Abstract
N6-methyladenosine (m6A) epitranscriptional modifications widely exist in RNA, which play critical roles in RNA metabolism and biogenesis processes. Long non-coding RNAs (lncRNAs) are class of non-coding RNAs longer than 200 nucleotides without protein-coding ability. LncRNAs participate in a large number of vital biological progressions. With the great improvement of molecular biology, m6A and lncRNAs are attracting more attention from researchers and scholars. In this review, we overview the current status of m6A and lncRNAs based on the latest research, and propose some viewpoints for future research perspectives.
Similar content being viewed by others
FACTS
-
1.
LncRNAs have a series of functions of mediating transcriptional and post-transcriptional gene expression in a wide range of mechanisms and diseases.
-
2.
Biological actions result in the particular transcriptome topology of m6A.
-
3.
NcRNAs can feedback to afflict the effect of m6A modifications, then complicating the outcomes of pathophysiological activities in vitro and in vivo.
Background
Given that a wide spectrum of protein-coding RNAs and non-coding RNAs (ncRNAs) can transcribe with the vast majority of genomic sequences, all organisms possess a far more complex transcriptional landscape than we imagined [1,2,3]. The transcription of ncRNAs is estimated to exceed 90% of the mammalian genome. Recently, evidence has emerged that these non-coding transcripts have biological significance and are indeed not the case of garbage or transcriptional noise [4, 5]. In the extended view of the genome and transcriptome, the current catalog of genetic elements is filled with long non-coding RNAs (lncRNAs) [6]. LncRNAs are ncRNAs with insufficient latency to code protein, which are transcripts of >200 nucleotides in length [7]. Less than 3% of lncRNAs have specified functions. LncRNAs partake in numerous vital biological phenomena, such as enzymatic activity, alternative forms of chromosome conformation, and functional structured RNA domains. Furthermore, unique codes of lncRNA expression function as scaffolds, decoys, or signals to harmonize cell proliferation, differentiation, apoptosis, and metabolism. Little is known about most of the functions of lncRNAs, and even a large quantity of lncRNAs may not have special functions [5, 8, 9].
MicroRNA (miRNAs/miRs), lncRNAs, and circular RNAs (circRNAs) belong to ncRNAs, all of which share a common characteristic that they originate from the transcribed genomes and demonstrate biological functions [10, 11]. CircRNAs are considered to be a category of single-stranded covalently closed RNA molecules that were first discovered in eukaryotes nearly 40 years ago [12]. Compared with lncRNAs, circRNAs can be deemed to a certain extent as a special kind of lncRNAs, which have been modified by an action called back-splicing [13].
N6-methyladenosine (m6A) epitranscriptional modifications are widely present in RNA, particularly in eukaryotic messenger RNAs (mRNAs), which play pivotal roles in mRNA metabolism and numerous biogenesis processes [8, 14,15,16]. With new technological advances in RNA modification catalysis, the universal of m6A has been published in eukaryotic cells besides its invertibility in mammalian cells [17, 18]. Recently, m6A has attracted widespread interest due to its significant influence on cell differentiation, proliferation, migration, invasion, and apoptosis [19, 20]. Three functional proteins, known as “writers”, “readers” and “erasers”, are involved in these internal modifications of m6A. All of these internal modifications refer to “epitranscriptomics” [21,22,23]. These genes that can be affected by m6A level changes can exert dramatic effects on cancer progression and growth. This suggests that m6A and its regulators may assume a crucial role in the diagnosis and treatment of cancers [7, 22].
Despite some recent substantial progress in the mechanism of m6A modifications of ncRNAs, little clarification has been obtained regarding the modification of lncRNAs. In this review, we summarize the role of m6A modifications in the mediation and function of lncRNAs and discuss its potential future application and research directions.
Three kinds of enzymes and detection techniques of m6A modifications
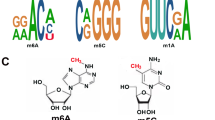
The vast bulk of m6A modifications result in the unanimity motif of RRm6ACH (in which R represents A or G, and H represents A, C, or U). Accompanied by site and cell/tissue explicitness, m6A modifications are generally concentrated in the 3′ untranslated regions (3′-UTRs), that is, they are adjacent to the stop-codons of mRNAs and within long internal exons, contributing to a distinctive m6A-derived transcriptome topology [24,25,26]. Almost all of the features of mRNA processing can be ascribed to the effect of m6A functions, like pre-mRNA splicing, export, translation, and stability, therefore exhibiting further influences on the development of humans and diseases [19].
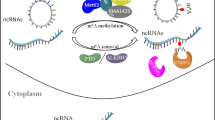
The m6A modification procedure is dynamic and invertible [1]. Enzymes working in the m6A modification regulation incorporate “writers” and “erasers” which respectively assemble and dislodge the methylation that can be identified by “readers” [14, 27]. In biological systems, these effector proteins have multiple significant features that can be exhibited by their complicated and changeable functions, which are strongly afflicted by the local environment (Fig. 1).
The m6A modification procedure is dynamic and invertible. The enzymes in m6A modification regulation incorporate “writers” and “erasers”, which respectively assemble and dislodge the methylation and can be identified by “readers”. In biological systems, these effector proteins have substantial signature features which can modify the physiological functions of lncRNAs, including transcription (promote or terminate), expression (advance or silence translation via diverse mechanisms), splicing (affect the performance), stability (enhance the structure or facilitate the degradation), and binding capacity. The entire metabolism can influence the functions of cells.
Writers
In epitranscriptomics, the writers comprise methyltransferase-like 3/14 (METTL3/14) that can form complexes to ensure the installation of m6A methylations into mRNAs [16, 21] and Wilms tumor 1-associating protein (WTAP) that assists METTL3/14 complexes in positioning nuclear spots and ensuring the stability of the complexes [23, 28,29,30]. The difference between groups of cell lines can indicate the spot effect of METTL3 which is impacted by the protein content proportion of METTL3 and METTL14 and other complexes in adapter subunits. Post-translational modifications (PTMs) proximate to the cytoplasmic presence can show the probability of adjusting the interactions between METTL3 and its partner proteins.
Furthermore, there are other kinds of writers of m6A modifications. For instance, Vir-like m6A methyltransferase associated (VIRMA, aka KIAA1499) that plays an obvious part in methylation deposition to the 3′-UTR [26], RNA binding motif protein 15/15B (RBM15/15B) that attracts the METTL3/14 complexes to methylate U-riched sites by relying on WTAP [31], and METTL16 that binds to pre-mRNAs and various ncRNAs expand our view of the active m6A methyltransferase installed on cellular RNAs [32, 33]. Meanwhile, zinc finger CCCH-type containing 13 (ZC3H13) promotes the “writer” to be located in the nuclear [31]. Recently, METTL5 and ZCCHC4 have been reported to act as modifiers for 18S and 28S ribosomal RNAs (rRNAs), respectively [34, 35]. METTL5 is stabilized by TRMT112, and the action of the METTL5-TRMT112 complex reveals a process very similar to DNA methyltransferases [34, 36]. In the meantime, ZCCHC4 pinpoints to the nucleolus, which assembles the ribosome. The proteins involved in RNA metabolism are overexpressed with the participation of ZCCHC4 [35, 37].
