Abstract
Genetic parasites, including viruses and transposons, exploit components from the host for their own replication. However, little is known about virus-transposon interactions within host cells. Here, we discover a strategy where human cytomegalovirus (HCMV) hijacks L1 retrotransposon encoded protein during its replication cycle. HCMV infection upregulates L1 expression by enhancing both the expression of L1-activating transcription factors, YY1 and RUNX3, and the chromatin accessibility of L1 promoter regions. Increased L1 expression, in turn, promotes HCMV replicative fitness. Affinity proteomics reveals UL44, HCMV DNA polymerase subunit, as the most abundant viral binding protein of the L1 ribonucleoprotein (RNP) complex. UL44 directly interacts with L1 ORF2p, inducing DNA damage responses in replicating HCMV compartments. While increased L1-induced mutagenesis is not observed in HCMV for genetic adaptation, the interplay between UL44 and ORF2p accelerates viral DNA replication by alleviating replication stress. Our findings shed light on how HCMV exploits host retrotransposons for enhanced viral fitness.
Similar content being viewed by others
Introduction
Retrotransposons are mobile genetic elements that replicate through retrotransposition, a process involving reverse transcription and insertion of retrotransposon RNA into their host genome. Among these elements, long-interspersed element-1 (LINE-1 or L1) is an active autonomous retrotransposon, accounting for approximately 17% of the human genome1. Retrotransposition-competent L1 encodes two proteins: ORF1p, a nucleic acid chaperone protein2, and ORF2p with endonuclease and reverse transcriptase domains3,4. ORF1p and ORF2p preferentially interact with their parent mRNA to form an L1 ribonucleoprotein (RNP) particle5. The L1 RNP serves as a functional unit for L1 retrotransposition, inducing DNA breaks and generating new L1 insertions in the genome via the mechanism of target-primed reverse transcription (TPRT)6. This genetic parasite poses a threat to genome integrity as well as contributes to host genome evolution7.
Human cytomegalovirus (HCMV) is a member of the Betaherpesvirinae subfamily with a global seroprevalence of up to 90%8,9, and it establishes latency and persists throughout the host lifespan. HCMV possesses a 230 kb double-stranded DNA genome that encodes hundreds of proteins and non-coding RNAs10,11. The large sets of HCMV genes engage in numerous interactions with the host, taking control of cellular machinery and disrupting normal functions12. In particular, HCMV alters the nuclear environment, repurposing chromatin modifiers and recruiting DNA repair machinery13,14,15. Moreover, HCMV activates genomic repetitive elements, including endogenous retroviruses and satellite repeat RNA16,17. However, little is known about the interplay between HCMV and genomic repetitive elements.
The role of transposons in viral infection has been increasingly recognized, with some transposons being involved in horizontal gene transfer18,19 or antiviral defense of the CRISPR-Cas system20. However, how viruses leverage host transposons during their life cycle has not been examined. Since HCMV has coevolved with the host over millions of years, shaping dynamic virus-host interactions21, we investigated HCMV as a model virus to address this knowledge gap.
In this study, we found that HCMV infection activates L1 expression by increasing the levels of transcription factors YY1 and RUNX3 expression, thereby inducing local chromatin remodeling at the L1 promoter. Furthermore, the HCMV DNA processivity factor, UL44, interacts with L1 RNP complexes and recruits them to replicate viral DNA. L1 ORF2p induces DNA damage responses enriched at viral replication compartments, promoting efficient replication. Our findings reveal a novel strategy for viruses to exploit host retrotransposons for enhanced replication.
Results
HCMV infection activates L1 retrotransposon expression
First, we analyzed the expression of human transposable elements (TEs) using a transcriptome of HCMV-infected fibroblasts16. We found that most TEs, including L1, were activated upon HCMV infection, whereas non-infected fibroblasts exhibited a tight suppression of L1 expression (Fig. 1a; Supplementary Fig. 1a, b). Notably, L1Hs, the youngest retrotransposition-competent subfamily of L1 elements, showed a higher expression level than other TEs (Fig. 1b). Using cytoplasmic RNA extracts from HCMV (Toledo clinical strain)-infected human glioblastoma cells (U373MG), we detected a clear increase in L1 RNA from the early to late phases of the HCMV life cycle [1–3 days post-infection (dpi)] using multiple qPCR primer sets to enhance coverage (Fig. 1c). Additionally, the L1 ORF1p protein accumulated in HCMV-infected U373MG cells and primary human foreskin fibroblasts (HFF) throughout the viral life cycle (Fig. 1d and Supplementary Fig. 1c). UV-inactivated HCMV, incapable of expressing viral genes yet able to stimulate cellular immune responses22, did not induce L1 protein expression (Fig. 1d), indicating that L1 expression is triggered by interplay with HCMV. Next, we tested whether there was a difference between clinical and laboratory strains of HCMV in inducing L1. Infection of laboratory strain Towne and clinical strain TB40/E also activated L1 expression, as seen in the HCMV Toledo strain (Supplementary Fig. 1d,e). These results suggest that HCMV infection triggers L1 expression through a mechanism conserved across different strains of HCMV.
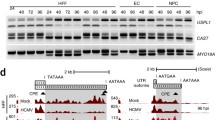
a Heat map showing the expression of transposable elements (TE) families upon HCMV infection. Color map indicates the Z-score of TE expression. b Volcano plot for differentially expressed gene (DEG) analyses of L1 subfamilies. The dataset of GSE12089016 was analyzed for (a) and (b). c Cytoplasmic L1 RNA expression levels in HCMV-infected U373MG cells. Four different L1 primer sets were used for L1 RNA detection. Bars represent mean ± SD of n = 4 independent samples. One-way ANOVA with Dunnett’s test. d Immunoblot assay of L1 ORF1p expression in HCMV-infected U373MG cells. IE1/2 are immediate-early proteins, and pp28 is a late tegument protein of HCMV. Representative blot of n = 3 independent replicates. Dunnett’s multiple comparisons test. e Correlation between a gene expression from RNA-seq data and a ratio of intact TEs from ATAC-seq data. Each dot represents the evolutionary age from L1PA12 to L1Hs with a color gradient. f Map of ATAC-seq read in L1Hs TSS region. The plot presents data from HCMV-infected U373MG in red and non-infection U373MG in gray. YY1 and RUNX3 consensus binding regions are indicated. g Immunoblot analysis for assessing the expression of YY1 and RUNX3 during HCMV infection in U373MG. Vinculin is the loading control. Representative blot of n = 2 independent replicates. h Immunoblot assay showing the effect of YY1 and RUNX3 knockdown in HCMV-infected HFFs at 3 dpi. Representative blot of n = 3 independent replicates. Source data are provided as a Source Data file.
Since L1 expression is epigenetically silenced in somatic cells23, L1 upregulation in HCMV-infected cells led us to test whether HCMV affects the epigenetic landscape of the genomic L1 regions. Using transposase-accessible chromatin sequencing (ATAC-seq)24, we profiled chromatin accessibility of the L1 subfamilies upon HCMV infection. Alignment of ATAC-seq reads to genomic L1 regions showed that evolutionarily young L1s were more accessible in HCMV-infected cells than in uninfected cells (Supplementary Fig. 2a and Supplementary Data 1). This chromatin accessibility pattern correlated with L1 subfamily gene expression levels from the RNA-seq data (Fig. 1e). This data suggests that HCMV unwraps the chromatin structure of the evolutionarily young L1 regions, leading to enhanced L1 transcription.
This preference prompted us to investigate transcription factors responsible for HCMV-mediated L1 activation. We performed motif enrichment analyses using ±500 bp region at the transcription start site (TSS) of L1Hs, which indicates the specific region for intact and full-length L1Hs, and selected 14 sequence motifs associated with several transcription factors (Supplementary Fig. 2b). Among these, YY1 and RUNX3 are known L1 transcription factors25,26, and showed binding motifs at approximately +13 bp and +92 bp relative to the L1 TSS in the L1 promoter, respectively (Supplementary Fig. 2c), and the loci showed enhanced ATAC-seq signals upon HCMV infection (Fig. 1f and Supplementary Fig. 2d). We also found that HCMV infection stimulated the expression of YY1 and RUNX3 in HCMV-infected HFF and U373MG cells (Fig. 1g and Supplementary Fig. 2e), confirming a positive correlation with the L1 expression. Depletion of YY1 or RUNX3 using siRNA reduced L1 ORF1p expression compared to that in control siRNA (Fig. 1h and Supplementary Fig. 2f). Thus, our data demonstrate that during HCMV infection, L1 expression is activated through the upregulation of L1 transcription factors YY1 and RUNX3 and the increased chromatin accessibility at the L1 promoter region.
Activated L1 expression promotes HCMV replication
To investigate the role of L1 in the HCMV life cycle, we introduced a small hairpin RNA (shRNA) targeting L1 mRNA into the HCMV genome, allowing the shRNA to function only in HCMV-infected cells (Fig. 2a). HCMV strains with two different L1 knockdown shRNAs (shL1-1 and shL1-2) effectively suppressed L1 ORF1p expression whereas wild-type (WT) and control shLacZ HCMV strains successfully induced L1 expression (Fig. 2b).
a Map of engineered HCMV genome and schematic representation of shRNA insertion. Functional constructs, including U6 promoter and shRNA, are inserted in the intergenic region of US1-2. b Western blot analysis showing L1 knockdown effect of shL1 RNA-inserted HCMV in U373MG at 5 dpi. Representative blot of n = 3 independent replicates. c ELISA assessing the production of secreted IFNβ from WT, shLacZ, shL1-1, or shL1-2 HCMV-infected HFFs. At 1 day post-infection (dpi), the culture medium of HCMV-infected HFFs was harvested and subjected to ELISA. Mean ± SEM of n = 6 independent samples. One-way ANOVA with Dunnett’s test. d Quantification of interferon-stimulated gene (ISG) expression. qRT-PCR assay using RNA from WT, shLacZ, shL1-1, or shL1-2 HCMV-infected HFFs. At 1 dpi, total RNA was extracted from HCMV-infected HFFs. Primers targeting ISG are indicated. Mean ± SEM of n = 4 independent samples. One-way ANOVA with Dunnett’s test. e, f Viral growth curves of WT and shRNA-encoded HCMV (shLacZ, shL1-1, and shL1-2). Primary HFFs were infected with the indicated viruses at a multiplicity of infection (MOI) of 0.1 (e) and 2 (f). Cell-free virus in the supernatant was titrated by limiting dilution analysis at the indicated time points. Mean ± SEM of n = 3 independent samples, Two-way ANOVA and Dunnett’s multiple comparisons test. HFF cells were infected with the cell-free viruses collected at 9 dpi from 0.1 MOI infection (e, right panel) and at 7 dpi from 2 MOI infection (f, right panel). HCMV IE1/2 (green) and DAPI (blue) images are shown. The scale bar represents 200 μm. Source data are provided as a Source Data file.
