Abstract
Objective
Gout, common metabolic disorders, have poorly understood links with blood metabolites. Exploring these relationships could enhance clinical prevention and treatment strategies.
Methods
We applied bidirectional two-sample Mendelian randomization (MR) analysis, using data from a genome-wide association (GWAS) study of 486 blood metabolites. Gout data was obtained from FinnGen R8 (7461 gout and 221,323 control cases). We implemented the inverse variance-weighted (IVW) method for main analytical approach. Extensive heterogeneity, pleiotropy tests, leave-one-out analysis, and reverse MR were conducted to validate the robustness of our findings. Both Bonferroni and False Discovery Rate (FDR) corrections were used to adjust for multiple comparisons, ensuring stringent validation of our results.
Results
Initial MR identified 31 candidate metabolites with potential genetic associations to gout. Following rigorous sensitivity analysis, 23 metabolites as potential statistical significance after final confirmation. These included metabolites enhancing gout risk such as X-11529 (OR = 1.225, 95% CI 1.112–1.350, P < 0.001), as well as others like piperine and stachydrine, which appeared to confer protective effects. The analysis was strengthened by reverse MR analysis. Additionally, an enrichment analysis was conducted, suggesting that 1-methylxanthine may be involved in the metabolic process of gout through the caffeine metabolism pathway.
Conclusion
Identifying causal metabolites offers new insights into the mechanisms influencing gout, suggesting pathways for future research and potential therapeutic targets.
Similar content being viewed by others
Introduction
Gout is caused by metabolic disorder (hyperuricemia), but it shares risk factors and pathophysiological mechanisms with other metabolic diseases. Its main characteristics include arthritis and the deposition of urate crystals. Studies indicate that the prevalence of gout in the general population ranges from 1% to 4%, and it is higher in certain regions (such as the Pacific Islands) and specific subgroups (such as the elderly, those with renal dysfunction, or metabolic syndrome). For example, the estimated prevalence of adult gout in the United States is approximately 3.9% [1,2,3]. Gout is closely related to metabolic processes, especially those involving urate metabolism. It is characterized by elevated serum uric acid levels, which crystallize and cause inflammation and acute gouty arthritis. This condition is closely associated with metabolic syndrome, which is linked to cerebrovascular disease, cardiovascular disease, and renal damage, thus imposing a significant economic burden on society [4]. Although various lifestyle and genetic factors associated with gout risk have been identified, current research results are inconsistent in terms of specific metabolic changes. For example, abnormal increases or decreases in the concentrations of metabolites such as hypoxanthine, uric acid, 2-deoxyadenosine, creatinine, homocysteine, D-glucose, DL-2-aminohexanoic acid, and adenosine may occur in the blood of gout patients [5,6,7,8,9]. The observational nature of studies limits our ability to establish a causal relationship between these metabolites and the development of gout, as such studies are prone to confounding factors and reverse causality. With the development of high-throughput technologies, such as multivariable genome-wide association studies (GWAS) and cluster analysis, advanced research methods allow us to measure and analyze blood metabolites more accurately, enhancing our understanding of disease mechanisms at the molecular level [10]. To further explore the causal relationship between gout and blood metabolites, this study plans to utilize Mendelian randomization (MR) methods based on GWAS data. By using genetic variants as natural experiments, MR methods can infer causality by analyzing the relationship between these variants and the disease. Compared to traditional observational studies, this method reduces research bias and provides more robust statistical results [11, 12]. This comprehensive research approach helps deepen our understanding of gout and related metabolic diseases, promoting the development of personalized medicine and precision treatment.
Materials and methods
Study data
We conducted MR analysis using publicly available GWAS data to assess the causal impact of exposure factors, blood metabolites, on the outcome factor, gout. We performed a bidirectional two-sample MR analysis to investigate the genetic causal relationship between blood metabolites and gout, following three key assumptions: (1) Metabolite instrumental variables (IVs) are associated with the outcome of gout, (2) Metabolite IVs are unrelated to confounders of gout, and (3) IVs affect gout only through the exposure. To ensure the robustness of the results, we performed careful IVs selection, as outlined in the flowchart (Fig. 1) and data table (Table 1) [13].
Exposure variable collection
This study utilized the genome-wide association study (GWAS) by Shin et al. in 2014, which included 7824 adults from two European cohorts, TwinsUK and KORA, encompassing approximately 2.1 million single nucleotide polymorphisms (SNPs) [14]. Metabolic profiling of fasting serum samples was conducted through non-targeted mass spectrometry and comprehensive high-performance liquid chromatography and gas chromatography-mass spectrometry, successfully identifying 486 serum metabolites (see details in Table S1). Of these, 309 were confirmed via the Kyoto Encyclopedia of Genes and Genomes (KEGG) database and categorized into eight major biochemical classes: amino acids, carbohydrates, cofactors and vitamins, energy, lipids, nucleotides, peptides, and xenobiotics. The remaining 177 unknown metabolites were not further analyzed due to undefined chemical properties. Complete statistical data of these genome-wide association summary datasets are publicly accessible through the Metabolomics GWAS server. Additionally, standardized identification, relative quantification, data reduction, and quality assurance processes conducted by Metabolon, Inc. ensured the accuracy and reliability of the data. This research not only provides an in-depth understanding of the biochemical pathways of known metabolites but also lays the groundwork for the functional exploration of unknown metabolites and future biomedical research.
Outcome data collection
The GWAS summary data for gout were obtained from the R8 version of the dataset in the Finngen biobank (www.finngen.fi/fi). This study included 462,933 adult samples of European ancestry from the UK Biobank, comprising 7461 gout patients and 221,323 healthy controls. All participants had obtained local ethical approval and provided informed consent, making additional ethical approval unnecessary for this study.
