Abstract
This study describes “lobular-like invasive mammary carcinomas” (LLIMCas), a group of low- to intermediate-grade invasive mammary carcinomas with discohesive, diffusely infiltrative cells showing retained circumferential membranous immunoreactivity for both E-cadherin and p120. We analyzed the clinical-pathologic features of 166 LLIMCas compared to 104 classical invasive lobular carcinomas (ILCs) and 100 grade 1 and 2 invasive ductal carcinomas (IDCs). Tumor size and pT stage of LLIMCas were intermediate between IDCs and ILCs, and yet often underestimated on imaging and showed frequent positive margins on the first resection. Despite histomorphologic similarities to classical ILC, the discohesion in LLIMCa was independent of E-cadherin/p120 immunophenotypic alteration. An exploratory, hypothesis-generating analysis of the genomic features of 14 randomly selected LLIMCas and classical ILCs (7 from each category) was performed utilizing an FDA-authorized targeted capture sequencing assay (MSK-IMPACT). None of the seven LLIMCas harbored CDH1 loss-of-function mutations, and none of the CDH1 alterations detected in two of the LLIMCas was pathogenic. In contrast, all seven ILCs harbored CDH1 loss-of-function mutations coupled with the loss of heterozygosity of the CDH1 wild-type allele. Four of the six evaluable LLIMCas were positive for CDH1 promoter methylation, which may partially explain the single-cell infiltrative morphology seen in LLIMCa. Further studies are warranted to better define the molecular basis of the discohesive cellular morphology in LLIMCa. Until more data becomes available, identifying LLIMCas and distinguishing them from typical IDCs and ILCs would be justified. In patients with LLIMCas, preoperative MRI should be entertained to guide surgical management.
Similar content being viewed by others
Introduction
Invasive mammary carcinoma of no special type, commonly referred to as invasive ductal carcinoma (IDC), and the special subtype of invasive lobular carcinoma (ILC) represent the two most common types of invasive breast carcinomas1. While IDCs typically show varying degrees of duct formation, ILCs are characterized by discohesive tumor cells with single-file infiltrative growth patterns dispersed in the fibrous stroma2. The differences between IDC and ILC, from clinicopathological features to prognostic outcomes, have been extensively reported in the literature, sometimes with conflicting results3,4,5. More recently, attention has turned to the molecular and evolutionary differences between the two entities and their precursor lesions, laying the foundations for personalized management of breast cancers6,7,8,9,10. Even though the diagnosis of IDC versus ILC is usually straightforward, cases with ambiguous histomorphology are not uncommon. Immunohistochemical (IHC) assessment of E-cadherin with or without p120 and beta-catenin is often used to assist in the diagnosis of such cases.
E-cadherin is a transmembrane adhesion glycoprotein encoded by the CDH1 gene (16q22.1). P120 catenin is a tyrosine kinase substrate anchored to the internal domain of E-cadherin in a juxtamembranous fashion11,12,13. Characteristically, ILC harbors biallelic inactivation of the CDH1 gene, often through a combination of pathogenic loss-of-function mutations coupled with loss-of-heterozygosity (LOH) of the CDH1 wild-type allele7,14. When E-cadherin is absent or nonfunctional, p120 catenin undergoes redistribution from the cell membrane to the cytoplasm. Although practice patterns among subspecialists in breast pathology are not uniform, conventionally, lack of E-cadherin membranous expression coupled with diffuse, intense cytoplasmic p120 catenin expression is diagnostic of lobular lesions, whereas distinctively crisp, intense membranous positivity for both E-cadherin and p120 catenin is characteristic of ductal phenotype. In the past decade, emerging evidence has revealed that a non-functional E-cadherin might present as an aberrant (i.e., lack of strong membranous reactivity) yet visible pattern immunohistochemically15,16,17,18,19. Most reported cases with aberrant E-cadherin immunoreactivity, however, displayed a corresponding disruption of the cadherin-catenin complex15,17,18,19,20,21.
The 5th edition of the WHO classification of tumors for breast carcinoma does not recommend the use of immunohistochemistry (IHC) for the diagnosis of ILC2. However, in our experience, the interobserver agreement is limited in classifying ILC with ambiguous morphology without IHC. Findings from a recent study by Christgen et al.22 also supported our observation. In their study, 35 pathologists were asked to classify specimens (using 2 sets of cases—set A with H&E section only and set B with H&E and E-cadherin IHC) as non‐lobular breast carcinoma versus mixed breast carcinoma versus ILC. Pairwise interobserver agreement was moderate in set A (median κ = 0.58) and substantial in set B (median κ = 0.75, p < 0.001). Agreement with the reference diagnosis was substantial in set A (median κ = 0.67) and almost perfect in set B (median κ = 0.86, p < 0.001). The authors of this study concluded that subtyping of breast cancer as ILC achieves almost perfect agreement with a pre‐defined reference standard if the assessment is supported by E‐cadherin IHC. To improve the standardization of lobular carcinoma diagnosis, it has been our practice at Magee-Womens Hospital since 2004–2005 to confirm the first-time diagnosis of ILC by E-cadherin (along with p120) IHC staining. Either complete absence or “aberrant” reactivity for E-cadherin combined with intense cytoplasmic p120 catenin expression is required for a diagnosis of ILC. Reactivity for E-cadherin is considered “aberrant” if it is partially membranous, beaded, perinuclear dot-like, or cytoplasmic. In the event of an equivocal E-cadherin staining, concurrent cytoplasmic p120 staining supports the diagnosis of ILC, whereas circumferential membranous p120 reactivity is indicative of ductal immunophenotype. When different combinations of E-cadherin/p120 immunoprofile are present, a diagnosis of mixed IDC and ILC is justified. Our approach, although different from WHO recommendation, results in a more accurate classification and is the basis of this study.
We have observed in our daily practice a group of invasive carcinomas predominantly characterized by dissociated tumor cells with low-to-intermediate grade, uniform nuclei, and lobular-like growth patterns but display distinct membranous IHC staining for both E-cadherin and p120. We refer to such cases as “lobular-like invasive mammary carcinoma” (LLIMCa), but it is unclear whether this terminology is appropriate and if such cases harbor biallelic inactivation of CDH1, akin to ILCs. To our knowledge, no previous study has defined the clinical-pathologic characteristics or addressed the clinical behavior of LLIMCa, even though multiple prior studies, including those analyzing “IDC with lobular features”, in fact, examined mixed ductal and lobular carcinomas with heterogeneous E-cadherin expression23,24,25,26,27.
The current study aims to (1) analyze the clinical, radiological, and pathological characteristics and prognostic outcomes of LLIMCa, (2) compare the characteristics of LLIMCa with those of classical ILC and typical IDC of no special type, (3) assess representative LLIMCa cases for biallelic alterations of CDH1, and (4) perform an exploratory, hypothesis-generating analysis of the repertoire of somatic genetic alterations comparing representative LLIMCa and classical ILC cases.