Since L1-derived nucleic acids are known to trigger cellular immune responses27, we tested if these sets of HCMV exhibit different immunogenicity to the host cell. By assessing the levels of secreted Interferon β (IFNβ) and RNA expression of interferon-stimulated genes (ISG), we found that there were no notable changes in the IFNβ and ISG expression levels between WT, shLacZ, and shL1 HCMV-infected cells (Fig. 2c, d).
We then evaluated the growth rate of each HCMV using primary HFF. Notably, while the replication kinetics of the WT and shLacZ HCMV was similarly efficient, the progeny production of shL1-1 and shL1-2 HCMVs was much slower and less prolific than that of the WT and shLacZ control viruses (Fig. 2e, f; ~100-fold decrease at 9 dpi of 0.1 MOI infection, and ~10-fold decrease at 7 dpi of 2 MOI infection). These results indicate the critical role of L1 in promoting HCMV replication without affecting host antiviral responses.
HCMV DNA processivity factor, UL44, interacts with L1 ORF2p of the L1 RNP complex
To probe the mechanism through which L1 promotes viral replication, we investigated the interactome of the L1 RNP complex during HCMV infection. We used a full-length L1 construct with FLAG-tagged ORF1p and generated L1ORF1-FLAG U373MG, a cell line expressing L1 in a doxycycline (dox)-dependent manner. (Fig. 3a). We immunoprecipitated the L1 RNP complex from HCMV-infected cells and performed liquid chromatography-tandem mass spectrometry (LC/MS) analyses (Fig. 3a and Supplementary Fig. 3a). We identified 44 ORF1p-interacting candidates, including 41 human and 3 HCMV proteins (Fig. 3b and Supplementary Data 2). The majority of the human proteins (36 out of 41) were known interactors of the L1 RNP complex28,29,30, validating our L1 RNP immunoprecipitation.
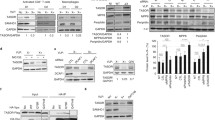
a A schematic representation of identifying L1 RNP interacting with HCMV proteins. pBL1ORF1p-FLAG U373MG were generated to express L1 with FLAG-tagged ORF1 upon dox treatment. b Average score from LC/MS analyses. Bars indicate human (black) and HCMV (red) proteins. The table shows the L1 RNP-interacting HCMV proteins and their function. c Co-IP assays using HCMV-infected pBL1ORF1p-FLAG U373MG. IP mixture was eluted with either FLAG peptides or RNase, as indicated. Representative blot of n = 2 independent replicates. d Endogenous L1 ORF1p Co-IP assay of U373MG cells at 4 dpi showing interactions between L1 RNP and UL44. Given differences in detection efficiencies, long and short exposures of L1 ORF1p and L1 ORF2p blots are presented. Flow-through was saved before IP wash. Representative blot of n = 2 independent replicates. e Co-IP assays showing the interaction of L1 ORF2p with UL44 via its PIP-box plug residues. HeLa cells were transfected with L1 ORF2p-3×FLAG and/or HA-UL44 plasmids. Co-IP assays were performed using HA antibody. Representative blot of n = 2 independent replicates. f Structural modeling of UL44 with PCNA binding region of L1 ORF2p (403–419). UL44 hydrophobic crevice composed of Val58, Val136, Leu251, and Phe266 (light blue) and L1 ORF2p hydrophobic plug Tyr414 and Tyr415 (red) are shown. g Co-IP assays using FLAG-UL44-transfected pBL1 HeLa with UL44 hydrophobic crevice mutations. Representative blot of n = 2 independent replicates. The band intensity of L1 ORF2p and L1 ORF1p are indicated below. Source data are provided as a Source Data file.
Among the three identified HCMV proteins, UL44 ranked the highest, and its interaction with L1 RNP was validated using a co-immunoprecipitation assay (Fig. 3b and Supplementary Fig. 3b). We next investigated whether UL44 binds directly to L1 ORF1p or interacts with the L1 RNP complex. For this, we tested the ribonuclease (RNase) sensitivity upon L1 ORF1p-FLAG immunoprecipitation (Fig. 3c left panel). With complete elution of the L1 RNP complex using excessive FLAG peptides, UL44 interacts with L1 ORF1p as well as its partner L1 ORF2p (Fig. 3c). However, RNase treatment disrupted UL44 and L1 ORF1p interaction, while L1 ORF1p was associated with L1 ORF2p. (Fig. 3c). This indicates that RNA is required for UL44 interaction with L1 ORF1p and L1 ORF2p. Furthermore, we examined whether mutations in ORF1p RNA-binding residues (R261-262A) affected their interaction with UL44. As expected, UL44 immunoprecipitation showed no interaction with the RNA-binding-defective ORF1p (Supplementary Fig. 3c). A co-immunoprecipitation assay of endogenous L1 ORF1p in HCMV-infected cells showed that endogenous L1 RNP also interacted with UL44 upon infection (Fig. 3d). Collectively, these data suggest that UL44 binds to L1 RNP, but not via direct interaction with ORF1p.
UL44 is the HCMV DNA polymerase processivity factor, and we were particularly interested in it because L1 ORF2p interacts with the human DNA polymerase processivity factor, PCNA30. The structural similarity between PCNA and UL4431 led us to hypothesize the potential interaction between L1 ORF2p and UL44. L1 ORF2p interacts with PCNA via its PCNA-interacting protein box (PIP-box; Qxx[V/L/M/I]xx[F/Y][F/Y])30. The hydrophobic “plug” formed by two aromatic residues in the PIP-box fits into the hydrophobic crevice of PCNA, located around the interdomain connecting loop32 (Supplementary Fig. 4a). This docking mechanism was also found in the X-ray crystal structure of UL4431 binding to the PIP-box of UL54, an HCMV polymerase catalytic subunit33 (Supplementary Fig. 4b). Based on these structures, we tested whether L1 ORF2p binds to UL44 via a conserved docking mechanism. We found that L1 ORF2p interacted with UL44, and this interaction was abolished by the mutation of two hydrophobic plug residues (Y414-415A) in the PIP-box (Fig. 3e). Additionally, we generated an in silico protein–peptide docking model of the L1 ORF2p PIP-box motifs and UL44 (Fig. 3f). Remarkably, this model revealed the molecular conformations of L1 ORF2p Y414 and Y415, fitting into the UL44 hydrophobic crevice, which is composed of V58, V136, L251, and F266 (Fig. 3f). To validate this model, we investigated whether mutations in the hydrophobic crevice affected the interaction between L1 ORF2p and UL44. All three V58A, V136A, and L251A mutations disrupted the interaction of UL44 with L1 ORF2p as well as the interaction with L1 ORF1p (Fig. 3g). These data show that HCMV UL44 interacts directly with L1 ORF2p via a docking mechanism similar to that of its host homolog, PCNA.
L1 induces DNA damage in HCMV DNA replication compartments
Because UL44 is highly enriched in replicating HCMV DNA, forming distinct nuclear compartments termed as viral replication compartments (VRCs)34, we speculated that UL44 recruited L1 retrotransposons to HCMV VRCs. UL44 in HCMV-infected L1ORF2-FLAG U373MG cells showed the formation of subnuclear structures separated from the host cell chromatin, which are canonical features of VRCs (Fig. 4a and Supplementary Fig. 5a). While L1 ORF2p-FLAG formed cytoplasmic foci in naïve L1ORF2-FLAG U373MG cells, during HCMV infection, L1 ORF2p-FLAG exhibited sparse localization within the nucleus, preferentially accumulating within the HCMV VRCs (Fig. 4a and Supplementary Fig. 5a). These findings suggest that UL44 recruits L1 ORF2p, a component of the L1 RNP, to the HCMV VRCs.
a Immunofluorescence assays using HCMV-infected pBL1ORF2p-3×FLAG U373MG at 48 hpi (left). At 24 hpi, cells were treated with 50 ng/mL dox until fixation. UL44 (green); L1 ORF2p-3×FLAG (magenta); DAPI (blue); and merged images are shown with the scale bar (5 μm). Violin plot for % colocalization values from 10 representative images is shown (right), with the horizontal dashed line indicating the median as well as 25th and 75th percentiles. b Immunoblot assay using 1 MOI of WT, shLacZ, shL1-1, and shL1-2 HCMV-infected HFFs at 4 dpi (left). Quantification of pH2AX intensities normalized to H2AX level (right). Mean ± SD of n = 3 independent samples. Two-sided Student’s t-test. c Immunofluorescence assay using WT, shLacZ, shL1-1, and shL1-2 HCMV-infected HFFs at 48 hpi. UL44 (green), pH2AX (red), and merged with DAPI (blue) images are indicated. The scale bar represents 20 μm. d Colocalization coefficients showing the numbers of UL44 and pH2AX-colocalized pixels from (c). Dots with mean ± SD indicate the values of 22 cells per sample from 3 independent experiments. Two-sided Kolmogorov–Smirnov test. e Immunofluorescence assay using dox-inducible L1 expressing U373MG (pBL1 U373MG) at 48 hpi. Mutations in L1 ORF2p and dox treatment are indicated. UL44 (green), pH2AX (red), and DAPI (blue) images are shown. The scale bar represents 10 μm. f Colocalization coefficients showing the numbers colocalized pixels of UL44 and pH2AX with intensity ≥ 60 from (e). Dots with mean ± SD indicate the values from 20 cells per sample from 3 independent experiments. Two-sided Kolmogorov–Smirnov test. Source data are provided as a Source Data file.
We investigated the effects of L1 RNP recruitment on the HCMV VRCs. Since L1 ORF2p is an endonuclease that induces DNA strand breaks35, we tested whether L1 induces a DNA damage response upon HCMV infection. We found that the phosphorylation of histone H2AX at serine 139 (pH2AX), an early marker of DNA strand breaks36, increased upon infection with WT and shLacZ HCMVs compared to that in the non-infected control (Fig. 4b). However, in shL1 HCMV-infected cells, there was no obvious activation of pH2AX (Fig. 4b). YY1 and RUNX3 knockdown significantly reduced the activation of pH2AX compared to that in the control cells, indicating that the activation of DNA damage response was dependent on L1 expression (Supplementary Fig. 5b). We then investigated whether our observed DNA damage markers also co-occupied the VRCs in an L1 retrotransposon-dependent manner. In WT and shLacZ HCMV-infected cells, pH2AX accumulated and formed foci in the periphery of UL44, but pH2AX signals were reduced in shL1 HCMV-infected cells (Fig. 4c, d), demonstrating that L1 knockdown attenuated the accumulation of DNA damage markers, specifically in the VRCs. These data suggest that L1 induces a DNA damage response in replicating HCMV DNA.
