Abstract
Compared to commercial chickens, local breeds exhibit better in meat quality and flavour, but the productivity (e.g., growth rate, body weight) of local chicken breeds is rather low. Genetic analysis based on whole-genome sequencing contributes to elucidating the genetic markers or putative candidate genes related to some economic traits, facilitating the improvement of production performance, the acceleration of breeding progress, and the conservation of genetic resources. Here, a total of 209 local chickens from 13 breeds were investigated, and the observation of approximately 91.4% high-quality sequences (Q30 > 90%) and a mapping rate over 99% for each individual indicated good results of this study, as confirmed by a genome coverage of 97.6%. Over 19 million single nucleotide polymorphisms (SNPs) and 1.98 million insertion-deletions (InDels) were identified using the reference genome (GRCg7b), further contributing to the public database. This dataset provides valuable resources for studying genetic diversity and adaptation and for the cultivation of new chicken breeds/lines.
Similar content being viewed by others
Background & Summary
Chickens are among the most important farm animals supplying eggs, meat, and other products to humans. After long-term domestication and human-driven selection, hundreds of distinctive chicken breeds are now cultivated worldwide, and chicken meat has been the largest meat resource since 20191. In particular, more than one hundred local breeds have been identified in China, constituting almost half of the broiler market. Compared to commercial broilers (e.g., Arbor Acre, Ross, and Cobb), local chickens exhibit markedly improved meat quality and flavour, adaptation to the environment, and disease resistance2,3. However, local breeds have not undergone intense artificial selection for productivity traits, which may explain their lower production performance compared to commercial breeds4,5. However, compared to pigs or cattle, chickens are more efficient and environmentally friendly livestock. Their feed conversion ratio is ten times that of cattle, and the carbon emissions of broilers are only 1/10 of those of cattle6. Therefore, it is necessary to investigate genomic markers and genetic mechanisms related to economic traits via whole-genome sequencing to bridge the gap in production performance and accelerate breeding progress7.
Simultaneously, domestic chickens are desirable models for investigating genetic adaptation and diversity and disease-related markers due to the advantages of their short reproductive cycle, small body size, and identical ancestors. After domestication from red jungle fowl (Gallus gallus spadiceus)8, chickens were cultivated with the aim of meeting various human demands, and meat-type (e.g., Xiaoshan chicken), egg-type (e.g., Xianju chicken), ornamental-type (e.g., Silkie chicken), and game fowl breeds (e.g., Luxi gamecocks, Henan gamecocks) were developed9,10,11,12, as well as the commercial broilers (e.g., broiler line B, Cobb)13. Therefore, it is relatively easy to unambiguously infer the driving factors of phenotypic or behavioural changes in these chickens. Additionally, domestic chickens can be used to generate special populations, such as F2 populations14,15 and advanced intercross lines16, which contributes to exploring the genetic mechanisms of chicken complex traits and provides new insights into genomic breeding.
Genomic analysis can reveal the demographic history of different chicken breeds and reconstruct gene flow among them, which contributes to a better understanding of the domestic history and potential mechanisms of some economic traits, such as breast muscle yield17, yellow skin4,18. The analysis of genome-wide variants in chickens distributed in different regions can be used to investigate genetic adaptation and diversity, especially adaptation to environmental conditions, such as altitude, temperature, and anoxic environments19,20,21,22. Moreover, selective sweep analysis based on single nucleotide polymorphisms (SNPs) is an effective method for identifying genetic markers and mechanisms underlying chicken production performance, reproduction, immunity, etc. This approach provides important insights into modern breeding systems20,21,22,23,24,25. In addition to SNPs and insertion‒deletions (InDels), genome-wide sequencing has also been used to identify copy number variants (CNVs)26 and structural variants (SVs)27, although the detection rate and efficiency of SV calling are relatively low compared to those of PacBio sequencing28. Therefore, genomic analysis using the whole-genome variants of local chickens is an effective approach for elucidating genetic diversity and selective signatures during long-term domestication.
