Abstract
We recently achieved the first-in-human transfusion of induced pluripotent stem cell-derived platelets (iPSC-PLTs) as an alternative to standard transfusions, which are dependent on donors and therefore variable in supply. However, heterogeneity characterized by thrombopoiesis-biased or immune-biased megakaryocytes (MKs) continues to pose a bottleneck against the standardization of iPSC-PLT manufacturing. To address this problem, here we employ microRNA (miRNA) switch biotechnology to distinguish subpopulations of imMKCLs, the MK cell lines producing iPSC-PLTs. Upon miRNA switch-based screening, we find imMKCLs with lower let-7 activity exhibit an immune-skewed transcriptional signature. Notably, the low activity of let-7a-5p results in the upregulation of RAS like proto-oncogene B (RALB) expression, which is crucial for the lineage determination of immune-biased imMKCL subpopulations and leads to the activation of interferon-dependent signaling. The dysregulation of immune properties/subpopulations, along with the secretion of inflammatory cytokines, contributes to a decline in the quality of the whole imMKCL population.
Similar content being viewed by others
Introduction
The first successful whole blood transfusion took place in 1818 by James Blundell1. This historic achievement was followed by Karl Landsteiner’s discovery of ABO blood types in 19012 and the development of a platelet isolation device for transfusion by Emil J. Freireich in the 1950s-1960s3. Since then, donor-dependent platelet transfusion has become a standard healthcare practice despite persistent challenges of supply shortages resulting from various factors including viral contamination, alloimmune transfusion refractoriness, and the recent COVID-19 pandemic. In this context, we have adopted the strategy of ex vivo blood pharming using human induced pluripotent stem cells (iPSCs). We developed immortalized MK progenitor cell lines (imMKCLs) as the starting cell source for ex vivo iPSC-derived platelet (iPSC-PLT) manufacturing4,5. imMKCLs exhibit sustained proliferation over several months in the presence of doxycycline (DOX) and shed iPSC-PLTs after DOX removal, enabling the generation of over 1011 iPSC-PLTs in a turbulent flow-based bioreactor6. Building upon these achievements, we initiated the first-in-human clinical trial, iPLAT17,8,9. While iPLAT1 demonstrated promising outcomes without significant adverse events, a transient increase in D-dimer levels and leukocyte count was observed after transfusion of the maximal dose7. These observations indicate the potential involvement of recently identified immune-skewed MKs10,11, raising concerns about the quality control of imMKCLs as master cells. Meanwhile, we noticed considerable variation in the quality of imMKCL clones with regards to their proliferation and iPSC-PLT production capacity12. Specifically, certain imMKCL clones displaying cellular senescence exhibited a diminished capacity to generate iPSC-PLTs, but the capacity could be restored through the knockdown of p53 and CDKN1A12. However, the molecular factors underlying this heterogeneity, which significantly hinder the efficiency and standardization of iPSC-PLT manufacturing, remain unknown.
MicroRNAs (miRNAs) are small non-coding RNAs that negatively regulate the stability and translation of the target mRNAs by binding with the complementary mRNA sequences. In hematopoietic cells, miRNAs have been identified as key players in cell fate decisions and functions13. Based on the evidence that DNA-based genetic circuits regulate protein expression depending on miRNA activities rather than miRNA expression14, we developed an innovative biotechnology method, miRNA switch, which enables the identification of specific cell types15,16. MiRNA switches can detect endogenous miRNA activity to distinguish heterogeneous cell populations without the need of antibody labeling. This technology has proven successful for various cell types, including iPSC-derived hepatocytes, endothelial cells, cardiomyocytes15, mouse embryonic stem cell (ESC)-derived neurons17, and undifferentiated human ESCs and iPSCs18. We therefore hypothesized that the miRNA switch can be employed to distinguish subpopulations among imMKCLs with heterogenous miRNA activities (i.e., a gradient variation in miRNA activity).
In this study, we screened a library containing 269 target miRNA switches, leading to the identification of let-7a-5p and let-7g-5p miRNAs with heterogenous activity among imMKCLs. By a transcriptional analysis of imMKCL subpopulations exhibiting either high or low let-7 activity (hereafter referred as let-7 high and let-7 low, respectively), we discovered that let-7 low imMKCLs exhibited an immune-skewed transcriptional signature. Further investigation unveiled the pivotal role of let-7a-5p and its downstream target, RAS like proto-oncogene B (RALB), in modulating the lineage determination of ‘immune’ MKs within imMKCLs. Importantly, we elucidated that the dysregulation of immune properties/subpopulations within imMKCLs, along with the secretion of inflammatory cytokines, results in a decline of their quality based on their arrested proliferation and deficient iPSC-PLT generation. This study provides valuable insights into the correlation between immune-biased MKs and their role in quality control during ex vivo iPSC-PLT manufacturing.
Results
Identification of miRNAs with endogenous activity in imMKCLs
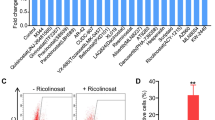
MiRNA switch technology describes synthetic mRNA-based genetic circuits that regulate transgene expressions through miRNA activity15,16. We hypothesized that miRNA switches can identify imMKCL subsets with distinct miRNA activities, providing a valuable tool for exploring the heterogeneity within imMKCLs. Figure 1a depicts a schematic illustration of miRNA switch technology. We designed synthetic mRNAs that contain complementary sequences of target miRNAs in the 5′ untranslated region (UTR) and encode a reporter fluorescent protein to monitor endogenous miRNA activity. A pair of mRNAs were synthesized and co-transfected into imMKCLs, including Azami-Green 1 (AG1)-coding mRNA as a transfection control and TagBFP-coding mRNA containing the target miRNA sequence (Fig. 1b). TagBFP expression levels were measured by flow cytometry in cells with endogenous activity for a specific miRNA (Fig. 1c). A miRNA switch-based screening was conducted using a library containing 269 miRNA switches (Supplementary Table 1) and resulted in the identification of 24 types of miRNAs with endogenous activity in imMKCLs (Fig. 1d).
a A schematic illustration of the design of miRNA switches. The mRNA is composed of the antisense sequence of the target miRNA and a reporter transgene. In the presence of an active target miRNA, the expression of the reporter gene is repressed. b Design of miRNA switches for imMKCLs. A pair of mRNAs, one encoding TagBFP containing the antisense sequence of the miRNA target in the 5’ UTR and the other of Azami Green protein, were synthesized in vitro. The two mRNAs were co-transfected into imMKCLs via lipofection. 24 h after the transfection, the cells were analyzed by flow cytometry to identify the miRNA activity in imMKCLs through a screening. c Representative dot plots of target miRNA activity in imMKCLs. Active miRNA showed less TagBFP expression. d Flow cytometry analysis of the target miRNAs showing endogenous activity in imMKCLs. A miRNA switch-based screening was performed from a library containing 269 kinds of miRNA switches (Supplementary Table 1), and 24 kinds of miRNAs were identified with endogenous activity in imMKCLs.