Results
Morphologic and immunohistologic features
Consistent with the definition of LLIMCa we put forth (refer to Methods for details), this group of tumors was found to harbor predominantly discohesive, uniform, small- to intermediate-size nuclei and lobular-like dissociated growth pattern, presenting as individual cells in single-files and cords within the fibrous stroma. Neoplastic cells were occasionally arranged in a concentric “targetoid” pattern around benign ducts and lobules. Nested or trabecular patterns were infrequently observed. Cytologically the tumor cells tended to have slightly larger and more angulated nuclei than classical-type ILC. The histomorphologic features were reflected in the analyzed Nottingham scores. Signet ring cells were not prominent in any of the LLIMCa cases. Scattered intracytoplasmic vacuoles were seen in some cases of LLIMCa. Overall, LLIMCas were morphologically difficult or even impossible to classify definitively as either lobular or ductal on H&E sections alone (Fig. 1a, b). Immunohistochemically, all tumor cells demonstrated distinct and circumferential membranous expression of both E-cadherin and p120 catenin, characteristic of ductal phenotype (Fig. 1c, d). In contrast, ILCs were characterized by either complete loss or aberrant expression of E-cadherin, coupled with predominantly cytoplasmic p120 catenin (Fig. 2).
In addition, β-catenin stain was performed on all 14 cases (7 LLIMCas and 7 ILCs) randomly selected for the exploratory genomic analysis. All 7 cases of LLIMCa demonstrated membranous expression. All 7 cases of ILC revealed granular cytoplasmic uptake of β-catenin (Supplementary Figs. 1 and 2). A recent study identified a group of ILCs with N-terminal-deficient E-cadherin while preserving E-cadherin C-terminus28. To confirm whether LLIMCas in the present study were, in fact, ILCs with N-terminal-deficient yet C-terminal-conserved E-cadherin protein, we subjected the above 14 cases to IHC staining of antibodies against E-cadherin N-terminus. All 7 LLIMCas showed membranous staining, while all 7 ILCs were negative for N-terminal E-cadherin (Supplementary Fig. 3).
Clinical characteristics
A comparison of the clinical characteristics of LLIMCa (n = 166), IDC (n = 100), and ILC (n = 104) (Table 1) revealed that patients with LLIMCa presented at a slightly older mean age (60.6 years) compared to those with IDC (57 years, p = 0.035), but were similar in age to ILC patients (61.6 years, p = 0.493). During the study period, LLIMCa patients more often (81%) underwent breast-conserving segmental resection as the first surgical procedure, compared to ILC (63%, p = 0.003) and IDC (52%, p < 0.0001) patients. In contrast, a significantly higher number of LLIMCa (83%, p < 0.0001) and ILC (75%, p < 0.0001) patients received adjuvant radiation therapy compared to IDC patients (47%), who had a significantly higher number of total mastectomies and avoided subsequent radiation. There were no differences in adjuvant systemic endocrine therapy or chemotherapy among the three tumor types. Albeit not statistically significant, the proportions of patients with LLIMCa developing locoregional recurrence appeared to be intermediate between ILC and IDC. In contrast, ILC was associated with a significantly higher frequency of locoregional recurrence compared to IDC. No difference in the patterns of distant metastasis was observed between different tumor types in our study.
Pathological characteristics
A comparison of the pathologic features of LLIMCa, ILC, and IDC (Table 2) demonstrated that the mean tumor size of LLIMCas (1.9 cm) was intermediate between IDCs (1.55 cm, p < 0.012) and ILCs (2.7 cm, p < 0.0001). As a result, LLIMCas presented at a higher pT stage than IDCs but a lower pT stage than ILCs.
LLIMCas was more likely to have positive or close margins (37%) in the first surgical procedure compared to IDCs (19%, p = 0.003) but was similar to ILCs (35%, p = 0.261). LLIMCas were found to have a significantly higher rate of lymphovascular space invasion (27%) than ILCs (11%, p = 0.002) but were similar to IDCs (22%, p = 0.465). No differences in multifocality, number of tumor foci, tumor-associated calcifications, lymph node status, or pN stage were observed between any of the tumor types.
Within the group of nuclear grade 1 and 2 tumors, as designed in the current study, LLIMCas exhibited a higher Nottingham grade and score than IDCs and ILCs. A substantial number of IDC were Nottingham grade I (grade I: 46%, grade II: 54%) due to the presence of tubule formation, whereas LLIMCas (grade I: 7%, grade II: 93%, p < 0.0001) and ILCs (grade I: 18%, grade II: 82%, p < 0.0001) were more likely to be grade II due to the lack of tubules. Additionally, LLIMCas frequently had higher Nottingham grades and scores compared to ILCs (p = 0.005), owing to the larger nuclei and more frequent mitoses.
Pathological–radiological correlation of tumor size
Analysis of the pathologic and radiologic tumor sizes at the time of diagnosis (Table 3) revealed that LLIMCas had a similar mean tumor size (1.35 cm) as IDCs (1.5 cm, p = 0.202) on radiologic measurement, which was smaller than that of ILCs (1.7 cm, p = 0.01). The final tumor size of LLIMCas (1.9 cm) upon pathologic examination of the resection specimen was intermediate between that of IDCs (1.55 cm, p = 0.012) and ILCs (2.7 cm, p < 0.0001). The majority of both LLIMCas (71%) and ILCs (75%) revealed larger pathologic tumor size compared to radiologic tumor size, whereas IDCs showed an even distribution of pathologic tumor size rather consistent with radiologic size. As a result, the pathologic to radiologic tumor size ratio of LLIMCas (1.6) was similar to ILCs (1.8) but significantly higher than that of IDC (1.1, p < 0.0001), indicating that the pathologic tumor size of either LLIMCas or ILCs was much larger than the tumor size estimated by imaging.
Prognostic biomarkers
LLIMCa displayed similar profiles of estrogen receptor (ER), progesterone receptor (PR), and Ki67 compared to ILC and IDC (Table 4), even though the mean ER H-score of LLIMCas (248) was slightly higher than that of ILCs (231, p = 0.046). LLIMCas appeared to show a higher rate of HER2 positivity (10%) compared to ILCs (1%, p = 0.004) and IDCs (1%, p = 0.004) in the study population. However, only a small percentage (3 of 16, 1.8%) of the HER2-positive LLIMCas were IHC 3+; the others were IHC 2+ and amplified by FISH with low HER2 copies but HER2/CEP17 ratio crossing the 2 cut-offs. The low HER2 positivity rate in IDC was attributed to the exclusion of nuclear grade 3 tumors. Both LLIMCas and ILCs demonstrated significantly higher Magee Equation 2 (ME2) scores compared to IDC (Table 4). The ME2 score distribution, however, was similar between LLIMCas and ILCs (33% of cases with a score <18 for both LLIMCas and ILCs compared to 50% of cases with a score <18 for IDCs).
