Abstract
Herein, we present the first high-quality long-read-based chromosome-level genome assemblies and gene annotations of the genomes of three endangered Tokudaia species: Tokudaia osimensis, Tokudaia tokunoshimensis, and Tokudaia muenninki. These species, which are endemic to different islands of the Ryukyu Islands, Japan, exhibited unique karyotypes and sex chromosomal characteristics. The genome assemblies generated using PacBio, Illumina, and Hi-C sequence data consisted of 13 (corresponded to 12 autosomes and one X chromosome), 23 (corresponded to 22 autosomes and one X chromosome), and 23 (corresponded to 21 autosomes and the neo- and ancestral X regions) chromosome-level scaffolds that contained 2,445, 2,477, and 2,661 Mbp of sequence data, respectively. Annotations of protein-coding genes were performed using RNA-Seq-based, homology-based, and Ab initio methods. BUSCO completeness values for every species exceeded 96% for genomes and 98% for genes. These data can be an important resource for contributing to our understanding of species genomes resulting from allopatric speciation and provide insights into mammalian sex-determination mechanisms and sex chromosome evolution.
Similar content being viewed by others
Background & Summary
Tokudaia is a genus of murine rodents consisting of only three species of spiny rat, Tokudaia osimensis, Tokudaia tokunoshimensis, and Tokudaia muenninki, all of which have been listed as endangered because of their declining populations in recent years1. The three Tokudaia species are endemic to Amami-Oshima, Tokunoshima, and Okinawa Islands of the Ryukyu Islands, Japan. These islands were probably formed by repetitive sea regressions and transgressions among the Central Ryukyu Islands because of sea-level changes caused by glacial–interglacial cycles since the early Pleistocene2. Phylogenetic relationships between the three Tokudaia species correspond to the geographical distances between the three islands3. The three spiny rat species exhibit different karyotypes and unique sex chromosome characteristics. For example, the diploid number of chromosomes (2n) for both sexes in T. osimensis is 25, in T. tokunoshimensis is 45, and in T. muenninki is 444,5,6. Moreover, both T. osimensis and T. tokunoshimensis have lost their Y chromosomes and Sry, a sex-determining gene in mammals7. Therefore, both males and females of these two species exhibit the XO/XO karyotype, with only one X chromosome;4,5 these species probably exhibit an Sry-independent sex-determination mechanism, different from that of common XX/XY-type mammals (Fig. 1). Various studies have been conducted to understand the sex-determination mechanism of these spiny rats, including experimental8,9,10 and genome sequence-based approaches. The results revealed that genes on the Y chromosome, other than Sry, have been conserved in the genome by translocation to the X chromosome11,12. Additionally, results showed a male-specific copy number variation (CNV) region upstream of Sox9 on autosomal chromosomes, including the Sox9 enhancer 14, which is involved in regulating Sox9 and Foxl2 expression, which are related to testicular differentiation13.
Habitat and phylogenetic relationships of three spiny rats. Karyotype information and evolutionary events assumed to have occurred in the ancestral species are shown. This map is based on a GSImap74 published by the Geospatial Information Authority of Japan. Photo courtesy: Kimiyuki Tsuchiya.
In contrast, T. muenninki exhibits an XX/XY sex chromosome configuration, similar to that of typical mammals; however, it contains a large neo-sex chromosome with autosomes, fused to the X and Y chromosomes14,15. A study showed that genes near the ancestral X and Y regions of the neo-X and neo-Y chromosomes accumulated male-specific mutations15. Unlike T. osimensis and T. tokunoshimensis, Sry has not been lost in T. muenninki and has multiple copies on the Y chromosome3,16. However, reportedly, Sry in T. muenninki is suspected to have lost its function as a sex-determining gene9,17. These three spiny rat species are suitable research subjects; however, their genome-based research has been limited to some RNA sequencing (RNA-Seq)-based15,18, fluorescence in situ hybridization-based14,19, and limited numbers of bacterial artificial chromosomes (BACs)-based16 studies, except for the T. osimensis genome, which has been studied by short-read sequencing13.
