Abstract
Several population-level studies have described individual clinical risk factors associated with suboptimal antibody responses following COVID-19 vaccination, but none have examined multimorbidity. Others have shown that suboptimal post-vaccination responses offer reduced protection to subsequent SARS-CoV-2 infection; however, the level of protection from COVID-19 hospitalisation/death remains unconfirmed. We use national Scottish datasets to investigate the association between multimorbidity and testing antibody-negative, examining the correlation between antibody levels and subsequent COVID-19 hospitalisation/death among double-vaccinated individuals. We found that individuals with multimorbidity ( ≥ five conditions) were more likely to test antibody-negative post-vaccination and 13.37 [6.05–29.53] times more likely to be hospitalised/die from COVID-19 than individuals without conditions. We also show a dose-dependent association between post-vaccination antibody levels and COVID-19 hospitalisation or death, with those with undetectable antibody levels at a significantly higher risk (HR 9.21 [95% CI 4.63–18.29]) of these serious outcomes compared to those with high antibody levels.
Similar content being viewed by others
Introduction
The COVID-19 vaccination programme in Scotland has rolled out up to five doses for some individuals to date (March 10, 2023)1 and the vaccines have been shown to be highly protective against severe outcomes of the associated disease2,3. However, immunological responses to these vaccines differ between individuals and there may be reduced vaccine clinical effectiveness in those with suboptimal immune responses.
Most individuals generate SARS-CoV-2-specific antibodies following COVID-19 vaccination or natural infection. While neutralising antibodies are widely accepted as markers of protection against future infection4, the total SARS-CoV-2-specific antibody titre is more practical to measure and can be deployed in large-scale studies5. Previous population-based studies have examined demographic, clinical and lifestyle factors associated with suboptimal post-vaccination IgG responses and observed lower antibody levels in older people, males, transplant recipients, obese individuals, smokers and those with specific comorbidities (e.g., cancer, depression, diabetes, hypertension, immunodeficiency, stroke, and kidney, liver, lung or neurological disease)6,7. Additionally, previous studies have shown a correlation between post-vaccination IgG and protection from subsequent SARS-CoV-2 infection5,8,9,10,11,12. To our knowledge, however, no studies have examined the association between multimorbidity and post-vaccination immune responses, and, crucially, questions remain as to the level of protection from severe COVID-19 sequelae (i.e., hospitalisation and death) among those with suboptimal post-vaccination immune responses. Answering these questions could illustrate the potential clinical utility of measuring IgG and help to identify individuals who should be targeted for modified vaccination strategies or COVID-19 therapeutics.
The COVID-19 vaccine programme began on 8 December 2020 in Scotland. The first doses of the ChAdOx1 vaccine were administered to care home residents and frontline health and social care workers. The eligibility for vaccination began with individuals aged 80 and over, progressing to younger age groups with a higher priority for those with underlying health conditions. Notably, in February 2021, the BNT162b2 vaccine was introduced and as the rollout progressed, by April 2021, mRNA-1273 was also added to the programme.
Here we use a novel, real-time, national linked dataset to investigate risk factors associated with testing negative for anti-SARs-CoV-2 IgG after completion of the primary COVID-19 vaccine course (at least two doses of ChAdOx1 [Oxford/AstraZeneca], BNT162b2 [Pfizer BioNTech] and/or mRNA-1273 [Moderna]). We also investigate the association between post-vaccination IgG levels and the risk of COVID-19 infection and hospitalisation or death.
Results
Participants
A total of 66,531 primary care samples were tested for anti-SARS-CoV-2 IgG antibodies between April 20, 2020, and March 28, 2022. (Fig. 1). After excluding samples with missing information (16%, n = 10,710) and restricting to those taken at least 14 days after completion of the primary vaccine course (i.e., two doses), 17,651 primary care samples (relating to 17,530 individuals) were available for analysis.
Descriptive data
Table 1 presents demographic and clinical characteristics overall and for the subset of individuals who tested IgG negative. In total, 7.3% (n = 1275) of individuals had at least one negative IgG test recorded after two or more vaccine doses. The percentage who tested IgG negative declined to 0.8% (n = 43) (range 0.6–1.0% depending on the vaccine product) after at least three doses of vaccine.
The percentage of double-vaccinated individuals who tested IgG negative increased with age (3.7% of primary care attendees aged 20–39 tested IgG negative, whereas 10.7% of those aged 60+ tested IgG negative). Among those with no documented clinical risks, 5.2% tested IgG negative, as compared to 20.6% of those with 5 or more risks. Individuals with haematological and respiratory cancers were found to have the highest percentage of individuals who tested IgG negative at 21.5% and 22.7%, respectively.
The most prevalent risk groups were asthma and severe mental health illnesses (Supplementary Fig. 6). Among the primary care cohort attendees with multimorbidity, certain risk groups were more commonly observed with greater multimorbidity and higher mean age. For instance, among those with five or more risks, 66% had coronary heart disease, 59% had type-II diabetes, and 50% had chronic kidney disease (Supplementary Fig. 6). Similarly, among those with five or more risks and who had tested IgG negative, the most common risk groups were coronary heart disease (59%), type-II diabetes (59%) and chronic kidney disease (55%) (Supplementary Table 8).
