was lower than that of the wild-type 0.7 kb mRNA in the RIP assay (Fig. 2D). In summary, ALYREF binds m5C-1291 and promotes the export of HBV mRNA.
A HBV transcripts are associated with ALYREF. Huh7 cells were transfected with Flag-ALYREF and then with HBV 1.1-mer at 24 h post-transfection. HBV transcripts were immunoprecipitated by anti-Flag antibody at 48 h post-transfection. The enriched HBV RNA was measured by qPCR (A top). Immunoprecipitated ALYREF examined by western blot. (A bottom). B ALYREF promotes HBV RNA export. Subcellular distribution of HBV transcripts in the cytoplasmic and nuclear fractions examined by qPCR after overexpression of ALYREF for 48 h (B top panel). ALYREF expression examined by western blot (B, bottom). C–E The m5C-1291 promotes 0.7 kb mRNA export. A point mutation of C1291T is shown, which was introduced to m5C-1291 in the cDNA reporter of HBV 0.7 kb mRNA (C bottom). Huh7 cells were co-transfected with Flag-ALYREF, and cDNA of 0.7 kb mRNA-wt or 0.7 kb mRNA-mt for 48 h, respectively. Subcellular distribution of 0.7 kb mRNA-wt or 0.7 kb mRNA-mt were examined as described in (B) (C top). HBV 0.7 kb mRNA wt or mt were then immunoprecipitated by anti-Flag antibody against Flag-ALYREF and measured by qPCR (D). E–G The m5C-1291 promotes 0.7 kb mRNA translation. Huh7 cells were transfected with cDNA of 0.7 kb mRNA-wt or 0.7 kb mRNA-mutant for 48 h. Cell lysates were hyper-centrifuged in a sucrose gradient after cycloheximide treatment, and RNA was extracted from the layer with maximum absorbance at OD260. Ribosome-associated HBV 0.7 kb mRNA was assessed by qPCR (F). GAPDH was used as a control (E). The 5′-UTR of 0.7 kb mRNA-wt or 0.7 kb mRNA-mt were fused to firefly luciferase (Fluc) reporter, respectively (G bottom). Huh7 cells were transfected with wild-type or mutant 5′-UTR Fluc reporter. The relative luciferase activities were calculated by dual luciferase assay at 24 h post-transfection (G top). Renilla luciferase (Rluc) served as an internal control. (H) HBV m5C mutants increase IFN-β mRNA expression. Huh7 cells were co-transfected with poly I:C and HBV 1.1-mer wild type or mutants. The expression of IFN-β was assessed by RT-qPCR at 18 h post-transfection. (I) Expression of five ISGs assessed by RT-qPCR at 18 h post-transfection. J The m5C-1291 blocks RIG-I recognition. RNA-oligo pulldown was performed with huh7 cell lysate using biotin-labeled oligos, which contained m5C-1291, m5C-1255, or non-m5C, respectively. The pulled-down proteins were immunoblotted by anti-RIG-I antibody. Pull down without oligos (Beads) served as the control. The bar represents the mean from three independent experiments. ***P < 0.001; **P < 0.01; *P < 0.05.
The HBV 0.7 kb mRNA potentially encodes the HBx protein. To investigate whether m5C-1291 is important for HBx translation, a ribosome-loading assay was performed to determine the association between the HBV 0.7 kb mRNA and ribosomes. GAPDH was used as a negative control (Fig. 2E). Approximately 4-fold more ribosomes were loaded onto wild-type 0.7 kb mRNA compared to mutant 0.7 kb mRNA (Fig. 2F). Furthermore, we inserted the wild-type and mutant 5′-UTR of the 0.7 kb mRNA into a luciferase reporter and evaluated the luciferase activity via dual-luciferase assay (Fig. 2G bottom) [41]. The wild type 5′-UTR exhibited 2-fold higher luciferase activity than the mutant (Fig. 2G top). Taken together, these data suggest that HBV m5C-1291 promotes 0.7 kb mRNA translation.
RNA modifications suppress toll-like receptor (TLR) and RIG-I triggered innate immune signaling [31, 37]. Transfection with m5C mutants induced upregulation of IFN-β (Fig. 2H and Fig. S1C) and certain ISG mRNAs (Fig. 2I) in poly I:C treated cells by qPCR assays. Similar results were obtained without poly I:C treatment (Fig. S1D, E). RIG-I can be activated by double-stranded RNA in the absence of a 5′-triphosphate terminus [42]; meanwhile, m5C-1291 was predicted to be located in the stem region of a folded 5′-UTR structure (nt 1250–1375 in the HBV genome) by RNA structure folding program (http://www.mfold.org/mfold/applications/rna-folding-form.php; ΔG = −36.40; Fig. S1F and Table S2). To explore whether m5C-1291 affects RIG-I recognition [36], an RNA oligo pulldown assay with Huh7 cell extract containing endogenous RIG-I was performed using an RNA oligo (nt 1250–1329) derived from the 5′-UTR of the 0.7 kb mRNA. Unmodified RNA oligos exhibited a stronger interaction with RIG-I than m5C-modified RNA oligos (Fig. 2J). Collectively, these results indicate that m5C-1291 is important for viral RNA to escape RIG-I recognition.
HBx interacts with EGR1 to downregulate NSUN2 promoter activity
HBV does not encode methyltransferases. Thus, the expression of host m5C “writers,” NSUN2 and DNMT2, and “readers,” ALYREF and YBX1 were analyzed by Western blot [13, 15, 16, 43, 44]. A decrease in NSUN2, DNMT2, and YBX1, and an increase in ALYREF were observed in HepG2.2.15 cells compared with parent HepG2 cells (Fig. 3A, left). The decrease in NSUN2 was not due to stable HBV expression from DNA integration in HepG2.2.15 cells, since this was also detected in HepG2-NTCP cells infected with HBV for 10 days (Fig. 3A middle and Fig. S2A), and Huh7 cells transiently transfected with HBV 1.1-mer for 72 h (Fig. 3A right). The global m5C levels in Huh7 cells was also significantly reduced by HBV-1.1mer transfection. (Fig. 3B). Low NSUN2 expression was consistently observed in HBV patient samples with acute liver failure [45] (Fig. S2B), indicating that HBV infection reduces NSUN2 expression in both cell culture and clinical samples.
A HBV infection downregulates NSUN2 expression. The endogenous expression of NSUN2, DNMT2, ALYREF, and YBX1 quantified by western blot in HepG2.2.15 cells or HepG2 cells (A left), HepG2-NTCP cells with or without HBV infection for 7 days (A middle), and Huh7 cells transiently transfected with HBV 1.1-mer or control vector for 72 h (A right). B The global m5C levels in Huh7 cells transiently transfected with HBV 1.1-mer or control vector for 72 h were examined by m5C dot blot. Methylene blue staining served as the RNA loading control. C, D Viral protein, HBx is responsible for NSUN2 downregulation by HBV infection. Huh7 cells were transfected with Flag-HBx or flag vector, respectively. Transfection reagent served as the MOCK control. The endogenous RNA level of NSUN2 was quantified by qPCR at 24 h post-transfection. GAPDH was used as an internal control (C). The expression of NSUN2 protein and exogenous HBx protein were accessed by western blot, and GAPDH served as the loading control (D). E Huh7 cells were co-transfected with HA-HBx and Flag-NSUN2. Co-IP with an anti-Flag was performed 48 h post-transfection. HBx protein was immunoblotted with an anti-HA. F HBV transfection or HBx expression suppresses NSUN2 promoter activity. A 2-kb DNA fragment (−1900 to +100) of the NSUN2 promoter was fused to Firefly luciferase (Fluc) reporter. Fluc reporter was co-transfected with the control vector, HBV 1.1-mer, flag vector, or Flag-HBx in Huh7 cells. The promoter activity was determined by dual luciferase assay 24 h post-transfection. Renilla luciferase (Rluc) served as a promoter control. G Mapping of the core promoter by 5′-end deletions in the Fluc reporter. The Huh7 cells were transfected with truncated Fluc reporter. The promoter activity was determined by dual luciferase assay at 24 h post-transfection. (H) Identification of HBx response region in the NSUN2 promoter. Two truncated Fluc reporters, −500 or −300 were co-transfected with Flag-HBx or flag vector in Huh7 cells. The promoter activity was determined by dual luciferase assay as above. I Fluc reporter was co-transfected with Flag vector or Flag-EGR1 in Huh7 cells. The promoter activity was determined by dual luciferase assay 24 h post-transfection. J Biotinylated double-stranded WT or MT oligos contained EGR1 or mutated motif, respectively. The pulled-down EGR1 protein by WT or MT oligos was accessed by western blot. The oligo containing YY1 motif served as the positive control. K Oligo pulled down was performed using cell lysates from Huh7 cells transfected with Flag-HBx or flag vector, respectively. EGR1 protein was accessed by western blot. The bar represents the mean from three independent experiments. ***P < 0.001; **P < 0.01; *P < 0.05.