Herein, we provide the first report of high-quality, long-read-based chromosome-level genome assemblies and gene annotations for the three Tokudaia species. Our results can provide a valuable foundation for future studies regarding mammalian sex-determination mechanisms and sex chromosome evolution. Moreover, the constructed dataset may provide a valuable resource for the direct comparison of the genomes of species formed because of allopatric speciation over the last several million years.
Methods
Sample collection and sequencing
Whole-genome shotgun sequencing was performed using the PacBio and Illumina sequencing platforms. The genomic DNAs of T. osimensis, T. tokunoshimensis, and T. muenninki were isolated from the muscle tissues of male specimens that had accidentally perished (recovered in December 2005, December 2011, and March 2013, respectively) and had been stored in a deep freezer using the smart DNA prep Kit (Anlytik Jena, Jena, Germany). Genomic DNA from T. osimensis and T. muenninki was sheared into 30–100 kb fragments using a g-tube device (Covaris Inc., MA, USA). Although the DNA of T. tokunoshimensis was slightly degraded, this procedure was omitted. Subsequently, a continuous long-read (CLR) single-molecule real-time (SMRT) bell library was prepared using the SMRTbell Express Template Prep Kit 2.0 (Pacific Bioscience, CA, USA) according to the manufacturer’s instructions. Three CLR libraries were size-selected using the BluePippin system (Saga Science, MA, USA) with a lower cutoff of 40 kb. For each species, the library was run on two SMRT Cell 8Ms with Binding Kit 2.0 and Sequencing Kit 2.0. Through PacBio sequencing, 254.2 Gb, 272.1 Gb, and 271.8 Gb of CLRs were obtained for T. osimensis, T. tokunoshimensis, and T. muenninki, respectively (Table 1). Additionally, genomic DNA was fragmented to an average size of 500–600 bp using Focused-ultrasonicator M220 (Covaris Inc., MA. USA). Paired-end libraries with 450–550 bp insert sizes were constructed using the TruSeq DNA PCR-Free Library Prep kit (Illumina, CA, USA) and size-selected on an agarose gel using the Zymoclean Large Fragment DNA Recovery Kit (Zymo Research, CA. USA). The final libraries were sequenced following the 2 × 150 bp paired-end protocol for HiSeq 2500 (T. osimensis) and NovaSeq 6000 (T. tokunoshimensis and T. muenninki) systems (Illumina, San Diego, CA, USA). After Illumina sequencing, we obtained 448.1 Gb, 267.2 Gb, and 334.3 Gb of paired-end reads for T. osimensis, T. tokunoshimensis, and T. muenninki, respectively (Table 1). The Omni-C library was prepared using the Dovetail Omni-C Kit (Dovetail Genomics, Scott Valley, CA, USA) according to the manufacturer’s protocol. 1 × 106 cultured cells were collected from the same T. osimensis individual from which the genomic DNA was extracted. The pulverized sample was processed into a proximity ligation library using the Omni-C Proximity Ligation Assay Mammalian Samples Protocol of the Omni-C Kit. The final library was sequenced using NovaSeq 6000 (Illumina) with a 2 × 150 bp read length, and 145.3 Gb of Omni-C reads were generated for T. osimensis (Table 1). The Arima Hi-C libraries were constructed from the same tissue (98.1 mg) of T. tokunoshimensis and cerebrum (177.4 mg) of the same T. muenninki individual from which genomic DNAs were extracted, using the Arima-HiC + Kit (Arima Genomics, CA, USA) according to the manufacturer’s instructions regarding Animal Tissues (A160132 v01) and library preparation (A160137 v00) using the TruSeq DNA PCR-Free Library Prep kit (Illumina, CA, USA). The obtained Hi-C libraries were run on the Illumina NovaSeq 6000 system with a 2 × 150 bp read length, and 153.8 Gb and 164.0 Gb of Arima Hi-C reads were generated for T. tokunoshimensis and T. muenninki, respectively (Table 1).