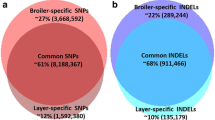
This study provides a whole-genome sequencing dataset from hundreds of local chickens (n = 209) of 13 local chicken breeds in China, including meat-type, egg-type, and ornamental-type chickens. A total of more than 19 million SNPs and 1.98 million InDels were identified on chromosomes 1–39 and one sex chromosome (Z) by aligning the sequencing reads against the most recent reference genome of chickens (assembly bGalGal1.mat.broiler. GRCg7b, GCF_016699485.2). The summarization of high-quality sequences (Q30 > 90%, mapping rate > 99%) confirmed the high accuracy and resolution of this genetic analysis. This dataset is expected to be useful in many studies, such as those aimed at identifying putative regions of positive selection, inferring demographic history, analysing gene flow, detecting candidate genes related to economic traits, determining genetic adaptation to local environmental factors, discovering breed-specific variants or markers, analysing genetic diversity, or developing SNP genotyping arrays for use in chicken breeding systems or species identification. Moreover, the whole-genome variants of most chicken breeds (except for Beijing You and Silky-feather chickens) included in the present study have not been reported. Therefore, this dataset provides an ideal resource for population genetics and evolutionary analyses. Furthermore, this database represents an important supplement to the chicken variant database and plays a crucial role in reconstructing the demographic and domestication history of chickens.
Methods
Sampling
Blood sampling from 13 chicken breeds was performed in Zhejiang Province, China. The following breeds were included (Supplement Table S1): BE, Baier chicken; BJY, Beijing You chicken; DXB, Dongxiang Black chicken; JSW, Jiangshan white-feathered chicken; LH, Luhua chicken; LY, Longyou chicken; SF, Silky-feather chicken; SYJ, Songyang Jin chicken; WL, Wenling chicken; XJ, Xianju chicken; XS, Xiaoshan chicken; XX, Xiaoxiang chicken; and YD, Yandang chicken. We collected a 1 ml blood sample from the wing vein of each individual and stored it in a 2 ml anticoagulation tube at −20 °C. All procedures associated with the chickens used in this study were consistent with the standards of the Laboratory Animal Guidelines for the Ethical Review of Animal Welfare and were approved by the Animal Use Committee of Zhejiang Academy of Agricultural Sciences (No. 2022ZAASLA68).
Genomic DNA extraction and quality control
The workflow from sampling to variant filtration is shown in Fig. 1. Genomic DNA was extracted from blood samples using the phenol‒chloroform method. DNA quality control was performed as follows: 1) DNA degradation and contamination were monitored on 1% agarose gels; 2) the OD 260/280 ratio was determined with a NanoDrop instrument to check the purity of the DNA; and 3) the DNA concentration was measured with a Qubit® DNA Assay Kit on a Qubit® 3.0 Fluorometer (Invitrogen, USA). Finally, more than 0.2 μg of DNA fragments with no degradation or contamination and an OD value of 1.8~2.0 were used for library construction.
Library preparation and sequencing
A sequencing library was created with the NEB Next UltraTM DNA Library Prep Kit for Illumina (NEB, USA). A mean size of 350 bp was achieved by shearing the genomic DNA. Before sequencing, the DNA fragments were subsequently submitted to end polishing, A-tailing, and adapter addition. PCR amplification and purification were then carried out using an AMPure XP system (Beckman Coulter, Beverly, USA). Using an Agilent 2100 Bioanalyzer and Qubit 3.0 Fluorometer (Invitrogen, USA), the quality of the library was evaluated based on the insert size and DNA concentration. Real-time PCR (>2 nM) was used to quantify the results. Ultimately, a flow cell containing the qualifying DNA nanospheres was filled, and a DNBSEQ-T7 platform was used for sequencing.
Data quality control, mapping, and variant calling
Clean reads were produced by trimming the raw reads using FASTP v0.21 software29, and reads meeting any of the following conditions were discarded: 1) one read of a pair contained adapter contamination (>10 nucleotides aligned to the adapter, allowing ≤10% mismatches); 2) more than 10% of bases were uncertain in either read; and 3) the proportion of low-quality (Phred quality <5) bases was over 50% in either read. The trimmed clean reads were evaluated by FastQC v0.12.1 software. Then, the clean reads were aligned against the most recent chicken reference genome (GRCg7b, GCF_016699485.2) using the Burrows‒Wheeler Aligner (BWA) v0.7.17 with the MEM algorithm30. The results were sorted, indexed, and converted to BAM files using SAMtools v1.12 software31. After removing polymerase chain reaction (PCR) duplicates, SNP calling was performed using the joint calling strategy, realized with the HaplotypeCaller, CombineGVCFs, and GenotypeGVCFs functions in the Genome Analysis Toolkit (GATK) v4.2.232. Next, we removed SNPs using GATK v4.2.2 according to specific standards: QualByDepth <2.0, FisherStrand >40.0, StrandOddsRatio >3.0, MappingQuality <40.0, and ReadPosRankSum <−8.0. A total of 19,820,641 biallelic SNPs and 1,980,710 biallelic InDels identified on autosomes after further quality control using VCFtools v0.1.1333 (--max-alleles 2 --min-alleles 2 --min-meanDP 3 --maf 0.01) were retained for subsequent analysis. The missing alleles were imputed by Beagle v5.1 software34 with the default parameters.
