Abstract
Fibrotic scar tissue formation occurs in humans and mice. The fibrotic scar impairs tissue regeneration and functional recovery. However, the origin of scar-forming fibroblasts is unclear. Here, we show that stromal fibroblasts forming the fibrotic scar derive from two populations of perivascular cells after spinal cord injury (SCI) in adult mice of both sexes. We anatomically and transcriptionally identify the two cell populations as pericytes and perivascular fibroblasts. Fibroblasts and pericytes are enriched in the white and gray matter regions of the spinal cord, respectively. Both cell populations are recruited in response to SCI and inflammation. However, their contribution to fibrotic scar tissue depends on the location of the lesion. Upon injury, pericytes and perivascular fibroblasts become activated and transcriptionally converge on the generation of stromal myofibroblasts. Our results show that pericytes and perivascular fibroblasts contribute to the fibrotic scar in a region-dependent manner.
Similar content being viewed by others
Main
Regeneration in the adult mammalian central nervous system (CNS) is limited. One major limiting factor is the formation of fibrotic scar tissue1,2,3,4. Initially, the recruitment of fibroblasts and deposition of fibrotic extracellular matrix (ECM) are required for wound closure, but they consequently result in the formation of persistent scar tissue3,5,6. Importantly, a moderate reduction of fibrotic scarring, which still prompts wound closure, promotes axon regeneration and functional recovery after spinal cord injury (SCI)1,3,7,8,9 and alleviates disease severity in experimental autoimmune encephalomyelitis (EAE)4. These findings suggest fibrotic scarring as a therapeutic target to improve regeneration after CNS lesions. Understanding the identity of scar-forming cells and fibrotic tissue formation in more detail may allow targeting the scar-forming process therapeutically.
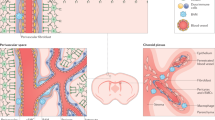
Using in vivo lineage tracing, we previously identified a small population of glutamate aspartate transporter (GLAST; gene name also known as Slc1a3)-expressing perivascular cells, named type A pericytes, as the cellular origin of scar-forming fibroblasts throughout the CNS in response to autoimmune demyelinating disease and traumatic and ischemic lesions6,10. GLAST+ cells represent roughly 10% of all platelet-derived growth factor receptor β (PDGFRβ)+ perivascular cells in the uninjured adult mouse brain and spinal cord6,10. Similarly, a subset of perivascular cells expressing GLAST and PDGFRβ can be found in the healthy human brain and spinal cord10. Single-cell transcriptomic studies of vascular cells in the mouse brain have defined perivascular fibroblasts11, which share Pdgfrb expression with pericytes but can be distinguished by the expression of Pdgfra and Col1a1 (refs. 4,11). Based on a Col1a1-green fluorescent protein (GFP) reporter line, which reflects Col1a1 transcriptional activity, perivascular fibroblasts have been suggested as the origin of fibrotic scar tissue after SCI and EAE4,12. Direct lineage-tracing evidence was limited to Col1a2-expressing cells4 after EAE, which, in addition to perivascular fibroblasts, include some pericytes and vascular smooth muscle cells (vSMCs) in the CNS11. Perivascular fibroblasts reside in the Virchow–Robin space along penetrating arterioles and large ascending venules in an abluminal position to vSMCs4,13. In contrast, pericytes are found in the microvasculature, embedded in the vascular basement membrane14. Furthermore, pericytes and fibroblasts show distinct gene expression profiles at the single-cell level, allowing them to be distinguished from one another11.
Based on single-cell RNA sequencing and lineage tracing in the context of SCI, we confined the fibrotic scar-forming capacity to a specific subset of GLAST-expressing pericytes and perivascular fibroblasts. Both cell populations are recruited locally in response to injury and inflammation. Given their anatomical location, pericytes predominantly contribute to focal lesions in the gray matter, whereas perivascular fibroblasts contribute more extensively to focal white matter lesions. Transcriptionally, we discerned five stromal cell populations at the lesion site, revealing functional diversity in fibrotic ECM deposition, revascularization and immune regulation. Furthermore, we reconstructed the molecular trajectories leading to fibrotic scar formation from pericytes and perivascular fibroblasts after SCI, demonstrating their convergence in generating stromal myofibroblasts. Our findings unveil previously unrecognized heterogeneity in the origin of fibrotic scar tissue, expanding our current understanding of the complexity underlying CNS scar formation4,15.
Results
Fibrotic scar tissue-forming perivascular cells
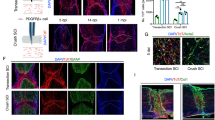
Lineage tracing of GLAST-expressing perivascular cells using GLAST-CreERT2 transgenic mice demonstrates up to 95% contribution to scar-forming fibroblasts after a complete spinal crush, a model of nonpenetrating SCI10. To define the molecular identity of GLAST-expressing perivascular cells, we crossed GLAST-CreERT2 mice to R26R-tdTomato Cre reporter mice and the Pdgfrb-eGFP reporter line, which drives enhanced GFP (EGFP) expression in all PDGFRβ+ perivascular cells (pericytes, vSMCs and fibroblasts). We then isolated equal numbers of GLAST+ (tdTomato+EGFP+) and GLAST− (tdTomato−EGFP+) perivascular cells from uninjured spinal cords (Fig. 1a,b and Supplementary Fig. 1a). Sorted cells were subjected to high-sensitivity single-cell gene expression profiling using Smart-seq16,17,18, in which 520 cells passed the downstream quality controls and were further analyzed.
a, GLAST-expressing perivascular cells can be labeled by tamoxifen-mediated genetic recombination in adult GLAST-CreERT2 transgenic mice carrying R26R-tdTomato reporter alleles, in combination with the Pdgfrb-eGFP reporter line. b, Left, both tdTomato+EGFP+ and tdTomato−EGFP+ cells are associated with blood vessels in the uninjured spinal cord of adult GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice. Right, thoracic spinal cord segments were dissected out and dissociated into a single-cell suspension, and tdTomato+EGFP+ and tdTomato−EGFP+ cells were isolated by fluorescence-activated single-cell sorting (FACS) for downstream single-cell RNA sequencing; the FACS plot is representative of four independent experiments. c, UMAP plot for dimension reduction of spinal cord single-cell data color-coded for 205 tdTomato−EGFP+ (green) and 315 tdTomato+EGFP+ (red) cells. d, The same UMAP plot as in c but color-coded for Seurat clustering of single cells into three distinct clusters. e, Heat map of single-cell RNA-sequencing gene expression data of perivascular cells from the uninjured spinal cord separated according to tdTomato−EGFP+ (green) and tdTomato+EGFP+ (red) origin of cells (selection) and clusters 1–3. All tdTomato+EGFP+ cells express established perivascular cell markers, such as vitronectin (Vtn), Pdgfrb and CD13 (Anpep), whereas Rgs5 is highly expressed in cluster 1 and lowly expressed in cluster 2. Other pericyte markers, such as Kcnj8, Abcc9 and Cd248, are mostly expressed in cluster 1. Fibroblast markers are largely expressed in cluster 2, and vSMC markers are mostly expressed in cluster 3. f, UMAP plot for dimension reduction of tdTomato+EGFP+ cells color-coded for cells of clusters 1 and 2. g, Heat map of differentially expressed genes between tdTomato+EGFP+ cells from clusters 1 and 2. h, Violin plots showing the expression levels of selected perivascular marker genes with similar or differential expression between tdTomato+EGFP+ cells in clusters 1 and 2. TAM, tamoxifen; PCs, pericytes; FBs, fibroblasts; vSMCs, vascular smooth muscle cells; Exp., expression. Scale bar: 10 μm (b).
To classify the sorted populations, we used marker profiles for pericytes, fibroblasts, vSMCs, astrocytes, oligodendrocytes and oligodendrocyte progenitor cells19 and compared marker gene expression between clusters. Cells in cluster 1 consisted of similar numbers of tdTomato+EGFP+ and tdTomato−EGFP+ cells and were identified as pericytes based on high expression of pericyte marker genes and low expression of fibroblast and vSMC markers (Fig. 1c–e). Astrocyte and oligodendroglia lineage markers were absent in cluster 1 (Supplementary Fig. 1b). Cluster 2 contained mainly tdTomato+EGFP+ cells, which were classified as perivascular fibroblasts (Fig. 1c–e) based on lower expression of the pericyte marker genes Cspg4, S1pr3, Cd248, Kcnj8, Abcc9 and Rgs5 and higher expression of the fibroblast markers Col1a1, Col3a1, Lum, Dcn and Pdgfra compared to cells in cluster 1 (Fig. 1e and Supplementary Fig. 1a). PDGFRα is also expressed by oligodendrocyte progenitor cells20, but cluster 2 cells did not express other oligodendroglia lineage marker genes (Supplementary Fig. 1a). We also detected the expression of the astrocyte marker genes Gja1 and Gjb6, encoding the gap junction proteins connexin 43 and connexin 30, respectively21,22, but no additional astrocyte marker genes were detected to be expressed (Supplementary Fig. 1a). Furthermore, we validated Col1a1, Pdgfra and Gjb6 expression in Pdgfrb+ perivascular cells (Supplementary Fig. 1c–e), in line with previous reports23,24,25. Cluster 3 contained mostly tdTomato−EGFP+ cells that were classified as vSMCs based on the high expression of Pdlim, Myh11, Mylk, Acta2 (encoding α-smooth muscle actin (αSMA)) and Tagln (encoding smooth muscle protein 22-α (SM22α)) (Fig. 1c–e). Taking these findings together, we found that most GLAST-expressing perivascular cells are transcriptionally classified as pericytes and that current clustering methods do not distinguish them from GLAST− pericytes (Fig. 1c), in line with a previous report11. Notably, we found a subset of GLAST+ cells clustering separately from pericytes and presenting a fibroblast-like gene expression profile, revealing heterogeneity within GLAST-expressing perivascular cells. GLAST+ cells were distinct from vSMCs (Fig. 1d,e). The cell-type classification was confirmed by integrating the gene expression profiles of all tdTomato+EGFP+ cells with a published single-cell dataset11 that molecularly defined mural cells and fibroblasts from the adult mouse brain (Supplementary Fig. 2a–c). Accordingly, 16% of the cells classified as pericytes by Vanlandewijck et al.11 showed expression of GLAST (Slc1a3) (Supplementary Fig. 3a). Slc1a3-expressing pericytes showed a similar topology to Spp1-, Itih5- and Apod-expressing cells, whereas Pdgfa and Casq2 expression reflected the Slc1a3− topology. These results indicate a gene expression gradient with vSMCs on one side and Slc1a3+ or Spp1+ pericytes on the other (Supplementary Fig. 3a). Consistent with this, we detected neuron–glial antigen 2 (NG2; encoded by Cspg4) expression but not Spp1 expression in tdTomato+EGFP+ capillary pericytes. Nonetheless, tdTomato+EGFP+ cells of the arteriole–capillary transitional zone with projections into the capillary bed expressed Spp1 (Supplementary Fig. 3b,c).
Reclustering of tdTomato+EGFP+ cells alone confirmed that GLAST+ cells encompass pericytes and fibroblasts (Fig. 1f,g). Gene Ontology (GO) enrichment analysis revealed gene expression related to vasculogenesis, transport across and maintenance of the blood–brain barrier, regulation of local adhesion assembly and regulation of phosphatidylinositol 3-kinase signaling in GLAST+ pericytes; in contrast, fibroblasts showed higher expression of genes related to ECM organization, regulation of insulin-like growth factor receptor signaling pathway, cell migration and fibroblast proliferation (Supplementary Fig. 4).
We identified fibroblast-enriched expression of Col1a1, Pdgfra and Fmod, among others (Fig. 1g,h). Owing to the absence of expression in other cell types of the adult CNS, we selected Col1a1 as a fibroblast-specific marker for downstream cell-specific targeting strategies.