Genome size and heterozygosity estimation
The genome size and heterozygosity of the three Tokudaia species were estimated using the Illumina sequencing data and a k-mer-based method. The sequenced reads from Illumina were filtered using Platanus_trim v1.0.7 (http://platanus.bio.titech.ac.jp/pltanus_trim) with default parameters. Using the trimmed reads, Jellyfish v2.3.020 was applied to extract and count the canonical k-mers at k = 32. Following this, GenomeScope 2.021 was used to estimate the haploid genome sizes and heterozygosities based on k-mer count data with parameters of “-k 32 -p 2” (Fig. 2). Thus, we estimated a haploid genome size of 2,472.5 Mbp with 0.53% heterozygosity for T. osimensis, 2,472.6 Mbp with 0.25% heterozygosity for T. tokunoshimensis, and 3,362.6 Mbp with 0.45% heterozygosity for T. muenninki (Fig. 2). The genome size estimation result that the T. muenninki has a genome size approximately 1 Gb larger than that of the other two species is consistent with a previous study showing that T. muenninki contains large heterochromatic blocks14.
K-mer analysis of three Tokudaia species genomes. (a) T. osimensis (b) T. tokunoshimensis (c) T. muenninki. The estimated haploid genome sizes for T. osimensis, T. tokunoshimensis, and T. muenninki were 2,472.5 Mbp with 0.53% heterozygosity, 2,472.6 Mbp with 0.25% heterozygosity, and 3,362.6 Mbp with 0.45% heterozygosity, respectively. T. muenninki, with a genome size approximately 1 Gb larger than that of the other two species, shows consistency with a previous study demonstrating the presence of large heterochromatic blocks.
De novo genome assembly
The PacBio sequenced reads were used for genome assembly using Canu v2.1.122, with the parameters of “corOutCoverage = 100 “batOptions = -dg 3 -db 3 -dr 1 -ca 500 -cp 50” -pacbio-raw”. The contigs were polished using two rounds of Arrow (Pacific Biosciences) and three rounds of NextPolish v1.4.023. Haplotigs that were considered redundant because of the separate construction of both haplotypes were removed using Purge_Dups v1.2.524. After removing redundant contigs, contigs with aberrant GC content (<2% or >98%), contaminant sequences derived from bacteria, parasitic organisms, such as Toxoplasma, and PacBio control sequences were removed based on the results of homology searches using BLASTN v2.11.0+25.
We performed Hi-C scaffolding using Hi-C datasets to obtain chromosome-level assemblies of the three Tokudaia species. The examination of input-purged assembly contigs before Hi-C scaffolding revealed the presence of sex chromosome-derived contigs that appeared to have been incorrectly removed by Purge_Dups. Therefore, Hi-C scaffolding was performed after reverting the sex chromosome-derived contigs, which were determined to have been erroneously deleted using the Hi-C read contact information. The Omni-C/Arima Hi-C reads were then mapped to the input contigs and processed to generate Hi-C contacts using Juicer v1.626. Thereafter, the Hi-C contacts files were used for scaffolding by 3D-DNA v18092227 with the following parameter settings: “–editor-coarse-resolution 50000–editor-coarse-region 250000–input 1000”. We visualized the Hi-C contact map using Juicebox v1.11.0828, checked the mapping information of sequence reads on Integrated Genomics Viewer (IGV)29, and performed extensive manual curation to fix mis-assemblies, mis-scaffoldings, and mis-foldings caused by duplicated regions. Additionally, the mitochondrial genome sequences were constructed from the Illumina short reads of each individual using GetOrganelle v1.7.5.030. All constructed mitochondrial sequences were circularized, and the obtained sequence lengths were 16,266, 16,263, and 16,250 bp for T. osimensis, T. tokunoshimensis, and T. muenninki, respectively.