Erasers
As a reversible chemical process of RNA modifications, erasers are a type of enzyme that can be demethylated during m6A modifications. Fat mass and obesity-associated protein (FTO), α-ketoglutarate-dependent dioxygenase homolog 5 (ALKBH5), and ALKBH3 are part of erasers, which significantly influence the biological processes of diseases with a train of complicated interactions that selectively identify m6A-labeled target mRNAs [28, 38,39,40]. So far, FTO and ALKBH5/3 are dominated by a family of α-ketoglutarate-dependent dioxygenases that demethylate m6A via Fe2+ and α-ketoglutarate [41]. Nevertheless, compared with m6A, FTO has been implicated in mRNAs and snRNAs with the demethylation of N6 and 2′-O-dimethyladenosine (m6Am) and in tRNAs with the demethylation of N1-methyladenosine (m1A) [42]. Especially in m6Am modifications, FTO indicates a tendency to be over-scaled to participate, which can preferentially weaken the stability of mRNAs in cells [43]. Prior research has elucidated that unstable mRNAs may be generated from different sites of FTO in a series of cell lines [38]. ALKBH5 is precisely located in the nucleus. The overexpression of ALKBH5 in cells vastly shrinks the processing of m6A in mRNAs [44]. A recent study has revealed that ALKBH3 is inclined to be demethylated on tRNA as opposed to on mRNA or rRNA. Furthermore, ALKBH3 is an undeveloped molecular marker in cancers for its ability as an enzyme to recover DNA damages [45].
Readers
The ‘readers’ are regarded as a series of proteins that can serve as chemical markers, manipulate RNA structure, or be recruited during the m6A modification. There exist several “readers”, like the YT521-B homology (YTH) domain family (including YTHDC1/2 and YTHDF1/2/3) [16, 39, 46], heterogeneous nuclear ribonucleoproteins (including HNRNPA2B1, HNRNPC, and HNRNPG) [47], insulin-like growth factor 2 mRNA binding proteins 1/2/3 (IGF2BP1/2/3) [21], NF-κB-associated protein (NKAP) [48], eukaryotic translation initiation factor 3 (eIF3), proline-rich coiled-coil 2A (Prrc2a), and zinc finger CCCH domain-containing protein 13 (ZC3H13) [49, 50].
The YTH domain has been reported as an m6A-binding domain, which was the earliest distinguished reader. YTHDF1 and YTHDF2 give impetus to the translation and decay of m6A-methylated mRNAs, respectively. In the cytoplasm, YTHDF3 dramatically increases the metabolism of m6A-methylated mRNAs as a group with YTHDF1 and YTHDF2 [46]. YTHDC1 mainly in the nuclear exhibits several functions, such as promoting the export of mRNAs, enhancing the deterioration of specific transcripts, and orchestrating the splicing of mRNAs using the ideal assembly of certain splicing factors [51]. YTHDC2 is related to the stabilization and translation of mRNAs, but the feature of binding domains of RNA remains ambiguous [52]. RNA-protein interactions can be mediated by sites near or around RNA structures that are transformed by m6A. Those interactions are called “m6A-switch”. HNRNPs, including HNRNPC, HNRNPG, and HNRNPA2B1, modulate selectively tagged transcripts during splicing or processing [28, 53]. Especially with HNRNPA2B1 which mediates the processing of RNAs by m6A in the nuclear and peculiarly identifies m6A-modified RNAs, eIF3 binds to the 5′-UTR of mRNAs modified by m6A to establish the translation expression [30, 54]. Meanwhile, IGF2BPs enhance the stabilization of target genes and related translations [55]. Prrc2a is a novel m6A reader, which prefers to bind to a methylated probe, aka the consensus GGACU motif in the coding sequence, to facilitate the stability of mRNA expression [56].
m6A modification detection methods
Analysis of m6A modifications
Dot blot
The dot blot (or slot blot) technology, as a study in molecular biology, is a semi-quantitative or quantitative detection of global changes in m6A levels in entire or individual RNA species. The dot blot technology remains the favored practice option because of its plain and inexpensive cost. The dot blot technology is simpler to be organized than western blotting, Northern blotting, or Southern blotting but is afflicted by the short susceptibility when low intrinsic specimens were utilized for m6A modifications. Nevertheless, with a novel method, Nagarajan et al. [57, 58] generated a technique through an immunoprecipitation-based enrichment of the RNA m6A modification step after the dot blot quantification. This technique can be utilized to not only investigate m6A modifications of total RNA but also to enrich specific RNAs for the analysis of RNAs in the individual species, such as mRNA, tRNA, rRNA, or miRNA. Nevertheless, regarding m6A detection, the dot blot technology can just detect the existence of m6A or compare the quantity of m6A among various groups [59].
The electrochemical immunosensor method
Most analytical techniques present challenges in terms of their achievement and final cost. A novel choice is supplied by the serviceability and susceptibility of the electrochemical immunosensor method that focuses on m6A-5′-triphosphate (m6ATP), the anti-m6A antibody, to track m6A. A previous study has manifested the repeatability, specificity, wide linear range, and lack of the detection limit of the immunosensor, which suggests that this method can be the technological basis for RNA and DNA tests for its advantages of being cheaper, convenient, and higher sensitive [60].
Quantification detection of m6A modifications
m6A sequencing (m6A-seq) and methylated RNA immunoprecipitation sequencing (MeRIP-Seq) are second-generation sequencing after catching m6A methylated RNA fragments by m6A specific antibodies for the immunoprecipitation technology [20, 27]. m6A-level and isoform-characterization sequencing (m6A-LAIC-seq) is basically the same as m6A-seq but widens the perception of m6A biological characteristics and depends on m6A-IP of full-length poly (A) + RNA [61, 62]. However, this method can be applied to measure the levels of m6A at each site but not stoichiometrically analyze the single modified nucleotide [7, 61]. All of these three methods are the most common biological techniques which have been adopted in the detection of m6A.
Site-specific cleavage and radioactive labeling followed by ligation-assisted extraction and thin-layer chromatography (SCARLET), is an accessible technique to research the biological functions of RNA modifications using reachable equipment and materials, together with 5-methylcytosine, pseudouridine, and 2′-O-methyl ribonucleosides. In m6A, SCARLET can be employed to estimate the exact sites and status of m6A modifications in any supposed locations of mRNAs/lncRNAs with an individual resolution of nucleotides. It can help m6A-seq, MeRIP-Seq, and m6A-LAIC-seq to precisely determine the single-base resolution of RNAs [7, 63, 64].
Besides immunoprecipitation enrichment, there are some other methods to detect m6A, including crosslinking and immunoprecipitation (CLIP) [65, 66]. The CLIP builds covalent bonds between proteins and RNAs with direct contact with ultraviolet (UV) light irradiation and is a state-of-the-art method that spread the practice to stabilize direct protein-RNA interactions. This technology, initially matured in Escherichia coli, is widely recognized as an essential procedure for researching dynamic protein interactions in multitudes of cellular processes in multifarious biological systems [67, 68]. CLIP consists of photo-crosslinking-assisted m6A sequencing (PA-m6A-Seq) [69], m6A individual-nucleotide-resolution cross-linking and immunoprecipitation (miCLIP) [70], and m6A-CLIP. The amplification of polymerase chain reaction (PCR) can advance the comparatively poor efficiency of this reaction, and the extremely purified protein-RNA complexes can potentially be obtained by the covalent binding. Arguello et al. advanced a chemical proteomics approach that depends on photo-cross-linking with synthetic diazirine-containing RNA probes and quantitative proteomics to profile RNA-protein interactions modulated by m6A. The proteins that can be used for combination are YTH domain-containing (reader) and ALKBH5 (eraser) [71]. However, the accuracy of the detection is limited by the fact that the crosslink is still feasible to be constructed in adjacent RNA sequences and that UV light has efficient penetrability.
The sequence-specific endoribonuclease MazF belongs to Escherichia coli toxins, which is engaged in growth mediation in response to stress. It includes MAZTER-seq and m6A-sensitive RNA-Endoribonuclease-Facilitated sequencing (m6A-REF-seq). Interestingly, with the specific capability against ACA sequences, endoribonuclease is sensitive to m6A to degrade mRNAs, decay protein synthesis, and enhance growth arrest. Theoretically, in the presence of MazF, every piece should start at the ACA site (5′ACA) and end downstream at the ACA site (3′ACA) [72, 73]. Mounting evidence unravels that the RNA cleavage reaction at the 5′-ACA-3′ site occurs only in single-stranded RNAs and never in double-stranded RNAs [74]. However, little is known about the susceptibility of MazF to the methylated presence of m6A bases in structured RNAs.
