Abstract
The Asian giant softshell turtle Pelochelys cantorii is one of the largest aquatic turtles in China and has been designated a First Grade Protected Animal in China. To advance conservation research, a combination of Illumina short-read, PacBio long-read, and Hi-C scaffolding technologies was used to develop a high-quality chromosome-level genome assembly for P. cantorii. A total of 262.77 Gb of clean data were produced (121.6 × depth) and then the genome was assembled into 2.16 Gb with a contig N50 of 41.44 Mb and scaffold N50 length of 120.17 Mb, respectively. Moreover, about 99.98% assembly genome sequences were clustered and ordered onto 33 pseudochromosomes. Genome annotation revealed that 21,833 protein-coding genes were predicted, and 96.40% of them were annotated. This new chromosome-level assembly will be an enabling resource for genetic and genomic studies to support fundamental insight into P. cantorii biology.
Similar content being viewed by others
Background & Summary
The Asian giant softshell turtle Pelochelys cantorii is one of the largest aquatic turtles and is widely distributed in Southeast Asia. With rapid economic development, the numbers of P. cantorii have sharply declined due to overhunting and habitat destruction in China. Although listed as a national first-class protected animal in China in 1989, wild P. cantorii individuals in China have been reported at a declining rate in the past 30 years1,2, and only 13 P. cantorii individuals from the wild are in captivity across 6 different locations. P. cantorii is extremely endangered in China1,3. The fate of another large soft-shelled turtle in China, the Yangtze giant softshell turtle, Rafetus swinhoei, is even more worrisome. No individuals were found in the wild, and two individuals (1 female and 1 male) reared in captivity failed in assisted reproduction after 2008. In 2019, the lone female R. swinhoei died during artificial insemination. If there are no R. swinhoei individuals in the wild, the fate of R. swinhoei is sealed. If the protection of P. cantorii is not strengthened, it is inevitable that the P. cantorii will not become the next R. swinhoei.
In 2014, 10 cantorii hatchlings were successfully bred in captivity from 2 turtle individuals (1 female and 1 male)4. From 2015 to 2021, four captive P. cantorii individuals (2 females and 2 males) successfully bred and raised 950 juveniles that are currently 1–7 years old. Knowledge of the artificial breeding of P. cantorii offers hope for the conservation of this species5. With the breakthrough of the artificial breeding of P. cantorii, related protection work has received attention from the government of China. P. cantorii is listed as one of the nine aquatic wild animals under the key management of the Ministry of Agriculture and Rural Affairs (MARA) of the People’s Republic of China. In 2019, the MARA of China issued the “Pelochelys cantorii Rescue Action Plan (2019–2035)”. In September 2020, the MARA of China organized and carried out the first wild adaptation protection test for P. cantorii. A total of 20 juvenile turtles with implanted PIT chips were released in a reservoir in Gaoming, Foshan. These turtles were between 4–5 years old and weigh 1.04–1.66 kg.
With the development of sequencing technology, conservation genomics was born, which overcomes the limitations of conservation genetics, such as the lack of markers to a large extent, and helps to solve some unresolved problems in the conservation biology of organisms, especially endangered species6. For example, de Novo sequencing and population resequencing analysis based on Yangtze finless porpoise Neophocaena Asiaeorientalis asiaeorientalis more accurately determined the phylogenetic relationships and population genetic structure, and confirmed the independent species status of the Yangtze finless porpoise7. Through the analysis of genomic data of hot spring snakes, the genes involved in DNA damage repair (FEN1) and hypoxia response (EPAS1) were identified, providing research ideas for the mechanism of environmental adaptation. With the continuous development and optimization of high-throughput sequencing technology, research on conservation genomes will continue to improve.
Third-generation sequencing technologies, such as Pacific Biosciences (PacBio) Single Molecule Real-Time (SMRT) and Oxford Nanopore sequencing, characterized by single-molecule sequencing, can overcome the shortcomings of second-generation sequencing methods8. Hi-C, based on the combination of chromatin conformation capture and high-throughput sequencing, reveals the interaction information between chromosome fragments through the analysis of sequencing data9,10,11,12.
