Abstract
The C-terminal domain of RPB1 (CTD) orchestrates transcription by recruiting regulators to RNA Pol II upon phosphorylation. With CTD driving condensate formation on gene loci, the molecular mechanism behind how CTD-mediated recruitment of transcriptional regulators influences condensates formation remains unclear. Our study unveils that phosphorylation reversibly dissolves phase separation induced by the unphosphorylated CTD. Phosphorylated CTD, upon specific association with transcription regulators, forms distinct condensates from unphosphorylated CTD. Functional studies demonstrate CTD variants with diverse condensation properties exhibit differences in promoter binding and mRNA co-processing in cells. Notably, varying CTD lengths influence the assembly of RNA processing machinery and alternative splicing outcomes, which in turn affects cellular growth, linking the evolution of CTD variation/length with the complexity of splicing from yeast to human. These findings provide compelling evidence for a model wherein post-translational modification enables the transition of functionally specialized condensates, highlighting a co-evolution link between CTD condensation and splicing.
Similar content being viewed by others
Introduction
The C-terminal domain of the largest subunit of RNA polymerase II (CTD) is a highly disordered region found in RNA polymerase II (Pol II), the workhorse responsible for transcribing all protein-coding mRNAs as well as some small nuclear and microRNAs in eukaryotes1,2. Different transcriptional regulatory proteins are recruited to RNA Pol II via the CTD to facilitate the progression of transcription3. The CTD recruits transcriptional proteins through extensive post-translational modifications (PTMs), with phosphorylation being the key modification during active transcription4,5.
The sequence of the CTD is surprisingly simple, with consensus heptads (historically numbered as Y1S2P3T4S5P6S7) repeated many times dependent on the species (e.g., 26 in Saccharomyces cerevisiae and 52 in humans)1. Despite of the simplicity, five of the seven residues in the heptad repeats are subject to phosphorylation, and the two proline residues can undergo isomerization, which can affect recognition by different CTD-interacting domains (CIDs) of diverse proteins that dynamically associate with Pol II throughout the transcription cycle6,7. Therefore, this simple repetitive sequence possesses an enormous capacity to encode information via combinatorial phosphorylation8. Different residues of the heptad repeats on CTD get phosphorylated at various stages of transcription, with Ser5 as the major species during initiation, recruiting capping enzymes9, and Ser2 at elongation/termination recruiting splicing and termination factors1. Instead of the traditional view of the CTD as scaffold for protein binding, the paradigm has shifted in recent years towards an ensemble view whereby Pol II functions within transcriptional condensates, the composition of which is governed in part by CTD phosphorylation10,11,12,13.
Mounting evidence shows that the CTD drives the Pol II participation into condensates10,11,12,14,15,16,17. At transcription initiation, RNA polymerase II, along with components of the Mediator complex such as MED1, accumulates through liquid-liquid phase separation (LLPS)10,11,12,15. A report observes RNA polymerase II also enters droplets characterized by spliceosome components in the same gene loci11. Mounting evidence suggests Pol II condensation via the CTD is not simply phenomenological, but evolutionarily adaptive. For example, aberrant CTD condensation properties lead to developmental failures in Drosophila, and cold tolerant fungi tune CTD condensation and its regulation for environmental adaptation10,18. Furthermore, the human CTD (52 repeats) is more prone to forming condensates compared to the yeast CTD (26 repeats)10,19, which suggests that the expansion of CTD length during evolution correlates with the increased complexity of transcription in eukaryotic cells.
Post-translational modifications (PTMs), such as phosphorylation, serve as crucial mechanistic triggers that alter the states of condensate. Diverse outcomes have been reported upon protein phosphorylation on intrinsically disordered regions (IDRs), which are frequent targets for PTMs. Both dissolution and nucleation of condensates have been demonstrated in response to PTMs20,21,22. Phosphorylation on the CTD is particularly interesting since its different phosphorylation states govern the recruitment of transcriptional regulators2,3,23. Stereospecific binding of regulators, and emerging view of the CTD as a modulator of transcriptional condensate formation, suggest a model of transcriptional condensates sorting and enriching CTD-interacting factors that in turn bind preferred heptad motifs with preferred patterns of phosphorylation. We thus focus on interplay between CTD phosphorylation, condensate formation, and CTD-interacting protein recruitment and their effects on transcription.
We employed a suite of CTD mutants, a series of CTDs with distinct post-translational modifications (PTMs) applied enzymatically, and a set of CTD-interacting proteins recognizing discrete patterns of CTD PTMs to derive rules of PTM and protein binding that govern CTD condensate formation and topological organization. Our findings reveal that CTD phosphorylation reversibly dissolves CTD condensates, yet binding by CTD-interacting partners antagonizes phosphorylation-mediated condensate dissolution. We demonstrate that phospho-CTD condensates are enabled by the binding of phospho-specific CTD-interacting proteins, and that factor binding can generate CTD condensates with layered topologies characterized by an unmodified CTD core and phospho-CTD shell harboring CTD-interacting partners. The transcriptomic analyses of human cells expressing RNA Pol II mutants harboring CTDs with varied abilities to form condensates reveal differences in promoter binding, alternative splicing, and ultimately growth defects that corroborate rules of CTD condensation derived from our biochemical assays. Our work establishes a biochemical principle for the maturation of RNA Pol II condensates associated with distinct transcriptional stages and support a model of RNA Pol II flux from promoter-associated initiation condensates to elongation condensates that support the co-transcriptional splicing. In addition, we also describe an intrinsic and essential function of the length of CTD in regulation of cell survival through altering exons skipping and intron retaining of alternative splicing events.
Results
Reversible phosphorylation of the CTD leads to phase transition
We employed a well-established system to investigate LLPS of RNA polymerase II in vitro12. This involved purifying a GST-tagged S. cerevisiae CTD domain containing 26 heptad repeats, predominantly composed of the consensus sequence (yCTD, Fig. S1a). Following purification of the recombinant GST-yCTD to homogeneity, a fluorophore was covalently attached to the N-terminus of the protein to enable visual detection. To ensure the robustness and rigor of our biochemical analysis, we ensured the purity of all protein samples used in the studies, including both the CTD variants and CTD binding proteins, through gel filtration chromatography, SDS-PAGE, differential interference contrast microscopy and light scattering. To provide further evidence of sample purity, the CTD binding proteins used in the study were able to produce diffracting quality crystals.
After labeling the GST-yCTD with a fluorophore for visual detection, confocal microscopy and phase diagram reveal that unphosphorylated CTD forms concentration-dependent condensates in the presence of a crowding agent, dextran (Fig. S1b), behaving as liquid droplets with the ability to fuse, as reported previously (Fig. S1c)12,14. We next conducted experiments to test whether phosphorylation by CTD kinases affects the ability of the CTD to form condensates24,25. Two kinases (Erk2 and Dyrk1a, respectively) were used to generate CTDs distinct patterns of phosphorylation of Ser526 (called pSer5 CTD in the rest of the paper) and Ser227(pSer2 CTD) in the context of the consensus sequence. Kinase specificity was confirmed with single, double, triplet, and 26-CTD repeats as substrates by Ultraviolet Photodissociation Mass Spectrometry (UVPD)26,27,28. We conducted the kinase reactions —treated by Erk2 or kinase domain of Dyrk1a (residue 127-485) — on CTD droplets in a time-dependent manner, simultaneously monitoring the CTD phosphorylation process and condensation states (Fig. 1a, b). The kinase phosphorylation of the CTD was monitored at each time point using Electrophoretic Mobility Shift Assay (EMSA) and Matrix Assisted Laser Desorption/Ionization – Time of Flight (MALDI-TOF) mass spectrometry (Figs. 1c and S1d, e). Simultaneously, we observed the condensate disappearance through visualization using fluorescence microscopy (Fig. 1a) and turbidity assays (Fig. 1b). The signal in turbidity assay decreased substantially more rapidly as an increasing amount of kinases resulted in faster phosphorylation (Fig. 1b). Notably, neither kinase by themselves exhibit any condensate formation under the reaction conditions (Fig. S1f). Furthermore, control conditions to exclude each phosphorylation reaction component (ATP, Mg2+ ion, and kinase) systematically reveal that condensate dissolves only when all kinase reaction components are present for phosphorylation to occur (Fig. S1g). Rigorously, we showed that the phosphate groups placed on the CTD induce condensate dissolution.
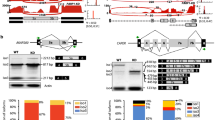
a Phosphorylation by ERK2 and kinase domain of DYRK1a gradually dissolves condensates formed by unphosphorylated CTD labeled with Texas Red-X fluorophore. Scale bars, 10 μm. b Time-course turbidity assay of unphosphorylated GST-yCTD samples treated with ERK2 (left) and DYRK1a (right) in the presence of 16% dextran. Data points represent mean values of three replicate experiments, and error bars show the standard error. c MALDI-TOF MS spectrum of ERK2 treated CTD samples (pSer5 CTD) taken from turbidity assay experiments at different timepoints. d Time-course confocal images of pSer5 CTD (labeled with Texas Red-X) treated by Ssu72 phosphatase (top) and Ssu72 C13D/D144N variant (bottom). Scale bars, 10 μm. e Time-course turbidity assay of pSer5 CTD samples treated with Ssu72 in the presence of 16% dextran. Data points represent mean values of three replicate experiments, and error bars show the standard error. f pSer5 CTD sample (top) and pSer2 CTD sample (bottom), both labeled with Texas Red-X, are treated by SCP1 overnight prior to confocal imaging. Scale bars, 10 μm. g Time-course turbidity assay of phosphorylated CTD samples treated with SCP1 in the presence of 16% dextran. Data points represent mean values of three replicate experiments, and error bars show the standard error. Source data are provided as a Source Data file.
To interrogate whether the disappearance of condensate due to phosphorylation is reversible, we used CTD phosphatases to remove the phosphate groups and monitored condensation status. Ssu72/Symplekin phosphatase complex is a component of the 3′-end cleavage and polyadenylation factor (CPF) complex conserved throughout eukaryotes29. Its dephosphorylation activity is highly specific to phospho-Ser5 of the CTD heptad with no activity against phospho-Ser230,31. While the Ssu72/Symplekin complex itself doesn’t form condensate (Fig. S1f), we directly observed droplets’ appearance and gradual accumulation in pSer5 CTD sample upon its treatment with Ssu72/Symplekin (Fig. 1d). The appearance of condensates coincides with the level of dephosphorylation (Fig. S1h). Control experiments using catalytically deficient phosphatase Ssu72 (C13D/D144N) exhibit no appearance of droplets under confocal microscopy, nor changes in absorbance at 350 nm (Fig. 1d, e). Phosphatase-regulated CTD condensation was also observed for a second enzyme, human SCP1, which displays high dephosphorylation activity against both Ser2 and Ser5 of CTD heptad32. While purified human SCP1 phosphatase domain (residue 78-263) are homogenous in solution (Fig. S1f), SCP1 treatment generates the condensate rapidly for both pSer5 CTD and pSer2 CTD (Fig. 1f), corresponding with a sharp increase in 350 nm absorbance in turbidity assay (Fig. 1g) coinciding with dephosphorylation as confirmed by EMSA and MALDI-TOF analyses, (Fig. S1i, j). Results from both phosphatase treatment experiments indicate that CTD phase transition caused by phosphorylation can be restored upon the removal of phosphate groups.
