Abstract
Hypoxia occurs in 90% of solid tumors and is associated with metastasis and mortality. Breast cancer cells that experience intratumoral hypoxia are 5x more likely to develop lung metastasis in animal models. Using spatial transcriptomics, we determine that hypoxic cells localized in more oxygenated tumor regions (termed ‘post-hypoxic’) retain expression of hypoxia-inducible and NF-kB-regulated genes, even in the oxygen-rich bloodstream. This cellular response is reproduced in vitro under chronic hypoxic conditions followed by reoxygenation. A subset of genes remains increased in reoxygenated cells. MUC1/MUC1-C is upregulated by both HIF-1α and NF-kB-p65 during chronic hypoxia. Abrogating MUC1 decreases the expression of superoxide dismutase enzymes, causing reactive oxygen species (ROS) production and cell death. A hypoxia-dependent genetic deletion of MUC1, or MUC1-C inhibition by GO-203, increases ROS levels in circulating tumor cells (CTCs), reducing the extent of metastasis. High MUC1 expression in tumor biopsies is associated with recurrence, and MUC1+ CTCs have lower ROS levels than MUC1- CTCs in patient-derived xenograft models. This study demonstrates that therapeutically targeting MUC1-C reduces hypoxia-driven metastasis.
Similar content being viewed by others
Introduction
Hypoxia is associated with metastasis and mortality in patients with breast cancer1. Despite decades of research on hypoxia and hypoxia inducible factor (HIF) target genes and their role in metastasis, it was unclear how intratumoral hypoxia directly contributes to the metastatic process, considering cells undergo some level of reoxygenation in the bloodstream. By developing a fluorescent hypoxia fate-mapping system, we previously determined that cells that experience intratumoral hypoxia are protected from reactive oxygen species (ROS) in the bloodstream, increasing their ability to form a metastasis2. Whether hypoxia-induced signaling stays active in disseminated cells remained elusive. One study demonstrated that of the patients with metastatic breast cancer in their study, 76% scored positive for HIF-1α expression in circulating tumor cells (CTCs)3, suggesting that HIF signaling might be preserved in cancer cells found in the circulation. Furthermore, the effectors driving the metastatic phenotype were yet to be identified.
Breast cancer cells have increased ROS levels compared to adjacent normal cells4. Hypoxia leads to ROS accumulation5,6,7, which can promote the activation of other stress responses. For instance, studies have shown that ROS accumulation leads to Nuclear Factor kappa-light-chain-enhancer of activated B cells (NF-kB) phosphorylation and activation8,9. Both HIF-1α10,11 and NF-kB12,13 have established roles in redox homeostasis. Mucin 1, encoded by the gene MUC1, is a heterodimeric protein located at the cell membrane, which is composed of MUC1-N (extracellular) and MUC1-C (cytoplasmatic) subunits. Under stress, while MUC1-N is shed, MUC1-C is released into the cytoplasm and can contribute to the expression of antioxidant effector enzymes14. Studies have established that MUC1-C can translocate to the nucleus15 and bind to transcription factors to regulate downstream transcriptional activity. Interestingly, it has been demonstrated that MUC1-C can bind to NF-kB p65 and transcriptionally regulate NF-kB16.
In this work, we optimized a protocol to combine fluorescent mapping of hypoxic cells (i.e., cells exposed to < 1% O2 express a green fluorescent protein (GFP)) with spatial transcriptomics. This approach allowed us to delineate the transcriptional profile of hypoxic cells, compared to cells localized in more well-oxygenated tumor regions (termed ‘post-hypoxic’), and cells that never experienced intratumoral hypoxia. In addition, we developed a CRISPR-based methodology to knock out any gene of interest when a cell undergoes hypoxia. Using this integrative approach, we identified a potential mechanism that may contribute to the metastatic potency of these cells. We demonstrate that GFP + cells are enriched for the expression of genes that play a role in hypoxia and NF-kB signaling in response to intratumoral hypoxia and ROS. The gene expression profile of GFP + cells is similar to that of cells exposed to chronic but not acute hypoxia in vitro. Post-hypoxic cells in vivo maintained an enrichment in hypoxia signaling which supports the notion of a “hypoxia memory'’ or HM. One of the HM genes, MUC1, is induced in a HIF-1α and NF-kB-dependent manner and confers resistance to ROS by promoting the expression of superoxide dismutase (SOD) enzymes. This antioxidant response enhances the survival of CTCs, which could contribute to metastasis. MUC1-C inhibitors, such as GO-203, currently in phase II clinical trials, could be considered in future studies to prevent metastasis by eliminating CTCs that were exposed to hypoxia.
Results
GFP + CTCs retain a memory of the hypoxic state
By designing a hypoxia fate-mapping system that utilizes Cre-recombination to trigger a switch from DsRed to GFP expression when cells experience intratumoral hypoxia, we found that GFP + cells were 5x more likely to form a metastatic lung lesion due to a ROS-resistant phenotype that promoted their survival in the bloodstream2 (Fig. 1a). To further investigate the mechanistic drivers of this phenotype, we performed RNA sequencing to transcriptionally profile GFP + and DsRed + cells that were fluorescently sorted (FACS) from the primary tumor or metastatic lung lesions. Both hypoxia and TNFα signaling via NF-kB activation pathways were highly enriched in a Hallmark gene set analysis that compared gene expression in GFP + versus DsRed + cells in the tumor and lung (Fig. 1b). Both hypoxia and NF-kB signaling have been shown to promote cancer progression and metastasis and are regulated in response to oxidative stress12,17.
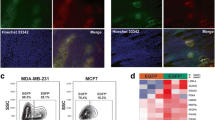
a Dual-vector hypoxia fate-mapping system. Cancer cells express DsRed under physiological O2 (6%) levels, or GFP after a Cre-mediated switch under hypoxia (0.5% O2) in vivo. Previous studies demonstrated that post-hypoxic MDA-MB-231 tumor cells had a 5-times higher probability of establishing a lung metastatic lesion in NSG mice due to a ROS-resistant phenotype that promoted survival in the bloodstream. b Cluster analysis of gene expression patterns from bulk RNA sequencing analysis to compare normalized enrichment scores (NES) of hallmark gene sets in GFP + versus DsRed + FACS-sorted tumor (TG/TR) or metastatic cells in the lung (LG/LR). c Spatial transcriptomics was applied to tumor sections obtained from mice bearing tumors from MDA-MB-231 hypoxia fate-mapping cells (left). The clustering analysis revealed gene expression clusters that were visually identified in hypoxic, post-hypoxic, or non-hypoxic regions of the tumor (right). d Magnified view of a tumor overlayed with gene expression clusters. e, f Top enriched hallmark gene sets in hypoxic (cluster #2) (e) and post-hypoxic (cluster #1) (f) clusters compared to non-hypoxic (cluster #0) tissue (adjusted p-value ****< 0.0001, **< 0.001, *< 0.05, versus normal). g Low input RNA sequencing was performed in DsRed + and matched GFP + CTCs isolated from the blood of tumor-bearing mice. Hallmark gene sets of interest enriched in GFP + versus DsRed + CTCs (BG/BR) are displayed. Source data are provided as a Source Data file. Cartoons adapted (altered cell colors) under Creative Commons Attribution 4.0 Unported License (https://creativecommons.org/licenses/by/4.0/).
To understand the role of hypoxia in metastatic progression, we sought to spatially map gene expression in hypoxic cells compared with cells that potentially migrated to more oxygenated tumor regions. We adapted the Visium spatial transcriptomics protocol to perform concurrent transcriptional profiling and fluorescent imaging of our hypoxia fate-mapping system (Fig. 1c and Supplementary Fig. 1a, b). We applied this technology to two primary tumor tissue sections and analyzed gene expression data using the R/Bioconductor package Seurat18. The spatial transcriptomic data of tumor #1 revealed three main gene expression clusters: cluster #0, located in the tumor periphery, and two distinct clusters (#1 and #2) identified within the GFP + tumor region, which could not be distinguished from the imaging data alone. One cluster was located immediately adjacent to the peri-necrotic tumor region that we termed ‘hypoxic’. The second cluster was located adjacent to the hypoxic cluster and dispersed within DsRed + regions, which we termed ‘post-hypoxic’ (Fig. 1c, d and Supplementary Fig. 1c, d). Gene set enrichment analysis (GSEA) showed that the most up-regulated pathways in tumor #1 cluster 0 (non-hypoxic) were proliferation-related, while the most down-regulated pathways were hypoxia, glycolysis, and NF-kB, in comparison to the other clusters (Supplementary Fig. 1e). This is consistent with a previous analysis using bulk RNA sequencing (Fig. 1b)2. Tumor #2 contained 6 unique clusters. Clusters #1 and #3 (non-hypoxic) showed similar enrichment patterns as cluster 0 in tumor #1 with negative enrichment for hypoxia and positive enrichment for proliferation pathways (Supplementary Fig. 1e). The UMAP embedding showed that the post-hypoxic cluster in tumor #2 (cluster #2) falls between the clusters that were enriched for hypoxia signaling (cluster #0 and #4) and the non-hypoxic clusters based on their transcriptional profile, spatial localization, and UMAP space (Supplementary Fig. 1c, d). To determine whether these cells are undergoing reoxygenation, we performed differential expression analysis of hypoxic and post-hypoxic populations (clusters #2 versus #1, respectively, in tumor #1). The hypoxic population was enriched for the expression of the hypoxia hallmark gene set (Supplementary Fig. 1f) whereas the post-hypoxic cells were enriched in proliferative pathways (Supplementary Fig. 1g). However, there were also similarities in hypoxic and post-hypoxic populations (clusters #2 and #1, respectively, in tumor #1) compared with the non-hypoxic population (cluster #0, tumor #1) (Fig. 1e, f). Taken together, the data is in line with the notion that reoxygenation can occur to some extent in a tumor, which reverses the expression of some but not all hypoxia-inducible gene products. Likewise, similar observations were found when comparing GFP + cells that metastasized to the lung (reoxygenated) versus GFP + cells collected from the tumor (Fig. 1b).
We previously determined that GFP + CTCs survive better than DsRed + CTCs in the bloodstream due to a ROS-resistant phenotype2. To determine whether the post-hypoxic tumor phenotype was retained following intravasation into the bloodstream, we isolated CTCs derived from tumor-bearing mice and FACS-sorted them into DsRed + or GFP + populations. We performed low-input RNA sequencing on the sorted CTC populations to further test whether the observed transcriptional changes induced by intratumoral hypoxia are preserved in CTCs. GSEA revealed that GFP + CTCs were also enriched in hypoxia, NF-kB, and glycolysis hallmark gene sets compared to the gene expression patterns in the DsRed + CTCs (Fig. 1g and Supplementary Table 1). Together, our results suggest that intratumoral hypoxia promotes hypoxia and NF-kB signaling, which is maintained to some extent even when GFP + cells (1) invade into more oxygenated tumor regions, (2) intravasate into the bloodstream, and (3) colonize the lung.
