Abstract
Acute myeloid leukemia (AML) is a clinically and genetically heterogeneous clonal disease associated with unmet medical needs. Paralleling the pathology of other cancers, AML tumorigenesis and propagation can be ascribed to dysregulated cellular processes, including apoptosis. This function and others are regulated by tumor suppressor P53, which plays a pivotal role in leukemogenesis. Opposing P53-mediated activities is the mouse double minute 2 homolog (MDM2), which promotes P53 degradation. Because the TP53 mutation rate is low, and MDM2 frequently overexpressed, in patients with leukemia, targeting the MDM2-P53 axis to restore P53 function has emerged as an attractive AML treatment strategy. APG-115 is a potent MDM2 inhibitor under clinical development for patients with solid tumors. In cellular cultures and animal models of AML, we demonstrate that APG-115 exerted substantial antileukemic activity, as either a single agent or when combined with standard-of-care (SOC) hypomethylating agents azacitidine (AZA) and decitabine (DAC), or the DNA-damaging agent cytarabine (Ara-C). By activating the P53/P21 pathway, APG-115 exhibited potent antiproliferative and apoptogenic activities, and induced cell cycle arrest, in TP53 wild-type AML lines. In vivo, APG-115 significantly reduced tumor burden and prolonged survival. Combinations of APG-115 with SOC treatments elicited synergistic antileukemic activity. To explain these effects, we propose that APG-115 and SOC agents augment AML cell killing by complementarily activating the P53/P21 pathway and upregulating DNA damage. These findings and the emerging mechanism of action afford a sound scientific rationale to evaluate APG-115 (with or without SOC therapies) in patients with AML.
Similar content being viewed by others
Introduction
Acute myeloid leukemia (AML) is a genetically and clinically heterogeneous clonal disease characterized by the rapid proliferation of immature myeloid progenitors, with suppression of hematopoiesis1,2. Allogeneic hematopoietic stem cell transplantation (allo-HCT) remains the most promising modality to effect cure. However, allo-HCT can be offered only to a minority of younger, fit patients, whereas AML primarily affects individuals older than 60 who are hence not suitable candidates3. Patients who have AML and are ineligible for allo-HCT and/or harbor high-risk karyotypes typically have poor outcomes, with an estimated median overall survival of only 1 year4.
For elderly patients with AML, the DNA hypomethylating agents azacytidine (AZA; 5-azacytidine) and decitabine (DAC; 5-aza-2′-deoxycytidine) have become standards of care5,6,7,8; however, rates of CR, together with CR and incomplete cell-count recovery in patients with newly diagnosed or advanced AML remains low, typically range from only 20 to 30%9.
The regimen of choice for induction in many patients with newly diagnosed AML is the DNA-damaging agent cytarabine (Ara-C), administered daily for 7 days, followed by daunorubicin for 3 days10,11. This so-called 7 + 3 regimen has been associated with high drug resistance and a low 5-year OS rate12,13. Even with recently approved molecularly targeted agents, the 5-year survival rate in patients younger than 65 with AML is only about 45%14. In addition, the number of newly diagnosed AML cases continues to rise, at an annual rate of 2% from 2007 to 201615. There is hence an urgent need to identify novel agents that enhance clinical outcomes in AML.
Evasion of apoptosis is a pathological hallmark of AML and other cancers3. Apoptosis and multiple other cellular functions are mediated by the tumor suppressor P53, which plays a pivotal role in leukemogenesis16. P53 is precisely controlled by the negative regulator mouse double minute 2 homolog (MDM2), which blocks transcriptional activity of P53 and promotes its proteasomal degradation. In various cancers, MDM2 overexpression is a prominent mechanism to impair P53 function. Studies of AML suggest that the TP53 mutation rate remains as low as 10%17,18,19,20, while MDM2 is frequently overexpressed21,22,23, rendering the P53-MDM2 axis an attractive pharmacologic target for small-molecule inhibitors.
In addition to monotherapy, combinations of MDM2 inhibitors with SOC agents could improve antileukemic efficacy24. A phase I/Ib clinical trial demonstrated that MDM2 inhibitor idasanutlin (RG-7388) combined with Ara-C conferred durable clinical efficacy, including CR in patients with r/r AML25. However, mechanisms underlying synergy between these therapies and SOC agents (e.g., DAC, AZA, and Ara-C) have yet to be elucidated.
We have developed a novel, potent MDM2 inhibitor (APG-115), which is being evaluated in patients with solid tumors (NCT02935907, NCT03611868). However, the effects of this small-molecule agent when administered with SOC and other treatments for AML are unknown. To address this knowledge gap, we established the following objectives for this study: (1) to evaluate the antileukemic activity of APG-115, administered alone or combined with AZA, DAC, or Ara-C, in preclinical models of AML; and (2) to assess synergy of APG-115 combined with these SOC therapies, and, if present, formulate underlying molecular mechanisms.
Results
APG-115 potently inhibits cellular proliferation and induces apoptosis and cell cycle arrest in TP53 wild-type AML cell lines
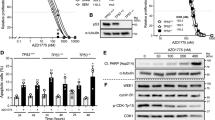
The in vitro antiproliferative activity of APG-115 was first evaluated in a panel of five distinct AML cultures, including three TP53 wild-types (TP53wt; MOLM-13, MV-4-11, OCI-AML-3), a TP53 null (TP53null; HL-60), and a TP53 mutant (TP53mut; SKM-1) cell lines (Fig. 1A). HL-60 is a cell line with documented TP53 gene deletion in both alleles, while SKM-1 is a cell line carrying a missense mutation (p.R248Q) in the TP53 gene. All three TP53wt cell lines were more sensitive (vs. TP53mut lines) to APG-115 treatment. Half-maximal inhibitory concentration (IC50) values of APG-115 were at least one order of magnitude lower (i.e., more active) in decreasing cellular proliferation in TP53wt, compared to TP53null and TP53mut cell lines. Mean (SD) values of IC50 for APG-115 were 26.8 (4.9) nM in MOLM-13, 165.9 (42.4) nM in MV-4-11, and 315.6 (97) nM in OCI-AML-3 cells (Supplementary Table 1). Compared to these TP53wt cells, HL-60 and SKM-1 cells with defected TP53 were resistant to APG-115 treatment, with mean (SD) IC50 values of 2558.3 (581.5) nM in HL-60 cells and 8,947.3 (569.6) nM in SKM-1 cells (Supplementary Table 1). APG-115 was also more potent than MDM2 inhibitor RG-7388 in both TP53wt (MOLM-13 and MV-4-11) and TP53mut (SKM-1) AML cell lines (Supplementary Table 1).
