Abstract
ANKRD11 (Ankyrin Repeat Domain 11) is a chromatin regulator and a causative gene for KBG syndrome, a rare developmental disorder characterized by multiple organ abnormalities, including cardiac defects. However, the role of ANKRD11 in heart development is unknown. The neural crest plays a leading role in embryonic heart development, and its dysfunction is implicated in congenital heart defects. We demonstrate that conditional knockout of Ankrd11 in the murine embryonic neural crest results in persistent truncus arteriosus, ventricular dilation, and impaired ventricular contractility. We further show these defects occur due to aberrant cardiac neural crest cell organization leading to outflow tract septation failure. Lastly, knockout of Ankrd11 in the neural crest leads to impaired expression of various transcription factors, chromatin remodelers and signaling pathways, including mTOR, BMP and TGF-β in the cardiac neural crest cells. In this work, we identify Ankrd11 as a regulator of neural crest-mediated heart development and function.
Similar content being viewed by others
Introduction
Congenital heart defects (CHDs) are the most common type of birth defect, with a global 1% incidence rate1,2. A significant portion of CHDs are conotruncal defects, which are caused by improper development of the heart outflow tract (OFT), an embryonic structure that connects the heart ventricles and pharyngeal arch arteries (PAAs)3,4. The OFT is remodeled from a single vessel into the base of the aorta and pulmonary artery during embryogenesis to separate the circulation of oxygenated and deoxygenated blood3,4. This process is orchestrated by interactions between different cell types of the embryonic heart: second heart field-derived cells, endocardial cells, and cardiac neural crest cells (CNCCs). Dysregulation of CNCC function is a primary cause of many conotruncal defects and contributes to many known multisystem developmental disorders4,5.
CNCCs originate at the dorsal neural tube border and migrate into the pharyngeal arches and OFT, where they facilitate the remodeling of the vessels and form the smooth muscle cell lining of the arteries3. CNCCs populating the OFT cushions fuse at the OFT midline to create the aorticopulmonary (AP) septum, which progresses from the distal edge of the OFT until it fuses with the interventricular septum, creating two distinct vessels of the aorta and pulmonary artery. In mice, OFT septation occurs between embryonic day (E) 11.5 and E13.56.
Neural crest development requires intricate spatiotemporal regulation of gene expression. Most of the research has focused on signaling pathways and transcription factors, while epigenetic regulation has been relatively understudied4,7,8,9. Nevertheless, several epigenetic regulators, which are implicated in various congenital disorders, including Coffin-Siris (BRG1), CHARGE (CHD7), and Williams (BAZ1B) syndromes, have been found to be essential for proper CNCC development7,10,11,12. Chromatin regulators are critical for the complex spatiotemporal regulation of gene expression that orchestrates the OFT remodeling, although much of the mechanism behind the epigenetic regulation of neural crest-mediated heart development is still largely unclear7.
ANKRD11 (Ankyrin repeat domain 11; previously known as ANCO1) is a chromatin regulator that interacts with histone acetylation modifying proteins, such as histone deacetylase HDAC3 and P/CAF (p300/CBP-associated factor) acetyltransferase complex subunits, to modulate gene expression13,14,15. Heterozygous pathogenic variants in ANKRD11 or 16q24.3 microdeletions containing ANKRD11 cause KBG syndrome (OMIM #148050), an autosomal dominant multisystem developmental disorder. Patients with KBG syndrome display global developmental delay, short stature, craniofacial defects, and intellectual disability16,17,18,19,20,21,22,23,24. About 40% of patients show cardiovascular defects, which include valve dysplasia and ventricular septal defects (VSD) as well as, less commonly, defects like aortic coarctation and patent ductus arteriosus23,25,26. Notably, these cardiovascular defects indicate a potential dysregulation of CNCC function4,5. However, the role of ANKRD11 in CNCCs or in cardiac development is not known.
Here, we demonstrate that ablation of Ankrd11 in the murine neural crest leads to OFT septation failure, a severe congenital cardiac defect termed persistent truncus arteriosus (PTA), as well as ventricular dilation and impaired ventricular contractility. We further show that deletion of Ankrd11 in the neural crest causes a delay in CNCC condensation in OFT cushions and failed AP septation as well as impairment of several signaling pathways, including mammalian target of rapamycin (mTOR), bone morphogenetic protein (BMP), and transforming growth factor-beta (TGF-β). This study identifies Ankrd11 as a crucial regulator of CNCC development and suggests CNCC dysregulation as a candidate cause for cardiac defects in KBG patients.
Results
Deletion of Ankrd11 in the neural crest leads to OFT defects
To investigate if Ankrd11 is expressed in CNCCs, we extracted Ankrd11 mRNA expression data from a murine embryonic day (E) 9.5-E10.5 trunk neural crest single-cell RNA sequencing (scRNAseq) dataset27. The results showed that Ankrd11 was broadly and robustly expressed in all neural crest cells, including early delaminating and non-cardiac Sox10+, Ets1+, Foxd3+ NCCs (Fig. 1a–c). Its expression was also observed in neural tube cells and central nervous system neurons, which supports and expands previous reports that demonstrate Ankrd11 expression in the brain and craniofacial tissues13,28 (Fig. 1b, c). Ankrd11 expression persisted in neural crest cells in which Foxd3 expression began to be downregulated, which signifies differentiation into cardiac derivatives (Fig. 1b). As all CNCCs showed the presence of Ankrd11 (Fig. 1c, d’), we further asked about the heterogeneity of its expression. Figure 1e shows that the majority ( ~ 64%) of CNCCs had a medium level of Ankrd11 expression, while 22% of CNCCs had the lowest and 14% of CNCCs had the highest expression. However, the high expression cells were mostly located at the more differentiated side of the CNCC cluster (Fig. 1d–d’), indicating a positive correlation of Ankrd11 expression along the pseudo time differentiation trajectory.
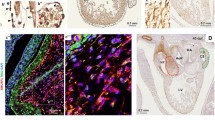
a UMAP embedding of mouse trunk neural crest and neural tube lineage at E9.5 and E10.5 (total 2121 cells). Data were re-analyzed from ref. 27. b mRNA expression of Ankrd11 and key markers for migratory and cardiac neural crest, abundance indicated by color intensity. c Ankrd11 expression level distribution in different cell types of mouse trunk neural crest and neural tube lineage at E9.5 and E10.5. d–d’ Expression of Ankrd11 and key markers in cardiac NCC cluster (92 cells). e Ankrd11 expression level distribution in cardiac NCC cluster. Dash-dotted, dotted, and dashed lines refer to the first (blue), second (gray), and third (red) quartiles (q), respectively. f, g Representative images of Ankrd11nchet (f) and Ankrd11ncko (g) embryos at E11.5 showing YFP fluorescence. h Schematic of OFT transverse section at E11.5. i–o Representative images of Ankrd11nchet (Ankrd11fl/WT;Wnt1Cre2, i–j”, m, n, and Ankrd11ncko (Ankrd11fl/fl;Wnt1Cre2, k–l”, o E11.5 OFT (i–m) or neural tube (n, o) sections processed for Basescope with Ankrd11 exon 7 mRNA probe (i–l”, n, o) and control negative probe (m) (red), and counterstained for cell nuclei (DAPI; blue). Solid outlines: OFT mesenchyme. * autofluorescent red blood cells. i’–l’ High magnification images of OFT mesenchyme (white dashed boxes in i–l). Dashed outlines: Ankrd11+ cells. i”–l” High magnification images of OFT myocardium (yellow dashed boxes in i–l). Dashed outlines: Ankrd11+ cells. n, o High magnification images of neural tube (NT) regions. Dashed outlines: Ankrd11+ cells. p Quantification for i-l for % Ankrd11+ cells in Ankrd11nchet (orange) and Ankrd11ncko (green) OFT mesenchyme. Two-tailed unpaired t-test (p = 0.0055). q Quantification of p for the distribution of Ankrd11+ puncta/cell in Ankrd11nchet OFT mesenchyme. **p < 0.01. Graphs represent mean ± s.e.m, n = 3 Ankrd11nchet and n = 3 Ankrd11ncko samples from two independent litters. Scale bars: 1 mm (f, g), 100 µm (i–l), 10 µm (i’-o). CNCC cardiac neural crest cells, CNS central nervous system, LV left ventricle, migr migratory, NCC neural crest cells, NT neural tube, OFT outflow tract, premigr pre-migratory, RV right ventricle. Source data are provided as a Source Data file.
To determine how the loss of Ankrd11 in the neural crest affects cardiac development, we knocked out Ankrd11 in the neural crest using the Ankrd11fl/fl; Wnt1Cre2 mouse model (herein referred to as Ankrd11ncko), which was previously shown to have aberrant craniofacial development28. In these mice, Cre is expressed in pre-migratory neural crest cells under the Wnt1 promoter, leading to the excision of Ankrd11 exon 7 in neural crest cells and their derivatives29,30. Neural crest cell lineage tracing was performed by breeding in the RosaYFPSTOP allele (Ankrd11fl/fl; RosaYFPSTOP/STOP; Wnt1Cre2) leading to STOP cassette excision and expression of YFP (yellow fluorescent protein)29,30. At E11.5 the YFP distribution in both the conditional heterozygote (Ankrd11nchet: Ankrd11fl/WT; Wnt1Cre2; RosaYFPSTOP/STOP) and conditional knockout (Ankrd11ncko: Ankrd11fl/fl; Wnt1Cre2; RosaYFPSTOP/STOP) embryos were characteristic of the neural crest lineage (Fig. 1f, g), which includes the derivatives of the pharyngeal arches as well as the dorsal root ganglia of the peripheral nervous system30.
To corroborate Ankrd11 expression in control CNCCs and confirm the loss of Ankrd11 in Ankrd11ncko CNCCs, we performed BaseScope, a single-molecule fluorescent in situ hybridization assay, with an Ankrd11-specific probe. In this assay, every detected fluorescent dot indicates a single Ankrd11 mRNA molecule. Transverse sections of E11.5 OFT mesenchyme, which is predominantly composed of CNCCs31, showed Ankrd11 expression in ~80% of Ankrd11nchet (control) and ~20% of Ankrd11ncko cells (p = 0.0055; Fig. 1i–l’, p). The residual Ankrd11 signal in Ankrd11ncko samples may be due to the non-CNCC-derived cells of the OFT mesenchyme or incomplete Cre-mediated recombination. Furthermore, the non-neural crest-derived OFT myocardium and neural tube tissue showed comparable Ankrd11 signals in control and Ankrd11ncko embryos (Fig. 1i”–l”, n, o). Notably, the negative probe did not yield a signal under similar experimental and imaging conditions (Fig. 1m). Ankrd11-expressing cells in the Ankrd11nchet OFT mesenchyme were further stratified based on the number of Ankrd11+ dots per cell, which showed a range of Ankrd11 expression (Fig. 1q). This is in line with the varied expression observed in the scRNAseq dataset (Fig. 1c–e). Together, these results support and extend our previous work, where we showed that Ankrd11 is expressed in cranial neural crest cells28.
Next, we examined how the loss of Ankrd11 in CNCCs affects heart morphology. Embryonic hearts at E18.5 were analyzed for anatomical anomalies using immunofluorescence analysis and micro-computed tomography (µCT). Homozygous ablation of Ankrd11 in the neural crest was perinatal lethal, in agreement with ref. 28, so only embryonic timepoints were analyzed. In comparison to Ankrd11WT embryos, which expressed wild-type Ankrd11 (Ankrd11WT/WT; Wnt1Cre2; RosaYFPSTOP/STOP, n = 9) and Ankrd11nchet embryos (n = 29), all Ankrd11ncko embryos (n = 40) exhibited a persistent truncus arteriosus (PTA), where the OFT failed to septate into the pulmonary trunk and aorta and remained a single vessel (Fig. 2a–c). This was accompanied by a ventricular septal defect (VSD). Since neural crest-specific ablation of one copy of Ankrd11 (Ankrd11nchet) did not cause apparent OFT septation defects, further analysis was done by comparing Ankrd11ncko to both Ankrd11nchet and/or Ankrd11WT embryos. Notably, the thymus, an organ located above the heart that receives neural crest contribution32, did not differ in size between Ankrd11nchet and Ankrd11ncko embryos (Fig. S1a–c). This demonstrates the specificity of neural crest-mediated ablation of Ankrd11 on distinct organ morphogenesis, such as the heart.