Phospho-specific association of proteins with CTD promotes the reformation of droplets
In addition to phosphorylation, another well-established post-translational modification on the RNA polymerase II is prolyl isomerization33,34. Proline residues are frequently found in intrinsically disordered regions. Yet, it was unclear whether the cis/trans proline configuration influences LLPS behavior. Human prolyl isomerase 1 (PIN1) binds the CTD only when the serine of the Ser-Pro motifs in the heptad is phosphorylated – preferably at Ser5 but also at Ser2 albeit with weaker affinity and activity35,36,37. Therefore, we investigated how PIN1’s association with phospho-CTD affects condensate formation. Quantification of the PIN1 interaction with CTD peptide containing two heptad repeats with one Ser5 phosphorylation site estimate a Kd of 21 ± 7 µM while non-phosphorylated CTD does not bind, consistent with previous reports (Fig. S2a)34. Neither PIN1 (Fig. S2b) nor pSer5 CTD (Fig. 1a) samples form droplets by themselves. When we added PIN1 into the pSer5 CTD solution, droplets appeared instantly (Fig. 2a, b). The condensate increased with more PIN1, as visualized in confocal microscopy and turbidity assay, suggesting a dose-dependent effect (Fig. 2a, b).
a Time-course turbidity assay of 10 μM pSer5 CTD samples treated with 10 μM PIN1 in the presence of 16% dextran. Addition of PIN1 at a specific timepoint (10 min) to the sample (black) was indicated with an arrow. The absorbance of control samples (pSer5 CTD only) is shown in red. Data points represent mean values of three replicate experiments, and error bars show the standard error. b Representative confocal microscopy images of 10 μM pSer5 CTD (Red, Texas Red-X) treated with different amounts of PIN1. Scale bars, 10 μm. c Representative confocal microscopy images of 10 μM pSer5 CTD (Red, Texas Red-X) treated with 10 μM of different PIN1 variants (wild-type, C113A, PPIase domain). Scale bars, 10 μm. d Representative confocal microscopy images of 10 μM pSer2 CTD and pSer5 CTD (Green, Alexa Fluor 488) treated with 10 μM of different CTD binding proteins. Scale bars, 10 μm. e Confocal microscopy of unphosphorylated CTD (Green, Alexa Fluor 488) mixed with equimolar amount of pSer5 CTD (Red, Texas Red-X) and PIN1. Scale bars, 10 μm. f Confocal microscopy of unphosphorylated CTD (Red, Texas Red-X) mixed with equimolar amount of pSer5 CTD (Green, Alexa Fluor 488) and SCAF4. Scale bars, 10 μm. g Confocal microscopy of unphosphorylated CTD (Red, Texas Red-X) mixed with equimolar amount of pSer2 CTD (Green, Alexa Fluor 488) and RPRD1b. Scale bars, 10 μm. h Confocal image of immiscible mixture of unphosphorylated CTD (Green, Alexa Fluor 488) or pSer5 CTD bound to PIN1 (Red, Texas Red-X) incubated with kinase DYRK1a or phosphatase SCP1 overnight. Scale bars, 10 μm. Source data are provided as a Source Data file.
Two non-exclusive possibilities can explain the droplet-inducing effect of PIN1 on homogenously phosphorylated CTD. First, the enzymatic activity of proline isomerization by human PIN1 may alter the local conformation of the CTD backbone to induce LLPS. Alternatively, the interaction of PIN1 with phosphorylated CTD promotes condensate formation – potentially through a mechanism that neutralizes or shields the negatively charged phosphate groups. To distinguish the two possibilities, we utilized a catalytic-deficient PIN1 with the nucleophilic cysteine mutated to alanine, C113A36. When we added the PIN1 mutant to the phospho-CTD, we observed rapid droplet formation identical to the behavior seen for wild-type PIN1 (Fig. 2c). On the other hand, when we preserved the PIN1 enzymatic domain but removed the substrate-recognizing WW domain (Fig. S2d)34, no condensate was observed (Fig. 2c). Thus, the binding of PIN1 to phosphorylated CTD is necessary and sufficient to induce condensate formation. This observation raises questions about whether the recruitment of other phospho-CTD-interacting factors could similarly counteract kinase-mediated CTD condensate dissolution, with implications for CTD condensate ‘switching’ or maturation as the CTD accumulates phosphates during the transcription cycle11,13.
The functional model of the CTD is to recruit different proteins to the RNA polymerase II at different stages of transcription based on its modification states3. Since the PIN1 association promotes condensate formation, we were curious if other proteins recruited to the phospho-CTD also induce phase separation through a similar binding-induced mechanism and investigated this directly. The CTD-interacting domain (CID) is the largest CTD binding module recognized and found in many RNA binding proteins involved in splicing or termination38. To dissect only the CTD binding function, we isolated the CID domains from SCAF4. SCAF4 is reported as a CTD-binding protein binding to both Ser2 and Ser5 with a role in transcription termination39, whose binding are confirmed with phosphorylated CTD species using fluorescence polarization assay (Fig. S2e). Increasing the concentration of SCAF4-CID and mixing it with either pSer5 CTD or pSer2 CTD led to the formation of droplets, which were observable through confocal microscopy and turbidity assays (Figs. 2d and S2f).
An intriguing inquiry arises regarding whether the condensate formation require a specific interaction between the phospho-CTD and the CID motif. To address this, we examined the purified CID domain of RPRD1b, an RNA-binding protein implicated in transcript elongation with specific binding to phospho-Ser2 but no association with phospho-Ser5 (Fig. S2g)40. The introduction of increasing amount of RPRD1b CID to the pSer2 CTD resulted in an increasing in droplet formation, whereas the same experiment using pSer5 CTD remains homogenous (Figs. 2d and S2h). This heightened concentration of RPRD1b induced more droplet scattering, as observed in turbidity assay (Fig. S2h). Notably, the CTD binding proteins (PIN1, SCAF4, RPRD1b), purified to homogeneity in all these experiments, exhibited no phase separation under DIC microscopy (Fig. S2b, i). Our experiments with CTD binding modules of recruited proteins (PIN1, SCAF4, RPRD1b) indicate that the association of phospho-CTD with binding partners enables condensate formation by the phosphorylated CTD.
Different CTD condensates remain distinct based on their physical properties
Our biochemical results show that both unphosphorylated CTD and phospho-CTD are capable of condensation. This effect echoes the cellular observation that the initiation and elongation condensates coexist without fusion on the same gene loci11. To test if CTD and phospho-CTD condensates remain distinct or undergo fusion in vitro, we used unphosphorylated CTD labeled with Texas Red (emission at 561 nm) to mimic the promoter-bound RNA polymerase II and Alexa Fluor 488-labeled phospho-CTD (emission wavelength 488 nm) to imitate an active transcribing RNA polymerase II CTD. These labeled proteins were utilized in various mixing experiments in the presence or absence of CTD binding partner PIN1. Separately, unphosphorylated CTD forms red droplets in the presence of crowding agent dextran, and phospho-CTD forms green condensates with PIN1 mixed it, consistent with our previous observation (Fig. 2e). Surprisingly, when we mixed the two CTD solutions in equal concentration, the two condensates remained immiscible, occasionally forming single condensates with a layered topology41,42,43 (Fig. 2e). Prolonged incubation did not lead to the blending of the two condensates.
Observing distinct phases formed by unphosphorylated and PIN1-bound phospho-CTD prompted us to question whether this behavior is unique to PIN1. To test this, we mixed the unphosphorylated CTD with the phospho-CTD at a 1:1 ratio under the conditions in which Texas Red labeled unphosphorylated CTD phase separates while Alexa Fluor labeled pSer5 CTD remained homogenous (Fig. 2F). The addition of the CID domains of SCAF4 induced droplet formation of the Alexa Fluor labeled pSer5 CTD, but no fusion between the unphosphorylated CTD and pSer5 CTD droplets was observed (Fig. 2f). In some instances, the pSer5 CTD formed layered condensates with the unphosphorylated CTD (Fig. 2f). Furthermore, we used a pSer2 CTD labeled with Alexa Fluor 488 to be mixed with unphosphorylated CTD (labeled with Texas Red). Adding pSer2-specific binding protein RPRD1b induced the green droplet formation but unphosphorylated CTD and pSer2 CTD droplets do not undergo fusion (Fig. 2g). Collectively, these experiments indicate unphosphorylated CTD droplets do not mix with protein-bound phosphorylated CTD droplets which coexist in distinct phases.
The biological implications of our results may be that RNA Pol II could be sorted between promotor-associated CTD condensates and elongation-associated phospho-CTD condensates as a function of phosphorylation state and factor association. Such a model requires the unphosphorylated CTD condensates to be both actionable substrates of CTD kinases and accessible to CTD-interacting factors. Thus, we tested if we could induce condensate fusion by changing the phosphorylation status and binding interactions of the unphosphorylated and phosphorylated CTDs. We mixed Alexa Fluor 488 labeled unphosphorylated CTD, Texas Red labeled pSer5 CTD and phospho-CTD binding protein PIN1; as expected, the condensates remained separated (Fig. 2h). However, upon incubation with the CTD kinase, DYRK1a or ERK2, the previously unphosphorylated Alexa Fluor 488 labeled CTD colocalizes with the Texas Red labeled pSer5 CTD (Figs. 2h and S2j). We next show the removal of CTD phosphorylation can induce similar fusion, by adding the CTD phosphatase SCP1. SCP1 dephosphorylates Texas Red labeled pSer5 CTD and enables its fusion with the Alex Fluor 488 labeled CTD condensates (Fig. 2h). Additionally, the fusion process was shown to be time-dependent (Fig. S2j, k), mirroring the kinetics observed in the phosphorylation or dephosphorylation processes (Fig. S1d, l). These results demonstrate that the PTM status of the CTD can dynamically dictate its partitioning between distinct condensate phases.