Let-7 miRNA switches enrich immune-skewed imMKCLs
Among the active miRNAs, we focused on let-7a-5p and let-7g-5p, as they showed heterogeneous activity among imMKCLs (Fig. 2a). We sorted subpopulations with high or low let-7 activity, with the let-7 low subset representing less than 5% of the total population. We confirmed that the let-7 activity levels in these populations correlated with their expression levels in imMKCL subpopulations (Supplementary Fig. 1a). To characterize molecular differences between the let-7 high and let-7 low subpopulations, we performed a bulk RNA-seq analysis using three different imMKCL clones (M35-1, clone 7, clone 7–3) in both the proliferation and maturation phases. The properties of clone 7 imMKCLs and the functionality of the derived iPSC-PLTs were reported previously5,6. Besides, clone 7 has been employed by multiple research groups for the investigation of megakaryopoiesis and thrombopoiesis19,20,21, highlighting its importance as a research tool. Clone 7–3, which is derived from clone 7 and displays an aged phenotype characterized by reduced proliferation and diminished iPSC-PLT production following repeated cultures, was also employed (Supplementary Fig. 1b). M35-1 was established from patient iPSCs for the iPLAT1 clinical trial due to its relatively superior expandability and iPSC-PLT productivity when compared with other competent patient-derived imMKCL clones8. Comparable iPSC-PLT generation was observed in the three clones for the let-7 low or let-7 high subpopulations, respectively (Supplementary Fig. 1c).
a Heterogenous responsive activities in imMKCLs were identified with let-7a-5p or let-7g-5p miRNA switches, resulting in let-7 low-responsive and let-7 high-responsive subpopulations. b A schematic illustration of the bulk RNA-seq sampling workflow. Let-7a-5p and let-7g-5p switches were transfected into imMKCLs, followed by the sorting of let-7 low- and high-responsive cells by flow cytometry. Three distinct imMKCL clones (clone 7, clone 7–3, M35-1) were employed for the bulk RNA-seq analysis. Gene set enrichment analysis (GSEA) were performed to compare let-7 low (n = 6 independent biological samples) and let-7 high (n = 6) subpopulations. Bar graphs showing the GSEA results of the top enriched immune-related gene sets in let-7 low-responsive imMKCLs in the proliferation phase (c) and maturation phase (d). GSEA plots showing typical enriched immune-related gene sets in the proliferation phase (e) and maturation phase (g). Representative enrichment plots from each group are displayed with the normalized enrichment score (NES), the determined nominal (non-adjusted) p-value, and the false discovery rate (FDR) derived from GSEA software. Heatmaps of the top differentially expressed TNF targets of let-7 low- or high-responsive imMKCLs in the proliferation phase (f) and maturation phase (h). Source data are provided as a Source Data file.
The sampling procedure for the bulk RNA-seq is illustrated in Fig. 2b. A differential gene expression analysis revealed many genes with altered expression profiles between let-7 high and let-7 low imMKCLs. To identify enriched transcriptional pathways in each subpopulation, we performed a gene set enrichment analysis (GSEA). Notably, we observed a series of immune-related gene sets that were significantly enriched in let-7 low imMKCLs in both the proliferation and maturation phases (Fig. 2c, d). Specifically, in the proliferation phase, gene sets in response to tumor necrosis factor (TNF), lipopolysaccharide (LPS), and enterotoxin were enriched in let-7 low cells, consistent with the reported properties of naïve mouse and human MKs with regard to sensing invading pathogens22,23. TNF-driven inflammation has also been suggested to induce platelet hyperreactivity in aging24. In the maturation phase, gene sets related to interferon signaling were significantly enriched in let-7 low cells. Some immune related properties were retained from the proliferation phase to maturation phase (Fig. 2e–h, Supplementary Table 2). These findings suggest that imMKCLs with lower let-7 activity preferentially enrich an immune-skewed MK phenotype. Our bulk RNA-seq analysis also identified the upregulation of several genes encoding chemokines and cytokines, including C-X-C motif chemokine ligand 10 (CXCL10), C-C motif chemokine ligand 2 (CCL2), and CCL3, in the immune-skewed let-7 low subpopulations of imMKCLs (Fig. 2f, h), indicating the consistency of multiple immune responses through the secretion of these molecules in human MKs25. Interestingly, elevated mRNA levels of these molecules have been reported in patients with severe acute respiratory syndrome (SARS) and middle east respiratory syndrome (MERS)26. Moreover, in COVID-19 patients, increased mRNA expression levels of CXCL10, CCL2, and TNF were correlated with an increased number of MKs27,28.
We further investigated whether immune MKs could be identified by let-7 miRNA switches in MK progenitors directly differentiated from human ESCs. We utilized CD34+ hematopoietic progenitor cells (HPCs) generated by our hPSC-sac method29,30 and performed a bulk RNA-seq analysis on two separate cell populations with different let-7a-5p and let-7g-5p activities (Supplementary Fig. 2a). A GSEA revealed similar enriched immune-related gene sets in let-7 low ESC-derived HPCs (Supplementary Fig. 2b, c, Supplementary Table 2), suggesting that the fate decision of immune-skewed MKs in MK development may occur as early as the hematopoietic progenitor stage.
scRNA-seq reveals the heterogeneity and functional diversity of imMKCLs
To further characterize the cellular heterogeneity of imMKCLs and explore the mechanism(s) underlying the enrichment of ‘immune’ imMKCLs by let-7 miRNA switches, we performed a single-cell (sc)RNA-seq analysis on let-7 low and high imMKCLs in the proliferation phase. Since both let-7a-5p and let-7g-5p miRNA switches demonstrated similar behaviors accordingly to the bulk RNA-seq analysis (Fig. 2), we focused our investigation on let-7a-5p. We identified five transcriptionally heterogeneous subpopulations of imMKCLs (Fig. 3a), with let-7 low imMKCLs being relatively concentrated in clusters 3 and 5 (Fig. 3b). We further determined the expression of the top differentially expressed genes (DEGs) and characterized the gene ontology (GO) terms enriched in each cluster (Fig. 3c, d).
a Visualization of imMKCLs (clone 7) by uniform manifold approximation and projection (UMAP) and colored according to let-7a-5p high/low-responsive imMKCLs and imMKCL sub-clusters. b The distribution of let-7a-5p low- and high-responsive cells in each cluster, and a bar plot showing the distribution of let-7a-5p low- and high-responsive cells in each cluster. c The relative expression level of the top ten DEGs in each cluster. d The representative terms of GO:BP enriched in each cluster. GO terms under the biological process category for upregulated differentially expressed genes (DEGs) in each cluster were analyzed by g:Profiler, p-values were determined by a default hypergeometric test and correction for multiple testing has been performed by the g:SCS algorithm. e Typical immune-related GO:BP terms in each cluster. f Violin plots showing the expression level of typical thrombopoiesis-related and immune-related genes in each cluster. Source data are provided as a Source Data file.
Cluster 1 exhibited an enrichment of GO:BP (Biological Process) terms associated with oxidative phosphorylation, indicating their role in energy supply. Conversely, clusters 2 and 4 represented subsets of cycling MK progenitors with enriched GO:BP terms related to cell cycle and mitotic nuclear division. The identification of proliferating cell nuclear antigen (PCNA) and tropomyosin 4 (TPM4), two marker genes of cycling MKs, is consistent with recent studies on human bone marrow (BM) cycling MKs31. Cluster 3 highly expressed genes associated with “platelet activation” and “blood coagulation”, suggesting a subset of MK progenitors dedicated to platelet emergence. In particular, thrombopoiesis-associated genes, such as thrombospondin 1 (THBS1)10, von Willebrand factor (VWF)32, and platelet glycoprotein IX (GP9)33, were highly expressed in cluster 3 compared to the other clusters (Fig. 3f). Moreover, cluster 3 was enriched with genes related to the immune system process, suggesting that it may represent a subset of MK progenitors with functional duality. For instance, platelet factor 4 (PF4, also known as CXCL4), one of the top DEGs enriched in cluster 3, promotes blood coagulation (Fig. 3c)34 and is a crucial regulator in innate immunity through its activity on macrophages35 and neutrophils36. Additionally, pro-platelet basic protein (PPBP, also known as CXCL7), an activator of neutrophils against bacteria37, was also significantly upregulated in cluster 3.