Chronic but not acute hypoxia causes NF-kB signaling
To confirm NF-kB activation in vivo and validate our transcriptomic studies, we transduced MDA-MB-231 hypoxia fate-mapping cells with a reporter that causes mKate2 expression upon NF-kB binding19 (Fig. 2a). The cells were implanted into the mammary fat pad of NSG mice, and blood, tumors, and lungs were harvested 4 weeks later. Flow cytometry analysis of mKate2 expression in DsRed + or GFP + cells revealed a significant increase in NF-kB activation in GFP + cancer cells (Fig. 2b and Supplementary Fig. 2a). In the primary tumor and at the metastatic site, the GFP and mKate2 signals colocalized (Fig. 2c, d and Supplementary Fig. 2b, c). Likewise, the immunolabeled expression of the phosphorylated p65 subunit of NF-kB (pNF-kB) overlapped with GFP + regions of orthotopic tumors derived from a syngeneic model of breast cancer (Supplementary Fig. 2d, e). Next, we stratified patients with breast cancer into high versus low protein expression of pNF-kB S536 using protein data available in the TCGA database generated by the reverse phase protein array high-throughput antibody-based technique20 (TCGA-RPPA, N = 873). NF-kB phosphorylation correlated with worse overall survival (Supplementary Fig. 2f). Together, the results show that intratumoral hypoxia causes NF-kB activation, which correlates with reduced overall survival in patients with breast cancer.
a Schematic of NF-kB-mKate2 reporter construct that drives mKate2 expression regulated by an NF-kB-responsive promoter. Fluorescent images of reporter MDA-MB-231 cells cultured in the absence or presence of 10 ng/mL TNFα. b Levels of mKate2 were assessed using flow cytometry in DsRed + and GFP + cells derived from tumor, blood, and lung samples of mice (mean ± SEM, N = 2: set 1 – n = 3 (blood) or n = 4 (tumor and lung) and set 2 – n = 7 (blood) or n = 8 (tumor and lung); GFP versus DsRed, RM two-way ANOVA with Fisher’s LSD post-test; The box extends from the 25th to 75th percentiles, the median is marked by the vertical line inside the box, and the whiskers represent the minimum and maximum points). c, d Representative fluorescent images displaying DsRed, GFP, and mKate2 expression in MDA-MB-231 tumor (c) and lung (d) sections. e Venn diagram displaying the overlap of the hallmark gene sets enriched in MDA-MB-231 cells exposed to hypoxia (1% O2) for 1 (acute) or 10 days (chronic). f Immunoblot assay of MDA‐MB‐231 cells cultured under 20% or 1% O2 for 4h, or 1, 3, 5, 7, and 10 days to assess the levels of HIF-1α, HIF-2α, pNF-kB p65, total NF-kB p65, pIKBα, and total IKBα. g Fluorescent images of MDA-MB-231 NF-kB.mKate2 reporter cells cultured under 20% O2 for 5 days and 1% O2 for 2 or 5 days. Quantification of fluorescence intensity measured by image analysis (mean ± SEM, N = 1, n = 20 fields of view; 1% versus 20%, unpaired t-test two-tailed). For boxplots, the box extends from the 25th to 75th percentiles, the median is marked by the vertical line inside the box, and the whiskers represent the minimum and maximum points. Source data are provided as a Source Data file. Cartoons adapted (altered cell colors) under Creative Commons Attribution 4.0 Unported License (https://creativecommons.org/licenses/by/4.0/).
Within a tumor, cells undergo varying durations and gradients of hypoxia21. We hypothesized that longer-term exposure to hypoxia might cause enrichment in the pathways observed in vivo. To address this, we performed RNA sequencing on the lysate from MDA-MB-231 cells cultured under 20% or 1% O2 for 1 day (acute) or 10 days (chronic). We verified that cells exposed to chronic hypoxia were enriched in hallmark gene sets highly expressed in GFP + tumor cells, such as NF-kB (Fig. 2e). To confirm NF-kB activation under chronic hypoxia, we evaluated the phosphorylation status of NF-kB and IKBα in cells exposed to 20% versus 1% O2 in a 10-day time course. After 5 days of hypoxia, we observed an increase in pNF-kB/NF-kB p65 and pIKBα/IKBα, which suggests activation of the canonical NF-kB signaling pathway (Fig. 2f). To verify the induction of well-known NF-kB effectors, we assessed superoxide dismutase 2 (SOD2)22 and cyclooxygenase-2 (COX-2, encoded by PTGS2)23 expression. The expression of SOD2 and PTGS2 was induced by TNFα and reduced by the NF-kB inhibitor BAY 11-7082 (Supplementary Fig. 2g, h). Both proteins showed induction starting at day 5 that increased with time to day 10 (Supplementary Fig. 2i). Similar findings were observed in MDA-MB-436 cells (Supplementary Fig. 2j). Overall mKate expression was also increased in the NF-kB.mKate2 reporter cells exposed to 1% versus 20% O2 (Fig. 2g). The observations suggested that 5 days of exposure to chronic hypoxia and/or responses that co-occur with chronic hypoxia (i.e., cell starvation, metabolic reprogramming, oxidative stress, etc) in vitro are sufficient to activate the canonical NF-kB pathway, which is in line with the results from our in vivo study.
HIF-1 and NF-kB regulate antioxidant genes in response to ROS
Previous studies have shown that hypoxia increases mitochondrial ROS levels, mainly via superoxide production through NADPH oxidase activation. Reoxygenation (hyperoxia) also causes ROS accumulation mainly via complex I deactivation, which can result in DNA damage5,24. We confirmed that cancer cells in the peri-necrotic region of tumors (GFP +) have excessive DNA damage caused by oxidative stress, as detected by 8-Oxoguanine + immunofluorescent staining (Fig. 3a). In addition to promoting HIF-1 expression25, ROS regulates NF-kB activity12. Thus, we hypothesized that intratumoral hypoxia promotes the generation of ROS, leading to NF-kB activation as a survival response. First, we demonstrated that cells experienced a surge of mitochondrial ROS in response to acute hypoxia (2 days), followed by a decrease measured at day 5 (Fig. 3b). Next, we measured both CellROX (cytoplasmatic ROS) and H2O2 production and found that unlike MitoROS they both increased with time under hypoxia (Supplementary Data Fig. 2k, l). This observation suggests that mitochondrial ROS (O2-) is metabolized to H2O2 under prolonged hypoxia. H2O2 can activate NF-kB activity26, which we confirmed using the NF-kB.mKate2 reporter in MDA-MB-231 cells treated with H2O2 (Supplementary Data Fig. 2m). To determine if hypoxia-induced ROS production promotes NF-kB activation, we exposed NF-kB.mKate2 reporter cells to 20% or 1% O2 for 5 days in the presence or absence of the ROS inhibitor, N-acetyl-L-cysteine (NAC). NAC treatment significantly decreased the levels of mKate2 reporter expression in both MDA-MB-231 cells (Fig. 3c, d). The NF-kB inhibitor, BAY 11-7082, was utilized as a control and reduced NF-kB.mKate2 activity to basal levels. Furthermore, NAC treatment reduced the level of pNF-kB, as well as both protein and mRNA levels of the NF-kB downstream targets, PTGS2/COX-2 and SOD226 (Fig. 3e and Supplementary Figs. 2n, o and 3a). Altogether, our results demonstrate that hypoxia-dependent mitochondrial ROS accumulation leads to H2O2 production, which causes NF-kB signaling under prolonged but not short-term hypoxia.
a MDA-MB-231 tumor section imaged to visualize DsRed, GFP, and 8-Oxoguanine + labeling with normalized intensity plots (N = 6). b MitoROS levels assessed using flow cytometry in MDA‐MB‐231 cells cultured under 20% versus 1% O2 for 2 or 5 days (N = 2; n = 3 except for set 1 d2 20% n = 2, Ordinary two-way ANOVA with Bonferroni post-test). Representative flow cytometry histograms (right). c Fluorescent images of NF-kB.mKate2 MDA-MB-231 or MDA-MB-436 cells cultured under 1% O2 for 5 days with or without NAC. d Quantification of mKate2 expression by flow cytometry in MDA-MB-231 cells treated with NAC or BAY 11-7082 (N = 6; versus control (C) 1%, RM one-way ANOVA with Fisher’s LSD post-test). e COX2, pNF-kB p65, NF-kB, and SOD2 protein expression in MDA‐MB‐231 cells cultured under hypoxia for 5 days with or without NAC treatment. f Venn diagram displaying the number of genes enriched in GFP + versus DsRed + tumor cells overlapping with genes enriched in MDA-MB-231 cells exposed to 1% O2 versus 20% O2 for 1 or 10 days of hypoxia. A number of upregulated (standard font), downregulated (underlined), or oppositely regulated (italic) genes for each comparison (Supplementary Table 2). g Sixteen ROS-protective genes were upregulated by both intratumoral and chronic hypoxia exposure and organized by reported or predicted HIF- and/or NF-kB-dependent regulation. h mRNA expression levels of genes (g) in MDA-MB-231 cells under acute (1d) or chronic (5d) hypoxia conditions. i MUC1, FN1, MTUS1, RORA, RAPGEF4, and CA9 mRNA expression levels in MDA-MB-231 control (NTC), HIF-1α (H1-1/H1-2), or HIF-2α (H2-1/H2-2) knockout cells cultured under 20% or 1% O2 for 5 days (N = 3, n = 3 except for RAPGEF4 n = 2; versus NTC 1%, Kruskal-Wallis test with Dunn’s post-test). j Relative MUC1, FN1, MTUS1, RORA, RAPGEF4, and CA9 mRNA expression in MDA-MB-231 control or RELA (R-1/R-2) knockout cells. k Fold-change of mRNA expression summarized from (i) and (j). Data displayed as mean ± SEM; N = biological and n = technical replicates. For boxplots, boxes are median centered and extend from 25th to 75th percentiles, with whiskers representing minimum and maximum points. Source data are provided as a Source Data file.
Studies have demonstrated that cells in hypoxic conditions survive in part by balancing ROS levels. Both HIF-1α10 and NF-kB12 have been implicated in antioxidant responses via transcriptional regulation of antioxidant gene products. This aligns with our hypothesis that intratumoral hypoxia causes ROS generation, which activates protective mechanisms such as a survival response, potentially via HIF-1 and/or NF-kB signaling. This adaptation may reduce the levels of mitochondrial ROS, selectively increasing the survival of GFP + CTCs when facing a re-exposure to ROS in the bloodstream, which occurs due to reoxygenation and physical/chemical stress27.
To determine possible downstream effectors of HIF-1α and/or NF-kB, we compared the intratumoral hypoxic gene expression changes from bulk RNA sequencing with the changes observed in vitro when cells are exposed to chronic hypoxia. This analysis identified 55 genes upregulated by both intratumoral and chronic in vitro hypoxic conditions (Fig. 3f and Supplementary Table 2). Twenty-one of the 55 genes have been reported to promote ROS (5 genes) or protect cells from ROS (16 genes). Twelve of the 16 gene products that protect cells from ROS have been reported or predicted to be regulated by HIF-1/2, NF-kB, or both (Fig. 3g and Supplementary Tables 3 and 4). To confirm the expression of the 16 anti-ROS genes, we performed RT-qPCR and measured the level of induction of each gene under acute and chronic hypoxia in MDA-MB-231 cells across three independent repeats. MUC1, FN1, MTUS1, RORA, RAPGEF4, and CA9 were robustly induced under chronic hypoxia (Fig. 3h and Supplementary Fig. 3b). The induction of this 6-gene panel was validated at the protein level (except MTUS1) and also observed in an independent cell line, MDA-MB-436 (Supplementary Fig. 3c, d).
To determine the requirement of HIFs in regulating the 6-gene panel, we utilized a CRISPR knockout approach targeting HIF1A (encoding HIF-1α) or EPAS1 (encoding HIF-2α), as well as a non-target control gRNA (NTC). MDA-MB-231 NTC cells were cultured under 20% or 1% O2 for 5 days, whereas HIF-knockout cells were only cultured under 1% O2. MUC1, FN1, RORA, RAPGEF4, and CA9 mRNA levels were significantly reduced in the HIF-1α knockout cells cultured under hypoxia, but not in the HIF-2α knockout cells (Fig. 3i). To determine the role of NF-kB in regulating this 6-gene panel, MDA-MB-231 cells were exposed to 20% or 1% O2 for 5 days in the presence or absence of 10 μM BAY 11-7082 or DMSO for the last 24 h of the experiment. BAY11-7082 treatment decreased the expression of MUC1, FN1, and MTUS1 under hypoxia (Supplementary Fig. 3e). Similar to BAY 11-7082 treatment, the CRISPR knockout of RELA (encoding NF-kB p65) significantly abolished the mRNA expression of MUC1, FN1 and MTUS1 upon chronic exposure to hypoxia (Fig. 3j). Taken together, HIF-1α, NF-kB or both regulate the expression of 6 genes that have previously been shown to prevent oxidative stress by decreasing ROS levels (Fig. 3k).