Characteristics associated with testing IgG negative after completing the primary vaccine course
Individuals with multimorbidity (i.e. those in five or more risk groups) had increased odds (Odds Ratio [OR] 1.94 [95% CI 1.45–2.60]) of testing IgG negative post-vaccination compared to those with no risk group (model A, Table 2, Fig. 2).
**Note: (1) additional variables for age, days since vaccination and the calendar period were used and adjusted for in the model (GAM) but are not displayed in this plot. (2) Missing data points indicate that the input variable was not included in the model due to lack of statistical power or was not appropriate.
For individuals with at least one risk (model B, Table 3, Fig. 3), the following clinical risk factors were associated with increased odds of testing negative: chronic kidney disease (1.38 [1.11–1.72]), cirrhosis (2.19 [1.35–3.57]), Type I diabetes (1.75 [1.00–3.08]), Type II diabetes (1.46 [1.21–1.77]), haematological cancer (2.49 [1.48–4.20]), and rare neurological conditions (2.45 [1.30–4.60]).
**Note: (1) additional variables for age, days since vaccination and the calendar period were used and adjusted for in the model (GAM) but are not displayed in this plot. (2) Missing data points indicate that the input variable was not included in the model due to lack of statistical power or was not appropriate. (3) ORs for the individual risk groups are calculated compared to individuals who are not in that risk group (as opposed to individuals in no risk groups at all).
Increased risks were observed for those who had been advised to remain indoors by the Scottish Government, due to medical fragility, subsequently referred to as “shielding” (1.68 [1.29–2.18]), and those who were severely immunosuppressed (2.25 [1.80–2.83]). Individuals who, at the time of the serology measurements, had received three or more vaccine doses or had at least one known prior SARS-CoV-2 infection, had decreased odds of testing IgG negative (Tables 2 and 3).
In model A, obesity (BMI \(\ge\) 30) was associated with an increased risk of testing IgG negative. Neither Scottish Index of Multiple Deprivation (SIMD) nor biological sex were found to have a statistically significant effect on the outcome. We observed the ORs for testing IgG negative increased with age (Supplementary Fig. 2a) and days since last vaccination (Supplementary Fig. 2b). In contrast, we observed a decreasing trend in ORs as a function of time since the first serology measurement (Supplementary Fig. 2c).
We conducted complementary analyses on serology samples extracted from blood donors to provide a comparison population; the results are detailed in Supplementary Note 1, Supplementary Tables 1-4. We conducted sensitivity analyses to examine the effect of including individuals who had received only one dose of any COVID-19 vaccine at the time of the antibody testing (Supplementary Note 2). We found individuals in additional risk groups (sickle cell disease or rheumatoid arthritis) had increased ORs of testing IgG negative after only one dose, but the odds were not significantly greater after completing the primary vaccine course. We also carried out an additional sensitivity analysis where we used more and less conservative definitions of a negative IgG test result; the former being one in which the quantitative IgG measurement could not detect any IgG, and the latter being one deemed to be a measurement of an arbitrary low level of IgG (Supplementary Note 3). The tighter definition yielded significantly higher ORs of a post-vaccination undetectable IgG test result for individuals in increasing number of risk groups (Supplementary Fig. 8), as well as individuals in at least one risk group (Supplementary Fig. 9) with: haematological cancer, pulmonary hypertension, a rare neurological disease, cirrhosis, a history of heart failure, or coronary heath disease.
Association between IgG levels and subsequent SARS-CoV-2 infection or severe COVID-19 outcomes
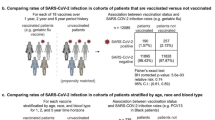
Thirteen percent (n = 2278) of individuals tested positive for SARS-CoV-2 (at least once) after a serology sample was taken. Risk of SARS-CoV-2 infection (model C, Table 4) was higher amongst post-vaccination IgG negatives, relative to IgG positives (Hazard Ratios [HR] of 1.50 [1.30–1.73]). Administration of additional vaccines between the date of IgG measurement and the outcome reduced the HR (Supplementary Fig. 3). Supplementary Note 1 additionally provides a comparison using samples extracted from blood donors for model C.
In total, 85 individuals (0.5%) were either hospitalised due to COVID-19 and/or died of a COVID-19-related death. Of those with severe COVID-19 outcomes, 31.8% had previously tested IgG negative after completing the primary vaccine course (Table 1).
The risk of hospitalisation or death was higher for individuals who had tested IgG negative post-vaccination (3.68 [2.28–5.94]) (model D, Table 5). Individuals with five or more risks were 13.37 [6.05–29.53] times more likely to be hospitalised or die from COVID-19 than an individual with no risks.