Tortoises are an ancient reptile with more than 300 species and have significant evolutionary and ecological value. No more than 10 turtle genome projects have been completed since the first investigation on turtle genomes were published in 2013 (Chinese soft-shell turtle and green turtle). Therefore, it is urgent to expand the diversity of the turtle genome databases.
Here, we sequenced and de novo assembled the genome of a young P. cantorii via a combination of long-read PacBio Sequel II platform and Hi-C sequencing technology. The high-quality reference genome constructed in this study will not only be of benefit to serve as the genetic basis for in-depth investigations of turtle evolution and biology but also offers a valuable genetic resource for turtle conservation.
Methods
Ethics statement
All the experimental procedures regarding the turtle involved in this experiment were approved by the Experimental Animal Care and Ethics Committee of the Pearl River Fisheries Research Institute, Chinese Academy of Fishery Sciences.
Sample preparation and genome sequencing
A healthy 2-year-old adult was obtained from the breeding center of the Pearl River Fisheries Research Institute, Guangzhou, Guangdong, China (Fig. 1). The turtle was anaesthetized using tricaine methanesulfonate (MS-222, Sigma) before sampling. The fresh muscle tissue was cut to small sizes (about 20 µg) and conducted for DNA extraction using a DNeasy Blood & Tissue Kit (Qiagen, Valencia, CA, USA), according to the manufacturer’s instructions. Only high-quality DNA can be used for the following library preparation, sequencing, and Hi-C library construction, so the quantity and quality of the genomic DNA were first evaluated by using Qubit 3.0 (Life Invitrogen, USA) and 1% agarose gel electrophoresis, respectively. The library was then constructed and sequenced on Illumina HiSeq4000 (Illumina, San Diego, CA, USA) and the PacBio Sequel II platform according to the manufacturer’s instructions.
Tissues from the heart, liver, kidney, muscle, eye, blood, and lung of the same turtle were used for RNA extraction by TRIzol Universal Reagent (TIANGEN Biotech, Beijing, China). Two micrograms of total RNA from each tissue were pooled for RNA sequencing on a PacBio Sequel II.
Genome size estimation and initial genome assembly
Genome size was estimated by k-mer analysis13 using filtered reads from the three 350 bp libraries constructed using genomic DNA and a k-mer size of 21. Genome size, repetition ratio, and heterozygosity were assessed by k-mer depth frequency distribution analysis (Table 1). The average k-mer depth was 56, and the total number of k-mers obtained from sequencing data was 131,112,165,086, further estimating the genome size of P. cantorii (Fig. 2 and Table 1). Sequences with a k-mer depth greater than 113 were repeated sequences, and those with a k-mer depth of approximately 28 were heterozygous sequences. After removing k-mers of abnormal depth, the genome size of P. cantorii was evaluated to be 2.20 Gb (based on the following formula: G = k-mer number/mean k-mer depth), with a low heterozygosity of approximately 0.14% and a repeat sequence content of 26.29% (Table 1). The PacBio sequencing platform generated a total of 262.77 Gb of clean data (approximately 121.60 × ), with a read N50 of 22.85 kb and an average read length of 14.62 kb (Table 2). The data were first assembled with WTDBG2, which generated a genome assembly with a total length of 2.16 Gb, 548 contigs, and 41.44 Mb contig N50 (Table 3).
A 21-mer distribution of the Illumina short reads to estimate genome size, ratio of repeat sequences and heterogeneity. The x-axis represents the sequencing depth of each unique 21-mer, and the y-axis represents the frequency of unique 21-mers. The k-mer depth value of the main peak is 56. After removing data with abnormal depth, a total of 131,112,165,086, 21-mers were used to further estimate the genome size; thus, we first estimated that the genome size of the Asian giant softshell turtle was 2.20 Gb.