Targeting of Col1a1-expressing perivascular fibroblasts
To specifically target Col1a1-expressing fibroblasts in the adult CNS, we used an inducible Col1a1-CreERT2 mouse line (Methods) and crossed it to the R26R-tdTomato reporter mouse line. In some cases, this line was further crossed to the Pdgfrb-eGFP reporter mouse line (Fig. 2a). Recombination of Col1a1-CreERT2;R26R-tdTomato mice led to faithful expression of tdTomato in Col1a1+ cells, as confirmed by RNAscope in situ hybridization (Fig. 2b). The average recombination efficacy was 71.2 ± 5.7% and 82.8 ± 3.7% (mean ± s.d.) in the gray and white matter, respectively (Supplementary Fig. 5a–d). The Col1a1-CreERT2 line recombined 6.9 ± 1.6 (mean ± s.d.) perivascular fibroblasts per section, whereas the GLAST-CreERT2 line, which targets GLAST+ pericytes and fibroblasts, recombined 28.5 ± 6.6 (mean ± s.d.) cells (Fig. 2c). Col1a1-CreERT2 tdTomato+ cells coexpressed Pdgfrb-EGFP and Pdgfra (Fig. 2d–f). We found that 98.3 ± 2.9% (mean ± s.d.) of the Col1a1-CreERT2 tdTomato+ fibroblasts expressed Slc1a3 (GLAST) in the uninjured spinal cord (Fig. 2g and Supplementary Fig. 5e,f). We also detected the expression of Gjb6 in tdTomato+ perivascular fibroblasts, validating the scRNA-sequencing results (Supplementary Fig. 6a). Within the population of GLAST-expressing cells (tdTomato+), Col1a1 expression was highest in perivascular cells on large-caliber penetrating blood vessels and significantly lower on smaller-caliber blood vessels. GLAST-CreERT2 tdTomato+ cells associated with the microvasculature had no detectable Col1a1 expression (Supplementary Fig. 6b–d). Together, these results indicate that Col1a1 and GLAST are coexpressed in perivascular fibroblasts and can be genetically labeled with the Col1a1-CreERT2 and GLAST-CreERT2 transgenic mouse lines, whereas capillary-associated pericytes do not express Col1a1 and are captured only in GLAST-CreERT2 mice.
a, Col1a1+ perivascular cells can be labeled by tamoxifen-mediated genetic recombination in adult Col1a1-CreERT2 transgenic mice carrying R26R-tdTomato reporter alleles, in combination with the Pdgfrb-eGFP reporter line. b, Detection of Col1a1 mRNA in tdTomato+ cells in the uninjured spinal cord of Col1a1-CreERT2;R26R-tdTomato mice. c, Quantification of the average number of tdTomato+ cells per transverse spinal cord section (20-μm-thick) in uninjured Col1a1-CreERT2;R26R-tdTomato (mean ± s.d., 6.9 ± 1.6) and GLAST-CreERT2;R26R-tdTomato (mean ± s.d., 28.5 ± 6.6) mice. d,e, In the uninjured spinal cord, Col1a1-CreERT2 tdTomato+ cells express Pdgfrb-EGFP (white arrow) (d) and surround the blood vessel wall (marked with podocalyxin, white arrowhead) together with Pdgfrb-EGFP+ mural cells (yellow arrow) (e). f, Col1a1-CreERT2 tdTomato+ cells coexpress Col1a1 and the fibroblast marker Pdgfra (white arrows). The Pdgfra expression level in tdTomato+Col1a1+ fibroblasts is lower than in neighboring tdTomato−Col1a1−;Pdgfra+ cells (white arrowhead), which are presumably oligodendrocyte precursor cells. g, Slc1a3 (GLAST) is expressed by tdTomato+EGFP+ perivascular fibroblasts (white arrows) in the spinal cord of Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice. In addition, Slc1a3 expression is detected in tdTomato−EGFP− cells (white arrowheads), which are mostly astrocytes, as well as in tdTomato−EGFP+ perivascular cells (yellow arrows). The insets show the close-up view of a tdTomato+EGFP+Slc1a3+ perivascular fibroblast. Pdx, podocalyxin; WM, white matter; GM, gray matter; tdTom, tdTomato. Scale bars: 100 μm (d), 20 μm (f, g), 10 μm (e, b) and 5 μm (insets in g). Data are shown as mean ± s.d. n = 5 (GLAST-CreERT2) and n = 7 (Col1a1-CreERT2) animals in c. ****P < 0.0001 by two-sided, unpaired Student’s t test in c. Dashed lines in d outline the spinal cord gray matter. Cell nuclei are labeled with 4′,6′-diamidino-2-phenylindole dihydrochloride (DAPI). Images are representative of two independent experiments. All images show transverse sections. Source data are provided as a source data file.
Anatomical distribution of fibroblasts and GLAST+ pericytes
To determine the distribution of Col1a1+ and GLAST+ cells along the spinal cord vasculature, we used optical tissue clearing and imaged the thoracic spinal cord segments of recombined GLAST-CreERT2;R26R-tdTomato and Col1a1-CreERT2;R26R-tdTomato mice using light-sheet microscopy. The blood vessel lumen was labeled by transcardial perfusion with Alexa Fluor 647-conjugated BSA. Arteries and arterioles were marked with an antibody against SM22α, and veins and venules were marked with an antibody against von Willebrand factor (vWF) (Fig. 3a–d, Extended Data Fig. 1a,b and Supplementary Fig. 7a,b). We found 26.1 ± 1.8% (mean ± s.d.) Col1a1-CreERT2 tdTomato+ cells on higher-order penetrating arterioles and arteriolar branches and 73.3 ± 2.7% (mean ± s.d.) on venules, whereas they were absent from the capillary network (Fig. 3a,b,f and Extended Data Fig. 1b–e). GLAST-CreERT2 tdTomato+ cells were found in the same positions along arterioles and venules (Fig. 3c,d), confirming the overlap of GLAST and Col1a1 expression in fibroblasts (Figs. 1c–h and 2f,g and Supplementary Fig. 6b–d). In addition, the GLAST-CreERT2 line targeted perivascular cells on smaller arteriolar branches and the capillary bed (Fig. 3c–e and Extended Data Fig. 1a,c–e). Overall, 42.7 ± 5.3% (mean ± s.d.) of GLAST-CreERT2 tdTomato+ cells were associated with venules, 45.5 ± 9.9% (mean ± s.d.) were distributed along the arteriolar tree including the postarteriolar transitional zone and 11.8 ± 4.7% (mean ± s.d.) were associated with capillaries (Fig. 3c–e and Extended Data Fig. 1a,c–e). Gray matter regions of the spinal cord contain most arteriolar branches and a higher density of capillaries than white matter regions, whereas penetrating arterioles and venules mostly run in the white matter. These anatomical characteristics result in a higher number of GLAST-CreERT2 tdTomato+ cells than Col1a1-CreERT2 tdTomato+ cells in gray matter regions (Fig. 3a–d and Extended Data Fig. 1f,g). Both lines showed a higher number of tdTomato+ cells in the ventral than in the dorsal side of the spinal cord (Fig. 3a,c).
a,c, Distribution of Col1a1-CreERT2 tdTomato+ cells (a) and GLAST-CreERT2 tdTomato+ cells (c) revealed by volumetric imaging of 500-μm-thick uninjured spinal cord thoracic segments costained with antibodies against SM22α and vWF for labeling of arterioles/arteries and venules/veins, respectively. b,d, Enlarged images of the spinal cord dorsal gray matter, ventral gray matter and white matter (from the boxed regions in a and c) showing that Col1a1-CreERT2 tdTomato+ cells (b) do not extend as far along arteriolar branches as GLAST-CreERT2 tdTomato+ cells (d). The blood vessel lumen was labeled by transcardial perfusion with Alexa Fluor 647-conjugated BSA in a gelatin solution. e, GLAST-CreERT2 tdTomato+ cells expressing Pdgfrb-EGFP can be found in close contact with endothelial cells on capillaries and small microvessels, on blood vessels of intermediate diameter and surrounding tdTomato−EGFP+ mural cells on larger blood vessels. f, Col1a1-CreERT2 tdTomato+ fibroblasts express Pdgfrb-EGFP and are preferentially located along large-diameter blood vessels, surrounding tdTomato−EGFP+ mural cells in the spinal cord gray matter (GM) and white matter (WM). The inner diameter of blood vessels in e and f was measured in formaldehyde-fixed, cryopreserved tissue sections. Scale bars: 200 μm (a, c), 100 μm (b, d) and 5 μm (e, f). Cell nuclei are labeled with DAPI. Images are representative of three independent experiments. All images show transverse sections.
Ultrastructural features of fibroblasts and GLAST+ pericytes
We used immunogold labeling to identify GLAST-CreERT2 tdTomato+ and Col1a1-CreERT2 tdTomato+ cells in the uninjured spinal cord at the ultrastructural level. Along large arterioles, both lines labeled cells embedded in the outermost layer of the basal lamina, surrounding vSMCs, with a light cytoplasm, an oval-shaped nucleus, large light vesicular organelles and basal lamina protrusions extending into astrocytic endfeet (Fig. 4a–f), classifying them as perivascular fibroblasts26. On the venular side, immunogold-labeled tdTomato+ cells from both transgenic lines adhered to the vascular basal lamina and presented a light cytoplasm and membrane protrusions toward the abluminal side. Frequently, we observed labeled cells directly juxtaposed to endothelial cells, only separated by the basement membrane (Fig. 4g–l). In addition, the GLAST-CreERT2 transgenic line labeled perivascular cells in the transitional zone between smaller arteriole branches and the capillary bed, as well as in the capillary bed (Fig. 3e). These cells were embedded within the vascular basal lamina, juxtaposed to endothelial cells (Fig. 4m)8,10. At positions where processes intersected, GLAST-CreERT2 tdTomato+ cells were commonly located abluminal to other, nonrecombined pericytes (Fig. 3e)8. When transitioning to the capillary bed, GLAST-CreERT2 tdTomato+ cells no longer intersected with other pericytes. A three-dimensional (3D) serial reconstruction of 13.2-µm capillary length revealed that GLAST-CreERT2 tdTomato+ cells were directly abutting the endothelial tube and shared the endothelial basement membrane, like neighboring nonrecombined pericytes (Fig. 4m–p). Interestingly, while nonrecombined pericytes presented a classic morphology with finger-shaped processes extending radially27, GLAST-CreERT2 tdTomato+ cells covered the endothelial surface more homogeneously (Fig. 4o–r). Moreover, the cytoplasm of GLAST-CreERT2 tdTomato+ pericytes was less electron dense compared to neighboring pericytes (Fig. 4m). In summary, both GLAST-CreERT2 and Col1a1-CreERT2 transgenic lines target perivascular fibroblasts with similar ultrastructural features and perivascular position on larger arterioles and venules. In addition, the GLAST-CreERT2 line targets a population of GLAST+Col1a1− cells, which are embedded in the vascular basement membrane along capillaries and the arteriole–capillary transitional zone. Based on their gene expression profile (Fig. 1e) and location within the vascular basal lamina juxtaposed to the endothelial cells (Fig. 4m–p), these cells can be classified as pericytes14.
a–l, Immunogold electron microscopy images showing GLAST-expressing (a–c, g–i) and Col1a1-expressing (d–f, j–l) cells on two different arterioles (a–f) and venules (g–l). Immunogold particles label GLAST-CreERT2 tdTomato+ and Col1a1-CreERT2 tdTomato+ perivascular cells (pseudocolored red). a,b,d,e, Both GLAST- and Col1a1-expressing cells have an oval nucleus and thin processes surrounding the vascular wall, including vSMCs. GLAST-expressing (a–c) and Col1a1-expressing (d–f) cells show similar structural features, including basal lamina protrusions (arrows), bright cytoplasm and coated vesicles (asterisk). High-magnification micrographs of the insets in a and d are shown in b and e, respectively. c,f, A portion of a GLAST-expressing cell (c) and a Col1a1-expressing cell (f) in two other arterioles. The same morphological features are observed in GLAST- and Col1a1-expressing cells on venules: oval to round nucleus, thin processes surrounding the vessel, bright cytoplasm (g, j); basal lamina protrusions (arrows) and coated vesicles (asterisk) (h, i, k, l). i,l, Specific for the venules are areas where GLAST- and Col1a1-expressing cells are directly apposed to endothelial cells with only the basal membrane separating them. m–r, 3D reconstruction of a GLAST-expressing cell on a capillary (tdTomato+ cell; pseudocolored red) in comparison to two pericytes (pseudocolored green). m,n, The 3D model spans a length of 13.2 µm of the capillary. o,p, Both the GLAST-expressing cell and the GLAST− pericytes are embedded in the vascular wall directly apposing the endothelial tube, occupying a mutually exclusive space with little overlap. At places where they overlap, the GLAST-expressing cell is always abluminal. q,r, Unwrapped GLAST-expressing and GLAST− pericytes to visualize process morphology. GLAST-expressing cells (q) have a distinctly wide appearance occupying a larger surface of the blood vessel wall and fewer, less well-defined finger-like extensions compared to neighboring GLAST− pericytes (r). N, nucleus; L, lumen; EC, endothelial cell; PC, GLAST− pericyte; IC, pericyte–vSMC intermediary cell; asterisk, coated vesicle; double arrow, basal lamina protrusion. Scale bars: 5 µm (a, d, g, j) and 1 µm (b, c, e, f, h, i, k, l, m). Images are representative of two independent experiments. All electron microscopy images show transverse sections.