Deamination-adjacent to RNA modification target sequencing (DART-seq), an antibody-free process to assess m6A locations, APOBEC1 expression leads to C-to-U deamination near the sites of m6A residues after the binding of the cytidine deaminase APOBEC1 to the YTH domain of m6A. It is also possible to isolate thousands of m6A sites in 10 ng entire RNA and to identify the quantification of m6A aggregation in cells. Furthermore, the long-read DART-seq can harvest the distribution of m6A among the individual transcripts. However, the functional quality of DART-seq is currently limited in vitro [75].
High-resolution melting (HRM) analysis is a high-throughput measurement method that can be utilized to check m6A modification as an elementary process. The RNA mixed-specimens can be modified stably from 100% methylation to 100% unmethylation with the HRM analysis. The presence of m6A can be easily characterized at the special position of RNA using entire RNA samples and qPCR machines by HRM. Although the monomethylation of the adenosine exocyclic amino group cannot change the hydrogen bonding in the nucleic acid duplex, it can influence the stacking interaction that can impact the melting properties. The HRM analysis may deepen the understanding of the dynamic modification of certain RNAs [76].
Recently, two innovative techniques have been developed for RNA modification detection. Wang et al. researched that with the help of dithiothreitol (DTT)-mediated thiol-addition chemical reactions, the unsteady N6-hydroxymethyladenosine (hm6A) can be transformed into the extra sturdy N6-dithiolsitolmethyladenosine (dm6A). An FTO-assisted m6A selective chemical labeling method (aka m6A-SEAL), which is associated with the action that hm6A in RNA can be oxidized by the FTO enzyme of m6A, is advanced to particularly identify the transcriptome-wide m6A [62]. In addition, although cap m6Am, a newly identified reversible RNA modification, can be oxidized to cap hm6Am, m6A-SEAL-seq simply oxidizes internal m6A to hm6A under the present FTO oxidation conditions.
Furthermore, m6A-label-seq is a metabolic labeling method to check the mRNA m6A transcriptome wide at base resolution. By feeding a methionine analog, Se-allyl-L-selenohomocysteine, the methyl group on the enzyme cofactor S-adenosyl-L-methionine (SAM), can be replaced by allyl cells. In this way, the assumed position of m6A-generating adenosine can be substituted by N6-allyladenosine (a6A) and modified. The advantages of this method are the clustered m6A locations and the ability to identify m6A modifications of nuclear nascent RNAs [77].
There is no perfect method to detect m6A modifications. Therefore, it is extremely necessary to develop more technologies at high levels of single-base and quantitative sequencing.
The lncRNAs and m6A modifications/regulators and diseases
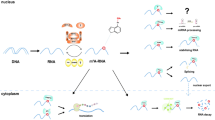
Numerous studies in recent years have elaborated that lncRNAs can be modified by m6A and orchestrate m6A regulators in disparate tissues or diseases due to the importance of m6A modification and the generous acceptance of next-generation sequencing technologies in combination with the escalation of bioinformatics (Fig. 2).
In head and neck squamous cell carcinoma, non-small-cell lung cancer, chronic kidney disease, thyroid cancer, human osteosarcoma, human renal cell carcinoma, epithelial ovarian cancer, breast cancer, lung adenocarcinoma, and cadmium-induced oxidative damage of pancreatic β-cells, the metabolism of m6A can accelerate tumor growth, migration, or invasion. However, in dendritic cells, cervical cancer, and pancreatic cancer, the functions of m6A regulators trigger the decrease of migration, invasion, or tumorigenicity. However, in the same disease, like hepatocellular carcinoma, colorectal cancer, and prostate cancer, different m6A regulators and lncRNAs can, to some extent, display dissimilar functions in the disease process, such as improving tumor growth or decay.
LncRNA regulation in diseases
Testis-associated highly conserved oncogenic long non-coding RNA (THOR)
THOR, a singular ultra-conserved cancer-specific lncRNA, mediates cell growth and is entirely expressed in testis and a wide variety of human cancers. In vitro and in vivo, the lack of THOR restrains the proliferation, migration, and invasion of a series of cancer cells [78, 79]. Meanwhile, highly expressed THOR increases IGF2BP1 mRNA expression, which can encourage the survival and proliferation of human osteosarcoma (OS) cells [80]. In addition, the overexpression of THOR also results in the upregulation of IGF2BP1 to enhance the survival and proliferation of human renal cell carcinoma (RCC) cells [81].
Metastasis-associated lung adenocarcinoma transcript 1 (MALAT1)
It is widely known that MALAT1 with various m6A sites is an extraordinary transcript modified by m6A with a large percentage of transcripts in several cell lines. As a competitive endogenous RNA, MALAT1 acts as a sponge of miR-1914-3p through Yes-associated protein (YAP) to accelerate the invasion and metastasis of non-small-cell lung cancer (NSCLC) cells [82]. Meanwhile, renal fibrosis plays a crux part in chronic kidney disease (CKD). MALAT1 leads to the promotion of renal fibrosis in patients with obstructive nephropathy (ON). The MALAT1/miR-145/focal adhesion kinase (FAK) pathway has been noted to assume a significant role in TGF-β1-induced renal fibrosis [83]. MALAT1 can aggressively bind competitively to miR-204 to upregulate MYC, thus facilitating the proliferation, migration, and invasion of thyroid cancer (TC) cells and reducing tumor growth and cell apoptosis [84]. In addition, MALAT1 is revealed to participate in cadmium-induced oxidative damage in pancreatic β-cells, which binds to proteins through m6A modifications [85].
Other lncRNAs
The high m6A level of nuclear paraspeckle assembly transcript 1 (NEAT1)-1 is correlated with the bone metastasis of prostate cancer (PCa). Ectopically expressed NEAT1-1 can induce cancer cell metastasis to the lung and bone, which is caused by NEAT1-1 with m6A site mutations. With the help of METTL3, the new CYCLINL1/CDK19/NEAT1-1 axis can be used as an undeveloped mechanism in the pathogenesis and growth of bone metastatic PCa [86]. CCAT1 and CCAT2, oncogenic lncRNAs, can be adopted as the independent predictors of poor prognosis and reduce the stability through VIRMA downregulation to depress the aggressive phenotypes of PCa cells [87].
In colorectal cancer (CRC) tissues, the direct interactions between growth arrest-specific transcript 5 (GAS5) and the WW domain of YAP facilitate the transferring of endogenous YAP from the nucleus to the cytoplasm and force YAP phosphorylation and afterwards ubiquitin-mediated degradation of YAP, ensuring the obstruction of CRC progression in vitro and vivo [88]. Furthermore, RP11 expression is elevated with increasing stage in CRC patients. RP11 is conclusively correlated with the migration, invasion, and epithelial mesenchyme transition (EMT) of CRC cells in vitro. RP11 can also promote liver metastasis in vivo [28].
LINC00958 is a lipogenesis-related lncRNA whose expression is upregulated in hepatocellular carcinoma (HCC) cell lines and tissues. The high level of LINC00958 can predict the low overall survival of HCC patients. After LINC00958 functions as a sponge of miR-3619-5p, hepatoma-derived growth factor (HDGF) is upregulated, which enhances the lipogenesis, progression, and malignant phenotypes of HCC in vitro and vivo [89]. Compared with adjacent normal tissues, KCNK15-AS1, which can suppress the migration and invasion of MIA PaCa-2 and BxPC-3 cells, reveal low expression in pancreatic cancer tissues [90].
RHPN1-AS1, a sponge of miR-596 to promote LETM1 expression and stimulate the FAK/PI3K/Akt pathway, augments the proliferation and metastasis of epithelial ovarian cancer (EOC) cells in functional experiments in vitro and in vivo [91]. Meanwhile, versus adjacent normal tissues, GAS5-AS1 expression is obviously diminished in cervical cancer (CC) tissues. GAS5-AS1 substantially subdues CC cell proliferation, migration, and invasion in vitro, and specifically weakens CC tumorigenicity and metastasis in vivo. As a result, the low expression of GAS5-AS1 is notably associated with tumor stage, distant and lymphatic metastasis, and poor prognosis in CC patients [92].