Chromosome assembly using Hi-C data
A high-throughput chromatin conformation capture (Hi-C) library was then created for sequencing from 2-year-old individual muscle tissue. The sample was fixed with paraformaldehyde and enzymatically digested with DnpII, generating sticky ends. The sticky ends can be repaired by ‘A’ or ‘C’ deoxynucleotides with biotin under the action of DNA polymerase. After labeling the 5′ overhang with a biotinylated residue, the DNA fragments were ligated to each other to form chimeric circles using DNA ligase. Then, the ligated DNAs were uncrosslinked, purified, and sheared to 300 bp-700 bp fragments, followed by capture with streptavidin beads and preparation for sequencing. Finally, the Hi-C libraries were quantified and sequenced using the Illumina HISeq X 10 platform in PE150 mode with paired-end methods.
To obtain high-quality Hi-C data, the joint information, low-quality bases, and undetected bases in the original sequencing data were first removed to generate clean reads. Second, BWA v0.7.10-r789 was applied to align the clean reads to the draft genome assembly with the default parameters. HiC-Pro v2.8.1 was performed to filter invalid read pairs, including dangling-end and self-cycle, re-ligation, and dumped products. The contigs of the primary genome assembly were corrected by splitting them into segments of 50 kb on average. After checking any two segments that showed inconsistent connection with information from the raw scaffold, LACHESIS (ligating adjacent chromatin enables scaffolding in situ) was used to produce chromosome-level scaffolds. The final assembled genome was 2.16 Gb and anchored 532 contigs. The scaffold N50 was 41.44 Mb with a fixed rate of 95.86% (Table 3). Among the sequences located on chromosomes, the length of the sequence that can determine the order and direction is up to 99.97%. The Hi-C scaffolding of the genome resulted in 33 chromosomes (Chr) (Figure S1), accounting for 99.98% of the total assembly (Table 4). In order to further determine the chromosome number of the Asian giant softshell turtle, about 1 mL blood was extracted from the neck blood vessels of four 1 to 3 years-old young individuals from the turtle domestication base of the Pearl River Fisheries Research Institute. After 72 h of culture, cells were collected for the preparation of chromosomes analysis as previously described14. A total of 43 metaphase mitotic cells with a higher number of chromosomes were obtained (Figure S2, Figure S3). Finally, we selected one of the chromosome division phases to make the standard karyotype map (Figure S4), and the karyotype formula was determined by the first eight division phase maps in Supplementary figure 1.
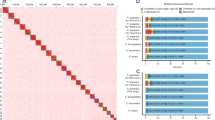
Subsequently, Juicer and JucieBox v.1.8.8 were implemented to construct an interactive map and correct assembly errors visually, respectively. The heat map of the Hi-C assembly interaction bins displayed stronger signals around the diagonal than that of other positions. The phenomenon revealed that intensity of the interaction between adjacent sequences (diagonal position) was high, while the intensity of the interaction signal between non-adjacent sequences (non-diagonal position) was weak, which was consistent with the principle of Hi-C assisted genome assembly, proving a high quality and completeness of a genome assembly (Fig. 3).
Assessment of the genome assemblies
Two methods were used to check the completeness and quality of the assembly. First, BWA v0.7.10-r78915 was used to compare the short sequences obtained by second-generation high-throughput Illumina sequencing with the reference genome, and the integrity of the assembled genome was evaluated by the alignment rate. Approximately 98.95% of clean reads were mapped to contigs, and 96.96% of clean reads were mapped to proper pairs (Table 5). Second, BUSCO v3.016 was used to search the 2,586 universal single-copy orthologous genes in vertebrata_OrthoDB v9 to evaluate the completeness, degree of fragmentation and missing genes of the genome assembly. Among the 2,586 prospective conserved core genes in the eukaryotic database, 2,478 (95.82%) and 54 (2.09%) were identified as complete and fragmented BUSCOs, respectively, indicating that the assembled genome had high integrity and validity, and could be used for further analysis (Fig. 4, Table 6).