A Mean (SD) cellular viability of MOLM-13 (TP53wt), MV-4-11 (TP53wt), OCI-AML-3 (TP53wt), HL-60 (TP53null), and SKM-1 (TP53mut) cells treated with increasing concentrations of APG-115 for 72 h (n = 3 replicates). B Total protein levels of MDM2, P53, P21, and the loading control β-actin by Western blot in MOLM-13 cells exposed to APG-115 at increasing concentrations for 4 h. C Representative flow cytometry results from two independent experiments (left panel) and mean (SD) percentage of MOLM-13 cells in apoptosis (right panel) presented as n = 4 technical replicates. *p < 0.0001; ns not significant; compared to dimethyl sulfoxide vehicle (DMSO) via two-tailed Student t tests for data from three independently repeated experiments. D Increased cell cycle arrest in the G0/G1 phase, as indicated by flow cytometry in AML cells exposed to increasing concentrations of APG-115 for 48 h. Data shown are representative images of two independent experiments.
In TP53wt MOLM-13 cells, APG-115 potently and dose-dependently induced rapid accumulation of MDM2 as well as P53 and its downstream protein P21 (Fig. 1B), with minimally effective concentrations of approximately 0.01–0.04 μM. APG-115 also induced dose-dependent cell apoptosis in MOLM-13 cells (Fig. 1C). Treatment with APG-115 for 48 h led to apoptosis in approximately 31% of cells at APG-115 0.04 μM, 55.1% at 0.11 μM, 80.2% at 0.33 μM, and 96.8% at 1 μM (each p < 0.0001 vs. DMSO vehicle). Moreover, APG-115 treatment of the three TP53wt AML cell lines reduced percentages of cells in the S phase, indicating cell cycle arrest in the G0/G1 phase (Fig. 1D). In the aggregate, these findings demonstrate that APG-115 effectively: (1) activated the P53/P21 pathway, (2) inhibited cellular proliferation, and (3) induced apoptosis and cell cycle arrest.
APG-115 significantly reduces the leukemic burden and prolongs survival in TP53 wild-type AML xenograft models
Figure 2 depicts the antileukemic activity of APG-115, including effects on survival in a systemic AML xenograft model derived from MOLM-13 cells, including the study schema (Fig. 2A). In brief, APG-115 treatment was initiated 3 days after cellular inoculation and, after 15 days of treatment, five or six mice were randomly selected from each group and euthanized to analyze human CD45+ cells and assess tumor burden. Compared to vehicle control, APG-115 treatment substantially depleted the proportion of human CD45+ leukemic cells, to <0.1% in both bone marrow (vs. 0.41‒16.7%), and spleen (vs. 0.39‒1.40%; each comparison p < 0.005; Fig. 2B, C). The remaining (no euthanized) mice were continually observed for survival rates after suspending the APG-115 dose. The median overall survival of mice in the control group was 18.5 days (range from 15 to 21 days), whereas APG-115 treatment significantly extended median survival time up to 37.0 days (range from 15 to 49 days (p < 0.0001, Fig. 2D).
A Study schema of the experimental paradigm to study tumor burden and animal survival in the systemic MOLM-13 AML model. For CD45 analysis, mice were treated with APG-115 for 15 days. For survival analysis, mice were treated with APG-115 for 21 days. B Representative flow cytometry results show tumor burden (i.e., engraftment of human CD45+ cells) in murine femoral bone marrow and spleen. Five or six mice treated as in A from each treatment group were euthanized and examined on Day 15. C Significantly lower proportions of human CD45+ cells in the bone marrow and spleen collected from mice exposed to APG-115 (treated as in A) compared to vehicle (**p < 0.01 by two-tailed Student tests). D Kaplan–Meier proportional-hazards plots show significantly increased percentage survival in mice receiving APG-115 per the dosing conditions shown in A (~50 days) compared to mice receiving vehicle. APG-115 prolonged median survival time by 18.5 days compared to vehicle control (37.0 vs. 18.5 days), p < 0.0001 by log-rank test; n = 9–10 mice. CTX cyclophosphamide, IP intraperitoneally, NOD SCID nonobese diabetic severe combined immunodeficient, PO orally, QOD every other day. E APG-115 confers significantly increased in vivo survival compared to RG-7388 in a systemic AML xenograft model derived from TP53 wild-type MOLM-13 cells. NOD SCID mice intravenously implanted with 1 × 107 MOLM-13 cells (n = 10/group) were treated 3 days after cell inoculation. APG-115 and RG-7388 were orally administered daily for 7 days. Kaplan–Meier curve depicts mouse survival. *p < 0.05 by log-rank test.
In another independent experiment conducted in the same model, treatment with oral APG-115 consistently exhibited potent, dose-dependent antitumor activities, prolonging median OS to 23 days (range from 22 to 29 days) at an alternate-day dosing schedule of 20 mg/kg for 21 days, 30.5 days (range from 25 to 33 days) at 50 mg/kg once daily for seven days, and 38.5 days (range from 36 to 47 days) at 100 mg/kg once daily for 7 days, compared to 22 days (range from 20 to 22 days) in controls (each p < 0.05; Supplementary Fig. S1). Compare to the alternate-day dosing schedule of APG-115 at a clinically relevant dose (i.e., 20 mg/kg) currently used in clinical trials in patients with solid tumors, a pulsed high-dose (i.e., 50 and 100 mg/kg) schedule successfully achieves more substantial effect in the preclinical AML models, indicating that identifying the optimal dose regimen for APG-115 in AML is necessary.
Under the same dosing conditions (PO, QD for 7 days), APG-115 significantly increased median survival compared to RG-7388 in mice bearing MOLM-13 xenografts: by 2 days at equivalent 50 mg/kg doses and by 6 days at 100 mg/kg (each p < 0.05 for APG-115 vs. RG-7388, Fig. 2E). These cell-derived-xenograft (CDX) findings extend the foregoing in vitro data, suggesting that APG-115 exerts potent in vivo antileukemic (survival-promoting) activity, which significantly exceeded that of MDM2 inhibitor RG-7388.
Combinations of APG-115 with AZA, DAC, or Ara-C exhibit synergistic antiproliferative and apoptogenic activity in TP53 wt AML cell lines
Antiproliferative activity against TP53wt MOLM-13 AML cell line was enhanced when APG-115 was combined with AZA, DAC, or Ara-C, as depicted in Fig. 3A. Similar effects were also observed when APG-115 was combined with SOC treatments in MV-4-11 and OCI-AML-3 cell lines (Supplementary Fig. 2).