LNCAROD, an oncogenic lncRNA, displays differential expression between head and neck squamous cell carcinoma (HNSCC) and normal samples, and the overexpression of LNCAROD is linked to the advanced T stage and poor overall survival of HNSCC. The decline in cell proliferation and mobility in vitro and tumorigenicity in vivo is attributed to the loss of LNCAROD expression, but the result is antithesis when LNCAROD is overexpressed [93].
LncRNA Dpf3 has the capability of manipulating dendritic cell (DC) migration which can be inhibited by CCR7. The DC migration is vital for the stabilization of immune homeostasis and the inauguration of protective immunity. The loss of DC-specific lncRNA Dpf3 can increase the CCR7-mediated DC migration, which contributes to overstating adaptive immune reactions and inflammatory injuries. In the meantime, the binding between lncRNA Dpf3 and HIF-1α can decrease the transcription of the HIF-1α-dependent glycolytic gene Ldha, thereby frustrating DC migratory capacity and glycolytic metabolism [94].
ABHD11-AS1 is highly expressed in NSCLC tissues and cells, and this upregulation in the ectopic area shared a tight association with the negative prognosis of NSCLC patients [95]. Just like ABHD11-AS1, KB-1980E6.3 also shows an association with the negative prognosis of breast cancer through its abnormal overexpression in breast cancer clinical tissues. In vitro and in vivo, the upregulated KB-1980E6.3 can modulate the tumorigenesis and self-renewal of breast cancer stem cells in the hypoxic microenvironment [96]. MEG3 overexpression constrains the proliferation, migration, and invasion of HCC cells. However, the anti-cancer effect of MEG3 can be negated by the high level of miR-544b [97]. Also, cell proliferation, migration, and invasion are reduced and cell apoptosis is increased in RMRP-knocked down lung adenocarcinoma cell lines. The findings indicate that the aberrant upregulation of RMRP is strictly related to the negative prognosis of patients with lung adenocarcinomas [98] (Table 1).
The interactions between lncRNAs and m6A modifications/regulators
THOR
With the high enrichment of m6A in THOR transcripts, YTHDF1 and YTHDF2 can mediate the stabilization and degradation of THOR. The interaction of these m6A-dependent RNA-proteins can ensure the stability of the oncogenic part of THOR. Meanwhile, the interaction between THOR and IGF2BP1 illustrates that the mRNA activity stability of IGF2BP1 is enhanced by THOR [78, 79]. Furthermore, the target mRNA of IGF2BP1 can also be downregulated through silenced THOR [80]. More importantly, IGF2BP1 expression can also be elevated by the overexpression of THOR, which can facilitate the survival and proliferation of human RCC cells [81].
MALAT1
In NSCLC, the high level of m6A modifications mediated by METTL3 can augment MALAT1 expression. In the meantime, the METTL3/YTHDF3 complex can stabilize MALAT1 [82]. METTL3 can be upregulated and considered to increase MALAT1 expression in TGF-β1-treated HK2 cells. Thus, m6A modifications can afflict MALAT1 expression and possibly exert effects on the MALAT1/miR-145/FAK pathway in renal fibrosis during CKD [83]. MALAT1 and IGF2BP2 are identified to be highly expressed in TC, which is correlated with lowly expressed miR-204. Because IGF2BP2 is confirmed as a target gene of miR-204, MALAT1 can competitively bind to miR-204 to reduce IGF2BP2 expression and elevate MYC expression through m6A modifications [84]. In addition, MALAT1, which lacks m6A modifications, inhibits the metastatic potential of cancer cells. The concatenated m6A residues on MALAT1 can allow YTHDC1 to obtain nuclear speckles for modifying the expression of key oncogenes to enhance the metastatic potential of cancer cells [99]. MALAT1 is determined to decrease with the positive relationship with m6A modifications (including METTL3 and FTO), proving that m6A modifications and m6A-modified lncRNAs potentially are implicated in oxidative damages [85].
X-inactive specific transcript (XIST)
XIST is a lncRNA that modulates the transcriptional silencing of genes on the X chromosome. The knockdown of METTL3 or RBM15 can damage the transcriptional silencing of XIST-mediated genes. RBM15 can regulate XIST to tie up the m6A-methylation complex and be linked to particular sites in RNA. Moreover, YTHDC1 prefers to detect m6A in XIST and maintains the XIST-mediated gene silencing with the loss of m6A [100]. XIST can also be documented as a downstream target of METTL14. With the reorganization by YTHDF2, XIST methylated by m6A can cause the decay of CRC [101].
Other lncRNAs
In CRC, it is worth noting that as a new target of YAP, YTHDF3 can be a linchpin in the YAP pathway which contributes to m6A-modified GAS5 degradation. In clinical, the protein levels of YAP and YTHDF3 are negatively related to GAS5 expression in cancer tissues from CRC patients [88]. The overexpression of METTL3 can enhance RP11 expression and the RNA-protein binding between RP11 and HNRNPA2B1. The expression of RP11 can be repressed by ALKBH5 overexpression. These results suggest that m6A mediates RP11 expression in CRC cells [28].
In PCa tissues, the m6A level of NEAT1-1 is an influential predictor of eventual death. A lack of NEAT1-1 or the decreased m6A of NEAT1-1 causes conspicuous damages to Pol II Ser-2p levels in the promoter of RUNX2. CYCLINL1 is not correlated with NEAT1-1 in P-18 cells, which knocks down METTL3. Because of the pivotal role of m6A on NEAT1-1 in modifying Pol II ser2 phosphorylation, NEAT1-1 has the potential to be an innovative and specific target for the treatment and diagnosis of bone metastasis cancers [86]. The knockdown of VIRMA can crucially diminish the level of m6A, which causes the low expression of CCAT1/2. Furthermore, the repression of VIRMA and m6A can reduce the stability and enrichment of CCAT1/2 transcripts. As a result, VIRMA is essential for maintaining the level of m6A, and malignant phenotypes can be suppressed by decaying CCAT1/2 transcripts [87].
In HCC, GATA3-AS, which can be transcribed from the antisense strand of the GATA3 gene, acts as a guidance element in the preferential interaction between the METTL3/METT14 complex and the GATA3 pre-mRNA. Ultimately, the growth and metastasis of liver cancer, which are driven by the METTL3/METT14 complex and GATA3-AS, are regulated by GATA3 [102]. LINC00958 can be upregulated by stabilizing its RNA transcript in the presence of METTL3-mediated m6A modification in HCC [89]. Meanwhile, ALKBH5, which can m6A-demethylate KCNK15-AS1 and mediate cell motility induced by KCNK15-AS1, exhibits low expression, thus considerably augmenting total RNA methylation in pancreatic cancer cells [90].
Following the knockdown of METTL3, the expression and stabilization of RHPN1-AS1 are decreased in EOC, suggesting that m6A modifications can improve the transcriptional stability and m6A level of RHPN1-AS1, which may explain the results that RHPN1-AS1 is remarkably overexpressed [91]. However, GAS5-AS1 stabilization can be enhanced through the interactions between GAS5-AS1 and ALKBH5 and the depression of GAS5-AS1 m6A modifications in CC [92]. More intriguingly, METTL3/14-mediated m6A modifications promote the stability of LNCAROD, and the dysregulation of m6A may cause the abnormal expression of LNCAROD in HNSCC [93].
In DC, the CCR7 chemokine receptor can accelerate the quick temporary migration to empty lymph nodes. CCR7 stimulation reduces RNA decay by decreasing m6A modifications, thus upregulating lncRNA Dpf3. Meanwhile, although CCR7 induction exerts no direct impact on YTHDF2, it can diminish the recognition of YTHDF2 to m6A sites of lncRNA Dpf3. In conclusion, CCR7 stimulation elevates lncRNA Dpf3 expression by the m6A demethylation and the reduction of m6A-related RNA deterioration of lncRNA Dpf3 [94].