HRs for model E are also shown in Table 5 (and Supplementary Fig. 4). Compared to those with a high IgG level, the risk of COVID-19 hospitalisation or death was: 9.21 [4.63–18.29] for those with an undetectable IgG level; 2.15 [1.08–4.26] for those with a very low IgG level; 1.57 [0.89–2.77] for those with a low IgG level; and 0.33 [0.15–0.74] for those with a very high IgG level. Of the risk groups, increased risk of COVID-19 hospitalisation or death was observed in individuals with haematological cancer (2.79 [1.15–6.76]), pulmonary hypertension (4.99 [1.95–12.75]), and chronic kidney disease (2.21 [1.33–3.65]).
Discussion
We found that individuals with multimorbidity were more likely to test IgG negative following at least 2 doses of COVID-19 vaccine. We also found that individuals who tested IgG negative post-vaccination were at greater risk of subsequent SARS-CoV-2 infection (approximately 1.5-fold) and, crucially, COVID-19 hospitalisation or death (nearly 4-fold) compared to those who had tested positive for IgG. In addition, post-vaccination IgG levels were associated with severe COVID-19 outcomes in a dose-dependent manner (i.e., increasing risk with decreasing IgG level), with those having the lowest IgG levels being at the greatest risk (2-fold and 9-fold increased risk among those with very low and undetectable antibodies, respectively, relative to those with high IgG levels).
We investigated the association between post-vaccination antibody response and risk of subsequent severe COVID-19 outcomes at a population level13,14. Other studies have examined the association between antibody levels and death from COVID-19, but these have focussed on small cohorts of patients with COVID-19 already admitted to hospital/critical care15,16,17. We demonstrated a dose-response relationship between post-vaccination antibody levels and hospitalisation or death from COVID-19, with decreasing antibody level associated with increasing risk of hospitalisation or death, and highlighting the group of individuals with undetectable post-vaccination IgG as especially vulnerable. We demonstrated that multimorbidity is a risk factor for insufficient IgG responses post-COVID-19 vaccination. We also showed that individuals with obesity, immunosuppression, and those advised to shield during the pandemic were at increased risk of testing IgG negative post-vaccination, as well as individuals with specific clinical risk factors (chronic kidney disease, cirrhosis, Type I diabetes, Type II diabetes, haematological cancer, and rare neurological conditions). Previous studies involving the EAVE-II cohort have identified similar factors (including multimorbidity and underlying health conditions such as individuals receiving immunosuppressants and those with chronic kidney disease) as being higher risk for severe COVID-19 outcomes13,14. Our findings suggest that IgG may mediate these associations. Individuals who are more likely to test IgG negative post-vaccination, indicative of a suboptimal post-vaccination immune response, may be eligible for COVID-19 therapeutics or modified vaccination strategies. However, it is notable that modified vaccination strategies, either through extra doses or by considering the timing of doses in relation to treatment, may benefit some, but potentially not all, as there may be persistent non-responders.
As in other studies12, our results demonstrated an increase in antibody levels with further vaccine doses, with a very small proportion (0.8%) testing IgG negative after three or more doses. It is possible that the proportion who remain seronegative would decrease even further after 4 or 5 vaccine doses. Among the individuals who had received ≥3 vaccine doses, and still tested IgG negative, 48% (n = 23) were in at least two risk groups. Of all individuals who had received ≥2 doses of the vaccine and tested IgG negative, 31.8% (n = 27) of these subsequently had severe outcomes.
Given that the data used in our analyses were derived from linkage of national administrative and surveillance datasets, the key strengths of our study include efficiency (using existing data as opposed to collecting primary data, which has significant time and cost implications), large sample sizes, national coverage and reduced risk of bias as they were not subject to the biases that can arise in traditional cohort studies – for example, biases resulting from participant attrition. However, our study also has limitations. Given the small number of individuals in some subgroups (for example individuals testing IgG negative after ≥3 vaccine doses), it was not possible to further stratify the data to provide information on the specific risk groups and vaccines used because of statistical disclosure requirements. There are potential biases in the study population: individuals attending primary care and getting a blood sample taken are more likely to have comorbidities than the general population (48% of individuals in our primary care cohort were in at least one risk group, as compared to 30% in the general population of Scotland). A reasonably large proportion (16%) of potentially eligible records were excluded because of missing information (e.g., on vaccination, risk categories, BMI, etc.). Bias may have been introduced if those with missing information were systematically different regarding the exposure and outcome variables.
Because the serology samples were taken at various time points after vaccination occurred, we do not know whether, among those who tested IgG negative, vaccines never induced seroconversion or whether levels waned over time, although we adjusted our models for time since vaccination to try to account for this. Our study was underpowered to detect differences between those who tested negative shortly after vaccination (i.e. never generated IgG) and those who tested negative at a much later time point (i.e. IgG waned) and future analyses should examine any differences in risk between these two groups. Given we did not follow up individuals over time, we also do not know if subsequent vaccines (e.g., three or more doses) will have induced seroconversion among those who tested negative after two doses. Further, the immune response to vaccination is complex and we have only assessed one component of humoral immunity (anti-spike IgG) and not the other types of humoral (e.g. IgA, IgM, neutralising antibodies) or cellular immunity18. For example, T-cells have been shown to play a significant role in protection against severe COVID-19 disease and death19, therefore we may be ’misclassifying’ individuals as having suboptimal post-vaccination responses (according to our measure of IgG) when they in fact have adequate T-cell responses. However, previous studies have found that IgG is highly correlated with neutralising antibody activity4, which is consistent with our results showing that the presence or levels of IgG are associated with post-vaccination COVID-19 outcomes (i.e., infection, hospitalisation and death), confirming that anti-S IgG is a good overall indicator of immune response. Additionally, S-based assays cannot distinguish between vaccine- or infection-induced antibodies, and we therefore do not know if an individual has additional immunity conferred by prior SARS-CoV-2 infection. We adjusted our models for prior infection; however, COVID-19 infection measured by Reverse Transcription Polymerase Chain Reaction (RT-PCR) may have been an underestimate as it did not include COVID-19 antigen tests done in the community and we will also have missed undiagnosed infections.