A Drug dose matrix of human MOLM-13 cells treated with increasing concentrations of APG-115, AZA, DAC, Ara-C or their combinations for 72 h. Cell viability was assessed using the CellTiter-Glo® assay. Representative results from three independent experiments are presented. The drug dose matrix indicates the percentage of cell viability of treated cells relative to vehicle controls. B Representative flow cytometry results from three independent experiments involving MOLM-13 cells treated with indicated concentrations of APG-115, DAC, AZA, and Ara-C (alone or in combination) for 48 h show enhanced apoptogenic effects. C Mean (SD) proportions of apoptotic cells (n = 3 replicates) show significant increases in cell lines exposed to APG-115 combined with SOC agents (vs. either alone). *p < 0.05, **p < 0.01, ***p < 0.001 by one-way ANOVA tests for data from three independently repeated experiments. DMSO, dimethylsulfoxide (vehicle); PI, propidium iodide.
Combination treatments including APG-115 further enhanced apoptogenic activity. Compared with single agents, the combination of APG-115 with AZA, DAC, or Ara-C led to significant increases in MOLM-13 cell apoptosis (Fig. 3B, C). On the other hand, only single-agent APG-115 consistently induced cell cycle arrest at the G0/G1 phase; concurrent treatment with APG-115 and concomitant SOC agents did not further enhance this effect on cell cycle arrest (Supplementary Fig. 3).
Collectively, these data suggest that APG-115, combined with the SOC agents AZA, DAC, and Ara-C, synergistically inhibits cellular proliferation and induces apoptosis in TP53wt AML cells. Given that cell cycle arrest was enhanced by single-agent APG-115, but not incrementally when combined with SOC agents, the antileukemic effects of APG-115 combination treatments are evidently primarily mediated by APG-115 induced inhibition of AML cellular growth and upregulated apoptosis.
The combination of APG-115 with AZA or DAC exerts superior antileukemic activity in TP53 wild-type AML xenograft models
When combined with hypomethylating agents, APG-115 exhibited superior in vivo antileukemic activity and augmented life expectancy in AML xenograft models. As shown in Fig. 4A, the median survival time of animals in the vehicle control group was 21 days (range from 17 to 24 days). Either APG-115 or AZA monotherapy significantly prolonged survival in mice bearing systemic MOLM-13 AML xenografts, to median values of 30 days (APG-115, rang from 23 to 33 days) and 29 days (AZA, range from 24 to 48 days, each p < 0.05 vs. vehicle control). Compared to either single agents or vehicle control, the combination of APG-115 with AZA significantly extended survival to a median of 41 days (range from 32 to 48 days, p < 0.05 vs. single-agent AZA, APG-115, or vehicle).
Three days after cell implantation, mice were treated with APG-115 (50 mg/kg, PO, QD), AZA (2 mg/kg, IV, QD), the combination, or vehicle for 7 days. A Kaplan–Meier curves show that APG-115 combined with AZA conferred significantly prolonged survival compared to any single agent in a systemic MOLM-13 AML xenograft model of NOD SCID mice (n = 10 per group): 41 days with APG-115-AZA compared to 30 days with APG-115, 29 days with AZA, and 21 days with the vehicle (*p < 0.05 for each comparison of APG-115‒Aza with single agents or vehicle by log-rank test). B APG-115 potentiates the growth-inhibitory effects of AZA and DAC in NOD SCID mice bearing subcutaneous OCI-AML-3 xenograft tumors. Mice were treated with APG-115 (50 mg/kg, PO, QOD for 15 days), AZA (2 mg/kg, IV, QD for 7 days), DAC (1 mg/kg, IV, QD for 7 days), or combinations as indicated (n = 5 per group). Tumor growth inhibition (TGI) was calculated at the endpoint.
In a subcutaneous OCI-AML-3 AML xenograft model, APG-115 with SOC agents potentiated tumor growth inhibition (TGI) compared to single agents: 55.1% on APG-115 with AZA, 43.4% with AZA, and 15.2% on APG-115 (each vs. vehicle; Fig. 4B). Comparable TGI values with DAC were 56.7% (APG-115 + DAC), 35.1% (DAC), and 15.2% (APG-115). These findings support the conclusion that APG-115 cooperatively augments antileukemic activity when combined with SOC therapies.
The combination of APG-115 with either AZA or Ara-C cooperatively activates the P53 pathway and downregulates genes involved in cell cycle progression and mismatch repair in TP53 wild-type AML cells
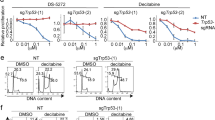
Compared to single agents, exposure of MOLM-13 cells to APG-115 combined with AZA or Ara-C resulted in more differentially expressed genes, implicating more marked effects of the combined treatments (Fig. 5A, B). Venn analysis of the intersection and union of differentiated expression genes between single agent and combination treatments are shown in Supplementary Fig. 4. The results showed that the largest number of differentially upregulated and downregulated genes appeared to be induced by the combined treatment with APG-115 and AZA or Ara-C, compared to either single-agent treatments. Interestingly, there are some but not many overlapped targets between the single drug vs. combinations.
MOLM-13 cells treated with APG-115 (40 nM), AZA (3 μM), Ara-C (100 nM), alone or in combination for 24 h in triplicates. Cells were collected for RNAseq analysis after treatment. A, B Numbers of differentially expressed genes after each treatment. C, D The most significantly enriched biological processes upregulated or downregulated in response to combined treatments: both AZA and Ara-C together with APG-115 upregulated P53, while the APG-115-AZA combination more markedly downregulated AML cell cycle, DNA replication, and mismatch repair. E, F Representative differentially expressed genes in response to APG-115, AZA, or Ara-C, alone or in combination. Combinations of APG-115 with either agent upregulated genes encoding the P53 pathway and downregulated those related to the cell cycle pathway, but only the APG-115-AZA combination markedly downregulated genes implicated in DNA repair. Expression levels are presented as normalized log2 counts per million. BBC3 Bcl-2-binding component 3 gene, CCNB1 Cdc2-cyclin B1 gene, DMSO dimethylsulfoxide vehicle, CDC20 cell division cycle 20 gene, CDKN1A cyclin-dependent kinase inhibitor 1A gene, GADD45 growth arrest and DNA damage-inducible 45, GDF15 growth differentiation factor 15 gene, KEGG Kyoto Encyclopedia of Genes and Genomes, MCM2/4 minichromosome maintenance proteins 2/4 gene, PLK-1 serine/threonine-protein kinase (also termed Polo-like kinase 1) gene, SESN1/2, sestrin 1 and 2 genes.