IVs selection and MR analysis
First, we retrieved SNPs associated with blood metabolites (P < 5e-5) as IVs, selected according to widely applied operational standards [15] to assess the correlation between metabolites and gout. Secondly, we filtered out linkage disequilibrium (LD) among the included IVs (r2 < 0.001, LD distance = 10,000 kb). Third, we excluded SNPs directly related to gout (P < 0.05). The strength of the IVs was estimated using the F-statistic, with F < 10 indicating exclusion of weak instrument bias.
To enhance the accuracy and robustness of MR analysis, we utilized the PhenoScanner V2 website to exclude SNPs significantly associated with known risk factors, including palindromic SNPs. This measure aims to eliminate SNPs that may bias the MR estimates, ensuring more stable and reliable results [16], see Table S2 for details.
In our study, we initially screened 31 potential metabolites associated with the development of gout (Table S3). Subsequently, we applied the Bonferroni correction algorithm (α = 0.05, PB = 0.05/486, PB = 1e-4) to correct the type I error rate in multiple hypothesis testing and identified a metabolite with a significant causal relationship with gout. Metabolites with a Bonferroni-adjusted P-value below 1e-4 were considered statistically significant. Two results had P-values less than 1e-4; However, due to the limited number of results, we adjusted the threshold to 1e-4 < PB < 0.05 [17]. The 29 metabolites with Bonferroni values between 1e-4 and 0.05 were classified as having potential statistical significance.
After Bonferroni correction, we conducted heterogeneity analysis using Cochran’s Q test to evaluate heterogeneity among the estimated SNP effects, with P > 0.05 indicating no significant heterogeneity. Horizontal pleiotropy was examined using MR Egger intercept tests and Mendelian randomization multivariable Effect Residual and Outlier (MR-PRESSO) methods, with P > 0.05 indicating no horizontal pleiotropy. Following sensitivity analysis, of the two results meeting the stricter significance criterion, only one remained significant. Moreover, out of the 29 metabolites, 22 retained their status of potential statistical significance after the analysis.
Following rigorous statistical analysis and sensitivity testing, the remaining 23 metabolites were confirmed to meet the criteria for potential statistical association with the development of gout. These metabolites demonstrated significant associations with gout development, supported by multiple analytical methods, and thus qualified as positive metabolites in the study.
Subsequently, we conducted reverse MR analysis on these 23 positive findings; this additional analysis was pivotal in determining whether the observed relationships could be due to reverse causation [18]. The Bonferroni-adjusted P-value below 2e-4 were considered statistically significant (0.05/23). Our primary analytical approach was the Inverse Variance-Weighted (IVW) method, which enhances statistical power by synthesizing results from multiple independent studies. Additionally, we utilized four other methods for sensitivity analysis: MR Egger, Simple mode, Weighted median, and Weighted mode, to further validate the robustness of our results.
The significance of these 23 metabolites in gout-related investigations was underscored. We used the MR Steiger filtering test to examine the causal directionality between each extracted SNP and the exposure and outcome, which is valuable for enhancing the accuracy of causal inference [19]. Additionally, we conducted leave-one-out analysis to assess the robustness of MR results by sequentially excluding individual SNPs to evaluate the sensitivity of the remaining SNPs’ effect estimates. We also employed scatterplot and funnel plot methods to visually inspect for potential outliers and assess the overall distribution and heterogeneity of the estimates [20, 21].
To accurately assess the strength of the association between the metabolites and gout, and to reduce the false discovery rate (FDR), we employed the Benjamin-Hochberg method, setting a significance threshold at PBH < 0.05. These findings were analyzed and visualized on the online platform “https://www.bioinformatics.com.cn” (last accessed on January 12, 2023), with details and citations documented in reference [22].
MR analyses and Steiger filtering analyses were performed in R Studio (version 4.2.1) using the TwoSampleMR package (version 0.5.6). Finally, we used funnel plots to detect and assess publication bias and visualize the distribution of study results, aiding in identifying potential confounding bias (analysis date: September 2023).
Metabolite pathway enrichment analysis
In our study, metabolic pathway analysis was initially conducted using the KEGG database (https://www.kegg.jp/) for 23 potential metabolites as specific causal relationships; however, no significant results were obtained. Consequently, the scope of our analysis was expanded to include 31 initial candidate metabolites for further pathway investigation. This analysis aimed to understand the functionality and interactions of metabolites, facilitating the identification of potential metabolic pathways [23]. Using metaconflict 5.0 (https://www.metaboanalyst.ca/), we analyzed blood metabolites via KEGG pathways [24], analyzed in July 2024.