a–c Representative images of Ankrd11WT (Ankrd11WT/WT;Wnt1Cre2, a), Ankrd11nchet (Ankrd11fl/WT;Wnt1Cre2, b) and Ankrd11ncko (Ankrd11fl/fl;Wnt1Cre2, c) E18.5 heart coronal sections, immunostained for CNCC derivatives (YFP; green) and counterstained for cell nuclei (Hoechst; magenta). a’–c’. High magnification images of OFT regions (white dashed boxes in a–c) from the same or adjacent sections immunostained for alpha-smooth muscle actin (αSMA; white) and CNCC derivatives (YFP; green). d–e’. Representative 3D reconstructions of µCT scans of E18.5 Ankrd11ctrl (Ankrd11fl/fl, Ankrd11fl/WT and Ankrd11nchet d) and Ankrd11ncko (e) hearts and magnified images (d’, e’) of the OFT regions (from white dashed boxes in d, e). f, g Quantification of d-e for total ventricular volumes (f) and ventricular wall thicknesses (g) from Ankrd11ctrl (blue) and Ankrd11ncko (green) embryos. Two-tailed unpaired t-test (f, p = 0.0057); two-tailed multiple t-tests with Holm-Sidak multiple comparisons test (g). h–k Representative B-mode long-axis images of in utero echocardiography of Ankrd11ctrl (Ankrd11fl/fl, Ankrd11fl/WT, and Ankrd11nchet, h, i) and Ankrd11ncko (j, k) embryos during systole (h, j) and diastole (i, k). Dashed lines outline the heart and ventricles in each image. l–n Quantification of the ventricular area during systole (l), diastole (m), and percent area change between systole and diastole (n) in the left (LV) and right (RV) ventricles from Ankrd11ctrl and Ankrd11ncko embryos. Two-tailed multiple t-tests with Holm-Sidak multiple comparisons test (l, LV p = 0.012655, RV p = 0.013974; n, LV p = 0.000121, RV p = 0.000974). ns: not significant, *p < 0.05, ** p < 0.01, ***p < 0.001. Graphs represent mean ± s.e.m; n = 7 Ankrd11ctrl and 8 Ankrd11ncko (f); n = 8 Ankrd11ctrl, 7 Ankrd11ncko LV, 5 Ankrd11ncko RV (g); n = 3 Ankrd11ctrl and n = 3 Ankrd11ncko (l–n) samples. Embryos were taken from 3 independent litters. Scale bars: 500 µm. Ao aorta, BCT brachiocephalic trunk, LCC left common carotid artery, LS left subclavian artery, LV left ventricle, OFT outflow tract, PA pulmonary artery, RCC right common carotid artery, RS right subclavian artery, RV right ventricle, TA truncal artery, V ventricle, VSD ventricular septal defect. Source data are provided as a Source Data file.
Additional analysis revealed that Ankrd11ncko embryos had dysplastic truncal valves, visualized by PDGFRα immunostaining33 (Fig. S1d–i), with 3 out of 5 analyzed hearts showing enlarged valves when compared to Ankrd11nchet (Fig. S1f, g). Notably, analysis of the αSMA + (alpha-smooth muscle actin) smooth muscle layer of the truncal artery did not show a deficiency of CNCC-derived smooth muscle cells compared to the aortic and pulmonary arteries (Fig. S1j–p). Although the Ankrd11ncko distal region showed a decreased proportion of smooth muscle cells that are CNCC-derived (% αSMA + YFP + /αSMA + ; p = 0.010763; Fig. S1n), this was not due to their decreased number (#YFP + αSMA+; Fig. S1o), but rather due to an increase in non-CNCC-derived smooth muscle cells (#YFP-αSMA + ; p = 0.034171; Fig. S1p).
µCT imaging and subsequent 3D reconstruction confirmed PTA in Ankrd11ncko hearts, with the truncal artery originating from the right ventricle in all cases (Fig. 2d, e). Ankrd11ncko ventricles were ~1.8-fold larger (p = 0.0057) than Ankrd11ctrl (Ankrd11fl/fl, Ankrd11fl/WT, and Ankrd11nchet) (Fig. 2d–f), although ventricular wall thickness was unchanged (Fig. 2g). More detailed analysis of vessel branching identified an interrupted aortic arch phenotype to accompany the PTA (Fig. 2d’, e’)34. These results show that Ankrd11ncko embryos had severe OFT and heart development defects.
Ankrd11 ncko hearts show impaired contractility and function
To investigate whether the observed abnormalities affected cardiac function, we performed in utero echocardiography at E18.535 (Fig. 2 & S1; Supplementary Movie 1 & 2). In agreement with wall thickness measurements through µCT analysis, echocardiography confirmed that there was no difference in wall thicknesses between Ankrd11ncko and control hearts (Fig. S1r). Fetal heart rate was also unchanged (Fig. S1q). Chamber size was increased in the Ankrd11ncko hearts compared to controls (Fig. 2h–m). Specifically, the left ventricular (LV) and right ventricular (RV) areas in systole were increased by ~3-fold (p = 0.012655) and ~2.6-fold (p = 0.013974), respectively (Fig. 2h–l). Furthermore, LV and RV diameter in systole and RV diameter in diastole was increased by ~1.9-fold (p = 0.018753), ~2.4-fold (p = 0.005594), and ~1.7-fold (p = 0.015199), respectively (Fig. S1s, t). LV and RV area in diastole showed a non-significant trending increase in Ankrd11ncko embryos compared to controls (Fig. 2m; p = 0.06) and LV diameter in diastole showed a non-significant trending increase in Ankrd11ncko embryos compared to controls (Fig. S1t; p = 0.18). Finally, measures of cardiac contractility were reduced in Ankrd11ncko hearts when compared to controls. LV and RV percent area change decreased by ~1.4-fold (p = 0.000121) and ~1.3-fold (p = 0.000974), respectively (Fig. 2n), and LV and RV fractional shortening decreased by ~1.7-fold (p = 0.0406) and ~1.9-fold (p = 0.0307), respectively (Fig. S1u, v). These results indicate that Ankrd11 ablation in the neural crest leads to cardiac functional defects, specifically impaired ventricular contractility.
Ankrd11 ncko CNCCs show a mild defect in migration into the OFT
To determine how the loss of Ankrd11 affects CNCC fates, we first analyzed CNCC migration, proliferation, and apoptosis in the OFT at E10.5, prior to OFT remodeling6 (Fig. S2). Analysis of CNCC distribution along the proximal-distal gradient in the E10.5 OFT revealed ~1.4-fold fewer (p = 0.038597) YFP+ CNCCs specifically in the medial region of Ankrd11ncko OFTs compared to control (Ankrd11nchet) (Fig. S2a–g). No changes were found in the proliferative index of CNCCs expressing Ki67, a proliferation marker28 (%Ki67+YFP + /YFP+; Fig. S2a’–f’, h), or in the proportion of apoptotic YFP + TUNEL+ (terminal deoxynucleotidyl transferase dUTP nick end labeling) CNCCs (Fig. S2a”–f”, i). This suggests that the deficit in CNCC numbers in Ankrd11ncko OFTs was most likely due to a modestly reduced CNCC migration into the OFT rather than abnormal proliferation or apoptosis.
Ankrd11 ncko OFTs show failed septation
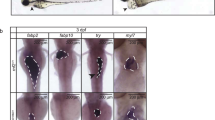
To determine the role of Ankrd11 in OFT remodeling, we analyzed the OFT at E11.5 and E12.5, when OFT septation begins6. At E11.5, OFTs in control hearts began to show significant remodeling, which was absent in the OFTs of Ankrd11ncko hearts (Fig. 3). In the distal region of E11.5 control OFTs, YFP+ CNCCs have formed the AP septum, which was densely filled by parallel F-actin filaments, visualized using phalloidin staining (Fig. 3a–c). In contrast, the AP septum was absent in the anatomically matched regions of E11.5 Ankrd11ncko OFTs (Fig. 3d, e). However, by E12.5, Ankrd11ncko OFTs formed an AP septum only at the very distal edge of the OFT, with the rest of the OFT unseptated (Fig. 3j–t).
a Schematic of OFT transverse sections undergoing different stages of remodeling along the proximal-distal axis from E11.5 to E12.5. In the proximal and medial OFT, CNCCs in green are condensed into tightly organized columns. In the distal region, streams of CNCCs from both OFT cushions migrated medially and fused to form the aorticopulmonary (AP) septum and two distinct vessels of the aorta (Ao) and pulmonary artery (PA). b–e Representative images of E11.5 Ankrd11nchet (b, c) and Ankrd11ncko (d, e) distal OFT transverse sections immunostained for CNCC and their derivatives (YFP; green) and F-actin (phalloidin; blue) and counterstained for nuclei (Hoechst; magenta). f–i’. Representative images of E11.5 Ankrd11nchet (f, g) and Ankrd11ncko (h, i) medial OFT sections. White dashed boxes are shown at higher magnification in f’–i’. * OFT cushion center. j–q Representative images of E12.5 Ankrd11nchet (j, k, n, o) and Ankrd11ncko (l, m, p, q) distal OFT sections. r–u’ Representative images of E12.5 Ankrd11nchet (r, s) and Ankrd11ncko (t, u) medial OFT sections. White dashed boxes are shown at higher magnification in r’–u’. v Schematic of Ankrd11nchet CNCC nuclei orientation within an OFT cushion at E11.5-E12.5. Nuclei oriented perpendicularly to the cushion center (pink cell) are closer to 90° while nuclei oriented in parallel (teal cell) are closer to 0°. Method adapted from ref. 36. Created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. w, x Quantification of CNCC nuclear orientation to OFT cushion center at E11.5 and E12.5 in medial regions between Ankrd11nchet (orange) and Ankrd11ncko (green) genotypes. Two-tailed multiple t-tests with Holm-Sidak multiple comparisons test (w, 90-70 p = 0.000226, 29-0 p = 0.000148). ns: not significant; p > 0.05 ***p < 0.001. Graphs represent mean ± s.e.m; n = 3 Ankrd11nchet and n = 3 Ankrd11ncko samples from at least 2 independent litters. Scale bars: 100 µm. Source data are provided as a Source Data file.
In medial regions of E11.5 control OFTs, CNCCs were organized into two tightly condensed columns within the cardiac cushions, visualized by spiral arrangements of CNCC nuclei and F-actin filaments in relation to the OFT center (Fig. 3f–g’). This condensation precedes CNCC migration to the OFT midline and their fusion to create the AP septum36. In contrast, in the anatomically matched regions of the Ankrd11ncko OFT, CNCCs showed less organization and condensation, with more varied orientation of nuclei and actin filaments in relation to the OFT cushion center compared to the control (Fig. 3h–i’). However, by E12.5 Ankrd11ncko medial region CNCCs showed more organized spiral nuclear and actin filament orientation that was more similar to control (Fig. 3r–u’). This was not observed in the unseptated distal region of E12.5 Ankrd11ncko OFTs, where the CNCC arrangement remained disorganized (Fig. 3n–q). Since the Ankrd11ncko OFTs remained unseptated by E18.5 (Fig. 2), this suggests that the septation process stalled in this region.
CNCC orientation to cushion center was quantified by binning the cells into categories by their nuclear orientation relative to the OFT cushion center (90°-70°, 69°-50°, 49°-30°, 29°-0°; Fig. 3v), adapted from ref. 36. Nuclei oriented perpendicularly to the cushion center are closer to 90° while nuclei oriented in parallel are closer to 0° (Fig. 3v). Since the remodeled distal region of control OFTs was morphologically distinct from the unremodeled Ankrd11ncko OFTs (Fig. 3b–e), we chose to compare medial regions for all subsequent quantifications, which had more comparable morphology (Fig. 3f–i). In control OFTs at E11.5 and E12.5, the 90°-70° bin contained the greatest number of YFP+ CNCCs (Fig. 3w–x). In contrast, all bins in the E11.5 Ankrd11ncko analysis had approximately similar numbers of YFP+ CNCCs (Fig. 3w). Compared to the control, E11.5 Ankrd11ncko OFTs had a notably decreased proportion of CNCCs in the 90°–70° bin (p = 0.000226) and an increased proportion of CNCCs in the 29°–0° bin (p = 0.000148; Fig. 3w). This indicates that Ankrd11ncko CNCCs showed impaired cell organization at E11.5. However, by E12.5, the nuclear orientation of Ankrd11ncko CNCCs was comparable to the control (Fig. 3x), suggesting that Ankrd11ncko CNCCs showed a delay in condensation. These results show that Ankrd11 ablation in the neural crest causes delayed CNCC organization in the OFT and failed AP septum formation.
Ankrd11 ncko CNCCs have impaired signaling pathways
To understand the mechanism behind the delay in the organization of Ankrd11-deficient CNCC and consequent AP septation failure, we sought to explore the signaling pathways that are known to regulate OFT development. BMP signaling has been shown to be activated along a distal-proximal gradient and impairment of its modulation dysregulated the timing of CNCC condensation36. To test for BMP signaling changes, we used immunostaining for phosphorylated Smad1/5/8, the intracellular effectors of Smad-dependent BMP signaling36,37,38. We observed pSmad1/5/8 signal in the medial and distal regions in control and Ankrd11ncko OFT cushions (Fig. 4a–p). In control medial OFT cushions at E11.5, we detected ~61% of YFP+ CNCCs with high pSmad1/5/8 signal (% pSmad1/5/8 + YFP + /YFP+ cells) (Fig. 4e–f, q). In contrast, we observed only ~33% of Ankrd11ncko pSmad1/5/8+ CNCCs in the medial region, a ~ 1.9-fold (p = 0.0035) decrease compared to control (Fig. 4g–h, q). However, by E12.5, we observed ~52% of Ankrd11ncko pSmad1/5/8+ CNCCs in the medial region, a ~ 1.6-fold (p = 0.035) increase compared to Ankrd11ncko medial region at E11.5 and statistically comparable (p = 0.9908) to control medial region at E12.5 (Fig. 4m–q). For the distal region, we only analyzed Ankrd11ncko cardiac cushions as controls have formed a septum in this area. At E11.5, we detected ~26% of Ankrd11ncko pSmad1/5/8+ CNCCs (Fig. 4c, d, r) compared to ~51% of Ankrd11ncko CNCCs at E12.5 in the unseptated distal region (Fig. 4k, l, r), which represents a ~ 1.9-fold (p = 0.0078) increase compared to the Ankrd11ncko distal region at E11.5. This suggests that Ankrd11ncko CNCCs show delayed BMP signaling, with reduced pSmad1/5/8 levels at E11.5 and normal levels at E12.5.