Region-dependent contribution to fibrotic scarring
To investigate the contribution of GLAST+ pericytes and perivascular fibroblasts to scar-forming fibroblasts in the context of SCI, we recombined GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP and Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice with tamoxifen, followed by a clearing period to ensure specificity3,28, and generated complete spinal crush lesions at a low thoracic level (Fig. 5a). GLAST-expressing pericytes and fibroblasts contributed 77.1 ± 8.2% and 89.7 ± 7.0% (mean ± s.d.) to all PDGFRβ+ scar-forming fibroblasts at 5 and 14 days after SCI, respectively (Fig. 5b–d), as observed previously10. In comparison, perivascular fibroblasts, targeted by the Col1a1-CreERT2 transgenic line, contributed to 37.0 ± 11.3% and 45.9 ± 15.6% (mean ± s.d.) of all PDGFRβ+ scar-forming fibroblasts at 5 and 14 days after SCI, respectively (Fig. 5b–d). These results indicate that, although virtually all scar-forming fibroblasts upregulate Col1a1 after SCI (Supplementary Fig. 8a–d), only a fraction of these cells are derived from Col1a1-expressing perivascular fibroblasts.
a, SCI lesion model (complete spinal crush at thoracic level 10 (T10)) and experimental timeline used for lineage tracing of GLAST- and Col1a1-expressing perivascular cells. b,c, Distribution of tdTomato+EGFP+ and tdTomato−EGFP+ cells in GLAST-CreERT2 and Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice at 5 days (b) and 14 days (c) after the complete spinal crush. The Pdgfrb-eGFP reporter line drives EGFP expression in PDGFRβ-expressing perivascular and stromal cells. d, Percentage of PDGFRβ+ cells that express tdTomato out of the total PDGFRβ+ stromal cells in the lesion core at 5 and 14 days after injury. e, Percentage of tdTomato+PDGFRβ+ cells out of the total PDGFRβ+ stromal cells in gray and white matter regions of the spinal cord at 14 days after injury. f, Lesion model (mild contusion at T9) and experimental timeline used for lineage tracing of GLAST- and Col1a1-expressing perivascular cells. g,h, Distribution of tdTomato+EGFP+ and tdTomato−EGFP+ cells in GLAST-CreERT2 and Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice at 5 days (g) and 14 days (h) after the mild spinal cord contusion. i, Percentage of PDGFRβ+ cells that express tdTomato out of the total PDGFRβ+ stromal cells in the lesion core at 5 and 14 days after injury. j, Percentage of tdTomato+PDGFRβ+ cells out of the total PDGFRβ+ stromal cells in gray and white matter regions of the spinal cord at 14 days after injury. All scale bars: 100 µm. Data are shown as mean ± s.d. Crush: GLAST-CreERT2 n = 3 (5 days) and n = 7 (14 days), Col1a1-CreERT2 n = 6 (5 days) and n = 5 (14 days) animals in d and n = 7 (GLAST-CreERT2 14 days) and n = 4 (Col1a1-CreERT2 14 days) animals in e. Contusion: GLAST-CreERT2 n = 5 (5 days) and n = 6 (14 days), Col1a1-CreERT2 n = 4 (5 days) and n = 4 (14 days) animals in i and n = 6 (GLAST-CreERT2 14 days) and n = 4 (Col1a1-CreERT2 14 days) animals in j. ***P = 0.0002, ****P < 0.0001 (d), *P = 0.0356, NS (no significance) = 0.1096 (e), ****P < 0.0001 (i), NS = 0.9804, ****P < 0.0001 (j) by one-way analysis of variance (ANOVA) followed by Šídák’s multiple-comparisons test. Dashed lines in b, c, g and h outline the gray matter–white matter border. Cell nuclei are labeled with DAPI. Images are representative of two independent experiments. All images show sagittal sections. Source data are provided as a source data file.
While GLAST-CreERT2 tdTomato+EGFP+ cells were distributed homogeneously throughout the fibrotic scar tissue, we observed that Col1a1-CreERT2 tdTomato+EGFP+ cells contributed significantly less to scar tissue in gray matter regions compared to white matter regions (Fig. 5b,c,e and Supplementary Fig. 8c,d). These observations suggest that perivascular fibroblasts, which are primarily located along large blood vessels in the spinal cord white matter, react locally to injury. In turn, GLAST+ pericytes, which are predominantly located in the spinal cord gray matter, contribute more extensively to gray matter scarring.
To validate the regional differences in scar contribution between GLAST+ pericytes and perivascular fibroblasts, we generated mild contusions (40 kdyn) at a low thoracic level, which caused fibrotic scar tissue formation in the central gray matter and immediately adjacent white matter regions (Fig. 5f–h). GLAST-CreERT2 tdTomato+ cells contributed to 94.7 ± 2.8% and 96.8 ± 2.4% (mean ± s.d.) of all PDGFRβ+ stromal fibroblasts at 5 and 14 days after contusion lesions, respectively (Fig. 5g–i). Again, GLAST-CreERT2 tdTomato+EGFP+ cells showed a similar contribution to fibrotic scarring in gray and white matter regions (Fig. 5g,h,j). In turn, fate-mapped Col1a1+ fibroblasts contributed to 35.8 ± 4.5% and 35.0 ± 2.5% (mean ± s.d.) of all PDGFRβ+ scar-forming fibroblasts at 5 and 14 days after contusion, respectively (Fig. 5g–i). Once more, we observed that the contribution of Col1a1-CreERT2 tdTomato+EGFP+ cells to fibrotic scarring was more pronounced in white than in gray matter regions (Fig. 5g,h,j).
Next, we asked whether the contribution of perivascular fibroblasts to fibrotic scar tissue can be compensated by GLAST+ pericytes. For this, we crossed Col1a1-CreERT2 mice to Rasless mice (Col1a1-CreERT2;Rasless), in which injury-induced proliferation of perivascular fibroblasts is inhibited through cell-specific deletion of floxed K-Ras in mice with H-Ras and N-Ras null alleles upon tamoxifen-induced genetic recombination3,6,10 (Extended Data Fig. 2a). Both Cre+ and CreWT animals were administered tamoxifen before injury, but CreWT animals did not undergo genetic recombination and served as controls.
Inhibition of proliferation in Cre+ animals reduced the density of scar-forming PDGFRβ+ fibroblasts by 29% compared to CreWT control animals (Extended Data Fig. 2b,c). We did not observe a complete abolishment of scar-forming stromal cell proliferation leading to improper wound healing and the formation of a tissue defect, as reported previously for similar experiments using GLAST-CreERT2;Rasless mice3,6. The partial inhibition of fibrotic scarring by rendering Col1a1+ perivascular fibroblasts unable to proliferate indicates their specific contribution to fibrotic scar tissue.
Together, these results suggest that scar-forming perivascular cells are recruited locally after SCI, with GLAST+ pericytes contributing to fibrotic scarring preferentially in gray matter regions and fibroblasts in the white matter of the spinal cord.
Local recruitment of fibroblasts and pericytes by inflammation
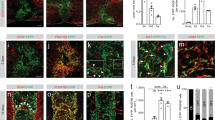
Fibrosis is mediated by inflammation4,29,30. To investigate the recruitment of GLAST+ pericytes and perivascular fibroblasts in response to local inflammation, we injected lipopolysaccharide (LPS), a potent activator of monocytes/macrophages and microglia31,32, into the spinal cord gray or white matter of recombined GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP and Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice (Fig. 6a). Focal injection of LPS resulted in comparable recruitment of MAC2+ macrophages/microglia and accumulation of Pdgfrb-EGFP+ stromal fibroblasts at the injection site in both gray and white matter regions (Fig. 6b–f). Interestingly, only GLAST-CreERT2 tdTomato+ cells contributed to stromal fibroblasts in the gray matter, whereas the contribution of Col1a1-CreERT2 tdTomato+ cells was negligible at 5 days after injection (Fig. 6b,c,g). In contrast, both lines contributed substantially to stromal fibroblasts after LPS injections into the white matter (Fig. 6d,e,g). The recruitment of pericytes and fibroblasts after gray and white matter LPS injections did not differ regarding cell proliferation, upregulation of Col1a1 or detachment from the vasculature (Supplementary Fig. 9a–g). In summary, perivascular fibroblasts and GLAST+ pericytes are recruited in response to focal inflammation. LPS injection caused minimal tissue disruption; under these circumstances, the local recruitment of fibroblasts and pericytes was restricted to white and gray matter regions, respectively, corroborating our results after SCI (Fig. 5).
a, Schematic outline and experimental timeline used for lineage tracing of GLAST- and Col1a1-expressing perivascular cells following LPS injection into the uninjured spinal cord gray matter and white matter. b,c, Stereotactic injection of LPS into the spinal cord gray matter of GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP (b) and Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP (c) mice triggers the recruitment of MAC2+ macrophages/microglia and local accumulation of tdTomato+EGFP+ stromal fibroblasts at 5 days after injection. b′, Close-up view of the boxed region in b showing that most Pdgfrb-EGFP+ stromal fibroblasts are tdTomato+ (yellow arrowheads), with only a few tdTomato−EGFP+ cells (white arrowhead) located off blood vessels (podocalyxin+). c′, Magnified image of the boxed region in c showing that almost all Pdgfrb-EGFP+ stromal fibroblasts are tdTomato− (white arrowheads), indicating that Col1a1-expressing perivascular fibroblasts (tdTomato+EGFP+ cells; yellow arrowheads) do not contribute substantially to gray matter scarring. d,e, Focal injection of LPS into the spinal cord ventral white matter of GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP (d) and Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP (e) mice leads to recruitment of MAC2+ macrophages/microglia and local accumulation of tdTomato+EGFP+ stromal fibroblasts at 5 days after injection. d′,e′, Magnified images of the boxed regions in d and e showing several tdTomato+EGFP+ stromal fibroblasts (yellow arrowheads) and tdTomato−EGFP+ stromal cells (white arrowheads) in GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP (d′) and Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eEGFP (e′) mice. f,g, Quantification of the inflammatory response by MAC2+ cell area (f) and percentage of tdTomato+PDGFRβ+ cells out of the total PDGFRβ+ stromal cells (g) at 5 days after intraspinal injection of LPS into the gray matter or white matter of GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP (GLAST-CreERT2) and Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP (Col1a1-CreERT2) mice. (b′, c′, d′, e′) Upper panel: tdTom, EGFP and Pdx labelling; lower panel: tdTom and Pdx labelling. Scale bars: 500 µm (b, c, d, e) and 100 µm (b′, c′, d′, e′). Data are shown as mean ± s.d. Col1a1-CreERT2: n = 3 (gray matter), n = 4 (white matter); GLAST-CreERT2: n = 4 (gray matter), n = 3 (white matter) animals in f. Col1a1-CreERT2: n = 3 (gray matter), n = 4 (white matter); GLAST-CreERT2: n = 7 (gray matter), n = 6 (white matter) animals in g. NS (gray matter) = 0.5719, NS (white matter) = 0.9788 (f) and NS = 0.1624, ***P = 0.007, ****P < 0.0001 (g) by one-way ANOVA followed by Tukey’s multiple-comparisons test. Dashed lines in b–e outline the spinal cord gray matter. Images are representative of two independent experiments. All images show transverse sections. Source data are provided as a source data file.