Both clusters 3 and 5 may represent subpopulations of MK progenitors with highly upregulated potential immune-related genes. The annotated immune genes in clusters 3 and 5 are involved in immune system process and immune response according to the GO analysis (Fig. 3e). Cluster 5 highly expressed genes sets in response to type I interferon, cytokine, and virus, whereas cluster 3 appeared to be responsive to myeloid leukocyte-mediated immunity. CCL5, a key proinflammatory chemokine38, was one of the top DEGs enriched in cluster 5 (Fig. 3f). Notably, CCL5 has been reported to promote proplatelet formation in thrombopoiesis39, and its expression was upregulated in cluster 3 compared to clusters 1, 2, or 4 (Fig. 3c, f). The interferon-stimulated gene for interferon-stimulated 15 (ISG15), a ubiquitin-like protein that can be covalently bound to host and viral proteins40, was identified as the top DEG in cluster 5. Given that the secretion of type 1 interferons from virus-infected cells is a hallmark of antiviral immunity41, our results indicated that cluster 5 represents the representative ‘immune’ MKs involved in antiviral functions. Recent in vivo models of naïve MKs have demonstrated the presence of ‘immune’ MKs, the hematopoietic stem cell (HSC) niche supporting MKs, and thrombopoiesis-biased MKs10,11,31. Notably, our results suggest that imMKCLs have a thrombopoiesis-biased subset (cluster 3) and a representative immune subset (cluster 5), similar to those found in endogenous human MKs.
The inhibition of let-7a-5p activity drives the development of ‘immune’ imMKCLs
We next investigated whether let-7 functionally drives the development of ‘immune’ subsets in imMKCLs by conducting loss-of-function experiments (Fig. 4a). We inhibited let-7a-5p expression/activity using a let-7a-5p inhibitor (Fig. 4b, c) and found enhanced gene expressions of several immune-related molecules identified in clusters 3 and 5, including PF4, PPBP, ISG15, and interferon induced protein with tetratricopeptide repeats 3 (IFIT3) (Fig. 4d). We also investigated whether the inhibition of let-7a-5p activity in imMKCLs alters the response to immune stimuli. The cells were stimulated with the pathogen receptor agonist LPS or control buffer. The supernatants were analyzed using a cytometric bead array kit by flow cytometry. We found that the inhibition of let-7a-5p in imMKCLs promoted the secretion of interleukin-8 (IL-8) (Fig. 4e), a critical proinflammatory chemokine42. IL-8 and its receptors have been shown to control MK proliferation and maturation43. These results are consistent with a previous report that found human cord blood-derived MKs produce IL-8 even in the absence of stimuli44. Collectively, we concluded that let-7a-5p is a functional player that modulates the development of ‘immune MKs’ among imMKCLs.
a A schematic illustration of the experimental workflow using a specific let-7a-5p inhibitor. b Let-7a-5p expression levels in imMKCLs (clone 7) treated with a negative control or let-7a-5p inhibitor. The expression levels were measured by qRT-PCR and normalized to an endogenous control, RNU6B (U6). c Let-7a-5p activity patterns were analyzed by FACS. d The inhibition of let-7a-5p in imMKCLs (clone 7) increased the mRNA expression levels of the marker genes identified in clusters 3 and 5 (Fig. 3). The mRNA expression levels were measured using qRT-PCR and normalized to GAPDH. e Let-7a-5p inhibition induced a greater secretion of IL-8 from imMKCLs at the proliferation stage (DOX-ON). After transfection of the let-7a-5p inhibitor or negative control, the imMKCLs were incubated for 24 h in the absence or presence of LPS (50 ng/mL) in the proliferation phase. Inflammatory molecules were measured by a cytometric bead array in the supernatant. Data are expressed as the mean ± SEM from three independent experiments. Unpaired two-tailed student’s t-tests were used to assess statistical significance. Source data are provided as a Source Data file.
RALB is a functional target of let-7a-5p in the development of immune-skewed imMKCLs
The preceding experiments revealed the critical role of let-7a-5p in immune-skewed imMKCLs. To further determine the underlying factors that contribute to immune-related outcomes, we conducted an ingenuity pathway analysis (IPA) to investigate the upstream regulators of cluster 3 and cluster 5 (let-7 low imMKCLs). Figure 5a shows a flowchart depicting the overall analysis design. IPA analysis identified the MK transcriptional factor GATA145 and the myeloid development regulator KLF2 as potential transcription activators in cluster 3 (Fig. 5b). Notably, KLF2 was shown to regulate host innate immune responses to polymicrobial infections46. On the other hand, known activators of virus-inducible cellular genes, such as interferon regulatory factor 7 (IRF7)47 and IRF348, were identified as potential upstream regulators in cluster 5 (Fig. 5b). We also compared common upstream regulators of clusters 3 and 5 with the predicted let-7a-5p targets (Fig. 5c). Our analysis identified twenty candidate upstream regulators, of which eight candidates showed detectable expression levels in imMKCLs (Fig. 5d). Based on their increased expression levels in the enriched let-7 low clusters (clusters 3 and 5), we suspected that cut-like homeobox 1 (CUX1) and RAS like proto-oncogene B (RALB) are targets of let-7a-5p (Fig. 5e) and may contribute to the observed immune-related outcomes. Accordingly, the inhibition of let-7a-5p upregulated the mRNA expression levels of CUX1 and RALB in both the proliferation and maturation phases (Fig. 5f). While the overexpression of CUX1 did not cause significant effects (Supplementary Fig. 3), RALB overexpression upregulated the mRNA expression of IRF7, ISG15, and IFIT3 (Fig. 5g, h, Supplementary Fig. 4a, b), but not the expression level of let-7a-5p (Supplementary Fig. 4c). On the other hand, despite being predicted as an upstream regulator of both clusters 3 and 5, RALB overexpression did not influence the expression of thrombopoiesis-related genes identified in Cluster 3 (Supplementary Fig. 4d), indicating that distinct let-7 targets may be responsible for regulating immune-related pathways and thrombopoietic pathways in imMKCLs. Combining our findings with the IPA (Fig. 5b), we propose that let-7a-5p tunes the immune properties of imMKCLs by targeting RALB (Fig. 5i).
a A flow chart of the upstream regulator analysis. b Bar plots showing the top 10 upstream regulators for cluster 3 and cluster 5 that were identified by the IPA based on scRNA-seq datasets. c The identification of eight potential upstream regulators by comparing the common upstream regulators of clusters 3 and 5 and predicted has-let-7a-5p targets. d Violin plots showing the expression levels of identified upstream regulators in each cluster. The regulators CUX1 and RALB showed increased expression levels in let-7a-5p low cells (clusters 3 and 5) and are colored red. The overlap p-values are calculated using Fisher’s Exact Test, and significance is generally attributed to p-values < 0.01. e The let-7a-5p binding site on CUX1 and RALB predicted by TargetScan. f Let-7a-5p inhibition induced the mRNA expression of CUX1 and RALB in imMKCLs. g The lentiviral-mediated overexpression of RALB elevated the mRNA expression of RALB at both the proliferation (DOX-ON) and maturation stages (DOX-OFF). h The overexpression of RALB induced the expression of interferon signaling genes. i A schematic illustration showing how the let-7 miRNA-RALB axis modulates interferon signaling in imMKCLs. Data are expressed as the mean ± SEM from three independent experiments. Unpaired two-tailed student’s t-tests were used to assess statistical significance. Source data are provided as a Source Data file.