Variant annotation
The identified variants were annotated using ANNOVAR software35, the SNPs/InDels were classified according to the genomic location, such as exonic, intergenic, downstream, upstream, and others.
Variant density, genetic diversity, and polymorphism information content (PIC) estimation
The variant density was calculated as the ratio of the total length of the genome (with N bases removed) to the variant number. The SNP/InDel density was calculated and visualized with the CMplot package36. The genetic diversity (π) was calculated using VCFtools v0.1.1333 with window size of 50 kb. The PIC was estimated using the following formula: PIC = 1 − pi2 − qi2, where pi indicates the frequency of the minor allele of SNPi and qi indicates the frequency of the major allele of SNPi.
Data Records
Raw FASTQ files for whole-genome sequencing were deposited in the NCBI Sequence Read Archive (SRA) and have been assigned BioProject accession number PRJNA942350 (https://identifiers.org/ncbi/insdc.sra:SRP426730)37. The raw VCF file containing SNPs and InDels was deposited in the EVA database with accession number PRJEB71347 (https://identifiers.org/ena.embl:PRJEB71347)38. The quality control results of raw reads and annotation files of variants have been deposited in the the Figshare database with the following digital object identifier: https://doi.org/10.6084/m9.figshare.24751956.v239. The relationship between the chicken ID in the VCF files and the SRA database was shown in Supplement Table S2.
Technical Validation
Quality control of sequencing results
For each individual, an average of 17.3~28.5 Gb of raw data from were obtained, with 91.4% of the data achieving a Phred quality score of 30 (indicating sequencing accuracy of 99.9%) on average (Table 1). A stable GC content (42.6%) was demonstrated for the sequence (Table 1, Supplement Table S3). A genome mapping rate greater than 99% and an average genome coverage of 97.1% (with N bases removed) were obtained (Table 2, Supplement Table S4).
Filtration of SNP and InDel datasets
The joint calling strategy was used in this procedure, and a total of 27 million raw SNPs and 2.75 million raw InDels were identified in the population of 13 chicken breeds. To exclude low-quality variants, we used the VariantFiltration function in GATK software32. The specific standards used are described in the Methods section above. After the first round of quality control, the SNPs and InDels were further filtered using VCFtools v0.1.13 software33, and only biallelic variants with a minor allele frequency of 0.01 and a minimum sequencing depth of 3 were retained. We calculated statistics for SNP types, and T:A > C:G-type mutations were mainly identified in this population (Fig. 2). Figure 3 shows the relationships among SNP quality, supported read number, SNP percentage, and neighbouring SNP distance. We detected a positive correlation between SNP quality and percentage, and most SNPs were supported by at least 20 reads (Fig. 3a,b). This indicates the high quality of the identified SNPs.
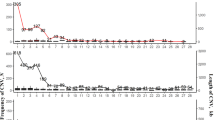
Summary statistics of SNPs and InDels across the whole genome
The high-quality SNPs and InDels were distributed across the genome with an average density of 1 SNP every 52 bases and 1 InDel every 521 bases (Table 3). Figures 3c, 4a show the high density of SNPs across the whole genome. The density of InDels was relatively low (Fig. 4b). Chromosomes 16, 29–32, and 34–38 were defined as dot chromosomes, while chromosomes 22, 25 and 33 were defined as microchromosomes40. We found that the SNP density on these chromosomes exhibited a polarized distribution (Table 3). Based on variants annotation, SNPs/InDels were mainly distributed in intronic and intergenic regions (Fig. 5). And more SNPs rather than InDels were located in exons (Fig. 5a). The PIC and genetic diversity π of each breed are shown in Fig. 6, the genomes of the SF and YD chickens exhibited higher polymorphism, and LH chickens were found significant lower genetic diversity.