Injury-induced myofibroblast generation
To investigate transcriptional changes underlying the fibrotic injury response, we lineage traced GLAST+ pericytes and fibroblasts using GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice and FACSorted tdTomato+EGFP+ and tdTomato−EGFP+ cells from lesion sites at 3 and 5 days after a complete spinal crush. Single cells were sequenced, and data were combined with the dataset from the uninjured spinal cord (Fig. 1). Uniform Manifold Approximation and Projection (UMAP) of the integrated data of all tdTomato+EGFP+ cells from the uninjured and injured spinal cord resulted in five connected clusters encompassing pericytes, activated pericytes, myofibroblasts, activated fibroblasts and fibroblasts (Fig. 7a,b and Extended Data Fig. 3a,b). The fibroblast marker Pi16 (ref. 33) was highly expressed in perivascular fibroblasts before injury (Fig. 7b,c and Extended Data Fig. 3a). Robust Rgs5 and Atp13a5 expression identified pericytes in uninjured tissue (Fig. 7b,c), whereas Acta2 expression identified myofibroblasts. All fibroblast populations expressed Lum and Col8a1 (Fig. 7b). Col8a1 was expressed at low levels in perivascular fibroblasts before injury and highly in activated fibroblasts after injury (Fig. 7b). Next, we performed pseudotime analysis, defining two starting points: uninjured pericytes (highest Atp13a5 expression) and uninjured fibroblasts (highest Pi16 expression) (Fig. 7c). The analysis revealed two trajectories that merged in the myofibroblast cluster (Fig. 7c), representing the fibroblast (Fig. 7d) and pericyte (Fig. 7e) branches. The common endpoint of the two branches was activated fibroblasts (Fig. 7c–e). The fibroblast branch showed high expression of the ECM-associated genes Fn1, Col5a1, Col8a1 and Postn, which gradually increased along the trajectory, whereas Acta2 expression peaked in the myofibroblast cluster (Fig. 7f and Extended Data Fig. 3c). Along the pericyte branch, pericyte marker gene expression (Atp13a5, Cspg4, Kcnj8) rapidly decreased during the transition from pericytes through activated pericytes to myofibroblasts, whereas the cells simultaneously acquired a fibroblast gene expression signature (Col1a1, Col8a1, Fn1) (Fig. 7g and Extended Data Fig. 3c). Despite this rapid change in gene expression, we found a small population of cells that coexpressed the pericyte marker genes Atp13a5 and Kcnj8 (ref. 11) and the fibroblast markers Col1a1 and Col5a1, representing a transition stage (Fig. 7h and Supplementary Fig. 10a–f). GLAST-expressing perivascular cells proliferate, detach from the blood vessel wall and migrate into the lesioned tissue3,6. Accordingly, a large fraction of activated pericytes and myofibroblasts expressed genes associated with the S and G2/M cell-cycle phases, whereas perivascular fibroblasts, pericytes and activated fibroblasts were mostly postmitotic (Extended Data Fig. 3d). With the detachment from the vascular wall, pericytes lose their pericyte identity and rapidly acquire a myofibroblast phenotype. In agreement, tdTomato+EGFP+ cells coexpressing pericyte and fibroblast markers were only sporadically detected associated with blood vessels close to the lesion, whereas Col1a1 was highly expressed by stromal myofibroblasts in the lesion core (Supplementary Fig. 11a,b).
a, UMAP plot of tdTomato+EGFP+ cells isolated from the uninjured and injured spinal cord of GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice at 3 and 5 days after injury grouped by injury state (left) and cell population (right). b, Violin plots showing the expression levels of cell population-specific genes. c, Left, feature plots showing the expression of the fibroblast marker Pi16 and pericyte marker Atp13a5. Right, pseudotime trajectory analysis of tdTomato+EGFP+ cells isolated from the uninjured and injured spinal cord at 3 and 5 days after injury. Dashed lines outline cell populations. High expression of Pi16 and Atp13a5 were used to identify pseudotime starting points in the fibroblast and pericyte populations, respectively. d,e, Highlight of the pseudotime trajectory of the fibroblast branch (d) and pericyte branch (e). f, Gene expression profile of Acta2, Col5a1 and Postn along pseudotime within the fibroblast branch. g, Gene expression profile of Atp13a5, Cspg4 and Col1a1 along pseudotime within the pericyte branch. h, Left, violin plots showing the gene expression levels of Atp13a5 and Col1a1 per cell population and coexpression graph of Col1a1 and Atp13a5. Right, violin plots showing the gene expression levels of Kcjn8 and Col5a1 per cell population and coexpression graph of Col5a1 and Kcnj8. Each dot in coexpression graphs represents a single cell, colored according to the injury state. i,j, GO analyses based on upregulated genes of activated pericytes and myofibroblasts (i) and upregulated genes of myofibroblasts and activated fibroblasts (j). Venn diagrams represent the overlap of common GO terms and unique GO terms between the indicated populations. Relevant GO terms per comparison are indicated below. k, Dot plot of upregulated marker genes per fibroblast population. d3, day 3; d5, day 5; Per, pericyte; aPer, activated pericyte; pFib, perivascular fibroblast; mFib, myofibroblast; aFib, activated fibroblast; NA, nonattributed; TNF, tumor necrosis factor; MHC, major histocompatibility complex; IGF, insulin-like growth factor; IGFBP, insulin-like growth factor-binding protein; SLC, solute carrier; HIF-1, hypoxia-inducible factor. Source data are provided as a source data file.
Activated pericytes and myofibroblasts are highly proliferative (Extended Data Fig. 3d) and commonly express genes related to transforming growth factor-β (TGFβ) regulation of ECM, brain-derived neurotrophic factor and epidermal growth factor receptor 1 signaling. GO analysis revealed 15 terms specifically enriched in activated pericytes, the most relevant of which were related to active immunity and lipid metabolism (Fig. 7i). In addition to oncostatin M and platelet activation, the GO term ‘protein metabolism’ was significantly enriched in myofibroblasts, indicative of cell activation, in line with higher expression of ribosomal genes (Fig. 7i and Extended Data Fig. 3e,f). Myofibroblasts consequently differentiated into activated fibroblasts (Fig. 7c), which showed more pronounced expression of genes related to collagen biosynthesis, cytokine interaction, angiogenesis and insulin-like growth factor-related signaling (Fig. 7j and Extended Data Fig. 3e). In turn, myofibroblasts showed higher expression of genes related to proliferation, protein metabolism, hypoxia and smooth muscle contraction. Myofibroblasts and activated fibroblasts shared GO terms related to TGFβ signaling and ECM organization (Fig. 7j and Extended Data Fig. 3e).
After injury, tdTomato+EGFP+ cells, which comprise pericytes and fibroblasts in the uninjured spinal cord, generated a new cell cluster of myofibroblasts and activated fibroblasts, resulting in gene expression changes in 2,038 genes (Extended Data Fig. 4a,b). In contrast, tdTomato−EGFP+ cells retained their pericyte and vSMC identity after injury, with some pericytes exhibiting an activated gene signature (Extended Data Fig. 4c). Accordingly, SCI led to expression changes in only 719 genes (Extended Data Fig. 4d). TdTomato−EGFP+ activated pericytes upregulated genes related to ECM deposition and angiogenesis and may, therefore, represent a population of pericytes that remain part of the blood vessel wall, promoting revascularization and contributing to fibrotic ECM production upon injury (Extended Data Fig. 4e,f and Supplementary Fig. 11a,b).
To differentiate perivascular fibroblasts, myofibroblasts and activated fibroblasts, we directly compared their expression of marker genes (Fig. 7k). Perivascular fibroblasts expressed elevated levels of Pi16, Lum, Dcn, Fbln1 and Mfap5. Myofibroblasts showed strong expression of Acta2, Tagln, Thbs2, Itga5 and Tnc, whereas activated fibroblasts displayed higher expression of Col3a1, Col5a1, S100a4 and Igfbp5 (Fig. 7k). Five days after SCI, most tdTomato+EGFP+ cells in the lesion expressed Col1a1, αSMA and SM22α, identifying them as myofibroblasts. Additionally, we identified a subset of tdTomato+EGFP+Col1a1+ cells with low αSMA and SM22α expression, which are presumably activated fibroblasts (Extended Data Fig. 5a–d and Supplementary Fig. 11a–c). VSMCs, also expressing αSMA and SM22α, were distinguished from myofibroblasts and activated fibroblasts by their lack of Col1a1 expression (Supplementary Fig. 11b).
Together, our data show that, upon SCI, GLAST-expressing pericytes and fibroblasts give rise to myofibroblasts and activated fibroblasts, which populate the lesion and produce abundant fibrotic ECM. Activated fibroblasts and myofibroblasts show distinct fibroblast marker expression, suggesting functional differences between these stromal cell populations during CNS wound repair.
Discussion
Fibrotic scar tissue formation is a common response to numerous CNS injuries and diseases in mice and humans5,10. Previous research has identified perivascular cells, including perivascular fibroblasts and GLAST-expressing perivascular cells (also known as type A pericytes), as primary contributors to stromal fibroblasts in CNS lesions3,4,6,10,12,34,35,36. Recent insights from single-cell transcriptomics data revealed that, beyond pericytes, GLAST (Slc1a3) is also expressed by a subset of fibroblasts in the adult mouse brain11, suggesting that GLAST-expressing perivascular cells may be more heterogeneous than previously assumed.
Indeed, our single-cell gene expression profiling revealed that GLAST-expressing perivascular cells can be transcriptionally defined as a subset of pericytes and a population of fibroblasts. GLAST+Col1a1− pericytes are located in the postarteriolar transitional zone and the capillary bed; they are most abundant in gray matter regions because, anatomically, the gray matter contains a larger capillary network compared to the white matter regions of the spinal cord37,38. However, GLAST+ pericytes are distinct from classic GLAST− pericytes at the ultrastructural level, exhibiting different morphological features and electron density. These observations are in line with reports describing morphological pericyte heterogeneity in the cerebral cortex39,40.
We observed local recruitment of pericytes and perivascular fibroblasts in response to inflammation and SCI. Perivascular fibroblasts, predominantly located around large arterioles and venules in the spinal cord white matter, contributed more to scarring in the white matter than in gray matter regions, consistent with the results of a previous study indicating the pivotal role of fibroblasts in white matter scarring following EAE4. However, accurately assessing the quantitative contribution of perivascular fibroblasts to fibrotic scarring in EAE may necessitate validation with an inducible Col1a1-CreER line, given the expression of Col1a2 in some CNS pericytes and vSMCs11. The Col1a1-CreERT2 line used in this study recombined, on average, 81.5% of all Col1a1-expressing cells, with higher efficacy observed in white than in gray matter regions (Supplementary Fig. 5d), potentially underrepresenting gray matter perivascular fibroblasts and their contribution to fibrosis.
While only a small subset of perivascular cells express Col1a1 under homeostatic conditions in the adult mouse spinal cord, we observed that virtually all scar-forming fibroblasts express Col1a1 after SCI (Fig. 2c and Extended Data Fig. 4), corroborating previous observations with Col1a1-GFP reporter mice12. However, our lineage tracing revealed that only approximately 40% of scar-forming fibroblasts originate from Col1a1-expressing perivascular fibroblasts. Conversely, GLAST-expressing perivascular cells, encompassing a subset of pericytes and perivascular fibroblasts, gave rise to most scar-forming fibroblasts after SCI, aligning with previous observations6,10 and implying the contribution of GLAST+Col1a1− pericytes to fibrotic scarring in the gray matter. This interpretation is supported by our single-cell expression analysis showing a transcriptional trajectory from GLAST+ pericytes to myofibroblasts and activated fibroblasts. Direct proof of a lineage relationship between GLAST+ pericytes and scar-forming fibroblasts is currently limited owing to the inaccessibility of inducible CreER lines that specifically and efficiently label the GLAST+ pericyte fraction. Readily available CreER lines targeting pericytes also label fibroblasts (Pdgfrb-CreERT2) or are inefficient (NG2-CreERT2)12. Another limitation is that GLAST is expressed in only a small subset of pericytes, and many pericyte markers are not homogeneously expressed by all pericytes (Fig. 1e,g and Extended Data Fig. 4f). A potential invasion of meningeal fibroblasts is rather limited in nonpenetrating CNS lesions but cannot be formally excluded. The ability of GLAST+ perivascular cells to generate stromal fibroblasts seems to be governed by cell-intrinsic factors, as GLAST− perivascular cells sharing the same microenvironment do not contribute to stromal fibroblasts after injury. Our single-cell transcriptomics data affirm that GLAST-expressing perivascular cells give rise to fibrotic ECM-producing myofibroblasts after SCI. Pseudotime analyses suggest the transcriptional convergence of GLAST+ pericytes and perivascular fibroblasts toward myofibroblast generation, corroborating our previous findings regarding the role of GLAST+ perivascular cell-derived progeny in wound contraction and fibrotic ECM deposition, which are crucial for wound closure after SCI3. Notably, GLAST+ fibroblasts express Pi16, indicative of a conserved fibroblast subset signature across organs33. Moreover, we identified a subset of GLAST− pericytes that activate a fibrotic gene signature after injury, highlighting the involvement of other perivascular cells, in addition to scar-forming myofibroblasts, in fibrotic ECM secretion.