Next, we investigated whether L1 ORF2p enzymatic activity is involved in HCMV-induced DNA damage responses. L1 ORF2p has two enzymatic activities, endonuclease (EN) and reverse transcriptase (RT), which are closely linked and contribute to creating breaks in the target DNA6,35,37. RT inhibitors stavudine (d4T) and lamivudine (3TC) did not affect pH2AX induction within HCMV VRCs (Supplementary Fig. 5c,d), indicating the RT independence of L1-induced DNA damage response to HCMV DNA. To further clarify L1 enzymatic functions in HCMV DNA damage responses, we utilized dox-inducible L1-expressing cells with two mutations in L1 ORF2p: EN-defective H230A and RT-defective D702A. Dox treatment increased pH2AX in VRCs of WT and RT-defective L1 mutants (Fig. 4e, f; Supplementary Fig. 5e). However, the EN-defective L1 did not induce pH2AX signals in the VRCs, regardless of dox treatment (Fig. 4e, f; Supplementary Fig. 5e). These findings indicate that the EN activity of L1 ORF2p activates DNA damage responses in replicating HCMV DNA.
We also examined whether the interaction between L1 ORF2p and UL44 is necessary for the induction of DNA damage responses. Using a proximity ligation assay (PLA), we assessed the spatial DNA damage responses in the periphery of UL44 in dox-inducible L1 and UL44-expressing HeLa cells. Remarkably, PLA signals of pH2AX and HA-tagged UL44 were observed only in L1-expressing cells (Supplementary Fig. 6a). However, the L1 ORF2p binding-deficient mutation of UL44 (UL44mut; V58A, V136A, and L251A) showed significantly fewer PLA signals in the nuclei than in WT UL44 (Supplementary Fig. 6a,b), suggesting that L1 ORF2p induces DNA damage in the vicinity of VRCs, and this effect is dependent on the interaction between UL44 and L1 ORF2p.
L1 retrotransposition and L1-mediated DNA damage do not contribute to HCMV genetic instability
Given the mobility of L1 and the involvement of L1 ORF2p in HCMV replication, we investigated whether endogenous L1 retrotransposition facilitates the genetic adaptation of HCMV during lytic infection. If certain L1 insertions were to promote the fitness of carrier HCMV, these insertions would be expected to be inherited by progeny viruses and amplified via clonal expansion in the virus population. To test this hypothesis, we first applied human active transposon-sequencing (HAT-seq)38, a PCR-based L1 capture method, to profile L1 insertions in the HCMV genomes (Fig. 5a; Supplementary Data 3). HCMV Toledo virus (WT, shLacZ, and shL1; >60 days culture in fibroblasts) and in vitro cultured laboratory strain Towne were analyzed. Although HAT-seq demonstrated high sensitivity in detecting L1 insertions from low-level human genomic DNA carryover (Fig. 5b; Supplementary Fig. 7a, b; Supplementary Data 4), we were unable to identify high-confidence clonal L1 insertions integrated into the HCMV genomes (Fig. 5c and Supplementary Fig. 7c). Further efforts to experimentally validate the predicted L1 insertions in the HCMV samples were unsuccessful (Supplementary Note 1, Supplementary Fig. 8, and Supplementary Data 5). We additionally performed ultra-deep Illumina whole-genome sequencing (WGS) to comprehensively identify all forms of L1-mediated retrotransposition, including Alu and SVA, in the HCMV genome. We found no evidence of putative clonal transposon insertions with an allele frequency greater than 0.1% in the viral population (Supplementary Note 2 and Supplementary Table S1). These extensive analyses suggest that, in our HCMV culture system, L1 and L1-mediated retrotransposition may not be the primary mechanism responsible for HCMV adaptation.
a Schematic representation of analyses of L1 insertion and mutation in HCMV genome. b, c Alignment of human active transposon-sequencing (HAT-seq) peaks for L1 insertions in the human and HCMV genomes, and integrative genomics viewer plots of representative L1 insertions in the human and HCMV genomes. b Non-reference L1Hs insertions supported by soft-clipped reads (with colored bases) that span the L1-genome junction. For each library, the upper track displays the alignment of the L1-trimmed 3′ flanking sequence, while the lower track shows intact but soft-clipped contigs spanning the integration site. c Putative L1 insertions in the HCMV genome supported by a single template with PCR duplicates. The red arrows indicate the start position of supporting contigs, which marks the number of different templates bearing the same insertion. d Mutational single-nucleotide polymorphism (SNP) burden across shL1 and control WT and shLacZ samples as detected by each variant-calling scheme: VarScan2 (top), Mutect2 (middle), and the intersection of VarScan2 and Mutect2 calls (bottom). Mutation calls are separated into Batch 1 (left) and Batch 2 (right). Each dot represents a single replicate. VAF variant allele frequency. e Circos plots of combined Batch 1 and Batch 2 SNPs. Inner rings show genomic coordinates in kb (brown) and major sequence features of HCMV (green). Middle tracks show SNP calls in WT (purple), shLacZ (blue), shL1-1 (red inner), and shL1-2 (red outer) samples. The outer ring shows GenBank HCMV Toledo sequence annotations. Source data are provided as a Source Data file.
Since HCMV exhibits high genetic diversity during natural infection39,40, we determined whether L1-mediated DNA damage leads to accumulated mutations in the HCMV genome, thereby conferring enhanced viral replication fitness. To this end, we investigated the underlying patterns of L1-mediated mutagenesis within the viral genomes of WT, shLacZ, and shL1 HCMVs by revisiting deep Illumina short-read WGS of HCMV. We developed a stringent single-nucleotide polymorphism (SNP) and indel variant-calling pipeline for high-confidence variant detection (Supplementary Fig. 9a,b). Considering that the variants of interest may be present in a small number of viral genomes relative to the sampled population, we designed our approach to achieve high sensitivity while maintaining robustness by combining Mutect241 and VarScan242. We used the HCMV Toledo BAC DNA as a control to detect mutations with low variant allele frequencies (VAFs) specifically present in the other HCVM samples, i.e., WT, shLacZ, or shL1, but absent in the control. Despite >60 days of passaging, very few high-confidence SNPs and no high-confidence indels were detected across all samples (Fig. 5d; Supplementary Fig. 9c, d). The mutational burden tended to differ between batches, but there was no overall significant enrichment of mutations across WT and shLacZ samples compared with shL1 samples (Fig. 5d). There were no apparent hotspots of mutation accumulation in the WT and shLacZ samples that could indicate key mutations mediated by L1 and coordinating proviral effects (Fig. 5e; Supplementary Fig. 9e, f, and Supplementary Table S2). Thus, our data suggest that accumulated mutations or newly emerging HCMV variants are not associated with enhanced HCMV replication by the L1 retrotransposon.
L1 retrotransposon-UL44 interaction accelerates viral DNA replication
Next, we investigated the role of L1 ORF2p in the life cycle of HCMV. Since the DNA damage response and HCMV growth rate showed a positive correlation with L1 expression levels (Figs. 2e, f and 4b), we hypothesized that the L1 retrotransposon participated in HCMV DNA replication. First, we found that WT and shLacZ HCMV accumulated a much greater number of HCMV genome DNA copies than shL1 HCMVs (Fig. 6a; Supplementary Fig. 10a, b). To assess the rate of viral DNA replication, we incubated cells with a pulse of 5-ethynyl-2′-deoxyuridine (EdU) and measured the intensity of incorporated EdU in HCMV-infected cells. As HCMV infection arrests the host cell cycle43, DNA replication in HCMV-infected cells is majorly ascribed to HCMV DNA. Using flow cytometry, we found that WT and shLacZ-infected cells showed higher EdU intensities than shL1 HCMVs-infected cells, indicating increased DNA replication in the presence of L1 (Supplementary Fig. 10c,d). We next examined the direct incorporation of EdU into HCMV VRCs. We found that EdU intensity is much higher in the UL44 area of WT and shLacZ HCMV than those of shL1 HCMVs (Fig. 6b, c). These findings indicate that L1 retrotransposons promote efficient HCMV DNA synthesis.
a Measurement of HCMV DNA copy number. HFF were infected with indicated HCMV (0.5 MOI). Viral was quantified via qPCR (UL86/7SK). Data are presented as mean ± SEM of n = 4 independent samples. Two-way ANOVA and Dunnett’s multiple comparisons test. b, c EdU incorporation assay. HFFs were infected with indicated HCMV (0.5 MOI). At 72 hpi, DNA was labeled using 10 μM EdU for 30 min. b Representative images of EdU incorporation into HCMV DNA. UL44 (orange) and EdU (green), and merged images with Hoechst (blue) are shown with the scale bar (10 μm). c Each dot in the box plot shows the EdU intensity within the UL44 area of an individual cell (n = 25), derived from 3 independent experiments. d EdU-BrdU dual labeling of HCMV DNA for fork stalling assay (left) and representative images (right). EdU (green), BrdU (magenta), and merged image with Hoechst (blue) are shown with the scale bar (5 μm). Gamma value 0.45 is applied to BrdU channel. e Each dot in the box plot indicates the BrdU intensity within the EdU area from an individual cell (n = 25) from 3 independent experiments. f Schematic of aniPOND (left). pBL1ORF2p-3×FLAG U373MG cells at HCMV 72 hpi were incubated with DMSO (Lane 1), 10 μM EdU (Lane 2), or 10 μM EdU followed by 10 μM Thymidine (Lane 3). The enrichment of UL44, L1 ORF2p-3×FLAG, and PCNA were analyzed by immunoblot (right, representative blot of n = 2 independent experiments). g, h pBL1ORF2p-3×FLAG U373MG cells expressing L1 ORF2p WT or Y414-415A were infected with HCMV, and incubated with 50 ng/mL of dox and fixed at 48 hpi. g Merged images of UL44 (green), pH2AX (red), and DAPI (blue) are shown with the scale bar (10 μm). h Colocalization coefficients were determined using the numbers of UL44 and pH2AX-colocalized pixels with both intensities ≥60. Dots with mean (the horizontal line) ± SD indicate the colocalization coefficient values (n = 25) from 3 independent experiments. c, e and h Two-sided Kolmogorov–Smirnov test. c, e The boxes extend from the 25th to 75th percentiles, with whiskers reaching the minimum and maximum values. The horizontal line within each box represents the median. Source data are provided as a Source Data file.
HCMV infection accompanies a feature of replication stress regarding stalled replication forks44, usually resolved by the recruitment of several endonucleases45,46. Combining our findings of (1) L1 accelerates viral DNA replication with induction of DNA damage response, and (2) L1 endonuclease activity is required to induce DNA damage response (Fig. 4), we hypothesized that L1 ORF2p could contribute to resolve replication stress at the replication forks in HCMV DNA. To test this hypothesis, we utilized Camptothecin (CPT) which directly targets topoisomerase I at the replication fork47, and adopted an EdU-BrdU dual labeling strategy to distinguish DNA replication before and after fork stalling at a single-cell level (Fig. 6d). We examined the HCMV DNA synthesis after recovery of stalled forks by measuring BrdU intensities within EdU area (Fig. 6d). Notably, BrdU intensities were much higher within VRCs of WT and shLacZ HCMV than shL1 HCMVs, indicating that shL1 HCMVs had defects in reinitiation of DNA replication after CPT stimulus (Fig. 6d, e). This data indicated that the delayed DNA replication of shL1 HCMVs could be due to stalled forks in HCMV DNA. We additionally tested whether L1 could resolve replication stress caused by other drugs disrupting fork progression by different mechanisms48. We observed that HCMV showed comparable levels of BrdU incorporation in response to Etoposide (ETP), hydrogen peroxide (H2O2), and hydroxyurea (HU) regardless of L1 expression (Supplementary Fig. 10e,f). These results indicate that L1 depletion sensitizes HCMV to replication stress, especially to fork stalling caused by topoisomerase I inhibition.