MUC1-C drives resistance to ROS via SODs
The spatial transcriptomics results demonstrate that the hypoxic cells located adjacent to the necrotic core have the highest enrichment in hypoxia signaling, which is maintained to some extent in hypoxic cells that migrate into more oxygenated regions of the tumor (post-hypoxic). The results are consistent with the notion that a subset of hypoxia-regulated genes are a part of the HM and remain elevated even when cells move to more well-oxygenated tumor regions or the bloodstream (Fig. 1g). To further investigate this observation, we cultured MDA-MB-231 cells in vitro under 1% O2 for 10 days, followed by 10 days of culture under 20% O2 conditions. RNA sequencing analysis demonstrated that 14 of 50 of the hypoxia-induced genes had an increase in expression upon re-exposure to 20% O2 conditions as compared to baseline (FC > 1.25) (Fig. 4a). Interestingly, MUC1 was the only gene in the 6-gene panel to retain increased expression after reoxygenation in vitro as compared to cells that never experienced hypoxia. To further investigate the spatial patterns of hypoxia-inducible genes, we selected CA9 and MUC1 from our 6-gene signature as markers of acute (CA9) and chronic (MUC1) response, respectively and performed immunofluorescent labeling. While CA9 expression is specific to the peri-necrotic region, MUC1 is found farther into the more oxygenated tumor regions (Fig. 4b, c).
a Heatmap displaying the fold-change expression of the 55-gene signature co-upregulated by both intratumoral and chronic hypoxia (9 genes did not have read counts in the RX group and were excluded; 4 additional genes were added (SNED1, LRRC24, GADD45G, TMEM45A) that were upregulated by acute and intratumoral hypoxia but not by chronic hypoxic conditions) (see Fig. 3f). 1%/20%: cells cultured for 10 days 1% versus 20% O2; RX/BL: cells cultured under 1% O2 for 10 days and then reoxygenated for 10 days (RX) versus cells cultured under 20% O2 (BL). Scale indicates Log2(FC). b Representative fluorescent images of full tumor cross-sections immunolabeled for CA9 or MUC1 c magnified region marked in (b). d Immunoblot assay was performed using lysates prepared from MDA-MB-231 control (NTC), HIF-1α (H1-1/H1-2), or HIF-2α knockout (H2-1/H2-2) cells cultured under 20% or 1% O2 for 5 days to assess the levels of MUC1-C and CA9. e Immunoblot assay was performed using lysates prepared from MDA-MB-231 control or RELA (R-1/R-2) knockout cells cultured under 20% or 1% O2 for 5 days to assess the levels of MUC1-C and SOD2. f Schematic of a suspension assay designed to mimic survival in the bloodstream. Cells are cultured under chronic hypoxia, followed by 4h of culture in liquid suspension with rotation after reoxygenation. g, h Relative levels of MitoROS (g) and Sytox (h) were assessed using flow cytometry in MDA‐MB‐231 NTC or MUC1 (M-1/M-2) knockout cells cultured as described in (f) (mean ± SEM, N = 4 biological repeats versus NTC, matched Friedman test with uncorrected Dunn’s post-test). Representative flow cytometry histograms (right). i, Relative mRNA expression of SOD1, SOD2 and SOD3 as measured by RT-qPCR in MDA-MB-231 NTC or MUC1 (M-1/M-2) knockout cells cultured under 20% or 1% O2 for 5 days (mean ± SEM, N = 3 biological repeats, n = 3 technical repeats except n = 2 for SOD3 set 1; versus NTC 1%, Ordinary one-way ANOVA with Dunnett’s post-test). j Immunoblot assay was performed using lysates prepared from the same MDA‐MB‐231 NTC or MUC1 (M-1/M-2) knockout cells cultured under 20% or 1% O2 for 5 days to assess the levels of the SOD enzymes. Source data are provided as a Source Data file. Cartoons adapted (altered cell colors) under Creative Commons Attribution 4.0 Unported License (https://creativecommons.org/licenses/by/4.0/).
The mechanism behind the hypoxia memory warrants further investigation as HIF-1α is rapidly degraded following reoxygenation to 20% O2. However, MDA-MB-231 cells exposed to 1% O2 for 5 days and then reoxygenated to 5% O2 (the highest oxygen concentration in tumors formed by MDA-MB-231 cells2) retain increased levels of both HIF-1α pNF-kB and MUC1-C as compared to cells cultured under 20% O2 (Supplementary Fig. 4a).
The role of the MUC1-C subunit in regulating antioxidant mechanisms has been well characterized28,29,30 (Supplementary Table 5). Moreover, the MUC1-C inhibitor, GO-203, is currently in clinical trials for acute myeloid leukemia with promising initial results (NCT02204085)31. Thus, we investigated whether MUC1-C is a potential therapeutic target for reducing metastatic spread caused by cells that experienced hypoxic in the primary tumor. MUC1 expression colocalized with GFP + expressing cells in hypoxia fate-mapping MDA-MB-231 tumors (Supplementary Fig. 4b). Furthermore, sorted GFP + tumor cells cultured under normal O2 conditions retained higher levels of MUC1/MUC1-C compared to DsRed + cells, supporting the hypothesis that MUC1 levels remain elevated after reoxygenation (Supplementary Fig. 4c–e). The CRISPR knockout of HIF-1α and NF-kB p65 prevented hypoxia-induced MUC1-C expression in MDA-MB-231 and MDA-MB-436 cell lines (Fig. 4d, e and Supplementary Fig. 5a, b). Treatment with NAC also abrogated MUC1-C induction under chronic hypoxia (Supplementary Fig. 5c, d).
To test the potential antioxidant role of MUC1, we first assessed the levels of MitoROS under chronic hypoxia. We verified that MDA-MB-231 and MDA-MB-436 cells transduced with CRISPR vectors targeting MUC1 had increased ROS accumulation compared to the NTC cells (Supplementary Fig. 5e, f). To simulate conditions in the bloodstream, we harvested cells after chronic exposure to hypoxia and cultured them in suspension under 20% O2 for 4h (CTC estimated circulation time32) (Fig. 4f). MUC1 knockout cells showed higher mitochondrial ROS and increased apoptosis compared to control cells (Fig. 4g, h and Supplementary Fig. 5g–i). We then utilized the same MUC1 knockout MDA-MB-231 cell lines to determine the mRNA expression levels of previously reported MUC1-C-regulated ROS scavengers28,29,30. Interestingly, while the expression levels of glutathione peroxidase 1 (GPX1), glutathione S‑transferase pi (GSTP1) and TP53 Induced Glycolysis Regulatory Phosphatase (TIGAR) were not affected, the levels of catalase (CAT) and superoxide dismutase 1 and 2 (SOD1 and SOD2) were significantly reduced by MUC1 deletion under chronic hypoxia (Supplementary Fig. 5j).
Our follow-up studies focused on the SOD enzymes since they detoxify superoxide anions, the most detrimental form of ROS for a cell33. In a time-course experiment, we observed that all SOD enzymes were significantly upregulated under chronic hypoxia (Supplementary Fig. 5k). Silencing MUC1 via CRISPR decreased the levels of SOD1-3 both at the mRNA and protein levels in MDA-MB-231 and MDA-MB-436 cells (Fig. 4i, j and Supplementary Fig. 5l). Our results suggest that MUC1/MUC1-C (1) is upregulated by both HIF-1α and NF-kB under chronic hypoxia, and (2) is required to control mitochondrial ROS levels in cells under chronic hypoxia or in a simulated bloodstream environment by promoting SOD1-3 expression.
MUC1 is a biomarker for TNBC metastasis
To investigate the potential role of MUC1 as a prognostic marker in cancer, we analyzed microarray data of cancer tissue from patients with breast cancer using the KMPlotter analysis tool to determine whether MUC1 levels were prognostic for distant metastasis-free survival (DMFS). Using two different probes that detect MUC1, we observed that high MUC1 expression was significantly associated with reduced DMFS only in basal breast cancer, with a similar trend in HER2 + breast cancer (Fig. 5a, b). Likewise, MUC1 expression was only correlated with an increase in hypoxia-inducible genes in breast tumors that were molecularly characterized as a basal subtype in the TCGA database, with a slight positive correlation in the HER2+ subtype (Fig. 5c, d and Supplementary Fig. 5m, n). The hypoxic score was evaluated by averaging the expression of genes in two previously reported hypoxia signature gene sets2,34.
a, b Kaplan–Meier analysis (https://kmplot.com) of distant metastasis-free survival (DMFS) of gene chip data from patients with breast cancer across each subtype – Luminal A, Luminal B, HER2 + and Basal – stratified by high or low MUC1 expression detected with probe 213693_s_at (a) or 211695_x_at (b). Log-rank p-value calculated by KMPlot. c, d Correlation analysis between MUC1 expression and hypoxia measured by a signature conserved across 31 breast cancer cell lines exposed to 1% O2 for 24h (Ye et al. 2018, MCR) (c) and a signature conserved between MDA-MB-231 cells exposed to intratumoral hypoxia and in vitro hypoxia (Godet et al. 2019, NC) (d) in breast cancer tumor samples from the TCGA database (https://xenabrowser.net) (r = Pearson’s coefficient; p = one-tailed p-value). e, f Correlation analysis between MUC1 expression and SOD enzyme expression in luminal A, luminal B, and HER2 + (e) and basal-like (f) breast cancer tumor samples from the TCGA database (https://xenabrowser.net) (r = Pearson’s coefficient; p = one-tailed p-value). g Fold change in the expression of MUC1 mRNA measured by qRT-PCR in the cell lines indicated cultured under 1% O2 for 5 days (mean ± SEM, N = 1 biological repeat, n = 3 technical repeats for all except n = 2 for MDA-MB-453 1%; 1% versus 20%, unpaired t test two-tailed when FC > 2). Source data are provided as a Source Data file.
In addition, basal tumors from patients represented in the TCGA database also displayed a positive correlation between MUC1 and the expression of SOD enzymes, which was not observed in other subtypes (Fig. 5e, f). Likewise, MUC1 was more prevalently increased under chronic hypoxia in TNBC cell lines but not in ER + breast cancer cell lines (Fig. 5g). Together, these results support the hypothesis that MUC1 is induced under hypoxia in TNBC and is associated with the expression of SOD enzymes, potentially driving metastatic disease.