CTD binding proteins colocalized with puncta formed by phosphorylated RNA polymerase II in cells
To investigate the dynamics of unphosphorylated and factor-associated phospho-CTD condensates in vivo during transcription, we analyzed the localization of endogenous CTD binding proteins relative to phosphorylated RNA polymerase II in cells. Notably, the endogenous PIN1 protein exhibited distinctive punctate formations localized to the nucleus (Fig. 3a). Concurrently, RNA polymerase II with Ser5 phosphorylation also appeared as puncta, which colocalized with those of PIN1 (Fig. 3a, b). To confirm that these puncta were dynamic and not merely aggregates, we employed fluorescence recovery after photobleaching (FRAP) on PIN1-YFP puncta in living cells. The partial recovery of fluorescence post-bleaching (Fig. S3a) suggests that PIN1 partitions into and associates with the phospho-CTD condensates dynamically and is not aggregated. Consistently, inhibiting Ser5 phosphorylation with a CDK7 inhibitor disrupted the formation of PIN1 puncta in vivo (Figs. 2a, b and S3b). Compounded on our and others’ previous findings35,36,37, these observations suggest that PIN1 phase separates specifically with phosphorylated RNA polymerase II. Further analysis of existing Chromatin Immunoprecipitation (ChIP)-Seq datasets [GSE231126, GSM4442247] revealed that PIN1 interacts with the CTD at the promoters of actively transcribing genes, with a sharp peak in the PIN1 profile at the transcription start site, coinciding with the Ser5 phosphorylation signal (Fig. S3c). These results from both ChIP-Seq analysis and immunofluorescence staining of endogenous proteins demonstrate that PIN1 interacts with pSer5 at gene loci around the promoter regions.
a U2OS cells were treated with DMSO or THZ1 (1 μM, 8 h), then fixed with paraformaldehyde and subjected to immunofluorescence staining. PIN1 (green) was co-stained with phospho-Ser5 CTD (red) using specific antibodies. Scale bars, 2 μm. b The average immunofluorescence intensity of puncta was analyzed, and corresponding line plots were generated for each immunofluorescence signal. c–g U2OS cells were fixed and subjected to triple immunofluorescence staining. PRPF8 (green), SRSF1 (green) and PIN1 (green) were co-stained with phospho-Thr4 CTD (red), phospho-Ser5 CTD (red) and MED1 (blue) using specific antibodies. Line plots were generated for PRPF8 (d), SRSF1 (e) and PIN1 (g) immunofluorescence signal separately. Scale bar, 2 μm. h U2OS cells were transfected with SCAF6-HA or SRSF2-Flag constructs and subjected to immunofluorescence staining by HA, Flag and phospho-Thr4 CTD antibody. Scale bars, 2 μm. Source data are provided as a Source Data file.
To understand phosphorylated RNA polymerase II condensates in other biologically relevant contexts, we analyzed its localization with spliceosome components. Core spliceosome components such as PRPF8 and SRSF1 have been reported to partially form condensates with phosphorylated RNA polymerase II11. Utilizing MED1 as a marker to identify the potential locations of the Mediator complex44, three-channel staining revealed that PRPF8 and SRSF1 formed puncta and colocalized with the pThr4 form of RNA Pol II. However, the MED1 puncta were distinctly separate from those of the spliceosome components (Fig. 3c–e). Additionally, we observed PIN1 formed puncta that colocalized with pSer5 of RNA Pol II, separate from MED1 puncta (Fig. 3f, g, left panel). Intriguingly, a transition from MED1 puncta to PIN1 puncta associated with pSer5 was observed in cells (Fig. 3f, g, right panel). This transition suggests that initiation-coupled CTD phosphorylation and unphosphorylated CTD exist in proximity during the initiation phase. Taken together, these imaging results indicate that CTD-binding proteins colocalize with puncta formed by phosphorylated RNA polymerase II in cells, distinct from those formed by Mediator complex relative to unphosphorylated CTD.
CTD binding proteins facilitated the formation of phosphorylated RNA polymerase II puncta in vivo
To corroborate our in vitro observation that association with CTD binding proteins can promote phospho-CTD condensate formation, we transfected U2OS cells with SCAF6, a protein known to directly interact with the CTD45. The transfected cells exhibited distinctive puncta formation localized to the nucleus (Fig. 3h). Overexpression of SCAF6 greatly enhanced the formation of phospho-CTD puncta compared to the empty vector, as demonstrated by RNA polymerase II with Thr4 phosphorylation that colocalized with SCAF6 (Fig. 3h). To further strengthen our notion, we also overexpressed SRSF2, a component of the spliceosome indirectly associated with phospho-CTD in vivo11, we observed in cell that SRSF2 molecules formed puncta, which exhibits significant overlap with phosphorylated Thr4 RNA polymerase II (Fig. 3h). Significantly, the overexpression of SRSF2 greatly enhance the condensation of phospho-Pol II. In contrast, overexpression of empty vector or Ssu72, which affect pSer5 level of Pol II, did not result in similar puncta formation (Figs. 3h and S3d), underscoring that the facilitation of phosphorylated RNA polymerase II puncta formation by CTD-binding proteins is specific. Thus, consistent with our biochemistry results that interactions of CTD-specific binding proteins induce the condensate formation of pCTD (Fig. 2), the imaging data also indicate that the overexpression of CTD binding proteins in cells greatly enhanced the condensation of phospho-CTD. These observations support a model of compositionally distinct Pol II condensates, governed at least in part by the phospho-specific associations of CTD-interacting factors.
Effect of electrostatic charges on CTD condensates
To link our biochemical observation to transcriptional function, we engineered the consensus CTD to mimic the phosphorylation state at different sites of the heptad. We hypothesize that reversible phosphorylation of the CTD dissolves the formed droplet because the repulsion between negatively charged CTD molecules counteracts attractive interactions, such as π-π interactions46. If this hypothesis holds, a negative charge installed at any position within each CTD heptad would disrupt condensate formation. To test this hypothesis, we employed phosphomimetic mutations, inserting glutamate at T4, S5, or S7 position of each heptad (Fig. S1a) and compared the condensation of each variant with wild-type CTD in vitro (Figs. 4a–c and S4a, b). Consistent with previous report10,19, the shorter CTD variant from yeast displayed a reduced capacity for condensate formation compared to the wild-type human CTD (Fig. 4a, b). Interestingly, unlike the wild-type CTD, no condensation was observed in phosphomimetic mutations even in the concentrations of protein greatly exceeding the saturation concentration for the wild-type CTD or with increased concentrations of crowding agent (Figs. 4c and S4a, b). This observation was corroborated by turbidity assays, which showed no absorbance indicative of light scattering by droplets (Fig. S4d). On the contrary, S7K spaced variant (positive charge) showed a similar tendency in phase separation as wild-type yeast CTD, with a slight shift toward less phase separation (Fig. S4c, d).
a Phase diagram of wild-type GST-hCTD. Black-filled circles indicate conditions where liquid-liquid phase separation is observed with DIC microscope. Phase diagram of GST-yCTD (b), GST-T4E yCTD (c), GST-T4E-spaced yCTD (d) and GST-S7E-spaced yCTD (e). f ChIP-Seq analyses of genomic binding of RPB1_52xCTD and RPB1_26xCTD. The heatmaps and profiles are generated using deepTools2 suite70. The heatmaps show bigWig scores over genomic intervals. The rows (genes) are sorted based on the mean score per region in the descending order. The profile above shows average signal over all the genes. The genome is binned to the size of 50 bp. The genes are scaled (either stretched or shrunken) to fit into 2000-bp window between TSS and TES. Also, a window 2 kb upstream from TSS and 2 kb downstream of TES are shown. g ChIP-Seq analyses of genomic binding of RPB1_26xCTD T4E (all), 26xCTD T4E (spaced) and 26xCTD S7E (spaced). h ChIP-Seq example illustrating the association of different RPB1 CTD constructs along with the active transcribing genes. i Boxplot of the pausing index in 26xCTD and 52xCTD (19,630 protein-coding genes). Shown is one representative paired biological replicate: CTD_26x vs CTD_52x. Boxplots represent median (50%) with interquartile range (25–75%), whiskers show minimal and maximal values, outliers not shown. Statistical comparison was performed using two-sided paired Wilcoxon test. ns not significant. Source data are provided as a Source Data file.
We then asked whether the frequency and spacing of the phosphomimetic negative charge affected phase separation. To test that, we mutated every other T4 or S7 to negatively charged glutamate residues (T4E-spaced or S7E-spaced) (Fig. S1a). These spaced variants are capable of forming liquid droplets (Fig. 4d, e), however, the concentrations of protein and dextran required to induce the liquid phase are significantly higher than those needed for the wild-type CTD. The effect of negative charges was dose dependent ― the wild-type CTD (no negative charge) (Fig. 4a, b), T4E-spaced and S7E-spaced (one negative charge every two heptads) (Fig. 4d, e), S5E, T4E, and S7E (one negative charge every heptad) (Figs. 4c and S4a, b) display a correlation between the density of negative charges to the loss of phase separation, regardless of their position within the heptad. These results suggest that threshold levels of negative charge impair CTD condensation and provide an experimental platform to relate CTD condensation properties in vitro with functional effects on RNA polymerase II transcription in vivo.
CTD condensation properties in vitro predict genomic locations of RNA polymerase II in vivo
The CTD of RNA polymerase II is crucial for eukaryotic transcription. However the functional and evolutionary significance of its variation, which exhibits diverse condensation properties, remains not fully understood. To study the effects of CTD condensation in the context of the full polymerase, we introduced RPB1 plasmids harboring 52xCTD (human wild-type), 26xCTD consensus heptads, and 26xCTD mutants heptads with YFP in HEK 293 T cells. Previously, multiple labs have shown that 52xCTD is more disposed to condensate formation than 26xCTD10,19. For 26xCTD mutants, we focused on T4E/S7E mutants to dissect the effects of CTD charges on transcription without the confounding variable of S2 and S5 phosphomimetic mutations that abolish transcription47. We performed ChIP-Seq analysis to identify genomic locations of Pol II harboring the CTD mutants with differing condensation properties. We first validated the relative expression of the ectopically expressed Pol II mutants in HEK 293 T cells (Fig. S4e, f), then preformed ChIP-Seq. Each Pol II mutant was immunoprecipitated using anti-GFP antibodies to specifically map distribution of the mutant Pol II isoforms. The genome-wide profiles of RPB1_52xCTD and 26xCTD looked very similar (Fig. 4f), consistent with a role for CTD condensation in targeting Pol II to promoters10,19. Intriguingly, the binding of RPB1_26xCTD T4E/S7E mutant was completely lost genome-wide, most notably in the TSS region (Fig. 4g). The distribution of RNA polymerase II along the gene body was altered exemplified in multiple genes (Fig. 4h). Consistent with these findings, subcellular localization studies showed that both 52xCTD and 26xCTD formed puncta in cells, whereas the T4E/S7E mutants remained diffused throughout the nucleus and did not form puncta (Fig. S4g, h). The degree of global impairment of Pol II distribution as a function of negative charge is consistent with a model of impaired phospho-Pol II recruitment to unphosphorylated Pol II condensates predicted by our in vitro results.