Gene prediction and functional annotation
With the help of LTR_FINDER17 and RepeatScout18, a repeat sequence database of the genome was constructed, based on the principle of structure prediction and ab initio prediction. PASTEClassifier19 was used to categorize different types of repetitive sequences. The ultimate repeat library was subsequently determined combined with Repbase20 database. At last, RepeatMasker21 software was conducted to predict the repeated sequences. We identified 993,573,618 bp sequences as repeats in the Repbase database, which covered 45.98% of the genome assembly (Table 7). The most common repetitive elements, RNA transposons (class I), account for 39.33% of the genome content, higher than DNA transposons (Class II) (Table 7). Long interspersed nuclear elements (LINEs) were the most abundant repeating element, followed by large retrotransposon derivatives (LARDs) and terminal inverted repeats (TIRs) (Table 7). In addition, several sequences classified as unknown repeats accounted for 2.48% of the genome assembly (Table 7).
A combination of three different strategies including ab initio prediction, homologous species prediction and transcriptome-based prediction were conducted to establish gene models. For gene predication based on ab initio, Genscan22, Augustus v2.423, GlimmerHMM v3.0.424, GeneID v1.425, and SNAP version 2006-07-2826 were employed to predict protein-coding genes. For homology-based prediction, GeMoMa v1.3.127,28 was used to align the assembled genome of P. cantorii with homologous species to predict the potential gene structures. Regarding RNA-Seq based prediction, Hisat v2.0.429 and Stringtie v1.2.330 were used for transcriptome assembly based on reference transcripts, and TransDecoder v2.0 and GeneMark-ST v5.131 were used for gene prediction. PASA v2.0.232 was used to predict Unigene sequences based on unreferenced assembly of transcriptome data. Finally, EVM v1.1.133 software was used to integrate the prediction results of the above three methods, and PASA v2.0.2 was used to modify the results (Table 8).
The predicted gene sequences were compared using BLAST V2.2.3134 (-evalue1e-5) with functional databases such as NR35, KOG36, GO37, KEGG38 and TrEMBL39. Gene functional annotation analysis, including KEGG pathway annotation analysis, KOG functional annotation analysis and GO functional annotation analysis, was performed. Finally, 21,833 protein-coding genes were successfully annotated. Of these predicted genes, 21,046 (~96.40%) were functionally annotated in at least one database, including GO, KEGG, KOG, TrEMBL and NR database (Table 9). Moreover, Blastn was used in the Rfam40 database for whole-genome comparison to identify microRNAs and rRNAs. tRNAscan-SE41 was used to identify tRNA. A total of 1595 tRNAs, 220 microRNAs and 86rRNAs were identified.
Data Records
The sequencing data have been deposited in the NCBI Sequence Read Archive database under the BioSample accession numbers. The accession number of coding sequences, Illumina, genome raw data, assembled genome data, gene annotation, exon annotation, Hi-C and full-length transcriptome sequences were SRR2271518942 (PRJNA910848), SRR22681424-SRR2268142643 (PRJNA911015), SRR2229639444 (PRJNA901634), SRR2417942545 (PRJNA955900), SRR2271519746 (PRJNA910853) and SRR2267465747 (PRJNA910849), SRR2304744248 (PRJNA922717) and SRR2304739349 (PRJNA922721) respectively. The genome assembly is available for public access at the NCBI GenBank under the accession number GCA_032595735.150. Genome annotations (.gff3), along with predicted coding sequences (.cds) and protein sequences (.pep), can be accessed through the Figshare51. Moreover, all the data were also stored in zenodo database52.
Technical Validation
Genomic integrity, fragmentation, and possible loss rates were measured using BUSCO V3. Among 2,586 prospective conserved core genes in the eukaryotic database, 2,478 (95.82%) and 54 (2.09%) were identified as complete BUSCOs and fragment BUSCOs, respectively, indicating that the assembled genome had high integrity and validity and could be used for further analysis (Fig. 4, Table 6).