Long-term survival analysis
For all three groups of patients, the median follow-up time for survival analysis was 10 years or longer (LLIMCa: 130 months, range 12.3–186.6 months; ILC: 128.7 months, range 28.8–185.1 months; IDC: 119.2, range 7.3–168.1 months). In the Kaplan–Meier survival analysis, although LLIMCas appeared to harbor an intermediate survival between IDCs and ILCs, statistical significance was not reached (Fig. 3) in either separate or combined sets of comparisons (Supplementary Data sets). Lower tumor grade (grade I), lower pT stage (pT1), lower pN stage (pN0 + pN1), and lower ME2 score (<18) were associated with significantly improved recurrence-free survival (RFS), distant RFS (DRFS), and breast cancer-specific survival (BCSS) (Table 5, Fig. 4 for BCSS, and Supplementary Data sets). Lower pT stage (pT1), lower pN stage (pN0 + pN1), and lower ME2 score (<18) were associated with significantly improved overall survival (OS) (Table 5 and Supplementary Data sets). Multivariable Cox proportional hazard regression analysis showed statistically significant improved RFS, DRFS, OS, and BCSS associated with lower pT stage (pT1) and lower ME2 score (<18) (Supplementary Data sets).
Recurrence-free survival or RFS (a), distant recurrence-free survival or DRFS (b), overall survival or OS (c), and breast cancer-specific survival or BCSS (d). Statistical significance is not reached based on different histologic types, p-values reported using log-rank test. LLIMCa lobular-like invasive mammary carcinoma, ILC invasive lobular carcinoma, IDC invasive ductal carcinoma.
Somatic genetic alterations and mutation signatures
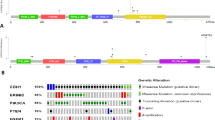
As an exploratory, hypothesis-generating analysis, we compared the repertoire of somatic genetic alterations of randomly selected 7 ILC and 7 LLIMCa cases from the current cohort (H&E and IHC stains shown in Supplementary Figs. 1 and 2) subjected to the Food and Drug Administration (FDA)-approved Memorial Sloan Kettering (MSK) Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT) targeted sequencing assay (Fig. 5, Supplementary Table 1). Copy number analysis revealed 16q LOH in all cases (Fig. 5, Supplementary Table 2), consistent with their ER-positive status and lower Nottingham grades. In addition to 16q LOH, this analysis revealed that all bona fide ILCs harbored CDH1 loss-of-function biallelic mutations (7/7), of which five were frameshift indel and two were splice site mutations uniformly coupled with LOH of the wild-type allele. Conversely, five of the seven LLIMCas did not harbor CDH1 mutations or genomic rearrangements. CDH1 alterations were identified in 2 LLIMCas: one (case DL09, Fig. 5) only harbored a subclonal CDH1 in-frame indel mutation coupled with LOH. This case mostly displayed membranous E-cadherin and p120 expression with focal areas of aberrant expression (cytoplasmic E-cadherin instead of membranous reactivity, supplementary fig. 4), which was identified only after careful re-review following the genomic analyses. The other CDH1-mutated LLIMCa (case DL11, Fig. 5) harbored a complex in-frame indel (i.e., one intronic deletion, one splice site mutation (p.X441_splice) and a frameshift indel (p.D443Gfs*10) in cis) with negligible impact on protein structure coupled with subclonal LOH. As expected, based on the genomic profile, DL11 displayed membranous expression of all four IHC markers (both N- and C-terminal-E-cadherin, p120, and β-catenin, Supplementary Figs. 1–3). Nevertheless, none of the seven LLIMCas harbored the loss-of-function mutations characteristic of ILCs, and none of the CDH1 alterations detected in the analyzed LLIMCas was biallelic. They are predicted not to affect gene function and are neither pathogenic nor clinically meaningful.
Non-synonymous somatic alterations identified in 7 ILCs and 7 LLIMCas detected by massively parallel sequencing targeting 515 cancer-related genes. Cases are shown in columns, and genes in rows. Clinicopathologic characteristics, including histology, E-cadherin and p120 staining by immunohistochemistry (IHC), tumor purity, CDH1 cancer cell fraction (CCF), mutational clonality, and status are depicted in phenobars (top). Somatic mutation types and copy number alterations are color-coded according to the legend. Loss of heterozygosity (LOH) is depicted by a square. HRD homologous recombination DNA repair defect, NA not available, LOH loss of heterozygosity.
Eleven of the 14 cases (78%) subjected to MSK-IMPACT had sufficient SNVs mutational signatures (Fig. 5, Supplementary Table 3) to be decomposed by SigMA, a machine learning-based algorithm optimized for mutational signature decomposition of formalin-fixed paraffin-embedded samples analyzed with targeted capture sequencing data. This analysis revealed that 75% (3/4) of ILCs displayed a dominant APOBEC mutational signature compared to 57% (4/7) of LLIMCas (p > 0.05, Fisher’s exact test). Homologous recombination deficiency-related signature (HRD) was found to be dominant in 25% (1/4) of ILCs, and 43% (3/7) of LLIMCas displayed mutational signatures attributed to aging.
Seven LLIMCas were subjected to CDH1 promoter methylation assessment by digital droplet PCR (ddPCR). This analysis revealed that four of the seven LLIMCas displayed CDH1 promoter methylation (Fig. 5, supplementary table 3), and these cases displayed slightly reduced E-cadherin expression (intensity of E-cadherin staining classified by one author (R.B.) prior to methylation testing), including the 2 LLIMCas that harbored CDH1 alterations (DL09 and DL11). Two of the remaining 3 LLIMCas were unmethylated, and in one case, the CDH1 methylation status was not accessible as the results did not meet the qualitative standards. The comparative analysis of the repertoire of somatic genetic alterations, patterns of gene copy number profiles, and mutational signatures present in LLIMCas and classical ILCs did not reveal any significant differences.
Discussion
Here we report a series of distinct breast carcinomas that we term as LLIMCa, characterized by low- to intermediate-grade uniform cells and exhibiting discohesive lobular-like growth patterns while maintaining intact adhesion molecule E-cadherin expression. In short, LLIMCa demonstrates some features similar to ILC but different from IDC, such as significantly larger pathologic tumor size than the size estimated on imaging, more frequent positive or close margins with breast-conserving surgery, higher Nottingham grade and score, and higher ME2 scores; some features similar to IDC such as higher rate of lymphovascular space invasion; and some features intermediate between ILC and IDC such as tumor size and pT stage. The study identifies several important findings worthy of close attention.
