Abstract
Cancer-associated fibroblasts (CAFs) play crucial roles in mediating tumor growth and metastasis via transferring exosomes to neighboring cells, whereas the mechanisms by which CAFs regulate the tumorgenesis of prostate cancer (PC) remain largely unknown. In this study, CAFs and normal fibroblasts (NFs) were isolated from PC tissues and adjacent normal tissues, respectively. Exosomes (NFs-Exo and CAFs-Exo) were then isolated from the supernatant of NFs and CAFs. Next, the differentially expressed miRNAs (DEMs) between NFs-Exo and CAFs-Exo were identified using RNA-sequencing. Cell viability, migration and invasion were detected with CCK-8 and Transwell assays. Protein expression was measured with western blot. We found that CAFs-Exo remarkably enhanced PC cell migration, invasion, stemness, epithelial-mesenchymal transition (EMT) and metastasis. Significantly, miR-1290 level was upregulated in CAFs-Exo compared to NFs-Exo. In addition, CAFs could transfer exosomes to PC cells, resulting in a marked increase of miR-1290 level in cells. Moreover, exosomal miR-1290 could inhibit GSK3β/β-catenin signaling by binding with the downstream target GSK3β mRNA. Meanwhile, miR-1290 antagomir notably reversed the effects of CAFs-Exo on PC cells through activating GSK3β/β-catenin signaling. Collectively, exosomal miR-1290 from CAFs could promote PC cell growth and metastasis via inhibiting GSK3β/β-catenin signaling, suggesting that miR-1290 may serve as potential therapeutic target for the treatment of PC.
Similar content being viewed by others
Introduction
Prostate cancer (PC) is the most common non-cutaneous cancer diagnosed in men with an estimated new 1,600,000 cases annually [1, 2]. Although the therapies for the treatment of PC including androgen deprivation therapy (ADT) surgical prostatectomy, radiotherapy and hormone therapy improve overall survival outcomes, severe side effects remain inevitable [3,4,5]. Therefore, identifying novel therapeutic targets for PC may promote the development of approaches for early detection and therapies.
Increasing evidences indicated that the cellular interaction within the tumor microenvironment (TME) exhibits key roles in reprograming tumor initiation, growth and metastasis [6, 7]. CAFs are the main stromal cells in the TME that facilitate the progression of human cancers via interacting with cancer cells [8, 9]. Recently, evidences have shown that CAFs could facilitate prostate tumor growth and invasion [10,11,12]. However, the mechanisms by which CAFs regulate the tumorgenesis of prostate tumor remain largely unknown.
CAFs could promote tumor growth and metastasis through communicating with cancer cells within TME [13]. In addition, the crosstalk between CAFs and cancer cells are often mediated by extra-cellular signals including exosomes [14]. Exosomes are 40–130 nm nano-sized extracellular vesicles that are secreted by cells [15]. Additionally, exosomes can act as mediators for intercellular crosstalk by the delivery of biomolecules, such as proteins, microRNAs (miRNAs), mRNA, DNA) and lipids [16, 17]. MiRNAs are able to negatively mediate gene expression [18, 19]. Furthermore, miRNAs are highly enriched in exosomes, and exosomes can transfer functional miRNA molecules from CAFs to cancer cells [20,21,22].
In the present study, our results indicated that miR-1290 level was significantly elevated in CAFs-Exo compared to NFs-Exo. In addition, CAFs led to significant increases in PC cell migration, invasion, stemness and metastasis through transferring exosomal miR-1290. The present findings demonstrating that miR-1290 may serve as potential therapeutic target for PC treatment.
Materials and methods
Isolation of primary human fibroblasts
PC tissues and matched adjacent non-cancerous tissues (10 pairs) were collected from the patients with PC who underwent surgery. This study was approved by the Ethics Committee of Peking University Cancer Hospital & Institute, and each participant gave the written informed consent. According to previously report, primary human CAFs and NFs were isolated from tumor tissues and matched non-cancerous tissues respectively [23]. Next, the morphology of CAFs and NFs were captured with a light microscope.
Cell culture and cell transfection
Human PC cell lines PC3, 22RV1 and LNCaP were obtained from American Type Culture Collection. These cell lines were authenticated by STR profiling and tested negative for mycoplasma contamination. PC cells were maintained in F-12K medium or in RPMI-1640 (Thermo Fisher Scientific) containing 10% FBS in a humidified 5% CO2 atmosphere at 37 °C. In addition, CAFs and NFs were maintained in DMEM/F12 medium containing 10% FBS at 37 °C with 5% CO2.
MiR-1290 agomir (50 nM), miR-1290 antagomir (anti-miR-1290, 100 nM) and negative control (NC) was purchased from RIBOBIO. Meanwhile, GSK3β was ligated into the pcDNA3.1 vector to obtain pcDNA3.1-GSK3β (GSK3β-OE) plasmids (GenePharma). Next, NC, miR-1290 agomir, anti-miR-1290 or GSK3β-OE plasmids were transfected into PC cells using Lipofectamine 2000.
Immunofluorescence (IF) assay
CAFs and NFs were probed with primary antibodies against vimentin (Abcam) and α-SMA (Abcam) overnight at 4 °C. Next, cells were incubated with secondary antibody conjugated with fluorescent Alexa Fluor® 594 (Abcam) in darkness. Finally, cells were captured with a fluorescence microscope. Nuclei were stained with DAPI for 15 min.
Exosome isolation and identification
CAFs and NFs were cultured in DMEM/F12 medium containing 10% FBS until about 80% confluent at 37 °C. Then, medium was replaced with DMEM/F12 medium without FBS. Next, exosomes were collected from the supernatant of CAFs and NFs using ultracentrifugation. Briefly, culture supernatant was centrifuged at 300 × g for 10 min, at 2000 x g for 10 min, and at 10,000 g for 30 min. Culture supernatant was then centrifuged at 100,00 x g for 70 min. The precipitates were collected and washed with sterilized PBS (ASPEN). Later on, the solution was centrifuged again at 100,00 × g for 70 min. The final pellet was resuspended in PBS (ASPEN). Next, exosomes were treated with 2.5% glutaraldehyde and then loaded on a carbon-coated copper grid. After that, 1% phosphotungstic acid was applied to stain the grid. Next, transmission electron microscope (TEM) was applied to examine the morphologies of the samples. Size distributions and quantification of exosomes were determined using a ZetaView nanoparticle tracking analyzer (Particle Metrix). PC3 and 22RV1 cells were treated with CAFs-Exo or NFs-Exo (50 μg/ml) for indicated times.