MUC1 is required for the survival of GFP + CTCs and metastatic dissemination
Next, we engineered a hypoxia-dependent CRISPR knockout system using a Cre-recombination approach. This system allowed us to selectively and permanently, knock out any gene of interest in cells experiencing hypoxia (Fig. 6a). We targeted NF-kB to validate our system since the levels of NF-kB p65 were the same under 20% or 1% O2 conditions in the non-target control (Cre-NTC) cells. In contrast, NF-kB p65 levels were abrogated in cells expressing a gRNA targeting RELA (Cre-R-1 or R-2) only when they were cultured under hypoxia (Fig. 6b, c). Using the same gRNA targeting RELA, we generated a constitutive, unconditional knockout of the NF-kB p65 (R-1) cell line as a control. NF-kB p65 expression was abolished in the R-1 cell line regardless of the O2 concentration (Fig. 6d). Overall, either 6-days of exposure to hypoxia (0.5% or 1% O2 concentrations), or treatment with Cre-recombinase caused GFP + expression and concomitantly abrogated NF-kB levels resulting in a reduction of MUC1 and SOD2 protein expression compared to the Cre-NTC cell line (Supplementary Fig. 6a).
a Schematic illustrating the design of the hypoxia-inducible MUC1 gene knockout lentiviral vector construct. b–d MDA‐MB‐231 hypoxia-inducible control (NTC) or RELA knockout (Cre-R-1/Cre-R-2) fate-mapping cells were cultured under 20%, 5% or 1% O2 for 6 days to assess the levels of pNF-kB and NF-kB p65 by immunoblot (b, c) or fluorescent imaging (d). e, f MDA‐MB‐231 NTC, Cre-R-1, or Cre-R-2 cells were orthotopically implanted into NSG mice (e) and re-isolated from tumors (f). g NF-kB staining was quantified in DsRed + or GFP + segmented cells (N = 3 mice except N = 2 for Cre-R-1, n = 3 fields of view; GFP versus DsRed, and Cre-NTC versus Cre-R-1 and Cre-R-2, RM two-way ANOVA with Fisher’s LSD post-test) (bottom). h Hypoxia fate-mapping MDA-MB-231 cells transduced with hypoxia-inducible NTC or MUC1 knockout vectors (Cre-M-1/Cre-M-2) were orthotopically implanted into NSG mice. Representative images of MUC1 immunolabeling (h) were quantified (i) (N = 3 mice n = 4 fields of view; versus NTC, Ordinary one-way ANOVA with Dunnett’s post-test). j MDA-MB-231 cells (h) were implanted into NSG mice. Whole blood was collected 35-40 days later to isolate CTCs. k MitoROS levels in DsRed + and GFP + CTCs were assessed using flow cytometry (N = 2 sets: set 1 – Cre-NTC n = 4, Cre-M-1 and Cre-M-2 n=2 mice; set 2 – Cre-NTC and Cre-M-2 n = 4, Cre-M-1 n = 5 mice; RM two-way ANOVA with Fisher’s LSD post-test for comparisons displayed). l MDA-MB-231 cells (h) were implanted into NSG mice, the tumor was resected on day 25, and lung metastasis was assessed on day 35. m The probability of a GFP + (DsRed) lung metastasis forming was determined by dividing the percentage of GFP + (DsRed) cells in the lung by the percentage of GFP + (DsRed) cells in the matched primary tumor by flow cytometry (N = 2 sets: set 1 – Cre-NTC n = 5; Cre-M-1 n = 3; Cre-M-2 n = 2 mice and set 2 – Cre-NTC n = 7; Cre-M-1 n = 5; Cre-M-2 n = 6 mice); GFP versus DsRed, and Cre-NTC versus Cre-M-1 and Cre-M-2; RM two-way ANOVA with Fisher’s LSD test) n Representative fluorescent images of lung tissue sections. Data displayed as mean ± SEM; N=biological replicates; n = technical replicates. For boxplots, boxes are median centered and extend from 25th to 75th percentiles, with whiskers representing the minimum and maximum points. Source data are provided as a Source Data file. Cartoons adapted (altered cell colors) under Creative Commons Attribution 4.0 Unported License (https://creativecommons.org/licenses/by/4.0/).
To test the hypoxia-dependent CRISPR knockout system in vivo, we generated orthotopic tumors from Cre-NTC, Cre-R-1, and Cre-R-2 cells. Immunolabeling of NF-kB in the tumor tissue sections or cells cultured from the primary tumor showed a significant reduction in NF-kB p65 expression in GFP + compared to DsRed + cells in both Cre-R-1/2 cell lines but not in the Cre-NTC cell line (Fig. 6e–g and Supplementary Fig. 6b). The results show that the hypoxia-dependent knockout construct depletes NF-kB expression exclusively in GFP + tumor regions.
Next, we adapted the same strategy to develop hypoxia-specific MUC1 knockout cell lines (Cre-M-1 and Cre-M-2) (Supplementary Fig. 6c, d). Immunolabeling of MUC1 in tissue sections from orthotopic tumors derived from Cre-NTC and Cre-M-1 or -2 lines demonstrated a significant abrogation of MUC1 levels by image analysis (Fig. 6h, i and Supplementary Fig. 6e). To investigate the effect of abrogating MUC1 expression in GFP + CTCs, orthotopic tumors derived from hypoxia-induced MUC1 knockout cells were generated in NSG mice (Fig. 6j). Blood was harvested approximately 4 weeks later, and MitoROS levels were measured in CTCs by flow cytometry. As previously reported, GFP + CTCs showed about 50% lower levels of MitoROS compared to DsRed + CTCs in the control group. Furthermore, the selective knockout of MUC1 in GFP + CTCs increased ROS levels (Fig. 6k and Supplementary Fig. 6f, g).
To investigate the role of increased MUC1 expression in GFP + cells during the course of metastasis, we altered our experimental strategy by removing tumors derived from hypoxia-induced MUC1 knockout cells or NTC (exclusive in GFP + cells) 25 days post-implantation and assessed lung metastasis at day 35. The contribution of GFP + cells to the overall metastatic burden was reduced by 43–50% in mice bearing hypoxia-induced MUC1 knockout cells (Fig. 6l–n and Supplementary Fig. 7a–f). The hypoxia-induced knockout of NF-kB p65 resulted in similar effects on lung metastasis (Supplementary Fig. 7g–m).
Targeting MUC1-C increases ROS in GFP + CTCs and reduces metastasis
To test the therapeutic potential of targeting MUC1-C, we treated MDA-MB-231 and 4T1 cells cultured under chronic hypoxia in the presence or absence of the MUC1 inhibitor (GO-203) in vitro, and we observed reduced levels of MUC1-C and SOD enzymes in treated cells (Supplementary Fig. 8a, b). In vivo, mice bearing orthotopic tumors were treated with 5 consecutive daily doses of GO-203, and whole blood was harvested. MitoROS levels in GFP + CTCs were significantly increased with GO-203 treatment as compared to the control group in both the immunocompromised MDA-MB-231 model as well as the syngeneic 4T1 model (Fig. 7a, b (MDA-MB-231) and Fig. 7c, d (4T1) Supplementary Fig. 8c–e).
a Hypoxia fate-mapping MDA-MB-231 cells were orthotopically implanted into NSG mice. When tumors reached ~ 500 mm3, GO-203 was I.V. administered for 5 days. b Levels of MitoROS in DsRed + and GFP + CTCs assessed by flow cytometry (N=2 sets: set 1 – n = 3 and set 2 – n = 5 mice). c Hypoxia fate-mapping 4T1 cells were orthotopically implanted into Balb/c mice. Fourteen days post-implantation, GO-203 was I.V. administered for 5 days. d Levels of MitoROS in DsRed and GFP CTCs assessed by flow cytometry (N = 1 set: DMSO n = 4 and GO-203 n = 5 mice; the percent reduction in the mean is displayed). e MDA-MB-231 cells were implanted into NSG mice. After 20 days, GO-203 was I.V. administered for 5 days, tumors were removed on day 25, and metastasis was assessed on day 35. f The probability of a GFP + lung metastasis forming was determined by dividing the percentage of GFP + cells in the lung by the percentage of GFP + cells in the matched primary tumor as assessed by flow cytometry (N = 1 set: DMSO n = 6 and GO-203 n = 5 mice). g Representative fluorescent images of lung tissue sections. h GO-203 selectively targets GFP + cells that express high levels of MUC1-C in response to hypoxia and ROS. i PDXs (HCl-001 and HCI-010) were implanted into NSG mice. When the tumor burden reached 1 cm3, CTCs were collected from whole blood. j Representative fluorescent images of MUC1 and CA9 immunolabeling in PDX-010 tumor. k Levels of MitoROS in MUC1low and MUC1high CTCs were assessed using flow cytometry (N = 1 set, HCI-001 - n = 6 or HCI-010 - n = 5; paired one-tailed t test). l–o HIF-1α (l), and MUC1 (n) immunofluorescent labeling were performed on adjacent core biopsy sections from patients with TNBC. Quantification based on digital pathology scores (Sup. Fig. 8k,l) for (m) HIF-1α and (o) MUC1 (N = 8 patients: 3-yr recurrence (N = 4) or no recurrence (N = 4); unpaired one-tailed t test). Data displayed as mean ± SEM; N = biological replicates; n = technical replicates. Statistical analysis for (b, d, f): RM two-way ANOVA with Fisher’s LSD post-test for comparisons displayed. For boxplots, the box is median-centered and extends from the 25th to 75th percentiles, and the whiskers represent the minimum and maximum points). Source data are provided as a Source Data file. Cartoons adapted (altered cell colors) under Creative Commons Attribution 4.0 Unported License (https://creativecommons.org/licenses/by/4.0/).
We also assessed lung metastasis in a neoadjuvant model of breast cancer by treating mice with 5-consecutive doses of GO-203 before tumor removal (Fig. 7e). Ten days after surgery, we assessed the lungs and found a 53% reduction in the contribution of GFP + cells to metastatic burden in treated mice. The reduction of GFP + cells at the metastatic site caused a reduction in metastasis overall (Fig. 7f, g and Supplementary Fig. 8f–i). Taken together, the results show that targeting MUC1-C with GO-203 can reduce metastasis (Fig. 7h).
In further support of this data, we orthotopically implanted two TNBC patient-derived xenograft (PDX) lines (HCI-001 and HCI-010, Welm lab35) into NSG mice (Fig. 7i). Like MDA-MB-231 derived tumors (Fig. 4a), CA9 expression was localized adjacent to the peri-necrotic region, while MUC1 expression was found dispersed in more oxygenated tumor regions (Fig. 7j). When the tumor burden reached 1 cm3, whole blood was collected. After red blood cells were lysed, ICAM1 staining was used to identify CTCs using flow cytometry. A MUC1 conjugated antibody was used to identify high and low MUC1 expressing CTCs. Overall, MUC1low CTCs showed 2x-higher levels of MitoROS as compared to matched MUC1high CTCs (Fig. 7k and Supplementary Fig. 8j).
Next, we investigated whether MUC1 expression in human tumor biopsies correlated with patient outcomes. We performed immunofluorescent staining of both MUC1 and HIF-1α on adjacent FFPE slides of eight core tumor biopsies collected from patients with TNBC at the time of diagnosis. Using digital pathology scores calculated from the % of tissue area with a fluorescent signal above an assigned threshold of MUC1 or HIF-1α, we determined that expression of both markers was significantly higher in patients who experienced recurrence 3 years after their initial diagnosis (Fig. 7l–o and Supplementary Fig. 8k–n). These studies should be expanded to further evaluate the prognostic potential of MUC1 as a biomarker. Unfortunately, our current observations are limited to small sample size (N = 8) and limited follow-up (3 years).
Discussion
It is well-accepted that hypoxia alters cancer cell signaling and phenotype. Our previous studies and others have demonstrated that HIF-induced gene products promote metastasis36,37,38,39, but HIF-1/2α levels quickly return to baseline upon reoxygenation (20% O2) in vitro. Therefore, the transcriptional changes that occur in response to hypoxia have previously been considered reversible. It remained unknown whether HIF-regulated gene products were still expressed and functional even after reoxygenation in vivo. Using multiplex imaging combined with spatial transcriptomics, we demonstrate that the response to hypoxia is durable for a subset of genes that we term the HM even after the cells are reoxygenated to the physiologic O2 concentrations of breast tissue. Many laboratories characterize the hypoxic effect by exposing cancer cells to 1% O2 for 24h to stabilize HIF-1/2α. However, the hypoxic tumor microenvironment is a dynamic niche where multiple stresses co-occur, causing transcriptional plasticity. Our studies demonstrate that exposure to chronic hypoxia in vitro more closely mimics cellular responses in vivo, perhaps because it causes other co-occurring stress factors that happen in tumors, such as ROS accumulation, nutrient starvation, acidosis, and metabolic reprogramming or that longer exposure time is necessary to cause the memory effect. It is not possible to decouple the individual and synergistic effects of the tumor microenvironment; thus, the complete biological mechanism behind the HM gene set enrichment remains elusive. We hypothesize that it is likely dependent on epigenetic changes that affect some but not all hypoxia-inducible genes. It is also possible that the selective pressure to survive in the hostile peri-necrotic tumor regions could cause a selection for more progenitor-like cells (cancer stem cells) with increased plasticity. Our study did highlight that unlike reoxygenation to supraphysiological levels (20% O2), cells that are reoxygenated to physiological O2 levels (i.e., 5% O2) in vitro retain increased HIF-1/2α and pNF-kB levels, and the expression of downstream effectors such as MUC1/MUC1-C.