The m6A modification can be increased by the transfection of METTL3 in NSCLC cells to increase the stability of ABHD11-AS1. Consequently, the overexpression of ABHD11-AS1 augments the effects of Warburg on NSCLC, including excess glucose, lactate production, and ATP accumulation, and the overexpression of ABHD11-AS1 prominently silences the protein level of KLF4 [95]. The KB-1980E6.3/IGF2BP1/c-Myc axis modifies the coding region instability determinant (CRD) mRNA of c-Myc to ensure the stabilization of c-Myc mRNA [96]. In addition, METTL3 silencing reduces the m6A modification in MEG3 and conspicuously diminishes the stabilization of MEG3 [97]. ALKBH5 can upregulate RMRP through the demethylation of m6A, and the tumorigenesis can be proscribed by ALKBH5 knockdown in vitro and in vivo, representing that ALKBH5 may be directly correlated with RMRP function to modulate lung adenocarcinoma tumor cells [98] (Table 2).
The circRNAs and m6A modifications/regulators and diseases
In contrast to linear RNAs (such as lncRNAs) with a 5′-UTR cap and a 3′-UTR tail, the special circular covalently closed structure of circRNAs can stabilize the architecture against the damages by exonucleases [103]. Moreover, similar to lncRNAs, circRNAs can also be methylated or demethylated by m6A modifications. It is widely accepted that circRNAs are critical for the occurrence, development, and prognosis of general diseases, such as cancers and immune disorders. For mechanisms, the functions of circRNAs, including particularly sponging and recruiting miRNAs [104], competitively binding to gene sites, and upregulating or downregulating and transcribing protein or mRNA expression, can mediate essential cellular processes, such as proliferation, differentiation, apoptosis, and metabolism [10, 105] (Fig. 3).
In fact, little is known about the mechanisms of the relations of m6A-circRNAs in cancers and diseases. In HCC cells, METTL3 or METTL14 silencing decreases the level of m6A, thus inactivating circRNA-SORE. The m6A modification enhances the stability of circRNA-SORE, a sponge of miR-103a-2-5p and miR-660-3p to isolate them. After that, the Wnt/β-catenin pathway can be activated competitively, hence inducing the drug fast of sorafenib in HCC cells [106]. CircNSUN2 expression is remarkably higher in tissues of CRC patients with liver metastasis (LM) than in original CRC tissues. Furthermore, the invasion and metastasis of CRC cells can be accelerated by the overexpression of circNSUN2, the m6A modification of circNSUN2, and the formation of a circNSUN2/IGF2BP2/HMGA2 RNA-protein ternary complex [107]. For patients with acute coronary syndrome (ACS), the levels of m6A-modified circ_0029589 and METTL3 are markedly increased by upregulating IFN regulatory factor-1 (IRF-1), which can cause circ_0029589 inhibition. Thus, macrophage pyroptosis and inflammation can be facilitated by IRF-1 in ACS and atherosclerosis (AS) [108]. CircFOREIGN exhibits the functions of producing antibodies and inducing anti-tumor immunity and antigen-specific T cell activation. However, due to the isolation of m6A-circRNA with YTHDF2, m6A modifications can restrict innate immunity in vivo [109]. Since circNDUFB2 upregulation can suppress the tumor growth and metastasis of NSCLC cells, circNDUFB2 is downregulated in NSCLC tissues and is inversely related to the malignant characteristics of NSCLC. As a framework, circNDUFB2 can facilitate the interplay between TRIM25 and IGF2BP1/2/3, which demonstrates a vigorous influence on tumor progression and metastasis [110]. CircDLC1 is negatively manipulated by KIAA1429, which shows a low level in HCC tissues and is associated with an obviously favorable prognosis [111] (Table 3).
Conclusion and further vision
In recent years, epigenetic modifications of RNA molecules have increased contemplation with an avalanche of publicity. As multiple key bridges of biological actions, RNA mediates the architecture of whole chromosomes and the functions of transcription, translation, enhancement, and inhibition. Compared with short RNAs, such as miRNAs, lncRNAs are widely regarded as RNA polymerase II (Pol II)-transcribed molecules with different types and dimensions >200 nt. Accumulating studies unveil that lncRNAs have a series of functions of mediating transcriptional and post-transcriptional gene expression in a wide range of mechanisms and diseases [5]. Meanwhile, the majority of circRNAs are ncRNAs, some of which in the cytoplasm can show the potential capability of peptide translation [11].
In a large number of primary and transformed cells, the dynamic reversibility of m6A can occur in profuse locations of mRNAs, lncRNAs, and circRNAs, even though m6A is not directly linked to the progression of tumors or diseases [18]. With m6A-seq, MeRIP-Seq, SCARLET, and innovatively developed detection methods, such as m6A-SEAL and m6A-label-seq, m6A-methylated RNA pieces are always recognized and m6A is noticeably expressed in the 3′-UTR near mRNA stop-codons and long internal exons in different cell/tissue special sites [24]. Thus, biological actions result in the particular transcriptome topology of m6A. With the supplementary modification of m6A, the functions of lncRNAs and circRNAs encompass an abundant diversity of significant biological phenomena, including the imprinting of genomes [4], the sponging or stabilization of miRNAs or proteins, and the upregulation or downregulation of related genes. Furthermore, ncRNAs can feedback to afflict the effect of m6A modification, then complicating the outcomes of pathophysiological activities in vitro and in vivo.
Regardless of how many DNAs, RNAs, proteins, and their complexes are involved, there still are only four steps required from genome to proteome: RNA production, RNA degradation, protein production, and protein degradation. Unfortunately, because of the limits of methods and other factors, we still only reveal the tip of the iceberg in the physiological and pathological activities of cells and organisms. There are a bunch of ncRNAs whose functions and interactions modulated by m6A modifications need more characterization studies and detections to be recognized, which may be the focus of future studies. In summary, with the deeper investigations into lncRNAs, circRNAs, and m6A, there may be some novel therapies, effective strategies, or sensitive detection methods to cure or screen for cancers, especially in combination with immunotherapies.
References
Berretta J, Morillon A. Pervasive transcription constitutes a new level of eukaryotic genome regulation. EMBO Rep. 2009;10:973–82.
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, et al. Landscape of transcription in human cells. Nature. 2012;489:101–8.
Keihani S, Kluever V, Bonn S, Gärtner A, Gomes LC, Kügler S, et al. The long noncoding RNA neuroLNC regulates presynaptic activity by interacting with the neurodegeneration-associated protein TDP-43. Sci Adv. 2019;5:eaay2670.
Robinson EK, Covarrubias S, Carpenter S. The how and why of lncRNA function: an innate immune perspective. Biochim Biophys Acta Gene Regul Mech. 2020;1863:194419.
Quinn JJ, Chang HY. Unique features of long non-coding RNA biogenesis and function. Nat Rev Genet. 2016;17:47–62.
Carlevaro-Fita J, Lanzós A, Hong C, Mas-Ponte D, Pedersen JS, Johnson R. Cancer LncRNA Census reveals evidence for deep functional conservation of long noncoding RNAs in tumorigenesis. Commun Biol. 2020;3:56.
Jacob R, Zander S, Gutschner T. The dark side of the epitranscriptome: chemical modifications in long non-coding RNAs. Int J Mol Sci. 2017;18:2387.
Liu N, Pan T. RNA epigenetics. Transl Res. 2015;165:28–35.
Ferrè F, Colantoni A, Helmer-Citterich M. Revealing protein-lncRNA interaction. Brief Bioinform. 2016;17:106–16.
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, et al. Extensive translation of circular RNAs driven by N(6)-methyladenosine. Cell Res. 2017;27:626–41.