Code availability
All commands and pipelines used in data processing were executed according to the manual and protocols of the corresponding bioinformatics software. The settings and parameters of these softwares are described below. (1) BWA v0.7.10-r789: aln, default parameters; (2) LACHESIS: CLUSTER_MIN_RE_SITES = 547; CLUSTER_MAX_LINK_DENSITY = 2; CLUSTER_NONINFORMATIVE_RATIO = 2; ORDER_MIN_N_RES_IN_TRUN = 1094; ORDER_MIN_N_RES_IN_SHREDS = 1076; (3) BUSCO v3.0: --evalue 1e-03 (E-value cutoff for BLAST searches), -sp human; (4) LTR_FINDER: default parameters; (5) RepeatScout: default parameters; (6) PASTEClassifier: default parameters; (7) RepeatMasker: -nolow -no_is -norna -engine wublast; (8) Genscan: default parameters; (9) Augustus v2.4: default parameters; (10) GlimmerHMM v3.0.4: default parameters; (11) GeneID v1.4: default parameters; (12) SNAP: version 2006-07-28, default parameters; (13) GeMoMa v1.3.1: default parameters; (14) Hisat v2.0.4: --max-intronlen 20000, --min-intronlen 20; (15) Stringtie v1.2.3: default parameters; (16) TransDecoder v2.0: default parameters; (17) GeneMark-ST v5.1: default parameters; (18) PASA v2.0.2: -align_tools gmap, -maxIntronLen 20000; (19) EVM v1.1.1: default parameters; (20) BLAST V2.2.31: -evalue 1e-5; (21) tRNAscan-SE: default parameters.
References
Gong, S. P. et al. Disappearance of endangered turtles within China’s nature reserves. Current Biology 27, R170–R171 (2017).
Wu, J., Wu, Y., Rao, D., Zhou, T. & Gong, S. China’s wild turtles at risk of extinction. Science (New York, N.Y.) 368, 838 (2020).
Hong, X. et al. Conservation status of the Asian giant softshell turtle (Pelochelys cantorii) in China. Chelonian Conservation and Biology 18, 68–74 (2019).
Zhu, X., Hong, X., Zhao, J., Liang, J. & Feng, Z. Reproduction of captive Asian giant softshell turtles, Pelochelys cantorii. Chelonian Conservation and Biology 14, 143–147 (2015).
Hong, X. et al. Status and analysis of artificial breeding and management of aquatic turtles in China. Biology 11, 1368 (2022).
Grueber, C. E. & Sunnucks, P. Using genomics to fight extinction. Science 376, 574–575 (2022).
Zhou, X. et al. Population genomics of finless porpoises reveal an incipient cetacean species adapted to freshwater. Nature Communication 9, 1276 (2018).
Bian, L. & Li, F. Chromosome-level genome assembly of the greenfin horse-faced filefish (Thamnaconus septentrionalis) using Oxford Nanopore PromethION sequencing and Hi-C technology. Molecular Ecology Resources 20, 1069–1079 (2020).
Sexton, T. et al. Three-dimensional folding and functional organization principles of the Drosophila genome. Cell 148, 458–472 (2012).
Burton, J. N. et al. Chromosome-scale scaffolding of de novo genome assemblies based on chromatin interactions. Nature Biotechnology 31, 1119–1125 (2013).
Selvaraj, S., Dixon, J. R., Bansal, V. & Ren, B. Whole-genome haplotype reconstruction using proximity-ligation and shotgun sequencing. Nature Biotechnology 31, 1111–1118 (2013).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Wang, H. et al. Estimation of genome size using k-mer frequencies from corrected long reads. arXiv: Genomics (2020).
Liu, X. et al. Characterization of the in vitro cultured ovarian cells in the Asian yellow pond turtle (Mauremys mutica). Biology 11, 1404 (2022).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Research 35, W265–268 (2007).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics (Oxford, England) 21(Suppl 1), i351–358 (2005).
Hoede, C. et al. PASTEC: an automatic transposable element classification tool. PLoS One 9, e91929 (2014).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenetic and Genome Research 110, 462–467 (2005).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Current Protocols in Bioinformatics Chapter 4, Unit 4.10 (2009).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. Journal of Molecular Biology 268, 78–94 (1997).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics (Oxford, England) 19(Suppl 2), ii215–225 (2003).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics (Oxford, England) 20, 2878–2879 (2004).