The results of our study demonstrate that cells that experience intratumoral hypoxia acquire a ROS-resistance mechanism at least initially driven by HIF-1α and NF-kB regulation. This is particularly useful for cancer cells that enter the bloodstream and face hyperoxia, as well as chemical, biological, and physical cues that cause a surge in ROS. Ros-resistence allows cells to overcome this stress and confer a survival advantage. The effects of transient/cycling hypoxia within the primary tumor, as well as while cancer cells pass through the lung capillaries, remain to be investigated.
Acute hypoxia causes stabilization of HIF-1α and accumulation of mitochondrial ROS (superoxide, O2−), which is converted into H2O2 after longer-term exposure to hypoxia, causing activation of NF-kB p65. HIF and NF-kB act in a concerted manner to drive an antioxidant response. Still, additional studies are warranted to investigate the crosstalk between these two signaling cascades. Moreover, it is likely that other antioxidant mechanisms might be at play to control ROS levels in the cell, such as the nuclear factor E2-related factor 2 (NRF2), which is a transcription factor that binds to the antioxidant response element (ARE) under oxidative stress40. In this work, we identified 16 genes within the HM with previously reported roles in antioxidant defense. Furthermore, we demonstrated that at least one of these HM genes (MUC1/MUC1-C) plays a role in the regulation of SOD enzymes, contributing to decreasing oxidative stress and promoting the metastasis of GFP + cells. On the other hand, the hypoxia-dependent knockout of NF-kB showed a more striking reduction of GFP + cells at the site of metastasis, suggesting that MUC1-C alone is not capable of driving the metastasis of hypoxia-exposed cells (GFP +).
MUC1 expression correlated with hypoxic signatures in TNBC tumors and MUC1 upregulation is more striking following chronic hypoxia in TNBC cell lines as compared to other subtypes. The mechanisms preventing MUC1-C upregulation in non-TNBC, as well as the extent to which this occurs, remain to be explored. However, it is important to note that there is a high degree of heterogeneity in response to hypoxia across cell lines and cancer subtypes. For example, we previously reported that only 42 genes had a conserved induction in expression across 31 breast cancer cell lines that were exposed to 20% or 1% O2 in comparison to over 1000 genes that were differentially expressed in any one cell line34. It is possible that the NF-kB and/or HIF-1/2 binding elements in the MUC1 promoter could be selectively methylated in ER-positive cell lines, preventing DNA binding and an increase in mRNA expression. We have previously shown that the methylation of the HRE-binding region in the EGFR gene only occurred in TNBC cell lines and prevented HIF-1/HRE binding under hypoxia41.
The finding of an HM has several implications for cancer treatment. For example, emerging therapeutic strategies aimed at targeting hypoxic cancer cells in the primary tumor may not kill cancer cells that escape the hypoxic microenvironment. We have demonstrated that GFP + cells that escape the primary tumor are chemoresistant and drive recurrence in mouse models42. In addition, systemic targeting of hypoxic cancer cells is challenging due to poor vascularization in these regions. Moreover, in many cases, the treatment plan for patients with solid tumors includes surgery to remove the primary tumor, thus removing hypoxic cancer cells, followed by treatment to prevent a recurrence. Our results suggest that cancer cells that experience low oxygen tensions and escape the primary tumor have a unique transcriptional profile due to their preconditioning upon exposure to hypoxia. Therefore, the HM provides a list of potential targets that can be used to design therapies that will be beneficial in both the neoadjuvant and adjuvant settings to eradicate cancer cells in the primary tumor, as CTCs and as disseminated cells that may remain after surgery. An important caveat to note is that the current study used a mix of in vitro studies and in vivo studies to define our genesets of interest, most importantly the reoxygenation condition. In the future, it will be important to design a system whereby one can identify cells undergoing short-term and long-term hypoxia from those that have been reoxygenated in vivo. For example, in our in vitro set-up, cells were exposed to 20% O2 as a baseline whereas in the tumor “normal” cells are exposed to lower O2 gradients (~5%) and are reoxygenated to different levels depending on the metastatic site.
In this study, we identified a subset of genes that we termed HM genes. Future work, including more sophisti`cated models will be needed to identify a complete and robust list of HM genes. We used GO-203 to pharmacologically target MUC1-C, a HM gene. It will be important to test the combination of GO-203 with standard-of-care agents to maximize therapeutic success. Targeting HM genes may provide an alternative treatment approach for patients with TNBC who typically have limited treatment options and are more likely to experience recurrance40,41.
Methods
Cell culture
Mycoplasma-free breast cancer cell lines, MDA-MB-231 (ATCC® HTB-26™), MDA-MB-436 (ATCC® HTB-130™), and 4T1 (ATCC® CRL-2539™), were obtained from the American Type Culture Collection (ATCC) and maintained in DMEM (MDA-MB-231 and MDA-MB-436) or RPMI (4T1) (Sigma-Aldrich or Corning) with 10% FBS (Corning) and 1% penicillin/streptomycin (Invitrogen). Hypoxic conditions were achieved using an InvivO2 workstation (Baker) with an ICONIC (Baker) electronically controlled gas mixing system maintained at 37 °C and 75% humidity, equilibrated at 0.5% or 1% O2, 5% CO2, and 94.5% or 94% N2. Since cancer cells cultured under 20% O2 are significantly more proliferative than cells cultured under chronic hypoxia, cell density was optimized to obtain equivalent confluences between dishes by plating 10 times more cells in hypoxic dishes.
Constructs and lentiviral delivery
Our hypoxia fate-mapping dual-vector system has been well described previously2. The methodology to utilize this system has also been described43. Briefly, vectors encoding CMV-loxp-DsRed-loxp-eGFP (#141148, Addgene) or vector encoding 4xHRE-MinTK-CRE-ODD (#141147, Addgene) were co-transfected with plasmid psPAX2 (#12260, Addgene) and plasmid pMD2.G (#12259, Addgene) into 293T cells using Polyjet (SL10088, Signagen). Filtered viral supernatant generated from lentiviral vector 1 was collected 48 h post-transfection and added to MDA-MB-231 or 4T1 cells with 8 μg/mL polybrene (Sigma–Aldrich) overnight. After 24 h in fresh media, zeocin (Invitrogen) was added to the medium of cells for selection (100–200 μg/mL). Following selection, cells were transduced with lentivirus from lentiviral vector 2 encoding 4xHRE-MinTK-CRE-ODD. The cell lines were sorted, single-cell cloned, and screened by image analysis and flow cytometry. To evaluate NF-kB activation, the construct NFKBRp_mKate2_2xNLS_p2a-puroR, a gift from Timothy Lu (#82024, Addgene; RRID:Addgene_82024), was transduced into MDA-MB-231 and MDA-MB-436 cells. Bacteria were grown in the presence of ampicillin overnight. Cultured bacteria were processed using the MaxiPrep kit (Sigma) following the manufacturer’s instructions to purify plasmid DNA, which was then co-transfected as already described. More details on the validation of this system have been previously demonstrated2. In the context of the current work, we confirmed that H2O2 treatment did not activate the Lox-Cre mechanism by both fluorescent microscopy and Cre mRNA expression analysis.
Animal studies
Female 5- to 7-week-old NOD-SCID Gamma (NSG) and BALB/C (Jackson Labs) mice were used according to protocols approved by the Johns Hopkins University Animal Care and Use Committee (JHU ACUC). Animal rooms were maintained at 30–70% relative humidity and a temperature of 18–26 °C with a minimum of 10 room air changes per hour. Cages were changed one time a week. Mice were fed a diet containing low fiber (5%), protein (20%) and fat (5–10%). Our findings are in relation to breast cancer, and because it affects significantly more women than men, we utilized female mice. The maximum allowable tumor size for a single spontaneous or implanted tumor that is visible without imaging is 2 cm in any dimension in adult mice. Our studies complied with this limit. Mice were anesthetized by the intraperitoneal injection (i.p.) of 100 mg/kg Ketamine 16 mg/kg Xylazine (Vet One).
To harvest sufficient numbers of CTCs, 2 x 106 MDA-MB-231 or 4T1 hypoxia fate-mapping cells, with or without the NF-kB reporter, were injected into the mammary fat pad (MFP) closest to the second nipple. Animals were sacrificed to harvest primary tumors, whole blood, and/or lungs 35–40 or 18 days post-implantation for MDA-MB-231 or 4T1 tumors, respectively. Volumes of blood ranging from 500–1000 μl were collected by cardiac puncture into an EDTA tube using a 26G syringe needle. Red blood cells were lysed using cold ACK Lysing Buffer (Quality Biological) and the remaining cells were washed, stained with MitoROS as described when indicated, and resuspended in FACS buffer.
To assess metastasis, tumor surgery was performed 25 days post-implantation of MDA-MB-231 cells. Mice were anesthetized and transferred to a heating pad. After hair removal and cleaning of the surgical area, an incision was made. The tumor was carefully detached from adjacent skin, and visible lymph nodes were removed. The wound was closed by using 9-mm autoclips (Braintree Scientific, Inc.) and cleaned with Betadine. Ophthalmic ointment was used to avoid eye desiccation. Ten days later, mice were sacrificed, and lungs were inflated with OCT (Fisher Scientific) and excised for both image and flow cytometry analysis. Half of the tumor and lung tissue was chopped and digested (collagenase 2 mg/mL (Sigma-Aldrich)) for 45 min to 1 h at 37 °C at 160 RPM. After passing through a 0.70 μm strainer, cells were washed with PBS and resuspended in FACS buffer. The other half was formalin-fixed, saturated in 30% sucrose, frozen in OCT, cryosectioned, mounted on charged super-frost slides, and imaged using Cytation5 (BioTek Instruments). This experimental set-up ends 10 days post-surgery to avoid the overgrowth of lung metastatic nodules that can potentially get hypoxic and develop GFP + cores. Mice with tumor re-growth were excluded from the study to avoid confounding results derived from recently hypoxic GFP + cells.
The TNBC PDX lines (HCI-001 and HCI-010)35 were provided by the Zahnow lab (Johns Hopkins) via the original creators, Welm lab (University of Utah). Tumor fragments were implanted as 2-3 mm chunks placed under the MFP closet to the 4th nipple via a small incision or as an organoid suspension injected directly into the MFP closest to the 2nd nipple.