We successfully constructed genomes of N50 = 234.0 Mbp, totaling 2,445.3 Mbp in T. osimensis; N50 = 125.1 Mbp, totaling 2,477.3 Mbp in T. tokunoshimensis; and N50 = 121.8 Mbp, totaling 2,660.8 Mbp in T. muenninki (Table 2). The length of constructed genomes of T. osimensis and T. tokunoshimensis were near their estimated genome sizes, 2,472.5 Mbp and 2,472.6 Mbp, respectively, whereas that of T. muenninki was only approximately 702 Mbp less than its estimated genome size of 3,362.6 Mbp. It was assumed to be because of the inability to construct the sequences of the heterochromatin region, which are highly repetitive. In T. osimensis, 99.79% of the assembled contigs were incorporated into 13 scaffolds, which corresponded to 12 autosomes and one X chromosome, suggesting that all 2n = 25 chromosomal karyotypes were constructed. Similarly, in T. tokunoshimensis, 99.87% of the assembled contigs were incorporated into 23 scaffolds, which corresponded to 22 autosomes and one X chromosome, suggesting that all 2n = 45 chromosomal karyotypes were constructed. Conversely, in T. muenninki, 91.31% of the assembled contigs were incorporated into 23 scaffolds, which were considered to correspond to 21 autosomes and the neo- and ancestral X regions in T. muenninki, with 2n = 44. For the ancestral Y region, the orientation and order of the candidate contigs could not be determined through Hi-C scaffolding, and chromosome-level scaffolds were not constructed. Therefore, sequences derived from the ancestral Y region were constructed as unplaced scaffolds. The unplaced scaffold accounted for 8.69% (231.3 Mbp) of the entire genome sequence, suggesting that it contained sequences of the heterochromatin region in addition to those of the ancestral Y region.
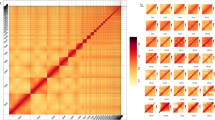
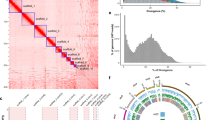
The chromosome numbers and orientations of the scaffolds were determined based on the synteny relationship examined among the three constructed chromosome-scale scaffolds and mouse chromosomes by extracting the best bidirectional alignments using minimap231 and corresponding them to the results of previous chromosome painting studies14,19. Figure 3 illustrates the Hi-C contact map constructed using the rearranged sequences. In all three species, the X chromosome had half of the coverage; therefore, the contacts were weak, whereas in T. muenninki, the fragmented Y chromosome-derived sequences were clustered in the lower-right corner of the contact map as unplaced scaffolds.
Genome-wide Hi-C contact map of three Tokudaia species genomes. The blue squares represent chromosomes. (a) Hi-C contact map of T. osimensis comprising 13 chromosome-level scaffolds (representing 12 autosomes and the X chromosome) (b) Hi-C contact map of T. tokunoshimensis comprising 23 chromosome-level scaffolds (representing 22 autosomes and the X chromosome) (c) Hi-C contact map of T. muenninki comprising 23 chromosome-level scaffolds (representing 21 autosomes and the neo- and ancestral X regions).
Gene structure prediction and functional annotation
Gene structures were predicted using the following methods: (1) an RNA-Seq-based method that predicts gene structures on the basis of transcriptome sequencing results, (2) a homology-based method that predicts gene structures on the basis of protein sequences of related species, and (3) an Ab initio method that learns the features of gene regions and predicts them directly from the genome sequence. The prediction results of the three methods were integrated using an integration tool. The three prediction methods are described below.
-
(1)
RNA-Seq-based method
For the RNA-Seq-based method, gene prediction was performed using both mapping and de novo methods using RNA-Seq data (T. osimensis: DRP00914932, T. tokunoshimensis: DRP01049433, and T. muenninki: DRP00343534 and DRP00413535). In the mapping-based method, HISAT2 v. 2.2.136 was used to map RNA-Seq reads to genomic sequences, and the mapped reads were assembled using StringTie v2.2.037. Next, de novo methods were used to assemble the RNA-Seq reads using the transcript assemblers Trinity v2.12.038 and Oases v2.0939. Redundant sequences were removed from the assembled contigs using CD-HIT v4.8.140 and the contigs were aligned to the genomic sequence using GMAP v2015–09–2941. Finally, open reading frames were determined using both mapping and de novo methods using TransDecoder (https://github.com/TransDecoder/TransDecoder) to predict gene structures.