To quantify the change of distribution of the polymerase over gene regions, we calculated the “Pausing Index” (PI) as the ratio of Pol II read density near the promoter (–50 to +300 bp of Transcription Starting Site Region/TSSR) over the remainder of the gene body (+300 downstream of the TSS to +3 kb past the Transcription Termination Site/TTS)48. The protein-coding genes (n = 19,630) were clustered into four groups based on the pausing index (Fig. S4i). Comparison of the pausing index of all the genes showed no statistically significant difference between 26xCTD and 52xCTD (Fig. 4I), with only slightly decreased PI (p = 1.611e−06) was observed in G3 cluster of most-paused genes in 26xCTD (Fig. S4j), consistent with previous report49. Taken together with our biochemical data (Fig. 4a–e), these results showed both 52xCTD and 26xCTD are competent to partition into unphosphorylated Pol II clusters at the promoters genome-wide. Conversely, negatively charged phospho-CTD Pol II mutants are impaired in their ability to partition into unphosphorylated Pol II condensates resulting in failed initiation that scales with negative charge.
Growth study and transcriptomic analysis of RNA polymerase II with different CTD variants
To interrogate the role of different CTD variants in vivo without the confounding variable of endogenous wild-type Pol II, we generated YFP_RPB1_52xCTD, 26xCTD and 26xCTD T4E/S7E constructs in the context of a mutant version of RPB1, N792D, conferring resistance to the potent Pol II inhibitor, α-amanitin50. Expression of the α-amanitin resistance Pol II mutants was similar, measured by both fluorescence intensity and western blotting (Figs. 5a and S5a). After 72 h of α-amanitin administration to degrade endogenous Pol II, we evaluated the physiological function of transfected Pol II mutants. The results showed that both RPB1_52xCTD and 26xCTD could sustain cell viability, with the RPB1_26xCTD exhibiting slowed growth49 (Fig. 5a, b). The T4E/S7E mutants exhibited a greater reduction of cell survival after endogenous RPB1 depletion by α-amanitin (Fig. 5a, b), consistent with impaired CTD condensation in vitro and Pol II promoter association in cells (Fig. 4c–e, g).
a Fluorescence and bright filed (BF) imaging of HEK 293 T which transfected with YFP_RPB1_52xCTD, 26xCTD, 26xCTD T4E (all), 26xCTD T4E (sp), and 26xCTD S7E (sp) constructs upon α-amanitin administration. Scale bars, 500 μm. b Cell density in transfected RPB1 with different CTD constructs upon α-amanitin administration. n = 3 independent experiments (mean ± SEM); ****p < 0.0001, comparison was performed using two-way ANOVA with Tukey correction. c U2OS cells were transfected with YFP_RPB1_52xCTD, 26xCTD, and then fixed and subjected to immunofluorescence staining. RPB1_52xCTD (green) and RPB1_26xCTD (green) were co-stained with SRSF1 (magenta) using GFP and SRSF1-specific antibodies. The numbers of RPB1 and SRSF1 positive puncta were quantified from more than 10 cells across three independent experiments (mean ± SEM), and plot was generated accordingly. **p = 0.0074, ****p < 0.0001, comparison was performed using two-way ANOVA with Tukey correction. Scale bar, 2 μm. d Types and absolute numbers of annotated alternative splicing events (ASE) that were statistically significant in RPB1_26xCTD vs. RPB1_52xCTD cells (FDR < 0.0005). In event type illustration, the constitutive exon is black, whereas alternatively spliced exons are striped. e, f Examples of Sashimi plots of TMEM106C and PIP5KL1 genes in RPB1_26xCTD compared to RPB1_52xCTD: read densities are shown on the y-axis. Arrow indicates the inclusion of an alternative exon or intron. Distribution of intron (g) or exon (h) sizes for alternative splicing events in RPB1_26xCTD (Tufte’s boxplots in blue: the dot indicates the median, blank space indicates interquartile range) in comparison with annotation (violin plots with median dot in red). Only high-confidence AS events (FDR < 0.0005) from two biological replicates (RPB1_26xCTD vs RPB1_52xCTD) are plotted. g: n = 84,050 for annotation and n = 356 for RPB1_26xCTD condition. h: n = 75,989 for annotation and n = 1859 for RPB1_26xCTD condition. Both Tufte’s boxplots (in blue) and violin plots (in red) represent median (50%, dots) with corresponding interquartile range (25-75%), whiskers show minimal and maximal values. i Calculated lengths of genes with skipped exons and retained introns in RPB1_26xCTD (with parameters FDR < 0.05; ILD inclusion level difference, ≥10%) compared to annotation. Bar plot representing the median gene lengths for each group. Source data are provided as a Source Data file.
To further dissect the role of different CTD repeats in transcription and cell viability, we initially compared the puncta formation of RPB1_52xCTD and RPB1_26xCTD in vivo, which, consistent with in vitro data (Fig. 4a, b), showed that 52xCTD forms more puncta in cells (Fig. 5c). Building on this observation, we also examined the assembly of the mRNA processing complex in cells expressing 52xCTD and 26xCTD. Notably, one of the spliceosome factors, SRSF1-positive puncta were fewer and more dispersed in cells with 26xCTD (Fig. 5c), suggesting that defects in condensate formation led to reduced spliceosome puncta formation in vivo. Subsequently, we conducted whole-transcriptome RNA-Seq to detect the polyadenylated mRNA in RPB1 N792D mutant with 52 or 26xCTD repeats, respectively. Differential expression analysis/DEG (Fig. S5b) in RPB1_26xCTD compared to 52xCTD revealed that among a total of 39,146 annotated expressed genes (counts > 0), there were only 675 genes upregulated (1.7%) and 742 genes (1.9%) downregulated in 26xCTD (log2FC cutoff = 0.58, p-adjusted cutoff = 0.05). This data suggests that 26xCTD only modestly alters transcription globally. Additionally, analyzing occupancy data in ‘DiffBind’ R package, we derived and overlapped 26xCTD and 52xCTD consensus ChIP-Seq peaks, as well as their shared peakset, with genomic locations of putative enhancers in HEK 293 T cells from EnhancerAtlas annotation51. As a result, nearly half of the shared peaks overlapped enhancer regions, and a total percentage of peaks overlapping enhancers was higher in 52xCTD (21.5%) compared to 26xCTD (15.4%) (Fig. S5c). These results are consistent with the previous observation that CTD shortening impairs enhancer transcription49.
The length of CTD alters alternative splicing
Our biochemical and cellular studies suggested that Pol II with impaired condensation could be associated with splicing defects (Figs. 4a, b and 5c). The truncated 26 heptad CTD recalls the length of CTD in S. cerevisiae, where splicing events are rare compared to human (52 heptad CTD). Using RPB1_26xCTD, which supports viability despite condensation defects, we investigated the splicing outcome of the truncated CTD. To test this, we used rMATS-turbo52 followed by SpliceTools53 suite to analyze splicing differences in shortened (26x) vs. full-length (52x) CTD. We identified substantial differences with a total number of 16,168 significant alternative splicing (AS) events in RPB1_26xCTD vs. RPB1_52xCTD (FDR < 0.05) (Fig. S5d). We then set an additional constraint [ILD ≥ 10%], where ILD (inclusion level difference) reflects the difference of the splicing pattern in RPB1_26xCTD vs. RPB1_52xCTD. As a result, a total of 9239 AS events were found statistically significant (FDR < 0.05, ILD ≥ 10%). Interestingly, CTD shortening preferentially leads to a decrease in exon inclusion levels (skipped exons/SE: 4350 events in 2812 unique genes) rather than an increase. The opposite trend was observed for intron retention (RI) events: 511 introns in 426 unique genes were retained in RPB1_26xCTD samples (Fig. S5e). When only high-confidence AS events (FDR < 0.0005) are considered, RPB1_26xCTD causes threefold more frequent exon skipping and 6-fold more frequent intron retention compared to alternative event (Fig. 5d). Specific examples of SE (ILD = 100%, FDR = 2.95e-06) (Fig. 5e) and RI (ILD = 51.4%, FDR = 1.8911e−10) (Fig. 5f) events in RPB1_26xCTD condition vs. RPB1_52xCTD are shown as Sashimi plots. Using the SpliceTools suite, we found that the global footprint of CTD shortening on AS of the 26xCTD cell resembles that of the knockdown of spliceosome components (Fig. S5f). It was estimated that increased exon skipping occurred in 9% (fraction = 0.0879) of all expressed genes with TPM ≥ 3, which was similar to the effect of SRFBP1 (RNA-binding protein/RBP) knockdown but somewhat lower than the effects of the core spliceosome knockdown (U2AF1 and U2AF2)53 (Fig. S5f).
Further analysis of the skipped exon/retained intron sizes in 26xCTD condition showed that the median size of retained introns upon 26xCTD was considerably shorter than the value derived from an input annotation file, p < 2.2e−16 (Fig. 5g). On the contrary, exons located upstream/downstream from the skipped exons in 26xCTD were not different from the annotation median (Fig. 5h). In addition, analysis of the length of the genes with SE or RI in 26xCTD revealed that exon skipping events were associated with longer genes and intron retention was associated with shorter gene lengths compared to hg38 annotation median (Fig. 5i). Therefore, comparison with annotation (19,966 unique protein-coding genes) suggest that 26xCTD has reduced accuracy when including exons in longer genes and/or cutting out introns from shorter genes. A comparison with other AS datasets from the SpliceTools collection (https://github.com/flemingtonlab/SpliceTools/tree/main/data/1_RBP_kd) showed that perturbation of two major pre-mRNA splicing factors - U2AF1 and U2AF2 - also led to frequent exon skipping: 2617 and 3267 events, respectively (FDR < 0.0005). The introns/exons alternatively spliced under 26xCTD condition did not differ in their sizes from those under U2AF1 and/or U2AF2 knockdown, suggesting that CTD shortening-induced AS pattern resembles patterns induced by splicing factors perturbation, rather than being CTD-length specific (Fig. S5g, h).