Das A, Gorospe M, Panda AC. The coding potential of circRNAs. Aging (Albany NY). 2018;10:2228–9.
Zhang L, Hou C, Chen C, Guo Y, Yuan W, Yin D, et al. The role of N(6)-methyladenosine (m(6)A) modification in the regulation of circRNAs. Mol Cancer. 2020;19:105.
Wu J, Qi X, Liu L, Hu X, Liu J, Yang J, et al. Emerging epigenetic regulation of circular RNAs in human cancer. Mol Ther Nucleic Acids. 2019;16:589–96.
Boccaletto P, Machnicka MA, Purta E, Piatkowski P, Baginski B, Wirecki TK, et al. MODOMICS: a database of RNA modification pathways. 2017 update. Nucleic Acids Res. 2018;46:D303–d307.
Gong J, Shao D, Xu K, Lu Z, Lu ZJ, Yang YT, et al. RISE: a database of RNA interactome from sequencing experiments. Nucleic Acids Res. 2018;46:D194–201.
Zhao W, Cui Y, Liu L, Ma X, Qi X, Wang Y, et al. METTL3 facilitates oral squamous cell carcinoma tumorigenesis by enhancing c-Myc stability via YTHDF1-mediated m(6)A modification. Mol Ther Nucleic Acids. 2020;20:1–12.
Yi YC, Chen XY, Zhang J, Zhu JS. Novel insights into the interplay between m(6)A modification and noncoding RNAs in cancer. Mol Cancer. 2020;19:121.
Lan Q, Liu PY, Haase J, Hüttelmaier S, Liu T, Bell JL. The critical role of RNA m(6)A methylation in cancer. Cancer Res. 2019;79:1285–92.
Alarcón CR, Lee H, Goodarzi H, Halberg N, Tavazoie SF. N6-methyladenosine marks primary microRNAs for processing. Nature. 2015;519:482–5.
Dominissini D, Moshitch-Moshkovitz S, Schwartz S, Salmon-Divon M, Ungar L, Osenberg S, et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature. 2012;485:201–6.
Li T, Hu PS, Zuo Z, Lin JF, Li X, Wu QN, et al. METTL3 facilitates tumor progression via an m(6)A-IGF2BP2-dependent mechanism in colorectal carcinoma. Mol Cancer. 2019;18:112.
Berulava T, Rahmann S, Rademacher K, Klein-Hitpass L, Horsthemke B. N6-adenosine methylation in MiRNAs. PLoS One. 2015;10:e0118438.
Wang X, Feng J, Xue Y, Guan Z, Zhang D, Liu Z, et al. Corrigendum: structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature. 2017;542:260.
Kane SE, Beemon K. Precise localization of m6A in Rous sarcoma virus RNA reveals clustering of methylation sites: implications for RNA processing. Mol Cell Biol. 1985;5:2298–306.
Wang X, Lu Z, Gomez A, Hon GC, Yue Y, Han D, et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature. 2014;505:117–20.
Yue Y, Liu J, Cui X, Cao J, Luo G, Liu J, et al. VIRMA mediates preferential m(6)A mRNA methylation in 3’UTR and near stop codon and associates with alternative polyadenylation. Cell Disco. 2018;4:10.
Meyer KD, Saletore Y, Zumbo P, Elemento O, Mason CE, Jaffrey SR. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell. 2012;149:1635–46.
Wu Y, Yang X, Chen Z, Tian L, Jiang G, Chen F, et al. m(6)A-induced lncRNA RP11 triggers the dissemination of colorectal cancer cells via upregulation of Zeb1. Mol Cancer. 2019;18:87.
Wang X, Feng J, Xue Y, Guan Z, Zhang D, Liu Z, et al. Structural basis of N(6)-adenosine methylation by the METTL3-METTL14 complex. Nature. 2016;534:575–8.
Choe J, Lin S, Zhang W, Liu Q, Wang L, Ramirez-Moya J, et al. mRNA circularization by METTL3-eIF3h enhances translation and promotes oncogenesis. Nature. 2018;561:556–60.
Knuckles P, Lence T, Haussmann IU, Jacob D, Kreim N, Carl SH, et al. Zc3h13/Flacc is required for adenosine methylation by bridging the mRNA-binding factor Rbm15/Spenito to the m(6)A machinery component Wtap/Fl(2)d. Genes Dev. 2018;32:415–29.
Warda AS, Kretschmer J, Hackert P, Lenz C, Urlaub H, Höbartner C, et al. Human METTL16 is a N(6)-methyladenosine (m(6)A) methyltransferase that targets pre-mRNAs and various non-coding RNAs. EMBO Rep. 2017;18:2004–14.
Nance DJ, Satterwhite ER, Bhaskar B, Misra S, Carraway KR, Mansfield KD. Characterization of METTL16 as a cytoplasmic RNA binding protein. PLoS One. 2020;15:e0227647.
van Tran N, Ernst FGM, Hawley BR, Zorbas C, Ulryck N, Hackert P, et al. The human 18S rRNA m6A methyltransferase METTL5 is stabilized by TRMT112. Nucleic Acids Res. 2019;47:7719–33.
Ren W, Lu J, Huang M, Gao L, Li D, Wang GG, et al. Structure and regulation of ZCCHC4 in m(6)A-methylation of 28S rRNA. Nat Commun. 2019;10:5042.
Ignatova VV, Stolz P, Kaiser S, Gustafsson TH, Lastres PR, Sanz-Moreno A, et al. The rRNA m(6)A methyltransferase METTL5 is involved in pluripotency and developmental programs. Genes Dev. 2020;34:715–29.
Ma H, Wang X, Cai J, Dai Q, Natchiar SK, Lv R, et al. N(6-)Methyladenosine methyltransferase ZCCHC4 mediates ribosomal RNA methylation. Nat Chem Biol. 2019;15:88–94.
Al-Serri A, Alroughani R, Al-Temaimi RA. The FTO gene polymorphism rs9939609 is associated with obesity and disability in multiple sclerosis patients. Sci Rep. 2019;9:19071.
Guo X, Li K, Jiang W, Hu Y, Xiao W, Huang Y, et al. RNA demethylase ALKBH5 prevents pancreatic cancer progression by posttranscriptional activation of PER1 in an m6A-YTHDF2-dependent manner. Mol Cancer. 2020;19:91.
Chen Z, Qi M, Shen B, Luo G, Wu Y, Li J, et al. Transfer RNA demethylase ALKBH3 promotes cancer progression via induction of tRNA-derived small RNAs. Nucleic Acids Res. 2019;47:2533–45.
Mathiyalagan P, Adamiak M, Mayourian J, Sassi Y, Liang Y, Agarwal N, et al. FTO-dependent N(6)-methyladenosine regulates cardiac function during remodeling and repair. Circulation. 2019;139:518–32.
Deng X, Su R, Stanford S, Chen J. Critical enzymatic functions of FTO in obesity and cancer. Front Endocrinol (Lausanne). 2018;9:396.
Wei J, Liu F, Lu Z, Fei Q, Ai Y, He PC, et al. Differential m(6)A, m(6)A(m), and m(1)A demethylation mediated by FTO in the cell nucleus and cytoplasm. Mol Cell. 2018;71:973–.e975.
Zhu H, Gan X, Jiang X, Diao S, Wu H, Hu J. ALKBH5 inhibited autophagy of epithelial ovarian cancer through miR-7 and BCL-2. J Exp Clin Cancer Res. 2019;38:163.
Dango S, Mosammaparast N, Sowa ME, Xiong LJ, Wu F, Park K, et al. DNA unwinding by ASCC3 helicase is coupled to ALKBH3-dependent DNA alkylation repair and cancer cell proliferation. Mol Cell. 2011;44:373–84.
Balacco DL, Soller M. The m(6)A writer: rise of a machine for growing tasks. Biochemistry. 2019;58:363–78.
