Abstract
Historically, intense forest hazards have resulted in an increase in the quantity of fallen wood in the Qinling Mountains. Fallen wood has a decisive influence on the nutrient cycling, carbon budget and ecosystem biodiversity of forests, and fungi are essential for the decomposition of fallen wood. Moreover, decaying dead wood alters fungal communities. The development of high-throughput sequencing methods has facilitated the ongoing investigation of relevant molecular forest ecosystems with a focus on fungal communities. In this study, fallen wood and its associated fungal communities were compared at different stages of decomposition to evaluate relative species abundance and species diversity. The physical and chemical factors that alter fungal communities were also compared by performing correspondence analysis according to host tree species across all stages of decomposition. Tree species were the major source of differences in fungal community diversity at all decomposition stages, and fungal communities achieved the highest levels of diversity at the intermediate and late decomposition stages. Interactions between various physical and chemical factors and fungal communities shared the same regulatory mechanisms, and there was no tree species-specific influence. Improving our knowledge of wood-inhabiting fungal communities is crucial for forest ecosystem conservation.
Similar content being viewed by others
Introduction
The Qinling Mountains in central China are an important climate boundary between the southern subtropics and the northern temperate zone where the typical types of vegetation from both climate zones assemble together, resulting in astonishingly high biodiversity in a veritable plant “kingdom”1. However, in recent decades, the Huoditang Forest Region in the Qinling Mountains has been constantly affected by wind and pests, resulting in increased quantities of fallen wood of two major species: Pinus tabulaeformis and Quercus aliena var. acuteserrata. Fallen wood is defined as downed or leaning deadwood (>45° from the vertical) with a minimum diameter ≥10 cm at the widest point and a length ≥1 m2,3. Fallen wood plays an important role in the forest ecosystem by providing wildlife habitats, facilitating nutrient cycling, and supplying carbon and nitrogen sources for micro-communities4,5,6. Fallen wood is also the substrate for multicellular forest species, acts as a seedbed and is a key component of forest biodiversity7,8,9,10.
The ecological function of fallen wood is primarily realized via the process of decomposition, during which fallen wood releases carbon, nitrogen, phosphorus and other nutrients11,12. Different trees undergo different wood decay processes. Decomposition is controlled by climatic factors as well as biome factors. Wood-decaying fungi primarily affect fallen wood decomposition at specific locations and are among the most common saproxylic organisms, comprising thousands of taxa13. Much of our knowledge of the process of wood decomposition in forest ecosystems involves wood-inhabiting fungi. Wood properties, including nutrients, lignin and stoichiometry, control or are controlled by fallen wood-inhabiting organisms14,15,16. To advance our understanding of modern forest ecosystems and the ecology of dead wood, it is essential to characterize the interaction between fallen wood and wood-inhabiting fungi.
Molecular studies focused on fungal communities associated with fallen wood at different stages of decomposition are being carried out with increasing frequency17. Modern molecular techniques improve our understanding of fallen wood decomposition. Sequence-based studies of DNA obtained directly from fallen wood have revolutionized our view of the wood-inhabiting fungal community18. The comparison of terminal restriction fragment length polymorphisms (RFLPs) targeting the internal transcribed spacer (ITS) regions of rRNA genes with high-throughput sequencing methods enables detailed, semiquantitative analysis of fungal communities in large sample nests and provides ecological information that extends far beyond terminal RFLPs in terms of both detail and magnitude17. Consequently, the use of high-throughput sequencing to study fungal communities increases accuracy and reliability.
Changes in wood structure and chemical composition during wood decomposition (primarily due to fungi) result in further changes to species composition and alter later decomposition processes. Fungal species differ during specific stages of decomposition, indicating that structural attributes are important when determining species assemblages, and fallen wood must be examined at different stages of decomposition to construct fungal assemblages19. Multilinear studies have examined fungal communities during the decomposition of fallen wood obtained from different trees, located in different regions, and demonstrating different levels of community diversity. For example, in a study of Norway spruce fallen wood in Sweden, fungal diversity tended to increase with sample decay stage20. Tree species diversity is an important factor in fungal species composition21, as species-rich communities exert negative impacts on the rate of decomposition22. However, species richness increases the decay rate due to different resource requirements and activity patterns among fungal species, as well as different environmental conditions23. There is still no consensus regarding the factors that impact fungal community diversity, and only a few publications have investigated differences in decay-associated fungi on different fallen wood hosts.