According to total RNA sequencing (RNASeq) analysis, exposure of MOLM-13 cells to combinations of APG-115 with AZA or Ara-C caused more marked activation of the P53 pathway and downregulated cell cycle progression and mismatch repair. Our analysis of the Kyoto Encyclopedia of Genes and Genomes (KEGG) demonstrated that the combination of APG-115 with AZA upregulated the expression of genes encoding constituents of the P53 signaling pathway and downregulated those involved in DNA replication, mismatch repair, and cell cycle progression (Fig. 5C). The combination of APG-115 with Ara-C mainly upregulated the expression of genes involved in the P53 pathway while downregulating those involved in cell cycle progression (Fig. 5D).
Upregulated P53-target genes included growth differentiation factor 15 (GDF15), which is a surrogate for P53 activation; and apoptogenic gene BBC3, which encodes P53 upregulated modulator of apoptosis [PUMA]), MDM2, BAX, and FAS. APG-115 combination treatments also upregulated expression of negative regulators of cell cycle progression genes, including cyclin-dependent kinase inhibitor 1A (CDKN1A), which encodes P21; and growth arrest and DNA damage-inducible 45 (GADD45), which plays a role in cell cycle control, DNA repair, and senescence (Fig. 5E, F). Both APG-115 combination treatments upregulated expression of sestrin genes 1 and 2 (SESN1 and SESN2), which are negative regulators of the mammalian target of rapamycin complex-1 (mTORC-1); this complex is, in turn, a sentinel of energy and redox status that regulates protein synthesis. Conversely, APG-115 combinations also significantly downregulated genes that encode oncoproteins involved in cell cycle division and progression, such as CDC20, CCNB1, and serine/threonine-protein kinase PLK1. Notably, treatment with APG-115 in tandem with Ara-C resulted in more substantial increases in P53-regulated genes compared to APG-115 with AZA, whereas only APG-115-AZA markedly reduced DNA replication by downregulating the expression of minichromosome maintenance protein 2/4 (MCM2, MCM4) genes. Taken together, these findings suggest that each combination of APG-115 with SOC AML treatments exerted antileukemic activity via largely overlapping, but also somewhat distinct, molecular pathways.
Combinations of APG-115 with DAC, AZA, or Ara-C coordinately induce DNA damage and upregulate expression of P53/P21 in TP53 wild-type AML cells
Further Western blot analyses revealed that combined APG-115 treatments significantly upregulated γ-H2AX in MOLM-13 cells. This histone H2A variant biomarker for DNA damage is indicative of DNA double-strand breaks, which can markedly attenuate genomic stability and compromise cell survival (Fig. 6A–C).
A, B MOLM-13 cells were treated with DAC (100 nM) or AZA (0.33 µM) for 24 h and then refreshed with DAC or AZA combined with APG-115 (40 nM) for an additional 24 h. Expression levels of γ-H2AX, a biomarker for DNA double-strand breaks, were assessed by Western blot. C Expression levels of γ-H2AX in MOLM-13 cells treated with Ara-C (100 nM) alone or in combination with APG-115 (40 nM) for 48 h. D–F Protein expression in MOLM-13 cells treated with APG-115 (40 nM), AZA (3 µM), Ara-C (100 nM), DAC (100 nM), or RG-7388 (40 nM), or their combinations as indicated for 48 h (D, F) or 6 h (E). β-actin was included as a loading control. Representative results from three independent experiments. G Proposed mechanism of action of APG-115 in combination with AZA, DAC, or Ara-C in AML cells. Cl-PARP-1, cleaved poly (ADP‒ribose) polymerase-1; DNMT1, DNA methyltransferase inhibitor; HDAC, histone deacetylase; γ-H2AX, H2A histone family member X; UHRF1, ubiquitin-like with PHD and RING finger domains 1.
APG-115 treatment also consistently stabilized P53 in MOLM-13 AML cells, leading to accumulation of P53 and P21 proteins (Fig. 6D–F). Perhaps most importantly, P53 and P21 proteins were further upregulated when APG-115 was combined with DAC, AZA, or Ara-C (vs. single SOC agents). Combined with DAC, AZA, or Ara-C, APG-115 also synergistically augmented output of a cleaved 89-kD catalytic fragment of poly (ADP-ribose-1; PARP-1), which is a prominent apoptotic signature. In combination treatments, APG-115 was also more potent than MDM2 inhibitor RG-7388 at the same concentrations. As anticipated, single-agent treatment with AZA or DAC substantially downregulated their key molecular target, DNMT1 protein. On the other hand, Ara-C significantly upregulated the expression of P21 while downregulating the expression of DNMT1.
A putative mechanism underlying the antileukemic activities of APG-115 combination treatments is depicted in Fig. 6G. In brief, APG-115 binds to MDM2 protein and releases its inhibitory effect on P53, activating the P53/P21 pathway by restoring the tumor-suppressor functions of P53. DAC, AZA, or Ara-C cause DNA damage, as evidenced by increased output of γ-H2AX, which further activates P53 and P21 signaling pathways. AZA and DAC also prevent inhibition of the DNMT1–UHRF1–HDAC1 complex on P2126,27 and hence contribute to the induction of P21 protein expression. Multidomain ubiquitin-like with PHD and RING finger domains 1 (UHRF1) leads to silencing of gene expression by bridging DNA and histone methylation, while HDAC1 (histone deacetylase 1) modulates P53 effects on apoptosis and cell growth.
Discussion
In the past four decades, bone marrow transplantation and chemotherapy have predominated the AML treatment landscape. Chemotherapy, especially the 7 + 3 regimen, continues to be the induction therapy of the first choice for many patients with newly diagnosed AML. Since 2017, several targeted therapeutics have been introduced to the AML treatment armamentarium. Despite these notable advances, the average life expectancy of patients with AML has not appreciably increased in recent years28,29. Because of the overall genetic and clinical heterogeneity of AML, meeting the critical unmet need of enhancing patient outcomes will require additional, novel, effective, and well-tolerated treatments.
A highly attractive treatment strategy for AML is to inhibit MDM2, restore the native tumor-suppressing functions of P53, and induce P53-dependent apoptosis in malignant cells. Preclinical and clinical studies of several MDM2 inhibitors have supported the viability of this approach to mediate antileukemic activity30,31,32,33.
Against this backdrop, we evaluated the ability of novel MDM2 inhibitor APG-115 to disrupt MDM2-P53 interactions in TP53wt AML. Not only did APG-115 exhibit antiproliferative cellular activity and induce both apoptosis and G0/G1 cell cycle arrest; it potentiated antileukemic effects when combined with SOC treatments in AML xenograft models. In the systemic MOLM-13 model, APG-115 augmented median survival by 18.5 days compared to vehicle control and significantly (by 2–6 days) compared to MDM2 inhibitor RG-7388.