First, LLIMCas display a discohesive lobular-like morphologic pattern in the presence of membranous staining by E-cadherin (with both C- and N-terminal antibodies), p120, and β-catenin. A dysfunctional E-cadherin/catenin complex due to genetic or epigenetic biallelic inactivation of CDH1 is the defining hallmark of ILC. The dynamic alteration of the E-cadherin/p120 complex is thought to be responsible for the changes in cell motility and, accordingly, the dispersed infiltrative morphology of ILC6,7,11,21,29,30,31,32,33. In the present study, the distinct membranous staining for all four currently used IHC markers of E-cadherin/catenin complex in LLIMCa suggests that the phenotypic appearance of discohesive, diffuse growth pattern observed in this particular type of invasive carcinoma has occurred in the presence of an intact E-cadherin/p120 junctional complex. Correspondingly, in the 14 randomly selected cases subjected to the exploratory genomic analyses, CDH1 loss-of-function biallelic mutations are identified in all 7 bona fide ILC cases. In contrast, none of the 7 LLIMCa cases harbors the pathogenic CDH1 loss-of-function mutation. Rather, the only CDH1 alterations detected in two of the LLIMCas consists of in-frame indels that are predicted not to affect gene function and are neither pathogenic nor clinically meaningful. It appears that the mechanisms driving the typical appearance of LLIMCa are not dependent on CDH1 biallelic loss-of-function mutations or E-cadherin/catenin complex genomic alterations. However, CDH1 promoter methylation assessment reveals that four of the seven LLIMCas are methylated. This result raises the tantalizing hypothesis that the abnormal (slightly reduced) E-cadherin expression pattern observed in these four cases, which is not observed in the unmethylated LLIMCas (strong membranous expression), might be associated with epigenetic mechanisms. Hence, the assessment of CDH1 methylation status, using a sensitive ddPCR method and tissue microdissection to ensure high tumor purity, should be considered in the presence of even slightly reduced E-cadherin expression in cancers lacking bona fide biallelic CDH1 pathogenic mutations. Our findings warrant further whole-genome sequencing, epigenomic and RNA-sequencing analyses to define the mechanistic basis for the phenotypic features of LLIMCas. Another finding worth noting is the presence of a dominant ABOBEC mutational signature in 57% (4 of 7 tested) of LLIMCas. ABOBEC signatures are more frequent in ILCs compared to IDCs and have been suggested as a biomarker of resistance to endocrine therapy34. This should also be further investigated.
Second, LLIMCas often harbor significantly larger pathologic tumor size than the size estimated on imaging and frequently positive or close margins at the first surgical procedure. This appears to be associated with lobular-like tumor morphology despite the preservation of the E-cadherin/catenin complex. In the past decade, multiple studies and consensus guidelines have recommended preoperative MRI for optimal management of patients with ILC, given its high sensitivity and better accuracy of measuring tumor size and multifocal disease35,36,37,38,39. Similar findings have been reported in “IDC with lobular features” on core biopsy40. Moreover, the percentage of aberrant E-cadherin and p120 expression was found to be irrelevant to additional diseases detected on MRI in such patients41. Given the findings in our study, we contend that distinguishing LLIMCas from the usual type IDCs in the biopsy report may be required and that preoperative MRI should be considered to guide surgical management for this type of invasive breast carcinoma.
Third, all three types of invasive carcinomas in our study cohort show essentially no statistically significant difference in hormone receptor status or prognosis, including long-term survival. Most of the previous studies included tumors of all nuclear grades23,24, whereas ILCs consist of significantly more low- to intermediate-grade Luminal A-type tumors and are more often hormone receptor-positive compared to IDCs. As a result, the differences reported in prior studies might have been differences between low- to intermediate-grade versus high-grade tumors rather than between ductal and lobular cancers. In our study design, we excluded all nuclear grade 3 tumors. Therefore, since we have a distinctively uniform low- to intermediate-grade tumor population with different phenotypes, the similar hormone receptor status, and survival outcome might be a reflection of the inherent nature of low-intermediate grade, ER-positive Luminal A-like tumors regardless of histologic phenotypes. Genomic studies of the mixed ductal and lobular carcinomas revealed that all components in such cases were frequently clonally related, suggesting shared origins of a common neoplastic clone42. Moreover, several lines of studies revealed that ILC shared common genetic alterations with low-grade IDC and direct clonal divergence from the ductal to the lobular phenotype occurred late in tumor evolution, where the aberrant E-cadherin expression appeared to be a key distinguishing switch30,43,44. Hence, the multistep model of breast cancer progression suggests that ILCs or tumors exhibiting lobular-like growth patterns may arise from the low-grade, ER-positive “ductal” pathway10,42,43,44,45,46. Whether LLIMCa represents an intermediate form in the phenotypic switch from ductal to lobular type is yet to be determined.
Lastly, although an accurate diagnosis of LLIMCa is required for proper pre-surgical evaluation and local therapy, the prognosis of these tumors remains defined by the traditional prognostic factors as well as the multivariable model ME2. Our survival analysis fails to show statistically significant differences when LLIMCa is compared to ILC or IDC, either by itself or in combined groups with IDC or ILC. In contrast, the traditional prognostic markers are associated with significantly improved survival. Interestingly, the ME2 score <18 is associated with significant improvement in all four survival measurements (RFS, DRFS, OS, and BCSS). Since ME2 only uses readily available histopathologic and IHC data from pathology reports at no additional cost, its use should be encouraged in routine practice.
Our study does have some limitations. The study cohort is retrospectively extracted from a single institution, a 6-year period archive, during which time we started routine E-cadherin/p120 stains on invasive tumors with lobular-like growth patterns. The retrospective nature and relatively small sample size could be the potential basis for the non-significant clinicopathologic and survival observations. In addition, only a limited number of cases are sequenced for the pilot molecular analyses. Ideally, multicenter prospective outcome studies along with whole-genome sequencing, epigenomic and RNA-sequencing analyses to define the mechanistic basis for the phenotypic features of LLIMCas would be useful to corroborate the findings.