The dysregulated immune properties/subsets are associated with the quality of imMKCLs
We previously reported that imMKCL clones exhibiting slower proliferation rates at the proliferation stage are less capable of generating iPSC-PLTs (Fig. 6a)12. However, the factor(s) responsible for inducing the low quality of imMKCLs, along with the cellular senescent/aging transcriptional signature (Supplementary Fig. 5a, b), remains unclear. Therefore, we reexamined our GSEA of the different qualities of MKCLs utilized in our previous study12. Similar to the GSEA of the let-7 low and high populations (Fig. 2), we found TNF signaling and interferon responsive gene sets were significantly enriched in intermediate-to-low (low-intermediate) quality MKCLs (Fig. 6b, Supplementary Fig. 5c). We also found that clone 7 and clone 7–3, despite being derived from the same iPSC clone with the same genetic background5,6, display different characteristics with regards to proliferation and iPSC-PLT production in both static and turbulent flow conditions (Fig. 6c–e) and let-7a-5p activity patterns (Supplementary Fig. 6a). Clone 7–3 showed enriched TNF signaling and interferon responsive signaling compared to clone 7 (Fig. 6f, Supplementary Fig. 6b). These results suggest that the induced immune-related pathways are associated with low-quality imMKCLs. The GSEA further demonstrated that downregulated senescent tumor protein 53 (TP53) targets are enriched in clone 7 (Supplementary Fig. 6c), indicating the cellular senescence of clone 7–3. The mRNA expression of RALB and IL-8 secretion of clone 7–3 was higher compared with clone 7 (Fig. 6g, h). It is known that inflammatory cytokines are secreted by senescent cells with persistent DNA damage49. In addition to IL-8 secretion (Fig. 6h), genes encoding inflammatory cytokines and chemokines (IFNB1, CXCL8, CXCL10, CXCL11) also showed higher expression levels in clone 7–3 (aged clone) compared with clone 7 (younger clone) (Supplementary Fig. 6d). Notably, the addition of recombinant IL-8 into imMKCL culture resulted in diminished iPSC-PLT production (Supplementary Fig. 7a–c). Blocking IL-8 signaling with Reparixin50, a specific CXCR1/2 inhibitor, improved iPSC-PLT production (Supplementary Fig. 7d–g) without affecting proliferation. Furthermore, the administration of recombinant interferon-α2a led to lower proliferation rates (Supplementary Fig. 8a) as well as impaired iPSC-PLT production in a dose-dependent manner (Supplementary Fig. 8b, d). Interferon treatment elevated mRNA expression of the senescence marker CDKN2A as well as the interferon-responsive genes ISG15 and IFIT3 in imMKCLs (Supplementary Fig. 8c). Taken together, these finding suggest that the dysregulation of immune properties/subpopulation within imMKCLs, along with the secretion of inflammatory cytokines, leads to arrested proliferation and deficient platelet generation of the whole imMKCL population, potentially due to the upregulation of RALB levels. Moreover, these results emphasize the effect of immune cytokines on ex vivo iPSC-PLT manufacturing, thus bearing implications for clinical applications.
a A schematic illustration showing the correlation of proliferation and iPSC-PLT production by a previous study12. b GSEA were performed to compare high quality clones (n = 5) and Low ~intermediate clones (n = 7). GSEA plots showing enriched TNF signaling and interferon responsive gene sets in low-intermediate quality clones compared with high quality clones. c The proliferation and fold change of the cell count at day 14 of clone 7 and of clone 7–3. d iPSC-PLT production of clone 7 and of clone 7–3 in static or turbulent flow conditions. e Representative flow cytometry plots of iPSC-PLTs generated from clone 7 and clone 7–3 in static or turbulent flow conditions. f GSEA plots showing enriched TNF signaling and interferon responsive gene sets in clone 7–3. Representative enrichment plots from each group are displayed with the NES, non-adjusted p-value, and FDR derived from GSEA software. g Clone 7–3 shows elevated RALB mRNA expression compared with clone 7. h The secretion of IL-8 from clone 7 and clone 7–3 at the proliferation stage. Data are expressed as the mean ± SEM from three independent experiments. Unpaired two wo-tailed student’s t-tests were used to assess statistical significance. Source data are provided as a Source Data file.
Intriguingly, our investigation of RALB revealed that the upregulation of RALB levels induced arrested proliferation, deficient iPSC-PLT production, and increased IL-8 secretion (Fig. 7a–d) as well as being accompanied by a cellular senescent/aging transcriptional signature (Supplementary Fig. 9a). We assumed that this mechanism could recapitulate the characteristics of low-quality imMKCLs. A bulk RNA-seq analysis revealed enriched TNF signaling and interferon response in RALB-overexpressing imMKCLs (Fig. 7e and Supplementary Fig. 9b), providing further support for our assumption. Additionally, RALB expression levels correlated with the expression levels of several interferon responsive genes in multiple imMKCL clones (Fig. 7f).
The lentiviral-mediated overexpression (O/E) of RALB leads to arrested proliferation (a) and deficient iPSC-PLT generation (b) in imMKCLs (Clone 7). c Representative flow cytometry plots of iPSC-PLTs generated from MOCK and O/E RALB in static or turbulent flow conditions. d IL-8 is secreted more by O/E RALB than by MOCK imMKCLs. e GSEA were performed to compare mock (n = 3) and RALB overexpressing cells (n = 3). GSEA plots showing enriched TNF signaling and interferon responsive gene sets in O/E RALB imMKCLs at the maturation stage. Representative enrichment plots from each group are displayed with the NES, non-adjusted p-value, and FDR derived from GSEA software. f A gene expression correlation analysis of RALB and immune-related genes. Data are expressed as the mean ± SEM from three independent experiments. Unpaired two-tailed student’s t-tests were used to assess statistical significance. Source data are provided as a Source Data file.
We employed two additional approaches to validate the effects of RALB. The small molecule inhibitor RBC8 (Supplementary Fig. 10a), which is described as a selective inhibitor against the GTPase Ral51, at 0.1 μM resulted in accelerated proliferation (Supplementary Fig. 10b) and ameliorated iPSC-PLT production (Supplementary Fig. 10c–e). siRNA-mediated RALB knockdown also enhanced iPSC-PLT production (Supplementary Fig. 10f, g). Although no significant difference was observed in IL-8 secretion (Supplementary Fig. 10h), RALB knockdown induced a decline in the mRNA expression levels of IRF7, ISG15, IFIT3 (Supplementary Fig. 10i). Finally, the iPSC-PLTs generated from low quality clone (clone 7–3) showed decreased PAC-1 binding and P-selectin expression (Supplementary Fig. 11). These results collectively suggest that dysregulated immune properties/subsets within imMKCLs are associated with diminished imMKCL proliferation and iPSC-PLT generation, a phenotype potentially attributed to the upregulation of RALB expression.
Lastly, we verified our findings in an in vitro differentiated MK model derived from cord blood CD34+ cells (Supplementary Fig. 12a). The inhibition of let-7 had no significant effect on interferon signaling or platelet production (Supplementary Fig. 12b–d). However, the overexpression of RALB resulted in the upregulation of interferon-responsive genes and a reduction in the production of platelet-like particles (Supplementary Fig. 12e–g). While our results suggest imMKCL and primary MKs respond differently to let-7 inhibition, they indicate a common effect of RALB, highlighting the pivotal role of RALB in determining immune properties/subsets in MK development.
Discussion
In this study, we applied miRNA switch technology to explore the heterogeneity of imMKCLs, a promising ex vivo source of iPSC-PLTs for transfusion therapy, at the proliferating stage. By screening a library containing 269 miRNA switches, we identified heterogenous let-7 miRNA activity among imMKCLs. By conducting bulk and scRNA-seq, we revealed that let-7 miRNA switches effectively enrich immune-skewed subpopulations. Our results demonstrate that the inability of let-7a-5p to suppress the expression of RALB, which lead to the activation of interferon-dependent signaling, is crucial for the lineage determination of immune-biased imMKCLs. While a senescence/aging signature is dominant in clone 7–3, we found the dysregulation of immune properties/subpopulations contributes to the secretion of inflammatory cytokines, resulting in a declined quality of the whole imMKCL population based on arrested proliferation and deficient iPSC-PLT generation. This senescence/aging related phenotype is potentially modulated by the let-7 microRNA/RALB/interferon signaling axis, as evidenced by RALB overexpression and knockdown. Consistently, we found the overexpression of RALB induced immune-skewed transcriptional phenotypes in a cord blood-derived MK model.