APG-115 induced accumulation of P53 and its transcriptional target P21 protein in our study. Combining AZA, DAC, or Ara-C with APG-115 synergistically potentiated its P53/P21-activating effects, which can most likely be ascribed to shared effects of these agents on the same targeted signaling pathways. In this context, activation of the P53/P21 pathway restores tumor suppression function, leading to more profound DNA damage and apoptosis. However, the synergistic effects of APG-115 combination treatments could not be attributed to increased cell cycle arrest.
Intriguingly, MOLM-13 and MV-4-11 AML cell lines carrying TP53wt and concurrent FLT3-ITDmut mutations were the most sensitive to APG-115-induced growth inhibition, compared to non-FLT3-mutated OCI-AML-3 cells. In addition, the MOLM-13 CDX model is a diffuse systemic tumor with a high degree of malignancy and disease progression. Therefore, our data demonstrate that APG-115 as a single agent is highly effective in reducing the leukemic burden and prolonging survival of mice bearing AML in vivo, and FLT3mut AML cells may be more sensitive to MDM2 inhibitors compared to wild-type FLT3.
Evidence from the present study also suggests that the antileukemic activity of APG-115 is more potent than RG-7388 both in vitro and in vivo. When combined with SOC agents, APG-115 induced apoptosis more markedly, as evidenced by increased cleavage of PARP-1 protein. Unknown at this juncture is whether these promising but preliminary findings with APG-115 can be extended to patients with AML. To address this question, multiple clinical trials of APG-115 and other MDM2 inhibitors combined with SOC agents have been undertaken in patients with AML and MDS32. A phase Ib, open-label, 3 + 3 dose-escalation study will evaluate the safety (dose-limiting toxicities), pharmacokinetics, and pharmacodynamics of APG-115, alone or combined with AZA or Ara-C, in patients with r/r AML and relapsed and high-risk MDS (NCT04275518).
Other clinical trials of MDM2 inhibitors in patients with AML are ongoing (clinicaltrials.gov). Despite being the focus of ongoing clinical inquiry to assess potentially synergistic antitumor effects, these agents have unclear complementary mechanisms. Both AZA and DAC are DNA hypomethylating agents with previously reported apoptosis-inducing effects in AML cells34. Each agent acts chiefly by depleting DNMT1 and inducing DNA hypomethylation and damage34. DNMT1 forms complexes with UHRF1 and HDAC1 proteins in tumor cells, reducing expression of tumor suppressor promoter genes (e.g., P16INK4A, P14ARF, and P21) and hence fostering tumor growth26,27. In this context, activation of the P53-mediated apoptosis pathway has been implicated in DAC-induced cell death35. Ara-C is a pyrimidine nucleoside analog that induces DNA damage, activating the ATM/ATR-P53/P21 signaling cascade, inducing cell cycle arrest, and markedly impeding cellular proliferation36. These combinations of DNA hypomethylating, and cytotoxic, agents may act via the P53/P21 signal-transduction pathway.
One other potentially viable combination of APG-115 is with immuno-oncologic (I-O) therapies. In addition to promoting the intrinsic (mitochondrial) pathway of apoptosis, APG-115 exhibits potential immunomodulatory properties, according to our previous preclinical research37. Future studies might assess the availability of appropriate AML models in immunocompetent animals. Potential immunomodulatory effects of APG-115 in tandem with the PD-1 inhibitor pembrolizumab are being evaluated in an ongoing phase I trial of patients with unresectable or metastatic melanoma or advanced solid tumors (NCT03611868).
Hematological toxicity due to on-target myelosuppression by continuous MDM2 inhibition remains a challenge for MDM2-targeted treatment in clinic38. However, myelosuppression is manageable in the clinic. An important approach to mitigate the toxicity is to optimize the doses and dosing schedules. Various doses and schedules were evaluated in order to achieve the optimal balance between efficacy and toxicity in the preclinical setting39,40. It is interesting to unveil that the high-dose pulse of an MDM2 inhibitor led to a rapid onset of apoptosis40. Consistently, a high-dose pulse daily dosing of RG7388 for 5 days demonstrated antitumor activity equivalent to chronic administration39. Accordingly, the application of a high-dose pulse schedule of RG7388 achieved clinical responses with manageable toxicity in patients with AML25. Such a 5- to 7-day high-dose pulse schedule was successfully adapted in multiple clinical trials to evaluate MDM2 inhibitors in cancer patients (clinicaltrails.gov).
In conclusion, our study affords consequential new insights into the in vitro and in vivo tumor-suppressing effects of the novel MDM2 inhibitor APG-115, alone and in concert with SOC agents, in AML cells and animal models. We have formulated plausible molecular mechanisms underlying the synergistic effects of our MDM2 inhibitor in tandem with DNA-hypomethylating and DNA-damaging agents in AML, including P53 and downstream signaling pathways. These data support further, clinical development of APG-115, especially in combination with these SOC agents, to enhance AML management. In this connection, phase I/II clinical trials of APG-115 combined with SOC agents in AML are underway (NCT04358393).
Materials and methods
Cell lines and reagents
Human AML cell lines, including MV-4-11, MOLM-13, and HL-60, were purchased from American Type Culture Collection (ATCC; Manassas, VA, USA). Human AML cell line Ontario Cancer Institute‒Acute Myeloid Leukemia-3 (OCI-AML-3) was sourced from Cobioer (Nanjing, China) and SKM-1 from the Japanese Collection of Research Bioresources Cell Bank (JCRB). MV-4–11 cells were propagated in Iscove’s Modified Dulbecco’s Medium (IMDM, Gibco, Grand Island, NY, USA Cat# C12440500BT) supplemented with 10% fetal calf serum (FCS, AUSGENEX, Loganholme, QLD, Australia, Cat# FBSSA500-S). Other cell lines were propagated in Roswell Park Memorial Institute (RPMI) 1640 medium (Gibco, Cat# C11875500BT) containing 10% FCS. All experiments utilized genetically authenticated, microbial-free cells in their exponential phases of growth.
APG-115 (produced by Ascentage Pharma, Jiangsu, China) was dissolved in dimethyl sulfoxide (DMSO, Sigma, St. Louis, MO, USA, Cat# D8418) for in vitro experiments or suspended in 0.2% hydroxypropyl methylcellulose (HPMC, Sigma, Cat# H7509-25G) for in vivo animal studies. DAC (Cat# S120009), AZA (Cat# S178206), and Ara-C (Cat# S164806) purchased from Selleckchem were dissolved in DMSO for in vitro assays and saline for in vivo studies. RG-7388 (idasanutlin, MDM2-P53 inhibitor, MedChemExpress, Cat# HY-15676) was dissolved in DMSO for in vitro assays.