Despite these limitations, we present a series of LLIMCas with morphologic features mimicking those of ILCs, but maintaining immunophenotypic membranous E-cadherin expression characteristic of IDCs, and typically spared of the pathogenic CDH1 loss-of-function gene mutations. Their morphologic discohesion could be partly explained by CDH1 promoter methylation in some but not all cases. One could potentially argue to classify all tumors with lobular-like growth patterns as ILCs regardless of E-cadherin expression in the current practice setting of breast cancer treatment. Yet, the interobserver agreement for the diagnosis of ILC without IHC is moderate at best. The holy grail of understanding different types of breast carcinoma is to develop more targeted and personalized clinical treatment; germane to this endeavor is the correct cataloging of distinct phenotypes in a systematic manner. We would contend that to facilitate future investigations in elucidating ductal versus lobular phenotypes at molecular, evolutional, functional, and therapeutic levels, accurately identifying LLIMCas and separating LLIMCas from the typical IDCs or ILCs would be justified. From a patient management standpoint, we suggest that preoperative MRI be entertained to guide the surgical management of patients with LLIMCa.
Materials and methods
Defining LLIMCa
Lobular-like IDCs (LLIMCas) in the current study were defined as invasive mammary carcinomas consisting of uniform discohesive cells with low to intermediate grade nuclei, dispersed growth pattern, and rare to absent tubule formation while maintaining moderate to strong uniform circumferential membranous reactivity for E-cadherin and p120 throughout the tumor.
Case selection and IHC studies
The study protocol was approved by the University of Pittsburgh Institutional Review Board with a waiver of the informed consent. All invasive mammary carcinomas with ambiguous histomorphology for ductal or lobular differentiation and, therefore, with IHC C-terminus E-cadherin stain (Clone: 36; Catalog # 790-4497; Vendor: Ventana, Tucson, AZ; Dilution: ready to use [RTU]; Pre-treatment: CC1-S 64 min; Detection: Ultraview; Staining platform: Ventana Benchmark Ultra) performed were extracted from a 6-year (2004–2009) archive at UPMC Magee-Womens Hospital. Among them, the cases from 2007 to 2009 also had concurrent p120 stain (Clone: 98; Catalog # 610134; Vendor: BD Biosciences, Franklin Lakes, NJ; Dilution: 1:200; Pre-treatment: CC1-S; Detection: Ultraview; Staining platform: Ventana Benchmark Ultra) performed. Cases were classified as LLIMCas only if they displayed the criteria mentioned previously. In contrast, cases were classified as ILCs if they displayed absent and/or aberrant E-cadherin staining along with cytoplasmic p120 staining. Fourteen cases submitted for sequencing were also stained for beta-catenin (Clone: B-catenin-1; Catalog # M3539, Vendor: Dako, Santa Clara, CA; Dilution: 1:250; Pre-treatment: CC1 24 minutes; Detection: Optiview; Staining platform: Ventana Benchmark Ultra) and N-terminus E-cadherin antibody (Clone: 36B5; Catalog # PA0387, Vendor: Leica Biosystems, Deer Park, IL; Dilution: RTU; Pre-treatment: ER2 20 min; Detection: DAB polymer refine; Staining platform: Leica Bond III).
The following cases were excluded from the extracted cohort to allow for more homogeneous comparison among different tumor groups and more accurate assessment of clinical outcomes: (1) mixed ductal and lobular carcinomas with either mixed components or indeterminate variable E-cadherin and p120 staining patterns; (2) high grade (nuclear grade 3) carcinomas; (3) tubulolobular carcinomas; (3) microinvasive carcinomas (≤1 mm); (4) concurrent multifocal tumor with different morphology; (5) patients with neoadjuvant treatment; (6) patients with previous cancer diagnosis; (7) cases with no in-house primary surgical resection or incomplete follow-up information.
Ultimately, 166 cases of LLIMCa and 104 cases of ILC were identified. Additionally, 100 cases of grade 1 or 2 typical IDC (carcinoma, no special type) were retrieved over the same period for comparison analysis.
In addition, β-catenin stain (Clone: β-catenin-1; Vendor: Agilent (Dako), Santa Clara, CA; Dilution: 1:250; Pre-treatment: CC1 24’; Detection: OptiView; Staining platform: Ventana Benchmark Ultra) and N-terminal-E-cadherin stain (Clone: 36B5; Vendor: Leica, Deer Park, IL; Dilution: RTU; Pre-treatment: ER2, 20’; Detection: DAB Polymer Refine; Staining platform: Leica Bond III) were performed on the 14 LLIMCa and ILC cases randomly selected for the exploratory genomic tests.
Tissue preparation and DNA extraction
Ten 8 μm-thick sections from each representative formalin-fixed paraffin-embedded (FFPE) tumor and matched normal tissue blocks of 7 LLIMCas and 7 classical ILCs were stained with nuclear fast red and subjected to microdissection using a sterile needle under a stereomicroscope (Olympus SZ61) to enrich tumor cell content, as previously described47,48. Genomic DNA was extracted from the tumor and matched normal tissue using the QIAamp DNA FFPE Tissue Kit (Qiagen) according to manufacturers’ instructions.
MSK-IMPACT sequencing
Tumor and normal DNA from each case were subjected to massively parallel sequencing targeting all coding regions of 505 cancer-related genes using the FDA-approved MSK-IMPACT assay as previously described47,49. The median depth of coverage of tumor and normal samples was 758× (range: 502×–1167×) and 389× (range: 145×–540×), respectively. In brief, reads were aligned to the reference human genome GRCh37 using the Burrows–Wheeler Aligner (BWA v0.7.15)50. The Genome Analysis Toolkit (GATK. V3.1.1)51 was employed for local realignment, duplicate removal, and base quality recalibration. Somatic single nucleotide variants (SNVs) were detected by MuTect (v1.0)52, insertions, and deletions (indels) by Strelka53, Varscan254, Scalpel55, and Lancet56. All mutations were manually inspected using the Integrative Genomics Viewer (IGV). The cancer cell fraction (CCF) of each mutation was inferred, as well as clonal probability, using ABSOLUTE57. Copy number alterations (CNAs) and LOH were determined using FACETS58. Mutations targeting hotspot loci were assigned according to Chang et al.59. Mutational signatures were inferred using Signature Multivariate Analysis (SigMA) based on all synonymous and nonsynonymous somatic mutations60. Exposure-based dominant mutational signatures obtained by SigMA, an algorithm previously validated for the analysis of formalin-fixed paraffin-embedded samples, were reported in cases with at least 5 SNVs (Supplementary Table 3), as previously described48. The repertoire of non-synonymous somatic mutations, mutational frequencies, and CNAs of classical ILCs were compared to genetic alterations affecting LLIMCas.
CDH1 promoter methylation assessment by digital droplet PCR
Following PicoGreen quantification, 0.2–9 ng bisulfite-treated genomic DNA was combined with locus-specific primers targeting the two CDH1 promoter CpG islands, FAM- and HEX-labeled probes (Supplementary Table 3), the restriction enzyme HaeIII, and digital PCR Supermix for probes (no dUTP). CpG Methylated DNA (ThermoFisher, Waltham, MA) and Universal Unmethylated DNA (Millipore, Burlington, MA) were used as positive and negative controls, respectively. All reactions were performed on a QX200 ddPCR system (Bio-Rad, Hercules, CA), and each sample was evaluated in two technical duplicates.