Non-coding RNAs, particularly miRNAs, play a crucial role in the post-transcriptional regulation of megakaryopoiesis52. For instance, miR-15053, miR-15554, and miR-146a55 have been proposed to act in MK lineage specification, while miR-125a-5p positively promotes proplatelet formation and platelet generation by targeting the expression of actin-bundling protein L-plastin56. Although the let-7 family is the most abundant miRNAs in megakaryocytic-erythroid progenitors57, its role in MK development has not been elucidated. Here we employed miRNA switch technology and observed the heterogeneous activities of let-7a-5p and let-7g-5p among imMKCLs. We discovered that immune-skewed phenotypes were preferentially enriched in imMKCLs with lower let-7 activity and characterized by the enhanced gene expression of immune-related molecules (i.e., ISG15, IFIT3; Fig. 3). The enriched gene sets in let-7 low imMKCLs (Fig. 2, Supplementary Table 2) partially overlapped with the enriched gene sets found in naïve lung MKs22,58,59. Interestingly, previous studies have suggested that the let-7 family is involved in the post-transcriptional control of innate immune responses to pathogenic agents60 and cytokine production by T lymphocytes in adaptive immunity61. In the present study, we observed that internal let-7 activity did not affect iPSC-PLT production (Supplementary Fig. 1c) and that the modulation of let-7 by mimics and inhibitors showed no discernible effect on iPSC-PLT production. These observations could be attributed to the intricate interplay of let-7 in regulating both the immune subset (cluster 5) and the thrombopoietic subset (cluster 3). We speculate that the inhibition of let-7 may compensate for any negative impact on iPSC-PLT production induced from the immune subset by influencing the thrombopoietic subset.
The classical view of megakaryopoiesis and thrombopoiesis has not provided insights about immune MKs in development. However, emerging evidence shows that lung-resident MKs in mice, particularly those in the non-vasculature area of the lungs, exhibit an immune-skewed transcriptional signature compared with BM-resident MKs22,58. Human embryonic MKs with immune characteristics were suggested to be generated along a distinct programmed trajectory10. Interestingly, our findings here show that heterogeneity is evident even within the immune-skewed MK population. We found imMKCLs contain ‘immune’ MKs in cluster 5, which displayed several immune programs (Fig. 3). The enriched gene sets in cluster 5 partially overlapped with those identified in human BM31. Cluster 3, on the other hand, exhibited functional duality in both thrombopoiesis and immunity. We suspect that cluster 3 contains two kinds of iPSC-PLT-generating MKs: one generating iPSC-PLTs for hemostasis specifically, and the other generating iPSC-PLTs with preferential immune functions. Cluster 3 may help explain why platelets have immune functions, while previously identified naïve immune MKs are mostly diploid and seemingly unable to produce platelets11,22.
Previous studies utilizing scRNA-seq have identified functionally distinct subsets of adult BM MKs11,31, including thrombopoiesis-biased MKs, immune MKs, and HSC niche-supporting MKs. However, the key molecular event(s) governing the development of the heterogenous ontology is poorly understood. The current study supports the functional diversity of imMKCLs, where lower let-7 activity enriches immune-skewed subpopulations. This heterogeneity may arise at or before the HPC stage (Supplementary Fig. 2). Furthermore, our findings reveal RALB as a functional target of let-7a-5p in modulating the lineage determination of ‘immune’ MKs within imMKCLs (Figs. 5, 7). RALB also drives the immune-skewed transcriptional phenotypes of cord blood-derived MKs (Supplementary Fig. 12), shedding light on the regulation of human immune MKs. RAL GTPases, encoded by RALA and RALB, are known for their roles in cell growth, granule secretion, and cancer metastasis62. Within the context of platelet biology, the activation of the RAL family in platelets occurs through α-thrombin-induced Ca2+/calmodulin binding to either RAL63. Additionally, the RAL family is recognized as critical regulators of P-selectin expression and platelet-leukocyte interactions in mice64. In cancer biology, evidence has highlighted the importance of the RALB/TBK1 pathway in cell autonomous survival by establishing a link between tumor formation and the innate immune response65. Considering the protective role of platelets in shielding cancer cells from immune surveillance by cytotoxic lympocytes66, it is reasonable to suspect that the RAL family serves as the molecular conduit connecting MKs/platelets and cancer. While another identified upstream regulator candidate, CUX1 (Fig. 5), is essential for modulating endothelial senescence67, our study highlighted the involvement of RALB in the senescence process during hematopoiesis. For ex vivo iPSC-PLT manufacturing, our findings indicate RALB may serve as a hallmark/predictor of imMKCL quality. Indeed, some low-quality imMKCL clones exhibited elevated RALB expression levels (Fig. 6g, Supplementary Fig. 5d). In addition, a decline in imMKCL quality, along with senescent phenotypes, was observed following the overexpression of RALB (Fig. 7).
The success of our ex vivo manufacturing strategy for iPSC-PLTs predominantly relies on maintaining high-quality imMKCLs preserved in a master cell bank (MCB)8,9. To meet the demand for platelet transfusion therapy, which typically requires 2–3 × 1011 platelets per standard dose68, it is mandatory to ensure a substantial quantity of imMKCLs. However, the heterogeneity of imMKCLs poses a considerable challenge to achieving efficient and standardized iPSC-PLT manufacturing and may result in the heterogenous functionality of iPSC-PLTs69. In the iPLAT1 study, which used patient-derived imMKCLs (originally an iPSC clone derived from T cells of the patient), a slightly elevated white blood cell count and D-dimer level, which were observed after the transfusion of 1011 iPSC-PLTs in cohort 37, indicated the possibility of immuno-thrombosis and the existence of immune subpopulations within imMKCLs. In the present study, we discovered a significant correlation between the immune properties/subpopulations of imMKCLs and their capacity to generate iPSC-PLTs. Whilst the immune subpopulations of MKs have been indicated as a genetically programmed ontogeny10, our findings suggest that the dysregulation of immune subpopulations/properties can have a significant impact on the overall imMKCL population, resulting in a decline of imMKCL quality. This decline is potentially mediated through the let-7 miRNA/RALB/interferon signaling axis, as evidenced by the immune-skewed transcriptional signatures and deficient iPSC-PLT production upon RALB overexpression (Figs. 5–7). Furthermore, our study revealed that utilization of the small molecule inhibitors RBC8 (Supplementary Fig. 10) and Reparxin (Supplementary Fig. 7) against RALB and IL-8 signaling favors iPSC-PLT production. The consistent efficacy of these small molecules across various clones suggests their universal functionality, highlighting their potential applicability in future clinical settings. Taken together, this study highlights the significance of considering the immune/senescent attributes of donor cells and allogenic imMKCL MCB in future iPSC-PLT transfusion therapies, as they profoundly impact both the quantity and quality of the produced iPSC-PLTs. Thus, this study provides important insights into the standardization of iPSC-PLT generation towards industrial-scaled manufacturing.
Methods
Cells
The human iPSC lines TkDN-Sev2 and T-1 were established in house70. imMKCLs (clone 7, clone 7–3, M35-1) were induced by DOX-inducible defined factors from human iPSCs and employed in previous studies5,6. The human ESC line KhES-3 was obtained from the Institute for Frontier Medical Sciences, Kyoto University (Kyoto, Japan). The cord blood-derived CD34+ cells were provided by the Japanese Red Cross Society Kanto-Koshinetsu Blook Blood Center. The use of all cells was approved by the ethics committees at Kyoto University and Chiba University.
Cell culture
The imMKCLs (clone 7, clone 7–3, M35-1) were cultured as described before6. Doxycycline was used to control the proliferation and differentiation stages.