Our findings confirm previously reported associations between the following risks or characteristics and a suboptimal antibody response following vaccination (individuals may fall into more than one risk group): cancers7,20,21, obesity7,22,23, use of immunosuppressant therapies6,20,21,24,25, being on the shielded patient list25 and age6,7,26. Our findings of a greater likelihood of testing negative for antibodies post-vaccination among specific clinical risk groups (those with kidney disease7,27, cirrhosis28, diabetes7,29, haematological cancers30,31,32, and neurological conditions7,33) were also consistent with the published literature, although in some of these conditions a diminished immune response may be attributable to therapies used to treat the condition rather than the condition itself. We did not however observe any association between socioeconomic status or sex and antibody response – the latter finding in contrast to previously published results, which have demonstrated higher SARS-CoV-2 IgG levels among females7,26. Prior SARS-CoV-2 infection was associated with very low likelihood of testing IgG negative, consistent with previous findings that those who are both infected and vaccinated have greater magnitude and persistence of antibody responses7,34,35. We are also aware of several studies that have directly examined the association between antibody responses to vaccination and subsequent SARS-CoV-2 infection5,8,9,10,11,12. One of these, a large cohort study (N > 4000) derived from a UK-based twin registry, found that those with the lowest 20% of antibody levels post-vaccination had 3-fold greater odds of SARS-CoV-2 infection12.
Our analysis spans a large timeframe, which includes the periods relating to the emergence and dominance of the Delta and Omicron variants. Studies have shown decreased vaccine effectiveness over time as new variants diverge from the original variants that were used to derive the vaccines, which may relate to the specificity of vaccine-induced antibodies3,36,37. We did not have information on the SARS-CoV-2 variant for relevant exposures or outcomes; therefore, we adjusted for the time elapsed since beginning of pandemic to account for the prevalence of different variants in Scotland (see Supplementary Fig. 5). However, this approach may not fully account for the risk associated with different variants, and we acknowledge this as a limitation.
Different vaccines have different immunogenic profiles; we attempted to account for this by adjusting for the vaccine product administered. We found that individuals who had received two doses of ChAdOx1 were significantly less likely to test positive for IgG than the reference category (two doses of BNT162b2 or mRNA-1273). This may be partly due to systematic differences among those who received ChAdOx1 because of vaccine programme prioritisation, and other unmeasured confounders; however, it may also be partly attributable to the known lower immunogenicity of the ChAdOx138.
In conclusion, we have demonstrated a strong association between IgG antibody levels and severe COVID-19 outcomes, illustrating the clinical utility of measuring IgG. We have also identified subgroups of the population who generated suboptimal serological responses post-COVID-19 vaccination – particularly those with multimorbidity – and therefore who remain at increased risk of serious COVID-19 outcomes. Consideration should now be given to extend the indications for COVID-19 therapeutics to include those living with multiple long-term conditions, rather than on the basis of single conditions alone. More research into COVID-19 therapeutics for those living with multimorbidity is required.
Methods
Study design and population
We used linked data from Early Pandemic Evaluation and Enhanced Surveillance of COVID-19 (EAVE II), a Scotland-wide cohort of 5.4 million people ( ~ 99% of the Scottish population), consisting of primary and secondary care, COVID-19 vaccination, SARS-CoV-2 testing, and mortality data39. Additionally, we used SARS-CoV-2 antibody testing data from the Enhanced Surveillance of COVID-19 in Scotland (ESoCiS), a national serological surveillance programme that sampled residual blood from multiple sources, including antenatal, blood donor, paediatric and primary care (i.e. general practice) settings, and tested these for the presence of SARS-CoV-2 IgG antibodies to the spike protein40,41.
Serology measurements used in this study were obtained from testing residual blood from people attending primary care settings for routine purposes. Samples were obtained from biochemistry laboratories (700 weekly samples) covering 11 regional health authorities, which represent >90% of the Scottish population. Additionally, primary care samples related to individuals aged 6 years and older and were chosen according to an age/sex/geographical sampling frame proportional to the Scottish general population. The serology results for primary care samples taken between April 20, 2020 and March 28, 2022 were linked with EAVE II data. All analyses were restricted to individuals who had received at least two doses of a COVID-19 vaccine prior to the serology sample date, unless otherwise specified.
We also undertook supplementary analyses of serology measurements from blood donors, similar to those applied to the primary care cohort. Methods and results of these analyses are described in Supplementary Note 1.