The unique effect of CPT is interesting, as this agent induces DNA breaks right at the site of replication fork49. Based on our finding of the contribution of L1 to resolve CPT-mediated replication stress, we suspected that L1 ORF2p recruited by UL44 directly functions at the HCMV replication fork. To investigate the spatial context of such interaction between L1 ORF2p and UL44, we performed accelerated nascent isolation of proteins on nascent DNA (aniPOND)50, a method to analyze the replication fork proteome, on HCMV-infected cells (Fig. 6f, left panel). By capturing nascent DNA that were mostly of viral origin (Supplementary Fig. 10g), we found that UL44 and L1 ORF2p were enriched at viral replication fork (Fig. 6f). EdU pulse followed by thymidine chase for purification of maturing chromatin reduced the enrichment of both UL44 and L1 ORF2p (Fig. 6f). This result demonstrates that L1 ORF2p and UL44 interact at the viral replication fork specifically.
We finally assessed the functional consequence of the L1 ORF2p-UL44 interaction at the viral replication fork. To this end, we generated dox-inducible L1 ORF2p Y414-415A mutant expressing U373MG cells in which L1 ORF2p did not bind to UL44 during HCMV infection (Supplementary Fig. 10h). We evaluated the pH2AX induction in VRCs, which positively correlated with HCMV DNA replication. Notably, expression of Y414-415A mutant L1 ORF2p did not induce pH2AX in VRCs (Fig. 6g, h; Supplementary Fig. 10i) compared to WT L1 ORF2p. This data suggests that L1 ORF2p recruitment to the viral replication fork by UL44 is critical for inducing the DNA damage response. Together, our data suggest that L1 retrotransposons contribute to alleviating replication stress at the viral replication fork, thereby increasing viral replication efficiency.
Discussion
In this study, we report a mechanism by which HCMV harnesses L1 retrotransposons for its replication. HCMV infection leads to the upregulation of L1 retrotransposons by opening chromatin and upregulating transcription factors YY1 and RUNX3. HCMV UL44 recruits L1 RNP to replicate viral DNA for prolific progeny production. HCMV exploits L1 retrotransposons to relieve DNA replication stress while maintaining the integrity of the viral genome (Supplementary Fig. 11).
Most TEs are epigenetically silenced in differentiated cells51. Thus, the global induction of various TEs (Fig. 1a) indicates that HCMV subverts cellular epigenetic regulation. We demonstrated YY1- and RUNX3-dependent L1Hs induction, but have not determined how HCMV enhances the chromatin accessibility of the L1Hs genomic regions. Notably, YY1 and RUNX3 act not only as traditional DNA binding transcription factors but also as regulators of chromatin accessibility52,53, which suggests their potential role in mediating multiple aspects of epigenetic regulation in HCMV infection. The other possible mechanism involves altered histone modifications. HCMV genes UL29, UL38, and UL123 can act to suppress histone deacetylases (HDACs)12,54. Since histone acetylation causes nucleosomes to unfold and increases access of transcription factors to genomic TE regions55, it could explain how HCMV induces not only the expression of L1 but also of other TEs.
L1-mediated retrotransposition occurs in the poxvirus genome, facilitating horizontal gene transfer from the host to the virus18,19. Since HCMV rapidly adapts to the host environment by acquiring diverse mutations, even in in vitro fibroblast culture56, we investigated L1-mediated HCMV evolution during >60 days in in vitro culture and passaging. However, we did not detect endogenous clonal retrotransposition events that could drive genetic adaptation. Both Fixsen et al.19 and Rahman et al.18 used engineered L1 constructs and conferred selective advantages to accelerated laboratory evolution. Similar to their scheme, the utilization of potent selection pressures, such as antiviral resistance, could induce a novel case of L1 retrotransposition in the HCMV genome. In addition, evolutionary pseudogene capture in herpesviruses implies the possibility of L1 retrotransposition into herpesviruses genomic DNA57.
Since the HCMV genome has a high proportion of guanine–cytosine nucleotide bonds (57% GC) and repeated sequences58, alleviating replication stress is critical for its efficient propagation. HCMV has been reported to exploit host DNA damage responses44,59. DNA repair proteins ATM and p53 are required for HCMV replication59,60,61, and DNA breaks marker, pH2AX, accumulate during infection without any exogenous DNA damage59,62,63. However, the mechanism of how accumulated DNA breaks contribute to reducing replication stress is a long-standing question. Our findings demonstrate that L1 endonuclease is required for HCMV-induced DNA damage and promotes efficient HCMV DNA replication. Given the broad substrate specificity of L1 ORF2p for AT-rich DNA3, it is unlikely that L1 ORF2p recognizes stalled forks. Instead, UL44 might play a role as the molecular bridge for L1 ORF2p to the viral replication forks. Since the endonuclease activity of L1 retrotransposons accumulates a wide range of DNA repair proteins at the host replication fork64,65, HCMV may exploit L1 retrotransposons for recruiting DNA repair proteins to the viral replication fork. Although it is still unclear if L1 endonuclease nicks stalled forks region directly and which DNA repair pathway proteins act at L1-induced DNA break sites during HCMV infection, our findings imply that HCMV has evolved a new strategy to exploit L1 machinery for enhanced fitness while mitigating potential L1-mediated genetic instability. Future studies should clarify the mechanistic role of L1 endonuclease in HCMV replication.
In summary, our findings provide a new perspective on L1 activity in DNA replication and how HCMV acquires the control of L1 retrotransposons. Considering the potent pathogenicity of L1, our study introduces a new pathogenic model for retrotransposon activation during viral infection. This mechanism may offer insights into novel therapeutic targets and improve our understanding of HCMV pathogenesis.
Methods
Cells
Primary HFFs, U373MG, and HeLa cells were grown in Dulbecco’s modified Eagle’s medium (DMEM; HyClone) with 10% fetal bovine serum (FBS; HyClone) and 1% (v/v) GlutaMAX-I (Gibco). Cells were incubated at 37 °C under 5% CO2 atmosphere.
Viruses
HCMV Toledo and AD169 BAC DNA were gifts from T. Shenk (Princeton University). HCMV TB40/E-mCherry BAC DNA was a gift from E. Murphy (Upstate Medical University). Infectious HCMVs were generated by transfecting HCMV BAC DNAs into primary HFFs using the Neon Transfection System (MPK5000; Invitrogen) via electroporation (1700 V, 20 ms, and 1 pulse). After 100% of cells showed cytopathic effects, culture media were harvested, and virus particles were collected via centrifugation at 40,000×g for 1 h at 4 °C. The pellet of virus particles was resuspended in DMEM with 10% FBS.
For titration of viral stocks, HFFs were infected with serially diluted viruses for 1 h and fixed with 3.7% formaldehyde at 24 hpi. Cells were then permeabilized with 0.1% Triton X-100, incubated with 2% bovine serum albumin (BSA) in phosphate-buffered saline (PBS), and stained with HCMV IE1/2 antibody (MAB810R; Millipore) and FITC-conjugated anti-mouse antibody (115-095-146; Jackson Laboratories). The number of HCMV IE1-positive cells was counted to determine the number of infectious units per milliliter of viral stock.
RNA-Seq
For the analyses of total RNA-seq data of HCMV-infected fibroblasts (GSE12089016), raw sequencing reads were preprocessed using Trim Galore! v0.6.766 and aligned to the hg38 reference genome using STAR v2.7.1a67. GENCODE v3768. Gene annotations and transposable element annotations69 (‘hg38_rmsk_TE.gtf’) were merged to generate STAR genome index tailored for TE expression quantification. We adopted the following TE-specific options for STAR alignment as follows: --outFilterMultimapNmax 100 --winAnchorMultimapMax 100 --outMultimapperOrder Random --runRNGseed 777 --outSAMmultiNmax 1 --outFilterType BySJout --alignSJoverhangMin 8 --alignSJDBoverhangMin 1 --outFilterMismatchNmax 999 --alignIntronMin 20 --alignIntronMax 1000000 --alignMatesGapMax 1000000. The alignments were indexed using Sambamba v0.8.170. Gene and TE expression levels were quantified using TEtranscripts v2.2.1, and differential expression analyses were performed using DESeq2 v1.34.071. Differentially expressed gene was defined as if the fold change was greater than two and its Benjamini–Hochberg adjusted P-value was less than 0.05.
RNA extraction and RT-qPCR
Total RNA was extracted using the TRIzol reagent (15596-018; Invitrogen). To remove the remaining DNA, total RNA was treated with recombinant DNase I (2270; Takara) for 1 h at 37 °C, followed by purification using a Nucleospin RNA Clean-up kit (Macherey-Nagel). RNA was reverse-transcribed using the ReverTra Ace qPCR RT kit (FSQ-101; Toyobo). Real-time PCR was performed using the TOPreal qPCR 2X SYBR premix (RT500M; Enzynomics). The primers used for qPCR are listed in Supplementary Table S3.
For cytoplasmic RNA extraction, HCMV-infected U373MG cells were lysed with subcellular fractionation buffer (250 mM sucrose; 20 mM HEPES, pH 7.4; 10 mM KCl; 1.5 mM MgCl2; 1 mM EDTA; 1 mM EGTA; and 1 mM DTT) supplemented with a complete protease inhibitor cocktail (Roche) and RNase inhibitor (Enzynomics). The lysates were agitated at 4 °C and then separated from the nuclei pellet by using centrifugation at 720 × g for 5 min at 4 °C. The supernatants were cleared via centrifugation at 10,000 × g at 4 °C for 10 min and subjected to RNA extraction using TRIzol LS reagent (Invitrogen).
ATAC-seq
ATAC-seq library preparation and sequencing were performed by Macrogen, Inc. A transposition reaction was performed immediately after lysis. Nuclei of 50,000 cells were resuspended in the transposition reaction mix and were incubated at 37 °C for 30 min. DNA was extracted using the MinElute PCR Purification kit (Qiagen). The transposed DNA fragments were amplified using the Nextera DNA Flex kit. Purified libraries were sequenced using a HiSeq platform (Illumina).
ATAC-seq data were preprocessed using Trim Galore! v0.6.7 and aligned to the hg38 reference genome using BWA v0.7.17-r118872. Alignment sorting and indexing were performed using Sambamba v0.8.1. Genome-wide read-depth signals (in Reads per Kilobase Million) were computed using deepTools v3.5.173. To avoid anomalous signals, we excluded ENCODE blacklist regions74 from the analysis. Aggregated signals around the intact L1 transposons were computed using deepTools. The locations of the L1 transposons were determined using transposable element annotations, and intact L1 transposons were defined as those longer than 5000 bp. MACS2 v2.2.7.175 was used to filter duplicate reads and call ATAC peaks.