Reactions were partitioned into ~41 K droplets per well using the QX200 droplet generator. Emulsified PCRs were run on a 96-well thermal cycler using the following cycling conditions: 95 °C 10’; 50 cycles of 94 °C 60’ and 54 °C 2’; 98 °C 10’. Plates were read and analyzed using the QuantaSoft software (Bio-Rad, Hercules, CA) to assess the number of droplets positive for CDH1 promoter methylated, unmethylated, both, or neither. Methylation Frequency (MF) was inferred as MF = 100 * Methylated/(Methylated + Unmethylated). Methylation of the CDH1 promoter was defined as higher than 35 methylated droplets.
Clinical characteristics
A review of patient’s electronic medical records was conducted to obtain the following information: (1) age at diagnosis; (2) date of first diagnosis; (3) type of first surgical procedure; (4) adjuvant therapies; (5) date of first recurrence; (6) type of recurrence and site of distant metastasis; (7) date and status at last follow-up (for patients who developed non-breast new malignancies, the last contact date was censored at the date of new malignancy diagnosis); (8) date and cause of death.
Pathologic characteristics
Pathology reports of the first surgical procedure were reviewed to obtain the following tumor pathologic characteristics: (1) multifocality and, if present, number of foci; (2) tumor size; (3) tumor Nottingham grade; (4) Nottingham score; (5) tumor-associated microcalcifications; (6) lymphovascular space invasion; (7) margin status at first surgery; (8) lymph node status; (9) pathologic stages of tumor (pT) and lymph node (pN).
Predictive and prognostic biomarkers
Results of the IHC stains for ER, PR, HER2, and Ki67 were extracted from the pathology reports. ER and PR were scored by the modified histologic score (H-score) method, calculated by multiplying the intensity of expression (0–3) by the percentage of cells showing that intensity (0–100%). The sum of these numbers was referred to as the H-score. H-score ≥ 1 was considered positive. HER2 was scored per the FDA-cleared interpretation guide for Ventana anti-HER2 antibody. We started performing Ki67 stain on invasive breast carcinomas in late 2007, and the proliferation index was scored as a percentage of cells with staining of any intensity.
ME2 scores were calculated for all cases. ME2 is one of the multivariable models developed to estimate the Oncotype DX® recurrence score61,62,63. ME1 and ME3 require a Ki-67 proliferation index, while ME2 does not. Since the Ki-67 proliferation index was unavailable for most of the cases, only ME2 scores were calculated.
Tumor size pathology-radiology correlation
The final tumor size was obtained by pathologic evaluation of the primary surgical resection. The size of the tumor estimated radiologically was recorded from the mammographic or ultrasound imaging reports at the time of the diagnostic core biopsy. Pathology-to-radiology tumor size ratio was calculated and used as one of the variables for comparing different tumor types.
Outcome and survival analysis
Clinical outcome analysis included recurrence rate, recurrence type, and sites of metastasis. Loco-regional recurrence was defined as a tumor arising in the treated breast/chest wall or within regional lymph nodes. Taking into account the findings published in the literature64,65,66, we separated the patterns of distant metastatic sites into lobular-like (gynecologic or gastrointestinal organs, regardless of any other site involvement), ductal-like (lung, pleura, liver, brain, lymph node, with or without bone involvement), and neutral (bone only).
Long-term survival data included RFS, DRFS, OS, and BCSS. RFS was defined as the time from diagnosis to first recurrence (local or distant) or the date of the last contact. DRFS was defined as the time from diagnosis to first distant recurrence or the date of the last contact. OS was defined as the time from diagnosis to death due to any cause or the date of the last contact. BCSS was defined as the time from diagnosis to death due to breast cancer or the date of the last contact.
Statistical analysis
For comparison of means, independent sample t-tests were performed. Univariable analysis was performed using χ2 and Fisher exact tests to compare the differences in percentages between groups. A p-value < 0.05 was considered significant. Long-term survival data, including RFS, DRFS, OS, and BCSS, were analyzed via Kaplan–Meier curves for different tumor types (all 3 subtypes simultaneously, LLIMCa versus ILC, LLIMCa versus IDC, ILC versus IDC) and after combining LLIMCa with IDC (i.e., ILC versus IDC + LLIMCa) or ILC (i.e., IDC versus ILC + LLIMCa). A log-rank test was used to compare Kaplan–Meier curves. Survival analysis was also performed for known prognostic variables (grade, nodal status, pT stage, pN stage) and ME2 score categories. The variables showing statistically significant differences in survival by log-rank test were included for multivariable Cox proportional hazard regression analysis. Statistical analysis was performed using IBM SPSS Statistics for Windows, Armonk, NY: IBM Corp.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The H&E and IHC datasets generated and analyzed during the current study are not publicly available but can be made available upon reasonable request, following ethics committee approval and a data transfer agreement, to guarantee the General Data Protection Regulation. All genomic data generated and analyzed during this study are available in the Sequencing Read Archive (SRA): SRP446067.
References
Breast Cancer Facts & Figures 2019-2020. (American Cancer Society, https://www.cancer.org/content/dam/cancer-org/research/cancer-facts-and-statistics/breast-cancer-facts-and-figures/breast-cancer-facts-and-figures-2019-2020.pdf).
Breast Tumours. WHO Classification of Tumours. 5th edn, Vol. 2 (International Agency for Research on Cancer, 2019).
Mersin, H., Yildirim, E., Gulben, K. & Berberoglu, U. Is invasive lobular carcinoma different from invasive ductal carcinoma? Eur. J. Surg. Oncol. 29, 390–395 (2003).
Pestalozzi, B. C. et al. Distinct clinical and prognostic features of infiltrating lobular carcinoma of the breast: combined results of 15 International Breast Cancer Study Group clinical trials. J. Clin. Oncol. 26, 3006–3014 (2008).
Wasif, N., Maggard, M. A., Ko, C. Y. & Giuliano, A. E. Invasive lobular vs. ductal breast cancer: a stage-matched comparison of outcomes. Ann. Surg. Oncol. 17, 1862–1869 (2010).
Weigelt, B. et al. The molecular underpinning of lobular histological growth pattern: a genome-wide transcriptomic analysis of invasive lobular carcinomas and grade- and molecular subtype-matched invasive ductal carcinomas of no special type. J. Pathol. 220, 45–57 (2010).
Ciriello, G. et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell 163, 506–519 (2015).
Adachi, Y. et al. Comparison of clinical outcomes between luminal invasive ductal carcinoma and luminal invasive lobular carcinoma. BMC Cancer 16, 248 (2016).