Results
Initiatory MR results
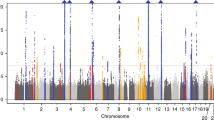
We selected 486 metabolites as IVs, with the number of IVs ranging from 4 to 631. The validity tests for all these IVs demonstrated that the minimum F-statistics exceeded 10 (see Table S3), indicating that they are sufficiently robust to support effective MR analyses for investigating associations with gout. Using the IVW method, we explored the causal relationships between known metabolites and gout. We identified 31 preliminary candidate metabolites with potential causal associations with gout, which included 2 amino acids, 1 carbohydrate, 5 lipids, 1 peptide, 2 xenobiotics, and 20 unknown metabolites. Among these, 19 metabolites were positively correlated with gout risk, while 12 were negatively correlated. All of candidate genetic metabolites met the criteria with FDR < 0.05 (Fig. 2). The top 5 positively correlated metabolites were as follows: mannoses (b = 1.42, 95% CI = 2.14 ~ 0.194, PFDR = 0.0006, P = 0.00003), X-11529 (b = 0.20, 95% CI = 1.11 ~ 1.35, PFDR = 0.0006, P = 0.00004), X-13429 (b = 0.37, 95% CI = 1.44 ~ 1.82, PFDR = 0.020, P = 0.002), X-11593 (b = 0.55, 95% CI = 1.18 ~ 2.55, PFDR = 0.021, P = 0.005), and undecanoate (b = 1.26, 95% CI = 1.45 ~ 8.57, PFDR c = 0.021, P = 0.005). The top 5 negatively correlated metabolites were: stachydrine (b = −0.31, 95% CI = 0.60 ~ 0.90, PFDR = 0.020, P = 0.003), bradykinin (b = −0.18, 95% CI = 0.74 ~ 0.94, PFDR = 0.020, P = 0.003), 1-linoleoylglycerol (b = −0.36, 95% CI = 0.53 ~ 0.93, PFDR = 0.026, P = 0.012), X-11530 (b = −0.28, 95% CI = 0.53 ~ 0.93, PFDR = 0.026, P = 0.012), and piperine (b = −0.33, 95% CI = 0.56 ~ 0.94, PFDR = 0.026, P = 0.014). Among these, 1 carbohydrate, 2 amino acids, 3 lipids, 1 xenobiotics, and 12 unknown metabolites increased the risk of gout, while 10 metabolites (2 lipids, 1 peptide, 1 xenobiotics, and 8 unknown metabolites) had a protective effect. In these two analyses, the highest odds ratio (OR) was for the carbohydrate mannoses (OR = 4.13, 95% CI = 2.14 ~ 0.194). In the MR Steiger test, the correct direction of causality was confirmed.
After the Bonferroni correction (P-value < 1e-4), two metabolites — mannose and X-11529 — remained statistically significant. When the significance threshold was adjusted to 1e-4 < P-value < 0.05, a total of 29 blood metabolites were found to potentially have a causal relationship with gout.
Upon further analysis for heterogeneity and pleiotropy, only X-11529 among these two metabolites maintained statistical significance. Of these 29 metabolites, 22 still showed significant associations. All the 23 potential metabolites met the criteria with FDR < 0.05 (Table S4 and Fig. 3).
The top 5 among 23 positively correlated metabolites were: X-11529 (b = 0.20, 95% CI = 1.11 ~ 1.35, PFDR = 0.0006, P = 0.00004), X-13429 (b = 0.37, 95% CI = 1.44 ~ 1.82, PFDR = 0.020, P = 0.002), X-11593 (b = 0.55, 95% CI = 1.18 ~ 2.55, PFDR = 0.021, P = 0.005), X-08402 (b = 0.61, 95% CI = 1.84 ~ 2.83, PFDR = 0.021, P = 0.006), and undecanoate (b = 1.26, 95% CI = 1.45 ~ 8.57, PFDR = 0.021, P = 0.005). The top 5 among 23 negatively correlated metabolites were consistent with the 31 candidate metabolites.
Sensitivity analysis
To assess the stability of the results, sensitivity analyses were performed on 31 preliminary selected candidate metabolites causally related to gout, including MR-Egger regression, Cochrane’s Q, MR-PRESSO, leave-one-out analysis, and funnel plots. As mentioned above, a total of 23 potential positive results were obtained.
The results of Cochrane’s Q showed no heterogeneity in the causal relationship between the 23 potential blood metabolites and gout (P > 0.05), and MR-PRESSO also indicated no pleiotropy (P > 0.05) (Table S4). After Bonferroni correction, we found one blood metabolites significantly associated with an increased risk of gout: X-11529, which is an unknown type (OR = 1.23, 95% CI = 1.11 ~ 1.35), leave-one-out analysis did not reveal single SNP bias (Fig. 4A), and the funnel plot was symmetric (Fig. 4B), confirming the reliability of the causal relationship in the result.
A Leave-one-out plot showing the effect of individual SNPs on the estimated association with gout. Points indicate effect estimates with the SNP omitted; horizontal lines are 95% confidence intervals. The red line is the overall effect size. B Scatter plot of MR analysis comparing two methods by plotting SNPs’ ratio estimates (x-axis) against precision (y-axis), with black for Inverse variance weighted and blue for MR Egger. B Scatter plot representing the MR analysis methods applied to assess the causal effect of metabolite X-11529 on gout. Each dot represents an SNP, plotted with its respective ratio estimate (β-IVW or β-Egger) on the x-axis and the precision of the estimate (1/standard error) on the y-axis. The IVW method is shown in light blue, while the MR Egger method is depicted in dark blue, with the vertical lines representing their respective overall causal estimates. Abbreviation:Mendelian randomization (MR), Odds Ratios (OR), single nucleotide polymorphisms (SNPs), Inverse variance weighted (IVW), confidence interval (CI), false discovery rate (FDR).
Exploring the causal relationship between gout to blood metabolites
Further investigation into the causal relationship between gout to the 23 potential blood metabolites was conducted using reverse MR analysis. The preliminary analysis, prior to statistical adjustment, indicated that levels of 15 metabolites were increasing, while 8 metabolites showed a decrease (Table S5 and Fig. S1). However, after applying the Bonferroni method to adjust for multiple comparison errors (adjusted significance threshold at P value = 0.05/23 = 0.002), the changes in these metabolites did not reach statistical significance. This suggests that under strict statistical standards, the impact of gout on these 21 metabolites could not be confirmed to have a causal relationship. This finding is consistent with the results from the forward MR analysis, indicating that more evidence is needed to interpret the relationship between gout and metabolites.