HOMER motif analysis
Transcription factor-binding motif enrichment analysis was performed using HOMER v4.1176. First, using the transposable element annotations, we defined the promoter regions for L1Hs transposons as ±500 bp regions centered at the TSS. Thereafter, enriched motifs were identified using “findMotifsGenome.pl” script with region size 200 bp and motif lengths 8, 10, and 12 from 351 L1Hs sequences. We selected 14 sequence motifs that are found in more than 75% of L1Hs with log P-value < −1000.
IFNβ ELISA
HFF were infected with HCMV at 1 MOI. At 24 hpi, culture medium was analyzed using VeriKine Human IFN beta ELISA kit (PBL ASSAY Science). The concentration of IFNβ secreted from HCMV-infected cells was calculated using human IFNβ standard (0–4000 pg/mL).
RNA interference
siRNAs targeting YY1 and RUNX3 or non-targeting siRNAs were purchased from Dharmacon (ON-TARGETplus, SMARTpool). All siRNA transfections were performed using DharmaFECT 1 transfection reagent (Dharmacon).
Generation of recombinant HCMV with shRNA construct
To generate HCMV encoding shRNA construct, we modified the HCMV Toledo BAC using a counter-selection BAC modification kit (GeneBridge), according to the manufacturer’s protocol. The intergenic region between US1 and US2 in the Toledo BAC was replaced with a prokaryotic selection cassette, rpsL-neoR, through homologous recombination. Next, the prokaryotic selection cassette was replaced with a shRNA-expressing cassette from the pLKO.1-TRC vector via homologous recombination. The shRNA-expressing cassette oligos were amplified using PCR. The primers used in BAC recombineering are listed in Supplementary Table S4. BAC clones with successful recombination were validated by whole plasmid sequencing. Sequences for all BAC clones are provided as a Source Data file. BAC DNAs were extracted using the NucleoBond Xtra BAC kit (740436; Macherey-Nagel). Purified BACs were transfected into primary HFFs to produce infectious viruses, as described in “Viruses.”
Generation of dox-inducible L1-expressing cells
U373MG or HeLa cells were seeded at 1.5 × 105 cells/well in a 6-well plate. The following day, the cells were transfected with 0.25 μg of Super PiggyBac Transposase Expression Vector (PB210PA-1; System Biosciences) and 1 μg of dox-inducible codon-optimized L1-expressing vectors (pBL1; gift from N. Liu) using Lipofectamine 3000 (L3000001; Invitrogen). After 5 days, the cells were selected with Blasticidin S (A1113903; Gibco).
LC-MS/MS analysis
A Q Exactive Hybrid Quadrupole-Orbitrap instrument (Thermo Scientific) equipped with a Dionex U 3000 RSLCnano HPLC system was used. Mass spectrometric analyses were performed using a Q Exactive Hybrid Quadrupole-Orbitrap mass spectrometer (Thermo Scientific). Fractions were reconstituted in solvent A [water/acetonitrile (ACN; 98:2 v/v) and 0.1% formic acid as the highly aqueous mobile phase] and then injected into the LC-nano ESI-MS/MS system. Samples were first trapped on an Acclaim PepMap 100 trap column (100 μm × 2 cm; nanoViper C18; 5 μm; 100Å; Thermo Scientific, part number 164564) and washed for 6 min using 98% solvent A [water/ACN (98:2 v/v), 0.1% Formic acid] at a flow rate of 4 μL /min, and then separated on an Acclaim PepMap 100 capillary column (75 μm × 15 cm; nanoViper C18; 3 μm; 100Å; Thermo Scientific, part number 164568) at a flow rate of 300 nL/min. The LC gradient was run at 2% to 40% solvent B for over 45 min, 40% to 95% solvent B for over 5 min, 95% solvent B (100% ACN and 0.1% formic acid) for 10 min, and finally 2% solvent B for 20 min. Xcaliber software version 4.3 was used to collect the MS data. The Orbitrap analyzer (Thermo Scientific) scanned the precursor ions with a mass range of 350–1800 m/z with 70,000 resolution at m/z 200. Mass data were acquired automatically using proteome discoverer 2.5 (Thermo Scientific). To identify potential interacting proteins of the L1 RNP, we filtered out the technical background, fragment proteins, and uncharacterized proteins. All data from replicated experiments were then filtered by using coverage and score (coverage ≥ 10, score ≥ 50). Proteins commonly identified in 2 independent experiments were selected, and the average scores of each protein were calculated.
Immunofluorescence assay and calculation of colocalization coefficient
Cells were washed twice with PBS, fixed with 3.7% formaldehyde in PBS for 30 min then permeabilized using 0.1% Triton X-100 in PBS for 10 min. Cells were incubated in blocking solution (2% BSA in PBS) for blocking and stained with HCMV UL44 mouse antibodies (CA006-1; Virusys) at a dilution of 1:400 and phosphor-histone H2A.X (Ser139; pH2AX) rabbit antibodies (9718; Cell Signaling Technology) at a dilution of 1:500 at 4 °C for 4 h. After three times washes, cells were incubated with Alexa Fluor-488-conjugated donkey anti-mouse (A-21202; Invitrogen) and Alexa Fluor-568-conjugated goat anti-rabbit (A-11011; Invitrogen) at a dilution of 1:200 at 4 °C for 4 h. Lastly, cells were washed three times with the blocking solution and mounted on glass slides using the Vectashield antifade mounting medium containing DAPI (H-1200; Vector Laboratories). Images were obtained by LSM700 confocal laser-scanning microscope (ZEISS) with C-Apochromat 40×/1.20NA water objective at a resolution of 512×512 pixels with a pixel size of 0.31 μm. The colocalization coefficient values were determined on a per-cell basis by counting the number of pH2AX pixels with an intensity ≥60 and overlapping with UL44 pixels having an intensity ≥60 using Zen Blue (ZEISS) [Eq. 1].
Immunoprecipitation
To identify the L1 RNP-interacting HCMV proteins, pBL1ORF1p-FLAG U373MG was used. We infected 2 × 107 cells with HCMV at 5 MOI. After 2 days, the cells were treated with 50 ng/mL dox. At 4 dpi, cells were harvested and resuspended in RIP lysis buffer [150 mM KCl; 10 mM HEPES, pH 7.6; 2 mM EDTA; 0.5% NP40; 1 mM DTT; complete protease inhibitor cocktail (Roche); and 400 unit/mL RNase inhibitor (Enzynomics)] for 10 min at 4 °C. Cells were then lysed by freezing and thawing using LN2, cleared using centrifugation (4 °C, 15,000×g for 15 min), and filtrated by passing through a 0.45 μm membrane syringe filter. For antibody-bead preparation, 15 μg FLAG-M2 antibody (Sigma-Aldrich) was diluted in 160 μL NT2 buffer [200 mM NaCl, 50 mM HEPES (pH7.6), 2 mM EDTA, and 0.05% NP40] and incubated with 4.8 mg Dynabead protein G for 30 min and rotation at room temperature. After incubation, the antibody-bead complexes were washed and resuspended in NT2 buffer, with a volume equivalent to the lysate volume. Lysates were then added to the bead-containing tube and incubated overnight on a rotating wheel at 4 °C. The beads were then washed five times with 1 mL ice-cold NT2 buffer. FLAG-ORF1p binding proteins were eluted using 3×FLAG peptide competition for 2 h at 4 °C (Sigma-Aldrich). The eluates were separated using SDS-PAGE, and the gels were excised for LC-MS/MS analyses.
For immunoblotting assays, we infected 5 × 106 cells of pBL1ORF1p-FLAG U373MG infected with HCMV at 5 MOI, treated 50 ng/mL dox at 2 dpi, and lysed cells using RIP lysis buffer at 4 dpi. Lysates were incubated with 3.8 μg of FLAG M2 overnight. Elution was performed using 20 μg of 3×FLAG peptide or 40 μg of Monarch® RNase A (New England BioLabs) for 2 h at 4 °C.
Endogenous ORF1p immunoprecipitations were performed using 2 × 107 cells of HCMV-infected U373MG (5 MOI). Cells were harvested at 4 dpi and lysed with RIP lysis buffer. After pre-clearing of lysates with beads on a rotating wheel for 1 h at 4 °C, lysates were incubated with 10 μg of antibodies [IgG (sc-2025; Santa Cruz) and L1 ORF1p (clone 4H1, MABC1152; Merck)] for 4 h at 4 °C. Supernatants were saved as flow-through control. Beads were washed five times with ice-cold NT2 buffer, resuspended in 2× Laemmli Sample buffer, and boiled for 10 min.
HCMV UL44–L1 ORF2p peptide complex structure modeling
The candidate interaction site of L1 ORF2p (residues 403–419) was determined to be near the PIP-box, which was previously known as the PCNA binding motif30. The ORF2p peptide (403-DPTEIQTTIREYYKHLY-419) docked to HCMV UL44 (input residues 10–270) was prepared using GalaxyPepDock77. The GalaxyPepDock program used the UL44–UL54 C-terminal peptide structure (PDB: 1YYP) as the template structure. Docking simulations showed 10 possible UL44–ORF2p peptide complex structures with the same scoring by GalaxyPepDock. All model structures were equivalent, and the most optimized structures were obtained from each initial sampling. We focused on the interaction between the UL44 hydrophobic crevice and hydrophobic plug (Tyr414 and Tyr415) of the L1 ORF2p peptide. Finally, the most plausible structure was selected as a representative model, as illustrated in Fig. 4f.
PLA
PLA was conducted using Duolink In Situ Detection Reagent Orange (DUO92007; Sigma-Aldrich) and Duolink In Situ PLA Probes [Anti-Mouse MINUS (DUO92004); Anti-Rabbit PLUS (DUO92002)]. First, pBL1ORF2p-3xFLAG HeLa cells were transfected with the HA-UL44 plasmid. Twenty-four hours after transfection, the cells were re-seeded on coverslips and treated with 200 ng/mL dox for an additional 2 days. Three days after transfection, cells were fixed with 3.7% formaldehyde and permeabilized with 0.1% Triton X-100. Cells were then blocked in 1X blocking solution (DUO92102; Sigma-Aldrich) for 1 h at 37 °C and incubated overnight in antibody solution (pH2AX rabbit antibodies (9718; Cell Signaling Technology)) at a dilution of 1:500 and anti-HA mouse antibody (ab18181; Abcam) at a dilution of 1:600 in Duolink Antibody Diluent at 4 °C. The next day, the coverslips were washed twice with 1X Wash buffer A and incubated with PLA rabbit PLUS and mouse MINUS probes for 1 h at 37 °C. Subsequent steps, including ligation and amplification, were performed according to the manufacturer’s instructions. Images were obtained by LSM700 confocal laser-scanning microscope (ZEISS) C-Apochromat 20×/0.8NA. The number of PLA puncta within the DAPI area was counted manually.