Hence, our objectives in this study were (i) to study the fungal species diversity and relative abundance of P. tabulaeformis and Q. aliena var. acuteserrata and (ii) to compare the physical and chemical factors within these two different fallen wood host species that affect the wood-inhabiting fungal community during decomposition. Knowledge of wood-inhabiting fungal communities, which also control the wood decay process, is important for both forest ecosystem conservation and to understand fungal community patterns within decaying dead wood.
Results
Overview of sequence assignments
A total of 371,130 sequencing reads were obtained from all samples, and we analysed 339,716 optimized, useful reads (Table 1). The reads obtained from P. tabulaeformis samples accounted for 159,221 of the total reads, with an average sequence length of 301.32 bp, while the 180,495 reads obtained from Q. aliena var. acuteserrata samples had an average sequence length of 301.686 bp.
Fungal community structure during decomposition succession
Fungal community members at different taxonomic levels and different stages of decomposition for two studied tree species are summarized in Table 2. These data provide an overview of the fungal community structures of these two species. We obtained 94–298 operational taxonomic units (OTUs) from Q. aliena var. acuteserrata samples and 125-263 OTUs from P. tabulaeformis samples at the species level. P. tabulaeformis samples exhibited higher fungal species diversity than Q. aliena var. acuteserrata samples at the family, genus and species levels. Thus, P. tabulaeformis supports a more diverse fungal community than Q. aliena var. acuteserrata.
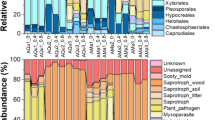
The majority of the fungal communities derived from both the P. tabulaeformis and Q. aliena var. acuteserrata samples consisted of the phyla Basidiomycota and Ascomycota (Fig. 1A). Fungal communities were categorized by class (Fig. 1B), and the relative abundance of each fungal community differed (Fig. 1). In Q. aliena var. acuteserrata samples, 5 fungal classes were observed: Agaricomycetes (41.72%), Eurotiomycetes (30.13%), Dothideomycetes (12.34%), Lecanoromycetes (8.28%) and Sordariomycetes (3.73%). In P. tabulaeformis samples, 3 classes were observed: Agaricomycetes (74.80%), Sordariomycetes (18.91%) and Eurotiomycetes (3.37%).
Fungal community composition heat maps at different stages of decomposition for these two wood species are displayed in Fig. 2, revealing relative abundance at the genus level. Most of Fig. 2 is coloured black or green. A few bars are colour-coded orange to red to indicate their abundance levels. Seven samples each contained a dominant fungal genus with a relative abundance of over 50%. For Q. aliena var. acuteserrata samples, the relative abundances of both the Lecania genus and the Galerina genus were 58% in RCL-II. An unclassified Agaricomycetes-no rank genus with a relative abundance of 69% was identified in RCL-IV samples. Among P. tabulaeformis samples, the Resupinatus genus had a relative abundance of 99% in YS-IV, the Ophiostoma genus had a relative abundance of 84% in YS-I, the Melanophyllum genus had a relative abundance of 67% in YS-V and the Sistotrema genus had a relative abundance of 63% in YS-III.
The rank abundance curves for OTUs from all samples provided relative abundance levels and visualized both fungal species richness and evenness (Fig. 3). Certain Q. aliena var. acuteserrata samples, specifically RCL-I, RCL-III and RCL-IV, contained fewer fungal species and exhibited low species evenness, particularly RCL-III. The others, RCL-II and RCL-V, had similar slopes for their best-fit lines, with the exception of the length of the X-axis for RCL-V. The two RCL-V samples contained more fungal species and demonstrated higher species evenness. The P. tabulaeformis samples exclusive to YS-II contained many fungal species types, similar to RCL-V, and exhibited similar high relative abundance, although RCL-V had the highest abundance among all samples.