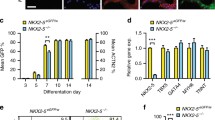
a–p Representative images of Ankrd11nchet (a, b, e, f, i, j, m, n) and Ankrd11ncko (c, d, g, h, k, l, o, p) distal and medial OFT cushions (outlined with white dashed lines) at E11.5 (a–h) and E12.5 (i–p) immunostained for CNCCs (YFP; green), pSmad1/5/8 (white) and counterstained for nuclei (Hoechst, magenta). q, r Quantification of a–p for percent of pSmad1/5/8+ CNCCs (% pSmad1/5/8 + YFP + /YFP+ cells) in Ankrd11nchet (orange) and Ankrd11ncko (green) genotypes in the medial region, and in the Ankrd11ncko genotype the distal region, at E11.5 and E12.5. Two-way ANOVA with Sidak’s multiple comparisons test (q, E11.5 Ankrd11nchet vs. E11.5 Ankrd11ncko p = 0.0035, E11.5 Ankrd11ncko vs. E12.5 Ankrd11ncko p = 0.035); two-tailed unpaired t-test (r, p = 0.0078). Graphs represent mean ± s.e.m; n = 3 Ankrd11nchet and n = 3 Ankrd11ncko samples from at least 2 independent litters. ns: not significant, *p < 0.05, **p < 0.01. Scale bars: 100 µm. Source data are provided as a Source Data file.
The mTORC1 signaling pathway has been recently implicated in OFT development, including the regulation of actin dynamics required for cell migration39. We analyzed phosphorylated ribosomal protein S6 (pS6) of the 40 s ribosomal subunit, a well-accepted readout of mTORC1 activation, and compared it to total S6 in E11.5 OFTs40,41. We observed pS6 and total S6 signals in the medial and distal regions in control and Ankrd11ncko OFT cushions (Fig. 5a–p, S3a–p). In control medial OFT cushions at E11.5, we observed ~63% pS6+ CNCCs (% pS6 + S6 + YFP + /S6 + YFP+ cells) (Fig. 5e, f, q). In contrast, we observed only ~37% pS6+ Ankrd11ncko CNCCs in the medial region, a ~ 1.7-fold (p = 0.0108) decrease compared to the control (Fig. 5e–h, q). However, by E12.5, we observed ~60% pS6+ Ankrd11ncko CNCCs in the medial region, a ~ 1.6-fold (p = 0.0182) increase compared to Ankrd11ncko medial region at E11.5, and statistically comparable (p = 0.2069) to control medial region at E12.5 (Fig. 5m–q). As described above, for the distal region, we only analyzed Ankrd11ncko cardiac cushions as controls have formed a septum in this area. In the distal region, we detected ~48% pS6+ Ankrd11ncko CNCCs at E11.5 (Fig. 5c, d, r), compared to ~76% pS6+ Ankrd11ncko CNCCs at E12.5 in the unseptated distal region (Fig. 5k, l, r), a ~ 1.6-fold (p = 0.0033) increase compared to E11.5. This suggests that Ankrd11ncko CNCCs show delayed mTORC1 signaling, with reduced pS6 levels at E11.5 and normal pS6 levels at E12.5.
a–p Representative images of Ankrd11nchet (a, b, e, f, i, j, m, n) and Ankrd11ncko (c, d, g, h, k, l, o, p) distal and medial OFT cushions (outlined with white dashed lines) at E11.5 (a–h) and E12.5 (i–p) immunostained for CNCCs (YFP; green), pS6 (white) and counterstained for nuclei (Hoechst, magenta). q, r Quantification of a–p for percent of pS6 CNCCs (% pS6 + S6 + YFP + /S6 + YFP+ cells) in Ankrd11nchet (orange) and Ankrd11ncko (green) medial OFT (q), and in the Ankrd11ncko distal OFT (r), at E11.5 and E12.5. Two-way ANOVA with Sidak’s multiple comparisons test (q, E11.5 Ankrd11nchet vs. E11.5 Ankrd11ncko p = 0.0108, E11.5 Ankrd11ncko vs. E12.5 Ankrd11ncko p = 0.0182); two-tailed unpaired t-test (r, p = 0.0033). Graphs represent mean ± s.e.m; n = 3 Ankrd11nchet and n = 3 Ankrd11ncko samples from at least 2 independent litters. ns: not significant, *p < 0.05, **p < 0.01. Scale bars: 100 µm. Source data are provided as a Source Data file.
The Smad2/3-dependent TGF-β pathway is also known to be involved in OFT development, with ablation of several pathway components causing OFT septation defects42,43,44,45. Therefore, to test for abnormalities in TGF-β signaling, we used immunostaining for phosphorylated Smad2/3, the intracellular effectors of Smad-dependent TGF-β signaling45. We observed pSmad2/3 signal throughout control and Ankrd11ncko OFT cushions (Fig. 6a–p). In control medial OFT cushions at E11.5, we observed that the parietal cushion (pc), positioned in the top-right of the OFT, had significantly higher pSmad2/3 signaling intensity compared to the septal cushion (sc), positioned in the bottom-left OFT, comprising ~55% CNCCs with high pSmad2/3 signal (% pSmad2/3 + YFP + /YFP+ cells) and ~33% pSmad2/3+ CNCCs, respectively (p = 0.030585; Fig. 6e, f, q). This asymmetrical pSmad2/3 staining persisted in E12.5 control medial OFT but was less pronounced compared to E11.5 (Fig. 6m, n, s). In contrast, we observed that parietal and septal cushions in the E11.5 Ankrd11ncko medial OFT had similar levels of pSmad2/3 signaling, with ~51% and ~48% pSmad2/3+ CNCCs in the parietal and septal cushions, respectively (Fig. 6g, h, q). However, by E12.5, Ankrd11ncko medial OFT showed a higher proportion ( ~ 63%) of pSmad2/3+ CNCCs in the parietal cushion compared to ~42% in the septal cushion (p = 0.002271; Fig. 6o, p, s). As described above, for the distal region, we only analyzed Ankrd11ncko cardiac cushions as controls have formed a septum in this area. In the distal region, at both E11.5 and E12.5, we detected comparable levels of pSmad2/3+ CNCCs between Ankrd11ncko parietal and septal cushions (Fig. 6c, d, k, l, r, t). These results suggest that Ankrd11ncko OFTs show delayed establishment of TGF-β signaling asymmetry, with abnormal symmetric pSmad2/3 pattern in parietal and septal cushions at E11.5, and normal asymmetric pattern at E12.5.
a–p Representative images of E11.5 Ankrd11nchet (a, b, e, f, i, j, m, n) and Ankrd11ncko (c, d, g, h, k, l, o, p) distal and medial OFT, in the parietal (PC) and septal (SC) cushions (outlined with white dashed lines) at E11.5 (a–h) and E12.5 (i–p) immunostained for CNCCs (YFP; green), pSmad2/3 (white) and counterstained for nuclei (Hoechst, magenta). q–t Quantification of a–p for percent of pSmad2/3 CNCCs (% pSmad2/3 + YFP + /YFP+ cells) in Ankrd11nchet (orange) and Ankrd11ncko (green) medial OFT (q, s) and in the Ankrd11ncko distal OFT (r, t), at E11.5 (q, r) and E12.5 (s, t), in the parietal and septal cushions. Two-tailed multiple t-tests with Holm-Sidak multiple comparisons test, medial (q, Ankrd11nchet p = 0.030585; s, Ankrd11nchet p = 0.004133, Ankrd11ncko p = 0.002271); two-tailed unpaired t-test, distal. Graphs represent mean ± s.e.m; n = 3 Ankrd11nchet and n = 3 Ankrd11ncko samples from at least 2 independent litters. ns: not significant, *p < 0.05, **p < 0.01. Scale bars: 100 µm. Source data are provided as a Source Data file.
We next sought to find other factors that may be asymmetrically expressed in OFT cushions. We focused on Cellular retinoic acid binding protein 2 (Crabp2), a protein that transports retinoic acid from the cytoplasm to the nucleus, which facilitates retinoic acid signaling46. In control medial OFT cushions at E11.5, we observed that the parietal cushion (pc) had lower Crabp2 levels compared to the septal cushion (sc), comprising 9% CNCCs with high Crabp2 signal (% Crabp2+YFP + /YFP+ cells) and ~19% Crabp2+ CNCCs, respectively (p = 0.046044; Fig. S4e, f, i). In contrast, Ankrd11ncko OFTs displayed asymmetric Crabp2 levels only in the distal region with ~22% Crabp2+ CNCCs in the parietal cushion compared to ~39% Crabp2+ CNCCs in the septal cushion (p = 0.0126; Fig. S4c, d, j). This suggests that Ankrd11ncko OFTs show established but spatially impaired Crabp2 protein level asymmetry at E11.5, and its failure to progress to the medial region may be a secondary consequence of AP septation failure.
Together, these results show that Ankrd11 ablation in the neural crest causes impaired BMP, mTORC1, and TGF-β signaling. Moreover, our results demonstrate that control OFTs exhibit asymmetric levels of expression of TGF-β and retinoic acid signaling pathway components and that this asymmetry is impaired in the Ankrd11ncko OFTs.
Ankrd11 ncko CNCCs show a diverse set of dysregulated targets
We next sought to explore the changes to known effectors of OFT development using a high throughput method. Since we have shown that the embryonic OFT has a high spatiotemporal restriction of signaling pathway activation (Figs. 4–6 and S4), we decided to employ Multiplexed error-robust fluorescence in situ hybridization (MERFISH), a single-cell spatial transcriptomic technique. We imaged 3 control and 3 Ankrd11ncko E11.5 anatomically matched embryo slices encompassing the medial OFT region using a custom panel of 140 RNA probes for genes (Supplementary Data 1) that are known to be important for OFT development and/or that were previously shown to be expressed in the OFT during the septation process in a scRNAseq dataset4,7,8,36,39,47,48,49,50,51,52. These include factors of important signaling pathways, extracellular matrix components, chemotactic ligands, transcription and chromatin remodeling factors, and cell type markers. Leiden analysis identified 10 clusters in the control and 13 clusters within the Ankrd11ncko OFT (Fig. 7a–c and Fig. S5). Clusters 11-13 contained a small number of cells that were only found in Ankrd11ncko OFTs (Fig. 7c). Clusters 2, 3, and 7 were identified to make up the OFT mesenchyme based on their location in the cushions and gene expression profiles (Fig. 7d–f). These three clusters showed high expression of the mesenchymal marker Postn, the mesenchymal and neural crest marker Pdgfra, and the neural crest marker Sox9[ 4,47,53,54. Clusters 2 and 3 also expressed the marker Penk (Fig. 7d, f), which was previously identified as a marker only expressed by CNCCs of a more mature phenotype that converge at the midline to form the AP septum53. Therefore, clusters 2, 3, and 7 were identified as CNCCs, although they may also include a small proportion of SHF-derived mesenchymal cells31. Ankrd11ncko OFTs had a smaller proportion of cells in cluster 2 and a larger proportion in cluster 3 when compared to control OFTs (Fig. 7e).
a Plot showing the spatial position of cells from representative transverse sections of E11.5 Ankrd11nchet (left) and Ankrd11ncko (right) embryos subjected to MERFISH with a custom 140 gene panel (Supplementary Data 1). A red bounding box defining the OFT is magnified and shown with cell clusters in the center. Cells in the OFT are colored according to Leiden clustering (clusters 1-13). b, c Plots showing the position of OFT cells from all samples in UMAP-space. Cells are colored according to Leiden clustering in b and according to genotype in c. d Heatmap plot showing scaled expression of Penk, Pdgfra, Sox9, and Postn in OFT cells from all samples. Cells are ordered according to their clustering. e Bar plot showing the proportion of all CNC-classified cells belonging to different genotypes and clusters (Ankrd11nchet: cluster 2, 518 cells; cluster 3, 306 cells; cluster 7, 152 cells. Ankrd11ncko: cluster 2, 475 cells; cluster 3, 632 cells; cluster 7, 249 cells). f Plots showing spatial positions of OFT cells colored according to normalized gene expression. Left column shows cells from representative Ankrd11nchet sample, right column shows cells from representative Ankrd11ncko sample. n = 3 Ankrd11nchet and n = 3 Ankrd11ncko samples from at least 2 independent litters. Source data are provided as a Source Data file.