Metabolic pathway
In our study, metabolic pathway analysis was conducted on 23 specific potential causal relationships. It was revealed that the gout-associated metabolite 1-methylxanthine is related to the caffeine pathway (P = 0.019), while 2-aminobutyrate is enriched in the cysteine and methionine metabolism pathway (P = 0.056), and isovalerate is enriched in the protein digestion and absorption pathway (P = 0.040), as illustrated in Supplementary Fig. S1 and Table S6.
Discussion
In this study, we utilized the causal associations between blood metabolites and gout through a bidirectional two-sample Mendelian Randomization (MR) analysis. We employed single nucleotide polymorphisms (SNPs) as instrumental variables (IVs) for this purpose. MR is a statistical methodology that utilizes genetic variants to establish causality, thereby enhancing our comprehension of the impact of specific genetic factors on disease susceptibility [12].
We initially identified 31 candidate metabolites associated with gout risk through comprehensive genome-wide association study (GWAS) and MR analysis. After applying stringent Bonferroni correction, Benjamin-Hochberg method, and sensitivity analyses, we confirmed 1 metabolite with a Bonferroni-corrected P-value less than 1e-4 as significantly associated with gout risk, while 22 metabolites with Bonferroni-corrected P-values between 1e-4 and 0.05 were identified as potentially associated with gout risk, enhancing the robustness of our findings. The identification of these metabolites unveils potential pathological mechanisms and may provide new biomarkers and therapeutic targets for the prevention and treatment of gout. Notably, our data revealed that certain metabolites, such as X-11529, awere significantly associated with increased gout risk; conversely, metabolites like stachydrine, bradykinin, and piperine were associated with reduced gout risk. This might reflect the regulatory roles of these metabolites under pathological conditions on gout. By conducting reverse MR analysis on these associations, we further validated the causal directions of these relationships, strengthening the credibility of our study. These results underscore the potential roles of specific metabolites in the pathological process of gout. Overall, our study not only enhances the understanding of the association between gout and blood metabolites but also verifies the authenticity of these relationships through various statistical methods. As far as we are aware, there is limited research on this subject.
Gout is an inflammatory arthropathy triggered by a disturbance in uric acid metabolism and is prevalent worldwide, with an increasing incidence in recent years. According to the Global Burden of Disease (GBD) Study, the prevalence of gout in the United States rose from 2007 to 2016 [25]. Gout is associated with socioeconomic status, being more common in lower-income populations. There are significant differences in prevalence and treatment patterns across different regions globally, with genetics, diet, and lifestyle all playing contributory roles. It is projected that by 2060, mortality related to gout may increase by 55%, posing a significant challenge to public health [26].
The complex role of metabolites in the development of gout has garnered significant attention, particularly through the application of metabolomic technologies such as proteomics and lipidomics. These methods have been instrumental in identifying potential biomarkers linked to gout. Recent metabolic research has identified essential pathways and specific metabolites that regulate serum uric acid levels, a critical determinant in gout. By analyzing blood and urine metabolites, biomarkers for pathological states, particularly gout and hyperuricemia, can be detected [27, 28]. Notably, metabolites including uric acid, hypoxanthine, xanthine, guanosine, adenosine, Kynurenic Acid (KYNA), 2-Aminoethylphosphonic acid (2PY), DL-2-aminoheptanoic acid (2AMIA), and 5-hydroxyindoleacetic acid (5-HIAALB4) are proposed as biomarkers for gout [29,30,31]. Recent studies have shown significant changes in lipid, carbohydrate, amino acid, and energy metabolism in gout patients, highlighting the link between metabolic dysregulation and gout [32]. These findings open new avenues for diagnosis, prevention, and treatment, although the bidirectional causal relationships between specific metabolites and gout are yet to be established.
Mannose, a carbohydrate closely related to glucose, plays a crucial role in protein glycosylation and energy provision. It is produced during the digestion of polysaccharides and glycoproteins, and notably influences the immune system by boosting antibody production against viruses [33]. Recent studies have established a significant link between mannose levels and gout. Metagenomic analyses have revealed notable differences in the gut microbiota composition between gout patients and healthy individuals, specifically highlighting an increased abundance of genes involved in mannose metabolism [34]. Additionally, the Chinese herbal medicine Wuwei Shexiang pill, recognized for its therapeutic potential in treating gout, may exert its effects by inhibiting several metabolic pathways including those of sugar and mannose, linoleic acid, and phenylalanine, thus mitigating symptoms of gout [35]. These findings underscore the critical role of mannose in the pathogenesis and management of gout, aligning with the risk effects observed in this study.
Piperine, extracted from plants, offers bioenhancing and anti-inflammatory benefits. It shows promise in gout treatment by inhibiting the NOD-like receptor family pyrin domain containing 3 (NLRP3) inflammasome, crucial for gout-related inflammation. This leads to reduced gout symptoms, prevents tophi, and protects cartilage. Piperine also decreases leukocyte infiltration, lipid peroxidation, and C-reactive protein production, aiding gout management [36]. Additionally, it increases cell membrane permeability, enhancing drug transport into cells [37]. Animal studies confirm that piperine alleviates urate-induced gouty arthritis in mice by reducing lysosomal enzymes, lipid peroxidation, tumor necrosis factor-alpha (TNF-α) levels, and boosting antioxidants [38]. Consistent with the analysis presented, piperine aligns as a protective factor against gout.