Western blot assay
Proteins were electrophoresed on 10% SDS-PAGE and electroblotted onto a PVDF membrane. Next, the membrane was probed with primary antibodies including anti-TSG101 (cat. no. ab125011, 1:1000), anti-CD9 (cat. no. ab236630, 1:1000), anti-E-cadherin (cat. no. ab231303, 1:1000, anti-N-cadherin (cat. no. ab76011, 1:1000), anti-Vimentin (cat. no. ab92547, 1:1000), anti-CD133 (cat. no. ab216323, 1:1000), anti-OCT4 (cat. no. ab200834, 1:1000), anti-Glycogen synthase kinase-3 beta (GSK3β; cat. no. ab93826, 1:1000), anti-β-catenin (cat. no. ab223075, 1:1000), anti-c-Myc (cat. no. ab32072, 1:1000), anti-cyclin D1 (cat. no. ab16663, 1:1000), anti-Calnexin (cat. no. ab133615, 1:1000) and anti-GAPDH (cat. no. ab8245, 1:1000) overnight at 4 °C. The membrane was then incubated with a secondary antibody for 1 h. Finally, the blots were developed with an enhanced chemiluminescence kit. All antibodies were provided from Abcam. GAPDH was used as an internal control.
Exosome labeling and uptake
NFs-Exo and CAFs-Exo were labeled with PKH26 membrane dye (Sigma). Next, PC3 cells were treated with above exosomes for 24 h, and then photographed by a fluorescence microscope. Phallodin was used to label F-actin of the cytoskeleton, DAPI was used to label nuclei.
RNA-sequencing (RNA-seq)
Total RNA was extracted from CAFs-Exo or NFs-Exo with the TRIpure Total RNA Extraction Reagent (ELK Biotechnology). RNA-seq libraries were prepared from all samples using the NEBNext Ultra Directional RNA Library Prep Kit for Illumina. Next, sequencing was performed by Illumina Hiseq sequencer (Illumina). The DEMs between CAFs-Exo and NFs-Exo were screened by the limma package. DEMs were obtained with |log2 (fold change)| > 1 and adjusted p-value < 0.05.
To predict the possible functions of the target genes of DEMs and the potential pathways they may participate in, gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis were conducted [24].
Cell-counting Kit-8 (CCK-8) assay
PC3 and 22RV1 cells were plated onto 96-well plates overnight. Cells were then treated with CAFs-Exo or NFs-Exo (50 μg/ml) for 48 h at 37 °C. Later on, each well was added with CCK-8 reagent (10 μL, Beyotime), and cells underwent 2 h of incubation at 37 °C. Subsequently, the absorbance of each well was measured by a microplate reader at the wavelength of 450 nm.
Transwell assays
PC3 and 22RV1 cells (100 µL) suspended in serum-free media were added to the 24-well Transwell inserts (Corning). The lower chamber contained the culture medium with 10% FBS. Next, cells on the bottom surface of the chamber were stained with 0.2% crystal violet at 24 h. Later on, the migrated or invaded cells were examined using a microscope. For detecting cell invasion ability, Transwell inserts were pre-coated with matrigel (BD Bioscience).
Real-time quantitative polymerase chain reaction (RT-qPCR) assay
Total RNA was isolated by TRIpure Total RNA Extraction Reagent and transcribed using EntiLink™ 1st Strand cDNA Synthesis Kit. Later on, qPCR was conducted by the EnTurbo™ SYBR Green PCR SuperMix kit and analyzed with the StepOne™ Real-Time PCR System. The level of each miRNA was normalized against U6, using 2–ΔΔCT method. U6 forward, 5’-CTCGCTTCGGCAGCACAT-3’ and reverse, 5’-AACGCTTCACGAATTTGCGT-3’; miR-135b-3p forward, 5’-CCGATGTAGGGCTAAAAGCC-3’ and reverse, 5’-CTCAACTGGTGTCGTGGAGTC-3’; miR-549a-3p forward, 5’-CCGGTGACAACTATGGATGAG-3’ and reverse, 5’-CTCAACTGGTGTCGTGGAGTC-3’; miR-1290 forward, 5’-GGCTGGATTTTTGGATCAGG-3’ and reverse, 5’-CTCAACTGGTGTCGTGGAGTC-3’; miR-224-5p forward, 5’-CAAGTCACTAGTGGTTCCGTTTAG-3’ and reverse, 5’-CTCAACTGGTGTCGTGGAGTC-3’; GSK3β forward, 5’-GCTGCACAGGAAAAACCACC-3’ and reverse, 5’-CCCCCTGGATCTCCCTCAAA-3’. All kits were provided from ELK Biotechnology.
Sphere-forming assay
PC3 and 22RV1 cells were cultured in DMEM/F12 medium containing 4 ng/ml insulin (Sigma), 2% B27 (Thermo Fisher Scientific), 20 ng/ml EGF (Sigma), 10 ng/ml FGF (Sigma). After 14 days of incubation, cells were observed under a light microscopy.
Luciferase reporter and TOPflash reporter assays
The fragment of 3′UTR of GSK3β containing the binding site of miR-1290 was inserted into the pGL6-miR‐based luciferase reporter vector. Next, PC3 cells were co-transfected with well-designed pGL6-miR‐based reporter plasmids, along with a miR-1290 agomir using Lipofectamine 2000 for 48 h. Subsequently, the Dual Luciferase Reporter Assay System (Beyotime) was used to measure luciferase activity in cell lysates. For the TOPflash reporter assay, PC3 cells were co-transfected with TOPflash reporter gene (TCF Reporter Plasmid; Millipore), along with anti-miR-1290. At 48 h post-transfection, the reporter activities were assayed by Dual Luciferase Reporter Assay System (Beyotime). Renilla luciferase activity was used for normalization.