Purification of HCMV DNA for sequencing
Viruses were propagated using approximately 1.2 × 107 HFFs. Primary HFFs were infected with 0.05 MOI of HCMV. After 100% of cells showed cytopathic effects, culture media were harvested, and virus particles were collected via centrifugation at 40,000 × g for 1 h at 4 °C. Viral DNA from HCMV particles was extracted using a Monarch HMW DNA extraction kit for cells and blood (T3050; New England BioLabs).
HAT-seq library construction and sequencing
Human active transposon-sequencing (HAT-seq) libraries were constructed as previously described38, with minor modifications. 500 ng of genomic DNA was sonicated using a Covaris S220 focused-ultrasonicator (Covaris, LLC), followed by end repair, A-tailing, and adapter ligation using KAPA HyperPrep Kit (KAPA Biosystems). L1 enrichment PCR was performed using 25 ng adapter-ligated genomic DNA fragments as templates. The PCR program consisted of initial denaturation at 95 °C for 5 min, followed by 5 cycles of linear amplification at 95 °C for 40 s, 61 °C for 15 s, and 72 °C for 15 s and a pause at 12 °C for the addition of the P5_extension primer. The program continued with 15 cycles of exponential amplification at 95 °C for 40 s, 61 °C for 15 s, and 72 °C for 15 s, ending with a final extension at 72 °C for 30 s and then held at 4 °C. PCR products (L1-genome junctions) were cleaned using AMPure XP beads (Beckman Coulter, Inc.), followed by four cycles of indexing PCR and one round of PCR to eliminate possible bubble products derived from over-amplification. Library fragments between 340 and 450 bp were selected using Pippin HT (Sage Science). Four HCMV HAT-seq barcoded libraries were equally pooled based on molar concentrations quantified using the KAPA Library Quantification Kit (KAPA Biosystems). The pooled library underwent quality control assays (Agilent TapeStation for size distribution and Roche LightCycle qPCR for concentration) at Macrogen, Inc. The final HAT-seq library was paired-end sequenced (2 × 150 bp) in one lane using a HiSeq X Ten sequencer (Illumina). The adapter and primer sequences are listed in Supplementary Data 3.
HAT-seq data processing and analysis
A total of 351,721,377 read pairs (2 × 150 bp) were generated, with an average of 87,930,344 read pairs per sample. Raw sequencing data were processed as follows: (1) de-multiplexing of raw data, (2) preprocessing of FastQ files to trim adapters and low-quality bases (Phred score <15), and (3) discarding reads shorter than 50 bp. The processed clean read pairs were filtered based on library structure and PCR targets. First, read 2 was checked to start with the L1 primer sequence (3′ end of P7_Ns_L1Hs “GGGAGATATACCTAATGCTAGATGACAC”). Second, the L1 primer was assessed to have annealed to the 3′ end of the L1Hs consensus sequence (with 95% sequence identity). These L1Hs-derived read 2 and their paired read 1 were merged into contigs using PEAR (version 0.9.6)78 and aligned to the human cytomegalovirus Toledo BAC genome (GenBank: AC146905.1) and the human reference genomes (version hg19), respectively. Alignments of the L1-genome junctions revealed that the majority of the junctions were amplified from low-level human gDNA carryover in the HCMV gDNA extracts from cell cultures, whereas a small proportion of contigs could be mapped to the HCMV genome. To accurately map putative L1 integrations, soft-clipped and non-uniquely mapped contigs were collected, and the L1 section and poly-Ts (TTTTTTTT) were trimmed, followed by re-alignment of the 3′ genomic flanking sequence to the HCMV genome for novel L1 insertion calling. Peak calling was performed on genomic regions with coverage depth greater than one. Putative L1 insertion peaks were annotated based on features such as peak height (number of supporting reads), peak width (length of peak coverage with depth ≥1), and template count (number of unique start positions of reads aligning to the peak). To distinguish putative low-level novel L1 insertions from artifacts (such as chimeric reads derived from mispairing during library construction and PCR amplification), peaks without supporting reads marked as PCR duplicates and with fewer than five supporting reads in total were rejected. This was based on the rationale that each L1H insertion in the template gDNA was amplified by 20 PCR cycles, and their PCR duplicates should be sequenced under such high throughput (75 M read pairs) per library, while technical artifacts induced by non-specific or chimeric PCR amplification were inevitable but generated at a much lower rate. Finally, the putative L1 insertions were manually inspected using the Integrative Genomics Viewer (IGV)79 to prioritize insertions for experimental validation.
Experimental validation of HAT-seq with nested junction PCR and amplicon sequencing
To validate the presence of putative L1 insertions in the HCMV genome, we performed 3′ and 5′ junction nested PCR assays as described by Zhao et al.38. Three different templates were used as inputs: adapter-ligated HCMV fragments, the HAT-seq library, and the original HCMV genome DNA extraction. The resulting PCR products were visualized on 2% agarose gels and compared to the expected sizes. Bands of the correct size were excised and subjected to downstream amplicon sequencing at GENEWIZ, Inc. Raw amplicon sequencing data (2 × 250 bp) were preprocessed, and clean reads were aligned to the HCMV genome using the BWA-MEM algorithm (version 0.7.17-r1194-dirty). The predicted L1 integration sites were inspected using IGV. The validation primer sequences are listed in Supplementary Data 5.
Deep Illumina WGS library construction, sequencing, and data analysis for TE insertions
We performed deep short-read Illumina WGS on the WT, shLacZ, shL1-1, and shL1-2 HCMV DNA. WGS libraries of HCMV strains were constructed and paired-end sequenced (2 × 150 bp) at Macrogen, Inc. Raw sequencing data were preprocessed as described above and aligned to the HCMV genome using the BWA-MEM algorithm. To identify TE insertions, we modified the SCRAMble software (https://github.com/GeneDx/scramble)80 to increase the sensitivity for detecting low-level clonal TE insertion events, as the tool was originally developed for germline TE insertion calling. Specifically, in the generation of clipped reads cluster step, we altered the requirement for the minimum number of soft-clipped reads (-s) from 5 to 1 to include all clusters supported by at least one soft-clipped read. In the cluster characterization step, we modified the minimum cluster size to analyze (-n) from 5 to 1 to retain all candidate TE insertions supported by a single read. Subsequently, candidates lacking the predicted insertional features, such as poly-A sequences and terminal site duplications, were excluded. Final clonal TE insertion candidates were manually evaluated using the IGV visualization tool.
Short variant calling on Illumina datasets
WGS datasets from two batches of WT, shLacZ, shL1-1, and shL1-2 samples (n = 2 biological replicates per sample within each batch for a total of 16 samples) were analyzed. The Toledo BAC genome was used as the control. Based on benchmarking by Deng et al.81, we developed a variant calling pipeline using Snakemake82 to detect low-level SNPs and indels in viral genomes (Supplementary Fig. S9A). Illumina FASTQ files were adapter and quality trimmed using fastp83. The quality of the sequencing reads was assessed using FastQC84. Trimmed reads were then checked for contamination against hg38 using FastQ Screen85, and only the reads that aligned uniquely to the HCMV genome were retained. Disordered reads or reads with missing mates were repaired using BBTools Repair86. The reads were aligned to the HCMV Toledo reference genome using BWA-MEM72 (v0.7.17) and sorted using SAMtools87. Duplicates were marked using Picard88. The depth and quality of the alignment were assessed using Qualimap89 and Mosdepth90. To compare the mutation burden across samples, aligned reads were downsampled to the lowest median coverage per batch using SAMtools (~16,000 × for batch 1 and ~26,000 × for batch 2). We performed variant calling using two callers, Mutect241 and VarScan242, in paired tumor and normal modes. In each case, we treated the samples as “tumor” and the Toledo BAC control sample as “normal.” We applied FilterMutectCalls and SelectVariants to the Mutect2 calls. For VarScan2, we increased the pileup depth to account for our high-depth sequencing by setting the SAMtools mpileup -d 1000000. We ran VarScan somatic and VarScan processSomatic with the parameter --min-tumor-freq 0.005. We filtered out variant calls that were present in the same sample groups of both batches, as the batches were cultured independently. To further account for any possible ancestral calls, we filtered out WT or shLacZ control sample variants from the shL1 variant call sets and vice versa, within each batch. To obtain high-confidence call sets, we took the intersection of calls between Mutect2 and VarScan2 using BCFtools91. We further imposed a VAF cutoff of 50%, effectively filtering out mutations likely present in the original constructs (Supplementary Fig. S9D). We manually inspected the putative variant calls in IGV79. Genome-wide maps depicting SNP calls were generated using Circos92.
Quantification of HCMV DNA copy number
HFF and U373MG were infected at 0.5 MOI of HCMV and harvested at indicated time points. Total DNA was purified using the QIAamp DNA Blood Mini Kit (Qiagen). For the qPCR analysis of the purified DNA, 40 ng genomic DNA was subjected to qPCR. Primers and probes were used to detect 7SK for total genomic DNA normalization and UL86 for the HCMV genome, as previously described13. To create a standard curve of HCMV copy number, we used 2 pg to 2 ng of HCMV Toledo BAC DNA diluted with HFF genomic DNA to a total amount of 20 ng. The copy number of the HCMV genome was calculated using the standard curve.
EdU labeling of the HCMV genome
Imaging of replicating HCMV DNAs was performed using the Click-It Plus EdU Cell Proliferation Kit for Imaging, Alexa Fluor 488 dye (C10637; Invitrogen). First, 2 × 105 Primary HFFs were seeded onto coverglass in a 12-well plate. After 2 days, cells were infected with 0.5 MOI of HCMV. At 72 hpi, cells were incubated with fresh media supplemented with 10 μM EdU for 30 min. Cells were then fixed with 3.7% formaldehyde for 15 min, washed twice with PBS, and permeabilized using 0.5% Triton X-100 for 20 min. Cells were washed twice with 2% BSA in PBS and incubated with 50 μL Click-iT Plus reaction cocktail (1X Click-iT reaction buffer, copper protectant, Alexa Fluor picolyl azide, and reaction buffer additive) for 30 min. After incubation, cells were washed twice and further stained with HCMV UL44 mouse antibodies (CA006-1; Virusys) at a dilution of 1:400, Alexa Fluor-568-conjugated goat anti-mouse antibodies (A-11004; Invitrogen) at a dilution of 1:200, and Hoechst. Images were obtained by LSM980 confocal laser-scanning microscope (ZEISS) with a Plan-Apochromat 63×/1.4NA oil objective using Airyscan2 SR mode with GaAsP Array PMT, followed by analysis using ImageJ. The region-of-interest (ROI) for UL44 in individual HCMV-infected cells was manually selected, and EdU intensities were measured in that ROI.