MicroRNA switches
miRNA-responsive mRNAs (miRNA switches) were generated using a MegaScript T7 kit (Ambion) as described previously15. The miRNA switches were encoded on modified mRNA to post-transcriptionally regulate TagBFP, a blue fluorescent protein, in response to the activity of an arbitrary miRNA (miR-X) expressed in imMKCLs. TagBFP-coding mRNA includes the target miR-X sequence in 5′ UTR such that the expression level of TagBFP is suppressed in response to increased miR-X activity. Azami Green 1 (hmAG1)-coding mRNA served as the transfection control (Fig. 1B).
miRNA switch-based screening
The miRNA switch-based screening was performed using an original constructed library (total of 269 miRNA switches). Two different mRNA coding for TagBFP and AG1, respectively, were co-transfected into the cells. The transfection of miRNA switches was performed as previously described15. Briefly, in the proliferation phase of imMKCLs, the reverse transfection of 150 ng mRNAs (75 ng each) was done with StemFect mRNA (Stemgent) in 50 μL of the cell suspension (1 × 106 cells/mL) for 30 min, followed by dilution to 200 μL using culture medium in 96-well plates. After 24 h, flow cytometry was performed using a LSRFortessa (BD Biosciences, San Jose, CA, USA). Candidate miRNA switches were selected based on the active responsive pattern of imMKCLs to the target miR-X (“active” in Fig. 1c). MiRNAs that did not show strong responsive activities in imMKCLs were defined as “inactive” (Fig. 1c). For let-7a-5p and let-7g-5p, we individually transfected the candidate switches with control hmAG1-coding mRNA into imMKCLs. Following a 24 h incubation, the cells were sorted using a BD FACSAria II and subjected to further investigation.
Cell sorting and flow cytometry
Cells were suspended in staining medium, incubated for 30 min with appropriate antibodies on ice in the dark, and sorted or analyzed using a BD FACSAria II. Platelet count was determined as previously described6. The following antibodies were used for the flow cytometry: allophycocyanin (APC)-conjugated anti-CD41a (integrin aIIbb3 complex: HIP8 clone) (Biolegend, San Diego, CA), phycoerythrin (PE)-conjugated anti-CD42b (GPIbα) (eBioscience, San Diego, CA), and PE-conjugated anti-CD41a (HIP8 clone) (Biolegend). The gating strategies are provided in the Supplementary Fig. 13.
Reverse transcription and real-time PCR
Total RNA was extracted using the microRNeasy Micro Kit or microRNeasy Mini Kit (Qiagen, Hilden, Germany) and reverse-transcribed using SuperScript VILO™ Master Mix (Thermo Fisher Scientific). qPCR was carried out with SYBR™ Green PCR Master Mix (Applied Biosystems, Foster City, CA, USA) using a StepOnePlus system (Thermo Fisher Scientific). GAPDH was used as the internal control. The primer sets used are listed in Supplementary Table 3. Let-7a-5p and let-7g-5p miRNA reverse transcription was performed using a TaqMan MicroRNA Reverse Transcription Kit (Thermo Fisher Scientific) and miRNA specific stem-loop RT primers according to the manufacturer’s instructions. qPCR was carried out with TaqMan Fast Advanced Master Mix (Thermo Fisher Scientific) using the StepOnePlus system. The let-7 expression levels were determined relative to RNU6B using specific TaqMan probes.
Analysis of cytokine secretion by imMKCLs
Cytokine secretion in culture supernatants was measured using the human inflammatory cytokine cytometric bead array kit (BD Biosciences) according to the manufacturer’s guidelines. Briefly, bead populations with distinct fluorescence intensities coated with capture antibody proteins were mixed with PE-conjugated detection antibodies and recombinant standards or test samples, and then incubated to form sandwich complexes. After acquiring sample data by flow cytometry, the cytokine concentrations were calculated using FCAP ArrayTM software (BD Biosciences).
Transfection of let-7a-5p inhibitor
An mirVana let-7a-5p-specific inhibitor and negative control were purchased from Thermo Fisher Scientific and used according to the manufacturer’s instructions. The transfection procedure was performed using a Stemfect RNA Transfection Kit (ReproCell, Yokohama, Japan).
Lentiviral production
The use of viral vectors was approved by committees at Kyoto University and Chiba University. The full-length coding sequences of human CUX1 and RALB were cloned into the lentiviral vector CS2-Ubic-IG-GFP or CS2-Ubic-IB. Lentiviral production using the 293 T system was described previously4.
Bulk RNA-sequencing analysis
Total RNA was extracted using the microRNeasy Micro Kit. RNA-seq libraries were prepared according to the manufacturer’s protocol. Briefly, ~10 ng of total RNA was used for the cDNA synthesis using a SMART-Seq v4 Ultra Low Input RNA Kit for sequencing (Takara Bio). cDNA was fragmented using an S220 Focused-ultrasonicator (Covaris, Woburn, MA, USA). The cDNA library was then generated using a NEBNext® Ultra™ DNA Library Prep Kit for Illumina (New England BioLabs, Beverly, MA, USA). Finally, the NEBnext library size was estimated using a bioanalyzer with an Agilent High Sensitivity DNA Kit. Sequencing was performed using a HiSeq2500 (Illumina) or a NextSeq 500 (Illumina) platform with a single-read sequencing length of 60 bp. TopHat (version 2.1.1) was used to map to the reference genome (UCSC/hg19) with annotation data from iGenomes (Illumina). Gene expression levels were quantified using Cuffdiff (Cufflinks version 2.2.1) and expressed as fragments per kilobase of exon per million mapped sequence reads (FPKM).
Gene sets enrichment analysis (GSEA)
A GSEA was applied to screen pathways enriched in (1) let-7 (let-7a-5p and let-7g-5p) low or high imMKCLs (M35-1, clone 7, clone 7–3), (2) let-7 low imMKCLs of clone 7–3 or of clone 7, and (3) MOCK or RALB-overexpressing imMKCLs (clone 7) based on bulk RNA-seq datasets. The plots were generated using GSEA software (version 4.2.3) from the Broad Institute. We also re-analyzed bulk RNA-seq datasets from high-, intermediate- and low-quality MKCL clones12. The normalized gene expression matrix was input for the GSEA, and FDR and P-values < 0.05 were considered statistically significant.
Chromium 10x single-cell RNA-seq library construction
imMKCLs (clone 7) with distinct let-7a-5p activity were sorted separately and resuspended at 1000 cells/μL in PBS with 0.4% BSA (Fig. 3). The cells were loaded onto a Chromium Next Gel Beads-in-EMulsion (GEMs) Chip G Single Cell Kit (10 x Genomics, USA). GEM generation and barcoding, reverse transcription, and cDNA generation and library construction were performed following the manufacturer’s protocol (Chromium Next GEM Single Cell 3’ Reagent Kits v3.1 Dual Index, 10 x Genomics). Dual-indexed, single-cell libraries were pooled and sequenced in paired-end reads on a Novaseq6000 (Illumina).
Bioinformatics analysis
10 x Genomics-derived datasets were collected, and quality control was performed to filter out low-quality and contaminated cells. Generally, reads were pre-processed with the Cell Ranger pipeline v.3.0.2 (10 x Genomics). Downstream analysis and visualization were done using Seurat (version 4.0.5) implemented in R (version 4.1.1). After inspection of the quality control metrics, cells with > 15% of mitochondrial content or <2500 detected genes were excluded from the downstream analysis. We normalized and scaled the unique molecular identifier (UMI) counts using regularized negative binomial regression. Afterward, we performed linear dimensionality reduction (principal component analysis) and used the top 20 principal components to perform the unsupervised Uniform Manifold Approximation and Projection (UMAP) and clustering, which were computed at a range of resolutions from 1.2 to 0.05. The let-7a-5p high and low imMKCL populations were combined for all subsequent analyses. Cell clusters were identified using the FindClusters function from Seurat. Five clusters for DOX-ON samples were identified (resolution 0.2). The FindAllMarkers function implemented in Seurat was used to identify DEGs between different clusters. The Wilcoxon test was performed on each gene, and the P-values and adjusted P-values for statistical significance were computed. Genes with adjusted P-values <0.01 were considered significant. The g:Profiler was used for the identification of enriched functional terms from the GO.