Differential gene expression analysis identified 40 differentially expressed genes (DEGs) in cluster 2, 28 DEGs in cluster 3, and 9 DEGs in cluster 7 (Fig. 8). In cluster 2, two of the downregulated DEGs in Ankrd11ncko cells were Penk and Cxcl12 (Fig. 8a–c and Supplementary Data 2). As CNCCs progress through development, they undergo a transition from a mesenchymal to a vascular smooth muscle cell (VSMC) state to create the AP septum and line the arterial walls47,53. During this transition, the CNCCs upregulate their expression of Cxcl12, a chemokine encoding gene and a marker of immature VSMCs47,53. To this end, we observed the highest proportion of Cxcl12 expression in cluster 2, with low proportions in clusters 3 and 7 of the control samples, in line with their expression of the maturity state marker Penk. In all three clusters, Cxcl12 was downregulated in Ankrd11ncko cells (Fig. 8a–c and Supplementary Data 2). Conversely, we found upregulation of mesenchymal markers in Ankrd11ncko cells, including Postn, Pdgfra, and Vim47,53,54,55, in all three clusters, and Twist1 in clusters 2 and 7. Furthermore, there was downregulation of Heyl, a transcription factor downstream of the Notch pathway, and upregulation of Tbx20, a transcriptional repressor, in clusters 2 and 3 in Ankrd11ncko cells (Fig. 8a–c and Supplementary Data 2)47. Heyl has been previously predicted as a positive regulator, and Tbx20 as a negative regulator, of the mesenchymal to VSMC transition in the OFT47. These results suggest a delayed mesenchymal to VSMC transition of Ankrd11ncko CNCCs.
a Dotplots showing statistically significantly differentially expressed genes (DEGs) between the Ankrd11nchet and Ankrd11ncko genotypes for each cluster. Dots are colored according to average log fold-change of expression from Ankrd11nchet genotype (negative change indicates expression is lower in Ankrd11ncko compared to Ankrd11nchet, and a positive change indicates higher expression). The dot radius corresponds to the percent of cells in each cluster that express (i.e., show the presence of at least one transcript) the gene of interest. Genes displayed are ordered according to absolute average log2 fold-change (highest at the top, lowest at the bottom). b Heatmap plot showing scaled expression of differentially expressed genes between the Ankrd11nchet and Ankrd11ncko genotypes for each cluster. Cells are ordered according to their clustering. Genes displayed are ordered according to absolute average log2 fold-change (highest at the top, lowest at the bottom). c Violin plots showing scaled expression of selected differentially expressed genes between the Ankrd11nchet and Ankrd11ncko for each cluster. Genes were selected by taking the 8 highest absolute average log2 fold-change values from each marker group. n = 3 Ankrd11nchet and n = 3 Ankrd11ncko samples from at least 2 independent litters. Source data are provided as a Source Data file.
We observed DEGs in several signaling pathways known to be important for OFT development, including Wnt (Fzd4, Vangl2, Fzd1, Ctnnb1), BMP (Id2, Bmp4, Bmpr2, Acvr1), TGF-β (Tgfb2), Notch (Heyl, Jag1, Notch1), Hippo (Yap1), JNK (Jun), Retinoic acid (Rxra, Rarg), FGF (Fgfr2), EGFR (Erbb3), and JAK-STAT (Stat3)4,7,8,36,39,47,48,49,50,51,52 (Fig. 8a–c and Supplementary Data 2). We found downregulated expression of several growth factors and chemotactic ligands (Sema3c, Efna5, Tgfb2)36,56,57,58,59,60,61. In cluster 2, one of the most downregulated DEGs in Ankrd11ncko cells was Sema3c, which we have corroborated with single-molecule FISH (RNAscope) (Fig. 8a–c, S6, Supplementary Data 2). Semaphorin 3c (Sema3c) is a chemoattractant and aggregation factor that plays a major role in CNCC migration into the OFT, their condensation in the OFT cushions, and their convergence to create the AP septum36,56,57,58,59. Furthermore, clusters 2 and 3 showed downregulation in neuropilin-1 (Nrp1), a known receptor for Sema3c4, which may amplify the effects of the dysregulation. Other factors included ephrin A5 (Efna5) and transforming growth factor-beta 2 (Tgfb2), which were shown to control neural crest migration60 and OFT morphogenesis61, respectively (Fig. 8a–c and Supplementary Data 2). DEGs also included other transcription and chromatin remodeling factors important for OFT development such as Egr1, Hoxa3, Ets1, Gata4, Sox11, Foxc1, and Chd7[ 51,53,55, and cytoskeletal or extracellular matrix factors, such as Cdh11, Mmp14, Adamts1, and Adam1962 (Fig. 8a–c and Supplementary Data 2). These results suggest that loss of Ankrd11 affects multiple signaling pathways, transcription factors, and chromatin remodelers important for CNCC function and OFT morphogenesis, further supporting and expanding our results in Figs. 4–6.
Discussion
In this study, we demonstrate that Ankd11, a chromatin regulator and a causative gene for KBG syndrome, is a critical regulator of cardiac neural crest cell biology during embryonic heart development. Ablation of Ankrd11 in the murine neural crest leads to dysregulation of CNCC organization and intracellular signaling, leading to aberrant OFT development and heart function.
We show that conditional knockout of Ankrd11 in the neural crest leads to a failure of AP septation by CNCCs, creating a cardiac defect termed persistent truncus arteriosus (PTA) and an associated VSD. We also observed ventricular dilation without changes in wall thickness and decreased ventricular contractility. Ventricular dilation occurring without changes in wall thickness is indicative of eccentric ventricular hypertrophy, a compensatory mechanism to the dysfunctional ejection of blood from the ventricles, and consequent volume overload63,64,65. Ventricular dilation is associated with PTA in patients and animal models65,66,67,68,69. This can be observed as early as the fetal period due to truncal valve insufficiency, which causes volume overload through blood leakage from the truncal artery into the ventricles. Since we also observed truncal valve dysplasia in Ankrd11ncko hearts, this may indicate impaired valve function70. Notably, our results also show greater right ventricle dilation compared to the left ventricle. We postulate that this may be caused by the right ventricular origin of the truncal artery, which may force the right ventricle to be disproportionately impacted by the volume overload from a regurgitant truncal valve. Therefore, our results suggest that the ventricular dilation in Ankrd11ncko embryos is a compensatory mechanism at least in part due to PTA.
PTA in patients leads to congestive heart failure or cyanosis without surgical intervention66,71. Genetic models of PTA die in utero or perinatally, although the causes may be difficult to determine due to the multisystemic effects of their mutation39,72,73,74. While it was previously suggested that Ankrd11ncko neonates die due to cleft palate28, our results suggest that a potentially fatal heart dysfunction may also contribute to their death.
At the cellular level, our results suggest that OFT septation failure in the Ankrd11ncko mice is primarily caused by impairment in CNCC organization within the OFT, and not by gross defects in CNCC migration into the OFT, proliferation, or survival. This differs from other genetic models of cardiac neural crest deficiency, where the cells are not able to infiltrate the most proximal regions of the OFT, indicating a severe migration defect69. Instead, we observed a delay in condensation in the Ankrd11ncko OFT cushions and failure to form the AP septum. These conclusions support and contrast other studies on Ankrd11. We have previously shown that Ankrd11ncko embryos display spatiotemporal impairment of cell organization and maturation of craniofacial neural crest-derived bone, suggesting that Ankrd11 ablation in the neural crest causes the most dysregulation at later stages of craniofacial bone remodeling28. However, we also showed a decrease in cell proliferation in some, but not all, craniofacial regions, with no difference in apoptosis28. Moreover, our work with Yoda mice, which carry a heterozygous splice site-like mutation in Ankrd11, or with mice that have Ankrd11 knocked down or out in neural stem cells, shows that Ankrd11 regulates embryonic neural stem cell proliferation13,75. We and others have also shown that Yoda mice or mice with Ankrd11 knockdown in cortical progenitors and their derivatives have impaired neurogenesis and neuron positioning, which may suggest altered neuronal migration13,76. This was recently corroborated in our study, which showed that Ankrd11 knockout in neural stem cells leads to severe neuroblast migration defect in the rostral migratory stream75. Together, the results presented in this study and previous reports suggest that Ankrd11 may have unique roles in each cell type it is expressed in and that Ankrd11’s function may depend on cellular identity and environment.
Ankrd11ncko CNCCs show dysregulation of several signaling pathways, which may contribute to the observed delay in condensation and failure to form the AP septum. The TGF-β superfamily plays an important role in OFT development. Neural crest conditional knockout of TGF-β receptors, including transforming growth factor-beta receptor type 1 (Tgfbr1) and type 2 (Tgfbr2) results in PTA with interrupted aortic arch as well as varying effects on cell number, apoptosis, and smooth muscle formation42,77. Of the three TGF-β ligands, only Tgfb2 global knockout results in cardiac defects including VSD and double outlet right ventricle61. Ablation of downstream factors Smad4 and Smad7 that are involved in the Smad2/3-dependent TGF-β and Smad1/5/8-dependent BMP pathways also causes OFT defects45,78,79. While Tgfb2 expression was downregulated in Ankrd11ncko CNCCs in MERFISH results, it is unclear how this contributes to pSmad2/3 signaling abnormalities in the Ankrd11ncko CNCCs. Here, we have identified asymmetric pSmad2/3 signaling by CNCCs in control medial OFTs. Notably, pSmad2/3 expression was symmetric in E11.5 and asymmetric in E12.5 medial, but not distal, Ankrd11ncko OFT. Since we did not analyze control distal OFT cushions just prior to septation, it is unknown whether pSmad2/3 asymmetry is normally established distally.
Since Ankrd11ncko CNCCs were able to establish an asymmetric distribution of Crabp2 and not of pSmad2/3 in distal E11.5 OFT, this suggests that loss of Ankrd11 in the neural crest impairs specific asymmetrically activated signaling pathways, rather than impairing all asymmetry within the CNCC population. Notably, as MERFISH results identified dysregulated expression of the Rxra and Rarg receptors in the Ankrd11ncko OFT, this suggests that retinoic acid signaling may be dysregulated through a different mechanism in Ankrd11ncko CNCCs.
However, nothing is known about the mechanism behind the pSmad2/3 and Crabp2 asymmetry within CNCCs or its downstream effects on AP septum formation. Left-right asymmetry is crucial for proper embryo development, and several factors are known to contribute to cardiac left-right asymmetry, including the TGF-β superfamily4. Activin-like TGF-β receptors Acvr2b and CFC1 and ligand Gdf1 are involved in left-right patterning in heart development80,81,82, and global knockouts of these genes lead to cardiac defects such as transposition of the great arteries. Our results show further evidence that the TGF-β pathway contributes to OFT asymmetry during OFT septation. There is also evidence of left-right asymmetry within the CNCC population. Gandhi et al.83 observed that ablation of the right cardiac neural folds in chick, which removes CNCCs from the right side of the embryo, produced a more severe septation impairment compared to ablation of the left cardiac neural folds.
A subfamily within the TGF-β superfamily, the BMP pathway is a well-known regulator of cardiac neural crest development, with manipulation of its different components also leading to variable phenotypes4. Neural crest-specific ablation of the Activin-type BMP receptor Acvr1 causes decreased CNCC migration into the OFT, failure of OFT septation, and impaired smooth muscle formation69, neural crest-specific heterozygous deletion of the BMP receptor Bmpr2 causes an overriding aorta84, while neural crest-specific deletion of Bmpr1a leads to septation failure with normal CNCC migration74. The downregulation of Bmp4, Bmpr2, and Acvr1 that was found in Ankrd11ncko MERFISH results may contribute to the decreased pSmad1/5/8 signaling.
Neural crest-specific ablation of Ctdnep1, a BMP inhibitor, causes an opposing phenotype to the defects observed in the Ankrd11ncko embryos, specifically premature CNCC condensation and asymmetric AP septum formation, as well as increased pSmad1/5/8 activity36. Furthermore, the BMP pathway was found to at least in part regulate Sema3c expression36, a crucial factor for CNCC condensation that was downregulated in the Ankrd11ncko OFT in MERFISH and RNAscope results. Sema3c is a glycoprotein that is secreted within the OFT and pharyngeal arches, and is considered a chemoattractant guidance signal to promote CNCC migration into these structures, their condensation within the cardiac cushions and their migration to create the AP septum36,56,57,58,59. Overactivation of BMP signaling in CNCCs causes increased Sema3c expression, contributing to an overcondensation phenotype36. Our findings support these results by showing that a reduction in BMP signaling in Ankrd11ncko OFT mesenchyme at E11.5 correlates with delayed CNCC condensation since both the pSmad1/5/8 signal and condensation were restored at E12.5. Together, this provides evidence that the fine-tuning of BMP signaling is important for OFT condensation.