Usage Notes
Whole-genome sequencing allows us to obtain the SNPs and InDels across the whole genome using the bioinformatic pipeline of this study. However, the CNVs and SVs were not included in the current study. As same as the previous report in human and chickens8,41,42,43, we did not consider the ploidy of variants in sex chromosome ZW. And the variants in chromosome W were discarded due to the uncertain gaps in chromosome. Although SNPs are more widely used for investigating investigate the genetic diversity and dissecting the genetic mechanism of economic traits, the heritability of traits of interest cannot be fully explained by SNPs, namely, missing heritability. These traits are also influenced by epigenetic factors. The effect of missing heritability could be relieved by increasing the sample size and sequencing depth and by considering the epistatic effects and SVs and CNVs, which also contributed to the identification of additional novel loci44. Additionally, we aligned the sequencing data to the reference genome (GRCg7b, GCF_016699485.2), which was assembled based on one broiler and did not include all the variants in the reference genome. Therefore, the data produced from this study may produce an incomplete explanation of the genetic background due to the missing alignment. Overall, our data provide insights into the evaluation of chicken population structure, and are an efficient dataset for identifying the CNVs and SVs. These data can be used in the construction of chicken pangenomes or graph pangenomes together with the Pacbio and Oxford Nanopore Technology data.
Code availability
The variant calling procedure was conducted in accordance with the standard bioinformatic workflow recommended by GATK, and all the steps were performed in the CentOS system. The detailed codes and parameters used in this study are provided as follows:
(1) Quality control for the raw reads:
Software: FASTP v0.21, FastQC v0.12.1
Code for trimming: fastp -I ${read1} -I ${read2} -o ${read1.qc} -O ${read2.qc} -q 5 -g -u 50 -n 15 -l 150 --min_trim_length 10 --overlap_diff_limit 1 --overlap_diff_percent_limit 10
Code for quality control: Fastqc -f fastq -t 6 -o ${sample.qc} ${read1.qc} ${read2.qc}
(2) Genome alignment:
Software: BWA-mem v0.7.17
Code: bwa mem -t 10 -M -R “@RG\tID:${individual}\tLB:${individual}\tPL:illumina\tSM:${individual}” /genome.index/bwa.index/gga7.${read1.qc} ${read2.qc} > ${individual}.sam
(3) Sorting, files converting, and indexing:
Software: SAMtools v1.12
Code for sorting and file converting: samtools sort -m 10 G -S ${individual}.sam -o ${individual}.sorted.bam -@ 10
Code for bam file indexing: samtools index -b ${individual}.sorted.bam ${individual}.sorted.bai
(4) Statistics for sequencing depth:
Software: Mosdepth v0.2.9
Code: mosdepth -t 6 ${individual.depth} ${individual.bam}
(5) Removing the duplicates
Software: Picard v2.26
Code: java -Xms100g -Xmx200g -jar picard.jar MarkDuplicates INPUT = ${individual}.sorted.bam OUTPUT = ${individual}.rmdup.bam M = ${individual}.metrices.txt REMOVE_DUPLICATES = true CREATE_INDEX = true
(6) Variants calling
Software: GATK v4.2.2
Code for the GVCF model generated using HaplotypeCaller: gatk --java-options “-Xmx60g -Xms20g” HaplotypeCaller --native-pair-hmm-threads 80 -R Gallus_gallus.bGalGal1.mat.broiler.GRCg7b.dna_sm.toplevel.fa -I ${individual}.rmdup.bam -ERC GVCF -O ${individual}.g.vcf
Code for combining GVCF files: gatk --java-options “-Xmx80g -Xms60g” CombineGVCFs -R Gallus_gallus.bGalGal1.mat.broiler.GRCg7b.dna_sm.toplevel.fa --variant individual1.g.vcf --variant individual2.g.vcf --variant individual3.g.vcf -O merge.g.vcf
Code for variant joint calling: gatk --java-options “-Xmx80g -Xms60g” GenotypeGVCFs -R Gallus_gallus.bGalGal1.mat.broiler.GRCg7b.dna_sm.toplevel.fa -V merge.g.vcf -O merge.vcf
(7) Variants extraction and hard filtration:
Software: GATK v4.2.2.0
Code for SNP extraction: gatk --java-options “-Xmx80g -Xms60g” SelectVariants -V merge.vcf -O raw.snp.vcf --select-type-to-include SNP
Code for InDel extraction: gatk --java-options “-Xmx80g -Xms60g” SelectVariants -V merge.vcf -O raw.indel.vcf --select-type-to-include INDEL --max-indel-size 50
Code for filtration: gatk --java-options “-Xmx80g -Xms60g” VariantFiltration -R Gallus_gallus.bGalGal1.mat.broiler.GRCg7b.dna_sm.toplevel.fa -V raw.snp.vcf -filter-expression QD < 2.0 || FS > 40.0 || SOR > 3.0 || MQ < 40.0 || ReadPosRankSum < −8.0” --filter-name filter -O filtered.snp.vcf
(8) Further filtration:
Software: VCFtools v0.1.13
Code: vcftools --vcf filtered.snp.vcf --max-alleles 2 --min-alleles 2 --min-meanDP 3 --remove-filtered-all –recode --out qc.snp
(9) Genotypes imputation:
Software: Beagle v5.1
Code: java -Xmx100g -Xms50g -jar beagle.jar gt = qc.snp.vcf impute = true nthreads = 20 out = qc.phased.snp
References
FAO. FAOSTAT Database https://www.fao.org/faostat/en/#data (2022).