RNA pull-down assay
The biotinylated-miR-1290 probe was incubated with Dynabeads™ M-280 (Thermo Fisher Scientific) at 4 °C for 30 min. PC3 cells were collected, lysed, and sonicated. After that, the biotin-beads complex was added to the lysates and incubated at 4 °C overnight with gentle mixing by vortex. Subsequently, the RNA mix bound to the beads was eluted and extracted with TRIzol reagent (Thermo Fisher Scientific). Later on, RNA samples were reverse-transcribed into cDNA using the EntiLink™ 1st Strand cDNA Synthesis Kit (ELK Biotechnology). Next, PCR was performed using the EntiLink™ PCR Master Mix kit (ELK Biotechnology). Subsequently, PCR products were examined by 1% agarose gel electrophoresis, and then visualized by ultraviolet (UV).
Animal models
Male BALB/c nude mice (18–22 g, 6 weeks old) were obtained from the Charles River. PC3 cells (1 × 107 cells in 200 μL PBS) were injected into the left flank of each mouse subcutaneously. When the tumor reached about 200 mm3, mice were divided into 4 groups (n = 6 per group; the sample size used in animal study is usually based on the experience in previous research) randomly: control, NFs-Exo, CAFs-Exo and CAFs-Exo + anti-miR-1290. After that, NFs-Exo or CAFs-Exo (1 mg/kg) was injected into the tumor twice a week. Meanwhile, 50 nM miR-1290 agomir was injected directly into the tumor twice per week. Meanwhile, tumor volume was calculated as (length x width2)/2. Animals were sacrificed on day 21, and the tumors were isolated and weighted.
Regarding as liver metastasis model, nude mouse was anesthetized with 1% pentobarbital sodium. After that, the spleen of mouse was sub-capsularly injection of PC3 cells (5 × 106 cells). Mice were sacrificed at day 21 post injection and the number of liver metastatic nodules in these mice was determined. This study was approved by the Ethics Committee of Peking University Cancer Hospital & Institute, and animals were maintained following the institutional guidelines. The investigator was blinded to the group allocation.
Immunohistochemistry (IHC) assay
The tumor tissues were embedded in paraffin and cut into 4 μm sections. Later on, tissues were probed with the primary antibody against Ki67 overnight at 4 °C. Next, sections were probed with HRP conjugated anti-rabbit second antibody for 30 min. Subsequently, tissues were visualized with DAB solution and photographed under a light microscope.
Statistical analyses
All data were repeated in triplicate. Data are expressed as the mean ± standard deviation (S.D.). Group comparisons among three or more groups were analyzed by One-way analysis of variance (ANOVA) and Tukey’s test. Differences between tumor tissues and adjacent normal tissues were analyzed using paired Student’s t-test (normal distribution). Data are reported including estimation of variation within each group. A P-value less than 0.05 was considered to have statistical significance.
Results
CAFs-Exo promoted PC cell migration and invasion
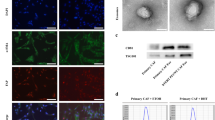
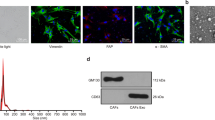
CAFs play key roles in tumor progression through transferring exosomes to neighboring cells [25]. To explore the role of CAFs in PC progression, NFs and CAFs were collected from patients with PC firstly. Then, cell morphologies were observed using a microscope. CAFs and NFs displayed the typical spindle-shape morphology (Fig. 1A). Additionally, CAFs and NFs were positive for vimentin, which was verified by IF assay (Fig. 1B). Meanwhile, α-SMA level was much higher in CAFs than that in NFs (Fig. 1C). Collectively, CAFs and NFs were isolated successfully.
Next, exosomes were isolated from the supernatant of CAFs (CAFs-Exo) and NFs (NFs-Exo), and the isolated vesicles were identified using TEM, nanoparticle tracking analysis (NTA) and western blot analysis. The results indicated that these vesicles (40–130 nm in diameter) were round and cup-shaped (Fig. 2A, B). In addition, exosomal markers TSG101 and CD9 were highly expressed, but exosomal negative marker calnexin was negative expressed in these vesicles. (Fig. 2C and Supplemental Material). These results suggested that the isolated vesicles were exosomes.
A, B Identification of exosomes (CAFs-Exo and NFs-Exo) derived from CAFs and NFs by TEM (Scale bar = 100 nm) and NTA analysis. Yellow arrow points at exosomes. C Western blot analysis of TSG101, CD9 and Calnexin expressions in CAFs-Exo and NFs-Exo. D PC3 cells were treated with PKH26-labeled CAFs-Exo and PKH26-labeled NFs-Exo for 24 h, and the red exosome signal was captured by fluorescence microscopy. Scale bar = 20 μm.
To determine if exosomes could be absorbed by PC3 cells, PKH26-labeled CAFs-Exo or PKH26-labeled NFs-Exo were added into PC3 cells. The results revealed that PKH26 dye was observed in CAFs-Exo- and NFs-Exo-treated PC3 cells, indicating that exosomes released by CAFs and NFs could be internalized by PC3 cells (Fig. 2D).
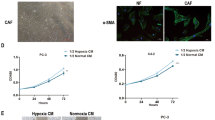
We next to explore the effects of CAFs-Exo on PC cells (PC3 and 22RV1 cells). Compared to NFs-Exo, CAFs-Exo remarkably enhanced PC cell proliferation, migration, and invasion (Fig. 3A–C). Collectively, CAFs-Exo facilitated PC cell migration and invasion.
A PC3 and 22RV1 cells were treated with CAFs-Exo or NFs-Exo (50 μg/ml) for 48 h. CCK-8 assay was used to determine cell viability. B, C PC3 and 22RV1 cells were incubated with CAFs-Exo or NFs-Exo (50 μg/ml) for 24 h. Transwell assays were used to determine cell migratory and invasive abilities. Scale bar = 50 μm. **P < 0.01. Data are presented as mean ± S.D. (n = 3). One-way ANOVA was used for comparing differences among three groups.