Recent studies have shown that mTOR signaling contributes to neural crest development. mTOR deletion causes decreased CNCC migration, proliferation, and differentiation, leading to PTA39. mTOR deletion in both cranial and cardiac neural crest causes decreased F-actin levels and consequent defects in cytoskeletal dynamics, which were attributed to mTORC1 pathway dysregulation39,85. mTORC1 is known to regulate cell migration in multiple contexts, with functions in cytoskeletal and extracellular matrix remodeling as well as focal adhesion formation86,87,88. Notably, Ankrd11 knockdown in murine embryonic cortical neuron cultures was previously shown to reduce pS6 levels, thus implicating mTORC1 signaling76.
Together, our study confirms mTORC1 and identifies TGF-β and BMP signaling pathways to be affected during OFT septation by the loss of Ankrd11. Furthermore, our MERFISH results identified other pathways that may be dysregulated in the Ankrd11ncko OFT, including Wnt, Notch, and Retinoic acid pathways.
From a clinical perspective, cardiac defects are detected in up to 40% of KBG syndrome patients23,25,26. Moreover, ANKRD11 variants are reported in large-scale exome and genome sequencing studies of gene‐disease associations in CHD, including conotruncal defects89,90,91,92. The reported defects include aortic coarctation, patent ductus arteriosus, valve dysplasia, and atrial and ventricular septal defects23,25,26. Such defects are also found in many other neural crest-mediated developmental disorders, including DiGeorge and Noonan syndromes5. Our results demonstrate a critical and direct role for ANKRD11 in regulating cardiac neural crest-mediated heart development and function. While our work has not tested the role of ANKRD11 in non-neural crest-derived heart tissues, our results suggest neural crest involvement in KBG syndrome cardiac defects.
Methods
Animal models
All animal use was approved by the Animal Care Committee of the University of Alberta in accordance with the Canadian Council of Animal Care policies. All mice were housed in a University of Alberta Animal Facility and serviced by Health Sciences Laboratory Animal Services (HSLAS). Mice were maintained on a 14/10 h light/dark cycle, at 21–23 °C room temperature, and 40–70% humidity. Food and water were provided ad libitum. Embryos at developmental ages between E10.5-18.5 and of either sex were used for all experiments.
Ankrd11fl/fl mice, where exon 7 of the Ankrd11 gene was flanked by LoxP sites, were derived from the Ankrd11TM1a(EUCOMM)Wtsi//IcsOrl (Ankrd11tm1a) sperm (EM:07651, the European Mouse Mutant Archive—Infrafrontier) as described in ref. 28. Ankrd11fl/fl mice were bred to hemizygous Wnt1Cre2+ mice (B6.Cg-E2f1Tg(Wnt1-cre)2Sor/J, stock # 022501, Jackson Laboratories)30, to create the Ankrd11fl/fl;Wnt1Cre[ 2 mice. These were used for µCT analysis. Neural crest cell lineage tracing was performed by breeding in the RosaYFPSTOP allele using the RosaYFPSTOP/STOP mice (B6.129X1-Gt(ROSA)26Sortm1(EYFP)Cos/J, stock # 006148, Jackson Laboratories)93. These mice were used for all other analysis. The mouse line was maintained by crossing Ankrd11WT/WT;RosaYFPSTOP/STOP;WntCre2 with Ankrd11fl/fl;RosaYFPSTOP/STOP mice. Resulting Ankrd11fl/WT;RosaYFPSTOP/STOP;WntCre2 were timed mated with Ankrd11fl/fl;RosaYFPSTOP/STOP or Ankrd11fl/WT;RosaYFPSTOP/STOP mice to generate Ankrd11WT/WT;RosaYFPSTOP/STOP;Wnt1Cre2+ (Ankrd11WT/WT), Ankrd11fl/fl;RosaYFPSTOP/STOP (Ankrd11fl/fl) and Ankrd11fl/WT;RosaYFPSTOP/STOP (Ankrd11fl/WT), Ankrd11fl/WT;RosaYFPSTOP/STOP;Wnt1Cre2 (Ankrd11nchet) as well as Ankrd11fl/fl;RosaYFPSTOP/STOP;Wnt1Cre2 (Ankrd11ncko). Mice were mated in the evenings and a plug was determined in the morning, with a positive plug considered as E0.25.
All mice were bred and genotyped using the following primers: Ankrd11: forward, 5’-CTGTCTCAGAGAGGAGAGTGAGGAGGAC-3’; reverse, 3’- TACCTTACACCCTGAGACGGCGTC-5’, 34 cycles of 94 °C–30 s, 62 °C–45 s, 72 °C–60 s. Pan-Cre: forward, 5’-TTCCCGCAGAACCTGAAGATG -3’; reverse, 3’- CCCCAGAAATGCCAGATTACG-5’; control forward primer 5’-AACAACAATGGCACAACCTAAT-3’; control reverses 3’-ACTTTCTCCCCACCCGTCTA-5’; 35 cycles of 94 °C–15 s, 60 °C–30 s, 72 °C–90 s. YFPSTOP: wild-type, 5’-GGAGCGGGAGAAATGGATATG-3’; common, 5’- AAAGTCGCTCTGAGTTGTTAT-3’; mutant, 5’-AAGACCGCGAAGAGTTTGTC-3’, 35 cycles of 94 °C–15 s, 65 °C–15 s, 72 °C–60 s.
Reagents
Primary antibodies and stains
Anti-Crabp2 (Proteintech, 10225-1-AP, Lot 00051203, 1:200), anti-eGFP (Abcam, ab13970, Lot GR3361051-16, 1:2000), anti-Ki67 (BD Pharmingen, 556003, Lot 1119219, 1:500), anti-αSMA (Sigma, A2547, Lot 099M4848V, 1:1000), anti-PDGFRα (R&D Systems, AF1062, Lot HMQ0220101, 1:500), anti-pSmad1/5/8 (Cell Signaling, 13820, Lot 4, 1:300), anti-pSmad2/3 (Cell Signaling, 8828, Lot 8, 1:300), anti-S6 (Cell Signaling, 2317, Lot 13, 1:600), anti-pS6 (phosphoSer240/244; Cell Signaling, 2215, Lot 18, 1:600), biotinylated IB4 (Vector Laboratories, VECTB1205, Lot ZB1017, 1:1000), Phalloidin-iFluor647 (Abcam, ab176759, Lot GR3279773-9, 1:1000). If the antibody was of mouse origin, then a M.O.M. Kit (Vector Biolabs) with appropriate streptavidin incubation (please see information below) was used according to instructions.
Secondary antibodies
Anti-chicken-Alexa488 (Jackson, 703-545-155, Lot 162189, 1:1000), anti-chicken-Alexa647 (Jackson, 703-605-155, Lot 153969, 1:1000), anti-goat-Alexa647 (Jackson, 705-605-147, Lot 154191, 1:1000), anti-rabbit-Alexa647 (Jackson, 711-605-152, Lot 154880, 1:1000), streptavidin-Cy3 (Jackson, 016-160-084, Lot 168280(2), 1:1000), streptavidin-Cy5 (Jackson, 016-170-084, Lot 151872, 1:1000).
Micro-computed tomography (µCT)
Embryos were dissected at E18.5 and fixed overnight in 4% PFA, followed by 48 h in Lugol’s iodine solution (Electron Microscopy Sciences, 26658-04), as per94. Embryos were imaged using a Milabs UHT-µCT scanner (Milabs, Utrecht, The Netherlands) of the School of Dentistry using the following parameters: step angle: 0.1 degrees, voltage: 50 kV, current: 0.24 mA, exposure time: 75 ms. Scans were exported as NII files, reconstructed with 15 µm voxel size, and analyzed using 3D Slicer (https://www.slicer.org/)95 (version 5.0.3) for ventricular volumes and wall thicknesses.
In utero echocardiography
Fetal cardiac structure and function were assessed by echocardiography using a 32 MHz linear array transducer (Vevo 3100, Visualsonics, Canada). Dams pregnant with E18.5 embryos were sedated with 2.5% isoflurane and maintained at 1.25–1.5% isoflurane using 0.8 L/min of oxygen. The abdomen was shaved and treated with depilatory cream. In a supine position, limbs were fixed to electrodes with tape and conductive electrode gel (SigmaGel, Parker Labs), and electrocardiogram, respiratory rate, and body temperature were continuously recorded. Body temperature was maintained with a heated table surface and heat lamp, and a heated ultrasound transmission gel was used. The total duration of all procedures did not exceed 1 h per dam and was performed by a single operator.
Each fetus was hand scanned using B-mode images to identify basic cardiac structure; fetuses with evident truncus arteriosus and ventricle septal defects (VSD) were preliminarily identified as Ankrd11ncko. Color Doppler images were then used to confirm VSD and truncus arteriosus. The area of the ventricular chambers was measured along the short and long-axis of the fetus using B-Mode images. Changes between the ventricular end-diastolic area and end-systolic area were used to calculate fractional area change (FAC) in the right and left ventricles. Using M-mode images in the short axis of the fetus, ventricular wall thicknesses and chamber diameters were measured in end-systole and end-diastole. Changes between the end-diastolic and end-systolic ventricular diameters were used to calculate fractional shortening. Finally, heart rate was calculated by counting heartbeats over two separate 10 s B-mode images. After imaging, the dam was euthanized with CO2, and fetuses were dissected to confirm the identification of the Ankrd11ncko and control (Ankrd11fl/fl, Ankrd11fl/WT and Ankrd11nchet) embryos.
Immunohistochemistry
Tissue preparation
Pregnant dams were euthanized with CO2 when embryos were at E10.5, E11.5, E12.5 and 18.5. Whole embryos (E10.5-E12.5) or dissected hearts (E18.5) were fixed in 4% paraformaldehyde (PFA; Electron Microscopy Sciences, 15710) for 24 h at 4 °C followed by three changes of 30% sucrose in 1xPBS every 24 h. Samples were embedded and flash frozen in an optimal cutting temperature (OCT) compound (ThermoFisher) and sectioned on a cryotome (CM1950, Leica Biosystems, Germany) to 14 µm slices for IHC, BaseScope or RNAscope assays and at 10 µm for MERFISH assay. The slices were transferred onto glass microscope slides and stored at –80 °C for IHC, BaseScope, or RNAscope assays or dehydrated with ethanol as per Vizgen® MERFISH manufacturer’s instructions and stored at 4 °C for a maximum 1 week.
Immunofluorescence and apoptosis detection
Sections were processed as described in96,97,98. Briefly, sections were dried at 37 °C, washed with phosphate-buffered saline (PBS), and blocked for 1 h (5% bovine serum albumin [BSA, Jackson ImmunoResearch, 001-000-162], 0.3% Triton-X100 in PBS), then incubated with primary antibodies listed in “Reagents” section (in 5% BSA) overnight at 4 °C. Sections were incubated with secondary antibodies listed in the “Reagents” section for 1 h at room temperature. If the antibody was of mouse origin, then a Vector Laboratories M.O.M. Kit was used according to instructions. Samples were counterstained with the nuclear stain Hoechst (33217; Riodel-De Haen Ag) at 1:1000 and mounted with Fluoromount-G mounting media (ThermoFisher, 00-4958-02).
Apoptotic cells were detected through the TUNEL (Terminal deoxynucleotidyl transferase dUTP nick end labeling; ApopTag Red In Situ Apoptosis Detection Kit, Millipore, S7165) assay according to the manufacturer’s protocol with modifications. After immunohistochemistry, sections were post-fixed in 4% PFA for 10 min at room temperature, followed by 5 min incubation in 2:1 ethanol:acetic acid at –20 °C. After washing, 50 µL per slide of equilibration buffer was applied for 10 s, followed by 100 µL of working strength TdT (terminal deoxynucleotidyl transferase) enzyme for 1 h at 37 °C. Slides were washed with 400 µL of stop/wash buffer for 10 min and 100 µL of anti-digoxigenin/rhodamine conjugate was applied for 30 min. Samples were counterstained with the nuclear stain Hoechst at 1:1000 and mounted with Fluoromount-G mounting media.
RNAscope and BaseScope
RNAscope was performed according to refs. 96,98 and manufacturer’s protocol with modifications (RNAscope Multiplex Fluorescent Reagent Kit, ACDBio). Briefly, sections were dried at 37 °C, dehydrated with ethanol, rehydrated with PBS, and incubated in 15 µl of 10% Protease IV (322340) diluted in PBS for 10 min at 37 °C. Sections were then washed and incubated with mRNA probes for 2 h at 40 °C, followed by consecutive incubations with AMP-1 to 3 (320852-320854) and AMP-4B (320856) or AMP-4C (320857). We used the Sema3c-C3 probe (441441-C3) and negative control provided by the manufacturer (320871). After RNAScope, samples were blocked for 1 h (5% BSA, 0.3% Triton-X100 in PBS), then incubated with primary antibodies listed in “Reagents” section (in 5% BSA) overnight at 4 °C. Sections were incubated with secondary antibodies listed in the “Reagents” section for 1 h at room temperature. Samples were counterstained with the nuclear stain Hoechst at 1:1000 and mounted with Fluoromount-G mounting media.