Our results reveal that GLAST+ pericytes and Col1a1+ perivascular fibroblasts contribute to fibrotic scar tissue in a region-dependent manner after SCI. This emphasizes the role of locally recruited perivascular cells in fibrotic tissue formation, which may be vital for designing therapeutic strategies for fibrotic diseases.
Methods
Transgenic mice
Generation of Col1a1-CreER T2 transgenic mice
C57BL/6J congenic mice that express a tamoxifen-dependent Cre recombinase under the control of the Col1a1 promoter (Tg(Col1a1-creERT2)6.1.Ics) were generated at PHENOMIN-Institut Clinique de la Souris, Strasbourg, France. The line was generated by microinjection of a circular murine BAC transgene (RP23-53E2) previously modified by recombineering at the ATG of the Col1a1 gene of a rabbit β-globin intron-CreERT2 cassette, followed by a flipped PGK-Tn5-Kana/NeoR cassette. The genetic background of the fertilized oocytes was C57BL/6J–SJL/J. The founder was then backcrossed in a C57BL/6J genetic background. The 6.1 subline was analyzed by reverse transcription–qPCR screening and cryopreserved. Available information on the characterization of the mouse line is accessible at http://mousecre.phenomin.fr/synthesis/152. This line was deposited in the Infrafrontier repository (https://www.infrafrontier.eu/) under the identifier EM:14947.
Crossing of reporter lines
GLAST-CreERT2 (ref. 41) or Col1a1-CreERT2 mice were crossed to the Rosa26-tdTomato Cre reporter line42 (B6.Cg-Gt(ROSA)26Sor tm14(CAG-tdTomato)Hze/J, JAX stock no. 007914) to obtain GLAST-CreERT2;R26R-tdTomato or Col1a1-CreERT2;R26R-tdTomato mice, respectively. For some experiments, these mice were further crossed to the Pdgfrb-eGFP reporter line43 (Tg(Pdgfrb-EGFP)JN169Gsat/Mmucd, RRID: MMRRC 031796-UCD) to generate GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP or Col1a1-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice. For one experiment, Col1a1-CreERT2 mice were further crossed to Rasless mice44 to generate Col1a1-CreERT2;Rasless mice. All Rasless mice used were homozygous for H-Ras and N-Ras null alleles and homozygous for floxed K-Ras alleles; Cre− mice were used as controls.
All strains were backcrossed to the C57BL/6J genetic background. Both male and female mice aged >8 weeks were included in the experiments, except when stated otherwise. All animals were housed in groups and kept in standardized individually ventilated cages (Scanbur), with controlled humidity (50%) and temperature (22 °C) and a 12-h/12-h light/dark cycle with water and food ad libitum. All experimental procedures were carried out in accordance with the Swedish and European Union law and guidelines and approved by the regional ethical committee (Stockholm Ethical Committee/‘Stockholms Djurförsöksetiska Nämnd’). Based on their genotype, mice were randomly assigned to the experimental groups.
Genetic labeling of transgenic mice
Recombination of CreERT2 transgenic lines was induced by a daily intraperitoneal (i.p.) injection of 2 mg of tamoxifen (Sigma-Aldrich, T5648, 20 mg ml−1 in 9:1 corn oil/ethanol) for five consecutive days, followed by a 7-day clearing period28.
Surgical procedures
For all injury models, mice were anesthetized with 4% isoflurane until sedated, followed by 2% isoflurane during surgery. All animals received preemptive and postoperative pain relief (Temgesic (buprenorphine), Schering-Plough, 0.1 mg kg−1 body weight and Rimadyl (carprofen), Pfizer, 5 mg kg−1 body weight; subcutaneous injection), as well as local anesthesia before spinal lesions or intraspinal injections (Xylocaine (lidocaine), AstraZeneca, 10 mg ml−1 and Marcaine (bupivacaine), AstraZeneca, 2 mg kg−1 body weight). Viscotears (carbomer) gel (2 mg g−1, Novartis) was topically applied to prevent the eyes from drying out. After surgery, the animals were placed in a prewarmed cage under close observation and transferred to their home cage thereafter.
Complete spinal cord crush
A laminectomy was conducted to expose the spinal cord at T10. After the application of local anesthesia, the spinal cord was fully crushed for 2 s with Dumont No. 5 forceps (11295-00, Fine Science Tools)10,45. Animals received postoperative analgesia as described above and were treated with antibiotics (Hippotrim vet., sulfadiazine 200 mg ml−1, trimethoprim 40 mg ml−1, 100 mg kg−1 body weight every 24 h; subcutaneous injection) to prevent bladder infection. To help the animals’ mobility, we supplemented the cage floor with a raised grid. Throughout the experiment, bladders were manually expressed two to three times per day to permit micturition. From 1 day before the surgery, the animals’ diet was supplemented with a high-energy nutritional supplement (DietGel Boost, Clear H2O) during the first week after surgery to support recovery and limit weight loss. The body weight was recorded before surgery, daily during the first week after injury and weekly thereafter. Only female mice were used for crush lesion experiments. Mice were killed at 3, 5, 7 and 14 days after injury.
Spinal cord contusion
For spinal cord contusion lesions, mice were placed on a stabilization platform for surgery. After laminectomy, the vertebral column was stabilized at T8 and T10 vertebrae with adjustable fine pitch forceps connected to three-joint surgery arms. After the application of local anesthesia, the mice received a 40-kdyn contusion centered on the T9 thoracic spinal cord using the Infinite Horizon impactor (IH-0400, PSI) with a 1.3-mm tip. Subsequently, the wounds were sutured, and mice were placed in a preheated cage for recovery and thereafter transferred to their home cage with an elevated floor grid. Bladders were manually expressed two to three times per day to permit micturition until mice regained bladder control (usually within 5 days after surgery in this mild contusion model). See above for postoperative analgesia.
Only female mice were used for contusion lesion experiments. Mice were killed at 5 and 14 days after injury.
Stereotactic injection of LPS into the spinal cord
LPS (from Escherichia coli, ALX-581-007, Enzo Life Sciences, 1 mg ml−1) was stereotactically injected into the low thoracic region (T12/T13) of the mouse spinal cord in deeply anesthetized uninjured adult animals. Mice were placed on a stabilization platform for surgery. After laminectomy, the vertebral column was stabilized with adjustable fine pitch forceps connected to three-joint surgery arms. Each mouse received three injections spaced 3 mm apart, with each injection delivering a total volume of 0.2 μl (200 ng of LPS per injection) using a 10-μl NanoFil syringe with a 36-gauge beveled needle tip (World Precision Instruments) coupled to a microinjector (UltraMicroPump III and Micro4 microsyringe pump controller, World Precision Instruments). After injection, the needle was left in place for an additional 2 min to allow diffusion and prevent backflow of the solution and then slowly withdrawn. Injections were carried out 0.35-mm lateral relative to the posterior median spinal vein and at 0.5-mm depth from the spinal cord surface to inject into the gray matter or at 1.3-mm depth to inject into the ventral white matter of the spinal cord. Both male and female mice were used for intraspinal injections. Mice were killed at 5 days after injection.
Isolation of different perivascular cell populations from the spinal cord
Preparation of single-cell cell suspensions and myelin removal were performed either using an adult brain dissociation kit (130-107-677, Miltenyi), followed by the Smart-seq3 and Smart-seq3xpress library preparation protocols, or adapted from previously published protocols, followed by the Smart-seq2 protocol45,46. In summary, GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP animals (uninjured and 3 and 5 days after spinal crush) were killed by i.p. injection of sodium pentobarbital (200 mg kg−1, 100 μl i.p., APL) and transcardially perfused with cold Hank’s balanced salt solution (HBSS) without calcium or magnesium (Invitrogen). The spinal cord was carefully dissected out of the vertebrae, and the crush injury segment was separated from the uninjured spinal cord tissue and treated separately thereafter. After the removal of the outer meningeal layers, the tissue was mechanically dissociated with a razor blade on a glass Petri dish and immediately transferred to cold HBSS. This step was followed by enzymatic digestion. Before Smart-seq2 library preparation, tissues were digested with papain at a final concentration of 8 U ml−1 with 80 Kunitz units of DNase I per ml (Sigma-Aldrich, D4263) in Ca2+/Mg2+-free piperazine-N,N′-bis(2-ethanesulfonic acid)/cysteine-based buffer (pH 7.4) for 40 min at 37 °C, 350 rpm on a ThermoMixer (Eppendorf). The tissue was carefully triturated and incubated for an additional 5 min. The cell suspension was passed through a 70-μm cell strainer (Corning), washed with minimum essential medium (MEM) with 1% BSA and spun down at 200g for 5 min at 4 °C. The cells were resuspended in MEM and centrifuged over a 90% Percoll gradient (GE Healthcare, 17-0891-01) at 250g for 15 min at 4 °C. Cells in the lipid layer and below were diluted five times in MEM with 1% BSA and centrifuged in a 15-ml tube at 250g for 10 min at 4 °C. All supernatants, including the lipid layer, were carefully removed. Before Smart-seq3 and Smart-seq3xpress library preparation, cells were enzymatically digested following the manufacturer’s instructions from the adult brain dissociation kit. Briefly, cells were incubated with enzymes for 15 min at 37 °C with agitation and then triturated by pipetting up and down with 1,000-μl tips and incubated for 10 min more. Digestion was stopped by adding 10 ml of MACS buffer (PBS with 0.5% FBS). Cells were filtered through a 70-μm cell strainer and centrifuged for 10 min at 300g at 4 °C.
Following the two protocols, pellets were resuspended in cold MACS buffer and magnetic myelin removal beads (Miltenyi Biotech, 130-096-433) and incubated for 15 min at 4 °C. The cells were washed and run over MACS magnetic separation columns (Miltenyi Biotech, 130-042-201) on a magnetic stand according to the manufacturer’s instructions. The cells in the flow through were collected, spun down, resuspended in FACS buffer (2% FBS in PBS) and kept on ice until further processing. For Smart-seq3xpress, cells were resuspended in pure PBS to avoid serum contamination in the lysis mix.
Single-cell sorting, library preparation, sequencing and quality control
The single-cell suspensions generated from GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP animals were subjected to FACS. Based on reporter gene expression, tdTomato+EGFP+ and tdTomato−EGFP+ cells were collected into 96- or 384-well plates with cell lysis buffer (Clonetech, 635013) or Smart-seq3/Smart-seq3xpress lysis buffer mix, followed by a brief centrifugation (1 min, 1,500 rpm) and stored at −80 °C until further processing. Cell sorting was performed on a FACSAria III cell sorter (BD Biosciences). Singlet discrimination was performed using plots for forward scatter (FSC-A versus FSC-H) and side scatter (SSC-W versus SSC-H), and dead cells were excluded by SYTOX Blue dead cell staining (Thermo Fisher Scientific, S34857). Single cells were processed to cDNA libraries according to the Smart-seq2, Smart-seq3 or Smart-seq3xpress protocols16,17,18. Smart-seq2 samples were sequenced on Illumina HiSeq 2500 (HiSeq Control Software 2.2.58/RTA 1.18.64) with a 1 × 51 setup using ‘HiSeq SBS Kit v4’ chemistry. Smart-seq3 and Smart-seq3xpress libraries were sequenced on an Illumina NextSeq 500 (Illumina NextSeq Control Software 2.2.0). Reads were then aligned to the mouse reference genome.
Single-cell gene expression analysis
All data were processed using Seurat (v4.3.0.1)47 and RStudio (v1.4.1717)48. Data from Smart-seq2, Smart-seq3 or Smart-seq3xpress sequencing were first preprocessed individually using Seurat package default parameters, with the first 20 principal components and 0.5 as the resolution. Cells expressing between 200 and 1,200 features and <20% of mitochondrial genes were retained for the analysis. The different datasets were then merged and integrated with Harmony (v0.1.1)49. To correct potential batch effects, we performed scaling with the feature number and percentage of mitochondrial and ribosomal content regression. Unbiased clustering was done using the first 30 principal components and 0.5 as the resolution. Astrocytes and immune cell contamination were removed by filtering data based on Gfap and Ptprc gene expression, respectively. Endothelial cells were removed based on high expression of Pecam1 and low expression of Pdgfrb.