This study involves human participants. The Public Benefit and Privacy Panel Committees of Public Health Scotland and Scottish Government approved the linkage and analysis of the deidentified datasets for this project (2021-0115).
Definition of outcomes
We defined testing IgG negative, to be a negative SARS-CoV-2 IgG test result obtained post-vaccination (at least 14 days after completion of the primary vaccine schedule). Assay manufacturers define a negative IgG test by quantitative IgG levels and stated cut-offs: <33.8 binding antibody units (BAU) per ml (tested using the Diasorin SARS-CoV-2 TrimericS IgG assay for antibodies to the S1/S2 protein)42. The outcome of a test result with undetectable IgG levels, as determined by the assay manufacturers, using a cut-off of <4.8 [BAU/ml], is presented in the sensitivity analysis in Supplementary Note 3. An arbitrary cut-off of <100 [BAU/ml] was also used in a sensitivity analysis to study a looser definition of a low post-vaccination IgG response.
For the analysis of the association between SARS-CoV-2 IgG antibody levels (hereafter referred to as “IgG levels”) and subsequent SARS-CoV-2 infection or severe COVID-19 outcomes, we considered SARS-CoV-2 infections, hospitalisations and deaths that occurred after the serology test date. SARS-CoV-2 infections were defined as positive RT-PCR test results; for multiple positive tests, the first test after the serology test was selected. COVID-19 hospitalisations were sourced from the Scottish Morbidity Record and were defined as hospital admissions where COVID-19 was listed as the primary reason for admission (International Classification of Diseases [ICD-10] codes U07.1 and U07.2); COVID-19 deaths were sourced from National Records of Scotland and defined as deaths where COVID-19 was mentioned on the death certificate (also ICD-10 codes U07.1 and U07.2).
Exposure definitions
For the analysis of factors associated with a negative IgG test, we used 26 comorbidity-based risk groupings43. We used either the total number of risk groups (0, 1, 2, 3-4, ≥5) for all individuals, or the specific risk groups as separate variables (Supplementary Table 5).
For the analysis of IgG levels and subsequent COVID-19 outcomes, the exposure was defined as either a positive or negative IgG test result, or as quantiles of the IgG. Quantiles were defined by analysis of the distribution of all measurements (Supplementary Fig. 1) as: very high (≥2000 BAU/ml), high (230–1999 BAU/ml), low (33.8–229 BAU/ml), very low (4.8–33.7 BAU/ml), and undetectable (< 4.8 BAU/ml).
Confounding factors
In all analyses, we accounted for potential confounding due to: biological sex; age; BMI; SARS-CoV-2 infection (i.e. prior to the serology sample date); vaccine dose and vaccine product last administered ( ≥ 14 days prior to the serology sample date); number of days between the last vaccine dose and the serology sample date; number of days since the start of the pandemic (March 1, 2020); classification variables – derived from vaccine records and patient data held by the national public health body relating to whether the individual was a care home resident, immunosuppressed, severely immunosuppressed, and/or on the Scottish shielded patient list; and socio-economic status measured in quintiles of SIMD.
In the analysis investigating the association between IgG levels and COVID-19 related outcomes, either the total number of risk groups (0, 1, 2, 3-4, ≥5), or specific risks associated with severe outcomes were included as confounders. We accounted for the administration of additional vaccine doses (or boosters) between the serology sample date and the COVID-19 related outcome. We also adjusted for Urban Rural Classification (URC) and SIMD.
Statistical analysis
To examine characteristics associated with testing IgG negative post-double vaccination, we fitted two generalised additive models (GAMs) (Fig. 4). To examine multimorbidity, model A included the total number of risk groups (e.g., 1, 2, 3-4, or 5 + ) as an exposure variable; model B included specific risk groups (e.g., asthma, chronic kidney disease, etc.) as exposure variables and were restricted to individuals with at least one risk. For more details of each specific model, please see Supplementary Table 7.
Given input datasets from EAVE-II and serology datasets from ESoCiS we constructed data containing serology measurements linked with individual demographics, GP records, SARs-COV-2 testing and hospitalisation data for individuals who had received at least two doses of any COVID-19 vaccine. Model A and Model B are Generalised Additive Models (GAMs) where the number of comorbidity-based risks is used as the exposure for the outcome of testing IgG negative after vaccination. Similarly Model B instead used individual risks as the exposure variable for all individuals with at least one risk. Models C, D and E are Cox Proportional Hazard Regression Models (CPHRMs) where the survival is defined as SARs-CoV-2 infection for Model C and hospitalisation/death for Models D and E. Testing negative for IgG is the exposure for models C, and D whereas quantiles of quantitative IgG measurements are used as the exposure for model E.
GAMs were used to account for the influence of multiple confounding variables, some of which exhibited non-linear relationships. Smoothed spline functions were used to address such confounding due to age, the time interval between the last vaccine administration and the serology sample date, as well as the date of the serology sample, to account for residual effects due to the period of the pandemic. Age was stratified by number of vaccine doses, type of product and product combinations, motivated by the timeline and prioritisation of vaccine rollout in Scotland.