-
(2)
Homology-based method
The protein sequences of eight mouse subfamily (Murinae) species annotated in NCBI RefSeq, Mus musculus42, Mus caroli43, Mus pahari44, Rattus norvegicus45, Rattus rattus46, Arvicanthis niloticus47, Grammomys surdaster48, Mastomys coucha49 were downloaded. They were splice-aligned to the genome sequence using Spaln v2.3.350 to predict gene structures.
-
(3)
Ab initio-based method
Augustus v3.351 was used to predict the gene structure. First, 1,000 randomly selected genes from the RNA-Seq-based predicted genes were selected to train the gene model, which was further used to predict the gene structure.
The results predicted by the three methods were integrated into the GINGER pipeline52. Additionally, predicted genes that satisfied all the requirements of the confidential M. musculus42 or R. rattus46 full-length homology-based results were extracted and integrated into the GINGER results. The requirements were as follows: Compared with mouse or rat orthologous genes, predicted genes should have (a) 80% or more identity and sequence coverage, (b) less than 3-amino acid length differences with all exons, and (c) all introns should be 0.5 to 2.0 times the length of the corresponding introns of the mouse or rat. If an exon overlapped between the newly predicted exons and the GINGER exon after integration, the exon with a longer CDS was used for the final result.
The statistics for the predicted gene sets are listed in Table 3. A total of 22,233, 22,106, and 22,240 protein-coding genes were predicted in T. osimensis, T. tokunoshimensis, and T. muenninki, respectively. These numbers were similar to those annotated in the closely related species mouse (22,173) and rat (22,215). Other statistics were similar to those for the mice and rats. Subsequently, the homology information of the mouse gene was assigned to the predicted genes as functional annotation. A homology search was performed using BLASTP v2.11.0+25 against the mouse RefSeq sequence42, and if a hit with more than 80% identity and coverage was found, the mouse annotation information was assigned to the corresponding predicted gene. Consequently, 18,469, 18,354, and 18,471 genes, which corresponded to approximately 83% of the total genes, were annotated for T. osimensis, T. tokunoshimensis, and T. muenninki, respectively.
Data Records
Genomic sequencing data (Illumina, PacBio, Hi-C) are available in the NCBI SRA database under BioProject ID PRJDB16410. The accession numbers of the Illumina sequencing data are DRR066821 and DRR06682253 (T. osimensis), DRR49570754 (T. tokunoshimensis), and DRR49571155 (T. muenninki). The accession numbers of the PacBio sequencing data are DRR49585156 and DRR49585257 (T. osimensis), DRR49570658 and DRR49570959 (T. tokunoshimensis), and DRR49571060 and DRR49571361 (T. muenninki). The accession numbers of the Hi-C sequencing data are DRR37886362 (T. osimensis), DRR49570863 (T. tokunoshimensis), and DRR49571264 (T. muenninki). The accession numbers of the assemblies are BTPL01000001– BTPL0100012365 (T. osimensis), BTHU01000001–BTHU0100015966 (T. tokunoshimensis), and BTHS01000001–BTHS0100152767 (T. muenninki). The accession number of mitochondrial genomes are LC778283.168 (T. osimensis), LC778284.169 (T. tokunoshimensis), and LC778282.170 (T. muenninki). The genome annotation files are available in the Figshare database71.
Technical Validation
DNA and RNA quality
Agarose gel electrophoresis was used to assess the quality of the extracted DNA. The main band was >20 kb, and the DNA spectrophotometer ratio (260 nm/280 nm) was >1.8. The quality of the purified RNA molecules was examined by 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA), and RNA integrity (RIN) was >7.0.
Assembly evaluation
The quality and completeness of the chromosome assembly were evaluated using two independent approaches. First, the QV value and completeness were estimated using Merqury v1.372 by comparing k-mers in the assembly with those found in the Illumina sequence reads. The results revealed that QV values of the assembly were 45.21, 47.85, and 45.32 for T. osimensis, T. tokunoshimensis, and T. muenninki, whereas completeness values were 93.71%, 93.72%, and 93.14%, respectively. Second, the completeness of the assembly was assessed using BUSCO v5.4.773 (genome mode) with 13,798 single-copy orthologs from the glires_odb10 database. The analysis revealed that 96.15%, 96.14%, and 96.30% of the complete BUSCOs were identified in the genomes of T. osimensis, T. tokunoshimensis, and T. muenninki, respectively.