Cell viability assay
AML cells were seeded at a density of 8000 cells in an opaque 96-well plate and incubated with increasing concentrations of APG-115, AZA, DAC, or Ara-C, or with vehicle (DMSO). After treatment, cell viability was measured by Promega® Cell-Titer Glo® 3D Cell Viability Assay luminescence assay kit according to instructions from the manufacturer (Promega, Madison, WI, USA, Cat# G7571). Luminescence signal was detected using a BioTek Synergy H1 Hybrid Multi-Mode (Microplate) Reader (BioTek, Shanghai).
To assess the synergy of combination regimens for AML, we used CalcuSyn software (v2.0, Biosoft, Cambridge, UK), which allowed us to compute values for half-maximal inhibitory concentration (IC50) and the combination index (CI). A CI value of less than 0.9 indicates a synergistic effect; a CI value equal to 0.9 indicates an additive effect, and a CI value exceeding 0.9 indicates an antagonistic effect41. Cellular proliferative (i.e., viability) curves were plotted using Graphpad Prism version 6.0 software (GraphPad Software, San Diego, CA, USA).
Cell cycle and apoptosis analysis
For cell cycle analysis, we exposed TP53 wild-type AML cell lines to APG-115, DAC, AZA, or Ara-C, either alone or combined, for 48 h. After treatment, cells were harvested and fixed with ice-cold 70% (v/v) ethanol (General reagent, Cat# G7353N) for 24 h and centrifuged at 200×g for five minutes. Cell pellets were then washed once with phosphate-buffered saline (PBS) and stained with propidium iodide (PI) using a cell cycle analysis kit (Fcmacs, Cat# FMS-CCC01) according to manufacturer instructions. A total of 20,000 events were acquired and proportions of cells in each phase of the cell cycle were calculated using FlowJo™ software v10.4.2 (BD Biosciences; San Jose, CA, USA).
For apoptosis analysis, AML cells were treated with APG-115, DAC, AZA, Ara-C, alone or in combination, for 48 h, then subjected to flow cytometry analysis using an Annexin V/fluorescein isothiocyanate (FITC)/PI (Annexin V-FITC-PI) Apoptosis Detection Kit (BD Biosciences Cat# 556547). In brief, cells were washed once with cold PBS and incubated for 15 min at room temperature with Annexin V-FITC-PI in binding buffer. Cells were then analyzed on an Attune NxT Flow Cytometer (Life Technologies, Thermo Fisher, Carlsbad, CA, USA) using FlowJo software. Results were expressed as percentages of Annexin V+ cells. Experiments were performed in duplicate three independent times.
Human CD45+ cell staining and analysis by flow cytometry
Bone marrow cells from two femurs and spleen cells were collected and lysed in 2 mL 1× red-blood-cell lysis buffer (BD Biosciences, Cat# 555899) at room temperature for 2 min, followed by centrifugation at 1000 rpm for 5 min. Supernatants were decanted and discarded. Cell pellets were washed twice with PBS, followed by dark incubation with live/dead antibody (eBioscience, San Diego, CA, USA, Cat# L34976) for 15 min. Cells were then washed twice with PBS and incubated with human CD45-APC antibody (BD Biosciences, Cat# 555485) diluted in Magnetic-Activated Cell Sorting (MACS®) buffer (Miltenyi Biotec, Auburn, CA, USA, Cat# 130-091-221) at 1:5 for 30 min on ice. The buffer consisted of PBS (pH 7.2), 0.5% bovine serum albumin, and 2 mM EDTA diluted in MACS® bovine serum albumin stock solution with autoMACS® rinsing solution. After incubation, cells were washed twice with PBS and processed for flow cytometry analysis on the Attune NxT Flow Cytometer. Data were analyzed using FlowJo software and reported as percentages of human CD45+ cells within live cells.
Western blot analysis
As described previously37, cells were collected and lysed in radioimmunoprecipitation assay lysis buffer (Beyotime, China, Cat# P0013B) containing a 1% protease inhibitor cocktail (Yeasen, China, Cat# 20124ES10), 1% InStabTM phosphatase inhibitor cocktail (Yeasen, Cat# 20109ES05), and 1% phenyl-methyl-sulfonyl fluoride (PMSF Yeasen, Cat# 20104ES08). Protein concentration was quantitated using a bicinchoninic acid protein assay kit (Beyotime, Cat# P0010).
Equal amounts of soluble protein were loaded and separated using 4–20% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (GenScript, Cat# M42010C), followed by transfer to a polyvinylidene difluoride membrane (Millipore, MA, USA, Cat# IPVH00010) and then exposed to immunoblotting antibodies against P21 (CST, MA, USA, Cat# 2947S), P53 (CST, Cat# 2524S), poly (ADP-ribose) polymerase (PARP-1, Cat# 9532S), DNA methyltransferase (DNMT1, CST, Cat# 5032S), mouse double-minute 2 homolog (MDM2, CST, Cat# 86934S), anti-γ-H2AX (CST, Cat# 2577S), and β-actin (CST, Cat# 4970S). Antirabbit or antimouse horseradish peroxidase-conjugated secondary antibodies purchased from Yeasen (Cat# 33101ES60 and 33201ES60) were used at 1:5000 dilution. Immunoreactive proteins were visualized using an Azure c300 Chemiluminescent Western Blot Imaging System (Azure Biosystems, Dublin, CA, USA). Western blots were at least repeated twice to ensure reproducibility.
RNA sequencing
MOLM-13 cells were treated for 24 h. Total RNA was extracted from 1 × 106 cells using Invitrogen TRIzol® RNA Isolation Reagent (Thermo Fisher, MA, USA, Cat# 15596018) and RNeasy MinElute spin columns based on silica-membrane technology according to manufacturer’s instructions (Qiagen, Cat# 74204, Germantown, MD, USA). The integrity of the total RNA was assessed using a 2100 Bioanalyzer Instrument (Agilent, Santa Clara, CA USA) and quantitated using NanoDrop® (Thermo Scientific). A total of 500 ng of a high-quality RNA sample was used to construct a sequencing library. RNA quality was determined according to an optical density [OD] 260/280 nm of 1.9–2.0 and an RNA integrity number [RIN] of at least 8, where integrity is graded from 1 (most degraded) to 10 (least degraded).
Messenger RNA-focused sequencing libraries from total RNA were prepared using a TruSeq mRNA Library Prep kit (Illumina, Cat RS-122-2001, San Diego, CA USA). Sequencing was performed using this system according to protocols provided by the manufacturer for 2 × 150 base-pair paired-end sequencing at WuXi NextCODE (Shanghai). Reads were aligned to a reference genome by using Spliced Transcripts Alignment to a Reference (STAR) software42 with default parameters. Differential expression analysis was performed using the edgeR package in R (Bioconductor, Stanford, CA, USA)43. Criteria for significantly differentially expressed genes comprised a false-discovery rate of less than 0.05 and a |fold change| of at least two. A set of significantly upregulated or downregulated genes was inputted to test the significance of gene enrichment in a KEGG (https://www.genome.jp/kegg/) path by using GOstat packages in R. The criteria value was p < 0.01.