Additional risk factors were chosen for inclusion in the models if the p-value calculated in a univariate model was 0.1, or if there was a clinical rationale for their inclusion (and if the group had five or more individuals with the outcome). We calculated adjusted (and unadjusted) ORs from the exponential of the coefficients of the parametric terms of the GAM, using a Wald statistic for the confidence intervals (at 95% CI).
To study the association between IgG levels and subsequent SARS-CoV-2 infection or severe COVID-19 outcomes, we fitted a further two models: one using infection as the outcome (model C), and two with severe COVID-19 as the outcome (models D and E). In model D, the number of risk groups was included as a confounder while in model E, individual risks that were associated with significantly higher risk of severe outcomes were decoupled from all other risk groups and included separately as confounders. In models C and D, we used a binary exposure variable (positive/negative IgG test result) with the number of risk groups as a confounder. In Model E, we used quantiles of IgG titres as the exposure to further investigate how IgG levels, rather than just positive/negative IgG tests, affected the HRs of severe outcomes. These models are also summarised in Supplementary Table 7.
The number of vaccinations post-serology sample date were adjusted for as a time-dependent covariate. Age, sex, and BMI were adjusted for in all models (C, D, and E), with the survival period defined with respect to the start of the pandemic, accounting for the severity of SARS-CoV-2 variants, virus abundance in the general population, lockdowns, and other changes in social relations. Further adjustments were made for care home residency, SIMD, and urban/rural classification in Model C.
The follow-up period extended up to three months after the last recorded IgG measurement (i.e., up to June 28, 2022). Individuals were censored if they died from non-COVID-19 reasons before the end of the study period. Hazard ratios (HR) were calculated from the exponent of the fitted coefficients of the Cox models, with 95% CIs obtained using Wald tests.
All statistical analyses were performed using R version 3.6.1. We followed a pre-specified study analysis plan (available from the authors on request). Results are reported according to the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) guidelines (Supplementary Table 6).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data cannot be shared publicly because it contains potentially identifiable and sensitive patient information and is legally restricted by Public Health Scotland and the Scottish and UK governments. Data were available for researchers who meet the criteria for access due to undertaking this work as part of a national Scottish surveillance program.
Code availability
A data dictionary covering the datasets used in this study can be found at https://eave-ii.github.io/data-dictionary/. All code used in this study is publicly available at https://github.com/EAVE-II/Serology-Analysis/tree/master/analysis. The data used to undertake these analyses are not publicly available because they are based on deidentified national clinical records. These data are available, subject to approval by the NHS Scotland Public Benefit and Privacy Panel, by application through the Scotland National Safe Haven.
References
Public Health Scotland. COVID-19 vaccine information and resources. https://publichealthscotland.scot/our-areas-of-work/conditions-and-diseases/covid-19/covid-19-vaccinations/covid-19-vaccine-information-and-resources/eligible-groups/ (2023).
Kerr, S., Vasileiou, E., Robertson, C. & Sheikh, A. COVID-19 vaccine effectiveness against symptomatic SARS-CoV-2 infection and severe COVID-19 outcomes from Delta AY.4.2: Cohort and test-negative study of 5.4 million individuals in Scotland. J. Glob. Health 12, 05025 (2022).
Florentino, P. T. V. et al. Vaccine effectiveness of two-dose BNT162b2 against symptomatic and severe COVID-19 among adolescents in Brazil and Scotland over time: a test-negative case-control study. Lancet Infect. Dis. 22, 1577–1586 (2022).
Khoury, D. S. et al. Neutralizing antibody levels are highly predictive of immune protection from symptomatic SARS-CoV-2 infection. Nat. Med. 27, 1205–1211 (2021).
Dimeglio, C. et al. Antibody Titers and Protection against Omicron (BA.1 and BA.2) SARS-CoV-2 Infection. Vaccines (Basel) 10, 1548 (2022).
Jolliffe, D. A. et al. Determinants of Antibody Responses to SARS-CoV-2 Vaccines: Population-Based Longitudinal Study (COVIDENCE UK). Vaccines (Basel) 10, 1601 (2022).
Ward, H. et al. Population antibody responses following COVID-19 vaccination in 212,102 individuals. Nat. Commun. 13, 907 (2022).
Gilbert, P. B. et al. Immune correlates analysis of the mRNA-1273 COVID-19 vaccine efficacy clinical trial. Science 375, 43–50 (2022).
Perry, J. et al. Does a humoral correlate of protection exist for SARS-CoV-2? A systematic review. PLoS One 17, e0266852 (2022).
Feng, S. et al. Correlates of protection against symptomatic and asymptomatic SARS-CoV-2 infection. Nat. Med. 27, 2032–2040 (2021).
Goldblatt, D. et al. Towards a population-based threshold of protection for COVID-19 vaccines. Vaccine 40, 306–315 (2022).
Cheetham, N. J. et al. Antibody levels following vaccination against SARS-CoV-2: associations with post-vaccination infection and risk factors in two UK longitudinal studies. Elife 12, e80428 (2023).