Gene annotation evaluation
The completeness of the annotated protein-coding genes was assessed using BUSCO v5.4.773 (protein mode) with 13,798 single-copy orthologs from the glires_odb10 database. The analysis revealed that 98.70%, 98.80%, and 99.01% of the complete BUSCOs were identified in the annotated genes of T. osimensis, T. tokunoshimensis, and T. muenninki, respectively.
Code availability
No custom code was used in this paper, and all of the programs used were publicly available.
References
International Union for Conservation of Nature and Natural Resources. IUCN 2023. The IUCN Red List of Threatened Species. Version 2022-2. https://www.iucnredlist.org/ (2023).
Iryu, Y. et al. Introductory perspective on the COREF Project. Island Arc. 15, 393–406 (2006).
Murata, C., Yamada, F., Kawauchi, N., Matsuda, Y. & Kuroiwa, A. Multiple copies of SRY on the large Y chromosome of the Okinawa spiny rat, Tokudaia muenninki. Chromosome Res. 18, 623–634 (2010).
Honda, T., Suzuki, H. & Itoh, M. An unusual sex chromosome constitution found in the Amami spinous country-rat, Tokudaia osimensis osimensis. Jpn. J. Genet. 52, 247–249 (1977).
Honda, T., Suzuki, H., Itoh, M. & Hayashi, K. Karyotypical differences of the Amami spinous country-rats, Tokudaia osimensis osimensis obtained from two neighbouring islands. Jpn. J. Genet. 53, 297–299 (1978).
Tsuchiya, K., Wakana, S., Suzuki, H., Hattori, S. & Hayashi, Y. Taxonomic study of Tokudaia (Rodentia: Muridae) I, genetic differentiation. Memoirs of the National Science Museum. 22, 227–234 (in Japanese) (1989).
Sutou, S., Mitsui, Y. & Tsuchiya, K. Sex determination without the Y chromosome in two Japanese rodents Tokudaia osimensis osimensis and Tokudaia osimensis spp. Mamm. Genome. 12, 17–21 (2001).
Kuroiwa, A. et al. Additional copies of CBX2 in the genomes of males of mammals lacking SRY, the Amami spiny rat (Tokudaia osimensis) and the Tokunoshima spiny rat (Tokudaia tokunoshimensis). Chromosome res. 19, 635–644 (2011).
Kimura, R., Murata, C. & Kuroiwa, A. Mutations in the testis-specific enhancer of SOX9 in the SRY independent sex-determining mechanism in the genus Tokudaia. PLoS One. 9, e108779 (2014).
Otake, T. & Kuroiwa, A. Molecular mechanism of male differentiation is conserved in the SRY-absent mammal, Tokudaia osimensis. Sci. Rep. 6, 32874 (2016).
Arakawa, Y., Nishida-Umehara, C., Matsuda, Y., Sutou, S. & Suzuki, H. X-chromosomal localization of mammalian Y-linked genes in two XO species of the Ryukyu spiny rat. Cytogene. Genome Res. 99, 303–309 (2002).
Kuroiwa, A., Ishiguchi, Y., Yamada, F., Abe, S. & Matsuda, Y. The process of a Y-loss event in an XO/XO mammal, the Ryukyu spiny rat. Chromosoma. 119, 519–526 (2010).
Terao, M. et al. Turnover of mammal sex chromosomes in the Sry-deficient Amami spiny rat is due to male-specific upregulation of Sox9. Proc. Natl. Acad. Sci. USA 119, e2211574119 (2022).
Murata, C., Yamada, F., Kawauchi, N., Matsuda, Y. & Kuroiwa, A. The Y chromosome of the Okinawa spiny rat, Tokudaia muenninki, was rescued through fusion with an autosome. Chromosome Res. 20, 111–125 (2012).