Cluster analysis based on Euclidean similarity distance, together with non-metric multidimensional scaling (NMDS) based on Bray-Curtis similarity distance, revealed similarities in species composition at the genus level (Fig. 4). The NMDS stress value (0.05) indicates excellent ordination representation. Fungal community composition significantly differed with the progression of decomposition. The largest differences in fungal community composition for both P. tabulaeformis and Q. aliena var. acuteserrata samples appeared during decomposition stage IV and then narrowed somewhat compared with the composition during initial state I. Nevertheless, the difference in fungal community composition for P. tabulaeformis narrowed only slightly between decomposition stage IV and V, unlike Q. aliena var. acuteserrata. Furthermore, both tree species exhibited a high degree of similarity in terms of fungal community composition during decomposition stages I and III and high discrepancy levels during stages IV and V.
Correspondence analysis
Correlations between the physical and chemical parameters of each fallen wood species and fungal community were discerned from a correspondence analysis, as shown in Figs 5 and 6, respectively. The length of a physical or chemical parameter arrow indicates the strength of that parameter with regard to the overall fungal community. The influence of each physical or chemical parameter on Q. aliena var. acuteserrata was approximate.
Density and moisture appeared to be the most important physical and chemical parameters. Carbon, potassium, phosphorus, magnesium, calcium and nitrogen were the secondary impact factors. For P. tabulaeformis, density and nitrogen were the most powerful physical and chemical parameters, while the influence of minor factors, i.e., carbon, potassium, phosphorus, magnesium, calcium and moisture, on the fungal community was relatively weak compared with their influence in the Q. aliena var. acuteserrata fungal community.
Discussion
We obtained sufficient sequencing outputs for the pine species P. tabulaeformis and the oak species Q. aliena var. acuteserrata. The rarefaction curves of fungal OTUs were plotted to test the reliability of the sequence outputs and to predict the maximum number of OTUs at a similarity level of 97% (Fig. 7). Some of the rarefaction curves plotted based on these results failed to reach saturation, but coverage estimators using Chao124,25 reflected the enormous diversity found in fallen wood samples. These samplings, RCL-II, YS-I and YS-V, initially stabilized their curves at 20,000 reads. When the number of reads reached 40,000, the curve of each sample became steady, indicating that the sequencing results were adequate and reasonable.
Fungal richness obviously increased during the final stage for Q. aliena var. acuteserrata, while only a slight increase occurred for P. tabulaeformis. Baldrian26 reported a slight increase during the final decomposition stage in temperate forests, which is similar to what we observed for P. tabulaeformis fallen wood. Fukasawa27 observed a peak fungal richness during the final stage in decayed fallen wood, which explains our findings in Q. aliena var. acuteserrata. We assume that the significant differences between the substrates provided by P. tabulaeformis and Q. aliena var. acuteserrata explain these differences in species richness, although further studies are necessary to investigate this issue.
Direct sequencing of fallen wood samples elucidated the fungal community at different stages of decomposition. Fallen wood during various stages of decomposition provides habitats for a variety of organisms, including fungal communities28. The phyla Basidiomycota and Ascomycota are the most frequently encountered wood-inhabiting fungi29,30 in fallen wood. Classes Agaricomycete, Sordariomycetes and Eurotiomycetes were all observed in both P. tabulaeformis and Q. aliena var. acuteserrata fallen wood. Arslanova suggested the existence of fungal communities in both coniferous and broad-leafed species31 in her dissertation. Decaying pine wood during all decomposition stages exhibits high diversity among wood-inhabiting fungi32. In her dissertation, Arslanova mentioned the existence of the fungal community in both coniferous and broad-leafed species31. The fungal communities identified at the class level in both pine and oak fallen wood samples aligned with the results of Kirker33. The unclassified genus belonging to Agaricomycetes, observed in RCL-IV with an abundance of 69%, was unexpected but explainable because the morphology of Agaricomycetes demonstrates an unparalleled diversity34. A similar taxonomic distribution was present across all of the samples from both tree species at the phylum and class levels. Visible differences in the taxonomic distribution first attracted our attention at the genus level. A distinct separation was observed for the species diversity of dead wood fungi based on their host tree. Host tree species are known to delimit wood-inhabiting fungi. Bîrsan35 found that this separation occurred even among three coniferous host trees; additionally, Küffer36 verified the distinction between coniferous and broadleaf host tree species. Differences in fungal species abundance in our study were consistent with these patterns.