Du, T. et al. Invasive lobular and ductal breast carcinoma differ in immune response, protein translation efficiency and metabolism. Sci. Rep. 8, 7205 (2018).
McCart Reed, A. E., Kalinowski, L., Simpson, P. T. & Lakhani, S. R. Invasive lobular carcinoma of the breast: the increasing importance of this special subtype. Breast Cancer Res. 23, 6 (2021).
Takeichi, M. Cadherin cell adhesion receptors as a morphogenetic regulator. Science 251, 1451–1455 (1991).
Ohkubo, T. & Ozawa, M. p120(ctn) binds to the membrane-proximal region of the E-cadherin cytoplasmic domain and is involved in modulation of adhesion activity. J. Biol. Chem. 274, 21409–21415 (1999).
Dabbs, D. J., Bhargava, R. & Chivukula, M. Lobular versus ductal breast neoplasms: the diagnostic utility of p120 catenin. Am. J. Surg. Pathol. 31, 427–437 (2007).
Pareja, F., Weigelt, B. & Reis-Filho, J. S. Problematic breast tumors reassessed in light of novel molecular data. Mod. Pathol. 34, 38–47 (2021).
Canas-Marques, R. & Schnitt, S. J. E-cadherin immunohistochemistry in breast pathology: uses and pitfalls. Histopathology 68, 57–69 (2016).
Choi, Y. J., Pinto, M. M., Hao, L. & Riba, A. K. Interobserver variability and aberrant E-cadherin immunostaining of lobular neoplasia and infiltrating lobular carcinoma. Mod. Pathol. 21, 1224–1237 (2008).
Da Silva, L. et al. Aberrant expression of E-cadherin in lobular carcinomas of the breast. Am. J. Surg. Pathol. 32, 773–783 (2008).
Rakha, E. A. et al. Clinical and biological significance of E-cadherin protein expression in invasive lobular carcinoma of the breast. Am. J. Surg. Pathol. 34, 1472–1479 (2010).
Sarrio, D. et al. Epigenetic and genetic alterations of APC and CDH1 genes in lobular breast cancer: relationships with abnormal E-cadherin and catenin expression and microsatellite instability. Int. J. Cancer 106, 208–215 (2003).
McCart Reed, A. E., Kutasovic, J. R., Lakhani, S. R. & Simpson, P. T. Invasive lobular carcinoma of the breast: morphology, biomarkers and 'omics. Breast Cancer Res. 17, 12 (2015).
Morrogh, M. et al. Cadherin-catenin complex dissociation in lobular neoplasia of the breast. Breast Cancer Res. Treat. 132, 641–652 (2012).
Christgen, M. et al. Inter-observer agreement for the histological diagnosis of invasive lobular breast carcinoma. J. Pathol. Clin. Res. 8, 191–205 (2022).
Biglia, N. et al. Clinical-pathologic features, long term-outcome and surgical treatment in a large series of patients with invasive lobular carcinoma (ILC) and invasive ductal carcinoma (IDC). Eur. J. Surg. Oncol. 39, 455–460 (2013).
Arps, D. P., Healy, P., Zhao, L., Kleer, C. G. & Pang, J. C. Invasive ductal carcinoma with lobular features: a comparison study to invasive ductal and invasive lobular carcinomas of the breast. Breast Cancer Res. Treat. 138, 719–726 (2013).
Park, J. S. et al. Comparison of clinicopathological features and treatment results between invasive lobular carcinoma and ductal carcinoma of the breast. J. Breast Cancer 18, 285–290 (2015).
Zengel, B. et al. Comparison of the clinicopathological features of invasive ductal, invasive lobular, and mixed (invasive ductal + invasive lobular) carcinoma of the breast. Breast Cancer 22, 374–381 (2015).
Xiao, Y. et al. Mixed invasive ductal and lobular carcinoma has distinct clinical features and predicts worse prognosis when stratified by estrogen receptor status. Sci. Rep. 7, 10380 (2017).
Yasui, H. et al. Expression of N-terminal-deficient E-cadherin protein in invasive lobular carcinoma of the breast. Am. J. Surg. Pathol. 46, 383–391 (2022).
Cancer Genome Atlas Research, N. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Gamallo, C. et al. Correlation of E-cadherin expression with differentiation grade and histological type in breast carcinoma. Am. J. Pathol. 142, 987–993 (1993).
Goldstein, N. S. Does the level of E-cadherin expression correlate with the primary breast carcinoma infiltration pattern and type of systemic metastases? Am. J. Clin. Pathol. 118, 425–434 (2002).
Sarrio, D. et al. Cytoplasmic localization of p120ctn and E-cadherin loss characterize lobular breast carcinoma from preinvasive to metastatic lesions. Oncogene 23, 3272–3283 (2004).
Shibata, T., Kokubu, A., Sekine, S., Kanai, Y. & Hirohashi, S. Cytoplasmic p120ctn regulates the invasive phenotypes of E-cadherin-deficient breast cancer. Am. J. Pathol. 164, 2269–2278 (2004).
Pareja, F. et al. The genomic landscape of metastatic histologic special types of invasive breast cancer. NPJ Breast Cancer 6, 53 (2020).
Johnson, K., Sarma, D. & Hwang, E. S. Lobular breast cancer series: imaging. Breast Cancer Res. 17, 94 (2015).
Mann, R. M., Kuhl, C. K., Kinkel, K. & Boetes, C. Breast MRI: guidelines from the European Society of Breast Imaging. Eur. Radiol. 18, 1307–1318 (2008).
Mann, R. M. et al. The value of MRI compared to mammography in the assessment of tumour extent in invasive lobular carcinoma of the breast. Eur. J. Surg. Oncol. 34, 135–142 (2008).
Oliveira, T. M. et al. Evolving concepts in breast lobular neoplasia and invasive lobular carcinoma, and their impact on imaging methods. Insights Imaging 5, 183–194 (2014).
Van Goethem, M. et al. Magnetic resonance imaging in breast cancer. Eur. J. Surg. Oncol. 32, 901–910 (2006).
Jafferbhoy, S. F. et al. Impact of preoperative MRI in invasive ductal carcinoma with lobular features on core biopsy. Clin. Breast Cancer https://doi.org/10.1016/j.clbc.2020.08.007 (2020).
El Sharouni, M. A., Postma, E. L. & van Diest, P. J. Correlation between E-cadherin and p120 expression in invasive ductal breast cancer with a lobular component and MRI findings. Virchows Arch. 471, 707–712 (2017).
McCart Reed, A. E. et al. Mixed ductal-lobular carcinomas: evidence for progression from ductal to lobular morphology. J. Pathol. 244, 460–468 (2018).