Differential gene expression analyses were done using the Wilcoxon test, and genes with P < 0.05 and log2(fold change) > 0.25 were used for subsequent GO analyses. GO term analyses were done using Enrichr50. GO functions with fewer than five genes and a P value of >0.05 were excluded, and analyses were performed using the BioPlanet library.
Pseudotime analyses were performed using Monocle3 (v1.3.1). Normalized data from the Seurat object were used as input for pseudotime analyses and were processed as described in the Monocle3 tutorial. To decipher the pseudotime trajectory, we first used the entire dataset by defining two starting points using the Pi16 expression for the fibroblast branch and the Rgs5 expression for the pericyte branch. The two branches were then divided manually using the choose_graph_segments function, and gene expression was plotted using the plot_genes_in_pseudotime function. UMAPs for the comparison of day 0 and day 5 SCI tdTomato+EGFP+ or tdTomato−EGFP+ datasets were generated using the first 15 principal components. A cluster of Slc1a3+Pdgfra+ fibroblasts in the tdTomato−EGFP+ dataset (nonrecombined GLAST+ cells) was excluded from the analysis. Heat maps were generated with the pheatmap package51.
For cell-type comparison, our dataset from the uninjured spinal cord was integrated with data from uninjured brain mural cells from ref. 11 (GSE98816) using the Seurat IntegrateData function. Both datasets were preprocessed by selecting cells with >350 features and <10% mtRNA, in which only genes present in more than three cells were considered. For normalization and variance stabilization, we used sctransform52. For plots with gene expression levels, only mural cells (pericytes, vSMCs)11 were included and shown as UMAP projections of the first 15 dimensions (determined based on elbow plots) and a resolution of 0.5. For analysis, we used the Seurat standard workflow, UMAP dimension reduction with 15 dimensions (determined based on elbow plots) and a resolution of 0.5. Cells annotated as endothelial cells were removed before data integration52.
Tissue processing and immunohistochemistry
Animals were killed by i.p. injection of an overdose of sodium pentobarbital and transcardially perfused with cold PBS, followed by 4% formaldehyde in PBS. Spinal cords were dissected out and postfixed in 4% formaldehyde in PBS overnight at 4 °C and then cryoprotected in 30% sucrose. After embedding in OCT mounting medium and freezing on dry ice, spinal cords were sectioned with a cryostat (Leica, CM1860). Cryosections (20 μm) were then collected on alternating slides and stored at –20 or –80 °C. For immunostaining, sections were incubated with blocking solution (10% normal donkey serum in PBS, with 0.3% Triton X-100) for 1 h at room temperature and then incubated at room temperature overnight in a humidified chamber with primary antibodies diluted in 10% normal donkey serum. The primary antibodies used were GFP (1:2,000, chicken, Aves Labs, GFP-1020; 1:2,000, sheep, Bio-Rad, 4745-1051), RFP (red fluorescent protein; 1:250, chicken, Novus Biologicals, NBP1-97371), PDGFRβ (1:200, rabbit, Abcam, ab32570; 1:100, rat, eBioscience, 14-1402-82), podocalyxin (1:200, goat, R&D Systems, AF1556), αSMA (1:200, rabbit, Abcam, ab5694), SM22α (also known as transgelin; 1:500, rabbit, Abcam, ab14106), Ki67 (1:2,000, rat, eBioscience, 14-5698), NG2 chondroitin sulfate proteoglycan (1:200, rabbit, Millipore, AB5320; no Triton X-100) and MAC2 (also known as galectin-3; 1:500, rat directly conjugated to biotin, Cedarlane Labs, CL8942B).
All secondary antibodies used for immunohistochemistry were F(ab′)2 fragment affinity-purified antibodies purchased from Jackson Immunoresearch and diluted at 1:500: Alexa Fluor 488 donkey anti-chicken immunoglobulin Y (IgY) (703-546-155), Cy3 donkey anti-chicken IgY (703-166-155), Alexa Fluor 647 donkey anti-goat IgG (705-606-147), Alexa Fluor 594 donkey anti-goat IgG (705-585-147), Alexa Fluor 680 donkey anti-goat IgG (705-625-147), Alexa Fluor 488 donkey anti-rabbit IgG (711-546-152), Alexa Fluor 647 donkey anti-rabbit IgG (711-606-152), Alexa Fluor 680 donkey anti-rabbit IgG (771-625-152), Alexa Fluor 488 donkey anti-rat IgG (712-546-153), Alexa Fluor 647 donkey anti-rat IgG (712-606-153) and Alexa Fluor 488 donkey anti-sheep IgG (713-546-147). Biotinylated secondary antibodies were revealed with Alexa Fluor 594-conjugated streptavidin (1:500, Jackson Immunoresearch, 016-580-084) or Alexa Fluor 680-conjugated streptavidin (1:500, Jackson Immunoresearch, 016-620-084).
Cell nuclei were visualized with DAPI (1 μg ml−1 in PBS, Sigma-Aldrich, D9542). Sections were mounted using VECTASHIELD antifade mounting medium (Vector Labs, H-1000).
In situ hybridization—RNAscope
For the detection of RNA molecules, we performed RNAscope on formaldehyde-fixed spinal cord cryosections stored at –80 °C. In situ hybridization was performed following a modified version of the RNAscope Multiplex Fluorescent Reagent Kit v2 Assay (ACD Bio-Techne, 323100). Briefly, tissue sections were allowed to dry and equilibrate to room temperature, washed in PBS and baked for 30 min at 60 °C before postfixation in 4% formaldehyde in PBS for 15 min at 4 °C. Slides were subsequently dehydrated in 50%, 70% and 100% ethanol for 5 min each at room temperature. After drying, slides were first boiled for 10 s in distilled water, followed by 5 min in antigen retrieval solution, washed in distilled water and 100% ethanol at room temperature, and allowed to dry again. For further antigen accessibility, sections were incubated with protease III (ACD Bio-Techne, 322337) at 40 °C for 30 min. The samples were washed in distilled water and incubated at 40 °C for 2 h with the following RNAscope probes (ACD Bio-Techne): Pdgfrb (Mm-Pdgfrb-C3, 411381-C3), Pdgfra (Mm-Pdgfra, 480661), Col1a1 (Mm-Col1a1, 319371), Slc1a3 (Mm-Slc1a3-C3, 430781-C3), Atp13a5 (Mm-Atp13a5-C2, 417211-C2) and Gjb6 (Mm-Gjb6, 458811). C3 probes were diluted 1:50 in C1 probes or probe diluent. Subsequent amplification and detection were performed following the assay protocol. Probes were detected with Opal dyes 520 (Perkin Elmer, FP1487A), 570 (Perkin Elmer, FP1488A) or 650 (Perkin Elmer, FP1496A). TdTomato labeling was recovered by immunohistochemistry with an antibody against RFP (1:250, chicken, Novus Biologicals, NBP1-97371). Nuclei were stained with DAPI (1:5,000, 1 μg ml−1 in PBS, Sigma-Aldrich, D9542) before mounting the slides. The protocol was run in 1 day to preserve sample quality.
Light-sheet fluorescence microscopy
Blood vessel labeling
Animals were first transcardially perfused with cold PBS and then with 4% formaldehyde in PBS, followed by 5 ml of a 2% gelatin (Sigma-Aldrich, G1890) solution in PBS containing 0.5 mg of BSA conjugated to Alexa Fluor 647 (Thermo Fisher Scientific, A34785). Subsequently, perfused mice were kept in ice-cold water for at least 15 min for the gelatin to solidify. The labeled tissue was kept in 4% formaldehyde in PBS for further histological processing.
Optical tissue clearing
The tissue clearing protocol was adapted from ref. 53. After fixation with 4% formaldehyde in PBS for 3 days, spinal cords were optically cleared with 4% SDS at 50 °C for 3–5 days in a rotating hybridization incubator with daily renewal of the clearing solution. The cleared spinal cords were embedded in 4% agarose and washed with PBS with 0.1% Tween 20 detergent (PBS-T) overnight. The embedded spinal cords were incubated with primary antibodies against SM22α (1:500, rabbit, Abcam, ab14106) and vWF (1:100, sheep, Abcam, ab11713) diluted in PBS-T for 2 days at room temperature. After being washed with PBS-T overnight, samples were incubated with species-specific secondary antibodies conjugated to Alexa Fluor 488 (1:200, Jackson ImmunoResearch) in PBS-T for 2 days. The spinal cords were then washed and stored in PBS-T at room temperature until imaging.
Light-sheet imaging of cleared tissues
The embedded cleared tissues were incubated in a customized refractive index (RI) matching medium (Omnipaque 350, GE Healthcare) supplemented with 1 M urea (Sigma-Aldrich, U5378) and 40% d-sorbitol (Sigma-Aldrich, S1876; RI = 1.48) overnight. After RI matching, the samples were imaged with a light-sheet microscope (Zeiss Z.1). The acquired images were processed to achieve an isotropic resolution of 2.5 μm and visualized with napari54.
Quantitative assessment of cleared tissues
To quantify the percentage of tdTomato+ cells in different vessel compartments and between gray matter and white matter, a 500-μm-thick uninjured thoracic spinal cord segment was used for each sample. Counting was done manually in an interactive 3D view, assisted by a custom-built napari plug-in. The classification of arterioles, venules and capillaries was based on the SM22α and vWF signals and the relative position and blood vessel size, which were determined together by two observers.
Electron microscopy
Transcardial perfusion
Uninjured GLAST-CreERT2;R26R-tdTomato and Col1a1-CreERT2;R26R-tdTomato animals were killed by i.p. injection of an overdose of sodium pentobarbital and initially transcardially perfused with ice-cold 2% dextran in 0.1 M phosphate buffer (20 s), followed by a mixture of 4% formaldehyde and 0.1% glutaraldehyde in 0.1 M phosphate buffer for 15 min. The spinal cords were removed, left in the fixative solution overnight and stored in a 1:10 dilution of the same solution in 0.1 M phosphate buffer.
Postembedding and immunogold electron microscopy
Small blocks were dissected out from perfusion-fixed spinal cords and subjected to a freeze-substitution procedure as described previously55. Briefly, tissue blocks were cryoprotected in 4% glucose overnight, and the tissues were suspended in graded glycerol solutions (10%, 20% and 30% glycerol in 0.1 M phosphate buffer for 30 min in each gradient). After cryoprotection, the tissue blocks were rapidly frozen in propane cooled to −170 °C using liquid nitrogen before being subjected to freeze substitution. Samples were later embedded in methacrylate resin (Lowicryl HM20) and polymerized by ultraviolet irradiation below 0 °C. Serial sections were cut at 90–100 nm using an ultramicrotome (Reichert Ultracut S, Leica) and placed on 300-mesh Ni grids until further use.
Immunogold cytochemistry was performed on every tenth ultrathin section as previously described56, and the remaining serial sections were only counterstained as described below. Briefly, the ultrathin sections were incubated with a primary antibody against RFP, which recognizes tdTomato (1:100, rabbit, Rockland, 600-401-379s), overnight at room temperature in a humidified chamber. The following day, the sections were washed and incubated with a secondary antibody (1:20, goat anti-rabbit IgG conjugated to 12-nm gold particles, Abcam, ab105298) for 2 h. The sections were counterstained using 2% uranyl acetate and 0.3% lead citrate for 90 s each. Images were acquired at 80 kV using a Tecnai 12 electron microscope (FEI Company) equipped with ITEM FEI version 5.1 software (Olympus Soft Imaging Solutions). Colored hue was added to images with Adobe Illustrator CS6 version 16.0 (Adobe Systems) to highlight cells of interest.