Decaying fallen wood provides a snapshot of the nutrients and organic substances available for wood-inhabiting microorganisms37. Fungi predominantly prefer middle and late decomposition succession stages38. We speculate that the lifestyle of the observed fungi is saprophytic, allowing them to invade heavily decayed wood. Wood decomposition represents a stabilizing force within dead wood39, and a new equilibrium is reached when decomposition reaches stage V. Changes such as decreased wood density, a decreased C/N ratio and increased mineral and nitrogen content develop during the decomposition succession of fallen wood. As density decreases, fungal richness peaks during the intermediate decomposition stages, such as decay classes III and IV40. A lower C/N ratio, together with higher mineral and nitrogen contents, in dead wood during late decomposition succession stages results in the availability of nitrogen sources for wood-inhabiting fungi, regardless of forest floor vegetation41. Thus, there are clear changes in fungal community diversity with the succession of decomposition.
Fungal communities from Q. aliena var. acuteserrata and P. tabulaeformis reflect physical and chemical factors. Correlations between physical and chemical factors are rarely tree species-specific42. Unique and mutual fungal OTUs derived from sequences play critical roles, and discrepancies in fungi explain the mildly different results. Density and moisture (precipitation) are factors that differ with different wood decomposition stages43 and exert the most significant effects. Analysis has indicated that nitrogen, phosphorus, and the C/N ratio are determining factors for fungal composition44. A meta-analysis of 36 studies from all forested continents revealed that nitrogen, phosphorus, and the C/N ratio correlate with the decomposition rates of angiosperms45. Wood decomposition results from the activities of decomposer organisms. The presence of fungal communities and their structure, dynamics, and diversity are explicit and susceptible to the status of decaying wood46. Thus, factors controlling the decomposition rates of angiosperms control the associated fungal communities.
This study focused on comparing fungal community diversity and richness during the decomposition of P. tabulaeformis and Q. aliena var. acuteserrata fallen wood. Furthermore, we analysed correlations among the fungal community, the physical and chemical factors impacting P. tabulaeformis and Q. aliena var. acuteserrata fallen wood and the stages of decomposition. Differing tree species were the major source of differences in fungal community structure during all stages of decomposition, and the fungal community reached its highest diversity levels during the intermediate and late decomposition stages. The physical and chemical factors and fungal communities of Q. aliena var. acuteserrata and P. tabulaeformis shared the same regulatory mechanisms, and there were no tree species-specific influences. Thus, this comparative study provides sufficient evidence to understand the relationship between the decomposition of fallen wood and fungal community dynamics in natural forests and will aid the accurate prediction of shifts in fungal community composition and function according to the decomposition stage and tree species of fallen wood. In addition, our study elucidated physical and chemical factors influencing fungal community structure during different decomposition stages in fallen wood, providing a theoretical basis for the protection of fungal resources.
Methodology
Study site
The total study area of 2037 ha is located in the Huoditang Experimental Forest Farm of Northwest A&F University in the Qinling Mountains, Shaanxi Province, China. The climate belongs to a warm temperate zone, with a mean annual temperature ranging from 8-10 °C, annual precipitation ranging from 900–1200 mm, and a frost-free period of 170 days. The abrupt and broken topography mainly consists of granite and gneiss with a mean slope of 35°. The soil is classified as loam with a mean soil depth of 45 cm. Human activities in this region have largely disappeared since the natural forest protection project was initiated in 1998.