Basescope was performed according to the manufacturer’s protocol with modifications (BaseScope v2 Kit-Red, ACDBio). Sections were taken directly from –80 °C and washed with 1x PBS 2 times; sections were left in the 3rd wash for 5 min, shaking gently. Sections were baked at 60 °C for 30 min and post-fixed with 4% paraformaldehyde (15710, Fisher) at 4 °C for 15 min. Sections were dehydrated for 5 min with 50% and 70% ethanol, and 2 × 5 min with 100% ethanol at RT. Ethanol was evaporated, and sections were treated with hydrogen peroxide (322381, ACDBio) for 10 min at RT. Sections were washed once with water, then boiled for 5 min in 1x Target Retrieval (322000, ACDBio). Sections were washed twice with water, shaking gently. Sections were treated with 100% ethanol for 5 min and dried at RT overnight. Samples were treated with 1:3 Protease III (322337, ACDBio) diluted 1x PBS at 40 °C for 25 min. Sections were washed twice with water.
Sections were incubated with either the probe Ankrd11-C1 (1091241-C1, ACDBio), positive or negative control probes supplied with the kit for 2 h at 40 °C. Slides were washed 2 × 2 min with 1X RNAScope Wash Buffer (310091, ACDBio), followed by consecutive incubations with AMP-1 to 8; Amps 1-2, 4, 5 were incubated for 30 min at 40 °C, Amps 3 and 6 were incubated for 15 min at 40 °C, Amp 7 was incubated for 30 min at RT, and Amp 8 was incubated for 15 min at RT (323911-323918, ACDBio). Sections were washed 2 × 2 min with 1x RNAScope Wash buffer in between each Amp incubation. FastRed solution (1:60 Fast B: Fast A, 322918, ACDBio) was used to detect the signal; FastRed solution was added to sections for 10 min at RT followed by 2 × 2 min 1x PBS wash. Samples were counterstained with nuclear stain DAPI (320858, ACDBio), washed for 3 × 5 min with 1x PBS, and mounted with Fluoromount-G mounting media.
MERFISH
Probe design and sample preparation
A total of 140 genes were identified and evaluated using the MERSCOPE® Gene Panel Design Portal available at Vizgen (portal.vizgen.com) to ensure that each gene was suitable in length for probe binding and that abundance was below the recommended threshold to avoid optical crowding during imaging.
Paraformaldehyde-fixed (PFA) frozen mouse embryos were embedded in optimal cutting temperature (OCT) compound and sectioned into 10 µm thick sections on a cryostat at –20 °C for placement on fiducial bead-coated MERSCOPE Slides (Vizgen 20400001). The tissue slices were washed three times with 5 mL 1x PBS and incubated with 70% ethanol at 4 °C overnight for tissue permeabilization.
Antibody staining and probe hybridization
Cell boundary staining was performed using Vizgen’s Cell Boundary Staining Kit (Vizgen 10400009) per the user guide for fresh- and fixed-frozen sample preparation. Briefly, samples were washed with 5 mL 1x PBS and then blocked for 1 h at room temperature in Blocking Buffer C Premix (Vizgen 20300100) with RNase inhibitor (New England Biolabs M0314L) added at 1:20 dilution. For primary antibody staining, samples were incubated for 1 hat room temperature with Cell Boundary Primary Stain Mix (Vizgen 20300010) at 1:100 dilution and RNase inhibitor (New England Biolabs M0314L) at 1:20 dilution in blocking buffer, followed by three washes with 5 mL 1x PBS. Samples were then incubated for 1 h at room temperature with Cell Boundary Secondary Stain Mix (Vizgen 20300011) at 1:33 dilution and RNase inhibitor (New England Biolabs M0314L) at 1:20 dilution in blocking buffer, then washed three times with 5 mL 1x PBS, post-fixed in 5 mL 4% paraformaldehyde in 1x PBS for 15 min at room temperature, and washed twice with 5 mL 1x PBS. Samples were then incubated in Formamide Wash Buffer (Vizgen 20300002) for 30 min at 37 °C followed by probe hybridization with 50 µL of a custom-designed MERSCOPE Gene Panel Mix (Supplementary Data 1) at 37 °C for 36–48 h.
Gel embedding and tissue clearing
Following incubation, the tissues were washed twice with 5 mL Formamide Wash Buffer (Vizgen 20300002) at 47 °C for 30 min, and embedded into a hydrogel using the Gel Embedding Premix (Vizgen 20300004), ammonium persulfate (Sigma, 09913-100G) and TEMED (N,N,N’,N’-tetramethylethylenediamine) (Sigma, T7024-25ML) from the MERSCOPE Sample Prep Kit (Vizgen 10400012). After a 1.5-h incubation at room temperature, the samples were cleared with a solution consisting of 50 µl of Proteinase K (NEB, P8107S) and 5 mL of Clearing Premix (Vizgen 20300003) at 37 °C for at least 24 h, or until the tissue became transparent.
Imaging
After removing the clearing solution and washing three times with Sample Prep Wash Buffer (Vizgen 20300001), the samples were stained with DAPI and Poly T Reagent (Vizgen 20300021) for 15 min at room temperature, washed for 10 min with 5 mL of Formamide Wash Buffer (Vizgen 20300002), then washed for 5 min with 5 mL of Sample Prep Wash Buffer (Vizgen 20300001). The imaging reagents and processed sample were loaded into the MERSCOPE and a low-resolution DAPI mosaic (10x magnification) was used to select the region of interest before high-resolution imaging on the MERSCOPE system (Vizgen 10000001). With a 60x oil immersion objective, DAPI and Poly T stains were imaged at 7 focal planes on the z-axis for each tiled field of view (FOV), followed by 6 rounds of 3-color imaging across all focal planes using 750 nm, 650 nm, and 560 nm laser illumination. Each imaging round was followed by incubation in extinguishing, rinse, hybridization, wash, and imaging buffers. Additionally, a single image of fiducial beads was acquired at each FOV using 488 nm illumination. The full instrumentation protocol is available at https://vizgen.com/resources/merscope-instrument/.
Image analysis
The MERlin image analysis pipeline v0.1.699 was used to analyze the raw image files from the MERSCOPE experiment to align image stacks from the different MERFISH rounds, filter out background noise, and enhance RNA spot detection. Individual RNA molecule barcodes were then decoded using a pixel-based algorithm and an adaptive barcoding scheme that corrects misidentified barcodes not matching the provided codebook. Next, cells were segmented using information from nuclear DAPI and cell membrane antibody stains with the Cellpose software package100. The decoded RNA molecules were then grouped into individual cells, generating single-cell RNA count matrices. The entire pipeline was run for each imaging FOV and then tiled over the entire sample imaging area, or up to 1 cm2.
Four samples (n = 2 Ankrd11nchet and n = 2 Ankrd11ncko) were processed in the Vizgen facility, and 2 samples (additional n = 1 Ankrd11nchet and n = 1 Ankrd11ncko) were processed at the University of Alberta, Advanced Cell Exploration Core facility. Altogether, n = 3 samples per genotype from 2-3 litters were processed and analyzed.
Microscopy
Phalloidin-stained as well as RNAscope and BaseScope sections were imaged with a Zeiss LSM700 confocal microscope with a photomultiplier tube (PMT) using the 40x objective. Z-stacks spanning ~10–14 µm were captured with an optical slice thickness of 0.85-1.5 μm. All other images were captured using a Zeiss Axio Imager M2 fluorescence microscope with ORCA-Flash LT sCMOS Camera using the 20x objective for OFT and heart sections and the 2.5x objective for whole embryos in a single plane. All image acquisition was performed using Zen software (Zeiss). Confocal images were processed into stacked Z-planes. The thymus was imaged with a dissection microscope using a Samsung A53 phone camera.
Image analysis
To analyze CNCC-derived smooth muscle cells in the arteries at E18.5, 1 proximal and 1 distal image was used from each heart. αSMA + YFP+ cells and total αSMA+ cells were counted within the αSMA+ ring of cells surrounding the artery lumen. For Ankrd11nchet, cell numbers represent the combined numbers from the aorta and pulmonary trunk.
To analyze CNCC number, proliferation, and apoptosis, at least 6 transverse OFT sections from E10.5 embryos were split into proximal, medial, and distal regions using morphological characteristics. At E10.5, the proximal region has a small lumen and dispersed CNCCs. The medial region has a large lumen, dispersed CNCCs, and a thickened bottom cushion. The distal region has a large lumen, more compacted CNCCs, and a thickened bottom cushion. YFP+ CNCCs were counted and averaged to show the mean average number of cells per OFT section in each region. The proliferative index is presented as % Ki67+YFP+ cells over total YFP+ cells. The apoptotic index is presented as % TUNEL+YFP+ cells over total YFP+ cells.
To analyze anatomically matched sections of Ankrd11nchet and Ankrd11ncko OFTs at E11.5 and E12.5, the Ankrd11ncko distal region was determined to be the equivalent region to the Ankrd11nchet septated region based on the number of OFT sections it occupied, and the medial sections used for analysis were adjacent to this boundary. This method was used to avoid relying on the differing morphology between Ankrd11nchet and Ankrd11ncko OFTs and between embryonic ages.
To determine CNCC nuclear orientation in the OFT, all YFP+ cells from three sections of the medial region per OFT were analyzed as per ref. 36 with slight modifications. Namely, the angle between the long-axis of the CNCC nucleus and the OFT cushion center was manually measured using the Angle tool in Fiji ImageJ and binned into the 90°-70°, 69°-50°, 49°-30°, 29°-0° categories. The OFT cushion center was visually identified as a cluster of CNCCs that acted as a focal point around which the rest of the CNCCs was organized in approximate spirals.
To investigate Sema3c mRNA expression via RNAscope, images from at least 3 medial sections were analyzed from each embryo, and due to the varying distal-proximal gradient of marker expression, sections with the highest level of expression were quantified. Cells with three or more RNAscope dots were considered positive for marker expression as described in refs. 101,102. In Ankrd11nchet OFT, % Sema3c+YFP+ over total YFP+ cells were graphed from one section closest to the AP septum. In Ankrd11ncko OFT, 3 medial sections were quantified, and one section with the highest % Sema3c+YFP+ over total YFP+ cells was graphed. An average of ~400 cells was counted per OFT section.
To investigate Ankrd11 mRNA expression via BaseScope, images from 1 medial section were analyzed from each embryo. Within the OFT mesenchyme, DAPI+ cells with at least 1 Ankrd11 probe count/cell were counted as Ankrd11+. Additionally for Ankrd11nchet, Ankrd11+ cells were stratified by the number of Ankrd11 probe counts.
To analyze pS6 activation in the OFT, the % pS6 + S6 + YFP+ over total S6 + YFP+ cells from two sections was averaged to show the mean percentage of cells per medial OFT section. To analyze pSmad1/5/8, pSmad2/3, and Crabp2 signal, a threshold was manually set to delineate cells with high marker signal and cells with low marker signal, and only YFP+ cells with high marker signal were counted and presented as % marker+YFP+/YFP+ cells. An average of ~400 cells was counted per OFT section. Two sections were counted per OFT region.
YFP+marker+ CNCCs were counted when YFP+ cell cytoplasm overlayed with a Hoechst+ nucleus and a marker+ signal. All cells were counted manually.
To analyze the E18.5 thymus, both lobes of the thymus gland were outlined by hand and the area was quantified using a reference scale.
Images were counted and analyzed for fluorescence intensity, and representative images were processed in Fiji ImageJ v1.53t software103. Figures were generated in Adobe Illustrator CC 2015.
Single-cell RNA sequencing data re-analysis
The mouse trunk NC datasets at E9.5 and E10.5, published in27, were loaded from GEO database (GSE129114) and re-processed by scanpy package pipeline (version 1.9.1)104. Each dataset was filtered out to remove the low-quality cells: only cells with 5500 ≤ genes ≥ 10,000, 75,000 ≤ transcripts ≥ 750,000, and ≤25% ERCC reads were kept for further analysis. The filtered datasets were integrated via the CCA (canonical correlation analysis) method from Seurat105. PCA (principal component analysis) was performed on the scaled count matrix, and PC (principal components) responsible for the cell cycle was removed. To compute a nearest-neighbor graph, first 10 PC components and 30 neighbors were used. Further clustering and embedding were performed using UMAP (Uniform Manifold Approximation and Projection)106 and Leiden algorithm107 respectively. The cell type assignment to the clusters was done based on a list of well-established marker genes27. Visualizations were done using Scanpy and Seaborn (version 0.12.1) packages108. Ankrd11 expression distributions in CNCCs cluster were analyzed using pandas v1.5.2 for statistical summaries109.
Statistical analysis
All immunofluorescence as well as RNAscope and BaseScope data were obtained using at least 3 embryos per genotype from at least two independent litters. All µCT and echocardiography data were obtained using at least 3 embryos per genotype from at least three independent litters. Sample sizes (n) indicated in the figure legends correspond to the number of embryos analyzed. All data are presented as mean ± s.e.m.