Stachydrine from Motherwort exhibits notable anti-inflammatory effects, crucial for gout management [39]. It reduces key inflammatory cytokines like interleukin-1β (IL-1β) and TNF-α, impedes platelet-neutrophil aggregation, and inhibits neutrophil extracellular traps (NETs) formation, which are important in the pathology of gout [40]. It also curtails oxidative stress, aiding in osteoclast inhibition, anti-fibrosis, and tissue protection [41, 42]. Moreover, stachydrine provides osmoprotective advantages, reducing the risk of gout-related kidney damage [43]. Our findings, alongside existing studies, highlight stachydrine’s potential as a therapeutic for gout-induced bone and tissue damage, although further clinical trials are needed to validate its clinical benefits.
1-Methylxanthine is a natural alkaloid akin to caffeine and theophylline,affects the central and cardiovascular systems. It is a caffeine metabolism byproduct [44]. Febuxostat inhibits xanthine oxidase (XO) to reduce uric acid levels, raising 1-methylxanthine’s excretion and effectiveness [45]. Studies confirm its importance in gout research; derivatives like 7-methylxanthine prevent urate crystal formation, potentially preventing gout attacks [46, 47]. A study found caffeine, as a prodrug of 1-Methylxanthine, links caffeine dose to 1-Methylxanthine/1-methyluric acid (1MU) ratio in urine, aiding gout treatment monitoring [48]. These insights offer new avenues for managing and treating gout.
Our study, utilizing MR analysis, discovered that bradykinin may have a protective role in gout, diverging from previous observational studies that identified it as primarily pro-inflammatory [49, 50]. This discrepancy may be attributed to an insufficient control of confounding factors in earlier research [51]. Furthermore, we observed that the bradykinin receptor 2 (B2 receptor) decreases leukocyte infiltration and tissue damage, underscoring its critical role in gout therapy and its potential as a therapeutic target [52]. More research might elucidate the role and therapeutic value of bradykinin at various stages of gout, with the potential to offer novel strategies for gout management.
A metabolomics study identified 23 serum metabolomic biomarkers differentially expressed across various stages of gout, predominantly associated with uric acid metabolism and oxidative stress [7]. The regulation of serum uric acid concentration is a complex process involving genetic, lifestyle, and environmental factors. Current data on the relationship between caffeine and gout or hyperuricemia are limited and controversial [53, 54]. Our results suggested that 1-methylxanthine might exacerbate the onset of gout through the caffeine pathway, consistent with prior literature. For instance, studies have indicated that excessive consumption of caffeine-containing coffee could elevate serum uric acid levels in healthy individuals, potentially due to caffeine influencing biochemical reactions (such as increasing the activity of xanthine oxidase) and modulating metabolic processes in the kidneys [55, 56]. A metabolomics analysis of Han Chinese individuals revealed that potential genetic targets related to gout are primarily enriched in the caffeine metabolism pathway, as well as the cysteine and methionine metabolism pathways [57]. However, the specific mechanisms and causal relationships require further investigation. This study identified two additional metabolic pathways closely related to gout: cysteine and methionine metabolism, and protein digestion and absorption. It highlights the critical role of amino acids in the production or excretion of uric acid, which is intimately connected to gout [58]. Furthermore, purines present in proteins can be metabolized into uric acid, affecting uric acid levels [59]. This research emphasizes the importance of further investigation into the metabolic pathways of caffeine and their relevance to biological functions associated with gout.
The strengths of our research: Our research utilized MR to establish causality, relying on publicly available data, thus obviating the need for individual-level datasets. This methodological choice significantly reduces both time and financial costs associated with data acquisition. Our analysis diverges from traditional methods by employing a comprehensive, large-scale GWAS dataset to elucidate the relationships between metabolites and gout, offering insights that are unique in the field. Although our findings align with previous studies, our research boasts a more rigorous methodology. An earlier MR study [60], did not employ the Steiger test to exclude IVs indicating reverse causation, not did they conduct reverse MR analysis, potentially affecting the robustness of their findings. By contrast, we believe our study makes a valuable contribution and extension to the field through the use of a more stringent methodological approaches. Importantly, MR’s advantage over observational studies lies in its ability to minimize confounding variables, thereby enhancing the validity and reliability of our findings regarding the causal effects of metabolites on gout. To further validate these associations, we conducted detailed sensitivity analyses. These analyses specifically addressed potential issues such as horizontal pleiotropy and genetic confounding, ensuring the robustness and reliability of the causal inferences drawn from our study.
However, there are limitations to our study. First, the GWAS data used are exclusively from European populations, which reduces the impact of population heterogeneity on the results but raises issues of generalizability. The conclusions may not necessarily apply to other populations. Further validation in more diverse populations using additional GWAS data is needed to confirm the generalizability of our results. Secondly, over half of the 31 candidate blood metabolites identified are unknown, indicating limited understanding and incomplete knowledge of gout pathogenesis. Further research is needed to elucidate their functions and biological roles. Thirdly, while our method is theoretically robust, further randomized controlled trials are crucial to verify causal relationships and translate findings into clinical applications, specifically in preventing or treating gout. Future research should address these limitations for a comprehensive understanding and reliable evidence.
Conclusion
A metabolome-wide MR scan identified 31 candidate metabolites associated with gout risk, particularly mannose, 1-methylxanthine. Piperine and stachydrine may protect against gout. 1-methylxanthine was linked through caffeine metabolism pathway. Further research is needed for biomarker validation and pathogenesis exploration.
Data availability
All data used in this study are available in publicly available datasets. The discovery dataset can be found from Finn biobank (https://storage.googleapis.com/finngen-public-data-r8/summary_stats/finngen_R8_M13_GOUT.gz/, consisting of 7461 cases of gout and 221,323 healthy control). The datasets produced and/or analyzed in the course of this study can be accessed by reaching out to the corresponding author upon making a reasonable request.