Integrated spatial profiling with Visium spatial transcriptomics and fluorescent hypoxia reporter imaging
The Visium spatial protocol (10X Genomics) was optimized to preserve the native fluorescence of the tumor tissue. Fixation and imaging times were optimized to preserve RNA integrity while acquiring acceptable fluorescent intensity using charged superfrost slides (Fisherbrand). Tissue permeabilization time was optimized to 3 min by utilizing the Visium Spatial Tissue Optimization Reagents Kits (#PN-1000193, 10X Genomics). To ultimately perform spatial transcriptomics, hypoxia fate-mapping MDA-MB-231 tumors were harvested 18 days post-implantation and immediately frozen in clear OCT (Fisherbrand) using 0.7 x 0.7 x 0.5 cm cryomolds (Fisherbrand) (N = 2). Using a cryotome (Leica), 10 μm tumor slices were placed within the fiducial frame of the Visium slide. The tissue was pre-fixed in cold methanol 99.9% (#34860, Sigma) for 15 min and dried at 37 °C for 30 sec. The slide was then imaged at 10X magnification using a Cytation5 (BioTek Instruments) in the RFP and GFP channels to acquire native fluorescence. The slide was immediately transferred to cold methanol 99.9% (#34860, Sigma) with DAPI at 1:400 dilution for 15 min. DAPI staining was imaged at 10X using Cytation5 after drying the slide. The Visium spatial gene expression protocol was then followed according to the manufacturer’s instructions for the Visium Spatial Gene Expression Kits (#PN-1000187, 10X Genomics). Briefly, tissue permeabilization for 3 min was followed by cDNA synthesis, second strand synthesis, and cDNA denaturation. The ideal cycle number was determined by qPCR, and cDNA was amplified and cleaned using SPRIselect (#B23317, Beckman Coulter). The amount of cDNA was measured to determine the cDNA input for library construction. Briefly, cDNA was fragmented and submitted to end repair and A-tailing. Adapter sequences were ligated, and the post-ligation product was cleaned and subjected to Index PCR. Overall, the P5, P7, i7, and i5 sample indexes, and TruSeq Read 2 were added. After a library QC test, samples were sequenced using Illumina
NovaSeq6000 with a sequence depth of > 100 k (tumor 1) or > 34 k (tumor 2) reads per spot for a total of 276 M (tumor 1) and 100 M (tumor 2) reads per sample. Sequencing data was preprocessed using the Space Ranger v1.3.0 software (10x Genomics) for demultiplexing and FASTQ conversion of barcodes and reads data, alignment of barcodes to the stained tissue image, and generation of read counts matrices. The processed sequencing data were inputs for the analyses using the Seurat pipeline v4.3.018. Data preprocessing with Seurat involved initial visualization of the counts onto the tissue image to discriminate technical variance from histological variance (e.g., collagen-enriched regions present lower cellularity that reflects in low counts), removal of necrotic regions as determined by percentage of mitochondrial reads (cutoff = 20%), and normalization with SCTransform44. Following filtering, dimensionality was reduced with PCA and then clustered with Leiden. Hypoxic status to cluster assignments was performed based on the most variable features (genes) and gene set enrichment analysis (GSEA). Differential expression analysis of tumor spots from hypoxia versus non-hypoxia regions was performed using the MAST test45. The gene set enrichment analysis was performed with fGSEA46 using the genes ranked according to the log-fold change (LFC) determined by the differential expression analysis. We evaluated the enriched expression of genes belonging to the MSigDB Hallmark Pathways47. Scripts and full datasets are available at github.com/FertigLab/hypoxiaROSanalysis with DIO reference 10.5281/zenodo.12640998
RNA sequencing and analysis of CTCs
Whole blood was harvested from NSG female mice bearing MDA-MB-231 hypoxia fate-mapping orthotopic tumors. After ACK lysis, 7000 to 25,000 CTCs were DsRed + or GFP + sorted directly into TRI-reagent (Zymo). RNA was extracted using the Direct-zol microRNA kit (Zymo), and samples with RIN>8 were submitted for low-input RNA sequencing (N = 2). Library preparation was conducted using an Ovation RNA-Seq System V2 kit, and samples were sequenced in a HiSeqX, 150 x 150 paired-end. Illumina's CASAVA 1.8.4 was used to convert BCL files to FASTQ files using default parameters. Alignment to hg38 reference genome and gene and transcript expression levels were ran using rsem-1.3.048. Normalized transcript per million data was utilized to perform pathway analysis with GSEA as described below.
RNA sequencing and analysis of hypoxic and reoxygenated cells
To compare acute versus chronic hypoxic conditions, MDA-MB-231 cells were cultured for 1 or 10 days under 20% or 1% O2. To investigate the effect of reoxygenation, cells cultured for 10 days under 1% O2 were reoxygenated (20% O2) for 10 days. RNA was extracted using the Direct-zol miniRNA kit (Zymo), and samples with RIN> 9 were submitted for library preparation, RNA sequencing, and preliminary analysis (N = 1). The reads were mapped to hg38 using the latest UCSC transcript set using Bowtie2 version 2.1.049, and the gene expression level was estimated using RSEM v1.2.1548. TMM (Trimmed Mean of M-values) was used to normalize gene expression. Differentially expressed genes were identified using the edgeR program50. Gene expression was compared within each data set. The Log2 fold change in mRNA expression in MDA-MB-231 cells exposed to 1% O2 compared to MDA-MB-231 cultured under 20% O2 for 1 or 10 days was determined. Genes with fold changes lower than -2 or greater than 2 were considered differentially expressed. Normalized transcript per million data was utilized to perform pathway analysis. The reoxygenation effect considered the Log2 fold change in mRNA expression in MDA-MB-231 cells exposed to 10 days under 1% O2 followed by 10 days under 20% compared to MDA-MB-231 cultured under 20% O2 for 1 day, which we termed ‘baseline’.
Pathway analysis
Gene Set Enrichment Analysis (GSEA)51 was conducted to determine the specific ‘Hallmark’ gene sets from The Molecular Signatures Database (MSigDB) collection v747 that showed statistically significance and concordant differences between GFP and DsRed in tumor and lung, as well as CTCs samples, using the class analysis tool in the GSEA desktop application v4.0.3. Briefly, GSEA is a computational methodology designed to determine if a defined set of genes has concordant differences between two biological conditions by estimating the significance of the over-representation of an independently defined set of genes in the highly correlated or anti-correlated genes in the ranked list between the two conditions. This tool was employed to investigate pathways in the gene expression clusters identified using spatial transcriptomics. Gene expression sets – Acute, Chronic, and Intratumoral hypoxic conditions - were compared using the VennPlex v1.0.0.2. tool52 to determine the number of genes that overlap between the different conditions (− 1.5 ≥ FC ≥ 1.5, p-value≤ 0.05). Gene Set Enrichment Analysis (GSEA)51 was also conducted to compare the level of enrichment in ‘Hallmark’ gene sets in acute and chronic hypoxic conditions by utilizing the pre-ranked analysis tool in the GSEA desktop application. This computational method analyzes gene expression data by taking a list of genes ranked by their expression differences across experimental conditions and assessing whether predefined gene sets, representing biological pathways, are enriched at the top or bottom of the list. GSEA calculates an Enrichment Score (ES) and determines its statistical significance through permutation testing. The ES for each gene set is normalized to accommodate for variations in set sizes, resulting in a normalized enrichment score (NES).
Analysis of third-party patient datasets
The distant-free metastasis (DMFS) analysis of patients with breast cancer was performed using the online tool KM Plotter (https://kmplot.com/analysis) after selecting the mRNA gene chip data for breast cancer53. The median cut-off of MUC1 expression using either probe 213693 (best jet-set probe) or 211695 (not the best for genes when using the A array) was used to stratify patients with high (above median) and low (below median) expression. The analysis was restricted to subtypes by selecting the PAM50 subtype of interest (Luminal A, Luminal B, HER2, basal). The correlation analysis between hypoxia signatures2,34 and MUC1 or SOD1-3 expression was performed using the UCSC Xena Browser54 (https://xenabrowser.net/heatmap/). After selecting breast cancer TCGA gene expression RNAseq – IlluminaHiSeq dataset (TCGA.BRCA.sampleMap/HiSeqV2), gene expression data was downloaded in Log2(norm_count + 1) format, and “PAM50_mRNA_nature2012” was used for phenotypic data. The mean of the expression of each gene in the hypoxia signature was compared to the MUC1 expression for each PAM50 subtype (Fig. 5c, d). SOD1, SOD2, and SOD3 expression were correlated with MUC1 expression in the same dataset.
Immunoblot assays
For protein extraction, 100,000 or 1,000,000 cancer cells were seeded overnight in 10 cm plates and exposed to 20% or 1% O2, respectively, for 5 days. For acute hypoxia exposure, 1,000,000 cancer cells were seeded in 10 cm plates overnight and cultured under 20% or 1% O2 for 48 h. Aliquots of whole cell lysates were prepared in NP-40 buffer (150 mM NaCl, 1% NP-40, 50 mM Tris-HCl, pH 8.0) and fractionated by 7.5% or 10% Sodium Dodecyl Sulfate–Polyacrylamide Gel Electrophoresis (SDS-PAGE). Proteins were transferred from the SDS gel to a nitrocellulose membrane for 30 min using a Trans-blot Turbo (Bio-Rad). The nitrocellulose membrane was blocked in 5% milk (% w/v) in Tris-buffered saline and 0.1% Tween-20 (TBS-T) for 30 min and incubated overnight with primary antibodies against HIF-1α (#610958, BD Biosciences), HIF-2α (#sc-46691, Santa Cruz), p-NF-kBp65 (#3033, Cell Signaling), NF-kBp65 (#8242, Cell Signaling or # A2547, Abclonal), p-IKBα (#2859, Cell Signaling), IKBα (#4814, Cell Signaling), COX-2 (#12282, Cell Signaling), CA9 (#sc-365900, Santa Cruz), Fibronectin (ab2413, Abcam), RAPGEF4/EPAC2 (#A4484, Abclonal) MUC1-C (#16564, Cell Signaling), MUC1 (#A19081, Abclonal) SOD1 (#A12537, Abclonal), SOD2 (#13141, Cell Signaling and #A19576, Abclonal) and SOD3 (#A6984, Abclonal). After three 10-min washes in TBS-T, the nitrocellulose membrane was incubated with the corresponding anti-mouse (#7076S, Cell Signaling) or anti-rabbit (#7074S, Cell Signaling) HRP-secondary antibody for 4 h at room temperature with orbital shaking. β-actin was used as loading control by utilizing the β-actin HRP-conjugated antibody (#HRP-60008, Proteintech). ECL (Enhanced Chemiluminescent Substrate) (Perkin Elmer or Prometheus Protein Biology Products) was used as the HRP-catalyzed detection substrate. The chemiluminescent signal was imaged using a c300 imager (Azure Biosystems). For each immunoblot experiment, an identical amount of sample was run on parallel gels. The membranes were cut according to the molecular weights of the protein of interest and incubated with the appropriate antibody. In most instances, actin was detected after stripping and reprobing the membrane.
Inhibitors
To inhibit ROS and further extract RNA or protein from cells, cells were plated and incubated overnight. On the following day, a volume of N-Acetyl-L-cysteine (NAC) (Sigma) or ultrapure water as vehicle was added to make a final NAC concentration of 3 mM, and cells were exposed to 20% or 1% O2 for 5 days. NAC or ultrapure water were refreshed on days 2 and 4. To test the effect of NF-kB inhibition, we treated cells that were exposed to 20% or 1% O2 for 5 days by adding a volume of BAY11-7082 (Sigma) or DMSO as a vehicle to reach the final BAY11-7082 concentration of 10 μM during the last 24 h of the experiment. Similarly, to test the effect of MUC1-C inhibition, we treated cells that were exposed to 20% or 1% O2 for 5 days with a volume of GO-203 (Selleckchem) or DMSO as vehicle during the last 24 h of the experiment to reach a final GO-203 concentration of 2.5 μM.
Flow cytometry analysis
For experiments in vitro, cells were trypsinized, fixed with 4% PFA (Fisher Scientific) for 10 min, washed with PBS, and resuspended in FACS buffer (PBS, 1% BSA, 0.5 mM EDTA, and 25 μg/ml DNAse). Fluorescence intensity was measured by using a CytoFLEX flow cytometer (Beckman Coulter). GFP was detected in the FITC channel, DsRed was detected in the PE channel, and mKate2 was detected in the APC channel. Data were analyzed via FlowJo V10 software (Tree Star, Inc.). Mitochondrial ROS levels were measured in the APC channel by staining with Elite Mitochondrial ROS Activity Kit—Deep Red Fluorescence (MitoROS) (e-Enzyme) following the manufacturer’s instructions. Cell viability was assessed by utilizing Sytox (Invitrogen) according to the manufacturer’s guidelines and using the FITC channel for detection in non-fluorescently labeled cells.