All data (except MERFISH, described below) were subjected to normality tests using the Anderson-Darling, D’Agostino & Pearson, and Kolmogorov–Smirnov tests and were found to be normal or have insufficient sample size for these tests and were thus considered normal, except Fig. S1c. For two group comparisons, two-tailed unpaired student’s t-test or two-tailed multiple t-tests with Holm-Sidak multiple comparisons were used to assess statistical significance between means, where a p-value < 0.05 was considered significant. For three or more group comparisons two-way ANOVA was followed by Sidak’s multiple comparisons test. For results that did not pass normality tests, the Mann-Whitney test was used. In all cases, Prism (version 8.0.2) was used. Sample size and statistical information are stated in the corresponding figure legends. In the figures, asterisks denote statistical significance marked by *p < 0.05; **p < 0.01; ***p < 0.001. The sample size was not predetermined through statistical tests. In Fig. 2f one Ankrd11ctrl data point and in Fig. 2g one Ankrd11ncko data point were excluded using the ROUT method with Q = 1% to identify outliers. As Ankrd11ncko samples were easily identifiable by gross heart morphological abnormalities, blinded analysis was not possible. Due to the full penetrance of the phenotype regardless of the sex of the embryo, sex-based analysis was not performed.
MERFISH data analysis
Data generated from MERSCOPE runs was first ingested in Python version 3.11.6 using the squidpy 1.3.1 library110 User input (via the matplotlib library111) was then utilized to achieve two tasks: first a rotation was applied to spatial coordinates of all presented samples such that the sample “head” was pointing upwards in plots; secondly, a polygonal bounding box was defined around the OFT samples, and all cells were classified as within or outside this bounding box. All previously described data was then saved to disk. The ingest.py script contains all relevant code for the above-described steps112.
Data saved to disk was then read using the Seurat 5.0.1 library in R version 4.3.3113. Only cells within the previously defined OFT bounding box were considered for further analysis. Raw RNA counts were then normalized and scaled, using the Seurat NormalizeData and ScaleData functions respectively. After the computation of principal components using the RunPCA function, a nearest-neighbor graph was constructed using the first 30 principal components via the FindNeighbors function. Cells were then blindly clustered using the Leiden algorithm107 via the FindClusters function. Finally, 2-dimensional UMAP-space embeddings were computed via the RunUMAP function, using the first 30 principal components.
For further analysis, it was of interest to identify cardiac neural crest cells. Using canonical CNCC markers Penk and Sox9 and mesenchymal markers Pdgfra and Postn4,47,53, clusters numbered 2, 3, and 7 were determined to be CNCCs. Differentially expressed genes between the Ankrd11nchet and Ankrd11ncko genotypes were then computed using the FindMarkers function separately for each of these three clusters using the Wilcoxon rank-sum test. The analysis.r script contains all relevant code for the above-described steps112.
Sema3c images were generated using the MERSCOPE Vizualizer Version: 2.3.3330.0.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The scRNAseq data from ref. 27 used in this study are available in the GEO database under accession code GSE129114. The MERFISH post segmentation cell/gene expression count matrices and associated per-cell metadata generated in this study have been deposited in the GEO database under accession code GSE258835. For further data exploration, please refer to our open-science website: [https://singlocell.openscience.mcgill.ca/NeuralCrestAnkrd11-KO] (AH59.5 = ncHET 1; AH59.6 = ncHET 2; AIO4.2 = ncHET 3; AH50.1 = ncKO 1; AH59.2 = ncKO 2; AI13.3 = ncKO 3). Source data are provided with this paper.
Code availability
The MERFISH analysis source code as well as processed and metadata is available in GitHub [https://doi.org/10.5281/zenodo.10998925].
References
Wu, W., He, J. & Shao, X. Incidence and mortality trend of congenital heart disease at the global, regional, and national level, 1990-2017. Medicine (Baltim.) 99, e20593 (2020).
van der Linde, D. et al. Birth prevalence of congenital heart disease worldwide: a systematic review and meta-analysis. J. Am. Coll. Cardiol. 58, 2241–2247 (2011).
Keyte, A. & Hutson, M. R. The neural crest in cardiac congenital anomalies. Differentiation 84, 25–40 (2012).
Neeb, Z., Lajiness, J. D., Bolanis, E. & Conway, S. J. Cardiac outflow tract anomalies. Wiley Interdiscip. Rev. Dev. Biol. 2, 499–530 (2013).
Vega-Lopez, G. A., Cerrizuela, S., Tribulo, C. & Aybar, M. J. Neurocristopathies: new insights 150 years after the neural crest discovery. Dev. Biol. 444, S110–S143 (2018).
Krishnan, A. et al. A detailed comparison of mouse and human cardiac development. Pediatr. Res. 76, 500–507 (2014).
Yan, S., Lu, J. & Jiao, K. Epigenetic regulation of cardiac neural crest cells. Front. Cell Dev. Biol. 9, 678954 (2021).
Stefanovic, S., Etchevers, H. C. & Zaffran, S. Outflow tract formation-embryonic origins of conotruncal congenital heart disease. J. Cardiovasc. Dev. Dis. 8, 42 (2021).
Martik, M. L. & Bronner, M. E. Regulatory logic underlying diversification of the neural crest. Trends Genet. 33, 715–727 (2017).
Li, W. et al. Brg1 governs distinct pathways to direct multiple aspects of mammalian neural crest cell development. Proc. Natl Acad. Sci. USA 110, 1738–1743 (2013).
Yan, S. et al. CHD7 regulates cardiovascular development through ATP-dependent and -independent activities. Proc. Natl Acad. Sci. USA 117, 28847–28858 (2020).
Barnett, C. et al. Williams syndrome transcription factor is critical for neural crest cell function in Xenopus laevis. Mech. Dev. 129, 324–338 (2012).
Gallagher, D. et al. Ankrd11 is a chromatin regulator involved in autism that is essential for neural development. Dev. Cell 32, 31–42 (2015).
Zhang, A. et al. Identification of a novel family of ankyrin repeats containing cofactors for p160 nuclear receptor coactivators. J. Biol. Chem. 279, 33799–33805 (2004).
Li, C. W., Dinh, G. K., Zhang, A. & Chen, J. D. Ankyrin repeats-containing cofactors interact with ADA3 and modulate its co-activator function. Biochem. J. 413, 349–357 (2008).
Sirmaci, A. et al. Mutations in ANKRD11 cause KBG syndrome, characterized by intellectual disability, skeletal malformations, and macrodontia. Am. J. Hum. Genet. 89, 289–294 (2011).
Low, K. et al. Clinical and genetic aspects of KBG syndrome. Am. J. Med. Genet. A 170, 2835–2846 (2016).
Ockeloen, C. W. et al. Further delineation of the KBG syndrome phenotype caused by ANKRD11 aberrations. Eur. J. Hum. Genet. 23, 1176–1185 (2015).
Willemsen, M. H. et al. Identification of ANKRD11 and ZNF778 as candidate genes for autism and variable cognitive impairment in the novel 16q24.3 microdeletion syndrome. Eur. J. Hum. Genet. 18, 429–435 (2010).
Gnazzo, M. et al. KBG syndrome: common and uncommon clinical features based on 31 new patients. Am. J. Med. Genet. A 182, 1073–1083 (2020).
Goldenberg, A. et al. Clinical and molecular findings in 39 patients with KBG syndrome caused by deletion or mutation of ANKRD11. Am. J. Med. Genet. A 170, 2847–2859 (2016).
Handrigan, G. R. et al. Deletions in 16q24.2 are associated with autism spectrum disorder, intellectual disability and congenital renal malformation. J. Med. Genet. 50, 163–173 (2013).
Digilio, M. C. et al. Congenital heart defects in molecularly confirmed KBG syndrome patients. Am. J. Med. Genet. A https://doi.org/10.1002/ajmg.a.62632 (2021).
Murray, N. et al. KBG syndrome: an Australian experience. Am. J. Med. Genet. A 173, 1866–1877 (2017).
Guo, L. et al. KBG syndrome: videoconferencing and use of artificial intelligence driven facial phenotyping in 25 new patients. Eur. J. Hum. Genet. 30, 1244–1254 (2022).
Kierzkowska, O. et al. Documentation and prevalence of prenatal and neonatal outcomes in a cohort of individuals with KBG syndrome. Am. J. Med. Genet. A 191, 2364–2375 (2023).
Soldatov, R. et al. Spatiotemporal structure of cell fate decisions in murine neural crest. Science 364, eaas9536 (2019).
Roth, D. M. et al. The chromatin regulator Ankrd11 controls palate and cranial bone development. Front. Cell Dev. Biol. 9, 645386 (2021).
Danielian, P. S., Muccino, D., Rowitch, D. H., Michael, S. K. & McMahon, A. P. Modification of gene activity in mouse embryos in utero by a tamoxifen-inducible form of Cre recombinase. Curr. Biol. 8, 1323–1326 (1998).
Lewis, A. E., Vasudevan, H. N., O’Neill, A. K., Soriano, P. & Bush, J. O. The widely used Wnt1-Cre transgene causes developmental phenotypes by ectopic activation of Wnt signaling. Dev. Biol. 379, 229–234 (2013).
Jiang, X. et al. Fate of the mammalian cardiac neural crest. Development 127, 1607–1616 (2000).
Handel, A. E. et al. Developmental dynamics of the neural crest–mesenchymal axis in creating the thymic microenvironment. Sci. Adv. 8, eabm9844 (2022).
Orr-Urtreger, A., Bedford, M. T., Do, M.-S., Eisenbach, L. & Lonai, P. Developmental expression of the α receptor for platelet-derived growth factor, which is deleted in the embryonic lethal Patch mutation. Development 115, 289–303 (1992).
Konstantinov, I. E. et al. Truncus arteriosus associated with interrupted aortic arch in 50 neonates: a Congenital Heart Surgeons Society study. Ann. Thorac. Surg. 81, 214–222 (2006).
Yu, Q., Leatherbury, L., Tian, X. & Lo, C. W. Cardiovascular assessment of fetal mice by in utero echocardiography. Ultrasound Med. Biol. 34, 741–752 (2008).
Darrigrand, J. F. et al. Dullard-mediated Smad1/5/8 inhibition controls mouse cardiac neural crest cells condensation and outflow tract septation. Elife 9, e50325 (2020).
Hegarty, S. V., O’Keeffe, G. W. & Sullivan, A. M. BMP-Smad 1/5/8 signalling in the development of the nervous system. Prog. Neurobiol. 109, 28–41 (2013).
Shi, Y. & Massague, J. Mechanisms of TGF-beta signaling from cell membrane to the nucleus. Cell 113, 685–700 (2003).
Nie, X., Ricupero, C. L., Jiao, K., Yang, P. & Mao, J. J. mTOR deletion in neural crest cells disrupts cardiac outflow tract remodeling and causes a spectrum of cardiac defects through the mTORC1 pathway. Dev. Biol. 477, 241–250 (2021).
Yang, L. et al. The mTORC1 effectors S6K1 and 4E-BP play different roles in CNS axon regeneration. Nat. Commun. 5, 5416 (2014).
Biever, A., Valjent, E. & Puighermanal, E. Ribosomal protein S6 phosphorylation in the nervous system: from regulation to function. Front. Mol. Neurosci. 8, 75 (2015).
Choudhary, B. et al. Cardiovascular malformations with normal smooth muscle differentiation in neural crest-specific type II TGFbeta receptor (Tgfbr2) mutant mice. Dev. Biol. 289, 420–429 (2006).
Wurdak, H. et al. Inactivation of TGFbeta signaling in neural crest stem cells leads to multiple defects reminiscent of DiGeorge syndrome. Genes Dev. 19, 530–535 (2005).
Jia, Q. et al. Smad signaling in the neural crest regulates cardiac outflow tract remodeling through cell autonomous and non-cell autonomous effects. Dev. Biol. 311, 172–184 (2007).
Chen, Q. et al. Smad7 is required for the development and function of the heart. J. Biol. Chem. 284, 292–300 (2009).
Napoli, J. L. Cellular retinoid binding-proteins, CRBP, CRABP, FABP5: Effects on retinoid metabolism, function and related diseases. Pharm. Ther. 173, 19–33 (2017).
Liu, X. et al. Single-cell RNA-Seq of the developing cardiac outflow tract reveals convergent development of the vascular smooth muscle cells. Cell Rep. 28, 1346–1361 e1344 (2019).
Zaffran, S., Robrini, N. & Bertrand, N. Retinoids and cardiac development. J. Dev. Biol. 2, 50–71 (2014).
Keyte, A. L., Alonzo-Johnsen, M. & Hutson, M. R. Evolutionary and developmental origins of the cardiac neural crest: building a divided outflow tract. Birth Defects Res C. Embryo Today 102, 309–323 (2014).
Ramsdell, A. F. Left-right asymmetry and congenital cardiac defects: getting to the heart of the matter in vertebrate left-right axis determination. Dev. Biol. 288, 1–20 (2005).
Nelms, B. L. & Labosky, P. A. Transcriptional Control of Neural Crest Development. (Morgan & Claypool Life Sciences, 2010).
Plein, A., Fantin, A. & Ruhrberg, C. Neural crest cells in cardiovascular development. Curr. Top. Dev. Biol. 111, 183–200 (2015).
Chen, W. et al. Single-cell transcriptomic landscape of cardiac neural crest cell derivatives during development. EMBO Rep. 22, e52389 (2021).