Alioto, T., Blanco, E., Parra, G. & Guigó, R. Using geneid to Identify Genes. Current Protocols in Bioinformatics 64, e56 (2018).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Keilwagen, J. et al. Using intron position conservation for homology-based gene prediction. Nucleic Acids Research 44, e89 (2016).
Keilwagen, J., Hartung, F., Paulini, M., Twardziok, S. O. & Grau, J. Combining RNA-seq data and homology-based gene prediction for plants, animals and fungi. BMC Bioinformatics 19, 189 (2018).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nature Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nature Biotechnology 33, 290–295 (2015).
Tang, S., Lomsadze, A. & Borodovsky, M. Identification of protein coding regions in RNA transcripts. Nucleic Acids Research 43, e78 (2015).
Campbell, M. A., Haas, B. J., Hamilton, J. P., Mount, S. M. & Buell, C. R. Comprehensive analysis of alternative splicing in rice and comparative analyses with Arabidopsis. BMC Genomics 7, 327 (2006).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biology 9, R7 (2008).
O’Driscoll, A. et al. HBLAST: Parallelised sequence similarity-A Hadoop MapReducable basic local alignment search tool. Journal of Biomedical Informatics 54, 58–64 (2015).
Marchler-Bauer, A. et al. CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Research 39, D225–229 (2011).
Koonin, E. V. et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biology 5, R7 (2004).
Dimmer, E. C. et al. The UniProt-GO Annotation database in 2011. Nucleic Acids Research 40, D565–570 (2012).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Research 28, 27–30 (2000).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Research 31, 365–370 (2003).
Griffiths-Jones, S. et al. Rfam: annotating non-coding RNAs in complete genomes. Nucleic Acids Research 33, D121–124 (2005).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Research 25, 955–964 (1997).
EBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP412853 (2022).
EBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP412460 (2022).
EBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP407988 (2022).
EBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP432763 (2023).
EBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP412855 (2022).
EBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP412374 (2022).
EBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP416946 (2023).
EBI Sequence Read Archive https://identifiers.org/insdc.sra:SRP416945 (2023).
Liu, X. Genbank https://identifiers.org/insdc.gca:GCA_032595735.1 (2023).
Liu, H. et al. A chromosome-level genome assembly of the Asian giant softshell turtle Pelochelys cantorii. Figshare https://doi.org/10.6084/m9.figshare.24174504.v1 (2023).
A chromosome-level genome assembly of the Asian giant softshell turtle Pelochelys cantorii. zenodo https://doi.org/10.5281/zenodo.8191629 (2023).
Acknowledgements
The work was funded by the National Key R&D Program of China (2018YFD0900201), Guangdong Natural Science Foundation (2022A1515011360), National Natural Science Foundation of China (32102789), Guangdong Basic and Applied Basic Research Foundation (2022A1515012274; 2020A1515110659), Science and Technology Program of Guangzhou (202206010070; 202102021070), Central Public-interest Scientific Institution Basal Research Fund, CAFS (2020TD35, 2020ZJTD01, 2021SJ-XT2), China-ASEAN Maritime Cooperation Fund (CAMC-2018F), Conservation and Utilization of Agricultural Resources (Z130135) and Guangdong Agricultural Research System (2019KJ150).
Author information
Authors and Affiliations
Contributions
X.Y.H., X.L.L. and X.P.Z. conceived the project and designed the experiments. H.Y.L., Y.K.W., K.K.W., L.Q.J. and C.Q.W. conducted the experiments. L.Y.Y., W.L., C.C., and M.Z.L. analyzed the results. X.L.L., X.Y.H. and X.P.Z. wrote the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hong, X., Liu, H., Wang, Y. et al. A chromosome-level genome assembly of the Asian giant softshell turtle Pelochelys cantorii. Sci Data 10, 754 (2023). https://doi.org/10.1038/s41597-023-02667-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02667-1
- Springer Nature Limited
This article is cited by
-
A chromosome-level genome assembly of the pig-nosed turtle (Carettochelys insculpta)
Scientific Data (2024)