To investigate which viral protein contributes NSUN2 downregulation, HBs HBc, and HBx were overexpressed in Huh7 cells, and endogenous NSUN2 expression was analyzed. HBx overexpression reduced NSUN2 mRNA (Fig. 3C and Fig. S2C, D) and protein levels (Fig. 3D). Notably, HBx did not interact with NSUN2 in the co-immunoprecipitation (co-IP) assay (Fig. 3E) or affect the stability of NSUN2 mRNA in the RNA stability assay (Fig. S2E).
Considering that HBx is a well-studied transcriptional regulator of HBV infection [46], we hypothesized that NSUN2 expression is transcriptionally regulated by HBx. To test this hypothesis, a ~2 kb DNA fragment containing the whole promoter of NSUN2 was inserted into luciferase reporter and the luciferase activities were evaluated using a dual-luciferase assay. Promoter activity was reduced by HBV 1.1-mer transfection or HBx expression (Fig. 3F). To explore the core promoter and cis-regulatory elements, promoter activities of the truncated sequence were tested. Although the −500 and −300 promoters exhibited similar activities (Fig. 3G), only the-500 promoter responded to HBx expression (Fig. 3H). Presumably, the promoter region between −500 nt and −300 nt contained HBx response elements. Moreover, based on our JASPAR database (https://jaspar.genereg.net/) analysis, EGR1, SP1, and ZNF148 may bind to two GC-enriched motifs (−476 to −463 and −350 to −333) in the NSUN2 promoter (Table S3). EGR1, which interacts with HBx [47], was identified as a transactivator of the NSUN2 promoter (Fig. 3I). EGR1 bound to the second GC-enriched motif via DNA oligo pulldown (Fig. 4J), while the expression and binding of EGR1 were decreased by HBx (Fig. 4K). Taken together, these data suggest that HBx suppresses NSUN2 promoter activity by interacting with EGR1 and preventing its binding to the NSUN2 promoter.
A–D NSUN2 serves as the methyltransferase of HBV m5C. A NSUN2 was associated with HBV mRNA. Huh7 cells were transfected with Flag-NSUN2 and HBV-1.1mer. HBV mRNA were then immunoprecipitated by Flag antibody and measured by qPCR at 48 h post-transfection (A top). The immunoprecipitated NSUN2 was examined by western blot (A bottom). Overexpression of NSUN2 enhanced the total level of HBV m5C determined by m5C-RIP (B). Knock-down of NSUN2 by two specific shRNAs decreased the total level of HBV m5C determined by m5C-RIP (C). Mutant NSUN2 (C321A) could not increase the level of HBV m5C modification compared to wild-type NSUN2 (D). E–G Overexpression of NSUN2 suppressed HBV antigen secretion and replication. Transfection with flag-NSUN2 and HBV 1.1-mer in Huh7 cells were performed as above. Secretion of viral proteins, HBeAg (E) and HBsAg (F), in cell culture supernatant were quantified by ELISA at 72 h post-transfection. The core-associated DNA was detected by Southern blot using a probe spanning nt 1 to nt 3182 at 72 h post-transfection (G). RC relaxed circular DNA; DL duplex-linear DNA; SS single-stranded DNA. H–J Knock-down of NSUN2 promotes HBV replication and antigen secretion. NSUN2 was knocked down by two shRNAs as described above. The secretion of HBeAg (H) and HBsAg (I), and the core-associated DNA (J), were examined. (K–M) Restoration of NSUN2 in NSUN2 knock-down cells reduces the enhanced HBV infection. Huh7 cells expressing NSUN2 shRNAs were transfected with shRNA-resistant NSUN2 cDNA by introducing mutations at shRNA seed regions. The secretion of HBeAg (K) and HBsAg (L), and the core-associated DNA (M), were examined as above. ***P < 0.001; **P < 0.01; *P < 0.05.
NSUN2 functions as the methyltransferase of HBV m5C while suppressing HBV infection
To explore which methyltransferase is responsible for the addition of m5C to HBV mRNAs, an RIP assay was performed to determine whether HBV mRNAs are associated with NSUN2 or DNMT2. HBV mRNAs co-immunoprecipitated with NSUN2, not DNMT2 (Fig. 4A and S3A). The overall level of m5C in HBV mRNA was increased by NSUN2 overexpression (Fig. 4B) and decreased by NSUN2 knockdown in m5C-RIP assay (Fig. 4C). However, overexpression or knockdown of DNMT2 did not impact m5C levels in HBV mRNA (Fig. S3B, C). Moreover, the NSUN2 C321A mutant, which disrupts the transient covalent bond to cytosine [28], failed to enhance HBV m5C levels compared with wild-type NSUN2 in m5C-RIP assay (Fig. 4D). In contrast to cytoplasmic replicating viruses [48], nuclear localization of NSUN2 was not altered by transient HBV transfection (Fig. S3D) or stable HBV ayw strain transfection (Fig. S3E). Thus, NSUN2 directly binds to HBV mRNA to catalyze cytosine methylation in hepatocyte nuclei.
Given that host RNA modification enzymes promote viral replication by modifying viral RNAs [28, 49,50,51], the role of NSUN2 in the HBV life cycle was investigated. A significant reduction in HBeAg and HBsAg secretion was observed in cells overexpressing NSUN2 (Fig. 4E, F). Core-associated DNA, including relaxed circular DNA, duplex-linear DNA, and single-stranded DNA, were suppressed by NSUN2 (Fig. 4G). Knockdown of endogenous NSUN2 using two shRNAs enhanced antigen secretion (Fig. 4H, I) and viral replication (Fig. 4J). Restoration of NSUN2 expression via transfection with shRNA-resistant cDNA reversed the enhanced antigen secretion (Fig. 4K, L) and viral replication (Fig. 4M). These data suggest that NSUN2 catalyzes HBV m5C addition, however, suppresses HBV replication.
The delicate balance of NSUN2 expression inhibits IFN production
To investigate the mechanism by which NSUN2 suppresses HBV replication, altered global m5C levels in host genes in HBV 1.1-mer-transfected cells were examined. We performed BS-Seq to analyze differentially methylated cellular RNAs. The sequencing data showed that the m5C modifications were primarily distributed in the 5′-UTR (Fig. 5A). Consistent with the globally reduced m5C levels (Fig. 3B), the average m5C density in CDS and 3′-UTR decreased following HBV 1.1-mer transfection (Fig. 5A and Table S4). A total of 20,148 differentially methylated regions (DMRs) were identified (Table S5), which were hierarchically clustered in a heat map (Fig. 5B). Moreover, the m5C levels were significantly altered in cellular RNA involved in innate immune responses, including those encoding IRF3, IFNB1, IFNAR1, IFNAR2, IFNGR2, and IFNGR1 (Table S5).