Identification of the DEMs between CAFs-Exo and NFs-Exo
As we know, exosomes could shuttle miRNAs to neighboring cells in the TME [26]. To explore the mechanisms how CAF-Exo exerts their pro-tumor effects on PC cells, the DEMs between CAFs-Exo and NFs-Exo was investigated using RNA sequencing assay. As shown in Fig. 4A, B, 7 downregulated miRNAs and 11 upregulated miRNAs were detected in CAF-Exo compared with NFs-Exo. In addition, GO results showed that the target genes of these 18 DEMs were primarily enriched in “regulation of cell communication”, “intracellular” and “cellular process” (Fig. 4C). KEGG pathway analysis revealed that the target genes of DEMs were mainly involved in “microRNAs in cancer” pathway (Fig. 4D).
CAFs secreted exosomal miR-1290 promoted PC cell migration, invasion, EMT, and stemness
With the aim of exploring the DEMs, cell experiments were conducted accordingly. Among these DEMs, miR-1290, miR-135b-5p and miR-224-5p level was notably elevated in CAFs-Exo compared to NFs-Exo (Fig. 5A). In addition, we found miR-1290 level was obviously elevated in PC tissues compared to adjacent normal tissues (Supplementary Fig. 1A). Moreover, high miR-1290 level correlated with shorter overall survival rates in patients with castration-resistant PC [27]. Meanwhile, miR-1290 level was obviously elevated in CAFs compared with that in NFs (Supplementary Fig. 1B). Thus, we focused our attention on miR-1290 in the next experiments, since the level of miR-1290 in CAFs-Exo was much higher that in NFs-Exo (Fig. 5A).
A RT-qPCR analysis of miR-135b-3p, miR-549a-3p, miR-1290, miR-135b-5p and miR-224-5p level in CAFs-Exo and NFs-Exo. B RT-qPCR analysis of miR-1290 level in PC3 and 22RV1 cells incubated with indicated exosomes (50 μg/ml) for 48 h. C RT-qPCR analysis of miR-1290 level in PC3 and 22RV1 cells transfected with anti-miR-1290. D PC3 and 22RV1 cells were treated with NFs-Exo, CAFs-Exo or CAFs-Exo + anti-miR-1290 for 48 h. Cell viability was determined using CCK-8 assay. E, F PC3 and 22RV1 cells were treated with NFs-Exo, CAFs-Exo or CAFs-Exo + anti-miR-1290 for 24 h. Cell migratory and invasive abilities were measured by Transwell assays. Scale bar = 50 μm. **P < 0.01. Data are presented as mean ± S.D.; n = 3. Student’s t-test and one-way ANOVA were used for comparison among the different groups.
Next, the result of RT-qPCR assay showed the level of miR-1290 was notably elevated in PC cells incubated with CAFs-Exo compared to cells treated with NFs-Exo (Fig. 5B). In addition, anti-miR-1290 remarkably lessened miR-1290 level in PC cells (Fig. 5C). Furthermore, CAFs-Exo significantly enhanced PC cell proliferation, migration, and invasion, whereas these phenomena were abolished by anti-miR-1290 (Fig. 5D–F). Moreover, CAFs-Exo remarkably increased the sphere-forming efficiency and notably elevated stemness markers CD133 and OCT4 expressions in PC cells; however, these phenomena were reversed by anti-miR-1290 as well (Fig. 6A–C and Supplemental Material). Meanwhile, CAFs-Exo markedly downregulated E-cadherin level and upregulated N-cadherin and vimentin level in PC cells compared to NFs-Exo, whereas the effects of CAFs-Exo were abolished by anti-miR-1290 (Fig. 6B, C and Supplemental Material). All these data illustrated that CAFs secreted exosomal miR-1290 was able to promote PC cell migration, invasion, EMT and stemness.
A PC3 and 22RV1 cells were treated with NFs-Exo, CAFs-Exo or CAFs-Exo + anti-miR-1290 for 2 weeks. The self-renewal of the PC cells was measured by a sphere-forming assay. Scale bar = 10 μm. B, C PC3 and 22RV1 cells were treated with NFs-Exo, CAFs-Exo or CAFs-Exo + anti-miR-1290 for 48 h. Western blot analysis of E-cadherin, N-cadherin, vimentin, CD133, and OCT4 expression in PC3 and 22RV1 cells. **P < 0.01. Data are presented as mean ± S.D.; n = 3. The difference among three groups was performed by one-way ANOVA.
GSK3β is a direct target of miR-1290 in PC cells
To explore the mechanism by which CAFs secreted exosomal miR-1290 promoted the growth and tumorgenesis in PC, the downstream targets of miR-1290 were predicted by TargetScan [28], miRWalk [29], and DIANA-microT databases [30], as described previously. These databases commonly indicated that GSK3β might be the direct target of miR-1290 (Fig. 7A). In addition, GSK3β has been found to exhibit a key role in the development of human cancers such as breast cancer, colorectal cancer, and PC [31,32,33]. RT-qPCR result showed that GSK3β level was notably downregulated in PC tissues compared to normal tissues (Supplementary Fig. 1C). Moreover, dual-luciferase reporter assay showed that the luciferase activity was lower in PC cells co-transfected with GSK3β‐WT and miR-1290 agomir (Fig. 7B). Furthermore, miR-1290 agomir markedly reduced GSK3β level in PC cells (Fig. 7C and Supplemental Material). Additionally, RNA pull-down assay results revealed that GSK3β could be pulled down by miR-1290 directly (Fig. 7D). Collectively, GSK3β is a direct binding target of miR-1290.
A Sequences of miR-1290 and the potential miR-1290-binding sites at the 3’UTR of GSK3β. B Effect of ectopic miR-1290 expression on the luciferase activity of GSK3β 3’UTR wild type. C PC3 and 22RV1 cells were transfected with agomir-NC or miR-1290 agomir. Western blot analysis of GSK3β expression in PC3 and 22RV1 cells. D RNA pull-down assay was used to explore the interaction between miR-1290 and GSK3β mRNA. E PC3 cells were treated with NFs-Exo, CAFs-Exo or CAFs-Exo + anti-miR-1290 for 48 h. Dual luciferase assay was used to determine TOP reporter activity in PC3 cells. F, G PC3 and 22RV1 cells were treated with NFs-Exo, CAFs-Exo or CAFs-Exo + anti-miR-1290 for 48 h. Western blot analysis of GSK3β, β-catenin, c-Myc and cyclin D1 expression in PC3 and 22RV1 cells. **P < 0.01. Data are presented as mean ± S.D.; n = 3. Student’s t-test and one-way ANOVA were used for comparison among the different groups.