3D modeling
The open-source software Reconstruct57 was used for 3D modeling. Consecutive immunogold electron microscopy images were traced to create the 3D reconstruction. All electron microscopy images used for the 3D modeling were taken with the same magnification to ease the calibration and alignment process. First, images were aligned in a semiautomatic manner using three to four corresponding reference points on two consecutive images, and this was repeated through the entire stack. Care was taken to ensure proper alignment and to avoid potential artifacts in the alignment process. Next, the structures of interest were traced using the wildfire mode, and all traces were inspected and corrected manually where needed. Rendering of the 3D model was done with the following software parameters: Generate; Botsonian surface, Normals; Vertex, Facets; 32. To unwrap selected cells from the endothelial tube, we exported isolated 3D models to the open-source software Blender by the Blender Foundation. Within Blender, cloth physics was applied to the isolated model, and the 3D model fell into the desired position.
Imaging and quantitative analysis
Images were acquired with a Leica TCS SP8X confocal microscope equipped with LAS X 3.5.7.23225 software. Image processing and assembly were performed with ImageJ/Fiji (v2.1.0/1.53c for Mac) and Adobe Illustrator (v25.04.1 for Mac).
Transverse (Figs. 2c and 6 and Supplementary Figs. 5, 9 and 10) or sagittal (Fig. 5, Extended Data Figs. 2 and 5, and Supplementary Figs. 8 and 11) spinal cord sections, spanning the injury site and 400-μm rostral and caudal to the injury site, were collected on 20 alternating slides at 20-μm thickness and used for quantifications. Matched segments of uninjured spinal cords were sectioned and collected similarly. All quantifications were done in at least three alternate sections per animal covering the lesion epicenter and spaced 400 μm apart. Except for Extended Data Fig. 2c, data collection and analysis were not performed blind to the conditions of the experiments.
The percentages of stromal PDGFRβ+ cells that express tdTomato (GLAST-CreERT2;R26R-tdTomato or Col1a1-CreERT2;R26R-tdTomato mice) upon spinal cord crush or contusion and upon intraspinal injection of LPS were assessed by manual counting. The number of tdTomato+PDGFRβ+ was divided by the total number of PDGFRβ+ cells in the lesion core (delimitated by the glia limitans). Only cells dissociated from the blood vessel wall within the lesion core (immunostained for the endothelial markers CD31 or podocalyxin) were considered. For gray and white matter comparisons, the percentages of recombined stromal cells were determined for gray and white matter regions separately. The area covered by the MAC2+ signal was thresholded and measured using ImageJ/Fiji software, and averaged values are presented as the percentage of MAC2+ area per hemisection. The recombination efficacy in Col1a1-CreERT2;R26R-tdTomato mice was determined by the ratio of tdTomato+Col1a1+ cells out of the total number of cells with Col1a1+ mRNA clusters (in situ hybridization with the Mm-Col1a1 RNAscope probe). The ratio was determined in transverse spinal cord tissue sections (n = 6 mice) in the gray matter and white matter. The ratio for the meninges was determined within an optical field spanning one-quarter of the meninges surrounding transverse sections.
The number of GLAST-CreERT2 tdTomato+ or Col1a1-CreERT2 tdTomato+ perivascular (PDGFRβ+) cells in the uninjured mouse spinal cord was counted and calculated over nine transverse sections (20-μm-thick) spanning three different thoracic spinal cord segments. To determine the expression level of Col1a1 mRNA in GLAST-CreERT2;R26R-tdTomato mice, nuclei of recombined (tdTomato+) cells expressing Pdgfrb+ mRNA were outlined (n = 4 mice, three tissue sections each) and the integrated density of the Col1a1 signal was measured using ImageJ/Fiji software. The analyzed cells were divided into tdTomato+ cells occupying large-caliber (>6-μm inner diameter) or small-caliber (<6-μm inner diameter) blood vessels, mainly in the gray matter of the spinal cord. The blood vessels were labeled with Lycopersicon esculentum lectin (B1175, Vector Laboratories).
Statistical analysis
Sample sizes were determined based on previous experience3,10. Data are presented as mean ± s.d., and individual data points are plotted in the graphs. The number of animals (sample size) and statistical tests used are noted in the figure legends. Normality tests were used to assess the Gaussian distribution of the datasets, followed by appropriate statistical tests. Statistical analyses were done with GraphPad Prism (v9.3.1). P values were calculated using two-tailed unpaired Student’s t test and one-way ANOVA followed by Šidáks or Tukey’s multiple-comparisons test. Differences were considered statistically significant when P < 0.05. Source data are provided as source data files.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that all data supporting the findings of this study are included in this published article and its supplementary information files. The publicly available dataset GSE98816 was used for comparisons in Extended Data Fig. 2, and the mouse reference genome used was GCF_000001635.20/27. Single-cell RNA-sequencing data are deposited in the Gene Expression Omnibus (GEO) database (accession nos. GSE229916, GSE266250 and GSE266251). Source data are provided with this paper.
References
Ruschel, J. et al. Axonal regeneration. Systemic administration of epothilone B promotes axon regeneration after spinal cord injury. Science 348, 347–352 (2015).
Brazda, N. & Müller, H. W. Pharmacological modification of the extracellular matrix to promote regeneration of the injured brain and spinal cord. Prog. Brain Res. 175, 269–281 (2009).
Dias, D. O. et al. Reducing pericyte-derived scarring promotes recovery after spinal cord injury. Cell 173, 153–165 (2018).
Dorrier, C. E. et al. CNS fibroblasts form a fibrotic scar in response to immune cell infiltration. Nat. Neurosci. 24, 234–244 (2021).
Dias, D. O. & Göritz, C. Fibrotic scarring following lesions to the central nervous system. Matrix Biol. 68–69, 561–570 (2018).
Göritz, C. et al. A pericyte origin of spinal cord scar tissue. Science 333, 238–242 (2011).
Hellal, F. et al. Microtubule stabilization reduces scarring and causes axon regeneration after spinal cord injury. Science 331, 928–931 (2011).
Yoshioka, N., Hisanaga, S. & Kawano, H. Suppression of fibrotic scar formation promotes axonal regeneration without disturbing blood–brain barrier repair and withdrawal of leukocytes after traumatic brain injury. J. Comp. Neurol. 518, 3867–3881 (2010).
Zhu, Y. et al. Hematogenous macrophage depletion reduces the fibrotic scar and increases axonal growth after spinal cord injury. Neurobiol. Dis. 74, 114–125 (2015).
Dias, D. O. et al. Pericyte-derived fibrotic scarring is conserved across diverse central nervous system lesions. Nat. Commun. 12, 5501 (2021).
Vanlandewijck, M. et al. A molecular atlas of cell types and zonation in the brain vasculature. Nature https://doi.org/10.1038/nature25739 (2018).
Soderblom, C. et al. Perivascular fibroblasts form the fibrotic scar after contusive spinal cord injury. J. Neurosci. 33, 13882–13887 (2013).
Lendahl, U., Nilsson, P. & Betsholtz, C. Emerging links between cerebrovascular and neurodegenerative diseases—a special role for pericytes. EMBO Rep. 20, e48070 (2019).
Armulik, A., Genové, G. & Betsholtz, C. Pericytes: developmental, physiological, and pathological perspectives, problems, and promises. Dev. Cell 21, 193–215 (2011).
Sofroniew, M. V. Inflammation drives fibrotic scars in the CNS. Nat. Neurosci. 24, 157–159 (2021).
Hagemann-Jensen, M., Ziegenhain, C. & Sandberg, R. Scalable single-cell RNA sequencing from full transcripts with Smart-seq3xpress. Nat. Biotechnol. 40, 1452–1457 (2022).
Hagemann-Jensen, M. et al. Single-cell RNA counting at allele and isoform resolution using Smart-seq3. Nat. Biotechnol. 38, 708–714 (2020).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
McKenzie, A. T. et al. Brain cell type specific gene expression and co-expression network architectures. Sci. Rep. 8, 8868 (2018).
Marques, S. et al. Oligodendrocyte heterogeneity in the mouse juvenile and adult central nervous system. Science 352, 1326–1329 (2016).
Zamboni, M., Llorens-Bobadilla, E., Magnusson, J. P. & Frisén, J. A widespread neurogenic potential of neocortical astrocytes is induced by injury. Cell Stem Cell 27, 605–617 (2020).
De Bock, M. et al. A new angle on blood–CNS interfaces: a role for connexins? FEBS Lett. https://doi.org/10.1016/j.febslet.2014.02.060 (2014).
Hirschi, K. K., Burt, J. M., Hirschi, K. D. & Dai, C. Gap junction communication mediates transforming growth factor-β activation and endothelial-induced mural cell differentiation. Circ. Res. 93, 429–437 (2003).
Ivanova, E., Kovacs‐Oller, T. & Sagdullaev, B. T. Domain‐specific distribution of gap junctions defines cellular coupling to establish a vascular relay in the retina. J. Comp. Neurol. 527, 2675–2693 (2019).
Mazaré, N., Gilbert, A., Boulay, A.-C., Rouach, N. & Cohen-Salmon, M. Connexin 30 is expressed in a subtype of mouse brain pericytes. Brain Struct. Funct. https://doi.org/10.1007/s00429-017-1562-4 (2018).
Lendahl, U., Muhl, L. & Betsholtz, C. Identification, discrimination and heterogeneity of fibroblasts. Nat. Commun. 13, 3409 (2022).
Krueger, M. & Bechmann, I. CNS pericytes: concepts, misconceptions, and a way out. Glia 58, 1–10 (2010).
Meletis, K. et al. Spinal cord injury reveals multilineage differentiation of ependymal cells. PLoS Biol. 6, e182 (2008).
Lee, S. B. & Kalluri, R. Mechanistic connection between inflammation and fibrosis. Kidney Int. Suppl. 78, S22–S26 (2010).
Mack, M. Inflammation and fibrosis. Matrix Biol. 68–69, 106–121 (2018).
Felts, P. A. et al. Inflammation and primary demyelination induced by the intraspinal injection of lipopolysaccharide. Brain 128, 1649–1666 (2005).
Batista, C. R. A., Gomes, G. F., Candelario-Jalil, E., Fiebich, B. L. & de Oliveira, A. C. P. Lipopolysaccharide-induced neuroinflammation as a bridge to understand neurodegeneration. Int. J. Mol. Sci. 20, 2293 (2019).
Buechler, M. B. et al. Cross-tissue organization of the fibroblast lineage. Nature 593, 575–579 (2021).
Eilken, H. M. et al. Pericytes regulate VEGF-induced endothelial sprouting through VEGFR1. Nat. Commun. 8, 1574 (2017).
Liu, J. et al. A human cell type similar to murine central nervous system perivascular fibroblasts. Exp. Cell Res. 402, 112576 (2021).
Milich, L. M. et al. Single-cell analysis of the cellular heterogeneity and interactions in the injured mouse spinal cord. J. Exp. Med. 218, e20210040 (2021).
Martirosyan, N. L. et al. Blood supply and vascular reactivity of the spinal cord under normal and pathological conditions. J. Neurosurg. Spine 15, 238–251 (2011).
Kirst, C. et al. Mapping the fine-scale organization and plasticity of the brain vasculature. Cell 180, 780–795 (2020).
Grant, R. I. et al. Organizational hierarchy and structural diversity of microvascular pericytes in adult mouse cortex. J. Cereb. Blood Flow. Metab. 39, 411–425 (2019).
Hartmann, D. A. et al. Pericyte structure and distribution in the cerebral cortex revealed by high-resolution imaging of transgenic mice. Neurophotonics 2, 041402 (2015).
Slezak, M. et al. Transgenic mice for conditional gene manipulation in astroglial cells. Glia 55, 1565–1576 (2007).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Gong, S. et al. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature 425, 917–925 (2003).
Drosten, M. et al. Genetic analysis of Ras signalling pathways in cell proliferation, migration and survival. EMBO J. 29, 1091–1104 (2010).
Lovatt, D. et al. The transcriptome and metabolic gene signature of protoplasmic astrocytes in the adult murine cortex. J. Neurosci. 27, 12255–12266 (2007).
Orre, M. et al. Isolation of glia from Alzheimer’s mice reveals inflammation and dysfunction. Neurobiol. Aging 35, 2746–2760 (2014).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 (2021).
RStudio Team. RStudio: Integrated Development Environment for R (RStudio, 2021).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Xie, Z. et al. Gene set knowledge discovery with Enrichr. Curr. Protoc. 1, e90 (2021).
Kolde, R. Pheatmap: pretty heatmaps. https://CRAN.R-project.org/package=pheatmap/index.html (2019).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Xu, N. et al. Fast free-of-acrylamide clearing tissue (FACT)—an optimized new protocol for rapid, high-resolution imaging of three-dimensional brain tissue. Sci. Rep. 7, 9895 (2017).