Statistical analysis
Statistical analysis was performed using GraphPad Prism software (GraphPad Software, La Jolla, CA). Data are expressed as the mean standard error of the mean (SEM). Values of P < 0.05 were considered significant. Details of the sample size used, statistics, and statistical significance are indicated in each figure legend.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The raw sequence data have been deposited in the DNA Data Bank of Japan (DDBJ) database under the association number PRJDB15883, composing of the following subseries: DRA016355, DRA016356, DRA016432. The project is also available at the NCBI BioProject [http://www.ncbi.nlm.nih.gov/bioproject/] under the same association number. The processed data sets were deposited with the association codes E-GEAD-615, E-GEAD-616, E-GEAD-617. Source data are provided with this paper. The authors declare that all data supporting the findings of this study are available within the article and its Supplementary Information file. Source data are provided with this paper.
References
Waller, C. Case of uterine hemorrhage, in which the operation of transfusion was successfully performed. Lond. Med. Phys. J. 54, 273–277 (1825).
Watkins, W. M. The ABO blood group system: historical background. Transfus. Med. 11, 243–265 (2001).
Kliman, A., Gaydos, L. A., Schroeder, L. R. & Freireich, E. J. Repeated plasmapheresis of blood donors as a source of platelets. Blood 18, 303–309 (1961).
Takayama, N. et al. Transient activation of c-MYC expression is critical for efficient platelet generation from human induced pluripotent stem cells. J. Exp. Med. 207, 2817–2830 (2010).
Nakamura, S. et al. Expandable megakaryocyte cell lines enable clinically applicable generation of platelets from human induced pluripotent stem cells. Cell Stem Cell 14, 535–548 (2014).
Ito, Y. et al. Turbulence activates platelet biogenesis to enable clinical scale ex vivo production. Cell 174, 636–648.e618 (2018).
Sugimoto, N. et al. iPLAT1: the first-in-human clinical trial of iPSC-derived platelets as a phase 1 autologous transfusion study. Blood 140, 2398–2402 (2022).
Sugimoto, N. et al. Production and nonclinical evaluation of an autologous iPSC-derived platelet product for the iPLAT1 clinical trial. Blood Adv. 6, 6056–6069 (2022).
Chen, S. J., Sugimoto, N. & Eto, K. Ex vivo manufacturing of platelets: beyond the first-in-human clinical trial using autologous iPSC-platelets. Int J. Hematol. 117, 349–355 (2023).
Wang, H. et al. Decoding human megakaryocyte development. Cell Stem Cell 28, 535–549.e538 (2021).
Sun, S. et al. Single-cell analysis of ploidy and the transcriptome reveals functional and spatial divergency in murine megakaryopoiesis. Blood 138, 1211–1224 (2021).
Sone, M. et al. Silencing of p53 and CDKN1A establishes sustainable immortalized megakaryocyte progenitor cells from human iPSCs. Stem Cell Rep. 16, 2861–2870 (2021).
Mehta, A. & Baltimore, D. MicroRNAs as regulatory elements in immune system logic. Nat. Rev. Immunol. 16, 279–294 (2016).
Mullokandov, G. et al. High-throughput assessment of microRNA activity and function using microRNA sensor and decoy libraries. Nat. Methods 9, 840–846 (2012).
Miki, K. et al. Efficient detection and purification of cell populations using synthetic microRNA switches. Cell Stem Cell 16, 699–711 (2015).
Fujita, Y. et al. A versatile and robust cell purification system with an RNA-only circuit composed of microRNA-responsive ON and OFF switches. Sci. Adv. 8, eabj1793 (2022).
Sunohara, T. et al. MicroRNA-based separation of cortico-fugal projection neuron-like cells derived from embryonic stem cells. Front. Neurosci. 13, 1141 (2019).
Parr, C. J. et al. MicroRNA-302 switch to identify and eliminate undifferentiated human pluripotent stem cells. Sci. Rep. 6, 32532 (2016).
Lo, R. W. et al. The endoplasmic reticulum protein SEC22B interacts with NBEAL2 and is required for megakaryocyte α-granule biogenesis. Blood 136, 715–725 (2020).
Ver Donck, F. et al. Ribosome dysfunction underlies SLFN14-related thrombocytopenia. Blood 141, 2261–2274 (2023).
Seo, H. et al. A β1-tubulin-based megakaryocyte maturation reporter system identifies novel drugs that promote platelet production. Blood Adv. 2, 2262–2272 (2018).
Pariser, D. N. et al. Lung megakaryocytes are immune modulatory cells. J. Clin. Invest 131, e137377 (2021).
Campbell, R. A. et al. Human megakaryocytes possess intrinsic antiviral immunity through regulated induction of IFITM3. Blood 133, 2013–2026 (2019).
Davizon-Castillo, P. et al. TNF-α-driven inflammation and mitochondrial dysfunction define the platelet hyperreactivity of aging. Blood 134, 727–740 (2019).
Cunin, P. & Nigrovic, P. A. Megakaryocytes as immune cells. J. Leukoc. Biol. 105, 1111–1121 (2019).
Channappanavar, R. & Perlman, S. Pathogenic human coronavirus infections: causes and consequences of cytokine storm and immunopathology. Semin. Immunopathol. 39, 529–539 (2017).
Ren, X. et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell 184, 5838 (2021).
Bernardes, J. P. et al. Longitudinal multi-omics analyses identify responses of megakaryocytes, erythroid cells, and plasmablasts as hallmarks of severe COVID-19. Immunity 53, 1296–1314.e1299 (2020).
Takayama, N. et al. Generation of functional platelets from human embryonic stem cells in vitro via ES-sacs, VEGF-promoted structures that concentrate hematopoietic progenitors. Blood 111, 5298–5306 (2008).
Yuzuriha, A. et al. Extracellular laminin regulates hematopoietic potential of pluripotent stem cells through integrin β1-ILK-β-catenin-JUN axis. Stem Cell Res. 53, 102287 (2021).
Liu, C. et al. Characterization of cellular heterogeneity and an immune subpopulation of human megakaryocytes. Adv. Sci. (Weinh.) 8, e2100921 (2021).
Ruggeri, Z. M. Von Willebrand factor, platelets and endothelial cell interactions. J. Thromb. Haemost. 1, 1335–1342 (2003).
Romo, G. M. et al. The glycoprotein Ib-IX-V complex is a platelet counterreceptor for P-selectin. J. Exp. Med. 190, 803–814 (1999).
Kowalska, M. A., Rauova, L. & Poncz, M. Role of the platelet chemokine platelet factor 4 (PF4) in hemostasis and thrombosis. Thromb. Res. 125, 292–296 (2010).
Scheuerer, B. et al. The CXC-chemokine platelet factor 4 promotes monocyte survival and induces monocyte differentiation into macrophages. Blood 95, 1158–1166 (2000).
Xiao, Z., Visentin, G. P., Dayananda, K. M. & Neelamegham, S. Immune complexes formed following the binding of anti-platelet factor 4 (CXCL4) antibodies to CXCL4 stimulate human neutrophil activation and cell adhesion. Blood 112, 1091–1100 (2008).
Laarman, A. J. et al. Staphylococcus aureus staphopain A inhibits CXCR2-dependent neutrophil activation and chemotaxis. EMBO J. 31, 3607–3619 (2012).
Marques, R. E., Guabiraba, R., Russo, R. C. & Teixeira, M. M. Targeting CCL5 in inflammation. Expert Opin. Ther. Targets 17, 1439–1460 (2013).
Machlus, K. R. et al. CCL5 derived from platelets increases megakaryocyte proplatelet formation. Blood 127, 921–926 (2016).
Perng, Y. C. & Lenschow, D. J. ISG15 in antiviral immunity and beyond. Nat. Rev. Microbiol. 16, 423–439 (2018).
Grandvaux, N., tenOever, B. R., Servant, M. J. & Hiscott, J. The interferon antiviral response: from viral invasion to evasion. Curr. Opin. Infect. Dis. 15, 259–267 (2002).
David, J. M., Dominguez, C., Hamilton, D. H. & Palena, C. The IL-8/IL-8R Axis: a double agent in tumor immune resistance. Vaccines (Basel) 4, 22 (2016).