Next, TOPflash reporter assay was used to examine the effect of exosomal miR-1290 on the activity of the Wnt pathway in PC3 cells. The result suggested that CAFs-Exo remarkably increased transcriptional activity of TOP, while anti-miR-1290 could reverse the enhancement of the transcriptional activity of TOP caused by CAFs-Exo (Fig. 7E). Meanwhile, CAFs-Exo significantly elevated β-catenin, c-Myc and Cyclin D1 level and reduced GSK3β level in PC cells compared to NFs-Exo, whereas these changes were reversed by anti-miR-1290 (Fig. 7F, G and Supplemental Material).
Furthermore, the effects of CAFs-Exo on cellular proliferation, migration and invasion were evaluated in an androgen-dependent PC cell line (LNCaP). As shown in Supplementary Fig. 2A, GSK3β level was notably reinforced in LNCaP cells when transfected with GSK3β-OE plasmids. Meanwhile, CAFs-Exo significantly facilitated LNCaP cell proliferation, migration, and invasion (Supplementary Fig. 2B, C). Consistently, anti-miR-1290 abolished the promotion effect of CAFs-Exo on LNCaP cell proliferation. Importantly, the effects of anti-miR-1290 on cells were all abolished when GSK3β was overexpressed (Supplementary Fig. 2B, C). To sum up, CAFs secreted exosomal miR-1290 was able to promote the growth and tumorgenesis of PC cells through inhibiting GSK3β/β-catenin signaling.
CAFs secreted exosomal miR-1290 promoted PC cell growth and metastasis in vivo through downregulation of GSK3β
Finally, we explored the effect of CAFs-Exo on the growth and metastasis of PC in vivo. Upon subcutaneous transplantation of PC3 cells, CAFs-Exo notably increased the tumor volume and weight compared to control or NFs-Exo group, whereas, that effects were significantly inhibited by anti-miR-1290 (Fig. 8A–C). In addition, IHC assay result revealed that the number of Ki67-positive tumor cells was much higher in CAFs-Exo-treated group than that in the control or NFs-Exo group; however, this phenomenon was reversed by anti-miR-1290 (Fig. 8D). Meanwhile, a mouse model of liver metastasis was established by subcapsular injection of the spleen method, and the data indicated that increased liver metastatic nodules were detected in CAFs-Exo group compared to control or NFs-Exo group. Similar to Ki67 stating, the promoting effect of CAFs-Exo on tumor metastasis was significantly inhibited by anti-miR-1290 (Fig. 8E).
A–C Tumor volume and tumor weight of xenograft tumors. D IHC analysis of Ki67 expression in tumor tissues. Scale bar = 50 μm. E The formation of metastasis nodules in liver tissues. F RT-qPCR analysis of miR-1290 level in tumor tissues. G Western blot analysis of GSK3β, β-catenin, c-Myc and cyclin D1 expression in tumor tissues. *P < 0.05; **P < 0.01. Data are presented as mean ± S.D.; n = 3. The significance among multiple groups was determined with one-way ANOVA.
Furthermore, CAFs-Exo remarkably increased miR-1290, β-catenin, c-Myc and Cyclin D1 level and reduced GSK3β level in tumor tissues; however, these phenomena were all reversed by anti-miR-1290 treatment (Fig. 8F, G and Supplemental Material). Taken together, CAFs secreted exosomal miR-1290 significantly promoted PC cell growth and metastasis in vivo through downregulation of GSK3β.
Discussion
CAFs in the TME have emerged as important player in the development of human cancers, including prostate cancer [9, 34, 35]. In addition, crosstalk between CAFs and cancer cells in the TME contributes to tumor invasion and metastasis in various cancers [36,37,38]. Recently, exosomes have been found to emerge as critical mediators for intercellular communication between cancer cells and CAFs through transporting biomolecules including miRNAs [22, 39]. A recent study reported that breast cancer cell-derived exosomes could accelerated the activation of CAFs [40]. Donnarumma et al reported that CAFs-Exo was able to promote the EMT process and breast cancer cell stemness [41]. Hu et al. found that CAFs-Exo could enhance colorectal cancer cell metastasis through enhancing cell stemness and EMT [42]. Yang et al. showed that CAFs-derived exosomal miR-210 was able to promote the migration and invasion of non-small cell lung cancer cells [43]. In the present study, we found that CAFs-Exo notably enhanced PC cell migration and invasion. Additionally, CAFs-Exo could induce the EMT phenotypes in PC cells via upregulation of N-cadherin and vimentin protein expressions. Meanwhile, CAFs-Exo promoted PC cell stemness via upregulation of CD133 and OCT4. All these data indicated that CAFs-Exo was able to enhance PC cell migration, invasion, EMT and stemness. However, the mechanisms by which CAFs-Exo promoted cancer growth and metastasis remain largely unclear. Thus, we focus on investigating the communication between CAFs and PC cells.
Recent studies revealed that exosomes contribute to tumor progression in cancer cells through delivering miRNAs [44, 45]. Thus, RNA-sequencing assay was performed to screen DEMs between NFs-Exo and CAFs-Exo. Our results indicated that miR-1290 level was highly expressed in CAFs-Exo compared to NFs-Exo. MiR-1290 was found to be abnormally upregulated in human cancers, such as non-small cell lung cancer, oral squamous cell carcinoma and gastric cancer [46,47,48]. The data from Starbase showed that there are no differences regarding as miR-1290 level between PC tissues (n = 5) and normal tissues (n = 2) (Supplementary Fig. 3A). However, the sample size was too small to draw a meaningful conclusion. Our results found that miR-1290 level was obviously upregulated in PC tissues. Similar to our result, exosomal miR-1290 was found to act as a prognostic marker in castration-resistant PC, and increased miR-1290 level was related to worse overall survival of patients with castration-resistant PC [27]. However, it is still unclear why miR-1290 level is increased in PC cells. Our results indicated that CAFs could transfer exosomes to PC cells, leading to the increase of miR-1290 in PC cells. Furthermore, CAFs-Exo promoted PC cell migration, invasion, EMT and stemness, whereas these effects were abolished by anti-miR-1290. EMT and cancer stem-like cells (CSCs) have emerged as important contributors to drive metastatic dissemination of carcinomas [49,50,51]. These data showed that CAFs secreted exosomal miR-1290 promoted EMT and stemness, leading to the metastasis of PC. Thus, we speculated that blocking the function of CAFs-secreted exosomal miR-1290 might serve as an alternative approach for the treatment of PC.