Furthermore, binding of U1 splice donor factors and U2 splice acceptor factors to initial transcripts depend on the quality of the splice junction sequences. Using SE/RI SpliceSiteScoring function from SpliceTools, we assessed the scores of all splice junction sequences for significantly altered SE and RI events (FDR < 0.0005) in RPB1_26xCTD, which showed that short CTD facilitated skipping of exons with weaker splice site scores (p < 1.268e−10) (Fig. S5i) and retaining of introns that are surrounded by “weaker-scored” donor and acceptor exons (p < 1.283e−05) (Fig. S5j). Taken together, these data imply that the short CTD likely has defects in spliceosome recruitment to RNA Pol II condensates leading to aberrant splicing.
To better understand the physiological impact of impaired splicing in the context of CTD mutations, we used ‘SETranslateNMD’ from the SpliceTools suite to understand the consequences of exon skipping caused by 26xCTD. Intriguingly, ~43% of alternative transcripts associated with a high-confidence (FDR < 0.0005) set of skipped exons by Pol II 26xCTD were predicted to undergo nonsense-mediated mRNA decay (NMD) (Fig. 6a). Therefore, 26xCTD-mediated exon skipping can frequently lead to NMD of transcripts, potentially decreasing their effective expression level. Overrepresentation analysis (Biological Process/BP, p.adjusted < 0.05) showed that frameshifted transcripts were enriched for genes playing a role in cell cycle transition (Fig. 6b), which was consistent with decreased proliferation of 26xCTD transfected cells. Some representative genes with exon skipping under 26xCTD (FDR < 0.0005), AURKB (ILD = −0.142, FDR = 1.56e−11) and CDK4 (ILD = −0.026, FDR = 3.96e−05) are demonstrated as examples of NMD-transcripts (Fig. 6c, d). These genes are known as master regulators of the cell cycle and DNA replication54,55, and previous report also showed alternative splicing governs cell cycle progression through NMD genes, including AURKB56. Indeed, one-fourth (100/463) of unique genes predicted to undergo NMD (Fig. 6e) were found to be downregulated by DEG analysis (FDR < 0.05). Furthermore, 7 genes out of these 100 belonged to GO:1901990 (“Regulation of mitotic cell cycle phase transition”) which was enriched among NMD-transcripts. To validate this analysis, we performed qPCR for two cell cycle progression-related genes, namely, CDK4 and ANAPC5 (Fig. 6f). As the experiments have shown, skipped isoforms of CDK4 and ANAPC5 were expressed significantly higher under 26xCTD condition (p < 0.05), whereas included isoforms were expressed at the same levels.
a NMD or non-NMD of transcripts detected in skipped exons in RPB1_26xCTD (with parameters FDR < 0.0005). b Gene ontology of biological processes enriched among NMD (nonsense-mediated mRNA decay) of genes in RPB1_26xCTD skipping exons. Overrepresentation analysis was performed using hypergeometric test, with BH (Benjamini-Hochberg) adjustment for multiple comparisons. c, d Examples of genes related to cell cycle in RPB1_26xCTD skipping exons, leading to frame shift and premature termination of gene transcription. e Venn diagram depicting unique genes predicted to undergo nonsense-mediated decay (NMD) compared to those identified in differential expression gene (DEG) analysis. f qPCR for two cell cycle progression-related genes: CDK4 and ANAPC5. Shown are results from three (n = 3) independent biological replicates (dots), each was performed in technical duplicates. Data are presented as mean ± SD. Statistical comparison was performed using unpaired one-tailed t-test. **p = 0.0063, ∗ p = 0.0229, ns not significant. g Comparison of the ability in condensate formation in vitro and biological function in vivo among multiple CTD variants. h Model for phase separation sorting of RNA polymerase II. h Created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. Source data are provided as a Source Data file.
Overall, these results support our hypothesis that the length of CTD affects functional transcriptional condensate assembly, is involved in the recruitment of splicing factors, and can lead to defects in recognizing precise splice sites in case of shortened CTD repeats.
Discussion
It was well accepted that a primary function of CTD was to facilitate pre-mRNA splicing by recruiting splicing factors23. However, recent high-resolution cryo-EM structures of spliceosomes have revealed no observable physical interaction with the CTD33,57,58,59,60. In this study, we provide a molecular mechanism for this apparent discrepancy. The interaction between phospho-CTD and splicing factors promotes the formation of splicing condensates, increasing the local concentration of splicing components. This, in turn, facilitates the spliceosome assembly and enhances the efficiency of pre-mRNA processing.
Through cellular imaging, we have observed that phosphorylated RNA polymerase II associates with spliceosome components, forming distinct puncta. Interestingly, the expansion of the CTD length during evolution seems to align with the increased frequency and complexity of splicing events in eukaryotic cells. Fine-tuning the capacity for condensate formation inside cells appears to play a crucial role in splicing precision. Notably, when we examined an RPB1 variant with shortened CTD comprising 26 heptad repeats, we observed profound differences in alternative splicing patterns (Fig. 5d). This result differs from a previous study where little difference was found when only long transcripts were sequenced49. In our study, the most noticeable difference lies in the short genes when the introns end up included up to six-fold more frequently. In longer genes, we saw a tendency of exon inappropriately skipped. Such errors were more pronounced at weaker splicing sites. We highlight that truncation of the CTD by a factor of two impairs splicing outcomes by a factor of 3 for skipped exons and 6 for retained introns. One implication of this non-linear impairment is that altered CTD phase separation, not simply a reduction of splicing factor binding sites by half contributes to splicing defects for Pol II mutants with truncated CTDs consistent with the ability of phase transitions to transform linear inputs (CTD length) into non-linear functional outputs with consequences for organismal fitness (splicing)61. Intriguingly, these alternative splicing events affect numerous genes that regulate cell growth. The alternatively spliced isoforms encoded shorter proteins with premature termination, providing an additional explanation for the observed deficiency in cellular growth associated with shortened CTD.
In our study, we generated multiple RPB1 variants with negative charges and consistently observed their inability to bind to the promoter for transcription initiation (Fig. 6g). This finding aligns with previous research indicating that promoter binding requires unphosphorylated RNA polymerase II62. The assembly of the PIC requires hydrophobic interaction facilitated by unphosphorylated CTD (Fig. 6h). The recently published structure of Mediator and RNA Pol II63 revealed that a residual CTD fragment bridges the interface between the Middle and Head of Mediator complex involving Y1 residues of CTD heptad making hydrophobic contacts64,65,66. The high local concentration of the Pol II at the site of transcription initiation can account for the phenomenon of transcriptional bursts. While dominated by hydrophobic forces like π-π stacking as seen in unphosphorylated CTD and Mediator, once transcription starts, CTD kinase-mediated phosphorylation of Pol II disrupts these hydrophobic interactions (Fig. 6h). As phosphorylation accumulates on the CTD, it recruits phospho-specific binding proteins, predominantly RNA- processing factors (Fig. 6h). In this scenario, phospho-CTD functions as the scaffold for multivalent interaction with the transcription regulatory proteins (clients), evolving the initiation condensate into one tuned for splicing. The existence of condensates with layered topologies in other membraneless organelles43, such as nucleolus, which contains subcompartments representing distinct coexisting condensates67. The layers of condensates give rise to a rational, organized factory for RNA synthesis, folding, and processing, facilitating the maturation of RNA. Similarly, layers of condensates have been observed in coexisting chromatin condensates68. Our in vitro and in vivo study of RNA polymerase II suggests a model where transcription initiation condensates including unphosphorylated CTD evolve to transcript-processing condensates as a function of Pol II phosphorylation and splicing factor recruitment. Our results shed new light of LLPS into the classic model of CTD function23 where phosphorylation acts to recruit transcriptional regulators not only via binding but by recruitment to functional condensates, highlighting how the CTD variations with different physical properties can affect splicing outcome perhaps by altering the local concentration of splicing factor clients of Pol II condensates.
Methods
Bacterial strains, cell lines, reagents, and antibodies
All Escherichia coli strains were grown in L.B. (Luria–Bertani) or Terrific Broth media at 37 °C as indicated below. HeLa, HEK 293 T and U2OS cells were from ATCC. No cell lines used in this study were found in the database of commonly misidentified cell lines maintained by ICLAC and NCBI Biosample. All cell lines were cultured in DMEM medium with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin solution at 37 °C in 5% CO2 (v/v).
Polyethyleneimine (PEI, Polysciences) and FuGENE HD (Promega) transfection reagents were used according to manual instructions and transfected plasmids in cultured cells. The α-amanitin (sigma) was used for endogenous RPB1 degradation, the concentration is 2.5 μg/mL. THZ1 (Ambeed) was used for Pol II CTD pSer5 inhibition.
The monoclonal antibodies anti-CTD pS5 (Millipore, 04-1572, 1:300 dilution), anti-CTD pT4 (Active Motif, 61361, 1:100 dilution), anti-MED1 (Abcam, ab313323, 1:100 dilution), anti-PRPF8 (Santa Cruz, sc-55533, 1:50 dilution), anti-SRSF1 (Santa Cruz, sc-33652, 1:50 dilution), PIN1 (Santa Cruz, sc-46660, 1:50 dilution), anti-HA (Cell Signaling Technology, #3724, 1:500 dilution), anti-Flag (Cell Signaling Technology, #14793, 1:500 dilution) were used for immunofluorescence staining. The anti-GFP (Proteintech, 50430-2-AP) was used for Chip experiment 5 μg/sample. The anti-rabbit IgG antibody was purchased from Invitrogen (08-6199). All of the antibodies used are listed in Supplementary Table 1.
Constructs
All 26x yCTD constructs (WT, S5E, S7E, S7E in every other repeat, T4E, T4E in every other repeat) were ordered as synthetic genes (Genscript), amplified, and cloned using ligation-independent cloning (SLIC)36. For bacterial protein expression, all constructs were cloned into a PET28a vector (Novagene) containing a 6x histidine tag and glutathione-S-transferase (GST) tag with a 3C protease cleavage site added after the two tags. For mammalian protein expression, YFP-RPB1-WT (52xCTD) was obtained from Addgene, YFP-RPB1-26xCTD and T4E/S7E mutants were generated by PCR-based cloning performed by a kit from ThermoFisher. For U2OS cell transfection & imaging, PIN1 were cloned into a pcDNA3 vector containing YFP tag with a 3C protease cleavage site. SRSF2 were obtained from Addgene and cloned into a pcDNA3 vector containing Flag and BFP tag. Ssu72 and SCAF6 were cloned into a pRK5 vector containing HA tag. All coding sequences were verified by DNA sequencing.