Qanbari, S. et al. Genetics of adaptation in modern chicken. PLoS. Genet. 15, e1007989 (2019).
van der Most, P. J., de Jong, B., Parmentier, H. K. & Verhulst, S. Trade‐off between growth and immune function: a meta-analysis of selection experiments. Funct. Ecol. 25, 74–80 (2011).
Zhang, C. et al. Widespread introgression in Chinese indigenous chicken breeds from commercial broiler. Evol. Appl. 12, 610–621 (2019).
Tan, X. D. et al. Large-Scale Whole Genome Sequencing Study Reveals Genetic Architecture and Key Variants for Breast Muscle Weight in Native Chickens. Genes (Basel). 13, 3–17 (2021).
IPCC. Climate Change 2021: The Physical Science Basis. Contribution of Working Group I to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change. (Cambridge University Press, 2021).
Tan, X. D. et al. Research progress and applications of genome‐wide association study in farm animals. Anim. Res. One. Health. 1, 56–77 (2023).
Wang, M. S. et al. 863 genomes reveal the origin and domestication of chicken. Cell. Res. 30, 693–701 (2020).
Muir, W. M. et al. Genome-wide assessment of worldwide chicken SNP genetic diversity indicates significant absence of rare alleles in commercial breeds. Proc. Natl. Acad. Sci. USA 105, 17312–17317 (2008).
Jiang, M. et al. Effects of balanced selection for intramuscular fat and abdominal fat percentage and estimates of genetic parameters. Poult. Sci. 96, 282–287 (2017).
Ren, X. et al. Systematic Selection Signature Analysis of Chinese Gamecocks Based on Genomic and Transcriptomic Data. Int. J. Mol. Sci. 24 (2023).
Ren, X. et al. Extensive intra- and inter-genetic admixture of Chinese gamecock and other indigenous chicken breeds revealed by genomic data. Poult. Sci. 102, 102766 (2023).
Tan, X. D. et al. Assessment the effect of genomic selection and detection of selective signature in broilers. Poult. Sci. 101, 101856 (2022).
Nadaf, J. et al. Identification of QTL controlling meat quality traits in an F2 cross between two chicken lines selected for either low or high growth rate. BMC Genomics. 8, 155 (2007).
He, Y. et al. Identification of New Genes and Genetic Variant Loci Associated with Breast Muscle Development in the Mini-Cobb F2 Chicken Population Using a Genome-Wide Association Study. Genes (Basel). 13, 2153–2170 (2022).
Wang, Y. et al. Multiple ancestral haplotypes harboring regulatory mutations cumulatively contribute to a QTL affecting chicken growth traits. Commun. Biol. 3, 472 (2020).
Tan, X. D. et al. Large-scale genomic and transcriptomic analyses elucidate the genetic basis of high meat yield in chickens. J. Adv. Res. S2090-1232(23)00067-X (2023).
Eriksson, J. et al. Identification of the yellow skin gene reveals a hybrid origin of the domestic chicken. PLoS. Genet. 4, e1000010 (2008).
Shi, S. et al. Whole genome analyses reveal novel genes associated with chicken adaptation to tropical and frigid environments. J. Adv. Res. 47, 13–25 (2022).
Gheyas, A. A. et al. Integrated Environmental and Genomic Analysis Reveals the Drivers of Local Adaptation in African Indigenous Chickens. Mol. Biol. Evol. 38, 4268–4285 (2021).