MiRNAs play key roles in cancer progression through regulating target gene expressions [52]. MiR-1290 could promote the proliferation of colorectal cancer cells via targeting INPP4B [53]. Additionally, Yan et al. showed that miR-1290 contributed to glioma cell migration and invasion through downregulating LHX6 [54]. Meanwhile, Wu et al. found that GSK3β was a direct target of miR-1290 in A549 cells [55]. Our results consistent with the previous research showing that GSK3β is a downstream target gene of miR-1290. GSK3β is a serine/threonine kinase involved in multiple cellular processes such as cell proliferation, cell cycle and metabolic pathways [56,57,58]. The data from TCGA database showed that there is no difference as for GSK3β level between PC tissues and adjacent normal tissue (Supplementary Fig. 3B). However, our results showed that GSK3β level was notably reduced in PC tissues. The reason for this disparity may be due to the individual differences in patients with PC.
Importantly, GSK3β was able to contribute to the degradation of β-catenin, which related to cell proliferartion [59,60,61]. Meanwhile, upregulation of the Wnt/β-catenin signaling pathway could promote prostatic cancer development [62, 63]. Thus, we focus on investigating the relationships among miR-1290, GSK-3β, and β-catenin in PC cells. MiR-1246 has been found to enhance lung cell migration and invasion by activating Wnt/β-catenin signaling via inhibiting GSK-3β expression [56]. In this study, we found that CAFs-Exo obviously reduced GSK3β level and elevated β-catenin, c-Myc and Cyclin D1 level in PC cells in vitro and in vivo; whereas these changes were reversed by anti-miR-1290. Collectively, CAFs secreted exosomal miR-1290 promoted PC cell growth and metastasis via downregulating GSK3β/β-catenin signaling.
It has been shown that one miRNA can regulate numerous mRNAs [64]; thus, more studies are needed to investigate whether exosomal miR-1290 affects the progression of PC via targeting other genes.
Conclusion
Collectively, we found that exosomal miR-1290 from CAFs could promote tumor growth and metastasis via downregulating GSK3β/β-catenin signaling, suggesting that inhibiting CAFs-derived exosomal miR-1290 may provide a new modality for the treatment of growth and metastasis in PC.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Wang G, Zhao D, Spring DJ, DePinho RA. Genetics and biology of prostate cancer. Genes Dev. 2018;32:1105–40.
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015;65:87–108.
Janiczek M, Szylberg Ł, Kasperska A, Kowalewski A, Parol M, Antosik P, et al. Immunotherapy as a promising treatment for prostate cancer: A systematic review. J Immunol Res. 2017;2017:4861570.
Gîngu C, Heidenreich A, Andreşanu A, Mihancea A, Sinescu I, Baston C. Treatment of the primary in metastatic prostate cancer. Curr Opin Urol. 2020;30:566–75.
Mokbel K, Wazir U, Mokbel K. Chemoprevention of prostate cancer by natural agents: Evidence from molecular and epidemiological studies. Anticancer Res. 2019;39:5231–59.
Jin MZ, Jin WL. The updated landscape of tumor microenvironment and drug repurposing. Signal Transduct Target Ther. 2020;5:166.
Corn KC, Windham MA, Rafat M. Lipids in the tumor microenvironment: From cancer progression to treatment. Prog Lipid Res. 2020;80:101055.
Erdogan B, Webb DJ. Cancer-associated fibroblasts modulate growth factor signaling and extracellular matrix remodeling to regulate tumor metastasis. Biochem Soc Trans. 2017;45:229–36.
Kubo N, Araki K, Kuwano H, Shirabe K. Cancer-associated fibroblasts in hepatocellular carcinoma. World J Gastroenterol. 2016;22:6841–50.
Yu S, Jiang Y, Wan F, Wu J, Gao Z, Liu D. Immortalized cancer-associated fibroblasts promote prostate cancer carcinogenesis, proliferation and invasion. Anticancer Res. 2017;37:4311–8.
Neuwirt H, Bouchal J, Kharaishvili G, Ploner C, Jöhrer K, Pitterl F, et al. Cancer-associated fibroblasts promote prostate tumor growth and progression through upregulation of cholesterol and steroid biosynthesis. Cell Commun Signal. 2020;18:11.
Zhang Y, Zhao J, Ding M, Su Y, Cui D, Jiang C, et al. Loss of exosomal miR-146a-5p from cancer-associated fibroblasts after androgen deprivation therapy contributes to prostate cancer metastasis. J Exp Clin Cancer Res. 2020;39:282.
Zhou Q, Wu X, Wang X, Yu Z, Pan T, Li Z, et al. The reciprocal interaction between tumor cells and activated fibroblasts mediated by TNF-α/IL-33/ST2L signaling promotes gastric cancer metastasis. Oncogene 2020;39:1414–28.
Lee TH, D’Asti E, Magnus N, Al-Nedawi K, Meehan B, Rak J. Microvesicles as mediators of intercellular communication in cancer-the emerging science of cellular ‘debris’. Semin Immunopathol. 2011;33:455–67.
Gupta D, Liang X, Pavlova S, Wiklander OPB, Corso G, Zhao Y, et al. Quantification of extracellular vesicles in vitro and in vivo using sensitive bioluminescence imaging. J Extracell Vesicles. 2020;9:1800222.
Li SP, Lin ZX, Jiang XY, Yu XY. Exosomal cargo-loading and synthetic exosome-mimics as potential therapeutic tools. Acta Pharm Sin. 2018;39:542–51.
Hough KP, Deshane JS. Exosomes in Allergic Airway Diseases. Curr Allergy Asthma Rep. 2019;19:26.
Zhao XQ, Ao Y, Chen HY, Wang H. [The role of miRNA in kidney development]. Yi Chuan. 2020;42:1062–72.