In the summer of 2013, we plotted P. tabulaeformis and Q. aliena var. acuteserrata forests into six sites. The plots were 60 m × 60 m rectangles with three replicates for both the P. tabulaeformis and Q. aliena var. acuteserrata forests. According to an investigation of fallen wood in each plot, most logs were e 20-30 cm in size. Thus, fallen wood with a diameter of 25 ± 5 cm was selected in each plot. Each plot was distributed on nearly flat terrain with similar site conditions. In the P. tabulaeformis forest, the altitude was 1484–1564 m, and the geographic coordinates were N33°25′11″−33°26′05″ and E108°26′21″−108°27′11″. The P. tabulaeformis forest was 60 years old and was dominated by P. tabulaeformis (75% of all trees), with a forest canopy density of 70%. The mean stand height, diameter at breast height (DBH) and stand density were 16 m, 24 cm and 1328 trees · ha−1, respectively. For the Q. aliena var. acuteserrata forest, the altitude was 1567-1621 m, and the geographic coordinates were N33°25′53″−33°26′21″ and E108°26′08″−108°26′21″. The Q. aliena var. acuteserrata forest was 50 years old and was dominated by Q. aliena var. acuteserrata (70% of all trees), with a forest canopy density of 75%. The mean stand height, DBH and stand density were 13 m, 18 cm and 1735 trees · ha−1, respectively.
Sampling
We conducted sampling in July 2013. Samples of fallen wood were collected and processed in the same manner47 and were assigned to one of five decay classes based on discrepancies in internal and external tissue characteristics48 with sufficient (at least three) replicates. A total of 100 samples were collected and cut using a handsaw, while late decomposition succession samples were simply transplanted into a sterile aluminium box. Within each site, upper parts of the cross-section were preferred to obtain soil microorganisms. Each piece fallen wood was sampled at both ends and in the middle and shipped to the laboratory immediately. A −80 °C refrigerator was used to store the samples.
DNA extraction and sequencing
Samples from the same pieces of fallen wood were mixed into one sample after liquid nitrogen grinding at a frequency of 30 Hz for 2 min. DNA was isolated using a PowerSoil® DNA Isolation Kit (MO BIO Corp.) according to the manufacturer’s instructions. High-throughput sequencing was performed with a GeneAmp® 9700 sequencer (ABI Corp.). Primers were designed based on the fungal ITS1 region. The forward primer was 1737F (GGA AGT AAA AGT CGT AAC AAG G), and the reverse primer was 2043 R (GCT GCG TTC ATC GAT GC), both with inserted barcodes. Each 20-μl PCR reaction consisted of 4 μl of 5 × FastPfu Buffer, 2 μl of dNTPs (2.5 mM), 0.4 μl of forward primer (5 μM), 0.4 μl of reverse primer (5 μM), 0.4 μl of FastPfu Polymerase, 10 ng of template DNA and ddH2O. The following protocol was used for PCR: initial denaturation at 95 °C for 2 min, followed by 30 cycles of 95 °C for 30 s, 50 °C for 30 s, and 72 °C for 30 s, with a final extension at 72 °C for 5 min. PCR products were recovered using an AxyPrepDNA Gel Recovery Kit (Axygen Corp.) and were quantitatively detected with a QuantiFluor™-ST fluorometer (Promega Corp.). A MiSeq library was constructed based on the combination of appropriately proportioned DNA fragments and was sequenced (2 × 300 BP) on an Illumina platform.