Agrawal, U. et al. COVID-19 hospital admissions and deaths after BNT162b2 and ChAdOx1 nCoV-19 vaccinations in 2·57 million people in Scotland (EAVE II): a prospective cohort study. Lancet Respir. Med. 9, 1439–1449 (2021).
Agrawal, U. et al. Severe COVID-19 outcomes after full vaccination of primary schedule and initial boosters: pooled analysis of national prospective cohort studies of 30 million individuals in England, Northern Ireland, Scotland, and Wales. Lancet 400, 1305–1320 (2022).
Martin-Vicente, M. et al. Low anti-SARS-CoV-2 S antibody levels predict increased mortality and dissemination of viral components in the blood of critical COVID-19 patients. J. Intern Med. 291, 232–240 (2022).
Liu, X. et al. Serum IgM against SARS-CoV-2 correlates with in-hospital mortality in severe/critical patients with COVID-19 in Wuhan, China. Aging (Albany NY) 12, 12432–12440 (2020).
Asif, S. et al. Weak anti-SARS-CoV-2 antibody response is associated with mortality in a Swedish cohort of COVID-19 patients in critical care. Crit. Care 24, 639 (2020).
Gaebler, C. et al. Evolution of antibody immunity to SARS-CoV-2. Nature 591, 639–644 (2021).
Alshammary, A. F. et al. T-Cell Subsets and Interleukin-10 Levels Are Predictors of Severity and Mortality in COVID-19: A Systematic Review and Meta-Analysis. Front. Med. (Lausanne) 9, 852749 (2022).
Monin, L. et al. Safety and immunogenicity of one versus two doses of the COVID-19 vaccine BNT162b2 for patients with cancer: interim analysis of a prospective observational study. Lancet Oncol. 22, 765–778 (2021).
Parry, H. et al. Antibody responses after first and second Covid-19 vaccination in patients with chronic lymphocytic leukaemia. Blood Cancer J. 11, 136 (2021).
Kara, Z. et al. Antibody Response to SARS-CoV-2 Vaccines in People with Severe Obesity. Obes. Surg. 32, 2987–2993 (2022).
Nasr, M. C., Geerling, E. & Pinto, A. K. Impact of Obesity on Vaccination to SARS-CoV-2. Front Endocrinol. (Lausanne) 13, 898810 (2022).
Liao, S., Gerber, A. N., Zelarney, P., Make, B. & Wechsler, M. E. Impaired SARS-CoV-2 mRNA Vaccine Antibody Response in Chronic Medical Conditions: A Real-World Analysis. Chest 161, 1490–1493 (2022).
Shrotri, M. et al. Spike-antibody waning after second dose of BNT162b2 or ChAdOx1. Lancet 398, 385–387 (2021).
Notarte, K. I. et al. Effects of age, sex, serostatus, and underlying comorbidities on humoral response post-SARS-CoV-2 Pfizer-BioNTech mRNA vaccination: a systematic review. Crit. Rev. Clin. Lab Sci. 59, 373–390 (2022).
Hou, Y., Lu, K. & Kuo, K. The efficacy of COVID-19 vaccines in chronic kidney disease and kidney transplantation patients: a narrative review. Vaccines (Basel) 9, 885 (2021).
Schinas, G. et al. COVID-19 Vaccination in Patients with Chronic Liver Disease. Viruses 14, 2778 (2022).
Ward, V. et al. SARS-CoV-2 antibody responses post-vaccination in UK healthcare workers with pre-existing medical conditions: a cohort study. BMJ Open 12, e066766 (2022).
Greenberger, L. M. et al. Antibody response to SARS-CoV-2 vaccines in patients with hematologic malignancies. Cancer Cell 39, 1031–1033 (2021).
Lee, A. R. Y. B. et al. Efficacy of covid-19 vaccines in immunocompromised patients: systematic review and meta-analysis. BMJ 376, e068632 (2022).
Lee, L. Y. W. et al. Association of SARS-CoV-2 Spike Protein Antibody Vaccine Response With Infection Severity in Patients With Cancer: A National COVID Cancer Cross-sectional Evaluation. JAMA Oncol. 9, 188–196 (2023).
Gombolay, G. Y., Dutt, M. & Tyor, W. Immune responses to SARS‐CoV‐2 vaccination in multiple sclerosis: a systematic review/meta‐analysis. Ann. Clin. Transl. Neurol. 9, 1321–1331 (2022).
Wei, J. et al. Antibody responses and correlates of protection in the general population after two doses of the ChAdOx1 or BNT162b2 vaccines. Nat. Med. 28, 1072–1082 (2022).
Karachaliou, M. et al. SARS-CoV-2 infection, vaccination, and antibody response trajectories in adults: a cohort study in Catalonia. BMC Med. 20, 347 (2022).
Andrews, N. et al. Covid-19 Vaccine Effectiveness against the Omicron (B.1.1.529) Variant. N. Engl. J. Med. 386, 1532–1546 (2022).
Nielsen, K. F. et al. Vaccine effectiveness against SARS-CoV-2 reinfection during periods of Alpha, Delta, or Omicron dominance: A Danish nationwide study. PLoS Med. 19, e1004037 (2022).