Smith, C. L. & Tallquist, M. D. PDGF function in diverse neural crest cell populations. Cell Adh Migr. 4, 561–566 (2010).
Yamagishi, H. Cardiac neural crest. Cold Spring Harb. Perspect. Biol. 13, a036715 (2021).
Plein, A. et al. Neural crest-derived SEMA3C activates endothelial NRP1 for cardiac outflow tract septation. J. Clin. Invest. 125, 2661–2676 (2015).
Toyofuku, T. et al. Repulsive and attractive semaphorins cooperate to direct the navigation of cardiac neural crest cells. Dev. Biol. 321, 251–262 (2008).
Kodo, K. et al. Regulation of Sema3c and the interaction between cardiac neural crest and second heart field during outflow tract development. Sci. Rep. 7, 6771 (2017).
Delloye-Bourgeois, C. et al. Microenvironment-driven shift of cohesion/detachment balance within tumors induces a switch toward metastasis in neuroblastoma. Cancer Cell 32, 427–443 e428 (2017).
McLennan, R. & Krull, C. E. Ephrin-as cooperate with EphA4 to promote trunk neural crest migration. Gene Expr. 10, 295–305 (2002).
Sanford, L. P. et al. TGFbeta2 knockout mice have multiple developmental defects that are non-overlapping with other TGFbeta knockout phenotypes. Development 24, 2659–2670 (1997).
Theveneau, E. & Mayor, R. Neural crest delamination and migration: from epithelium-to-mesenchyme transition to collective cell migration. Dev. Biol. 366, 34–54 (2012).
Cantor, E. J., Babick, A. P., Vasanji, Z., Dhalla, N. S. & Netticadan, T. A comparative serial echocardiographic analysis of cardiac structure and function in rats subjected to pressure or volume overload. J. Mol. Cell Cardiol. 38, 777–786 (2005).
Müller, A. L. & Dhalla, N. S. in Cardiac Adaptations: Molecular Mechanisms (eds. Ostadal, B. & Dhalla, N. S.) 147–166 (Springer New York, 2013).
Momma, K., Ando, M., Takao, A. & Miyagawa-Tomita, S. Fetal cardiovascular morphology of truncus arteriosus with or without truncal valve insufficiency in the rat. Circulation 83, 2094–2100 (1991).
Buckvold, S. M., Yosowitz, T. & Hoffman, J. F. in Heart Diseases in Children: A Pediatrician’s Guide (ed. Ra-id Abdulla) 235–247 (Springer US, 2011).
Chikkabyrappa, S., Mahadevaiah, G., Buddhe, S., Alsaied, T. & Tretter, J. Common arterial trunk: physiology, imaging, and management. Semin. Cardiothorac. Vasc. Anesthesia 23, 225–236 (2018).
Tandon, R., Hauck, A. J. & Nadas, A. S. Persistent truncus arteriosus. Circulation 28, 1050–1060 (1963).
Kaartinen, V. et al. Cardiac outflow tract defects in mice lacking ALK2 in neural crest cells. Development 131, 3481–3490 (2004).
Jain, R. et al. Cardiac neural crest orchestrates remodeling and functional maturation of mouse semilunar valves. J. Clin. Invest 121, 422–430 (2011).
Martínez-Quintana, E. & Portela-Torrón, F. Truncus arteriosus and truncal valve regurgitation. Transl. Pediatr. 8, 360–362 (2019).
Singh, N. et al. Histone deacetylase 3 regulates smooth muscle differentiation in neural crest cells and development of the cardiac outflow tract. Circ. Res. 109, 1240–1249 (2011).
Chapnik, E., Sasson, V., Blelloch, R. & Hornstein, E. Dgcr8 controls neural crest cells survival in cardiovascular development. Dev. Biol. 362, 50–56 (2012).
Stottmann, R. W., Choi, M., Mishina, Y., Meyers, E. N. & Klingensmith, J. BMP receptor IA is required in mammalian neural crest cells for development of the cardiac outflow tract and ventricular myocardium. Development 131, 2205–2218 (2004).
Goodkey, K. et al. Olfactory bulb anomalies in KBG syndrome mouse model and patients. BMC Med. 22, 158 (2024).
Ka, M. & Kim, W. Y. ANKRD11 associated with intellectual disability and autism regulates dendrite differentiation via the BDNF/TrkB signaling pathway. Neurobiol. Dis. 111, 138–152 (2018).
Wang, J. et al. Defective ALK5 signaling in the neural crest leads to increased postmigratory neural crest cell apoptosis and severe outflow tract defects. BMC Dev. Biol. 6, 51 (2006).
Nie, X., Deng, C. X., Wang, Q. & Jiao, K. Disruption of Smad4 in neural crest cells leads to mid-gestation death with pharyngeal arch, craniofacial and cardiac defects. Dev. Biol. 316, 417–430 (2008).
Delot, E. C., Bahamonde, M. E., Zhao, M. & Lyons, K. M. BMP signaling is required for septation of the outflow tract of the mammalian heart. Development 130, 209–220 (2003).
Rankin, C. T., Bunton, T., Lawler, A. M. & Lee, S.-J. Regulation of left-right patterning in mice by growth/differentiation factor-1. Nat. Genet. 24, 262–265 (2000).
Gaio, U. et al. A role of the <em>cryptic</em> gene in the correct establishment of the left–right axis. Curr. Biol. 9, 1339–1342 (1999).
Oh, S. P. & Li, E. The signaling pathway mediated by the type IIB activin receptor controls axial patterning and lateral asymmetry in the mouse. Genes Dev. 11, 1812–1826 (1997).
Gandhi, S., Ezin, M. & Bronner, M. E. Reprogramming axial level identity to rescue neural-crest-related congenital heart defects. Dev. Cell 53, 300–315 e304 (2020).
Beppu, H. et al. BMP type II receptor regulates positioning of outflow tract and remodeling of atrioventricular cushion during cardiogenesis. Dev. Biol. 331, 167–175 (2009).
Nie, X., Zheng, J., Cruciger, M., Yang, P. & Mao, J. J. mTOR plays a pivotal role in multiple processes of enamel organ development principally through the mTORC1 pathway and in part via regulating cytoskeleton dynamics. Dev. Biol. 467, 77–87 (2020).
Liu, L. & Parent, C. A. Review series: TOR kinase complexes and cell migration. J. Cell Biol. 194, 815–824 (2011).
Sakakibara, K., Liu, B., Hollenbeck, S. & Kent, K. C. Rapamycin inhibits fibronectin-induced migration of the human arterial smooth muscle line (E47) through the mammalian target of rapamycin. Am. J. Physiol. Heart Circ. Physiol. 288, H2861–2868, (2005).
Liu, L. et al. Rapamycin inhibits cytoskeleton reorganization and cell motility by suppressing RhoA expression and activity. J. Biol. Chem. 285, 38362–38373 (2010).
Reuter, M. S. et al. The Cardiac Genome Clinic: implementing genome sequencing in pediatric heart disease. Genet Med. 22, 1015–1024 (2020).
Jin, S. C. et al. Contribution of rare inherited and de novo variants in 2,871 congenital heart disease probands. Nat. Genet. 49, 1593–1601 (2017).
Chui, M. M. C. et al. Evaluating high-confidence genes in conotruncal cardiac defects by gene burden analyses. J. Am. Heart Assoc. 12, e028226 (2023).
Sifrim, A. et al. Distinct genetic architectures for syndromic and nonsyndromic congenital heart defects identified by exome sequencing. Nat. Genet. 48, 1060–1065 (2016).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4–4 (2001).
Degenhardt, K., Wright, A. C., Horng, D., Padmanabhan, A. & Epstein, J. A. Rapid 3D phenotyping of cardiovascular development in mouse embryos by micro-CT with iodine staining. Circ. Cardiovasc. Imaging 3, 314–322 (2010).
Fedorov, A. et al. 3D Slicer as an image computing platform for the Quantitative Imaging Network. Magn. Reson. Imaging 30, 1323–1341 (2012).
Watson, A. E. S. et al. Fractalkine signaling regulates oligodendroglial cell genesis from SVZ precursor cells. Stem Cell Rep. 16, 1968–1984 (2021).
Li, Y. et al. Hepatoma derived growth factor enhances oligodendrocyte genesis from subventricular zone precursor cells. ASN Neuro 14, 17590914221086340 (2022).
de Almeida, M. M. A. et al. Fractalkine enhances oligodendrocyte regeneration and remyelination in a demyelination mouse model. Stem Cell Rep. https://doi.org/10.1016/j.stemcr.2022.12.001 (2023).
Zhang, M. et al. Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598, 137–143 (2021).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Voronova, A. et al. Migrating interneurons secrete fractalkine to promote oligodendrocyte formation in the developing mammalian brain. Neuron 94, 500–516 e509 (2017).
Zahr, S. K. et al. A translational repression complex in developing mammalian neural stem cells that regulates neuronal specification. Neuron 97, 520–537 e526 (2018).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902.e1821 (2019).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: Uniform Manifold Approximation and Projection. J. Open Source Softw. 3, https://doi.org/10.21105/joss.00861 (2018).
Traag, V. A., Waltman, L. & van Eck, N. J. From Louvain to Leiden: guaranteeing well-connected communities. Sci. Rep. 9, 5233 (2019).
Waskom, M. Seaborn: statistical data visualization. J. Open Source Softw. 6, 3021 (2021).
McKinney, W. in Proceedings of the 9th Python in Science Conference (SciPy, 2010).
Palla, G. et al. Squidpy: a scalable framework for spatial omics analysis. Nat. Methods 19, 171–178 (2022).
Hunter, J. D. Matplotlib: a 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Afanasiev, E. The chromatin regulator Ankrd11 controls cardiac neural crest cell-mediated outflow tract remodeling and heart function. Github, https://doi.org/10.5281/zenodo.10998925 (2024).
Hao, Y. et al. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat. Biotechnol. 42, 293–304 (2023).
Acknowledgements
Y.K. was supported by University of Alberta Motyl Scholarship in cardiac sciences, Women and Children’s Health Research Institute (WCHRI) Graduate studentship and CIHR CGS-D; N.L.D. was supported by Brad Mates E Drive Studentship in Parkinson’s Disease and Movement Disorders and NSERC PGS-D; K.G. by NSERC CGS-M and WCHRI graduate studentship; AESW by CIHR CGS-D and IP by Marie Skłodowska-Curie Innovative Training Networks (ITN) Program Neu-Crest (grant agreement No. 860635). A.V. was also supported by the Canada Research Chair in Neural Stem Cell Biology and Sloan Fellowship in Neuroscience awards. This work was supported by private donations from families affected by KBG syndrome (A.V.), Gilbert Winter K. Fund from the University of Alberta Hospital Foundation operating grant (A.V.), KBG Foundation seed funding (A.V.), Stollery Children’s Hospital Foundation and the Alberta Women’s Health Foundation through the Women and Children’s Health Research Institute grant 3925 (A.V.), European Research Council Synergy grant KILL-OR-DIFFERENTIATE 856529 (I.A.), Knut och Alice Wallenbergs Stiftelse consortium grant (IA), Vetenskapsrådet (grant 2023-02161, I.A.), Cancerfonden Project Grant (20240101, I.A.), Paradifference foundation grant (I.A.), Bertil Hållstens Forskningsstiftelse (I.A.), Göran Gustafssons Stiftelse för Naturvetenskaplig och Medicinsk Forskning (I.A.) and Austrian Science Fund Project Grant (P34136, I.A.) and Consortium Grant (SFB-F78, I.A.). The authors thank Sarah Hughes, Heather McDermid, Jonathan Epp, Dylan Terstege, Karim Fouad, Pamela Raposo, Daniela Roth, Tim Footz, Sana Bibi, and Meghan Riddell for technical assistance and helpful discussions, Alexis Allot for hosting datasets on an open science platform and Sarah Hughes for access to the confocal microscope. Finally, we thank the Advanced Cell Exploration Core led by Mike Wong at the University of Alberta for their help with MERFISH assays.
Author information
Authors and Affiliations
Contributions
Conceptualization, Y.K. and A.V.; Methodology, Y.K., E.A., J.M., A.E.S.W., R.N., M.A., I.P., N.D., A.G., K.G., J.U., I.A., D.G., S.B., J.S., A.V.; Formal analysis, Y.K., E.A., A.E.S.W., R.N., I.P., K.G.; Resources, A.V.; Writing—original draft, YK; Writing—review & editing, Y.K. and A.V.; Supervision, A.V., J.U., I.A., D.G., S.B., J.S.; Funding acquisition, A.V.
Corresponding author
Ethics declarations
Competing interests
The authors declare the following competing interests: J.M. is an employee of Vizgen. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kibalnyk, Y., Afanasiev, E., Noble, R.M.N. et al. The chromatin regulator Ankrd11 controls cardiac neural crest cell-mediated outflow tract remodeling and heart function. Nat Commun 15, 4632 (2024). https://doi.org/10.1038/s41467-024-48955-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-48955-1
- Springer Nature Limited
This article is cited by
-
Insights into the ANKRD11 variants and short-stature phenotype through literature review and ClinVar database search
Orphanet Journal of Rare Diseases (2024)