A–C BS-seq was performed with total RNA isolated from Huh7 cells transfected with HBV 1.1-mer at 18 h post-transfection. The average m5C density in each gene structure, 5′-UTR, CDS and 3′-UTR was calculated in HBV 1.1-mer transfected or un-transfected cells (A). The heatmap shows the differentially methylated region identified in six samples (B). The line graph shows the m5C density in each gene structure, promoter, 5′-UTR, exon, intron, 3′-UTR and downstream of several genes related to RIG-I signaling (RIG-I, MAVS, TRIM25, IRF3, and IFNB1) (C). D validation of the m5C level of genes related to RIG-I signaling by m5C-RIP assay in HBV 1.1-mer transfected or un-transfected cells. E–H HBV 1.1-mer transfection inhibits poly I:C induced IFN-α and IFN-β production. Huh7 cells were transfected with poly I:C and HBV 1.1-mer, or poly I:C and control vector for 18 h, respectively. The production (E) and mRNA expression (F) of IFN-α and IFN-β were assessed by qPCR or ELISA, respectively. The RNA stability of IFNA1 (G) and IFNB1 (H) RNA was reduced by HBV-1.1 mer transfection. ***P < 0.001; **P < 0.01; *P < 0.05.
Given that the HBV-induced immune response correlates with RIG-I recognition [36], we assessed the m5C densities of molecules related to RIG-I signaling, including RIG-I, MAVS, TRIM25, TBK1, IRF3, IFNA1, IFNA4, IFNB1, and IFNG. Notably, the m5C density in the 5′-UTR of IRF3 and the exon of IFNB1 was significantly decreased (Fig. 5C). Additionally, according to the m5C-RIP analysis, m5C levels were decreased in IFNA1 and IFNB1 RNAs (Fig. 5D). In addition, HBV 1.1-mer transfection inhibited poly I: C-induced IFNA1 and IFNB1 mRNA in qPCR assays (Fig.5E) and protein secretion in ELISA (Fig. 5F), which was associated with reduced RNA stability (Fig. 5G, H). These data imply that HBV transfection suppresses type I IFN production by reducing m5C levels and RNA stability.
Negligible levels of type I IFN have been detected in patients with HBV; however, the mechanism associated with its downregulation is poorly understood [52]. To explore the correlation between NSUN2 and inhibited IFN production, NSUN2 was depleted or overexpressed, followed by measurement of IFN production by ELISA and IFN expression by qPCR assay. As expected, NSUN2 knockdown decreased the poly I: C-induced IFN-α and IFN-α production (Fig. 6A, B). In contrast, overexpression of NSUN2 increased IFN-α and IFN-β production (Fig. 6C, D). NSUN2 depletion also significantly decreased virus-induced IFN production (Fig. 6E). Restoration of NSUN2 in HBV 1.1-mer-transfected cells rescued the inhibited IFN-α and IFN-β production (Fig. 6F, G), as well as the reduced ISGs (ISG20, TRIM14, CBFB, IFI6 and IFI27; Fig. 6H). Taken together, IFN-α and IFN-β production is closely correlated to the m5C levels of their mRNA and NSUN2 expression.
A, B Knock-down of NSUN2 decreases poly I:C-induced IFN-α and IFN-β production. The NSNUN2 or NC shRNA stably expressing cells were treated with poly I:C for 18 h. The protein (A) and mRNA expression (B) of IFN-α and IFN-β were assessed by ELISA or qPCR, respectively. C, D Overexpression of NSUN2 increases poly I:C-induced IFN-α and IFN-β production. Huh7 cells were transfected with poly I:C and Flag-NSUN2, or poly I:C and flag vector for 18 h. The protein (C) and mRNA expression (D) of IFN-α and IFN-β were assessed. E NSUN2 positively regulates IFN-β expression in uninfected, EV71-infected, and VSV-infected cells. Huh7 cells were infected with EV71 or VSV, or without virus for 24 h, and IFN-β mRNA expression was assessed. F–H HBV-mediated IFN inhibition is dependent on NSUN2. Poly I:C was co-transfected with the control vector, HBV 1.1-mer, and flag vector, or HBV 1.1-mer and Flag-NSUN2 in Huh7 cells for 18 h. The protein (F) and mRNA expression (G) of IFN-α and IFN-β were assessed. The expression of several ISGs was accessed by qPCR (H). ***P < 0.001; **P < 0.01; *P < 0.05.
Discussion
Host RNA modification enzymes are reportedly essential for viral replication and pathogenesis by modifying viral RNA [28, 49,50,51]. In the present study, we found that NSUN2 plays an important role in the HBV life cycle (Fig. 7). NSUN2 expression is downregulated by HBx which decreases the binding of EGR1 to the NSUN2 promoter. NSUN2 catalyzes the addition of m5Cs in HBV mRNAs, and functional m5Cs promote HBV RNA export and translation while inhibiting the innate immune response by decreasing RIG-I binding. In contrast, decreased NUSN2 expression reduces m5Cs in IFN RNA, resulting in decreased IFN production. Thus, maintaining a delicate balance in NSUN2 expression during HBV infection is important for the viral life cycle.
During the HBV infection, NSUN2 is transcriptionally repressed by HBx through inhibition of EGR1 binding to its promoter in the nucleus. The reduced NSUN2 catalyzes m5C modifications in HBV mRNAs to promote viral RNA export. However, the m5C modifications in the RNAs of IFN-α and IFN-β were decreased due to the repressed level of NSUN2. After the viral and cellular mRNAs are delivered into cytosol, HBV m5Cs promote viral RNA translation and inhibit RIG-I recognition. The reduced m5Cs in IFN-α and IFN-β RNAs result in low expression of IFN-α and IFN-β, which ultimately leads to efficient HBV replication.
RNA modifications, including m6A, m5C, and ac4C, of viral RNA are indispensable for viral replication as they regulate viral RNA splicing, stability, and translation [28, 51]. Certain functional m5Cs were identified in HBV mRNA by DRS-seq and confirmed by BS-seq. These m5Cs have multiple roles in the HBV lifecycle. First, introducing mutations in HBV m5Cs abolishes viral replication and decreases antigen secretion. Second, HBV m5C modifications promote viral mRNA export by binding to ALYREF [53]. Third, m5C-1291 in the 5′-UTR of the 0.7 kb mRNA promotes HBx translation, which further promotes HBV transcription and influences IFN production [24,25,26,27]. Finally, m5C-1291 attenuates the immune response by decreasing RIG-I binding to the 0.7 kb HBV mRNA. Owing to the importance of this single nucleotide in HBV infection, a specific targeting strategy combined with HBV therapy may improve the efficiency of HBV infection clearance.
Host RNA modification enzymes are often elevated and hijacked by viral proteins to promote the infection [48, 50, 54, 55]. Depletion of the methyltransferase of m6A modification, METTL3, suppresses the production of EV71 [55] and SARS-CoV-2 [48] by disrupting viral RNA stability and translation. However, the expression of NSUN2 was downregulated in HBV-infected cell cultures and patient samples by HBx. To understand the cause of this discrepancy, genome-wide m5C levels in cellular RNAs were investigated in HBV DNA-transfected cells. HBV infection led to reduced m5C in IFN RNA and decreased IFN secretion, likely mediated by reduced NSUN2 expression. Thus, a balance in NSUN2 expression results in decreased m5C levels in IFN RNA, which appears necessary for the virus to complete its life cycle.
NSUN2 expression was moderately downregulated by viral encoded HBx through inhibiting EGR1 binding to NSUN2 promoter. NSUN2 catalyzed the m5C addition to both HBV and IFN mRNAs. The repressed NSUN2 expression decreased m5C levels and RNA stabilities of IFN genes, which led to impair the immune responses; while the existing amount of NSUN2 was sufficient to catalyze the m5C addition in HBV mRNA, which promoted viral RNA exportation and translational efficiency but inhibited the RIG-I recognition of viral RNAs. Thus, we speculated the delicate balance in NSUN2 expression was important during HBV replication.