Shishkin SS, Kovalev LI, Pashintseva NV, Kovaleva MA, Lisitskaya K. Heterogeneous nuclear ribonucleoproteins involved in the functioning of telomeres in malignant cells. Int J Mol Sci. 2019;20:745.
Zhang J, Bai R, Li M, Ye H, Wu C, Che X, et al. Excessive miR-25-3p maturation via N(6)-methyladenosine stimulated by cigarette smoke promotes pancreatic cancer progression. Nat Commun. 2019;10:1858.
Liao S, Sun H, Xu C. YTH domain: a family of N(6)-methyladenosine (m(6)A) readers. Genomics Proteom Bioinforma. 2018;16:99–107.
Zhu S, Wang JZ, Chen D, He YT, Meng N, Chen M, et al. An oncopeptide regulates m(6)A recognition by the m(6)A reader IGF2BP1 and tumorigenesis. Nat Commun. 2020;11:1685.
Xiao W, Adhikari S, Dahal U, Chen YS, Hao YJ, Sun BF, et al. Nuclear m(6)A reader YTHDC1 regulates mRNA splicing. Mol Cell. 2016;61:507–19.
Hsu PJ, Zhu Y, Ma H, Guo Y, Shi X, Liu Y, et al. Ythdc2 is an N(6)-methyladenosine binding protein that regulates mammalian spermatogenesis. Cell Res. 2017;27:1115–27.
Alarcón CR, Goodarzi H, Lee H, Liu X, Tavazoie S, Tavazoie SF. HNRNPA2B1 is a mediator of m(6)A-dependent nuclear RNA processing events. Cell. 2015;162:1299–308.
Meyer KD, Patil DP, Zhou J, Zinoviev A, Skabkin MA, Elemento O, et al. 5' UTR m(6)A promotes cap-independent translation. Cell. 2015;163:999–1010.
Huang H, Weng H, Sun W, Qin X, Shi H, Wu H, et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol. 2018;20:285–95.
Wu R, Li A, Sun B, Sun JG, Zhang J, Zhang T, et al. A novel m(6)A reader Prrc2a controls oligodendroglial specification and myelination. Cell Res. 2019;29:23–41.
Nagarajan A, Janostiak R, Wajapeyee N. Dot blot analysis for measuring global N(6)-methyladenosine modification of RNA. Methods Mol Biol. 2019;1870:263–71.
Zhang W, Yu G, Zhang Y, Tang F, Lv J, Tian G, et al. Quantitative Dot Blot (QDB) as a universal platform for absolute quantification of tissue biomarkers. Anal Biochem. 2019;576:42–47.
Chen Y, Peng C, Chen J, Chen D, Yang B, He B, et al. WTAP facilitates progression of hepatocellular carcinoma via m6A-HuR-dependent epigenetic silencing of ETS1. Mol Cancer. 2019;18:127.
Yin H, Wang H, Jiang W, Zhou Y, Ai S. Electrochemical immunosensor for N6-methyladenosine detection in human cell lines based on biotin-streptavidin system and silver-SiO(2) signal amplification. Biosens Bioelectron. 2017;90:494–500.
Molinie B, Wang J, Lim KS, Hillebrand R, Lu ZX. m(6)A-LAIC-seq reveals the census and complexity of the m(6)A epitranscriptome. Nat Methods. 2016;13:692–8.
Wang Y, Xiao Y, Dong S, Yu Q, Jia G. Antibody-free enzyme-assisted chemical approach for detection of N(6)-methyladenosine. Nat Chem Biol. 2020;16:896–903.
Liu N, Parisien M, Dai Q, Zheng G, He C, Pan T. Probing N6-methyladenosine RNA modification status at single nucleotide resolution in mRNA and long noncoding RNA. Rna. 2013;19:1848–56.
Liu N, Pan T. Probing N6-methyladenosine (m6A) RNA modification in total RNA with SCARLET. Methods Mol Biol. 2016;1358:285–92.
Ule J, Jensen KB, Ruggiu M, Mele A, Ule A, Darnell RB. CLIP identifies Nova-regulated RNA networks in the brain. Science 2003;302:1212–5.
Kennedy EM, Bogerd HP, Kornepati AV, Kang D, Ghoshal D, Marshall JB, et al. Posttranscriptional m(6)A editing of HIV-1 mRNAs enhances viral gene expression. Cell Host Microbe. 2016;19:675–85.
Nakamura S, Ishino K, Fujimoto K. Photochemical RNA editing of C to U by using ultrafast reversible RNA photo-crosslinking in DNA/RNA duplexes. Chembiochem. 2020;21:3067–70.
Miyazaki R, Akiyama Y, Mori H. A photo-cross-linking approach to monitor protein dynamics in living cells. Biochim Biophys Acta Gen Subj. 2020;1864:129317.
Chen K, Lu Z, Wang X, Fu Y, Luo GZ, Liu N, et al. High-resolution N(6)-methyladenosine (m(6) A) map using photo-crosslinking-assisted m(6) A sequencing. Angew Chem Int Ed Engl. 2015;54:1587–90.
Linder B, Grozhik AV, Olarerin-George AO, Meydan C, Mason CE, Jaffrey SR. Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods. 2015;12:767–72.
Arguello AE, DeLiberto AN, Kleiner RE. RNA chemical proteomics reveals the N(6)-methyladenosine (m(6)A)-regulated protein-RNA interactome. J Am Chem Soc. 2017;139:17249–52.
Garcia-Campos MA, Edelheit S, Toth U, Safra M, Shachar R, Viukov S, et al. Deciphering the “m(6)A code” via antibody-independent quantitative profiling. Cell. 2019;178:731–747.e716.
Zhang Z, Chen LQ, Zhao YL, Yang CG, Roundtree IA, Zhang Z. Single-base mapping of m(6)A by an antibody-independent method. Sci Adv. 2019;5:eaax0250.
Imanishi M, Tsuji S, Suda A, Futaki S. Detection of N(6)-methyladenosine based on the methyl-sensitivity of MazF RNA endonuclease. Chem Commun (Camb). 2017;53:12930–3.
Meyer KD. DART-seq: an antibody-free method for global m(6)A detection. Nat Methods. 2019;16:1275–80.
Golovina AY, Dzama MM, Petriukov KS, Zatsepin TS, Sergiev PV, Bogdanov AA, et al. Method for site-specific detection of m6A nucleoside presence in RNA based on high-resolution melting (HRM) analysis. Nucleic Acids Res. 2014;42:e27.
Shu X, Cao J, Cheng M, Xiang S, Gao M, Li T, et al. A metabolic labeling method detects m(6)A transcriptome-wide at single base resolution. Nat Chem Biol. 2020;16:887–95.
Liu H, Xu Y, Yao B, Sui T, Lai L, Li Z. A novel N6-methyladenosine (m6A)-dependent fate decision for the lncRNA THOR. Cell Death Dis. 2020;11:613.
Hosono Y, Niknafs YS, Prensner JR, Iyer MK, Dhanasekaran SM, Mehra R, et al. Oncogenic role of THOR, a conserved cancer/testis long non-coding RNA. Cell. 2017;171:1559–.e1520.
Chen W, Chen M, Xu Y, Chen X, Zhou P, Zhao X, et al. Long non-coding RNA THOR promotes human osteosarcoma cell growth in vitro and in vivo. Biochem Biophys Res Commun. 2018;499:913–9.
Ye XT, Huang H, Huang WP, Hu WL. LncRNA THOR promotes human renal cell carcinoma cell growth. Biochem Biophys Res Commun. 2018;501:661–7.
Jin D, Guo J. m(6)A mRNA methylation initiated by METTL3 directly promotes YAP translation and increases YAP activity by regulating the MALAT1-miR-1914-3p-YAP axis to induce NSCLC drug resistance and metastasis. J Hematol Oncol. 2019;12:135.
Liu P, Zhang B, Chen Z, He Y, Du Y, Liu Y, et al. m(6)A-induced lncRNA MALAT1 aggravates renal fibrogenesis in obstructive nephropathy through the miR-145/FAK pathway. Aging (Albany NY). 2020;12:5280–99.