Schmidt, T. et al. Immunogenicity and reactogenicity of heterologous ChAdOx1 nCoV-19/mRNA vaccination. Nat. Med. 27, 1530–1535 (2021).
Mulholland, R. H. et al. Cohort Profile: Early Pandemic Evaluation and Enhanced Surveillance of COVID-19 (EAVE II) Database. Int. J. Epidemiol. 50, 1064–1074 (2021).
Palmateer, N. E. et al. National population prevalence of antibodies to SARS-CoV-2 in Scotland during the first and second waves of the COVID-19 pandemic. Public Health 198, 102–105 (2021).
Public Health Scotland. Enhanced Surveillance of COVID-19 in Scotland. https://publichealthscotland.scot/publications/enhanced-surveillance-of-covid-19-in-scotland/enhanced-surveillance-of-covid-19-in-scotland-population-based-seroprevalence-surveillance-3-august-2022/ (2022).
DiaSorin. LIAISON SARS-CoV-2 TrimericS IgG assay: A quantitative assay for immune status monitoring with an accurate correlation of neutralizing IgG antibodies. https://int.diasorin.com/en/immunodiagnostics/infectious-diseases/sars-cov-2-diagnostic-solutions (2023).
Clift, A. K. et al. Living risk prediction algorithm (QCOVID) for risk of hospital admission and mortality from coronavirus 19 in adults: national derivation and validation cohort study. BMJ 371, m3731 (2020).
Acknowledgements
We thank Dave Kelly from Albasoft (Inverness, UK) for his support with making primary care data available; the Scottish National Serology Strategy Group for their strategic oversight of the serology programme; Steven Kerr and Tristan Millington for validating the QCOVID risk groups; and Wendy Inglis-Humphrey, Lana Woolford, Gabriella Linning, Vicky Hammersley, Calder Hudson, Natalia Reglinska-Matveyev, Laura Brook (University of Edinburgh, Edinburgh, UK) and the EAVE II Public Advisory Group for their support with project management, administration, public involvement and communications work. This study is part of the EAVE II project. EAVE II is funded by the MRC (MC_PC_19075) with the support of BREATHE—The Health Data Research Hub for Respiratory Health (MC_PC_19004), which is funded through the UK Research and Innovation Industrial Strategy Challenge Fund and delivered through Health Data Research UK. Additional funding for this work was received by National Core Studies Immunity. This research is part of the Data and Connectivity National Core Study, led by Health Data Research UK in partnership with the Office for National Statistics and funded by UK Research and Innovation (grant ref MC_PC_20058). Additional support has been provided through Public Health Scotland, the Scottish Government Director General Health and Social Care and the University of Edinburgh. The original EAVE project was funded by the National Institute for Health Research (NIHR) Health Technology Assessment programme (11/46/23). The views expressed are those of the authors and not necessarily those of the NIHR, the Department of Health and Social Care, or the UK government. SVK acknowledges funding from the Medical Research Council (MC_UU_00022/2), the Scottish Government Chief Scient Office (SPHSU17) and a NHS Research Scotland Senior Clinical Fellowship (SCAF/15/02).
Author information
Authors and Affiliations
Contributions
CM, NP, JM, AM, AS were involved in the conception of the research questions. NP, AM, CM, LL, TH, SSH, and EH developed the statistical analysis plan. CM, LL and CR developed the statistical analysis methods. CM conducted the statistical analysis. KJ checked the analyses and code. CM, NP, and JM wrote the first draft of the manuscript. CM and NP are both listed as co-first authors due to their contributions to the manuscript. All authors read, revised and approved the final manuscript. AS is PI of EAVE II, was involved in conceiving this analysis, and commented on drafts of the manuscript. JM and CR conceived of the serology testing programme. AM, NP, SM and LJ were involved in running the serology programme. CM, NP, AM, LL, PG, JM, ZG, LD, HT, AF, CS, CM, BS, SVK IR, LR, CC, PM, CR and AS participated in regular discussions and problem solving with analyses and data linkage throughout the project.
Corresponding authors
Ethics declarations
Competing interests
AS and CR are members of the Scottish Government Chief Medical Officer’s COVID-19 Advisory Group and AS its Standing Committee on Pandemics. AS is also a member of the New and Emerging Respiratory Virus Threats Advisory Group (NERVTAG) Risk Stratification Subgroup. AS was a member of AstraZeneca’s Thrombotic Thrombocytopenic Taskforce. All AS’ roles are unremunerated. CR is a member of SPI-M. JM was a member of the National Incident Management Team COVID-19, and lead of Enhanced Surveillance of COVID-19 in Scotland during this project. IR is a member of the Scientific Council on COVID-19 pandemic of the Government of the Republic of Croatia and the co-Editor-in-Chief of the Journal of Global Health. All other authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Macdonald, C., Palmateer, N., McAuley, A. et al. Association between antibody responses post-vaccination and severe COVID-19 outcomes in Scotland. npj Vaccines 9, 107 (2024). https://doi.org/10.1038/s41541-024-00898-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-024-00898-w
- Springer Nature Limited