Zhang B, Pan X, Cobb GP, Anderson TA. microRNAs as oncogenes and tumor suppressors. Dev Biol. 2007;302:1–12.
Wang X, Luo G, Zhang K, Cao J, Huang C, Jiang T, et al. Hypoxic Tumor-Derived Exosomal miR-301a Mediates M2 Macrophage Polarization via PTEN/PI3Kγ to Promote Pancreatic Cancer Metastasis. Cancer Res. 2018;78:4586–98.
Hannafon BN, Trigoso YD, Calloway CL, Zhao YD, Lum DH, Welm AL, et al. Plasma exosome microRNAs are indicative of breast cancer. Breast Cancer Res. 2016;18:90.
Xu G, Zhang B, Ye J, Cao S, Shi J, Zhao Y, et al. Exosomal miRNA-139 in cancer-associated fibroblasts inhibits gastric cancer progression by repressing MMP11 expression. Int J Biol Sci. 2019;15:2320–9.
Li YY, Zhou CX, Gao Y. Interaction between oral squamous cell carcinoma cells and fibroblasts through TGF-β1 mediated by podoplanin. Exp Cell Res. 2018;369:43–53.
Ye Y, Li SL, Wang SY. Construction and analysis of mRNA, miRNA, lncRNA, and TF regulatory networks reveal the key genes associated with prostate cancer. PLoS One. 2018;13:e0198055.
Sun LP, Xu K, Cui J, Yuan DY, Zou B, Li J, et al. Cancer‑associated fibroblast‑derived exosomal miR‑382‑5p promotes the migration and invasion of oral squamous cell carcinoma. Oncol Rep. 2019;42:1319–28.
Fang T, Lv H, Lv G, Li T, Wang C, Han Q, et al. Tumor-derived exosomal miR-1247-3p induces cancer-associated fibroblast activation to foster lung metastasis of liver cancer. Nat Commun. 2018;9:191.
Huang X, Yuan T, Liang M, Du M, Xia S, Dittmar R, et al. Exosomal miR-1290 and miR-375 as prognostic markers in castration-resistant prostate cancer. Eur Urol. 2015;67:33–41.
Jia H, Hao C. Exploring dysregulated miRNAs in cryptorchidism: A systematic review. J Int Med Res. 2021;49:300060521999950.
Dweep H, Gretz N, Sticht C. miRWalk database for miRNA-target interactions. Methods Mol Biol. 2014;1182:289–305.
Paraskevopoulou MD, Georgakilas G, Kostoulas N, Vlachos IS, Vergoulis T, Reczko M, et al. DIANA-microT web server v5.0: service integration into miRNA functional analysis workflows. Nucleic Acids Res. 2013;41:W169–173.
Vijay GV, Zhao N, Den Hollander P, Toneff MJ, Joseph R, Pietila M, et al. GSK3β regulates epithelial-mesenchymal transition and cancer stem cell properties in triple-negative breast cancer. Breast Cancer Res. 2019;21:37.
Wu S, Yue Y, Gu Y, Wang Q, Liu T, Li L, et al. MUC15 loss facilitates epithelial-mesenchymal transition and cancer stemness for prostate cancer metastasis through GSK3β/β-catenin signaling. Cell Signal. 2021;84:110015.
Li Q, Wang G, Tao J, Chen W. RNF6 promotes colorectal cancer invasion and migration via the Wnt/β-catenin pathway by inhibiting GSK3β activity. Pathol Res Pr. 2021;225:153545.
Dvořánková B, Lacina L, Smetana K Jr. Isolation of normal fibroblasts and their cancer-associated counterparts (CAFs) for biomedical research. Methods Mol Biol. 2019;1879:393–406.
Ippolito L, Morandi A, Taddei ML, Parri M, Comito G, Iscaro A, et al. Cancer-associated fibroblasts promote prostate cancer malignancy via metabolic rewiring and mitochondrial transfer. Oncogene 2019;38:5339–55.
Jang I, Beningo KA. Integrins, CAFs and mechanical forces in the progression of cancer. Cancers (Basel). 2019;11:721.
Hurtado P, Martínez-Pena I, Piñeiro R. Dangerous Liaisons: Circulating Tumor Cells (CTCs) and Cancer-Associated Fibroblasts (CAFs). Cancers (Basel). 2020;12:2816.
Ren J, Ding L, Zhang D, Shi G, Xu Q, Shen S, et al. Carcinoma-associated fibroblasts promote the stemness and chemoresistance of colorectal cancer by transferring exosomal lncRNA H19. Theranostics 2018;8:3932–48.
Ingenito F, Roscigno G, Affinito A, Nuzzo S, Scognamiglio I, Quintavalle C, et al. The role of exo-mirnas in cancer: A focus on therapeutic and diagnostic applications. Int J Mol Sci. 2019;20:4687.
Scognamiglio I, Cocca L, Puoti I, Palma F, Ingenito F, Quintavalle C, et al. Exosomal microRNAs synergistically trigger stromal fibroblasts in breast cancer. Mol Ther Nucleic Acids. 2022;28:17–31.
Donnarumma E, Fiore D, Nappa M, Roscigno G, Adamo A, Iaboni M, et al. Cancer-associated fibroblasts release exosomal microRNAs that dictate an aggressive phenotype in breast cancer. Oncotarget 2017;8:19592–608.
Hu JL, Wang W, Lan XL, Zeng ZC, Liang YS, Yan YR, et al. CAFs secreted exosomes promote metastasis and chemotherapy resistance by enhancing cell stemness and epithelial-mesenchymal transition in colorectal cancer. Mol Cancer. 2019;18:91.
Yang F, Yan Y, Yang Y, Hong X, Wang M, Yang Z, et al. MiR-210 in exosomes derived from CAFs promotes non-small cell lung cancer migration and invasion through PTEN/PI3K/AKT pathway. Cell Signal. 2020;73:109675.
Zhang H, Deng T, Liu R, Ning T, Yang H, Liu D, et al. CAF secreted miR-522 suppresses ferroptosis and promotes acquired chemo-resistance in gastric cancer. Mol Cancer. 2020;19:43.
Dou D, Ren X, Han M, Xu X, Ge X, Gu Y, et al. Cancer-associated fibroblasts-derived exosomes suppress immune cell function in breast cancer via the miR-92/PD-L1 pathway. Front Immunol. 2020;11:2026.