Chiu, C.-L. & Clack, N. napari: a Python multi-dimensional image viewer platform for the research community. Microsc. Microanal. 28, 1576–1577 (2022).
Lunde, L. K. et al. Postnatal development of the molecular complex underlying astrocyte polarization. Brain Struct. Funct. 220, 2087–2101 (2015).
Prydz, A. et al. Subcellular expression of aquaporin-4 in substantia nigra of normal and MPTP-treated mice. Neuroscience 359, 258–266 (2017).
Fiala, J. C. Reconstruct: a free editor for serial section microscopy. J. Microsc. 218, 52–61 (2005).
Acknowledgements
We thank J. Frisén, U. Lendahl and members of the Göritz laboratory for valuable comments on the manuscript. C.G. is a Hållsten Academy, Knut and Alice Wallenberg Academy and STIAS fellow. D.O.D. was supported by the Strategic Network for Neuroscience (STRATNEURO) at Karolinska Institutet. S.S. was supported by the European Union’s Horizon 2020 research and innovation program under the Marie Skłodowska-Curie grant agreement no. 707449 (PERICYTEStroma). A.J. was supported by a postdoctoral fellowship from Svenska Sällskapet för Medicinsk Forskning. Research in the Göritz laboratory was supported by grants to C.G. from the Knut and Alice Wallenberg Foundation (2018.0159), Swedish Research Council (2018-02798 and 2022-01244), Swedish Brain Foundation (FO2020-0093 and FO2022-0206), Anna-Stina och John Mattsons Minnesstiftelse för sonen Johan, Swedish Cancer Foundation (21-1768Pj01H), Ming Wai Lau Centre for Reparative Medicine, the Strategic Network for Stem Cells and Regenerative Medicine (STRATREGEN) at Karolinska Institutet, and Wings for Life Foundation (WFL-SE-10/20). We acknowledge the support of the Government of Canada’s New Frontiers in Research Fund (NFRF) (NFRFT-2020-00238) to C.G. We also acknowledge the support by the research fund to the Center for Neuromusculoskeletal Restorative Medicine from the Health@InnoHK program launched by the Innovation and Technology Commission, the Government of the Hong Kong Special Administrative Region of the People’s Republic of China to C.G. We thank the Advanced Light Microscopy Unit at KTH-Royal Institute of Technology and the National Microscopy Infrastructure (NMI) (VR-RFI 2019-00217) for helping with microscopy, as well as R. M. L. Cabezas for her support with tissue clearing and imaging. We are grateful for the Bioinformatics Long-term Support (WABI) from the National Bioinformatics Infrastructure Sweden at SciLifeLab (financed by the Knut and Alice Wallenberg Foundation).
Funding
Open access funding provided by Karolinska Institute.
Author information
Authors and Affiliations
Contributions
D.H., W.F.H., A.J., S.B., J.K., S.S., D.O.D. and E.L.-B. performed experiments and/or analyses. D.H., D.O.D. and C.G. designed experiments. Y.H. and G.P. generated Co1a1-CreERT2 mice. M.A.-M. supervised the electron microscopy analysis. C.G. conceptualized the study and supervised experiments. D.H., D.O.D. and C.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Neuroscience thanks Vittorio Gallo, Julie Siegenthaler and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Perivascular fibroblasts and GLAST−expressing pericytes distribute differently along the vascular tree and between grey matter and white matter in the uninjured spinal cord.
Distribution of GLAST-CreERT2 tdTomato+ (a) and Col1a1-CreERT2 tdTomato+ (b) cells along arterioles (white arrowheads/dots), venules (yellow arrowheads/dots) and capillaries (green arrowheads/dots) revealed by volumetric imaging of 500 μm thick uninjured spinal cord thoracic segments co-stained with antibodies against SM22α (arterioles/arteries) and vWF (large ascending venules/veins) (upper panel) and 200 μm thick enlarged image of the dorsal grey matter (lower panel). The blood vessel lumen was labeled by transcardial perfusion with Alexa Fluor 647-conjugated bovine serum albumin in a gelatin solution. (c) Transverse sections of the spinal cord (same sections as shown in (a) and (b)) showing the location of GLAST-CreERT2 tdTomato+ cells and Col1a1-CreERT2 tdTomato+ cells within a 500 μm thick uninjured spinal cord thoracic segment (each dot represents a cell, color-coded for the vessel type the cell resides on). Solid lines outline the spinal cord parenchyma, dashed lines outline the grey matter. (d) Distribution of tdTomato+ perivascular cells along arterioles (including the arteriole-capillary transitional zone), venules and capillaries, represented as a percentage of total tdTomato+ perivascular cells. (e) Quantification of the number of tdTomato+ perivascular cells per 500 μm thick uninjured spinal cord thoracic segments. (f) Quantification of the number of tdTomato+ perivascular cells within the grey and white matter per 500 μm thick uninjured spinal cord thoracic segments. (g) Distribution of tdTomato+ perivascular cells in the spinal cord grey and white matter, represented as a percentage of total tdTomato+ perivascular cells. There are significantly more Col1a1-CreERT2 tdTomato+ cells in the white matter than in the grey matter. Scale bars: 200 μm (a and b upper panel, c) and 50 μm (a and b lower panel). Data shown as mean + SD. n = 3 (GLAST-CreERT2), n = 3 (Col1a1-CreERT2) animals in (d-g). ns=0.688, **p = 0.006 (arterioles-capillaries), **p = 0.0017 (venules-capillaries), ****p < 0.0001 (d), **p = 0.001, ns=0.1705, *p = 0.0214 (e), **p = 0.0021, ns=0.1008 (f), ns=0.3832, **p = 0.004 (g) by two−sided, unpaired Student’s t test. All images show transverse sections. Images are representative of three independent experiments. Source data are provided as a Source Data file.
Extended Data Fig. 2 Genetic inhibition of perivascular fibroblast proliferation reduces the total number of PDGFRβ+ stromal fibroblasts upon SCI.
(a) Genetic strategy to inhibit injury-induced proliferation of Col1a1-CreERT2 perivascular fibroblasts through cell-specific deletion of floxed K-Ras in mice with H-Ras and N-Ras null alleles (Col1a1-CreERT2;Rasless mice) upon tamoxifen-induced genetic recombination. (b) Sagittal view of the lesion showing the distribution of PDGFRβ+ stromal fibroblasts and blood vessels (podocalyxin+, Pdx) in CreWT and Cre+ Col1a1-CreERT2;Rasless mice, 7 days after complete spinal cord crush. Both Cre+ and CreWT animals were administered tamoxifen prior to injury, but CreWT animals did not undergo genetic recombination and served as control. (c) The density of scar-forming PDGFRβ+ fibroblasts is reduced in Cre+ (248.4 ± 50.6 cells/mm2, mean ± SD) compared to CreWT (350.4 ± 67.9 cells/mm2, mean ± SD) Col1a1-Rasless mice. TAM, tamoxifen. Scale bars: 100 µm (b). Data shown as mean + SD in (c) n = 3 (CreWT), n = 9 (Cre+ ) animals. *p = 0.0182 by two-sided, unpaired Student’s t test in (c). Cell nuclei are labeled with DAPI. Images are representative of two independent experiments. Source data is provided as a Source Data file.
Extended Data Fig. 3 SCI induces GLAST+ pericyte and fibroblast activation and the generation of myofibroblasts.
(a) Heatmap showing single-cell RNA-sequencing data of tdTomato+EGFP+ cells from the uninjured and injured spinal cord (3 and 5 days after complete spinal crush) of GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice subdivided by clusters based on Fig. 7a. Expression level of selected genes relevant for fibrosis are shown; scale: log counts. (b) Contribution of cells from different conditions (uninjured, 3 and 5 days after complete spinal crush) to each cluster (labelled in a), clustering based on Fig. 7a. (c) Expression of Col8a1, Fn1 and Kcnj8 genes (note scale on y-axis) along pseudotime within the fibroblast (left) and pericyte (right) branches. (d) UMAP plot based on Fig. 7a showing tdTomato+EGFP+ cells color-coded for cell cycle phases (G1, G2/M, S), determined with the CellCycleScoring function in Seurat. (e) UMAP representation of the percentage of ribosomal genes detected in comparison to total gene count, assessed with the PercentageFeatureSet function in Seurat. (f) Violin plots of genes relevant for GO terms based on Fig. 7i and j.
Extended Data Fig. 4 GLAST+ pericytes and fibroblasts and a subpopulation of GLAST−pericytes get activated upon SCI.
(a) UMAP representation of tdTomato+EGFP+ cells isolated from the uninjured and injured spinal cord of GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice at 5 days after complete spinal crush, color-coded by injury condition. Dashed lines outline pericytes, fibroblasts, as well as activated fibroblasts and myofibroblasts based on marker expression. (b) Volcano plot of differentially expressed genes (DEG) between uninjured and 5 days post injury (dpi) tdTomato+EGFP+ cells. (c) UMAP projection of tdTomato−EGFP+ cells isolated from the uninjured and injured spinal cord of GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice at 5 days after complete spinal crush, color-coded by injury condition. Dashed lines outline pericytes, vascular smooth muscle cells (vSMCs) and activated pericytes based on marker expression. (d) Volcano plot of DEG between uninjured and 5 dpi tdTomato−EGFP+ cells. (e) UMAP color-coded for Seurat cluster analysis, separating tdTomato−EGFP+ pericytes (cluster 0) and vSMCs (cluster 1). (f) Heatmap of tdTomato-EGFP+ cells shown in (c-e) separated by cluster and injury condition, showing gene expression of selected pericyte, fibroblast, extracelullar matrix (ECM), angiogenesis, and vSMC marker genes. Activated tdTomato−EGFP+ pericytes at 5 dpi do not cluster separately (c), but expression of genes related to ECM and angiogenesis indicate activation of a subset of cells (f).
Extended Data Fig. 5 Activated pericytes, fibroblasts and myofibroblasts populate the lesion site after SCI.
(a) Overview images of the lesion site of GLAST-CreERT2;R26R-tdTomato;Pdgfrb-eGFP mice at 5 days after complete spinal cord crush. The majority of tdTomato+EGFP+ scar-forming cells expresses high levels of smooth muscle protein 22-alpha (SM22α, gene name Tagln), a marker of myofibroblasts and vascular smooth muscle cells (vSMCs). (b) Close-up of the grey matter lesion border. TdTomato−EGFP+SM22α+ cells associated with podocalyxin+ (Pdx) blood vessels (vSMCs or activated pericytes; yellow arrowhead) are mainly located in the lesion border intermingled with tdTomato+EGFP+SM22α+ myofibroblasts (white arrowheads). (c) Close-up of the grey matter lesion core showing that myofibroblasts are mostly derived from tdTomato+EGFP+ perivascular cells (white arrowhead). (d) Close up of the lesion in the white matter. The cellular composition and distribution, and the expression level of SM22α in the white matter injury compartment is comparable to the grey matter. Magnified boxed region on the right side shows blood vessel-associated tdTomato−EGFP+SM22α+ vSMCs or activated pericytes surrounded by tdTomato+EGFP+SM22α+ myofibroblasts (white arrowhead). Scale bars: 500 μm (a) and 50 μm (b-d). Cell nuclei are labeled with DAPI. Images are representative of two independent experiments. All images show sagittal sections.
Supplementary information
Supplementary Information
Supplementary Figs. 1–11.
Supplementary Data
Source data for Supplementary Figs. 4–6 and 9.
Source data
Source Data Fig. 2
Individual data points/counted values.
Source Data Fig. 5
Individual data points/counted values.
Source Data Fig. 6
Individual data points/counted values.
Source Data Fig. 7
Full list of GO terms.
Source Data Extended Data Fig. 1
Individual data points/counted values.
Source Data Extended Data Fig. 2
Individual data points/counted values.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Holl, D., Hau, W.F., Julien, A. et al. Distinct origin and region-dependent contribution of stromal fibroblasts to fibrosis following traumatic injury in mice. Nat Neurosci 27, 1285–1298 (2024). https://doi.org/10.1038/s41593-024-01678-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-024-01678-4
- Springer Nature America, Inc.
This article is cited by
-
Each scar whispers a story
Nature Neuroscience (2024)