Protein purification
E. coli (DE3) cells were used as the protein expression system for the proteins used in the study. The transformation was carried out by thawing the competent cells on ice for 5 min, adding the DNA to cells and incubating on ice for 30 min, heat shocking at 42 °C for 90 s, and finally cooling the cells on ice for 3 min. The cells were recovered in SOC medium for 1 h at 37 °C and were plated on Luria-Bertani agar plates containing 50 μg/mL kanamycin for selection. Individual colonies were grown in 50 mL of Luria-Bertani medium at 37 °C containing 50 μg/mL kanamycin. 1 L of terrific broth medium (Thermo Fisher) was inoculated with 10 mL of inoculum, and the culture was grown to an O.D. of 0.4–0.6. 0.5 mM IPTG was added to each culture to induce the protein expression. The cultures were pelleted by centrifugation after overnight growth (20 h at 16 °C), and the cells were lysed through sonication in a lysis buffer (50 mM Tris-Cl pH 8.0, 500 mM NaCl, 10% glycerol, 0.1% Triton-X 100, 20 mM imidazole, and 10 mM BME). Sonication of cell pellets were carried out on ice at 90 A for 3 min per cycle (1 s on and 5 s off) for five cycles with a 3 min break between each cycle. The lysate was cleared by centrifugation at 27000 g for 45 min at 4 °C. The supernatant was purified through affinity column chromatography using Ni2+/NTA beads (Qiagen). The column was equilibrated with lysis buffer. Then, the cleared lysate supernatant was run through the column. The column was washed with 10 times column volume of wash buffer (50 mM Tris-Cl pH 8.0, 500 mM NaCl, 10% glycerol, 20 mM imidazole, and 10 mM BME) and eluted with an elution buffer (50 mM Tris-Cl pH 8.0, 500 mM NaCl, 10% glycerol, 250 mM imidazole, and 10 mM BME). Proteins were dialyzed in a gel filtration buffer (50 mM Tris-Cl pH 8.0, 500 mM NaCl, 10 mM BME) at 4 °C overnight. Proteins were concentrated using centrifugal concentrator (Sartorius) and further purified with size exclusion chromatography using a Superdex 200 column (GE Life Sciences). The purity of each protein fractions was assessed by polyacrylamide gel electrophoresis (Coomassie Brilliant Blue Staining).
Covalent labeling of the CTD molecules
Two different kinds of succinimidyl ester probe (Invitrogen) that contain different fluorescent dye (Texas RedTM and Alexa Flour 488) were purchased and stored at −20 °C as powder samples. One milligram of each dye was dissolved in 100 μL of DMSO and mixed with 1 mL of 10 mg/mL GST-yCTD protein sample. Each protein-dye mixture was incubated for 1 h at 25 °C with continuous stirring. Then the protein-dye conjugate was separated from the unreacted dye by using size-exclusion chromatography. A Superdex 200 column (GE Life sciences) was equilibrated with PBS, and the reaction mixture was separated by using PBS as the gel filtration buffer. The fraction corresponding to conjugated GST-yCTD were collected and concentrated for storage at −80 °C.
CTD phosphorylation
GST-yCTD was phosphorylated using either Homo sapiens ERK2 or GST-tagged H. sapiens kinase domain of DYRK1a. 5 mg/mL GST-yCTD were incubated with 0.125 mg/mL ERK2 or 0.5 mg/mL GST-DYRK1a, supplemented with 50 mM TRIS pH 7.5, 5 mM ATP, and 5 mM MgCl2. After overnight incubation at 30 °C, the phosphorylation reaction was quenched by adding EDTA to a final concentration of 5 mM. After completion of overnight reaction, phosphorylated GST-yCTD was mixed at a final concentration of 10 μM into 16% dextran, 50 mM TRIS (pH 7.5), 150 mM NaCl, 10% glycerol, and 1 mM DTT in order to check condensate formation with microscopy or turbidity assay (as described below).
Kinase and phosphatase treatment assay on condensates
Before kinase treatment, GST-yCTD droplets were formed by mixing 10 μM GST-yCTD in 50 mM TRIS (pH 7.5), 150 mM NaCl, 10% glycerol, 1 mM DTT with 16% dextran. After droplet formation, 2 mM ATP, 2 mM MgCl2, and appropriate amount of kinase (ERK2 or DYRK1a) were mixed and incubated at 30 °C with continuous shaking. For phosphatase treatment experiments, phosphorylated GST-yCTD sample was mixed at a final concentration of 10 μM into 16% dextran, 50 mM TRIS (pH 7.5), 150 mM NaCl, 10% glycerol, and 1 mM DTT, then appropriate amount of phosphatase (SCP1 or Ssu72) was added and incubated at 30 °C with continuous shaking. Droplet disruption or formation was monitored by microscopy or turbidity assay (as described below). Phosphorylation of GST-yCTD samples was confirmed by using gel shift assay (EMSA) and MALDI-TOF mass spectrometry (as described below).
Turbidity assay
Turbidity assays were carried out in 50 μL samples containing 50 mM TRIS (pH 7.5), 150 mM NaCl, 10% glycerol, 1 mM DTT, indicated concentrations of GST-yCTD or GST-yCTD variants, and indicated concentration of dextran. Each solution was prepared in 96 well plate (Thermo Scientific) and absorbance at 350 nm readings were taken in a plate reader (Tecan) using default Absorbance settings. For kinase and phosphatase treatment assay, absorbance at 350 nm was measured at 30 °C with continuous shaking.
MALDI-TOF mass spectrometry and EMSA
Five microliters of phosphorylated GST-yCTD samples were taken out from reaction batch for measuring molecular weight. The samples were desalted over Ziptip C18 resins (MilliporeSigma) using standard protocols. Mass spectrometric analysis of phosphorylated GST-yCTD was carried out in an AB Voyager-DE PRO MALDI-TOF (Brunker Corporation) with the 1:1 DHB matrix (Thermo Fisher Scientific).
GST-yCTD samples treated with kinases were analyzed by mobility shift assays. Five microliters of GST-yCTD samples were taken from kinase reactions, then loaded and separated on 8% denaturing Tris-glycine polyacrylamide gels and stained with Coomassie solution. Stained gels were imaged with Gel Doc XR+ Gel Documentation System (Biorad).
In vitro confocal microscopy
GST-yCTD samples were fluorescently labeled as described in above. Fluorescently labeled GST-yCTD samples were mixed with crowding reagent (dextran) and buffer. Then, 10 μL of samples were directly loaded onto glass slides, covered with 22 mm coverslips. Fluorescent images were acquired with a Nikon W1 Spinning Disk Confocal Microscope with either 60× objective (water immersion) or 100x objective (oil immersion). Fluorescent images were processed using NIS-Elements Viewer (Nikon).
Differential interference contrast (DIC) microscopy
Wild-type GST-yCTD samples or GST-yCTD variants were mixed with various amounts of dextran, and droplet formation was monitored by DIC microscopy. Ten microliters of each sample were applied to slide glass and covered with 22 mm coverslip DIC images were acquired with Nikon eclipse Ni Compound Microscope with 60× objective. DIC images were processed using NIS-Elements Viewer (Nikon).
In vivo immunofluorescence and microscopy
U2OS cells were transfected with Pin1-YFP, SRSF2-Flag, SCAF6-HA, Ssu72-HA or RPB1aAmr-YFP plasmids for 24 h before harvest, fixed in 4% paraformaldehyde in PBS for 10 min at room temperature, permeabilization with PBS containing 0.1% Triton X-100 for 10 min at room temperature, then blocked in 2% bovine serum albumin (BSA) in PBS for 1 h, and incubated sequentially with different primary antibodies for overnight at 4 °C and Alexa-labeled secondary antibodies (Invitrogen: A11006, A11077 and A11011, 1:1000 dilution; Jackson: 115-545-062, 115-585-062 and 111-545-003, 1:500 dilution) for 1 h at room temperature with extensive washing. Slides were stained with DAPI (Sigma, MBD0015) and mounted with anti-Fade fluorescence mounting media (Abcam, ab104135). Immunofluorescence images were obtained and analyzed using the Zeiss LSM710 confocal microscope and ImageJ software.
Fluorescence recovery after photobleaching (FRAP)
Pin1-YFP was transfected in U2OS cells for 36 h and the condensates were photobleached and imaged with a 405 nm laser using Zeiss LSM710 confocal microscope. At each time point, fluorescence intensity within the bleaching spot was divided by the intensity of a neighboring unbleached area of the same size to correct the changes.
Fluorescence polarization
CTD peptides with double repeats were labeled with fluorescein isothiocyanate (FITC) and purchased from Biomatik. Protein and peptide concentrations were determined according to their absorbance at 280 nm. Fluorescence polarization values were collected on a Tecan F200 plate reader in buffer (50 mM Tris pH 8.0, 300 mM NaCl) at room temperature. Samples were excited with vertically polarized light at 485 nm and at an emission wavelength of 535 nm. Recombinant Pin1, RPRD1B-CID or SCAF4-CID proteins were titrated into a reaction mixture containing buffer supplemented with 100 nM of FITC-peptide. Measurements were taken in triplicates and the experimental binding isotherms were analyzed in GraphPad Prism v8 using a 1:1 binding mode to obtain Kd values.
Chromatin immunoprecipitation (ChIP) and ChIP-Sequencing
HEK 293 T cells were seeded in 15 cm dishes and transfected YFP-RPB1_52xCTD, 26xCTD and mutant plasmids. After 24 h transfection, cells were fixation with 1% formaldehyde for 8 min at room temperature. Crosslinking was quenched with 0.125 M glycine for 5 min. Cells were successively lysed in lysis buffer LB1 (50 mM HEPES-KOH, pH 7.5, 140 mM NaCl, 1 mM EDTA, 10% glycerol, 0.5% NP-40, 0.25% Triton X-100, 1× PI), LB2 (10 mM Tris-HCl, pH 8.0, 200 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, 1× PI) and LB3 (10 mM Tris-HCl, pH 8.0, 100 mM NaCl, 1 mM EDTA, 0.5 mM EGTA, 0.1% Na-deoxycholate, 0.5% N-lauroylsarcosine, 1×PI). Chromatin was sonicated to an average size of ~200–500 bp using Q800R3 Sonicator (30 s on and 30 s off for 25 min). A total of 5 μg of GFP antibody (Proteintech, 50430-2-AP) that was pre-mixed in a 50 μL volume of Dynabeads protein A (Invitrogen) was added to each sonicated chromatin sample with 1% Triton X-100 and incubated overnight at 4 °C. The chromatin-bound beads were washed two times with low salt wash buffer (0.1% Na Deoxycholate, 1% Triton X-100, 1 mM EDTA, 50 mM HEPES pH 7.5, 150 mM NaCl), once with high salt wash buffer (0.1% Na Deoxycholate, 1% Triton X-100, 1 mM EDTA, 50 mM HEPES pH 7.5, 500 mM NaCl), once with LiCl wash buffer (250 mM LiCl, 0.5% NP-40, 0.5% Na-Deoxycholate, 1 mM EDTA, 10 mM Tris-Cl pH 8.0) and twice in TE buffer. The chromatin was reverse crosslinked overnight at 65 °C with shaking at 750 rpm in cross-linking buffer (1% SDS and 0.1 M NaHCO3). After DNA extraction using phenol-chloroform, the DNA was resuspended in 10 mM Tris-HCl pH 8.0. The purified DNA was subjected to qPCR to confirm target region enrichment before moving on to deep sequencing library preparation. For sequencing, the extracted DNA was used to construct the ChIP-Seq library using the NEBNext Ultra II DNA Library Prep Kit, followed by sequencing with an Illumina NovaSeq 6000 system by Novogene.