AML xenograft models
All experiments were conducted in the animal facility of GenePharma (Suzhou, China). Protocols and experimental procedures involving the care and use of animals were approved by the GenePharma Institutional Animal Care and Use Committee. Animals could acclimate to the environment for at least three days then transferred to a temperature- (20–26 °C) and humidity-controlled (40–60%, relative humidity) SPF room with a 12-h light/12-h dark cycle during the experimental period. Animals were housed in cages, with no more than five mice per cage. Animals had free access to sterile drinking water and food.
For the systemic AML model, 6- to 8-week-old female nonobese diabetic severe combined immunodeficient mice (NOD SCID) were pretreated with cyclophosphamide (150 mg/kg, intraperitoneally) for two consecutive days. The MOLM-13 systemic AML xenograft model was then established by intravenous inoculation of 1 × 107 cells. After 3 days, tumor-bearing mice were randomized to treatment with vehicle, APG-115, or AZA, alone and in combination. Mice were monitored daily for the development of hind-limb paralysis or abdominal swelling resulting from disease progression, as well as more than a 20% body weight loss. These clinical manifestations served as humane endpoints.
For the subcutaneous model, OCI-AML-3 cells (1 × 106) were injected subcutaneously into the right flanks of six- to eight-week-old nonobese diabetic severe combined immunodeficient (NOD SCID) mice. Approximately, 10 days after cellular inoculation, mice were randomly grouped based on primary body weight and tumor volume, which reached a mean of approximately 100–150 mm3. APG-115, AZA, and DAC were then administered alone or in combination. Tumor volumes were measured by caliper twice weekly and expressed in mm3 using the following formula:
where a was the long diameter of the tumor and b was the short diameter. Mice were humanely euthanized when their body weight loss >20% or a subcutaneous tumor volume reached 2000 mm3.
As a measurement of efficacy, a treatment/control (T/C%) value was calculated as the mean RTV of treated tumors (T) divided by the mean RTV of control tumors (C) × 100%, where TRTV was the relative tumor volume (RTV) of the treatment group and CRTV was the RTV of the control group. RTV is expressed as the ratio Vt/V1, where Vt is the average tumor volume at a certain time point (Day t) and V1 is the average tumor volume on the first day of treatment (Day 1).
Statistical analysis
One-way analysis of variance, followed by Games–Howell’s posttest to adjust for multiple comparisons44, was conducted to assess the statistical significance of differences between multiple treatment groups. In the MOLM-13 systemic AML model, the date of the last mouse death was used to analyze median overall survival, with the generation of Kaplan–Meier plots. Survival curves of different treatment groups were compared using the log-rank test with Bonferroni’s test for multiple comparisons. All data were analyzed in SPSS version 18.0 (IBM PASW Statistics for Windows; Chicago, IL USA). GraphPad Prism version 6.0 was used for graphic presentation.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Lindsley, R. C. et al. Acute myeloid leukemia ontogeny is defined by distinct somatic mutations. Blood 125, 1367–1376 (2015).
Grove, C. S. & Vassiliou, G. S. Acute myeloid leukaemia: a paradigm for the clonal evolution of cancer? Dis. Model Mech. 7, 941–951 (2014).
Ossenkoppele, G. & Lowenberg, B. How I treat the older patient with acute myeloid leukemia. Blood 125, 767–773 (2015).
Klepin, H. D. Myelodysplastic syndromes and acute myeloid leukemia in the elderly. Clin. Geriatr. Med 32, 155–173 (2016).
Ritchie, E. K. et al. Decitabine in patients with newly diagnosed and relapsed acute myeloid leukemia. Leuk. Lymphoma 54, 2003–2007 (2013).
Malik, P. & Cashen, A. F. Decitabine in the treatment of acute myeloid leukemia in elderly patients. Cancer Manag. Res. 6, 53–61 (2014).
Itzykson, R. et al. Azacitidine for the treatment of relapsed and refractory AML in older patients. Leuk. Res. 39, 124–130 (2015).
Fenaux, P. et al. Azacitidine prolongs overall survival compared with conventional care regimens in elderly patients with low bone marrow blast count acute myeloid leukemia. J. Clin. Oncol. 28, 562–569 (2010).
Kim, T. K., Gore, S. D. & Zeidan, A. M. Epigenetic therapy in acute myeloid leukemia: current and future directions. Semin. Hematol. 52, 172–183 (2015).
Rai, K. R. et al. Treatment of acute myelocytic leukemia: a study by cancer and leukemia group B. Blood 58, 1203–1212 (1981).
The AML Collaborative Group. A systematic collaborative overview of randomized trials comparing idarubicin with daunorubicin (or other anthracyclines) as induction therapy for acute myeloid leukaemia. Br. J. Haematol. 103, 100–109 (1998).
Burnett, A., Wetzler, M. & Lowenberg, B. Therapeutic advances in acute myeloid leukemia. J. Clin. Oncol. 29, 487–494 (2011).
Tallman, M. S., Gilliland, D. G. & Rowe, J. M. Drug therapy for acute myeloid leukemia. Blood 106, 1154–1163 (2005).
Howlader N. et al. SEER Cancer Statistics Review (CSR) 1975-2016. https://seer.cancer.gov/csr/1975_2016/ (2018).
American Cancer Society. Cancer Facts & Figures 2020. (American Cancer Society, Atlanta, 2020).
Vousden, K. H. & Prives, C. Blinded by the light: the growing complexity of p53. Cell 137, 413–431 (2009).
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Rucker, F. G. et al. TP53 alterations in acute myeloid leukemia with complex karyotype correlate with specific copy number alterations, monosomal karyotype, and dismal outcome. Blood 119, 2114–2121 (2012).
Amelio, I. How mutant p53 empowers Foxh1 fostering leukaemogenesis? Cell Death Discov. 5, 108 (2019).
Falcicchio, M., Ward, J. A., Macip, S. & Doveston, R. G. Regulation of p53 by the 14-3-3 protein interaction network: new opportunities for drug discovery in cancer. Cell Death Discov. 6, 126 (2020).
Bueso-Ramos, C. et al. The human MDM-2 oncogene is overexpressed in leukemias. Blood 82, 2617–2623 (1993).