For in vivo experiments using the hypoxia fate-mapping system, cells were resuspended in FACS buffer (PBS, 1% BSA, 0.5 mM EDTA, and 25 mM HEPES, pH 8) after tissue processing. Samples were analyzed by using a SH800 cytometer (Sony). GFP was detected in the FITC channel, and DsRed was detected in the PE channel. The NF-kB.mKate2 reporter or MitoROS was detected in the APC channel, and Sytox was measured in the Pacific-Blue channel. Samples were also sorted by using a SH800 cytometer (Sony) cell sorter cytometer into (GFP+/DsRed−) or DsRed + expressing populations directly into media. DsRed and GFP-only cell lines were used as controls for compensations.
For in vivo experiments using PDX models, after washing, lysed blood extracts were blocked for 10 min on ice. Immunolabeling with conjugated antibodies anti-ICAM1-FITC (CD54, # 353107, Biolegend) and anti-MUC1-PE (CD227, Cat# 355603, Biolegend) was performed in MitoROS (APC) staining buffer on ice for 30 min. A CytoFLEX flow cytometer (Beckman Coulter) was utilized for data acquisition. Unlabeled blood extracts were used as controls.
Immunofluorescent staining
Tissue cryo-sections were subjected to immunolabeling after permeabilization with 1% Triton-X (Sigma-Aldrich) for 5 min and blocking with 2% BSA for 30 min at room temperature (RT). Slides were then incubated overnight at 4 °C with primary antibodies at dilution 1:100 against p-NF-kBp65 (#3033, Cell Signaling), NF-kBp65 (#8242, Cell Signaling or # A2547, Abclonal), MUC1 (#A19081, Abclonal), CA9 (#sc-365900, Santa Cruz), and Anti-8-oxoG DNA Lesion Antibody (#sc-130914, Santa Cruz). For intranuclear staining, the permeabilization step was performed for 10 min. All slides were then incubated with the anti-mouse (#A21237) or anti-rabbit (#A21245) secondary Alexa Fluor 647™ antibodies (Invitrogen) at 1:1000 dilution for 1.5 h at RT. DAPI staining at dilution 1:1000 was performed for 15 min at RT. Finally, slides were mounted with anti-fade solution (90% glycerol, 20 mM Tris, pH 8.0, and 0.5% N-prolyl gallate) and imaged using a Cytation 5 (BioTek Instruments).
Reverse transcription (RT) and quantitative real-time PCR (qPCR)
For RNA extraction, 50,000 or 500,000 cancer cells were seeded overnight in 6 cm plates and cultured under 20% or 1% O2, respectively, for 5 days. For acute hypoxia exposure, 500,000 cancer cells were seeded overnight in 6 cm plates and cultured under 20% or 1% O2 for 24 h. Total RNA was extracted from cells using a Direct-zol miniRNA kit (Zymo) with DNase I treatment. 1000 ng of RNA was used for first-strand DNA synthesis with the GoScript™ Reverse Transcriptase cDNA Synthesis kit (#A2791, Promega). qPCR was performed using specific human primers and iTaq Universal SYBR Green Fast qPCR mix (#RK21203, Abclonal). The expression of each target mRNA relative to 18S RNA was calculated based on the threshold cycle (Ct) as 2−Δ(ΔCt), where ΔCt = Cttarget − Ct18S and Δ(ΔCt) = ΔCttest − ΔCtcontrol. Primer sequences are available in Supplementary Table 6.
Knockout by CRISPR/Cas9
HIF-1α, HIF-2α, NF-kB, and MUC1 knockout by CRISPR/Cas9 were performed as previously described37. Insert oligonucleotides that include a guide RNA sequence were designed and annealed. The duplex oligonucleotides were inserted into the BsmBI cloning site. After confirmation by Sanger sequencing, plasmids were transformed into stbl3 cells. Cultured bacteria were processed using the MaxiPrep kit (Sigma) following the manufacturer’s instructions to purify plasmid DNA, which was then co-transfected with plasmid psPAX2 (#12260, Addgene) and plasmid pMD2.G (#12259, Addgene) into 293T cells using Polyjet (SL10088, Signagen). Filtered viral supernatant was added to the MDA-MB-231 or MDA-MB0436 cells with 8 μg/mL polybrene (Sigma–Aldrich) overnight. After 24 h in fresh media, puromycin (Corning) was added to the medium of cells for selection (100 μg/mL). Sequences of guide RNA oligonucleotides are available in Supplementary Table 7.
Hypoxia-inducible CRISPR construct design
The loxp-3xSV40-loxp sequence was PCR amplified from the plasmid pCAG-loxPSTOPloxP-ZsGreen (#51269, Addgene) by using primers designed to contain sticky-end sequences for XbaI and AgeI restriction enzyme sites. The amplified PCR fragment was agarose-gel purified. The lentiCRISPR_v2 vector was cut with XbaI and AgeI and gel purified. The In-Fusion Snap Assembly kit (#638945, Takara) was utilized to ligate the fragmented DNA into the lentiCRISPR_v2 vector following the manufacturer’s instructions. The vector loxp-3xSV40-loxp-lentiCRISPR was then utilized to introduce guide RNA sequences of interest as described in the previous section. To test this system under hypoxia, cells were cultured for 6 days to allow for the chronic hypoxic downstream regulation to occur (5 days), but with one extra day to activate the system.
In vitro assays to assess ROS levels
To assess ROS levels under hypoxia, 5000 or 50,000 cancer cells were seeded overnight in 24-well plates and cultured under 20% or 1% O2, respectively, for 5 days. Cells were then trypsinized, washed with PBS, and incubated with Elite™ Mitochondrial ROS (MitoROS) Activity Assay Kit (Deep Red Fluorescence) (e-Enzyme) or CellROX™ Deep Red (Thermo Fisher Scientific) following manufacturer’s instructions. Secreted hydrogen peroxide levels were determined using the OxiRed™ probe, a peroxidase fluorogenic substrate (Abcam), following the manufacturer’s instructions. To mimic the bloodstream, 250,000 cancer cells were seeded overnight, followed by a culture under 1% O2 for 5 days. Cells were then resuspended in 5 ml of fresh media in a 15 ml conical tube. Cells were kept in suspension culture with rotation for 4 h under normal O2 conditions to recapitulate the estimated circulation time of a CTC. After 4 h, cells were pelleted, washed in PBS, and stained with MitoROS as described. Cell viability was measured by adding Sytox (Invitrogen). Both MitoROS and Sytox levels were measured in the APC and FITC channels, respectively, using a CytoFLEX flow cytometer (Beckman Coulter).
GO-203 treatment
Fourteen or 20 days post-implantation of 4T1 or MDA-MB-231 cells, respectively, animals were subjected to 5 consecutive doses of GO-203 treatment. GO-203 (4 mg/kg) or DMSO was delivered intravenously in normal saline. Briefly, mice were warmed for 5 min with an overhead heat lamp to dilate the tail veins. Friction was also applied to the tail. Animals were restrained, and 300 μl of drug solution was delivered into the tail vein.
Human study
Core biopsy tissue collected from patients diagnosed with triple-negative breast cancer (TNBC) was utilized according to the availability of tissue in the Johns Hopkins Pathology Archives. Tissue samples belonged to a single institution retrospective cohort study, including patients who had received care at the Johns Hopkins Medical Institute (JHMI). Donors provided written informed consent for providing samples. This was approved by the Johns Hopkins IRB.
Fluorescent immunohistochemistry staining
Immunofluorescent staining of FFPE sections of biopsy cores was performed as previously described55. Briefly, tissue sections were subjected to deparaffinization with xylenes after pre-warming the slides at 60 °C for 20 min. After serially hydrating the tissues, antigen retrieval was performed using citrate buffer (pH 6.2). Peroxidase activity was quenched by incubating for 15 min with 3% H2O2 prepared in methanol, followed by PBS-T washes. The samples were blocked for 1 hr with blocking buffer (0.05% Tween, 1% Casein, 2% BSA, and 5% Goat serum, in PBS). Primary antibody against HIF-1α (#ab179483, Abcam) or MUC1 (#A19081, Abclonal) was added to adjacent tissue sections at 1:1000 dilution, followed by incubation at 4 °C overnight. On the following day, all slides were washed in PBS-T buffer, followed by a wash in 1% Casein-PBS, before incubating for 1 hr with HRP goat anti-rabbit (#20403, Biotium) secondary antibody at 1:200 dilution prepared in blocking buffer. The slides were then subjected to signal amplification using a tyramide amplification kit (#33018, Biotium). DAPI staining at dilution 1:1000 was performed for 15 min at RT. Finally, slides were mounted with the anti-fade solution and imaged using a Cytation 5 (BioTek Instruments).
Statistical analysis
All data is presented as Mean ± SEM, and statistical analysis was performed using GraphPad Prism 10 with statistical tests appropriate for each experimental set-up as indicated in the figure captions. Biological replicates for in vitro assays are listed as N and for in vivo analysis using mice, as set. Technical repeats are listed as n. Mice that died during surgery had tumor regrowth, or needed to be euthanized before the endpoint of the experiment were not considered when assessing metastatic burden. All statistical analysis was performed using the total number of samples analyzed, n. Repeated measures analysis was used when the number of GFP versus DsRed were analyzed from the same tumor sample.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The RNAseq datasets generated and/or analyzed during the current study were deposited into the Gene Expression Omnibus (GEO) database under the following accession numbers: GSE126609 (tumor), GSE136372 (lung), GSE240127 (CTCs), and GSE240211 (acute vs chronic hypoxia in vitro experiment). The spatial transcriptomics datasets generated and analyzed during this study were also deposited into the GEO database under the accession number: GSE240212. The survival data utilized in this study is available in third-party public databases: KMplot (https://kmplot.com) and XENA Browser (https://xenabrowser.net).
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE126609
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE136372
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE240127
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE240211
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE240212. Source data are provided with this paper.
Code availability
Code scripts and full datasets for the spatial transcriptomics are available at github.com/FertigLab/hypoxiaROSanalysis with DIO reference https://doi.org/10.5281/zenodo.1264099856.
References
Vaupel, P., Höckel, M. & Mayer, A. Detection and characterization of tumor hypoxia using pO2 histography. Antioxid. Redox Signal 9, 1221–1236 (2007).
Godet, I. et al. Fate-mapping post-hypoxic tumor cells reveals a ROS-resistant phenotype that promotes metastasis. Nat. Commun. 10, 4862 (2019).
Kallergi, G. et al. Hypoxia-inducible factor-1α and vascular endothelial growth factor expression in circulating tumor cells of breast cancer patients. Breast Cancer Res. 11, R84 (2009).
Hecht, F. et al. The role of oxidative stress on breast cancer development and therapy. Tumor Biol. 37, 4281–4291 (2016).
Marshall, C., Mamary, A. J., Verhoeven, A. J. & Marshall, B. E. Pulmonary artery NADPH-oxidase Is activated in hypoxic pulmonary vasoconstriction. Am. J. Respir. Cell Mol. Biol. 15, 633–644 (1996).
Turrens, J. F. Mitochondrial formation of reactive oxygen species. J. Physiol. 552, 335 (2003).
Wang, Q. S. et al. Role of mitochondrial reactive oxygen species in hypoxia-dependent increase in intracellular calcium in pulmonary artery myocytes. Free Radic. Biol. Med. 42, 642–653 (2007).
Takada, Y. et al. Hydrogen peroxide activates NF-κB through tyrosine phosphorylation of IκBα and serine phosphorylation of p65. J. Biol. Chem. 278, 24233–24241 (2003).
Song, C. et al. Oxidative stress-mediated NFκB phosphorylation upregulates p62/SQSTM1 and promotes retinal pigmented epithelial cell survival through increased autophagy. PLoS ONE 12, e0171940 (2017).