Abdel-Fatah, T. M. et al. Morphologic and molecular evolutionary pathways of low nuclear grade invasive breast cancers and their putative precursor lesions: further evidence to support the concept of low nuclear grade breast neoplasia family. Am. J. Surg. Pathol. 32, 513–523 (2008).
Moll, R., Mitze, M., Frixen, U. H. & Birchmeier, W. Differential loss of E-cadherin expression in infiltrating ductal and lobular breast carcinomas. Am. J. Pathol. 143, 1731–1742 (1993).
Simpson, P. T., Reis-Filho, J. S., Gale, T. & Lakhani, S. R. Molecular evolution of breast cancer. J. Pathol. 205, 248–254 (2005).
Weigelt, B. & Reis-Filho, J. S. Histological and molecular types of breast cancer: is there a unifying taxonomy. Nat. Rev. Clin. Oncol. 6, 718–730 (2009).
da Silva, E. M. et al. Mesonephric and mesonephric-like carcinomas of the female genital tract: molecular characterization including cases with mixed histology and matched metastases. Mod. Pathol. 34, 1570–1587 (2021).
da Silva, E. M. et al. TERT promoter hotspot mutations and gene amplification in metaplastic breast cancer. NPJ Breast Cancer 7, 43 (2021).
Selenica, P. et al. APOBEC mutagenesis, kataegis, chromothripsis in EGFR-mutant osimertinib-resistant lung adenocarcinomas. Ann. Oncol. 33, 1284–1295 (2022).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Narzisi, G. et al. Accurate de novo and transmitted indel detection in exome-capture data using microassembly. Nat. Methods 11, 1033–1036 (2014).
Narzisi, G. et al. Genome-wide somatic variant calling using localized colored de Bruijn graphs. Commun. Biol. 1, 20 (2018).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131 (2016).
Chang, M. T. et al. Accelerating discovery of functional mutant alleles in cancer. Cancer Discov. 8, 174–183 (2018).
Gulhan, D. C., Lee, J. J., Melloni, G. E. M., Cortes-Ciriano, I. & Park, P. J. Detecting the mutational signature of homologous recombination deficiency in clinical samples. Nat. Genet. 51, 912–919 (2019).
Bhargava, R. et al. Magee equations and response to neoadjuvant chemotherapy in ER+/HER2-negative breast cancer: a multi-institutional study. Mod. Pathol. 34, 77–84 (2021).
Flanagan, M. B., Dabbs, D. J., Brufsky, A. M., Beriwal, S. & Bhargava, R. Histopathologic variables predict Oncotype DX recurrence score. Mod. Pathol. 21, 1255–1261 (2008).
Klein, M. E. et al. Prediction of the Oncotype DX recurrence score: use of pathology-generated equations derived by linear regression analysis. Mod. Pathol. 26, 658–664 (2013).
Lamovec, J. & Bracko, M. Metastatic pattern of infiltrating lobular carcinoma of the breast: an autopsy study. J. Surg. Oncol. 48, 28–33 (1991).
Arpino, G., Bardou, V. J., Clark, G. M. & Elledge, R. M. Infiltrating lobular carcinoma of the breast: tumor characteristics and clinical outcome. Breast Cancer Res. 6, R149–R156 (2004).
Mathew, A. et al. Distinct pattern of metastases in patients with invasive lobular carcinoma of the breast. Geburtshilfe Frauenheilkd. 77, 660–666 (2017).
Acknowledgements
We thank Ms. Morgan DeBerry, HTL(ASCP)CM, QIHCCM for her excellent technical assistance. This work is supported by internal department funds (UPMC) and in part by a Cancer Center Support Grant of the National Cancer Institute (NCI)/National Institutes of Health (NIH) [P30CA008748]-MSKCC. J.S.R.-F. is funded in part by the Breast Cancer Research Foundation, from Susan G. Komen through a Komen Scholar Leadership grant, and in part by an NIH/NCI P50 CA247749 01 grant. F.P. is funded in part by an NIH/NCI P50 CA247749 01 grant. S.O. is funded in part by the Breast Cancer Research Foundation and from Susan G. Komen through a Komen Scholar Leadership grant. A.L. is funded in part by the Breast Cancer Research Foundation. Findings from this study were presented in part at the 2022 San Antonio Breast Cancer Symposium and the 2023 United States and Canadian Academy of Pathology Annual Meeting.
Author information
Authors and Affiliations
Contributions
J.Y.: Conceptualization, investigation, analysis, writing—original draft, and writing—review and editing. E.M.d.S.: Investigation, analysis, and writing—review and editing. H.-S.L.: Data acquisition, investigation, and writing—review and editing. B.Z.C.: Investigation and writing—review and editing. J.L.F: Investigation and writing—review and editing. G.J.C.: Investigation and writing—review and editing. T.M.V.: Investigation and writing—review and editing. T.R.S.: Investigation and writing—review and editing. A.V.L.: Investigation and writing—review and editing. S.O.: Investigation and writing—review and editing. T.B.: Investigation and writing—review and editing. J.B.-H.: Investigation and writing—review and editing. P.S.: Investigation and writing—review and editing. Q.Y.: Investigation and writing—review and editing A.D.C.P.: Investigation and writing—review and editing. H.D.: Investigation and writing—review and editing. A.G.: Investigation and writing—review and editing. A.M.: Investigation and writing—review and editing. F.P.: Investigation, analysis, and writing—review and editing. J.S.R.-F.: Investigation, analysis, supervision, and writing—review and editing. R.B.: Conceptualization, acquisition of departmental funds, investigation, supervision, analysis, and writing—review and editing.
Corresponding authors
Ethics declarations
Competing interests
None of the authors have conflict of interest directly related to this manuscript. J.S.R.-F reports receiving personal/consultancy fees from Goldman Sachs, Bain Capital, REPARE Therapeutics and Paige.AI, membership of the scientific advisory boards of VolitionRx, REPARE Therapeutics and Paige.AI, membership of the Board of Directors of Grupo Oncoclinicas, and ad hoc membership of the scientific advisory boards of Astrazeneca, Merck, Daiichi Sankyo, Roche Tissue Diagnostics and Personalis, outside the scope of this study. R.B. reports receiving consultancy fee from Agilent Technologies Inc. and ImmunoGen. A.L. is a member of the Susan G Komen Scientific Advisory Board. S.O. is an Associate Editor and J.S.R.-F. is a Deputy Editor-in-Chief of npj Breast Cancer and played no role in the peer review or decision to publish this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yu, J., da Silva, E.M., La, HS. et al. Clinicopathologic and genomic features of lobular like invasive mammary carcinoma: is it a distinct entity?. npj Breast Cancer 9, 60 (2023). https://doi.org/10.1038/s41523-023-00566-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41523-023-00566-7
- Springer Nature Limited