Data analysis
Biological statistical analysis of OTUs was conducted at the 97% similarity level using Usearch (version 7.1 http://qiime.org/)49,50. The RDP Bayesian algorithm was used to classify the representative sequences, and the community composition of each sample was statistically analysed at various levels (phylum, class, order, family, genus and species). Fungal ITS region sequences were identified based on reference sequences in the public database Unite (Release 5.0 http://unite.ut.ee/index.php). Taxonomy analysis was performed using Qiime51 (http://qiime.org/scripts/assign_taxonomy.html) and RDP Classifier52 (version 2.2 http://sourceforge.net/projects/rdp-classifier/). Mothur53 was used for diversity analysis, and cluster analysis and NMDS were conducted for community similarity analysis. Relative graphic progress was tracked using R i386 3.1.2. Correspondence analysis was performed to investigate the effects of physical and chemical factors on the fungal community at different stages of decomposition and was visualized using PC-ORD 5.0. Q. aliena var. acuteserrata was abbreviated to RCL, and P. tabulaeformis was abbreviated to YS, according to Mandarin pronunciations. I, II, III, IV and V represent different decomposition stages, i.e., I means the initial stage, while V means the final stage.
References
Yuan, J. et al. Characteristics in coarse woody debris mediated by forest developmental stage and latest disturbances in a natural secondary forest of Pinus tabulaeformis. Acta Ecologica Sinica 34, 232–238 (2014).
Ringvall, A. & Ståhl, G. Field aspects of line intersect sampling for assessing coarse woody debris. Forest Ecol. Manag. 119, 163–170 (1999).
Yuan, J. et al. Impacts of CWD on understory biodiversity in forest ecosystems in the Qinling Mountains, China. Pak. J. Bot. 47, 1855–1864 (2015).
Harmon, M. E. et al. Ecology of coarse woody debris in temperate ecosystems. Adv. Ecol. Res. 15, 133–302 (1986).
Wilcke, W. et al. Coarse woody debris in a montane forest in Ecuador: mass, C and nutrient stock, and turnover. Forest Ecol. Manag. 205, 139–147 (2005).
Freedman, B. et al. Biodiversity implications of changes in the quantity of dead organic matter in managed forests. Environ. Rev. 4, 238–265 (1996).
Kuuluvainen, T. & Juntunen, P. Seedling establishment in relation to microhabitat variation in a windthrow gap in a boreal Pinus sylvestris forest. J. Veg. Sci. 9, 551–562 (1998).
Allmér, J., Vasiliauskas, R., Ihrmark, K., Stenlid, J. & Dahlberg, A. Wood-inhabiting fungal communities in woody debris of Norway spruce (Picea abies (L.) Karst.) as reflected by sporocarps mycelial isolations and T-RFLP identification. FEMS Microbiol. Ecol. 55, 57–67 (2006).
Nally, R. M., Parkinson, A., Horrocks, G., Conole, L. & Tzaros, C. Relationships between terrestrial vertebrate diversity, abundance and availability of coarse woody debris on south-eastern Australian floodplains. Biol. Conserv. 99, 191–205 (2001).
Stevenson, S. K., Jull, M. J. & Rogers, B. J. Abundance and attributes of wildlife trees and coarse woody debris at three silvicultural systems study areas in the Interior Cedar-Hemlock Zone, British Columbia. Forest Ecol. Manag. 233, 176–191 (2006).
Mackensen, J., Bauhus, J. & Webber, E. Decomposition rates of coarse woody debris - A review with particular emphasis on Australian tree species. Aust. J. Bot. 51, 27–37 (2003).
Yuan, J. et al. Decay and nutrient dynamics of coarse woody debris in the Qinling Mountains, China. Plos One 12, e0175203 (2017).
Junninen, K., Similä, M., Kouki, J. & Kotiranta, H. Assemblages of wood‐inhabiting fungi along the gradients of succession and naturalness in boreal pine‐dominated forests in Fennoscandia. Ecography 29, 75–83 (2006).
Shorohova, E. & Ekaterina, K. Influence of the substrate and ecosystem attributes on the decomposition rates of coarse woody debris in European boreal forests. Forest Ecol. Manag. 315, 173–184 (2014).
Lindahl, B. O., Taylor, A. F. & Finlay, R. D. Defining nutritional constraints on carbon cycling in boreal forests–towards a less ‘phytocentric’ perspective. Plant Soil 242, 123–135 (2002).