Zhong, H. A. et al. Microevolutionary mechanism of high-altitude adaptation in Tibetan chicken populations from an elevation gradient. Evol. Appl. 15, 2100–2112 (2022).
Elbeltagy, A. R. et al. Natural Selection Footprints Among African Chicken Breeds and Village Ecotypes. Front. Genet. 10, 376 (2019).
Ding, J. et al. A significant quantitative trait locus on chromosome Z and its impact on egg production traits in seven maternal lines of meat-type chicken. J. Anim. Sci. Biotechnol. 13, 96 (2022).
Psifidi, A. et al. Genome-wide association studies of immune, disease and production traits in indigenous chicken ecotypes. Genet. Sel. Evol. 48, 74 (2016).
Suminda, G. G. D. et al. High-throughput sequencing technologies in the detection of livestock pathogens, diagnosis, and zoonotic surveillance. Comput. Struct. Biotechnol. J. 20, 5378–5392 (2022).
Chen, X. et al. Population Genomic Sequencing Delineates Global Landscape of Copy Number Variations that Drive Domestication and Breed Formation of in Chicken. Front. Genet. 13, 830393 (2022).
Geibel, J., Praefke, N. P., Weigend, S., Simianer, H. & Reimer, C. Assessment of linkage disequilibrium patterns between structural variants and single nucleotide polymorphisms in three commercial chicken populations. BMC Genomics. 23, 193 (2022).
Zhang, J. et al. Comprehensive analysis of structural variants in chickens using PacBio sequencing. Front. Genet. 13, 971588 (2022).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Vasimuddin, M., Misra, S., Li, H. & Aluru, S. in 2019 IEEE International Parallel and Distributed Processing Symposium (IPDPS). 314–324 (IEEE).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome. Res. 20, 1297–1303 (2010).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics. 27, 2156–2158 (2011).
Browning, B. L., Zhou, Y. & Browning, S. R. A One-Penny Imputed Genome from Next-Generation Reference Panels. Am. J. Hum. Genet. 103, 338–348 (2018).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic. Acids. Res. 38, e164 (2010).
Yin, L. et al. rMVP: A Memory-efficient, Visualization-enhanced, and Parallel-accelerated Tool for Genome-wide Association Study. Genom. Proteom. Bioinf. 19, 619–628 (2021).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP426730 (2023).
European Variation Archive https://identifiers.org/ena.embl:PRJEB71347 (2023).
Tan, X. D. Annotation for genomic variants from 209 local chickens. figshare. figshare https://doi.org/10.6084/m9.figshare.24751956.v2 (2023).
Huang, Z. et al. Evolutionary analysis of a complete chicken genome. Proc. Natl. Acad. Sci. USA 120, e2216641120 (2023).
Wang, K. et al. The Chicken Pan-Genome Reveals Gene Content Variation and a Promoter Region Deletion in IGF2BP1 Affecting Body Size. Mol. Biol. Evol. 38, 5066–5081 (2021).
Wang, M. S. et al. An Evolutionary Genomic Perspective on the Breeding of Dwarf Chickens. Mol. Biol. Evol. 34, 3081–3088 (2017).
Davis, E. J. et al. Sex-Specific Association of the X Chromosome With Cognitive Change and Tau Pathology in Aging and Alzheimer Disease. JAMA. Neurol. 78, 1249–1254 (2021).
Visscher, P. M., Brown, M. A., McCarthy, M. I. & Yang, J. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
Acknowledgements
This study was funded by grants from the Zhejiang Science and Technology Major Program on Agricultural New Variety Breeding (No. 2021C02068-9), Major Program of Agricultural Biological Breeding (2023ZD0406403), and the Key Research and Development Program of Zhejiang Province (No. 2021C02026-2).
Author information
Authors and Affiliations
Contributions
X.T. and D.W. conceived the research program. J.Z., Q.L., H.W., L.B. and M.C. collected blood samples. X.T. conducted the bioinformatic analysis. X.T. wrote the draft. J.D., M.H., Z.Z. and S.Y. edited the manuscript. All authors contributed critically to the draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information(The order of supplementary table should be changed, the table S1 should be arranged to the front)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tan, X., Zhang, J., Dong, J. et al. Whole-genome variants dataset of 209 local chickens from China. Sci Data 11, 169 (2024). https://doi.org/10.1038/s41597-024-02995-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-02995-w
- Springer Nature Limited