RNA isolation, library preparation, and RNA-Sequencing
Total RNA was isolated from HEK 293 T cells (at least ~106 cells/sample) using DirectZol RNA Miniprep kit (Zymo Research, Irvine, CA, product number #R2050). Poly (A) enrichment RNA-Seq was performed by Novogene, mRNA was purified from total RNA using poly-T oligo-attached magnetic beads. After fragmentation, the first strand cDNA was synthesized using random hexamer primers, followed by the second strand cDNA synthesis using dUTP for directional library. The library was checked with Qubit and real-time PCR for quantification and bioanalyzer for size distribution detection. Quantified libraries will be pooled and sequenced on NovaSeq 6000 instrument (paired-end 2 × 150, 100 cycles). A minimum number of reads was set to 40 × 106 per sample.
qPCR
A total of 0.5 µg RNA was used for reverse transcription using the AzuraQuant™ cDNA Synthesis Kit, #AZ-1995 (Azura, Raynham, MA, USA) using manufacturer’s guidelines. Relative expression of alternatively spliced isoforms was measured using AzuraQuant™ Green Fast qPCR Mix, Lo-Rox (Azura) and normalized to overall transcript expression in the sample using 2-dCt method. Amplification was performed using the ViiA 7 Real-Time PCR System (Applied Biosystems, Waltham, MA). Specificity of amplification was controlled with melting curves / primer efficiency calculation. Primer sequences (designed to span exon-exon juctions or to be separated by a relatively large intron) and qPCR conditions are shown in Supplementary Table 2.
Analyses of ChIP-Seq data and calculation of Pausing Index (PI)
Initial quality assessment showed high library complexities with low level of duplication. Adapter sequences and low-quality read ends were trimmed off by TrimGalore! v.0.6.7 with default parameters. Paired-end reads were aligned to human reference genome, GRCh38 version, using Bowtie2 v.2.4.569 with default parameters. Mapping stats confirmed high alignment rates (>80–90% of reads aligned concordantly exactly 1 time). Next, coverage bigwig files normalized by Input (IgG-control) were generated out of bam files for every sample using log2 of the number of reads ratio (mapq > 10)70. DeepTools v.3.5.1 were also used to prepare score matrices and plotting metagene Pol II ChIP-Seq profiles (over subset of protein-coding genes, n = 19,984 regions). Next, CTD_26x and CTD_52× filtered reads (mapq > 10) were used to call ‘broad’ peaks (p < 0.005) with MACS2 peak caller v.2.2.7.1 keeping one duplicate tag at the exact same location (--keep-dup 1). Obtaining consensus peaksets and occupancy analysis was performed using ‘DiffBind’ v.3.0.15 pipeline71 in R. The absolute majority of consensus peaks in both conditions (CTD_26x/52x) reached IDR-threshold (IDR/irreproducible discovery rate <0.05): 84% and 89% peaks, respectively. Published ChIP-Seq data on PIN1 and pSer5 binding were accessed at GSE231126 and GSM4442247, respectively, and re-analyzed using provided supplementary files (bigWig, bed). ChIP-Seq signal from provided bigWig files was used to prepare score matrices followed by metagene profile/heatmap plotting (hg38 or hg19 genome version, 50-bp or 10-bp bins where applicable). IDR-thresholded peaks were annotated with ‘Chippeakanno’ R package.
To calculate Pausing Index (PI) as the measure of promoter-proximal pausing of RNA Pol II under CTD_26x vs CTD_52x condition, we used Input-normalized read count files in bigwig format. PI was defined as follows:
Where TSSR (transcription start site region) is (−50 bp to +300 bp around TSS), and the gene body is (+300 bp downstream of the TSS to +3 kb past the TES). L1 and L2 are the corresponding lengths of the regions48. The read densities were calculated using Bwtool (https://github.com/CRG-Barcelona/bwtool, “summary” function) which sums up signal in normalized bigwig files over the defined genomic regions in a bed file containing corresponding to “numerator” and “denominator” coordinates of human protein-coding genes (n = 19,984) derived from gencode hg38 annotation.gtf. Further analysis was conducted in R, where the genes were ranked depending on their average PI across the conditions and then clustered into four groups: G0 cluster with PI = 0, G1 cluster with PI < 25% quartile, G2 cluster with 25% <PI < 75% quartile; and G3 cluster with PI > 75% quartile (most paused genes have PI ≥ 248) (https://github.com/tailana703/CTD_length_PolII_pausing_alternative_splicing). ChIP-Seq data was deposited in GEO under the accession number GSE252261 and GSE252258.
Analyses of RNA-Seq data and alternative splicing events (ASE)
As with raw ChIP-Seq reads, adapter sequences and low-quality read ends were trimmed off by TrimGalore! v.0.6.7 with default parameters. Trimmed reads were aligned to human reference genome, GRCh38 version, using HISAT2 fast aligner v.2.2.1 with default parameters, except Reverse (RF) --rna-strandedness. Gencode v38 gtf file was used as annotation gtf. Lastly, mapped fragments were quantified by featureCounts v.2.0.1 in Galaxy72.
Differential expression in CTD_26x vs CTD_52x was analyzed using raw unnormalized counts in DESeq2 v.1.30.1 in R; genes with adjusted p-value < 0.05 and |log2FC| > 0.58 were considered as differentially expressed73. rMATS turbo v.4.1.2 was employed for detection of alternatively spliced events upon CTD_26x vs. WT-CTD_52x52. As input files for rMATS, we used alignment.bam files from HISAT2 mapper (two biological replicates per condition) and gencode v38 annotation gtf. Downstream analysis of rMATS output files containing JCEC counts (Junction Counts and Exon Coverage) was performed in SpliceTools suite with hg38.fasta and.bed12 annotation53. Gene lengths’ comparison was done using custom script in R. Overrepresentation analysis of gene clusters (gene ontology) was performed using Bioconductor R package ‘clusterProfiler’ v.3.18.1 with org.Hs.eg.db as a genome-wide annotation for human genes. We used ‘BH’ (Benjamini–Hochberg) method for adjusting p-values and a cut-off of q < 0.05 to report significantly enriched terms. RNA-Seq data was deposited in GEO under the accession number GSE252260.
Statistics and reproducibility
Statistical analyses were performed using Orgin Pro 9.1, RStudio v.4.0.5 and GraphPad Prism 8.0. Two-tailed or one-tailed (where applicable), independent sample t-test was used to compare the two groups. Non-parametrically distributed data were compared using Kruskal-Wallis test and Wilcoxon rank test/post-hoc pairwise tests. p < 0.05 was considered statistically significant. BH p-value adjustment was performed where applicable. For cell survival analysis, two-way ANOVA was performed to determine p-values. Correlations were assessed using two-tailed Pearson r coefficients. All statistical source data and raw blot data can be found in Source Data.
For in vitro biochemistry experiments (Figs. 1a, d, f, 2b–h and S1b, c, f, g, S2b, i, j, k), three independent experiments containing the indicated mix was imaged and at least five independent viewpoints were acquired for each slide. For cell imaging experiments (Figs. 3a, c, f, h and 5a and S3a, d, S4g, h), three independent experiments were conducted, and at least five independent viewpoints were acquired for each slide. For the ChIP-Seq and RNA-Seq experiments, two replicates were used for each condition.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The data supporting the findings of this study are available from the corresponding authors upon request. The sequencing data generated in this study have been deposited in GEO under the accession numbers: GSE252261, GSE252258 (ChIP-Seq, HEK293T), GSE252260 (RNA-Seq, HEK293T). Source data are provided with this paper.
Code availability
We have deposited the code we used to analyze our data in https://github.com/tailana703/CTD_length_PolII_pausing_alternative_splicing.
References
Heidemann, M., Hintermair, C., Voss, K. & Eick, D. Dynamic phosphorylation patterns of RNA polymerase II CTD during transcription. Biochim Biophys. Acta 1829, 55–62 (2013).
Jeronimo, C., Collin, P. & Robert, F. The RNA polymerase II CTD: the increasing complexity of a low-complexity protein domain. J. Mol. Biol. 428, 2607–2622 (2016).
Corden, J. L. RNA polymerase II C-terminal domain: tethering transcription to transcript and template. Chem. Rev. 113, 8423–8455 (2013).
West, M. L. & Corden, J. L. Construction and analysis of yeast RNA polymerase II CTD deletion and substitution mutations. Genetics 140, 1223–1233 (1995).
Chapman, R. D. et al. Transcribing RNA polymerase II is phosphorylated at CTD residue serine-7. Science 318, 1780–1782 (2007).
Harlen, K. M. et al. Comprehensive RNA polymerase II interactomes reveal distinct and varied roles for each phospho-CTD residue. Cell Rep. 15, 2147–2158 (2016).
Harlen, K. M. & Churchman, L. S. Subgenic Pol II interactomes identify region-specific transcription elongation regulators. Mol. Syst. Biol. 13, 900 (2017).
Buratowski, S. The CTD code. Nat. Struct. Biol. 10, 679–680 (2003).
Ho, C. K. & Shuman, S. Distinct roles for CTD Ser-2 and Ser-5 phosphorylation in the recruitment and allosteric activation of mammalian mRNA capping enzyme. Mol. Cell 3, 405–411 (1999).
Lu, F., Portz, B. & Gilmour, D. S. The C-terminal domain of RNA polymerase II is a multivalent targeting sequence that supports Drosophila development with only consensus heptads. Mol. Cell 73, 1232–1242.e1234 (2019).
Guo, Y. E. et al. Phosphorylation regulates a switch between transcriptional and splicing condensates. Nature 572, 543–548 (2019).
Boehning, M. et al. RNA polymerase II clustering through carboxy-terminal domain phase separation. Nat. Struct. Mol. Biol. 25, 833–840 (2018).
Portz, B. & Shorter, J. Switching condensates: The CTD code goes liquid. Trends Biochem. Sci. 45, 1–3 (2020).