References
Zhu Y, Pandya BJ, Choi HK. Prevalence of gout and hyperuricemia in the US general population: the National Health and Nutrition Examination Survey 2007–2008. Arthritis Rheum. 2011;63:3136–41. https://doi.org/10.1002/art.30520.
Dalbeth N, Merriman TR, Stamp LK. Gout. Lancet. 2016;388:2039–52. https://doi.org/10.1016/S0140-6736(16)00346-9.
Parisa N, Kamaluddin MT, Saleh MI, Sinaga E. The inflammation process of gout arthritis and its treatment. J Adv Pharm Technol Res. 2023;14:166–70. https://doi.org/10.4103/japtr.japtr_144_23.
Li Y, Han X, Tong J, Wang Y, Liu X, Liao Z, et al. Analysis of metabolites in gout: a systematic review and meta-analysis. Nutrients. 2023;15:3143. https://doi.org/10.3390/nu15143143.
Zhang J, Jin C, Ma B, Sun H, Chen Y, Zhong Y, et al. Global, regional and national burdens of gout in the young population from 1990 to 2019: a population-based study. RMD Open. 2023;9:e003025. https://doi.org/10.1136/rmdopen-2023-003025.
Lyu S, Rao Y, Liu P, Yang S, Chen W, Yang H, et al. Metabolomics analysis reveals four biomarkers associated with the gouty arthritis progression in patients with sequential stages. Semin Arthritis Rheum. 2022;55:152022. https://doi.org/10.1016/j.semarthrit.2022.152022.
Liu S, Wang Y, Liu H, Xu T, Wang MJ, Lu J, et al. Serum lipidomics reveals distinct metabolic profiles for asymptomatic hyperuricemic and gout patients. Rheumatology. 2022;61:2644–51. https://doi.org/10.1093/rheumatology/keab743.
Chen J, Zhou J, Wei S-S, Li HC, Wen CP, Xu GW. Analysis of serum metabolic characteristics in patients with gout using gouts chromatography—mass spectrometry. J Instrum Anal 2016;35:137–42.
Wang W, Kou J, Zhang M, Wang T, Li W, Wang Y, et al. A metabonomic study to explore potential markers of asymptomatic hyperuricemia and acute gouty arthritis. J Orthop Surg Res. 2023;18:96. https://doi.org/10.1186/s13018-023-03585-z.
Loos RJF. 15 years of genome-wide association studies and no signs of slowing down. Nat Commun. 2020;11:5900. https://doi.org/10.1038/s41467-020-19653-5.
Zeng H, Zhou K, Zhuang Y, Li A, Luo B, Zhang Y. Unraveling the connection between gut microbiota and Alzheimer’s disease: a two-sample Mendelian randomization analysis. Front Aging Neurosci. 2023;15:1273104. https://doi.org/10.3389/fnagi.2023.1273104.
Yang T, Bi S, Zhang X, Yin M, Feng S, Li H. The impact of different intensities of physical activity on serum urate and gout: a Mendelian randomization study. Metabolites. 2024;14:66. https://doi.org/10.3390/metabo14010066.
Thompson W, Swain S, Zhao SS, Kamps A, Coupland C, Kuo C, et al. Causal association between subtypes of osteoarthritis and common comorbidities: a Mendelian randomisation study. Osteoarthr Cartil Open. 2023;5:100414. https://doi.org/10.1016/j.ocarto.2023.100414.
Shin SY, Fauman EB, Petersen AK, Krumsiek J, Santos R, Huang J, et al. An atlas of genetic influences on human blood metabolites. Nat Genet. 2014;46:543–50. https://doi.org/10.1038/ng.2982.
Burgess S, Small DS, Thompson SG. A review of instrumental variable estimators for Mendelian randomization. Stat Methods Med Res. 2017;26:2333–55. https://doi.org/10.1177/0962280215597579.
Kamat MA, Blackshaw JA, Young R, Surendran P, Burgess S, Danesh J, et al. PhenoScanner V2: an expanded tool for searching human genotype-phenotype associations. Bioinformatics. 2019;35:4851–3. https://doi.org/10.1093/bioinformatics/btz469.
Zhi-Gang Y, Han-Dong W. A causal link between circulating leukocytes and three major urologic cancers: a Mendelian randomization investigation. Front Genet. 2024;15:1424119. https://doi.org/10.3389/fgene.2024.1424119.
Xu M, Li C, Xiang L, Chen S, Chen L, Ling G, et al. Assessing the causal relationship between 731 immunophenotypes and the risk of lung cancer: a bidirectional Mendelian randomization study. BMC Cancer. 2024;24:270. https://doi.org/10.1186/s12885-024-12014-1.
Lingyu M, Hongguang L, Mingdong Z, Na L, Yahui L. Aminotransferases as causal factors for metabolic syndrome: a bidirectional Mendelian randomization study. PLoS ONE. 2024;19:e0302209. https://doi.org/10.1371/journal.pone.0302209.
Verbanck M, Chen CY, Neale B, Do R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet. 2018;50:693–8. https://doi.org/10.1038/s41588-018-0099-7.
Bowden J, Davey Smith G, Burgess S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol. 2015;44:512–25. https://doi.org/10.1093/ije/dyv080.
Tang D, Chen M, Huang X, Zhang G, Zeng L, Zhang G, et al. SRplot: a free online platform for data visualization and graphing. PLoS ONE. 2023;18:e0294236. https://doi.org/10.1371/journal.pone.0294236.
Jin Z, Sato Y, Kawashima M, Kanehisa MKEGG. tools for classification and analysis of viral proteins. Protein Sci. 2023;32:e4820. https://doi.org/10.1002/pro.4820.