Ye M, Dong S, Hou H, Zhang T, Shen M. Oncogenic role of long noncoding RNAMALAT1 in thyroid cancer progression through regulation of the miR-204/IGF2BP2/m6A-MYC signaling. Mol Ther Nucleic Acids. 2021;23:1–12.
Qu T, Mou Y, Dai J, Zhang X, Li M, Gu S, et al. Changes and relationship of N(6)-methyladenosine modification and long non-coding RNAs in oxidative damage induced by cadmium in pancreatic β-cells. Toxicol Lett. 2021;343:56–66.
Wen S, Wei Y, Zen C, Xiong W, Niu Y, Zhao Y. Long non-coding RNA NEAT1 promotes bone metastasis of prostate cancer through N6-methyladenosine. Mol Cancer. 2020;19:171.
Barros-Silva D, Lobo J, Guimarães-Teixeira C, Carneiro I, Oliveira J, Martens-Uzunova ES, et al. VIRMA-dependent N6-methyladenosine modifications regulate the expression of long non-coding RNAs CCAT1 and CCAT2 in prostate cancer. Cancers (Basel). 2020;12:771.
Ni W, Yao S, Zhou Y, Liu Y, Huang P, Zhou A, et al. Long noncoding RNA GAS5 inhibits progression of colorectal cancer by interacting with and triggering YAP phosphorylation and degradation and is negatively regulated by the m(6)A reader YTHDF3. Mol Cancer. 2019;18:143.
Zuo X, Chen Z, Gao W, Zhang Y, Wang J, Wang J, et al. M6A-mediated upregulation of LINC00958 increases lipogenesis and acts as a nanotherapeutic target in hepatocellular carcinoma. J Hematol Oncol. 2020;13:5.
He Y, Hu H, Wang Y, Yuan H, Lu Z, Wu P, et al. ALKBH5 inhibits pancreatic cancer motility by decreasing long non-coding RNA KCNK15-AS1 methylation. Cell Physiol Biochem. 2018;48:838–46.
Wang J, Ding W, Xu Y, Tao E, Mo M, Xu W, et al. Long non-coding RNA RHPN1-AS1 promotes tumorigenesis and metastasis of ovarian cancer by acting as a ceRNA against miR-596 and upregulating LETM1. Aging (Albany NY). 2020;12:4558–72.
Wang X, Zhang J, Wang Y. Long noncoding RNA GAS5-AS1 suppresses growth and metastasis of cervical cancer by increasing GAS5 stability. Am J Transl Res. 2019;11:4909–21.
Ban Y, Tan P, Cai J, Li J, Hu M, Zhou Y, et al. LNCAROD is stabilized by m6A methylation and promotes cancer progression via forming a ternary complex with HSPA1A and YBX1 in head and neck squamous cell carcinoma. Mol Oncol. 2020;14:1282–96.
Liu J, Zhang X, Chen K, Cheng Y, Liu S, Xia M, et al. CCR7 chemokine receptor-inducible lnc-Dpf3 restrains dendritic cell migration by inhibiting HIF-1α-mediated glycolysis. Immunity. 2019;50:600–615.e615.
Xue L, Li J, Lin Y, Liu D, Yang Q, Jian J, et al. m(6) A transferase METTL3-induced lncRNA ABHD11-AS1 promotes the Warburg effect of non-small-cell lung cancer. J Cell Physiol. 2021;236:2649–58.
Zhu P, He F, Hou Y, Tu G, Li Q, Jin T, et al. A novel hypoxic long noncoding RNA KB-1980E6.3 maintains breast cancer stem cell stemness via interacting with IGF2BP1 to facilitate c-Myc mRNA stability. Oncogene. 2021;40:1609–27.
Wu J, Pang R, Li M, Chen B, Huang J, Zhu Y. m6A-induced LncRNA MEG3 suppresses the proliferation, migration and invasion of hepatocellular carcinoma cell through miR-544b/BTG2 signaling. Onco Targets Ther. 2021;14:3745–55.
Yu H, Zhang Z. ALKBH5-mediated m6A demethylation of lncRNA RMRP plays an oncogenic role in lung adenocarcinoma. Mamm Genome. 2021;32:195–203.
Wang X, Liu C, Zhang S, Yan H, Zhang L, Jiang A, et al. N(6)-methyladenosine modification of MALAT1 promotes metastasis via reshaping nuclear speckles. Dev Cell. 2021;56:702–715.e708.
Patil DP, Chen CK, Pickering BF, Chow A, Jackson C, Guttman M, et al. m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature. 2016;537:369–73.
Yang X, Zhang S, He C, Xue P, Zhang L, He Z, et al. METTL14 suppresses proliferation and metastasis of colorectal cancer by down-regulating oncogenic long non-coding RNA XIST. Mol Cancer. 2020;19:46.
Lan T, Li H, Zhang D, Xu L, Liu H, Hao X, et al. KIAA1429 contributes to liver cancer progression through N6-methyladenosine-dependent post-transcriptional modification of GATA3. Mol Cancer. 2019;18:186.
Paramasivam A, Vijayashree Priyadharsini J. Novel insights into m6A modification in circular RNA and implications for immunity. Cell Mol Immunol. 2020;17:668–9.
Zhao W, Cui Y, Liu L, Qi X, Liu J, Ma S, et al. Splicing factor derived circular RNA circUHRF1 accelerates oral squamous cell carcinoma tumorigenesis via feedback loop. Cell Death Differ. 2020;27:919–33.
Prats AC, David F, Diallo LH, Roussel E, Tatin F, Garmy-Susini B, et al. Circular RNA, the key for translation. Int J Mol Sci. 2020;21:8591.
Xu J, Wan Z, Tang M, Lin Z, Jiang S, Ji L, et al. N(6)-methyladenosine-modified CircRNA-SORE sustains sorafenib resistance in hepatocellular carcinoma by regulating β-catenin signaling. Mol Cancer. 2020;19:163.
Chen RX, Chen X, Xia LP, Zhang JX, Pan ZZ, Ma XD, et al. N(6)-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis. Nat Commun. 2019;10:4695.
Guo M, Yan R, Ji Q, Yao H, Sun M, Duan L, et al. IFN regulatory Factor-1 induced macrophage pyroptosis by modulating m6A modification of circ_0029589 in patients with acute coronary syndrome. Int Immunopharmacol. 2020;86:106800.
Chen YG, Chen R, Ahmad S, Verma R, Kasturi SP, Amaya L, et al. N6-Methyladenosine modification controls circular RNA immunity. Mol Cell. 2019;76:96–109.e109.
Li B, Zhu L, Lu C, Wang C, Wang H, Jin H, et al. circNDUFB2 inhibits non-small cell lung cancer progression via destabilizing IGF2BPs and activating anti-tumor immunity. Nat Commun. 2021;12:295.
Liu H, Lan T, Li H, Xu L, Chen X, Liao H, et al. Circular RNA circDLC1 inhibits MMP1-mediated liver cancer progression via interaction with HuR. Theranostics. 2021;11:1396–411.
Funding
This work was supported by National Natural Science Foundation of China (No. 82002889), Tianjin Medical University Stomatological Hospital Foundation (No: 2020YKY01).
Author information
Authors and Affiliations
Contributions
JL and XW wrote the main manuscript text. LZ and WZ prepared Fig. 1–3. XW and WZ were responsible for the designing and funding. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent for publication
All the authors agree with the publication.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, J., Zhao, W., Zhang, L. et al. The emerging roles of N6-methyladenosine (m6A)-modified long non-coding RNAs in human cancers. Cell Death Discov. 8, 255 (2022). https://doi.org/10.1038/s41420-022-01050-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-022-01050-0
- Springer Nature Limited
This article is cited by
-
The roles of m6A methylation in cervical cancer: functions, molecular mechanisms, and clinical applications
Cell Death & Disease (2023)