EdU incorporation analysis using flow cytometry
HCMV-infected cells (2 × 105) cells at 72 hpi were incubated with 10 μM EdU for 3 h. Cells were washed with PBS twice, detached, and then resuspended in 200 μL of 3.7% formaldehyde in PBS for 2 min, subsequently permeabilized by slowly adding 1 mL of ice-cold 70% ethanol and incubated in 4 °C for 10 min. Cells were washed for 3 times with wash buffer (0.5% Tween-20 and 1% BSA in PBS), and incubated in 100 μL Click-iT Plus reaction cocktail (1X Click-iT reaction buffer, copper protectant, Alexa Fluor 488 picolyl azide, and reaction buffer additive) for 30 min with protection from light. After the click reaction, cells were washed 4 times in wash buffer and incubated in wash buffer supplemented with HCMV UL44 antibodies (CA006-1; Virusys) at a dilution of 1:2000 at 4 °C for overnight. The next day, cells were washed twice and incubated in wash buffer supplemented with anti-mouse Alexa Fluor 647 antibodies at a dilution of 1:2000 for 1 h at RT, washed twice, and then analyzed using a Flow-Activated Cell Sorter Canto II (BD Bioscience) and Flowjo software (Flowjo).
EdU and BrdU dual labeling of HCMV DNA for analysis of fork stalling
We adopted a previously described protocol with minor modification62. Primary HFFs (3 × 105) were seeded on coverslips and incubated until they reached confluent density. The cells were infected with 0.5 MOI HCMV and supplemented with 1 μM EdU at 48 hpi for 16 h. During the last 6 h of EdU labeling, cells were treated with 50 mM hydroxyurea. Alternatively, during the last 1 h of EdU labeling, cells were treated with 1 μM CPT, 10 μM Etoposide, or 100 μM hydrogen peroxide. After the removal of EdU and drugs, cells were incubated in the growth medium with 10 μM BrdU. After 3 h, cells were fixed with 3.7% formaldehyde and denatured using 4 N hydrochloric acid for 10 min. Cells were permeabilized using 0.5% Triton X-100 for 20 min and blocked using 2% BSA in PBS. After washing twice using 2% BSA in PBS, cells were incubated with 100 μL Click-iT Plus reaction cocktail for 30 min. Cells were then stained with BrdU antibody (5292; Cell Signaling Technology) at a dilution of 1:500 and AlexaFluor-568-conjugated goat anti-mouse antibodies (A-11004; Invitrogen) for 2 h at 37 °C. The cells on the coverslips were mounted on glass slides using the Vectashield antifade mounting medium containing DAPI. Images were obtained by confocal laser-scanning microscope LSM700 (ZEISS) with a C-Apochromat 40×/1.20NA water objective, followed by analysis using ImageJ. The ROI for EdU in an individual HCMV-infected cell was manually selected, and BrdU intensities were measured in that ROI.
Acceleration of native isolation of protein on nascent DNA (aniPOND)
AniPOND was conducted according to previously described methods with minor modifications13,50. First, 2.1 × 107 cells of pBL1ORF2p-3×FLAG U373MG cells were seeded and infected with 5 MOI HCMV the next day. At 72 hpi, cells (5 × 107) were labeled with DMSO (no EdU), 10 μM EdU for 15 min (EdU+), or 10 μM EdU and additional 30 min with 10 μM thymidine (Thy+). Nuclei fractions were then collected by incubation in a Nuclear Extraction Buffer (20 mM HEPES pH 7.2, 50 mM NaCl, 3 mM MgCl2, 300 mM Sucrose, and 0.5% IGEPAL CA630) for 20 min at 4 °C, followed by scraping and centrifugation at 800 × g for 10 min at 4 °C. Nuclei pellets were washed and incubated in Click reaction cocktail (5 μM Biotin Azide, 5 mM sodium ascorbate, and 2 mM CuSO4 in PBS) at a density of 2 × 107 cells per mL for 1 h at 4 °C. After Click reaction, Nuclei pellets were washed with ice-cold PBS, frozen and thawed by brief incubation at −80 °C for 5 min and incubated on ice. Pellets were then resuspended in 500 μL of ice-cold Buffer B1 (50 mM Tris-HCl pH 8.0, 25 mM NaCl, 2 mM EDTA, and 1% IGEPAL CA630) and incubated at 4 °C for 15 min. Pellets were lysed by sonication for 30 min (25% Amplitude; 10 s ON and OFF; Diagenode Bioruptor). After centrifugation at 13,000 × g for 10 min at 4 °C, supernatant fractions were collected and supplemented with an additional 500 μL of Buffer B1. Lysates were then incubated with 100 μL of Dynabeads MyOne Strepavidin T1 Bead (65601; Invitrogen) at 4 °C for 16 h. Beads were washed three times with Buffer B1. For immunoblot, 90% of beads were mixed with Lammeli sample buffer and boiled at 98 °C for 10 min. The remaining 10% of beads were incubated in DNA elution buffer (1% SDS and 100 mM NaHCO3) at 65 °C overnight, followed by DNA extraction using the MinElute PCR Purification kit (Qiagen). To assess the enrichment of host or HCMV DNA, Input and Capture DNA were subjected to PCR reactions using HCMV UL84 and human 7SK primers (Supplementary Table S3).
Statistical analysis
GraphPad Prism 7.00 was used for statistical analysis. A two-sided Student’s t-test was used for unpaired data. A two-sided Kolmogorov–Smirnov test was used to assess the quantification of the number and intensity of fluorescence signals. For multiple comparisons, one-way analysis of variance (ANOVA) with Tukey’s or Dunnett’s multiple comparison test was used. P-values < 0.05 were considered significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Genomic sequencing and HAT-seq data have been deposited at the National Center for Biotechnology Information Sequence Read Archive (NCBI SRA) under the BioProject PRJNA1144403. ATAC-seq has been deposit at NCBI under accession number GSE217152. All data supporting the findings of this study are provided in the accompanying Supplementary Information and Source Data files. Source data are provided with this paper.
References
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Hohjoh, H. & Singer, M. F. Cytoplasmic ribonucleoprotein complexes containing human LINE-1 protein and RNA. EMBO J. 15, 630–639 (1996).
Feng, Q., Moran, J. V., Kazazian, H. H. Jr. & Boeke, J. D. Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell 87, 905–916 (1996).
Mathias, S. L., Scott, A. F., Kazazian, H. H. Jr, Boeke, J. D. & Gabriel, A. Reverse transcriptase encoded by a human transposable element. Science 254, 1808–1810 (1991).
Kulpa, D. A. & Moran, J. V. Ribonucleoprotein particle formation is necessary but not sufficient for LINE-1 retrotransposition. Hum. Mol. Genet. 14, 3237–3248 (2005).
Cost, G. J., Feng, Q., Jacquier, A. & Boeke, J. D. Human L1 element target-primed reverse transcription in vitro. EMBO J. 21, 5899–5910 (2002).
Goodier, J. L. & Kazazian, H. H. Jr Retrotransposons revisited: the restraint and rehabilitation of parasites. Cell 135, 23–35 (2008).
Dowd, J. B., Aiello, A. E. & Alley, D. E. Socioeconomic disparities in the seroprevalence of cytomegalovirus infection in the US population: NHANES III. Epidemiol. Infect. 137, 58–65 (2009).
Weller, T. H. The cytomegaloviruses: ubiquitous agents with protean clinical manifestations. N. Engl. J. Med. 285, 203–214 (1971).
Weekes, MichaelP. et al. Quantitative temporal viromics: an approach to investigate host-pathogen interaction. Cell 157, 1460–1472 (2014).
Stern-Ginossar, N. et al. Decoding human cytomegalovirus. Science 338, 1088–1093 (2012).
Nobre, L. V. et al. Human cytomegalovirus interactome analysis identifies degradation hubs, domain associations and viral protein functions. ELife 8, e49894 (2019).
Manska, S. & Rossetto, C. C. Identification of cellular proteins associated with human cytomegalovirus (HCMV) DNA replication suggests novel cellular and viral interactions. Virology 566, 26–41 (2022).
Terhune, S. S. et al. Human cytomegalovirus UL29/28 protein interacts with components of the NuRD complex which promote accumulation of immediate-early RNA. PLoS Pathog. 6, e1000965 (2010).
Rauwel, B. et al. Release of human cytomegalovirus from latency by a KAP1/TRIM28 phosphorylation switch. Elife 4, e06068 (2015).
Nogalski, M. T. et al. A tumor-specific endogenous repetitive element is induced by herpesviruses. Nat. Commun. 10, 90 (2019).
Assinger, A., Yaiw, K.-C., Göttesdorfer, I., Leib-Mösch, C. & Söderberg-Nauclér, C. Human cytomegalovirus (HCMV) induces human endogenous retrovirus (HERV) transcription. Retrovirology 10, 132 (2013).
Rahman, M. J. et al. LINE-1 retrotransposons facilitate horizontal gene transfer into poxviruses. Elife 11, e63327 (2022).
Fixsen, S. M. et al. Poxviruses capture host genes by LINE-1 retrotransposition. Elife 11, e63332 (2022).
Krupovic, M., Makarova, K. S., Forterre, P., Prangishvili, D. & Koonin, E. V. Casposons: a new superfamily of self-synthesizing DNA transposons at the origin of prokaryotic CRISPR-Cas immunity. BMC Biol. 12, 36 (2014).
Tyl, M. D., Betsinger, C. N. & Cristea, I. M. Virus-host protein interactions as footprints of human cytomegalovirus replication. Curr. Opin. Virol. 52, 135–147 (2022).
Browne, E. P., Wing, B., Coleman, D. & Shenk, T. Altered cellular mRNA levels in human cytomegalovirus-infected fibroblasts: viral block to the accumulation of antiviral mRNAs. J. Virol. 75, 12319–12330 (2001).
Coufal, N. G. et al. L1 retrotransposition in human neural progenitor cells. Nature 460, 1127–1131 (2009).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Yang, N., Zhang, L., Zhang, Y. & Kazazian, H. H. Jr An important role for RUNX3 in human L1 transcription and retrotransposition. Nucleic Acids Res. 31, 4929–4940 (2003).
Athanikar, J. N., Badge, R. M. & Moran, J. V. A YY1-binding site is required for accurate human LINE-1 transcription initiation. Nucleic Acids Res. 32, 3846–3855 (2004).
Choi, J., Hwang, S.-Y. & Ahn, K. Interplay between RNASEH2 and MOV10 controls LINE-1 retrotransposition. Nucleic Acids Res. 46, 1912–1926 (2018).
Moldovan, J. B. & Moran, J. V. The zinc-finger antiviral protein ZAP inhibits LINE and Alu retrotransposition. PLoS Genet. 11, e1005121 (2015).
Goodier, J. L., Cheung, L. E. & Kazazian, H. H. Jr. Mapping the LINE1 ORF1 protein interactome reveals associated inhibitors of human retrotransposition. Nucleic Acids Res. 41, 7401–7419 (2013).
Taylor, M. S. et al. Affinity proteomics reveals human host factors implicated in discrete stages of LINE-1 retrotransposition. Cell 155, 1034–1048 (2013).