Samanta, D. & Semenza, G. L. Maintenance of redox homeostasis by hypoxia-inducible factors. Redox Biol. 13, 331–335 (2017).
Kirito, K., Hu, Y. & Komatsu, N. HIF-1 prevents the overproduction of mitochondrial ROS after cytokine stimulation through induction of PDK-1. Cell Cycle 8, 2844–2849 (2009).
Morgan, M. J. & Liu, Z. G. Crosstalk of reactive oxygen species and NF-κB signaling. Cell Res. 21, 103–115 (2010).
Djavaheri-Mergny, M., Javelaud, D., Wietzerbin, J. & Besançon, F. NF-κB activation prevents apoptotic oxidative stress via an increase of both thioredoxin and MnSOD levels in TNFα-treated Ewing sarcoma cells. FEBS Lett. 578, 111–115 (2004).
Mehla, K. & Singh, P. K. MUC1: A novel metabolic master regulator. Biochim. Biophys. Acta Rev. Cancer 1845, 126–135 (2014).
Leng, Y. et al. Nuclear import of the MUC1-C oncoprotein is mediated by nucleoporin nup62. J. Biol. Chem. 282, 19321–19330 (2007).
Ahmad, R. et al. MUC1-C oncoprotein functions as a direct activator of the nuclear factor-kappaB p65 transcription factor. Cancer Res. 69, 7013 (2009).
Lingappan, K. NF-κB in oxidative stress. Curr. Opin. Toxicol. 7, 81–86 (2018).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Perli, S. D., Cui, C. H. & Lu, T. K. Continuous genetic recording with self-targeting CRISPR-Cas in human cells. Science 353, aag0511 (2016).
Ősz, Á., Lánczky, A. & Győrffy, B. Survival analysis in breast cancer using proteomic data from four independent datasets. Sci. Rep. 11, 1–15 (2021).
Liu, Q., Palmgren, V. A. C., Danen, E. H. & Le Dévédec, S. E. Acute vs. chronic vs. intermittent hypoxia in breast Cancer: a review on its application in in vitro research. Mol. Biol. Rep. 49, 10961–10973 (2022).
Xu, Y. et al. An intronic NF-kappaB element is essential for induction of the human manganese superoxide dismutase gene by tumor necrosis factor-alpha and interleukin-1beta. DNA Cell Biol. 18, 709–722 (1999).
Yamamoto, K., Arakawa, T., Ueda, N. & Yamamoto, S. Transcriptional roles of nuclear factor kappa B and nuclear factor-interleukin-6 in the tumor necrosis factor alpha-dependent induction of cyclooxygenase-2 in MC3T3-E1 cells. J. Biol. Chem. 270, 31315–31320 (1995).
Zorov, D. B., Juhaszova, M. & Sollott, S. J. Mitochondrial reactive oxygen species (ROS) and ROS-induced ROS release. Physiol. Rev. 94, 909–950 (2014).
Tafani, M. et al. The interplay of reactive oxygen species, hypoxia, inflammation, and sirtuins in cancer initiation and progression. Oxid. Med. Cell Longev. 2016, 3907147 (2016).
Oliveira-Marques, V., Marinho, H. S., Cyrne, L. & Antunes, F. Role of hydrogen peroxide in NF-κB activation: From inducer to modulator. Antioxid. Redox Signal 11, 2223–2243 (2009).
Park, H. A., Brown, S. R. & Kim, Y. Cellular mechanisms of circulating tumor cells during breast cancer metastasis. Int J. Mol. Sci. 21, 1–19 (2020).
Yin, L., Li, Y., Ren, J., Kuwahara, H. & Kufe, D. Human MUC1 carcinoma antigen regulates intracellular oxidant levels and the apoptotic response to oxidative stress. J. Biol. Chem. 278, 35458–35464 (2003).
Hagiwara, M. et al. MUC1-C activates the PBAF chromatin remodeling complex in integrating redox balance with progression of human prostate cancer stem cells. Oncogene 40, 4930–4940 (2021).
Yin, L., Kosugi, M. & Kufe, D. Inhibition of the MUC1-C oncoprotein induces multiple myeloma cell death by down-regulating TIGAR expression and depleting NADPH. Blood 119, 810–816 (2012).
Liegel, J. et al. Phase I/Ib trial of the MUC1 inhibitor GO-203-2C alone and in combination with decitabine for acute myeloid leukemia. Blood 130, 2659–2659 (2017).
Galanzha, E. & Zharov, V. Circulating tumor cell detection and capture by photoacoustic flow cytometry in vivo and ex vivo. Cancers 5, 1691–1738 (2013).
Che, M., Wang, R., Li, X., Wang, H.-Y. & Zheng, X. F. S. Expanding roles of superoxide dismutases in cell regulation and cancer. Drug Discov. Today 21, 143–149 (2016).
Chae Ye, I. et al. Molecular portrait of hypoxia in breast cancer: A prognostic signature and novel HIF-regulated genes. Mol. Cancer Res. 16, 1889–1901 (2018).
Derose, Y. S. et al. Tumor grafts derived from women with breast cancer authentically reflect tumor pathology, growth, metastasis and disease outcomes. Nat. Med. 17, 1514–1520 (2011).
Ju, J. A. et al. Hypoxia selectively enhances integrin α5β1 receptor expression in breast cancer to promote metastasis. Mol. Cancer Res. 15, 723–734 (2017).
Ju, J. A., Godet, I., DiGiacomo, J. W. & Gilkes, D. M. RhoB is regulated by hypoxia and modulates metastasis in breast cancer. Cancer Rep. 3, e1164 (2020).
Wang, R. et al. Hypoxia-inducible factor-dependent ADAM12 expression mediates breast cancer invasion and metastasis. Proc. Natl. Acad. Sci. USA 118, e2020490118 (2021).
Gilkes, D. M. & Semenza, G. L. Role of hypoxia-inducible factors in breast cancer metastasis. Future Oncol. 9, 1623–1636 (2013).
Buendia, I. et al. Nrf2–ARE pathway: An emerging target against oxidative stress and neuroinflammation in neurodegenerative diseases. Pharm. Ther. 157, 84–104 (2016).
Mamo, M. et al. Hypoxia alters the response to anti-EGFR therapy by regulating EGFR expression and downstream signaling in a DNA methylation-specific and hif-dependent manner. Cancer Res. 80, 4998–5010 (2020).
Godet, I., Mamo, M., Thurnheer, A., Rosen, D. M. & Gilkes, D. M. Post-Hypoxic cells promote metastatic recurrence after chemotherapy treatment in TNBC. Cancers 13, 5509 (2021).
Weinstein, A. G., Gilkes, D. M. & Godet, I. Mapping the fate of hypoxic cells using an irreversible fluorescent switch. Methods Mol. Biol. 2755, 49–61 (2024).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 1–15 (2019).
Finak, G. et al. MAST: A flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA sequencing data. Genome Biol. 16, 1–13 (2015).
Korotkevich, G. et al. Fast gene set enrichment analysis. Preprint at bioRxiv https://doi.org/10.1101/060012 (2021).
Liberzon, A. et al. The molecular signatures database hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Li, B. & Dewey, C. N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC. Bioinformatics 12, 1–16 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Robinson Mark, D., McCarthy Davis, J. & Smyth Gordon, K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Cai, H. et al. VennPlex-A novel venn diagram program for comparing and visualizing datasets with differentially regulated datapoints. PLoS ONE 8, e53388 (2013).
Győrffy, B. Survival analysis across the entire transcriptome identifies biomarkers with the highest prognostic power in breast cancer. Comput. Struct. Biotechnol. J. 19, 4101–4109 (2021).
Goldman, M. J. et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 38, 675–678 (2020).
Oza, H. H. & Gilkes, D. M. Multiplex immunofluorescence staining protocol for the dual imaging of hypoxia-inducible factors 1 and 2 on formalin-fixed paraffin-embedded samples. Methods Mol. Biol. 2755, 167–178 (2024).
mconsid3 & Johnson, J. FertigLab/hypoxiaROSanalysis: v.1.0.1. Preprint at https://doi.org/10.5281/zenodo.12647476 (2024).
Acknowledgements
We thank Dr. Edward Gabrielson, MD, and Dr. Ashley Cimino-Mathews, MD, for reviewing the tissue sections and identifying areas of cancer cells. We thank Dr. Denis Wirtz, PhD and Dr. Andrew Ewald, PhD for their feedback over the course of this study. We thank Dr. Cynthia Zahnow, PhD for kindly sharing the PDX lines HCI-001 and HCI-010 that were originally gifted by Dr. Alana Welm, PhD. We are grateful for being selected for the 2018 Core Coins grant program offered by the Johns Hopkins Experimental and Computational Genomics Core (ECGC). We thank Linda Orzolek of the Johns Hopkins Medical Institutions Deep Sequencing and Microarray Core Facility for technical assistance. We are also grateful to David Mohr and the Genetic Resource facilities for providing guidance and SpaceRanger Output for the Spatial Transcriptomic Studies. Dr. Hanhvy Bui (IIC Flow Cytometry Core) and Dr. Jessica Gucwa (SKCCC Flow Cytometry Core) assisted with flow cytometry. Research in the Gilkes Lab is funded by The Jayne Koskinas Ted Giovanis Foundation for Health and Policy and the NCI/SKCCC Core grant number P50CA006973 (D.G.). Research in the Fertig lab is funded by NCI grant number 5U01CA253403-03 (E.F.). Ines Godet was supported by a postdoctoral fellowship from the National Cancer Center. The Cartoons included in the display items of this work were adapted from Servier Medical Art, licensed under a Creative Commons Attribution 4.0 Unported License (https://creativecommons.org/licenses/by/4.0/).
Author information
Authors and Affiliations
Contributions
I.G. and D.M.G. were responsible for research conceptualization and design, including the design of a Cre-dependent CRISPR knockout vector. I.G. performed all in vivo work with assistance from N.S.J., A.G.W., and H.H.O. I.G. performed in vitro studies and prepared all samples for spatial and bulk RNA sequencing. H.H.O. developed and executed the fluorescent staining protocol for FFPE samples and assisted with cloning and plasmid preparation. J.J., L.K., and M.C. used the Seurat pipeline to perform quality control testing of the spatial transcriptomics data followed by clustering and pathway enrichment analysis. R.X. assisted with qRT-PCR experiments. S.D. assisted with LoupeBrower and GSEA analyses. D.M. assisted with validating the Cre-inducible CRISPR system via immunoblot and immunofluorescence analysis. C.A.S. provided annotated human breast cancer samples and advised on the interpretation of immunohistochemistry staining. I.G., Y.S., H.H.O., S.T., and J.Y. performed experiments for the revisions requested by the reviewers. G.S., L.K., and E.J.F. advised on the design, analysis, and interpretation of the spatial transcriptomics data. L. K. performed the initial processing of the low-input RNA sequencing of CTCs. I.G. and D.M.G. performed data interpretation manuscript preparation and revision. D.M.G. was responsible for the supervision and acquisition of funding.
Corresponding author
Ethics declarations
Competing interests
C.A.S. has received research funds from AstraZeneca, GSK/Tesaro, Merck, Gilead, Celldex, BMS, and Pfizer and has received consulting fees from Seattle Genetics. E.J.F. serves on the Scientific Advisory Board of Resistance Bio and is a consultant for Merck and Mestag Therapeutics and received research funding from Abbvie, Inc. and Roche/Genentech. The other authors have no competing interests to disclose.
Peer review
Peer review information
Nature Communications thanks Nicholas Denko, Jun Liu and the other anonymous reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Godet, I., Oza, H.H., Shi, Y. et al. Hypoxia induces ROS-resistant memory upon reoxygenation in vivo promoting metastasis in part via MUC1-C. Nat Commun 15, 8416 (2024). https://doi.org/10.1038/s41467-024-51995-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-51995-2
- Springer Nature Limited