Rajala, T., Peltoniemi, M., Pennanen, T. & Mäkipää, R. Fungal community dynamics in relation to substrate quality of decaying Norway spruce (Picea abies [L.] Karst.) logs in boreal forests. FEMS Microbiol. Ecol. 81, 494–505 (2012).
Lindahl, B. D. et al. Fungal community analysis by high‐throughput sequencing of amplified markers–a user’s guide. New Phytol. 199, 288–299 (2013).
Head, I. M., Saunders, J. R. & Pickup, R. W. Microbial evolution, diversity, and ecology: a decade of ribosomal RNA analysis of uncultivated microorganisms. Microb. Ecol. 35, 1–21 (1998).
Crites, S. & Dale, M. R. Diversity and abundance of bryophytes, lichens, and fungi in relation to woody substrate and successional stage in aspen mixedwood boreal forests. Can. J. Bot. 76, 641–651 (1998).
Kubartová, A., Ottosson, E., Dahlberg, A. & Stenlid, J. Patterns of fungal communities among and within decaying logs, revealed by 454 sequencing. Mol. Ecol. 21, 4514–4532 (2012).
Heilmann-Clausen, J., Aude, E. & Christensen, M. Cryptogam communities on decaying deciduous wood–does tree species diversity matter? Biodivers. Conserv. 14, 2061–2078 (2005).
Jousset, A., Schmid, B., Scheu, S. & Eisenhauer, N. Genotypic richness and dissimilarity opposingly affect ecosystem functioning. Ecol. Lett. 14, 537–545 (2011).
Gessner, M. O. et al. Diversity meets decomposition. Trends Ecol. Evol. 25, 372–380 (2010).
Hughes, J. B., Hellmann, J. J., Ricketts, T. H. & Bohannan, B. J. Counting the uncountable: statistical approaches to estimating microbial diversity. Appl. Environ. Microb. 67, 4399–4406 (2001).
Chao, A. Estimating the population size for capture-recapture data with unequal catchability. Biometrics 43, 783–791 (1987).
Baldrian, P. et al. Fungi associated with decomposing deadwood in a natural beech-dominated forest. Fungal Ecol. 23, 109–122 (2016).
Fukasawa, Y., Osono, T. & Takeda, H. Microfungus communities of Japanese beech logs at different stages of decay in a cool temperate deciduous forest. Can. J. Forest Res. 39, 1606–1614 (2009).
Maser, C. & Trappe, J. M. The seen and unseen world of the fallen tree. USDA-FS Pacific Northwest Forest and Range Experiment Station. General Technical Report. (1984).
Kahl, T., Baber, K., Otto, P., Wirth, C. & Bauhus, J. Drivers of CO2 emission rates from dead wood logs of 13 tree species in the initial decomposition phase. Forests 6, 2484–2504 (2015).
Rubino, D. L. & McCarthy, B. C. Composition and ecology of macrofungal and myxomycete communities on oak woody debris in a mixed-oak forest of Ohio. Can. J. Forest Res. 33, 2151–2163 (2003).
Arslanova, T. Diversity and ecological properties of Dothideomycetes and Sordariomycetes in alluvial black alder forests and pine forest affected by cormorants. Doctoral dissertation, Vilnius University, Vilnius (2014).
Renvall, P. Community structure and dynamics of wood-rotting Basidiomycetes on decomposing conifer trunks in northern Finland. Karstenia 35, 1–51 (1995).
Kirker, G. T., Wagner, T. L. & Diehl, S. V. Relationship between wood-inhabiting fungi and Reticulitermes spp. in four forest habitats of northeastern Mississippi. Int. Biodeter. Biodegr. 72, 18–25 (2012).
Hibbett, D. S. et al. 14 Agaricomycetes in Systematics and evolution (eds. McLaughlin, D. J. & Spatafora, J. W.) 373-429 (Springer Berlin Heidelberg, 2014).
Bîrsan, C., Tanase, C., Mardari, C. & Cojocariu, A. Diversity and ecological determinants of dead wood fungi in tree natural reserves of broad leaved forests from Suceava County. J. Plant Develop. 21, 153–160 (2014).