In previous reports, increased methyltransferases were found to be post-translationally regulated by viral proteins [48, 51, 55]. However, our data revealed a different mechanism of NSUN2 down-regulation during HBV infection, which was transcriptionally suppressed by HBx. We also identified an HBx response region in the NSUN2 promoter. Given that HBx is not a DNA-binding protein, interaction with other host factors or activation of signal transduction is necessary for HBx to target the NSUN2 promoter [56]. In fact, DNA-binding proteins, including DDB1 [24], CBFβ [57], p53 [58] and TFIIB [59] interact with HBx. HBx also physically interacts with EGR1, EGR2, and EGR3, but only transactivates EGR2 and EGR3, not EGR1, by recruiting the coactivator CBP [60]. In this study, we found that EGR1 transactivates the NUSN2 promoter by binding to the second GC-rich motif while HBx interacts with EGR1 and inhibits its binding, resulting in suppressed NSUN2 transcription.
IFN production and signaling pathways are compromised by HBV infection [61,62,63,64], which may account for the failure of IFN-α therapy in many patients with chronic HBV infection [65]. Here we demonstrated that NUSN2 positively regulates the expression of IFN-α and IFN-β production in HBV-infected cells. Thus, we hypothesize that HBV-mediated IFN inhibition occurs through the downregulation of NSUN2. In fact, restoring NSUN2 expression in HBV cell cultures rescued the inhibition of IFN production. However, the opposite function was recently reported for NSUN2; Zhang et.al. found that NSUN2 depletion transcriptionally upregulates polymerase (Pol) III-transcribed ncRNAs, which serve as RIG-I substrates to induce an IFN response in A549 cells. However, the mechanism of NSUN2-mediated transcriptional regulation is unclear [66]. Considering that hepatocytes were used in the current study, we hypothesize that NUSN2-mediated innate immune responses differ in a cell context-dependent manner, however, this requires further confirmation.
Although our data suggest that m5C is preferentially localized in the common region of the four HBV mRNAs, the sequencing reads in our current DRS were not sufficiently long to map all HBV transcripts. Accordingly, we were unable to distinguish transcript-specific m5Cs. Moreover, a previous report found that m5C modifications generated by NSUN2 are typically localized in GC-rich regions of mRNAs [14], but no preferential sequences were identified around the m5C sites by our motif analysis. The motifs of NSUN2-mediated m5C modifications differ between ncRNAs and tRNA [67, 68]. Bohnsack et. al. reviewed the motif of NSUN2-mediated m5C modifications and proposed that NSUN2 targets the secondary structure of RNA substrates rather than a specific RNA sequence [22]. This hypothesis is supported by the notion that NSUN2-mediated m5C34 is dependent on the elongated anticodon stem structure of pre-tRNALeu [68]. Although this study provides valuable insights into the role of m5C and NSUN2 in HBV pathogenesis, their precise functions require further investigation in animal models and clinical patient cases [69,70,71].
In summary, m5Cs and the host methyltransferase NSUN2 are key factors in HBV-mediated IFN inhibition, indicating that they are promising targets for the development or improvement of HBV therapy.
Materials and methods
Cell culture and transfection
Huh7, HepG2, HepG2-NTCP, and HepG2.2.15 cells, which stably express HBV, were cultured in Dulbecco’s modified Eagle’s medium supplemented with 10% fetal bovine serum at 37 °C in a 5% CO2 environment. Plasmid transfection was carried out using Lipofectamine 2000 reagent (Invitrogen, Carlsbad, CA, USA; catalog no. 11668-019) according to the manufacturer’s instructions.
Nanopore direct RNA sequencing (DRS-seq)
PolyA+ RNA was purified from 1 mg of total RNA from HepG2.2.15 cells or AAV-HBV transduced mouse liver [38, 39] using a GenEluteTM mRNA Miniprep Kit (Sigma-Aldrich). Independent experiments of DRS-seq were performed according to the manufacturer’s instructions (Oxford Nanopore DRS protocol; SQKRNA002). Briefly, the sequencing library prepared from PolyA+ RNA was loaded onto the FLO-MIN106D flow cell, followed by a 72 h sequencing run on a MinION device (Oxford Nanopore Technologies). Multi-fast5 reads were base called guppy (v3.1.5) and then converted to single-read fast5 using the command multi_to_single_fast5 in the Oxford Nanopore Technologies API, ont_fast5_api (v3.0.2). Single-fast5 data were mapped to the reference genome of HBV (NC_003977.2) using Tombo (v1.5.1). The specific modifications were detected by the Tombo detect_modifications command, and the all-context alternate model was used to identify 5-methylcytosine in any sequence by running “-alternate-bases 5mC.” Subsequently, the “text_output” command was applied to obtain the methylation score and coverage rate at each position.
Bisulfite sequencing (BS-seq)
Total RNA was extracted from Huh7 cells transfected with Poly I:C and HBV 1.1-mer at 18 h post-transfection using TRIzol™ Reagent (Thermo Fisher Scientific). PolyA+ RNA was then selected and the library was constructed using the EZ RNA Methylation™ Kit (ZYMO RESEARCH) and KC UMI RNA Library Kit (Wuhan SeqHealth Tech) according to the manufacturer’s instructions. Sequencing of three samples in each condition was performed using a MGISEQ-T7 sequencing system (BGI). Sequencing data was uploaded to the GEO repository (GSE246879). Raw reads were first processed using fastp (version 0.23.0) to remove residual adaptor sequences and low-quality reads [72]. The clean reads were then mapped to the reference genome, and duplicated reads were removed using Bismark (version 0.22.3) [73]. The chromosome depth and coverage were calculated using MosDepth (version 0.3.1) and RSeQC (version 4.0.0) [74]. Methylation detection along the whole genome was conducted using Bismark, which calculates the proportion of methylated reads at each genomic site. DMRs between different groups were detected using methylene (version 0.2–8) [75].
Plasmid constructs
NSUN2, DNMT2, ALYREF, HBx, HBsAg, and HBcAg expression plasmids were cloned into the PXJ40-Flag/HA vector. The Flag-C321A plasmid was constructed as described previously [28]. shRNA-resistant cDNAs of NSUN2 plasmids were constructed by introducing four site mutations into the NSUN2 expression plasmids. C1204T, C705T, C1235G, C1291T, and M-ALL plasmids were generated by introducing site mutations into the pch9-3091 plasmid. The HBV 0.7 kb mRNA wt and mt plasmids were generated by inserting the 5′-UTR, CDS and 3′-UTR region of HBx from the pch9-3091 plasmid into the PXJ40-HA vector. The 5′-UTR-wt and 5′-UTR-mt plasmids were generated by inserting the HBx wt 5′-UTR and m5C mutant 5′-UTR into the PGL3-basic plasmid. The NSUN2 promoter luciferase reporter plasmid and truncated plasmids were constructed by inserting the −190 to +100, −1000 to +100, −500 to +100, −300 to +100, −200 to +100, and-100 to +100 promoter regions of the NSUN2 genome from huh7 cells into the vector PGL3-basic.
Dot blot
Ten micrograms of total RNA isolated from huh7 cells were denatured at 75 °C for 5 min and then added dropwise onto the Hybound N+ membrane (GE Healthcare). The membrane was then dried and exposed to UV light to crosslink the RNA samples. The membrane was blocked in 5% skimmed milk and incubated with primary antibody against m5C modification (cat. no. ab10805; Abcam, Cambridge, UK) overnight at 4 °C. After three washes, the membranes were incubated with secondary antibodies [horseradish peroxidase (HRP) goat anti-mouse IgG (H + L) 115-035-003] for 1 h at room temperature. Next, 1 mL of the prepared chemiluminescent reagent mix was added to the membrane after three washes. Immunoblot signals were detected using the Tanon-5200 ChemiDoc MP imaging system (Tanon Science & Technology, Shanghai, China). Three independent experiments of Dot blots were performed, and one representative result was shown Fig. 3B.