Appleton, B. A., Loregian, A., Filman, D. J., Coen, D. M. & Hogle, J. M. The cytomegalovirus DNA polymerase subunit UL44 forms a C clamp-shaped dimer. Mol. Cell 15, 233–244 (2004).
Gulbis, J. M., Kelman, Z., Hurwitz, J., O’Donnell, M. & Kuriyan, J. Structure of the C-terminal region of p21WAF1/CIP1 complexed with human PCNA. Cell 87, 297–306 (1996).
Appleton, B. A. et al. Crystal structure of the cytomegalovirus DNA polymerase subunit UL44 in complex with the C terminus from the catalytic subunit: differences in structure and function relative to unliganded UL44. J. Biol. Chem. 281, 5224–5232 (2006).
Strang, B. L. et al. Human cytomegalovirus UL44 concentrates at the periphery of replication compartments, the site of viral DNA synthesis. J. Virol. 86, 2089–2095 (2012).
Gasior, S. L., Wakeman, T. P., Xu, B. & Deininger, P. L. The human LINE-1 retrotransposon creates DNA double-strand breaks. J. Mol. Biol. 357, 1383–1393 (2006).
Rogakou, E. P., Pilch, D. R., Orr, A. H., Ivanova, V. S. & Bonner, W. M. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J. Biol. Chem. 273, 5858–5868 (1998).
Khadgi, B. B., Govindaraju, A. & Christensen, S. M. Completion of LINE integration involves an open ‘4-way’ branched DNA intermediate. Nucleic Acids Res. 47, 8708–8719 (2019).
Zhao, B. et al. Somatic LINE-1 retrotransposition in cortical neurons and non-brain tissues of Rett patients and healthy individuals. PLoS Genet. 15, e1008043 (2019).
Cunningham, C. et al. Sequences of complete human cytomegalovirus genomes from infected cell cultures and clinical specimens. J. Gen. Virol. 91, 605–615 (2010).
Renzette, N. et al. Limits and patterns of cytomegalovirus genomic diversity in humans. Proc. Natl Acad. Sci. USA 112, E4120–E4128 (2015).
Van der Auwera, G. A., O’Connor, B. D. & Safari, A. O. R. M. C. Genomics in the Cloud: Using Docker, GATK, and WDL in Terra 1st edn (O’Reilly Media, 2020).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Salvant, B. S., Fortunato, E. A. & Spector, D. H. Cell cycle dysregulation by human cytomegalovirus: influence of the cell cycle phase at the time of infection and effects on cyclin transcription. J. Virol. 72, 3729–3741 (1998).
Merchut-Maya, J. M. et al. Human cytomegalovirus hijacks host stress response fueling replication stress and genome instability. Cell Death Differ. 29, 1639–1653 (2022).
Zheng, L. et al. Novel function of the flap endonuclease 1 complex in processing stalled DNA replication forks. EMBO Rep. 6, 83–89 (2005).
Hanada, K. et al. The structure-specific endonuclease Mus81 contributes to replication restart by generating double-strand DNA breaks. Nat. Struct. Mol. Biol. 14, 1096–1104 (2007).
Hsiang, Y. H., Lihou, M. G. & Liu, L. F. Arrest of replication forks by drug-stabilized topoisomerase I-DNA cleavable complexes as a mechanism of cell killing by camptothecin. Cancer Res. 49, 5077–5082 (1989).
Zellweger, R. et al. Rad51-mediated replication fork reversal is a global response to genotoxic treatments in human cells. J. Cell Biol. 208, 563–579 (2015).
Berniak, K. et al. Relationship between DNA damage response, initiated by camptothecin or oxidative stress, and DNA replication, analyzed by quantitative 3D image analysis. Cytom. A 83, 913–924 (2013).
Leung, K. H., Abou El Hassan, M. & Bremner, R. A rapid and efficient method to purify proteins at replication forks under native conditions. Biotechniques 55, 204–206 (2013).
Molaro, A. & Malik, H. S. Hide and seek: how chromatin-based pathways silence retroelements in the mammalian germline. Curr. Opin. Genet Dev. 37, 51–58 (2016).
Verheul, T. C. J., van Hijfte, L., Perenthaler, E. & Barakat, T. S. The why of YY1: mechanisms of transcriptional regulation by Yin Yang 1. Front. Cell Dev. Biol. 8, 592164 (2020).
Wang, D. et al. The transcription factor Runx3 establishes chromatin accessibility of cis-regulatory landscapes that drive memory cytotoxic T lymphocyte formation. Immunity 48, 659–674.e656 (2018).
Nevels, M., Paulus, C. & Shenk, T. Human cytomegalovirus immediate-early 1 protein facilitates viral replication by antagonizing histone deacetylation. Proc. Natl Acad. Sci. USA 101, 17234–17239 (2004).
Grunstein, M. Histone acetylation in chromatin structure and transcription. Nature 389, 349–352 (1997).
Dargan, D. J. et al. Sequential mutations associated with adaptation of human cytomegalovirus to growth in cell culture. J. Gen. Virol. 91, 1535–1546 (2010).
Farré, D., Engel, P. & Angulo, A. Novel role of 3’UTR-embedded Alu elements as facilitators of processed pseudogene genesis and host gene capture by viral genomes. PLoS ONE 11, e0169196 (2016).
Ravichandran, S. et al. Genome-wide analysis of regulatory G-quadruplexes affecting gene expression in human cytomegalovirus. PLoS Pathog. 14, e1007334 (2018).
E, X. et al. An E2F1-mediated DNA damage response contributes to the replication of human cytomegalovirus. PLoS Pathog. 7, e1001342 (2011).
Casavant, N. C. et al. Potential role for p53 in the permissive life cycle of human cytomegalovirus. J. Virol. 80, 8390–8401 (2006).
Castillo, J. P. et al. Human cytomegalovirus IE1-72 activates ataxia telangiectasia mutated kinase and a p53/p21-mediated growth arrest response. J. Virol. 79, 11467–11475 (2005).
Schilling, E. M., Scherer, M., Rothemund, F. & Stamminger, T. Functional regulation of the structure-specific endonuclease FEN1 by the human cytomegalovirus protein IE1 suggests a role for the re-initiation of stalled viral replication forks. PLoS Pathog. 17, e1009460 (2021).
E, X. F. et al. RNA interference-mediated targeting of human cytomegalovirus immediate-early or early gene products inhibits viral replication with differential effects on cellular functions. J. Virol. 86, 5660–5673 (2012).
Ardeljan, D. et al. Cell fitness screens reveal a conflict between LINE-1 retrotransposition and DNA replication. Nat. Struct. Mol. Biol. 27, 168–178 (2020).
Mita, P. et al. BRCA1 and S phase DNA repair pathways restrict LINE-1 retrotransposition in human cells. Nat. Struct. Mol. Biol. 27, 179–191 (2020).
Krueger, F., James, F., Ewels, P., Afyounian, E. & Schuster-Boeckler, B. TrimGalore: v0.6.7. Zenodo. https://doi.org/10.5281/zenodo.5127898 (2021).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2012).
Frankish, A. et al. GENCODE 2021. Nucleic Acids Res. 49, D916–D923 (2020).
Jin, Y., Tam, O. H., Paniagua, E. & Hammell, M. TEtranscripts: a package for including transposable elements in differential expression analysis of RNA-seq datasets. Bioinformatics 31, 3593–3599 (2015).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 1–21 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Amemiya, H. M., Kundaje, A. & Boyle, A. P. The ENCODE blacklist: identification of problematic regions of the genome. Sci. Rep. 9, 1–5 (2019).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Lee, H., Heo, L., Lee, M. S. & Seok, C. GalaxyPepDock: a protein-peptide docking tool based on interaction similarity and energy optimization. Nucleic Acids Res. 43, W431–W435 (2015).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 30, 614–620 (2014).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Torene, R. I. et al. Mobile element insertion detection in 89,874 clinical exomes. Genet. Med. 22, 974–978 (2020).
Deng, Z. L. et al. Evaluating assembly and variant calling software for strain-resolved analysis of large DNA viruses. Brief Bioinform. 22, bbaa123 (2021).
Koster, J. & Rahmann, S. Snakemake—a scalable bioinformatics workflow engine. Bioinformatics 28, 2520–2522 (2012).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (2010).
Wingett, S. W. & Andrews, S. FastQ Screen: a tool for multi-genome mapping and quality control. F1000Res 7, 1338 (2018).
Bushnell, B., Rood, J. & Singer, E. BBMerge—accurate paired shotgun read merging via overlap. PLoS ONE 12, e0185056 (2017).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Broad Institute. Picard Toolkit. http://broadinstitute.github.io/picard (2019).
Garcia-Alcalde, F. et al. Qualimap: evaluating next-generation sequencing alignment data. Bioinformatics 28, 2678–2679 (2012).
Pedersen, B. S. & Quinlan, A. R. Mosdepth: quick coverage calculation for genomes and exomes. Bioinformatics 34, 867–868 (2018).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Acknowledgements
We are grateful to Prof. Thomas Shenk (Princeton University) for providing HCMV Toledo and AD169 BAC and Prof. Nian Liu (Tsinghua University) for providing pBL1 plasmid. We would like to express sincere gratitude to all members of Ahn and Lee laboratories for discussion and technical support. This work was supported by the Institute for Basic Science of the Ministry of Science Grant IBS-R008-D1 and the National Research Foundation of Korea (NRF) grant funded by the Korean government NRF-2020R1A2C3011298 and NRF-2020R1A5A101808 (K.A.). This work was also supported by the National Institute of Health (NIH) (DP2 AG072437), the Suh Kyungbae Foundation, and the Allen Discovery Center program, a Paul G. Allen Frontiers Group advised program of the Paul G. Allen Family Foundation (E.A.L., D.D., B.Z.). B.Z. was supported by the Manton Center Pilot Project Award and Rare Disease Research Fellowship. The funders had no role in the study design, data collection, analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
S.-Y.H., H.K., B.Z., D.D., E.A.L. and K.A. contributed to the conceptualization and designed the experiments. S.-Y.H., H.K., J.J., K.P., D.J., and S.P. performed the biochemical and cell biology experiments. S.-Y.H., H.K., and J.P. performed image analyses. D.L. and S.K. analyzed RNA-seq and ATAC-seq data. B.Z. performed HAT-seq library construction, data analysis, and experimental validation. D.D., B.Z. and E.A.L. analyzed the deep-sequencing data and performed mutation burden analysis. J.K. and H.-J.C. performed the structural modeling. S.-Y.H., H.K., B.Z. and D.D. wrote the manuscript with input from all authors. E.A.L. and K.A. supervised the overall study and revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Mary Hummel and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hwang, SY., Kim, H., Denisko, D. et al. Human cytomegalovirus harnesses host L1 retrotransposon for efficient replication. Nat Commun 15, 7640 (2024). https://doi.org/10.1038/s41467-024-51961-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-51961-y
- Springer Nature Limited