Faderl, S. et al. The prognostic significance of p16(INK4a)/p14(ARF) locus deletion and MDM-2 protein expression in adult acute myelogenous leukemia. Cancer 89, 1976–1982 (2000).
Seliger, B. et al. Analysis of the p53 and MDM-2 gene in acute myeloid leukemia. Eur. J. Haematol. 57, 230–240 (1996).
Canon, J. R., Osgood, T., Saiki, A. Y. & Oliner, J. D. The MDM2 inhibitor AMG 232 causes tumor regression and potentiates the anti-tumor activity of MEK inhibition and DNA-damaging cytotoxic agents in preclinical models of acute myeloid leukemia. Cancer Res. 76, 3761 (2016). Abstracts of the 107th Annual meeting of American Association for Cancer Research.
Yee, K. et al. Phase 1/1b study of RG7388, a potent MDM2 antagonist in acute myelogenous leukemia patients. Blood 124, 116 (2014).
Unoki, M., Brunet, J. & Mousli, M. Drug discovery targeting epigenetic codes: the great potential of UHRF1, which links DNA methylation and histone modifications, as a drug target in cancers and toxoplasmosis. Biochem. Pharm. 78, 1279–1288 (2009).
Vijayaraghavalu, S., Dermawan, J. K., Cheriyath, V. & Labhasetwar, V. Highly synergistic effect of sequential treatment with epigenetic and anticancer drugs to overcome drug resistance in breast cancer cells is mediated via activation of p21 gene expression leading to G2/M cycle arrest. Mol. Pharm. 10, 337–352 (2013).
Dombret, H. et al. International phase 3 study of azacitidine vs conventional care regimens in older patients with newly diagnosed AML with >30% blasts. Blood 126, 291–299 (2015).
Lancet, J. E. Is the overall survival for older adults with AML finally improving? Best. Pract. Res. Clin. Haematol. 31, 387–390 (2018).
Andreeff, M. et al. Results of the phase I trial of RG7112, a small-molecule MDM2 antagonist in leukemia. Clin. Cancer Res. 22, 868–876 (2016).
Kojima, K. et al. MDM2 antagonists induce p53-dependent apoptosis in AML: implications for leukemia therapy. Blood 106, 3150–3159 (2005).
Tisato, V., Voltan, R., Gonelli, A., Secchiero, P. & Zauli G. MDM2/X inhibitors under clinical evaluation: perspectives for the management of hematological malignancies and pediatric cancer. J. Hematol. Oncol. https://doi.org/10.1186/s13045-017-0500-5 (2017).
Weisberg, E. et al. Inhibition of wild-type p53-expressing AML by the novel small molecule HDM2 inhibitor CGM097. Mol. Cancer Ther. 14, 2249–2259 (2015).
Hollenbach P. W., et al. A comparison of azacitidine and decitabine activities in acute myeloid leukemia cell lines. PLoS ONE https://doi.org/10.1371/journal.pone.0009001 (2010).
Brodska, B., Otevrelova, P. & Holoubek, A. Decitabine-induced apoptosis is derived by Puma and Noxa induction in chronic myeloid leukemia cell line as well as in PBL and is potentiated by SAHA. Mol. Cell Biochem. 350, 71–80 (2011).
Wang, H. et al. An ATM- and Rad3-related (ATR) signaling pathway and a phosphorylation-acetylation cascade are involved in activation of p53/p21Waf1/Cip1 in response to 5-aza-2’-deoxycytidine treatment. J. Biol. Chem. 283, 2564–2574 (2008).
Fang D. D., et al. MDM2 inhibitor APG-115 synergizes with PD-1 blockade through enhancing antitumor immunity in the tumor microenvironment. J. Immunother. Cancer https://doi.org/10.1186/s40425-40019-40750-40426 (2019).
Khurana, A. & Shafer, D. A. MDM2 antagonists as a novel treatment option for acute myeloid leukemia: perspectives on the therapeutic potential of idasanutlin (RG7388). Onco Targets Ther. 12, 2903–2910 (2019).
Higgins, B. et al. Preclinical optimization of MDM2 antagonist scheduling for cancer treatment by using a model-based approach. Clin. Cancer Res. 20, 3742–3752 (2014).
Jeay, S. et al. Dose and schedule determine distinct molecular mechanisms underlying the efficacy of the p53-MDM2 inhibitor HDM201. Cancer Res. 78, 6257–6267 (2018).
Chou, T. C. Drug combination studies and their synergy quantification using the Chou-Talalay method. Cancer Res. 70, 440–446 (2010).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Games, P. A. & Howell, J. F. Pairwise multiple comparison procedures with unequal n’s and/or variances: a Monte Carlo Study. J. Educ. Stat. 1, 113–125 (2016).
Kilkenny C., Browne W. J., Cuthill I. C., Emerson M. & Altman D. G. Improving bioscience research reporting: the ARRIVE Guidelines for Reporting Animal Research. PLoS Biol. https://doi.org/10.1371/journal.pbio.1000412.t1000001 (2010).
Acknowledgements
We thank Ascentage CMC and Analytic Center colleagues for synthesizing APG-115 and providing quality control. Assistance in paper preparation was provided by Stephen W. Gutkin with Ascentage. Preparation of this study report was informed by Animal Research: Reporting In Vivo Experiments (ARRIVE) guidelines for preclinical research45. The research was partially supported by grants from the National Science and Technology Major Project (2018ZX09301015-001) and the Major Innovation Team of Soochow (ZXT2018004). The research costs were also funded by Ascentage Pharma (Suzhou) Co., Ltd., where all authors were employees at the time the study was conducted.
Author information
Authors and Affiliations
Contributions
D.D.F. and Q.T. designed experiments, interpreted the results, and drafted the paper, with assistance from Stephen W. Gutkin; Y.K., T.R., Q.W., N.L., X.F., J.G., D.X., Y.Y performed the experiments and analyzed the results; D.X. analyzed RNA-seq data; J.D. designed cell viability assays and interpreted the results; D.Y. and Y.Z. supervised the research and reviewed and finalized the paper. All authors read and approved the final paper and agreed to be responsible for the integrity of its content.
Corresponding authors
Ethics declarations
Ethics statement
This article contains studies of animals performed by the authors. The studies have already been approved by the local research ethics committee. For this type of study, formal consent is not required.
Conflict of interest
The authors are full-time employees of and stockholders in Ascentage Pharma.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by A. Rufini
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fang, D.D., Tang, Q., Kong, Y. et al. MDM2 inhibitor APG-115 exerts potent antitumor activity and synergizes with standard-of-care agents in preclinical acute myeloid leukemia models. Cell Death Discov. 7, 90 (2021). https://doi.org/10.1038/s41420-021-00465-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-021-00465-5
- Springer Nature Limited