Western blotting
Cells were lysed on ice for 30 min with lysis buffer (P0013, Beyotime), centrifuged at 14,000 rpm for 10 min, and denatured at 100 °C. Proteins were separated by SDS-PAGE and transferred to nitrocellulose membranes. The following steps were performed as described previously [51]. The primary antibodies included Flag (cat. no. F1804-1 MG; Sigma-Aldrich), GAPDH (cat. no. 60004-1-lg; Proteintech, Rosemont, IL, USA), NSUN2 (cat. no. 20854-1-AP; Proteintech, Rosemont, IL, USA), DNMT2 (cat. no. 19221-1-AP; Proteintech, Rosemont, IL, USA), ALYREF (cat. no. 16690-1-AP; Proteintech, Rosemont, IL, USA), HA (cat. no. 51064-2-AP; Proteintech, Rosemont, IL, USA), and PreS2 (cat. No. sc-23944; Santa Cruz Biotechnology, Dallas, TX, USA). The anti-HBsAg antibody was gifted by Dr. Bing Yan from the Wuhan Institute of Virology, CAS. At least three independent experiments for each western blot were performed. Original western blots are shown in Supplementary Material.
Immunofluorescence
Cells were seeded in microplates at 50% confluence and then transfected with the HBV 1.1-mer plasmid, pch9-3091, as described above. The following day, cells were fixed with 3.7% paraformaldehyde for 30 min at room temperature 72 h post-transfection and then permeabilized in 0.5% Triton-X100 on ice for 10 min. After blocking in 3% BSA for 1 h, the cells were incubated with primary antibodies at a dilution of 1:50 for 1 h at room temperature and then incubated with secondary antibodies (A-11001 or A-11011, Invitrogen) for 1 h at room temperature. Finally, the samples were stained with Hoechst for 5 min. Images were obtained using a VoX confocal microscope (PerkinElmer, Waltham, MA, USA).
M5C-methylated RNA immunoprecipitation (m5C-RIP) and RT-qPCR
Total RNA was extracted from HepG2.2.15 cells using TRIzol reagent (Invitrogen, Carlsbad, CA, USA). m5C-RIP was performed as previously described [55]. Briefly, 300 μg of RNA were resuspend in immunoprecipitation (IP) buffer (150 mmol/L NaCl, 0.1% NP-40, 10 mmol/L Tris-HCl pH 7.4) and incubated with an anti-m5C antibody (ab10805, Abcam) or normal rabbit/mouse IgG antibody (Proteintech) overnight at 4 °C. The RNA and antibody mixture was then incubated with 35 μL of magnetic beads (New England Biolabs) for 2 h at 4 °C. Beads were washed with IP buffer six times and then incubated with 300 μL of elution buffer (5 mmol/L Tris-HCl pH 7.5, 1 mmol/L EDTA pH 8.0, 0.05% SDS, 4.2 μL 20 mg/mL proteinase K) at 50 °C for 1.5 h. Eluted RNA was purified using phenol/chloroform. Immunoprecipitated RNA was used for cDNA synthesis using the HiScript 1st Strand cDNA Synthesis Kit (Vazyme) according to the manufacturer’s protocol. The relative RNA level was measured by quantitative PCR (qPCR) using Hieff® qPCR SYBR® Green Master Mix (Yeasen Biotech Co., Shanghai, China) on a CFX Connect real-time system (Bio-Rad Laboratories, Hercules, CA, USA). The primers used for RT-qPCR were listed in Table S6. At least three samples in each qPCR analysis were prepared, and three independent experiments were performed.
Formaldehyde-crosslinked RNA-immunoprecipitation
Cells (1 × 107) seeded in 10-cm plates were cross-linked by 1% formaldehyde at 37 °C for 10 min. Next, 2.5 mol/L glycine was added to the plate at a final concentration of 0.125 mol/L to stop the cross-linking reaction. The cells were washed three times with phosphate-buffered saline (PBS) and scraped from the plate. After centrifugation at 800 ×g for 3 min at 4 °C, the cells were resuspended with 800 μL of RIP buffer (150 mmol/L KCl, 25 mmol/L Tris-HCl pH 7.4, 5 mmol/L EDTA, 0.5 mmol/L DTT, 0.5% NP40, 100 U/mL RNase inhibitor, 100 μmol/L PMSF, 1 μg/mL proteinase Inhibitors) and incubated on ice for 30 min. The cell lysates were centrifugated at 13000 ×g for 10 min; 100 μL of the supernatant were collected as an input control. The remaining lysates were divided into two aliquots and stored at −80 °C until further analysis.
The lysates were incubated with anti-Flag antibody (Sigma-Aldrich) or IgG antibody (Proteintech) overnight at 4 °C. Protein-G agarose beads were washed three times with wash buffer (300 mmol/L KCl, 25 mmol/L Tris-HCl pH 7.4, 5 mmol/L EDTA, 0.5 mmol/L DTT, 0.5% NP40, 100 U/mL RNase inhibitor, 100 µmol/L PMSF, 1 μg/mL proteinase inhibitors) and added to the cell lysate and antibody mixture, and incubated at 4 °C for 2 h. The beads were washed three times with RIP buffer and three times with wash buffer. RNA was extracted using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA) for qRT-PCR.
Enzyme-linked immunosorbent assay (ELISA)
Huh7 cells were transfected with the HBV 1.1-mer plasmid, pch9-3091, using Lipofectamine 2000 as described above. Cell culture supernatants were harvested 72 h post-transfection. HBsAg and HBeAg were detected using an enzyme-linked immunosorbent assay (ELISA) kit (Kehua Bio-Engineering, Shanghai, China) according to the manufacturer’s instructions. IFN-β was assayed at 24 h post-transfection with a human IFN-β/IFNB ELISA Kit (MM-1641H1, MEIMIAN) according to the manufacturer’s instructions.
Southern blotting
HBV DNA was extracted from huh7 cells 72 h post-transfection, as described previously [76]. In brief, 5 × 106 cells were lysed with 800 μL HBV DNA lysis buffer (50 mmol/L Tris-HCl pH7.4, 1 mmol/L EDTA, 1%NP-40) on ice for 10 min. The cells were scraped off and centrifugated at 14,000 rpm for 3 min. The supernatants were mixed with 8 μL DNaseI(10 mg/mL) and 8 μL 1 mol/L MgCl2, and incubated at 37 °C for 30 min. Then, 40 μL of 0.5 mol/L EDTA was added to stop the reaction. HBV capsids were digested at 55 °C for 2 h after adding 20 μL of 20 mg/mL proteinase K and 80 μL of 10% SDS to cell lysates. Core-associated DNA was extracted using a phenol/chloroform mixture. The extracted DNA was separated on a 1% agarose gel for 4 h at 50 V, followed by denaturation and transfer to Hybound N+ membranes (GE Healthcare) in 20× SSC (3 mol/L NaCl, 0.3 mol/L sodium citrate). The membrane was UV-crosslinked and hybridized with a DIG-labeled probe (DIG HIGH PRIME Kit II). Viral DNA was detected as previously described [77].
Nuclear and cytoplasmic fractionation
The cells (1 × 106) were harvested 24 h post-transfection. The cytoplasmic and nuclear fractions were extracted using the Cytoplasmic and Nuclear RNA Purification Kit (21000, NORGEN), respectively, according to the manufacturer’s instructions. Purified RNA was reverse-transcribed for qPCR, as described above. U6 and GAPDH serve as internal controls in nucleus and cytoplasm respectively.
Immuno-precipitation (IP)
IP was performed as previously described [48]. Briefly, cells were washed with cold PBS and then lysed with IP buffer (50 mmol/L Tris-HCl pH 7.5, 1 mmol/L EGTA, 1 mmol/L EDTA, 1% Triton X-100, 150 mmol/L NaCl, 2 mmol/L DTT, 100 μmol/L PMSF, 1 μg/mL proteinase inhibitors) on ice for 30 min. Next, 10% of the cell lysates were collected and used as input control after being centrifuged at 13,000 rpm and 4 °C for 10 min. The remaining lysates were incubated with primary or normal IgG antibodies overnight at 4 °C respectively. Protein-G agarose (catalog number) was added to the mixture and incubated for 1 h at 4, followed by six washes with IP buffer. Immunoprecipitated proteins were eluted in SDS loading buffer. All samples were heat-denatured before SDS-PAGE and western blotting was performed.