Murata, C. et al. Initiation of recombination suppression and PAR formation during the early stages of neo-sex chromosome differentiation in the Okinawa spiny rat, Tokudaia muenninki. BMC Evol. Biol. 15, 234 (2015).
Murata, C., Kuroki, Y., Imoto, I. & Kuroiwa, A. Ancestral Y-linked genes were maintained by translocation to the X and Y chromosomes fused to an autosomal pair in the Okinawa spiny rat Tokudaia muenninki. Chromosome Res. 24, 407–419 (2016).
Ogata, Y. et al. Spiny rat SRY lacks a long Q-rich domain and is not stable in transgenic mice. Dev. Dyn. 248, 784–794 (2019).
Zushi, H., Murata, C., Mizushima, S., Nishida, C. & Kuroiwa, A. Unique XCI evolution in Tokudaia: initial XCI of the neo-X chromosome in Tokudaia muenninki and function loss of XIST in Tokudaia osimensis. Chromosoma. 126, 741–751 (2017).
Nakamura, T. et al. Comparative chromosome painting map between two Ryukyu spiny rat species, Tokudaia osimensis and Tokudaia tokunoshimensis (Muridae, Rodentia). Chromosome Res. 15, 799–806 (2007).
Marcais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat. Commun. 11, 1432 (2020).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736 (2017).
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics. 36, 2253–2255 (2020).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics. 36, 2896–2898 (2020).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics. 10, 421 (2009).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Robinson, J. T. et al. Integrated genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Jin, J. J. et al. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21, 241 (2020).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics. 34, 3094–3100 (2018).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRP009149 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRP010494 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRP003435 (2017).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRP004135 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Schulz, M. H., Zerbino, D. R., Vingron, M. & Birney, E. Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics. 28, 1086–1092 (2012).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics. 28, 3150–3152 (2012).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics. 21, 1859–1875 (2005).
FASTA format sequences of the protein products annotated on the Mus musculus genome assembly version GRCm39 (NCBI Mus musculus Annotation Release 109) https://ftp.ncbi.nlm.nih.gov/genomes/all/annotation_releases/10090/109/GCF_000001635.27_GRCm39/GCF_000001635.27_GRCm39_protein.faa.gz (2020)
FASTA format sequences of the protein products annotated on the Mus caroli genome assembly version CAROLI_EIJ_v1.1 (NCBI Mus caroli Annotation Release 100) https://ftp.ncbi.nlm.nih.gov/genomes/all/annotation_releases/10089/100/GCF_900094665.1_CAROLI_EIJ_v1.1/GCF_900094665.1_CAROLI_EIJ_v1.1_protein.faa.gz (2019).
FASTA format sequences of the protein products annotated on the Mus Pahari genome assembly version PAHARI_EIJ_v1.1 (NCBI Mus pahari Annotation Release 100) https://ftp.ncbi.nlm.nih.gov/genomes/all/annotation_releases/10093/100/GCF_900095145.1_PAHARI_EIJ_v1.1/GCF_900095145.1_PAHARI_EIJ_v1.1_protein.faa.gz (2019).
FASTA format sequences of the protein products annotated on the Rattus norvegicus genome assembly version mRatBN7.2 (NCBI Rattus norvegicus Annotation Release 108) https://ftp.ncbi.nlm.nih.gov/genomes/all/annotation_releases/10116/108/GCF_015227675.2_mRatBN7.2/GCF_015227675.2_mRatBN7.2_protein.faa.gz (2021).
FASTA format sequences of the protein products annotated on the Rattus rattus genome assembly version Rrattus_CSIRO_v1 (NCBI Rattus rattus Annotation Release 100) https://ftp.ncbi.nlm.nih.gov/genomes/all/annotation_releases/10117/100/GCF_011064425.1_Rrattus_CSIRO_v1/GCF_011064425.1_Rrattus_CSIRO_v1_protein.faa.gz (2020).