Jin JJ, Liu YH, Si JM, Ni R, Wang J. Overexpression of miR-1290 contributes to cell proliferation and invasion of non small cell lung cancer by targeting interferon regulatory factor 2. Int J Biochem Cell Biol. 2018;95:113–20.
Qin WJ, Wang WP, Wang XB, Zhang XT, Du JD. MiR-1290 targets CCNG2 to promote the metastasis of oral squamous cell carcinoma. Eur Rev Med Pharm Sci. 2019;23:10332–42.
Lin M, Shi C, Lin X, Pan J, Shen S, Xu Z, et al. sMicroRNA-1290 inhibits cells proliferation and migration by targeting FOXA1 in gastric cancer cells. Gene 2016;582:137–42.
Li Y, Lv Z, Zhang S, Wang Z, He L, Tang M, et al. Genetic fate mapping of transient cell fate reveals N-Cadherin activity and function in tumor metastasis. Dev Cell. 2020;54:593–607.e595.
Bakir B, Chiarella AM, Pitarresi JR, Rustgi AK. EMT, MET, plasticity, and tumor metastasis. Trends Cell Biol. 2020;30:764–76.
Babaei G, Aziz SG, Jaghi NZZ. EMT, cancer stem cells and autophagy; The three main axes of metastasis. Biomed Pharmacother. 2021;133:110909.
Wu M, Wang G, Tian W, Deng Y, Xu Y. MiRNA-based Therapeutics for Lung Cancer. Curr Pharm Des. 2018;23:5989–96.
Ma Q, Wang Y, Zhang H, Wang F. miR-1290 contributes to colorectal cancer cell proliferation by targeting INPP4B. Oncol Res. 2018;26:1167–74.
Yan L, Cai K, Sun K, Gui J, Liang J. MiR-1290 promotes proliferation, migration, and invasion of glioma cells by targeting LHX6. J Cell Physiol. 2018;233:6621–9.
Wu L, Liu T, Xiao Y, Li X, Zhu Y, Zhao Y, et al. Polygonatum odoratum lectin induces apoptosis and autophagy by regulation of microRNA-1290 and microRNA-15a-3p in human lung adenocarcinoma A549 cells. Int J Biol Macromol. 2016;85:217–26.
Yang F, Xiong H, Duan L, Li Q, Li X, Zhou Y. MiR-1246 promotes metastasis and invasion of A549 cells by targeting GSK-3β‒Mediated Wnt/β-catenin pathway. Cancer Res Treat. 2019;51:1420–9.
Walz A, Ugolkov A, Chandra S, Kozikowski A, Carneiro BA, O’Halloran TV, et al. Molecular pathways: Revisiting glycogen synthase kinase-3β as a target for the treatment of cancer. Clin Cancer Res. 2017;23:1891–7.
Pecoraro C, Faggion B, Balboni B, Carbone D, Peters GJ, Diana P, et al. GSK3β as a novel promising target to overcome chemoresistance in pancreatic cancer. Drug Resist Updat. 2021;58:100779.
Vallée A, Lecarpentier Y, Guillevin R, Vallée JN. Effects of cannabidiol interactions with Wnt/β-catenin pathway and PPARγ on oxidative stress and neuroinflammation in Alzheimer’s disease. Acta Biochim Biophys Sin (Shanghai). 2017;49:853–66.
Reabroi S, Chairoungdua A, Saeeng R, Kasemsuk T, Saengsawang W, Zhu W, et al. A silyl andrographolide analogue suppresses Wnt/β-catenin signaling pathway in colon cancer. Biomed Pharmacother. 2018;101:414–21.
Rubinfeld B, Albert I, Porfiri E, Fiol C, Munemitsu S, Polakis P. Binding of GSK3beta to the APC-beta-catenin complex and regulation of complex assembly. Science 1996;272:1023–6.
Xi M, Cheng L, Hua W, Zhou YL, Gao QL, Yang JX, et al. MicroRNA-95-3p promoted the development of prostatic cancer via regulating DKK3 and activating Wnt/β-catenin pathway. Eur Rev Med Pharm Sci. 2019;23:1002–11.
Sha J, Han Q, Chi C, Zhu Y, Pan J, Dong B, et al. PRKAR2B promotes prostate cancer metastasis by activating Wnt/β-catenin and inducing epithelial-mesenchymal transition. J Cell Biochem. 2018;119:7319–27.
Kim VN, Nam JW. Genomics of microRNA. Trends Genet. 2006;22:165–73.
Acknowledgements
This study was supported by the Science Foundation of Peking University Cancer Hospital-2021-7.
Author information
Authors and Affiliations
Contributions
SW, YC, and JM made major contributions to the conception, design, investigation, and manuscript drafting of this study. XY, ZY, and YY were responsible for investigation, data analysis, data interpretation. PD made substantial contributions to conception and design of the study and revised the manuscript critically for important intellectual content. All authors agreed to be accountable for all aspects of the work. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All procedures were approved by the ethics committee of the Peking University Cancer Hospital & Institute.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, S., Du, P., Cao, Y. et al. Cancer associated fibroblasts secreted exosomal miR-1290 contributes to prostate cancer cell growth and metastasis via targeting GSK3β. Cell Death Discov. 8, 371 (2022). https://doi.org/10.1038/s41420-022-01163-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-022-01163-6
- Springer Nature Limited
This article is cited by
-
The miR-1290/OGN axis in ovarian cancer-associated fibroblasts modulates cancer cell proliferation and invasion
Journal of Ovarian Research (2024)
-
Exosomal miRNAs: the tumor's trojan horse in selective metastasis
Molecular Cancer (2024)
-
The role of cancer-associated fibroblasts and exosomal miRNAs-mediated intercellular communication in the tumor microenvironment and the biology of carcinogenesis: a systematic review
Cell Death Discovery (2024)
-
The biological function of extracellular vesicles in prostate cancer and their clinical application as diagnostic and prognostic biomarkers
Cancer and Metastasis Reviews (2024)
-
Cancer-associated fibroblasts and its derived exosomes: a new perspective for reshaping the tumor microenvironment
Molecular Medicine (2023)