Dual luciferase reporter assay
Huh7 cells were seeded in a 12-well plate and transfected with luciferase vectors using Lipofectamine 2000 (Invitrogen) as described above. Dual-luciferase reporter assays (Promega #E1910) were performed according to the manufacturer’s instructions 24 h post-transfection. Briefly, 100 Μl Passive Lysis Buffer was added to each PBS-washed well. Then, 20 μL of cell lysates was transferred to an Optiplate-96 following a 15 min incubation at room temperature. Firefly and Rellina luciferase activities were assayed using a GloMax Luminometer (Promega). The relative luciferase activity of each sample was calculated as the ratio of firefly to Renilla luciferase activity.
HBV virus production and infection
HepG2.2.15 cells were cultured for more than three days. The culture supernatants were then harvested and centrifuged at 1000 × g and 4 °C for 15 min to remove cell debris. The clarified supernatants was mixed with 5% (w/v) PEG8000 and incubated at 4 °C overnight. The mixture was centrifuged at 4000 rpm and 4 °C for 30 min. The pellets were dissolved in culture medium containing 1% of the starting volume. HepG2-NTCP cells were seeded in 12-well plates prior to HBV infection. The cells were incubated with the collected virus in the presence of 4% PEG8000 and 2% DMSO at 37 °C for 24 h. The cells were washed twice and cultured in fresh medium containing 2% DMSO for more than seven days.
Ribosome loading
Huh7 cells were cultured and transfected as described above; 2 × 107 cells were treated with 5 mg/mL cycloheximide at 37 °C for 10 min 24 h post-transfection. The cells were then washed three times with PBS and resuspended in 1 mL of ribosome lysis buffer (10 mmol/L Tris-HCl pH 7.4, 5 mmol/L MgCl2, 100 mmol/L KCl, 1% Triton X, Protease inhibitor, 2 mmol/L DTT, 100 mg/mL cycloheximide, and RNase inhibitor). RNA was extracted from 10% of the cell lysates and used as the input control. The remaining lysates were loaded into a gradient sucrose solution ranging from 5% to 50% (5% increments) in an SW41 transparent ultracentrifuge tube. The sucrose solution was prepared and stored at 4 °C for 24 h to form a continuous gradient. Sucrose (1 mL) was collected by ultracentrifugation at 30,000 × g for 2 h at 4 °C. The OD 260/320 ratio of each fraction was measured using a NanoDrop One. RNA was extracted from the fraction with the highest OD 260/320 nm. The amount of HBV mRNA in this fraction was measured by qPCR.
RNA oligo pull down
huh7 cells (8 × 107) were harvested and lysed in 2 mL of RIPA buffer (50 mM Tris-HCl, pH 8.0, 0.5 mM EDTA, 0.1% SDS, 1% NP-40, 150 mM NaCl). The cell lysates were centrifuged at 13000 rpm for 15 min at 4 °C. Then, the supernatant was collected as INPUT, and 50 μL of neutravidin beads (29200, Thermo) were washed twice with Tris-buffered saline (TBS). Next, 5 μg of m5C modified oligo RNA (Beijing Tsingke Biotech Co., Ltd) or control oligo RNA were added to the beads, the volume was adjusted to 500 μL and rotated at 4 °C for 2 h. The beads were washed three times, and 100 μL of huh7 cell lysate was added to the beads and rotated at 4 °C overnight. The beads were washed again with cold TBS three times, and 40 μL 2x LDS Laemmli buffer was added. Pull-down proteins were analyzed by western blotting.
RNA stability assay
huh7 cells (1 × 104) were seeded into a 12-well plate. Cells were harvested at 0 h, 2 h, 4 h, 6 h, and 8 h following treatment with 4 μg/mL Actinomycin D (Sigma) at 24 h post-transfection. Total RNA was extracted from huh7 cells at different time points using TRIzol reagent. The same amount of RNA was reverse-transcribed for qPCR, as described above. The proportion of target gene expression relative to 0 h was calculated, and a one-step decay curve was drawn to calculate the half-life.
Double-stranded DNA oligo pull down
Five micrograms of sense and anti-sense biotin-labeled oligo DNA were incubated at 95 °C for 10 min, and cooled at room temperature. Next, 80 μL of NeutrAvidinTM Agarose (29200, thermo) was washed three times with wash buffer (20 mmol/L Tris-HCl pH 7.5, 100 mmol/L NaCl, 1 mmol/L MgCl2, 0.5 mmol/L EDTA, 0.5 mmol/L DTT). The beads were resuspended in 1 mL of wash buffer, annealed double-stranded oligonucleotide DNAs were added, and the beads were incubated for 2 h at room temperature. The beads were then washed three times with wash buffer and resuspended with 900 μL of pull down buffer (20 mmol/L Tris-HCl pH 7.5, 100 mmol/L NaCl, 1 mmol/L MgCl2, 0.5 mmol/L EDTA, 0.5 mmol/L DTT, 4% Glycerol, 10 µg/mL Poly dI-Dc,1× Roche’s protease inhibitor cocktail). Subsequently, 100 μL of cell lysates in RIPA buffer were added to the bead and oligo mixture and incubated overnight at 4 °C. The beads were washed with wash buffer six times and resuspended in 40 μL 2× LDS Laemmli buffer. Proteins were analyzed by western blotting.
Statistical analysis
Comparisons between two groups were performed using students’ unpaired t-tests, while comparisons of the decay curves were performed using two-way ANOVA. P ≤ 0.05 is considered statistically significant. All statistical analyses were conducted in GraphPad prism software.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Liu Y, Yang Y, Wu R, Gao CC, Liao X, Han X, et al. mRNA m(5)C inhibits adipogenesis and promotes myogenesis by respectively facilitating YBX2 and SMO mRNA export in ALYREF-m(5)C manner. Cell Mol Life Sci. 2022;79:481.
Liu Y, Zhao Y, Wu R, Chen Y, Chen W, Liu Y, et al. mRNA m5C controls adipogenesis by promoting CDKN1A mRNA export and translation. RNA Biol. 2021;18 sup2:711–21.
Xu T, He B, Sun H, Xiong M, Nie J, Wang S, et al. Novel insights into the interaction between N6-methyladenosine modification and circular RNA. Mol Ther Nucleic Acids. 2022;27:824–37.
Liu M, Guo G, Qian P, Mu J, Lu B, He X, et al. 5-methylcytosine modification by Plasmodium NSUN2 stabilizes mRNA and mediates the development of gametocytes. Proc Natl Acad Sci USA. 2022;119:e2110713119.
Jing C, Hu J, Amara U, Park SJ, Li Y, Jeong D, et al. FIONA1 is an N6-methyladenosine methyltransferase that plays a vital role in floral transition via affecting the splicing of FLC and the stability of SPL3 and SEP3 in Arabidopsis. J Exp Bot. 2022;74:864–77.
Shi B, Liu WW, Yang K, Jiang GM, Wang H. The role, mechanism, and application of RNA methyltransferase METTL14 in gastrointestinal cancer. Mol Cancer. 2022;21:163.
Navarro IC, Tuorto F, Jordan D, Legrand C, Price J, Braukmann F, et al. Translational adaptation to heat stress is mediated by RNA 5-methylcytosine in Caenorhabditis elegans. EMBO J. 2021;40:e105496.
Delaunay S, Pascual G, Feng B, Klann K, Behm M, Hotz-Wagenblatt A, et al. Mitochondrial RNA modifications shape metabolic plasticity in metastasis. Nature. 2022;607:593–603.
Shen H, Ontiveros RJ, Owens MC, Liu MY, Ghanty U, Kohli RM, et al. TET-mediated 5-methylcytosine oxidation in tRNA promotes translation. J Biol Chem. 2021;296:100087.
Nachtergaele S, He C. Chemical modifications in the life of an mRNA transcript. Annu Rev Genet. 2018;52:349–72.