FASTA format sequences of the protein products annotated on the Arvicanthis niloticus genome assembly version mArvNil1.pat.X (NCBI Arvicanthis niloticus Annotation Release 100) https://ftp.ncbi.nlm.nih.gov/genomes/all/annotation_releases/61156/100/GCF_011762505.1_mArvNil1.pat.X/GCF_011762505.1_mArvNil1.pat.X_protein.faa.gz (2020).
FASTA format sequences of the protein products annotated on the Grammomys surdaster genome assembly version NIH_TR_1.0 (NCBI Grammomys surdaster Annotation Release 100) https://ftp.ncbi.nlm.nih.gov/genomes/all/annotation_releases/491861/100/GCF_004785775.1_NIH_TR_1.0/GCF_004785775.1_NIH_TR_1.0_protein.faa.gz (2019).
FASTA format sequences of the protein products annotated on the Mastomys coucha genome assembly version UCSF_Mcou_1 (NCBI Mastomys coucha Annotation Release 100) https://ftp.ncbi.nlm.nih.gov/genomes/all/annotation_releases/35658/100/GCF_008632895.1_UCSF_Mcou_1/GCF_008632895.1_UCSF_Mcou_1_protein.faa.gz (2019).
Gotoh, O. A space-efficient and accurate method for mapping and aligning cDNA sequences onto genomic sequence. Nucleic Acids Res. 36, 2630–2638 (2008).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics. 19, ii215–ii225 (2003).
Taniguchi, T. et al. GINGER: An integrated method for high-accuracy prediction of gene structure in higher eukaryotes at the gene and exon level. DNA Res. dsad017 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR066822 (2022).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495707 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495711 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495851 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495852 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495706 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495709 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495710 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495713 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR378863 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495708 (2023).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:DRR495712 (2023).
NCBI GenBank https://identifiers.org/ncbi/insdc:BTPL01000000 (2023).
NCBI GenBank https://identifiers.org/ncbi/insdc:BTHU01000000 (2023).
NCBI GenBank https://identifiers.org/ncbi/insdc:BTHS01000000 (2023).
NCBI GenBank https://identifiers.org/ncbi/insdc:LC778283.1 (2023).
NCBI GenBank https://identifiers.org/ncbi/insdc:LC778284.1 (2023).
NCBI GenBank https://identifiers.org/ncbi/insdc:LC778282.1 (2023).
Okuno, M. et al. Dataset for “Chromosomal-level assembly of Tokudaia osimensis, Tokudaia tokunoshimensis, and Tokudaia muenninki genomes”. FigShare https://doi.org/10.6084/m9.figshare.24105600.v1 (2023).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245 (2020).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO Update: Novel and Streamlined Workflows along with Broader and Deeper Phylogenetic Coverage for Scoring of Eukaryotic, Prokaryotic, and Viral Genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
GSImap published by the Geospatial Information Authority of Japan https://maps.gsi.go.jp/.
Acknowledgements
We thank the technical staff at the National Institute of Genetics and the Itoh Laboratory for library preparation, sequencing, and data management. This work was supported by JSPS KAKENHI, Grant Numbers 16H06279 (PAGS), 22H04925(PAGS), 22H02598, and 23K18093.
Author information
Authors and Affiliations
Contributions
Miki Okuno, Asato Kuroiwa, and Takehiko Itoh conceived the study. Takamichi Jogahara collected all the samples. Atsushi Toyoda performed the DNA extraction and sequencing. Luisa Matiz-Ceron performed the RNA extraction. Miki Okuno, Kentaro Matsuoka, Yuta Mochimaru, Takahiro Yamabe, and Takehiko Itoh performed the analysis. Miki Okuno, Asato Kuroiwa, and Takehiko Itoh wrote the manuscript. All authors read, edited, and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Okuno, M., Mochimaru, Y., Matsuoka, K. et al. Chromosomal-level assembly of Tokudaia osimensis, Tokudaia tokunoshimensis, and Tokudaia muenninki genomes. Sci Data 10, 927 (2023). https://doi.org/10.1038/s41597-023-02845-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02845-1
- Springer Nature Limited