Küffer, N., Gillet, F., Senn-Irlet, B., Job, D. & Aragno, M. Ecological determinants of fungal diversity on dead wood in European forests. Fungal Divers. 30, 83–95 (2008).
Cornelissen, J. H. C. et al. Controls on coarse wood decay in temperate tree species: birth of the LOGLIFE experiment. Ambio 41, 231–245 (2012).
Sippola, A. L. & Renvall, P. Wood-decomposing fungi and seed-tree cutting: a 40-year perspective. Forest Ecol. Manag. 115, 183–201 (1999).
Ausmus, B. S. Regulation of wood decomposition rates by arthropod and annelid populations. Ecological bulletins 25, 180–192 (1977).
Ódor, P. et al. Diversity of dead wood inhabiting fungi and bryophytes in semi-natural beech forests in Europe. Biol. Conserv. 131, 58–71 (2006).
Rajala, T., Peltoniemi, M., Hantula, J., Mäkipää, R. & Pennanen, T. RNA reveals a succession of active fungi during the decay of Norway spruce logs. Fungal Ecol. 4, 437–448 (2011).
Hart, D. D. Community organization in streams: the importance of species interactions, physical factors, and chance. Oecologia 91, 220–228 (1992).
Zhou, L., Dai, L. M., Gu, H. Y. & Zhong, L. Review on the decomposition and influence factors of coarse woody debris in forest ecosystem. J. Forestry Res. 18, 48–54 (2007).
Weedon, J. T. et al. Global meta‐analysis of wood decomposition rates: a role for trait variation among tree species? Ecol. Lett. 12, 45–56 (2009).
Boddy, L. & Watkinson, S. C. Wood decomposition, higher fungi, and their role in nutrient redistribution. Can. J. Bot. 73, 1377–1383 (1995).
Rayner, A. D. M. & Boddy, L. Fungal communities in the decay of wood. Adv. Microb. Ecol. 10, 115–166 (1988).
Harmon, M. E. & Sexton, J. Guidelines for Measurements of Woody Detritus in Forest Ecosystems. U.S. LTER Network Office, University of Washington, Seattle (1996).
Yan, E. R., Wang, X. H., Huang, J. J., Zeng, F. R. & Gong, L. Long-lasting legacy of forest succession and forest management: Characteristics of coarse woody debris in an evergreen broad-leaved forest of Eastern China. Forest Ecol. Manag. 252, 98–107 (2007).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Kõljalg, U. et al. Towards a unified paradigm for sequence‐based identification of fungi. Mol. Ecol. 22, 5271–5277 (2013).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microb. 73, 5261–5267 (2007).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microb. 75, 7537–7541 (2009).
Acknowledgements
We are grateful to the Qinling National Forest Ecosystem Research Station for providing some data and the experimental equipment. This research was funded by the project of “Technical management system for increasing the capacity of carbon sink and water regulation of mountain forests in the Qinling Mountains” (201004036) from the State Forestry Administration of China.
Author information
Authors and Affiliations
Contributions
J.Y. and S.Z. managed the project; J.Y., F.C. and S.Z. designed the experiments and prepared the samples; F.C. and J.Y. conducted the experiments. F.C. and X.Z. led the data analysis, X.Z. and J.Y. drafted the manuscript; L.H. and J.L. made the figures and tables. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yuan, J., Zheng, X., Cheng, F. et al. Fungal community structure of fallen pine and oak wood at different stages of decomposition in the Qinling Mountains, China. Sci Rep 7, 13866 (2017). https://doi.org/10.1038/s41598-017-14425-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-14425-6
- Springer Nature Limited
This article is cited by
-
Influence of scion cultivar on the rhizosphere microbiome and root exudates of Phaseolus vulgaris in grafting system
Plant and Soil (2024)
-
Resistance of in natura and torrefied wood chips to xylophage fungi
Scientific Reports (2019)
-
Compositional response of Phaseolus vulgaris rhizomicrobiome to a changing soil environment is regulated by long-distance plant signaling
Plant and Soil (2019)