Zhao BS, Roundtree IA, He C. Post-transcriptional gene regulation by mRNA modifications. Nat Rev Mol Cell Biol. 2017;18:31–42.
Gilbert WV, Bell TA, Schaening C. Messenger RNA modifications: form, distribution, and function. Science. 2016;352:1408–12.
Chen YS, Yang WL, Zhao YL, Yang YG. Dynamic transcriptomic m(5) C and its regulatory role in RNA processing. Wiley Interdiscip Rev RNA. 2021;12:e1639.
Yang X, Yang Y, Sun BF, Chen YS, Xu JW, Lai WY, et al. 5-methylcytosine promotes mRNA export - NSUN2 as the methyltransferase and ALYREF as an m(5)C reader. Cell Res. 2017;27:606–25.
Shi H, Chai P, Jia R, Fan X. Novel insight into the regulatory roles of diverse RNA modifications: Re-defining the bridge between transcription and translation. Mol Cancer. 2020;19:78.
Guo G, Pan K, Fang S, Ye L, Tong X, Wang Z, et al. Advances in mRNA 5-methylcytosine modifications: Detection, effectors, biological functions, and clinical relevance. Mol Ther Nucleic Acids. 2021;26:575–93.
Cui X, Liang Z, Shen L, Zhang Q, Bao S, Geng Y, et al. 5-Methylcytosine RNA methylation in Arabidopsis thaliana. Mol Plant. 2017;10:1387–99.
David R, Burgess A, Parker B, Li J, Pulsford K, Sibbritt T, et al. Transcriptome-wide mapping of RNA 5-methylcytosine in Arabidopsis mRNAs and noncoding RNAs. Plant Cell. 2017;29:445–60.
Wei Z, Panneerdoss S, Timilsina S, Zhu J, Mohammad TA, Lu ZL, et al. Topological characterization of human and mouse m(5)C epitranscriptome revealed by bisulfite sequencing. Int J Genomics. 2018;2018:1351964.
Tang Y, Gao CC, Gao Y, Yang Y, Shi B, Yu JL, et al. OsNSUN2-mediated 5-methylcytosine mRNA modification enhances rice adaptation to high temperature. Dev Cell. 2020;53:272–86.e7.
Amort T, Rieder D, Wille A, Khokhlova-Cubberley D, Riml C, Trixl L, et al. Distinct 5-methylcytosine profiles in poly(A) RNA from mouse embryonic stem cells and brain. Genome Biol. 2017;18:1.
Bohnsack KE, Hobartner C, Bohnsack MT. Eukaryotic 5-methylcytosine (m(5)C) RNA methyltransferases: mechanisms, cellular functions, and links to disease. Genes. 2019;10:102.
Tsukuda S, Watashi K. Hepatitis B virus biology and life cycle. Antiviral Res. 2020;182:104925.
Decorsiere A, Mueller H, van Breugel PC, Abdul F, Gerossier L, Beran RK, et al. Hepatitis B virus X protein identifies the Smc5/6 complex as a host restriction factor. Nature. 2016;531:386–9.
Kumar M, Jung SY, Hodgson AJ, Madden CR, Qin J, Slagle BL. Hepatitis B virus regulatory HBx protein binds to adaptor protein IPS-1 and inhibits the activation of beta interferon. J Virol. 2011;85:987–95.
Wei C, Ni C, Song T, Liu Y, Yang X, Zheng Z, et al. The hepatitis B virus X protein disrupts innate immunity by downregulating mitochondrial antiviral signaling protein. J Immunol. 2010;185:1158–68.
Tsunematsu S, Suda G, Yamasaki K, Kimura M, Izumi T, Umemura M, et al. Hepatitis B virus X protein impairs alpha-interferon signaling via up-regulation of suppressor of cytokine signaling 3 and protein phosphatase 2A. J Med Virol. 2017;89:267–75.
Courtney DG, Tsai K, Bogerd HP, Kennedy EM, Law BA, Emery A, et al. Epitranscriptomic addition of m(5)C to HIV-1 transcripts regulates viral gene expression. Cell Host Microbe. 2019;26:217–27.e6.
Henry BA, Kanarek JP, Kotter A, Helm M, Lee N. 5-methylcytosine modification of an Epstein-Barr virus noncoding RNA decreases its stability. RNA. 2020;26:1038–48.
Eckwahl M, Xu R, Michalkiewicz J, Zhang W, Patel P, Cai Z, et al. 5-Methylcytosine RNA modifications promote retrovirus replication in an ALYREF reader protein-dependent manner. J Virol. 2020;94:e00544–20.
Kariko K, Buckstein M, Ni H, Weissman D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity. 2005;23:165–75.
Li N, Hui H, Bray B, Gonzalez GM, Zeller M, Anderson KG, et al. METTL3 regulates viral m6A RNA modification and host cell innate immune responses during SARS-CoV-2 infection. Cell Rep. 2021;35:109091.
Lu M, Zhang Z, Xue M, Zhao BS, Harder O, Li A, et al. N(6)-methyladenosine modification enables viral RNA to escape recognition by RNA sensor RIG-I. Nat Microbiol. 2020;5:584–98.
Kim GW, Imam H, Khan M, Siddiqui AN. (6)-Methyladenosine modification of hepatitis B and C viral RNAs attenuates host innate immunity via RIG-I signaling. J Biol Chem. 2020;295:13123–33.
Thomsen MK, Nandakumar R, Stadler D, Malo A, Valls RM, Wang F, et al. Lack of immunological DNA sensing in hepatocytes facilitates hepatitis B virus infection. Hepatology. 2016;64:746–59.
Sato S, Li K, Kameyama T, Hayashi T, Ishida Y, Murakami S, et al. The RNA sensor RIG-I dually functions as an innate sensor and direct antiviral factor for hepatitis B virus. Immunity. 2015;42:123–32.
Durbin AF, Wang C, Marcotrigiano J, Gehrke L. RNAs containing modified nucleotides fail to trigger RIG-I conformational changes for innate immune signaling. mBio. 2016;7:e00833–16.
Teng Y, Xu Z, Zhao K, Zhong Y, Wang J, Zhao L, et al. Novel function of SART1 in HNF4alpha transcriptional regulation contributes to its antiviral role during HBV infection. J Hepatol. 2021;75:1072–82.
Zhao K, Guo F, Wang J, Zhong Y, Yi J, Teng Y, et al. Limited disassembly of cytoplasmic hepatitis B virus nucleocapsids restricts viral infection in murine hepatic cells. Hepatology. 2023;77:1366–81.
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 2009;37:W202–8.
Saito I, Oya Y, Shimojo H. Novel RNA family structure of hepatitis B virus expressed in human cells, using a helper-free adenovirus vector. J Virol. 1986;58:554–60.
Jiang F, Ramanathan A, Miller MT, Tang GQ, Gale M Jr, Patel SS, et al. Structural basis of RNA recognition and activation by innate immune receptor RIG-I. Nature. 2011;479:423–7.
Nombela P, Miguel-Lopez B, Blanco S. The role of m(6)A, m(5)C and Psi RNA modifications in cancer: novel therapeutic opportunities. Mol Cancer. 2021;20:18.
Boo SH, Kim YK. The emerging role of RNA modifications in the regulation of mRNA stability. Exp Mol Med. 2020;52:400–8.
Nissim O, Melis M, Diaz G, Kleiner DE, Tice A, Fantola G, et al. Liver regeneration signature in hepatitis B virus (HBV)-associated acute liver failure identified by gene expression profiling. PLoS ONE. 2012;7:e49611.
Levrero M, Zucman-Rossi J. Mechanisms of HBV-induced hepatocellular carcinoma. J Hepatol. 2016;64 1 Suppl:S84–S101.
Yoo YD, Ueda H, Park K, Flanders KC, Lee YI, Jay G, et al. Regulation of transforming growth factor-beta 1 expression by the hepatitis B virus (HBV) X transactivator. Role in HBV pathogenesis. J Clin Investig. 1996;97:388–95.