Emadi, S. et al. IL-8 and its CXCR1 and CXCR2 receptors participate in the control of megakaryocytic proliferation, differentiation, and ploidy in myeloid metaplasia with myelofibrosis. Blood 105, 464–473 (2005).
Higuchi, T. et al. Megakaryocytes derived from CD34-positive cord blood cells produce interleukin-8. Br. J. Haematol. 99, 509–516 (1997).
Orkin, S. H., Shivdasani, R. A., Fujiwara, Y. & McDevitt, M. A. Transcription factor GATA-1 in megakaryocyte development. Stem Cells 16, 79–83 (1998).
Mahabeleshwar, G. H. et al. The myeloid transcription factor KLF2 regulates the host response to polymicrobial infection and endotoxic shock. Immunity 34, 715–728 (2011).
Ning, S., Pagano, J. S. & Barber, G. N. IRF7: activation, regulation, modification and function. Genes Immun. 12, 399–414 (2011).
Kubota, T. et al. Virus infection triggers SUMOylation of IRF3 and IRF7, leading to the negative regulation of type I interferon gene expression. J. Biol. Chem. 283, 25660–25670 (2008).
Rodier, F. et al. Persistent DNA damage signalling triggers senescence-associated inflammatory cytokine secretion. Nat. Cell Biol. 11, 973–979 (2009).
Gorio, A. et al. Reparixin, an inhibitor of CXCR2 function, attenuates inflammatory responses and promotes recovery of function after traumatic lesion to the spinal cord. J. Pharm. Exp. Ther. 322, 973–981 (2007).
Yan, C. et al. Discovery and characterization of small molecules that target the GTPase Ral. Nature 515, 443–447 (2014).
Zhang, L., Sankaran, V. G. & Lodish, H. F. MicroRNAs in erythroid and megakaryocytic differentiation and megakaryocyte-erythroid progenitor lineage commitment. Leukemia 26, 2310–2316 (2012).
Lu, J. et al. MicroRNA-mediated control of cell fate in megakaryocyte-erythrocyte progenitors. Dev. Cell 14, 843–853 (2008).
Norfo, R. et al. miRNA-mRNA integrative analysis in primary myelofibrosis CD34+ cells: role of miR-155/JARID2 axis in abnormal megakaryopoiesis. Blood 124, e21–e32 (2014).
Opalinska, J. B. et al. MicroRNA expression in maturing murine megakaryocytes. Blood 116, e128–e138 (2010).
Bhatlekar, S. et al. miR-125a-5p regulates megakaryocyte proplatelet formation via the actin-bundling protein L-plastin. Blood 136, 1760–1772 (2020).
Edelstein, L. C. et al. MicroRNAs in platelet production and activation. J. Thromb. Haemost. 11, 340–350 (2013).
Lefrançais, E. et al. The lung is a site of platelet biogenesis and a reservoir for haematopoietic progenitors. Nature 544, 105–109 (2017).
Yeung, A. K., Villacorta-Martin, C., Hon, S., Rock, J. R. & Murphy, G. J. Lung megakaryocytes display distinct transcriptional and phenotypic properties. Blood Adv. 4, 6204–6217 (2020).
Gilles, M. E. & Slack, F. J. Let-7 microRNA as a potential therapeutic target with implications for immunotherapy. Expert Opin. Ther. Targets 22, 929–939 (2018).
Wells, A. C. et al. Modulation of let-7 miRNAs controls the differentiation of effector CD8 T cells. Elife 6, e26398 (2017).
Bodemann, B. O. & White, M. A. Ral GTPases and cancer: linchpin support of the tumorigenic platform. Nat. Rev. Cancer 8, 133–140 (2008).
Clough, R. R., Sidhu, R. S. & Bhullar, R. P. Calmodulin binds RalA and RalB and is required for the thrombin-induced activation of ral in human platelets. J. Biol. Chem. 277, 28972–28980 (2002).
Wersäll, A. et al. Mouse platelet ral GTPases control P-selectin surface expression, regulating platelet-leukocyte interaction. Arterioscler. Thromb. Vasc. Biol. 38, 787–800 (2018).
Chien, Y. et al. RalB GTPase-mediated activation of the IkappaB family kinase TBK1 couples innate immune signaling to tumor cell survival. Cell 127, 157–170 (2006).
Schmied, L., Höglund, P. & Meinke, S. Platelet-mediated protection of cancer cells from immune surveillance—possible implications for cancer immunotherapy. Front. Immunol. 12, 640578 (2021).
Jiang, D. et al. Post-GWAS functional analysis identifies CUX1 as a regulator of p16INK4a and cellular senescence. Nat. Aging 2, 140–154 (2022).
Kaufman, R. M. et al. Platelet transfusion: a clinical practice guideline from the AABB. Ann. Intern. Med. 162, 205–213 (2015).
van der Meijden, P. E. J. & Heemskerk, J. W. M. Platelet biology and functions: new concepts and clinical perspectives. Nat. Rev. Cardiol. 16, 166–179 (2019).
Ohnuki, M., Takahashi, K. & Yamanaka, S. Generation and characterization of human induced pluripotent stem cells. Curr. Protoc. Stem. Cell. Biol. https://doi.org/10.1002/9780470151808.sc04a02s9 (2009).
Acknowledgements
The authors thank Dr. Y. Fujita for technical support and critical discussion, Dr. M. Fukuyo, Dr. B. Rahmutulla, Dr. M. Koyama, Ms. K. Okita, Ms. K. Tsujimura, and Ms. I. Yoshino for technical support, and Dr. P. Karagiannis for critical reading. We thank the Japanese Red Cross Society Kanto-Koshinetsu Blook Blood Center for providing cord blood-derived CD34+ cells. This study was supported in part by the Project for Core Center for iPS Cell Research (JP22bm0104001) and the Project for Regenerative / Cellular Medicine and Gene Therapies under Grant Number JP22bm0704051(K.E.) and JP23bm1123028 (N.S.) from Japan Agency for Medical Research and Development (AMED); the CiRA Foundation Fund (K.E.); the iPS Cell Research Fund (S.J.C.); and a grant-in-aid for scientific research (S) (21H05047, K.E.), a grant-in-aid for early-career scientists (22K13124, S.J.C.), and a grant-in-aid for challenging research (23K18299, K.E.) from the Japan Society for the Promotion of Science (JSPS).
Author information
Authors and Affiliations
Contributions
S.J.C. designed and performed the majority of the experiments, analyzed the data, interpreted the results, prepared figures, and wrote the manuscript. K. Hashimoto and K.F. designed and performed experiments regarding miRNA switch screening. K. Hayashi prepared the miRNA switches. S.K.P., A.Y., W.-Y.Q. E.N., and M.A.K. assisted with the experiments. A.K. sequenced the bulk RNA-seq libraries and provided intellectual contributions. T.Y. and M.K. contributed to the bioinformatics analysis of the scRNA-seq. S.N. provided intellectual contributions. N.S. provided intellectual contributions and edited the manuscript. H.S. provided guidance on miRNA switch-related experiments and edited the manuscript. N.T. provided guidance on the experimental design, contributed to the bioinformatics analysis, and edited the manuscript. K.E. managed the overall project, proposed ideas, contributed to the experimental design and data interpretation, and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
S.J.C. and K.E. have applied a patent related to this study. K.E. was a founder of Megakaryon Co. Ltd. but with no stock currently. H.S. is a founder and an outside director of aceRNA Technologies Ltd. The interests of K.E. and H.S. were reviewed and are managed by Kyoto University in accordance with its competing interest policies. The other authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Alastair Poole and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, S.J., Hashimoto, K., Fujio, K. et al. A let-7 microRNA-RALB axis links the immune properties of iPSC-derived megakaryocytes with platelet producibility. Nat Commun 15, 2588 (2024). https://doi.org/10.1038/s41467-024-46605-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-46605-0
- Springer Nature Limited