Zhong Y, Yang C, Zhang B, Chen Y, Cai W, Wang G, et al. Causal impact of human blood metabolites and metabolic pathways on serum uric acid and gout: a Mendelian randomization study. Front Endocrinol. 2024;15:1378645. https://doi.org/10.3389/fendo.2024.1378645.
Dehlin M, Jacobsson L, Roddy E. Global epidemiology of gout: prevalence, incidence, treatment patterns and risk factors. Nat Rev Rheumatol. 2020;16:380–90. https://doi.org/10.1038/s41584-020-0441-1.
Mattiuzzi C, Lippi G. Recent updates on worldwide gout epidemiology. Clin Rheumatol. 2020;39:1061–3. https://doi.org/10.1007/s10067-019-04868-9.
Albrecht E, Waldenberger M, Krumsiek J, Evans AM, Jeratsch U, Breier M, et al. Metabolite profiling reveals new insights into the regulation of serum urate in humans. Metabolomics. 2014;10:141–51. https://doi.org/10.1007/s11306-013-0565-2.
Ohashi Y, Ooyama H, Makinoshima H, Takada T, Matsuo H, Ichida K. Plasma and urinary metabolomic analysis of gout and asymptomatic hyperuricemia and profiling of potential biomarkers: a pilot study. Biomedicines. 2024;12:300. https://doi.org/10.3390/biomedicines12020300.
Lyu S, Ding R, Yang S, Chen W, Rao Y, OuYang H, et al. Establishment of a clinical diagnostic model for gouty arthritis based on the serum biochemical profile: a case-control study. Medicine. 2021;100:e25542. https://doi.org/10.1097/MD.0000000000025542.
Wu X, You C. The biomarkers discovery of hyperuricemia and gout: proteomics and metabolomics. PeerJ. 2023;11:e14554. https://doi.org/10.7717/peerj.14554.
Zhang Y, Zhang H, Chang D, Guo F, Pan H, Yang Y. Metabolomics approach by 1H NMR spectroscopy of serum reveals progression axes for asymptomatic hyperuricemia and gout. Arthritis Res Ther. 2018;20:111. https://doi.org/10.1186/s13075-018-1600-5.
Suh YS, Noh HS, Kim HJ, Cheon YH, Kim M, Lee H, et al. Differences in clinical and dietary characteristics, serum adipokine levels, and metabolomic profiles between early- and late-onset gout. Metabolites. 2021;11:399. https://doi.org/10.3390/metabo11060399.
Davis JA, Freeze HH. Studies of mannose metabolism and effects of long-term mannose ingestion in the mouse. Biochim Biophys Acta. 2001;1528:116–26. https://doi.org/10.1016/s0304-4165(01)00183-0.
Chu Y, Sun S, Huang Y, Gao Q, Xie X, Wang P, et al. Metagenomic analysis revealed the potential role of gut microbiome in gout. NPJ Biofilms Microbiomes. 2021;7:66. https://doi.org/10.1038/s41522-021-00235-2.
Lang J, Li L, Quan Y, Tan R, Zhao J, Li M, et al. LC-MS-based metabolomics reveals the mechanism of anti-gouty arthritis effect of Wuwei Shexiang pill. Front Pharmacol. 2023;14:1213602. https://doi.org/10.3389/fphar.2023.1213602.
Jati GAK, Assihhah N, Wati AA, Salasia SIO. Immunosuppression by piperine as a regulator of the NLRP3 inflammasome through MAPK/NF-κB in monosodium urate-induced rat gouty arthritis. Vet World. 2022;15:288–98. https://doi.org/10.14202/vetworld.2022.288-298.
Boonyarattanasoonthorn T, Kongratanapasert T, Maiuthed A, Hamlin R, Kijtawornrat A, Khemawoot P. Bioenhancing effects of piperine and curcumin on triterpenoid pharmacokinetics and neurodegenerative metabolomes from Centella Asiatica extract in beagle dogs. Sci Rep. 2022;12:20789. https://doi.org/10.1038/s41598-022-24935-7.
Sabina EP, Nagar S, Rasool M. A role of piperine on monosodium urate crystal-induced inflammation-an experimental model of gouty arthritis. Inflammation. 2011;34:184–92. https://doi.org/10.1007/s10753-010-9222-3.
Cheng F, Zhou Y, Wang M, Guo C, Cao Z, Zhang R, et al. A review of pharmacological and pharmacokinetic properties of stachydrine. Pharmacol Res. 2020;155:104755. https://doi.org/10.1016/j.phrs.2020.104755.
Sun X, Zhou M, Pu J, Wang T. Stachydrine exhibits a novel antiplatelet property and ameliorates platelet-mediated thrombo-inflammation. Biomed Pharmacother. 2022;152:113184. https://doi.org/10.1016/j.biopha.2022.113184.
Tan H, Li Z, Zhang S, Zhang J, Jia E. Novel perception of neutrophil extracellular traps in gouty inflammation. Int Immunopharmacol. 2023;115:109642. https://doi.org/10.1016/j.intimp.2022.109642.
Liu L, Shan L, Wang H, Schauer C, Schoen J, Zhu L, et al. Neutrophil extracellular trap-borne elastase prevents inflammatory relapse in intercritical gout. Arthritis Rheumatol. 2023;75:1039–47. https://doi.org/10.1002/art.42431.
Meng J, Zhou C, Zhang W, Wang W, He B, Hu B, et al. Stachydrine prevents LPS-induced bone loss by inhibiting osteoclastogenesis via NF-κB and Akt signalling. J Cell Mol Med. 2019;23:6730–43. https://doi.org/10.1111/jcmm.14551.
Prathap L, Jayaraman S, Roy A, Santhakumar P, Jeevitha M. Molecular docking analysis of stachydrine and sakuranetin with IL-6 and TNF-α in the context of inflammation. Bioinformation. 2021;17:363–8. https://doi.org/10.6026/97320630017363.