Kwon, I. et al. Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155, 1049–1060 (2013).
Cho, W. K. et al. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 361, 412–415 (2018).
Appel, L. M. et al. PHF3 regulates neuronal gene expression through the Pol II CTD reader domain SPOC. Nat. Commun. 12, 6078 (2021).
Lu, H. et al. Phase-separation mechanism for C-terminal hyperphosphorylation of RNA polymerase II. Nature 558, 318–323 (2018).
Palumbo, R. J. et al. Coevolution of the Ess1-CTD axis in polar fungi suggests a role for phase separation in cold tolerance. Sci. Adv. 8, eabq3235 (2022).
Quintero-Cadena, P., Lenstra, T. L. & Sternberg, P. W. RNA Pol II length and disorder enable cooperative scaling of transcriptional bursting. Mol. Cell 79, 207–220.e208 (2020).
Monahan, Z. et al. Phosphorylation of the FUS low-complexity domain disrupts phase separation, aggregation, and toxicity. EMBO J. 36, 2951–2967 (2017).
Tsang, B. et al. Phosphoregulated FMRP phase separation models activity-dependent translation through bidirectional control of mRNA granule formation. Proc. Natl Acad. Sci. USA 116, 4218–4227 (2019).
Kim, T. H. et al. Phospho-dependent phase separation of FMRP and CAPRIN1 recapitulates regulation of translation and deadenylation. Science 365, 825–829 (2019).
Hsin, J. P. & Manley, J. L. The RNA polymerase II CTD coordinates transcription and RNA processing. Genes Dev. 26, 2119–2137 (2012).
Tee, W. W., Shen, S. S., Oksuz, O., Narendra, V. & Reinberg, D. Erk1/2 activity promotes chromatin features and RNAPII phosphorylation at developmental promoters in mouse ESCs. Cell 156, 678–690 (2014).
Di Vona, C. et al. Chromatin-wide profiling of DYRK1A reveals a role as a gene-specific RNA polymerase II CTD kinase. Mol. Cell 57, 506–520 (2015).
Mayfield, J. E. et al. Mapping the phosphorylation pattern of Drosophila melanogaster RNA polymerase II carboxyl-terminal domain using ultraviolet photodissociation mass spectrometry. ACS Chem. Biol. 12, 153–162 (2017).
Ramani, M. K. V. et al. Structural motifs for CTD kinase specificity on RNA polymerase II during eukaryotic transcription. ACS Chem. Biol. 15, 2259–2272 (2020).
Mayfield, J. E. et al. Tyr1 phosphorylation promotes phosphorylation of Ser2 on the C-terminal domain of eukaryotic RNA polymerase II by P-TEFb. Elife 8, e48725 (2019).
Xiang, K. et al. Crystal structure of the human symplekin-Ssu72-CTD phosphopeptide complex. Nature 467, 729–733 (2010).
Irani, S. et al. Structural determinants for accurate dephosphorylation of RNA polymerase II by its cognate C-terminal domain (CTD) phosphatase during eukaryotic transcription. J. Biol. Chem. 294, 8592–8605 (2019).
Luo, Y. et al. Novel modifications on C-terminal domain of RNA polymerase II can fine-tune the phosphatase activity of Ssu72. ACS Chem. Biol. 8, 2042–2052 (2013).
Zhang, Y. et al. Determinants for dephosphorylation of the RNA polymerase II C-terminal domain by Scp1. Mol. Cell 24, 759–770 (2006).
Liu, S. et al. Structure of the yeast spliceosomal postcatalytic P complex. Science 358, 1278–1283 (2017).
Verdecia, M. A., Bowman, M. E., Lu, K. P., Hunter, T. & Noel, J. P. Structural basis for phosphoserine-proline recognition by group IV WW domains. Nat. Struct. Biol. 7, 639–643 (2000).
Lu, P. J., Zhou, X. Z., Liou, Y. C., Noel, J. P. & Lu, K. P. Critical role of WW domain phosphorylation in regulating phosphoserine binding activity and Pin1 function. J. Biol. Chem. 277, 2381–2384 (2002).
Ranganathan, R., Lu, K. P., Hunter, T. & Noel, J. P. Structural and functional analysis of the mitotic rotamase Pin1 suggests substrate recognition is phosphorylation dependent. Cell 89, 875–886 (1997).
Zhang, Y. et al. Structural basis for high-affinity peptide inhibition of human Pin1. ACS Chem. Biol. 2, 320–328 (2007).
Venkat Ramani, M. K., Yang, W., Irani, S. & Zhang, Y. Simplicity is the ultimate sophistication-crosstalk of post-translational modifications on the RNA polymerase II. J. Mol. Biol. 433, 166912 (2021).
Gregersen, L. H. et al. SCAF4 and SCAF8, mRNA Anti-terminator proteins. Cell 177, 1797–1813.e1718 (2019).
Ni, Z. et al. RPRD1A and RPRD1B are human RNA polymerase II C-terminal domain scaffolds for Ser5 dephosphorylation. Nat. Struct. Mol. Biol. 21, 686–695 (2014).
Lafontaine, D. L. J., Riback, J. A., Bascetin, R. & Brangwynne, C. P. The nucleolus as a multiphase liquid condensate. Nat. Rev. Mol. Cell Biol. 22, 165–182 (2021).
Schmidt, H. B. & Rohatgi, R. In vivo formation of vacuolated multi-phase compartments lacking membranes. Cell Rep. 16, 1228–1236 (2016).
Fare, C. M., Villani, A., Drake, L. E. & Shorter, J. Higher-order organization of biomolecular condensates. Open Biol. 11, 210137 (2021).
Lyons, H. et al. Functional partitioning of transcriptional regulators by patterned charge blocks. Cell 186, 327–345 e328 (2023).
Moreno, R. Y. et al. Distinctive interactomes of RNA polymerase II phosphorylation during different stages of transcription. iScience 26, 107581 (2023).
Flores-Solis, D. et al. Driving forces behind phase separation of the carboxy-terminal domain of RNA polymerase II. Nat. Commun. 14, 5979 (2023).
Zhang, J. & Corden, J. L. Identification of phosphorylation sites in the repetitive carboxyl-terminal domain of the mouse RNA polymerase II largest subunit. J. Biol. Chem. 266, 2290–2296 (1991).
Day, D. S. et al. Comprehensive analysis of promoter-proximal RNA polymerase II pausing across mammalian cell types. Genome Biol. 17, 120 (2016).
Sawicka, A. et al. Transcription activation depends on the length of the RNA polymerase II C-terminal domain. EMBO J. 40, e107015 (2021).
Nguyen, V. T. et al. In vivo degradation of RNA polymerase II largest subunit triggered by alpha-amanitin. Nucleic Acids Res. 24, 2924–2929 (1996).
Gao, T. & Qian, J. EnhancerAtlas 2.0: an updated resource with enhancer annotation in 586 tissue/cell types across nine species. Nucleic Acids Res. 48, D58–D64 (2020).
Shen, S. et al. rMATS: robust and flexible detection of differential alternative splicing from replicate RNA-Seq data. Proc. Natl Acad. Sci. USA 111, E5593–E5601 (2014).
Flemington, E. K. et al. SpliceTools, a suite of downstream RNA splicing analysis tools to investigate mechanisms and impact of alternative splicing. Nucleic Acids Res. 51, e42 (2023).
Baker, S. J., Poulikakos, P. I., Irie, H. Y., Parekh, S. & Reddy, E. P. CDK4: a master regulator of the cell cycle and its role in cancer. Genes Cancer 13, 21–45 (2022).
Goldenson, B. & Crispino, J. D. The aurora kinases in cell cycle and leukemia. Oncogene 34, 537–545 (2015).
Dominguez, D. et al. An extensive program of periodic alternative splicing linked to cell cycle progression. Elife 5, e10288 (2016).
Galej, W. P. et al. Cryo-EM structure of the spliceosome immediately after branching. Nature 537, 197–201 (2016).
Plaschka, C., Lin, P. C., Charenton, C. & Nagai, K. Prespliceosome structure provides insights into spliceosome assembly and regulation. Nature 559, 419–422 (2018).
Wan, R., Bai, R., Yan, C., Lei, J. & Shi, Y. Structures of the catalytically activated yeast spliceosome reveal the mechanism of branching. Cell 177, 339–351.e313 (2019).
Zhan, X., Yan, C., Zhang, X., Lei, J. & Shi, Y. Structure of a human catalytic step I spliceosome. Science 359, 537–545 (2018).
Riback, J. A. et al. Stress-triggered phase separation is an adaptive, evolutionarily tuned response. Cell 168, 1028–1040.e1019 (2017).
Komarnitsky, P., Cho, E. J. & Buratowski, S. Different phosphorylated forms of RNA polymerase II and associated mRNA processing factors during transcription. Genes Dev. 14, 2452–2460 (2000).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell 175, 1842–1855.e1816 (2018).
Chen, X. et al. Structures of the human mediator and mediator-bound preinitiation complex. Science 372, eabg0635 (2021).
Abdella, R. et al. Structure of the human Mediator-bound transcription preinitiation complex. Science 372, 52–56 (2021).
Schilbach, S., Wang, H., Dienemann, C. & Cramer, P. Yeast PIC-Mediator structure with RNA polymerase II C-terminal domain. Proc. Natl Acad. Sci. USA 120, e2220542120 (2023).
Feric, M. et al. Coexisting liquid phases underlie nucleolar subcompartments. Cell 165, 1686–1697 (2016).
Gibson, B. A. et al. Organization of chromatin by Intrinsic and regulated phase separation. Cell 179, 470–484.e421 (2019).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ramirez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Ross-Innes, C. S. et al. Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481, 389–393 (2012).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
We thank Dr. Jack E Dixon and Tanja Mittag for advice on the manuscript and the National Institutes of Health (R01GM104896, R01GM125882 and R35GM148356 to Y.J.Z.) for supporting our research. We also thank funds from L. Leon Campbell Professorship. Thanks also to technical assistance by the core facilities of the University of Texas at Austin. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
Q.Z. and W.K. carried out most experiments and helped with experimental design. S.P. conducted the bioinformatics analysis, J.M. and B.P. helped with experimental design and discussions. Y.J. Z. conceived the study and experimental design, Y.J. Z., Q.Z., W.K. and B.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Maria Carmo-Fonseca and the other, anonymous, reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Q., Kim, W., Panina, S.B. et al. Variation of C-terminal domain governs RNA polymerase II genomic locations and alternative splicing in eukaryotic transcription. Nat Commun 15, 7985 (2024). https://doi.org/10.1038/s41467-024-52391-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-52391-6
- Springer Nature Limited