Zhang X, Hao H, Ma L, Zhang Y, Hu X, Chen Z, et al. Methyltransferase-like 3 modulates severe acute respiratory syndrome coronavirus-2 RNA N6-methyladenosine modification and replication. mBio. 2021;12:e0106721.
Gokhale NS, McIntyre ABR, McFadden MJ, Roder AE, Kennedy EM, Gandara JA, et al. N6-Methyladenosine in flaviviridae viral RNA genomes regulates infection. Cell Host Microbe. 2016;20:654–65.
Imam H, Khan M, Gokhale NS, McIntyre ABR, Kim GW, Jang JY, et al. N6-methyladenosine modification of hepatitis B virus RNA differentially regulates the viral life cycle. Proc Natl Acad Sci USA. 2018;115:8829–34.
Hao H, Liu W, Miao Y, Ma L, Yu B, Liu L, et al. N4-acetylcytidine regulates the replication and pathogenicity of enterovirus 71. Nucleic Acids Res. 2022;50:9339–54.
Ortega-Prieto AM, Skelton JK, Wai SN, Large E, Lussignol M, Vizcay-Barrena G, et al. 3D microfluidic liver cultures as a physiological preclinical tool for hepatitis B virus infection. Nat Commun. 2018;9:682.
Eckwahl M, Xu R, Michalkiewicz J, Zhang W, Patel P, Cai Z, et al. 5-Methylcytosine RNA modifications promote retrovirus replication in an ALYREF reader protein-dependent manner. J Virol. 2020;94:e00544-20.
Rubio RM, Depledge DP, Bianco C, Thompson L, Mohr I. RNA m(6) A modification enzymes shape innate responses to DNA by regulating interferon beta. Genes Dev. 2018;32:1472–84.
Hao H, Hao S, Chen H, Chen Z, Zhang Y, Wang J, et al. N6-methyladenosine modification and METTL3 modulate enterovirus 71 replication. Nucleic Acids Res. 2019;47:362–74.
Sivasudhan E, Blake N, Lu Z, Meng J, Rong R. Hepatitis B viral protein HBx and the molecular mechanisms modulating the hallmarks of hepatocellular carcinoma: a comprehensive review. Cells. 2022;11:741.
Song H, Xu F, Xiao Q, Tan G. Hepatitis B virus X protein and its host partners. Cell Mol Immunol. 2021;18:1345–6.
Wang XW, Forrester K, Yeh H, Feitelson MA, Gu JR, Harris CC. Hepatitis B virus X protein inhibits p53 sequence-specific DNA binding, transcriptional activity, and association with transcription factor ERCC3. Proc Natl Acad Sci USA. 1994;91:2230–4.
Haviv I, Shamay M, Doitsh G, Shaul Y. Hepatitis B virus pX targets TFIIB in transcription coactivation. Mol Cell Biol. 1998;18:1562–9.
Yoo YG, Lee MO. Hepatitis B virus X protein induces expression of Fas ligand gene through enhancing transcriptional activity of early growth response factor. J Biol Chem. 2004;279:36242–9.
Cheng X, Xia Y, Serti E, Block PD, Chung M, Chayama K, et al. Hepatitis B virus evades innate immunity of hepatocytes but activates cytokine production by macrophages. Hepatology. 2017;66:1779–93.
Wang H, Ryu WS. Hepatitis B virus polymerase blocks pattern recognition receptor signaling via interaction with DDX3: implications for immune evasion. PLoS Pathog. 2010;6:e1000986.
Wieland S, Thimme R, Purcell RH, Chisari FV. Genomic analysis of the host response to hepatitis B virus infection. Proc Natl Acad Sci USA. 2004;101:6669–74.
Wieland SF, Chisari FV. Stealth and cunning: hepatitis B and hepatitis C viruses. J Virol. 2005;79:9369–80.
Brunetto MR, Bonino F. Interferon therapy of chronic hepatitis B. Intervirology. 2014;57:163–70.
Zhang Y, Zhang LS, Dai Q, Chen P, Lu M, Kairis EL, et al. 5-methylcytosine (m(5)C) RNA modification controls the innate immune response to virus infection by regulating type I interferons. Proc Natl Acad Sci USA. 2022;119:e2123338119.
Hussain S, Sajini AA, Blanco S, Dietmann S, Lombard P, Sugimoto Y, et al. NSun2-mediated cytosine-5 methylation of vault noncoding RNA determines its processing into regulatory small RNAs. Cell Rep. 2013;4:255–61.
Brzezicha B, Schmidt M, Makalowska I, Jarmolowski A, Pienkowska J, Szweykowska-Kulinska Z. Identification of human tRNA:m5C methyltransferase catalysing intron-dependent m5C formation in the first position of the anticodon of the pre-tRNA Leu (CAA). Nucleic Acids Res. 2006;34:6034–43.
Du Y, Broering R, Li X, Zhang X, Liu J, Yang D, et al. In vivo mouse models for hepatitis B virus infection and their application. Front Immunol. 2021;12:766534.
Yi J, Lei X, Guo F, Chen Q, Chen X, Zhao K, et al. Co-delivery of Cas9 mRNA and guide RNAs edits hepatitis B virus episomal and integration DNA in mouse and tree shrew models. Antiviral Res. 2023;215:105618.
Xu Z, Zhao L, Zhong Y, Zhu C, Zhao K, Teng Y, et al. A novel mouse model harboring Hepatitis B Virus Covalently Closed Circular DNA. Cell Mol Gastroenterol Hepatol. 2022;13:1001–17.
Chen S, Zhou Y, Chen Y, Gu J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics. 2018;34:i884–90.
Krueger F, Andrews SR. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics. 2011;27:1571–2.
Pedersen BS, Quinlan AR. Mosdepth: quick coverage calculation for genomes and exomes. Bioinformatics. 2018;34:867–8.
Juhling F, Kretzmer H, Bernhart SH, Otto C, Stadler PF, Hoffmann S. metilene: fast and sensitive calling of differentially methylated regions from bisulfite sequencing data. Genome Res. 2016;26:256–62.
Lin Y, Wu C, Wang X, Liu S, Zhao K, Kemper T, et al. Glucosamine promotes hepatitis B virus replication through its dual effects in suppressing autophagic degradation and inhibiting MTORC1 signaling. Autophagy. 2020;16:548–61.
Hao S, Zhang J, Chen Z, Xu H, Wang H, Guan W. Alternative polyadenylation of human bocavirus at Its 3’ end is regulated by multiple elements and affects capsid expression. J Virol. 2017;91:e02026–16.
Acknowledgements
This study was supported by the Strategic Priority Research Program of the Chinese Academy of Sciences [XDB0490000], National Natural Science Foundation of China [31970168], Key R&D Program of Hubei Province [2021BCD004], Hubei Science and Technology Major Project [2021ACB004], Wuhan Knowledge Innovation Special Project [2023020201020303] and Emergency Key Project of Guangzhou Laboratory [EKPG21-30-2]. Total RNA isolated from HBV AAV-HBV transduced mouse liver was kindly provided by Dr. Yuchen Xia (TaiKang Medical School, Wuhan University).We thank the Core Facility and Technical Support in the Wuhan Institute of Virology (WIV), Chinese Academy of Sciences (CAS), especially Lei Zhang and Ding Gao, for their assistant with ultracentrifugation and confocal microscopy.
Author information
Authors and Affiliations
Contributions
SD, HL, FZ, and WG designed research; SD, ZC, MZ, Lishi Liu, XZ, HH and Z performed research; SD, HL, Lijuan Liu, LM, FH, FZ, and WG analyzed data; and SD, HL, FZ, and WG wrote the paper. All authors commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Mauro Piacentini
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ding, S., Liu, H., Liu, L. et al. Epigenetic addition of m5C to HBV transcripts promotes viral replication and evasion of innate antiviral responses. Cell Death Dis 15, 39 (2024). https://doi.org/10.1038/s41419-023-06412-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-023-06412-9
- Springer Nature Limited