Chambers ST, Kunin CM. Isolation of glycine betaine and proline betaine from human urine. Assessment of their role as osmoprotective agents for bacteria and the kidney. J Clin Invest. 1987;79:731–7. https://doi.org/10.1172/JCI112878.
Youn H, Hee Kook Y, Oh ET, Jeong SY, Kim C, Kyung Choi E, et al. 1-Methylxanthine enhances the radiosensitivity of tumor cells. Int J Radiat Biol. 2009;85:167–74. https://doi.org/10.1080/09553000902741190.
Tsai M, Wu JT, Gunawardhana L, Naik H. The effects of xanthine oxidase inhibition by febuxostat on the pharmacokinetics of theophylline. Int J Clin Pharmacol Ther. 2012;50:331–7. https://doi.org/10.5414/cp201648.
Costa-Bauza A, Grases F. 7-methylxanthine inhibits the formation of monosodium urate crystals by increasing its solubility. Biomolecules. 2023;13:1769. https://doi.org/10.3390/biom13121769.
Birkett DJ, Miners JO, Valente L, Lillywhite KJ, Day RO. 1-Methylxanthine derived from caffeine as a pharmacodynamic probe of oxypurinol effect. Br J Clin Pharmacol. 1997;43:197–200. https://doi.org/10.1046/j.1365-2125.1997.53711.x.
Damas J, Remacle-Volon G, Adam A. Inflammation in the rat paw due to urate crystals. Involvement of the kinin system. Naunyn Schmiedebergs Arch Pharmacol. 1984;325:76–9. https://doi.org/10.1007/BF00507057.
Phelps P, Prockop DJ, McCarty DJ. Crystal induced inflammation in canine joints. 3. Evidence against bradykinin as a mediator of inflammation. J Lab Clin Med. 1966;68:433–44.
Kalincik T, Butzkueven H. Observational data: understanding the real MS world. Mult Scler. 2016;22:1642–8. https://doi.org/10.1177/1352458516653667.
Prado GN, Taylor L, Zhou X, Ricupero D, Mierke DF, Polgar P. Mechanisms regulating the expression, self-maintenance, and signaling-function of the bradykinin B2 and B1 receptors. J Cell Physiol. 2002;193:275–86. https://doi.org/10.1002/jcp.10175.
Choi HK, Curhan G. Coffee, tea, and caffeine consumption and serum uric acid level: the third national health and nutrition examination survey. Arthritis Rheum. 2007;57:816–21. https://doi.org/10.1002/art.22762.
Rozza F, Trimarco V, Izzo R, Grassi D, Ferri C. Effects of a novel fixed combination of nutraceuticals on serum uric acid concentrations and the lipid profile in asymptomatic hyperuricemic patients: results from the PICONZ-UA study. High Blood Press Cardiovasc Prev. 2016;23:381–6. https://doi.org/10.1007/s40292-016-0168-x.
Amer MG, Mazen NF, Mohamed AM. Caffeine intake decreases oxidative stress and inflammatory biomarkers in experimental liver diseases induced by thioacetamide: biochemical and histological study. Int J Immunopathol Pharmacol. 2017;30:13–24. https://doi.org/10.1177/0394632017694898.
Towiwat P, Tangsumranjit A, Ingkaninan K, Jampachaisri K, Chaichamnong N, Buttham B, et al. Effect of caffeinated and decaffeinated coffee on serum uric acid and uric acid clearance, a randomised within-subject experimental study. Clin Exp Rheumatol. 2021;39:1003–10. https://doi.org/10.55563/clinexprheumatol/gd9nia.
Liu H, Xie R, Dai Q, Fang J, Xu Y, Li B. Exploring the mechanism underlying hyperuricemia using comprehensive research on multi-omics. Sci Rep. 2023;13:7161. https://doi.org/10.1038/s41598-023-34426-y.
He S, Qiao X, Zhang S, Xia J, Wang L, Liu S. Urate oxidase from tea microbe Colletotrichum camelliae is involved in the caffeine metabolism pathway and plays a role in fungal virulence. Front Nutr. 2023;9:1038806. https://doi.org/10.3389/fnut.2022.1038806.
Xie Y, Li Y, Zhang J, Chen Y, Ren R, Xiao L, et al. Assessing the causal association between human blood metabolites and the risk of gout. Eur J Clin Invest. 2023:e14129. https://doi.org/10.1111/eci.14129.
Funding
This work was financially supported in part by research grants from Shenzhen Science and Technology Project (JCYJ20210324111805014). Scientific research of Guangdong Provincial Bureau of Traditional Chinese Medicine Project (20221343) with Ye Zhang.
Author information
Authors and Affiliations
Contributions
Conception and design: HQ Z, JD L, and W L. Data curation: all authors. Formal analysis: JD L, ZH L, and HQ Z. Investigation: all authors. Methodology: all authors. Project administration: Y Z, ZH L, and W L. Resources: all authors. Data analysis: HQ Z, ZH L, and JD L. Supervision: W L,ZH L, and Y Z. Writing—original draft: HQ Z, JD L, and W L. Writing—review & editing: Y Z, HQ Z, and W L. Approval of final manuscript: all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The research conducted to support this study received approval from the appropriate ethics committees. Furthermore, all participants in the original studies had given their informed written consent.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zeng, H., Lai, J., Liu, Z. et al. Specific blood metabolite associations with Gout: a Mendelian randomization study. Eur J Clin Nutr (2024). https://doi.org/10.1038/s41430-024-01497-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41430-024-01497-7
- Springer Nature Limited